Allelic Variation in Taste Genes Is Associated with Taste and Diet Preferences and Dental Caries
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects
2.2. Genotyping of Target Genes
2.3. Recording of Taste Threshold (TT) and Taste Preference (TP)
2.4. Recording of Food Preferences and Food Intake
2.5. Caries Scoring
2.6. Statistical Analyses
2.7. Ethical Approval
3. Results
3.1. Study Group Characteristics
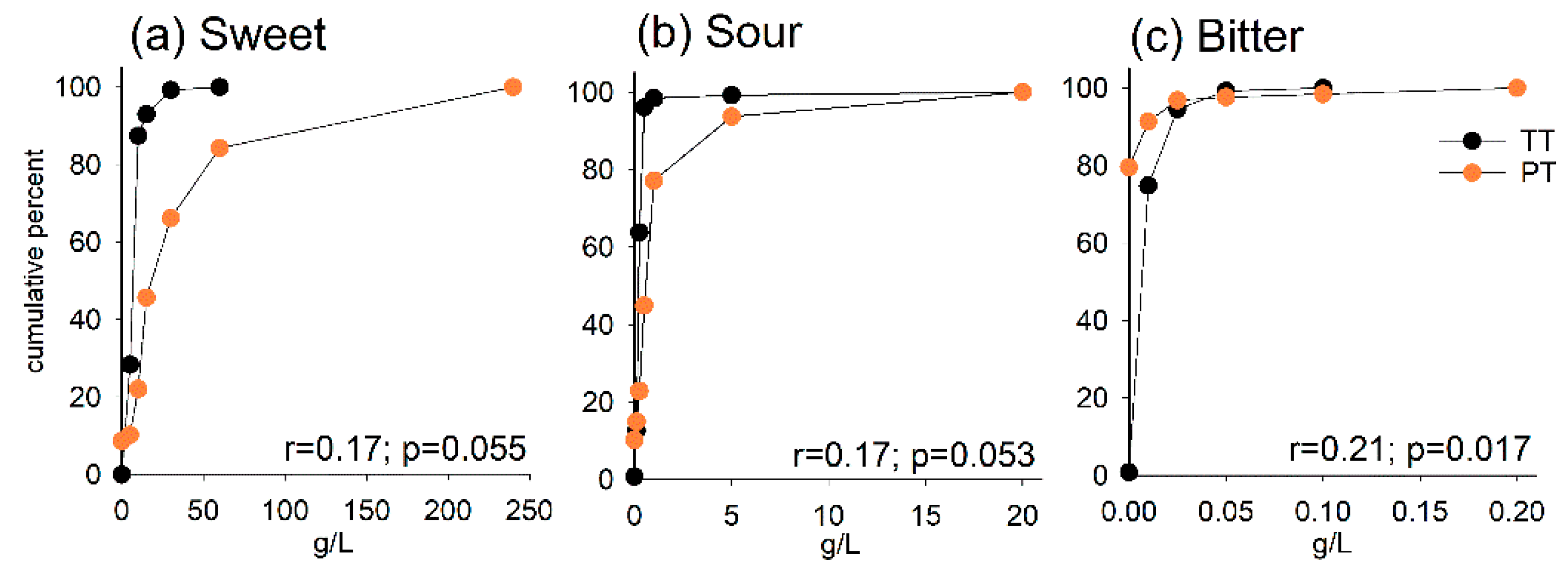
3.2. Taste Threshold (TT) and Preferred Taste (PT)
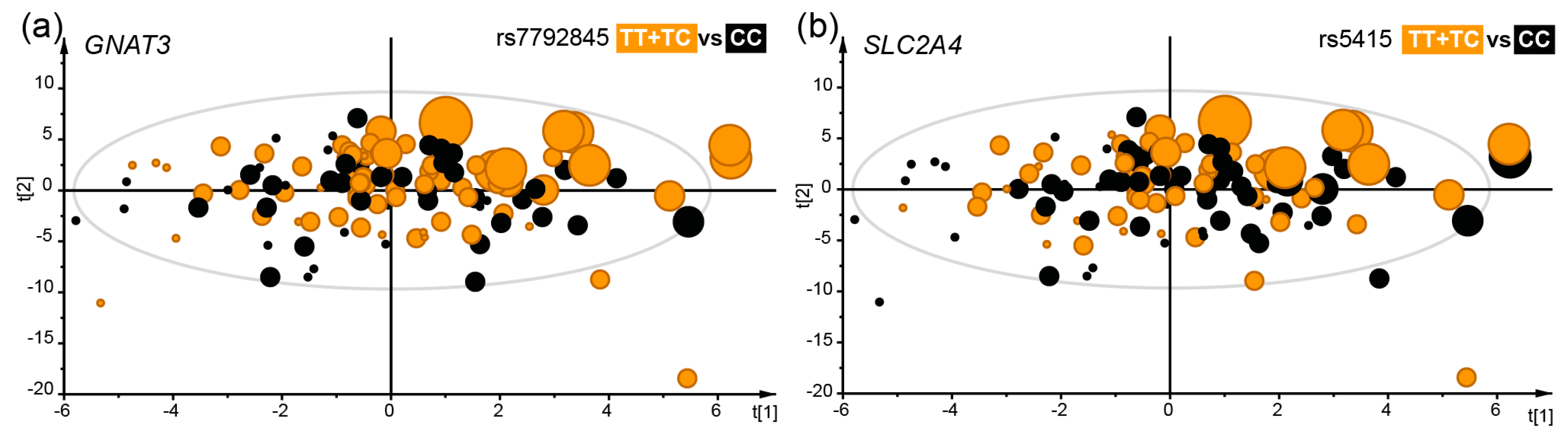
3.3. Taste Receptor Gene Variation and Taste Threshold and Preference
3.4. Food Preference and Food Intake in Taste Categories
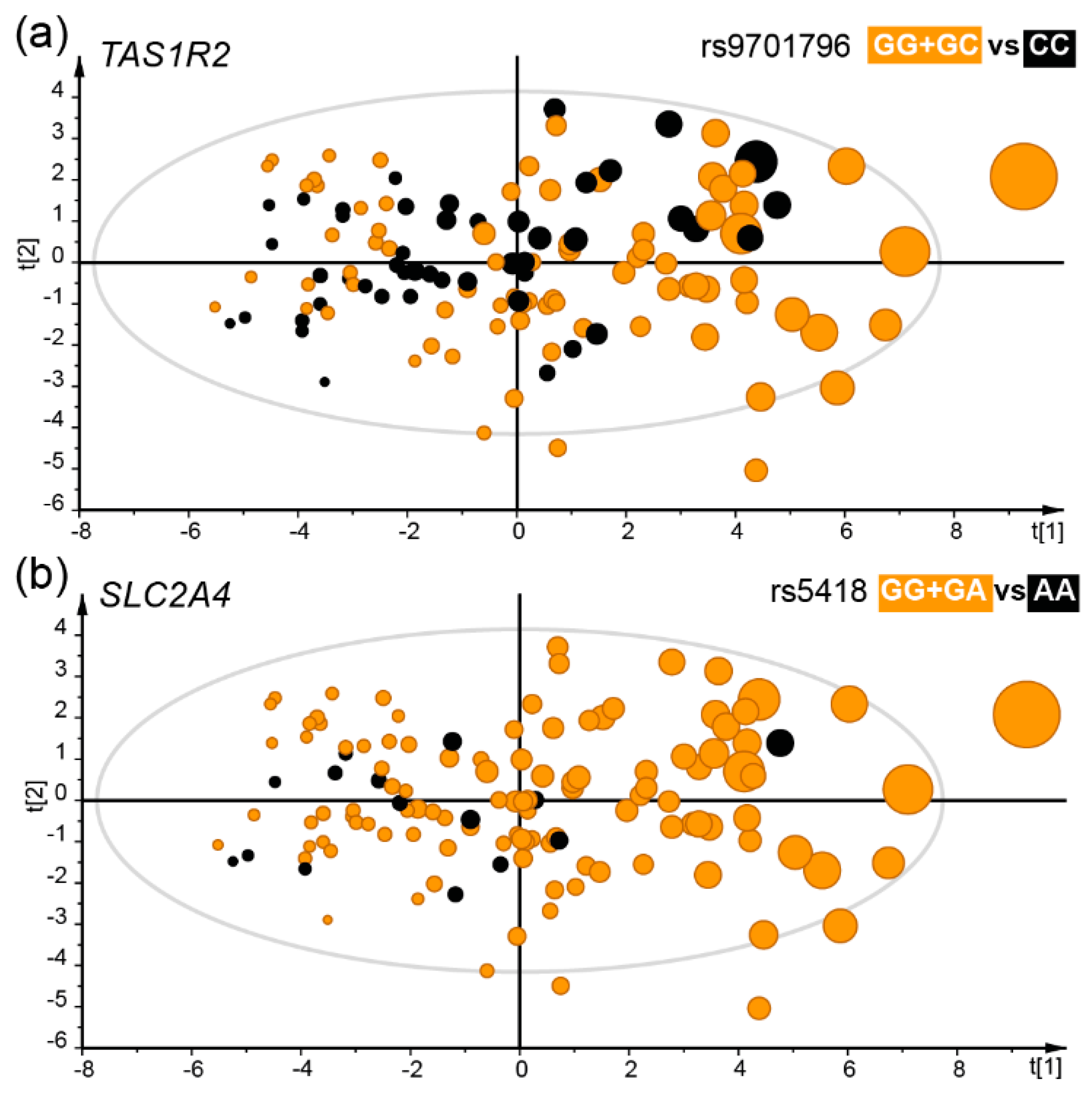
3.5. Taste Receptor Gene Variation and Food Preference and Selection
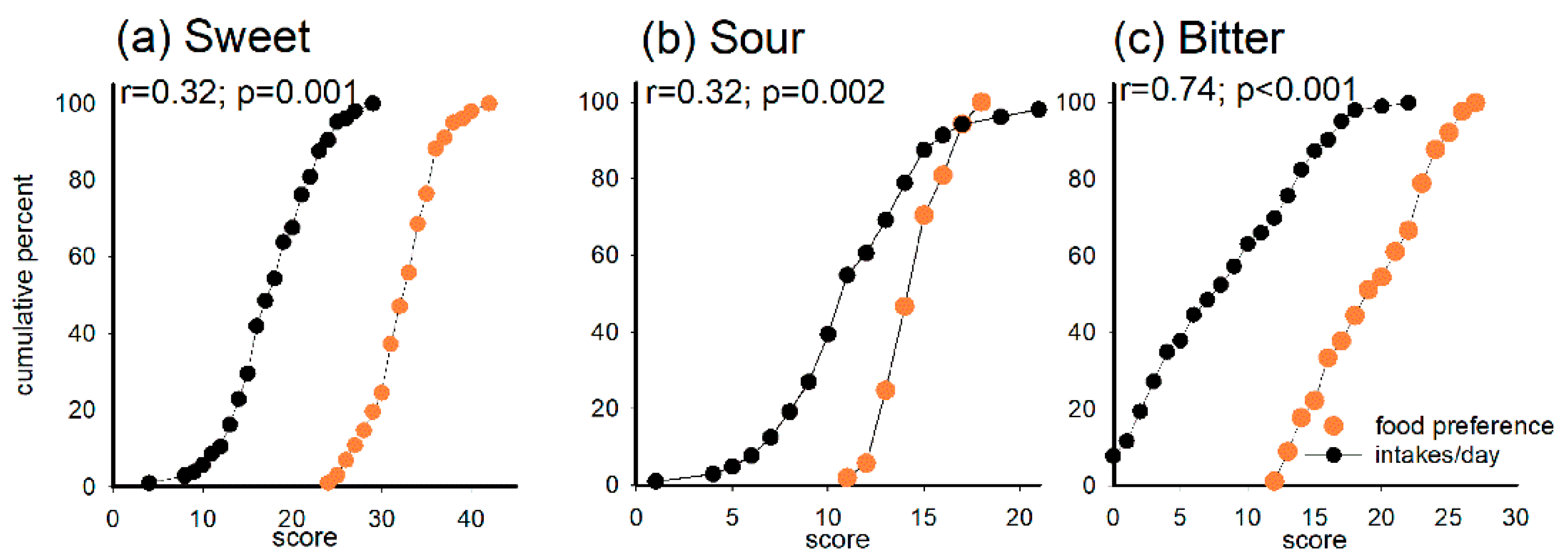
3.6. Correlations between Taste Perception and Preference and Food Preferences and Intake
3.7. Taste Receptor Gene Variation and Caries Status
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Forouzanfar, M.H.; Alexander, L.; Anderson, H.R.; Bachman, V.F.; Biryukov, S.; Brauer, M.; Burnett, R.; Casey, D.; Coates, M.M.; Cohen, A.; et al. Global, regional, and national comparative risk assessment of 79 behavioural, environmental and occupational, and metabolic risks or clusters of risks in 188 countries, 1990–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 2015, 386, 2287–2323. [Google Scholar] [CrossRef]
- Murakami, K.; Livingstone, M.B. Associations between meal and snack frequency and overweight and abdominal obesity in US children and adolescents from National Health and Nutrition Examination Survey (NHANES) 2003–2012. Br. J. Nutr. 2016, 115, 1819–1829. [Google Scholar] [CrossRef] [PubMed]
- Björck, L.; Rosengren, A.; Winkvist, A.; Capewell, S.; Adiels, S.; Bandosz, P.; Critchley, J.; Boman, K.; Castillo Guzman, M.; O’Flaherty, M.; et al. Changes in dietary fat intake and projections for coronary heart disease mortality in Sweden: A simulation study. PLoS ONE 2016, 11, e0160474. [Google Scholar] [CrossRef] [PubMed]
- Drewnowski, A. Taste preferences and food intake. Annu. Rev. Nutr. 1997, 17, 237–253. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.Y.; Tucker, R.M. Sweet Taste as a Predictor of Dietary Intake: A Systematic Review. Nutrients 2019, 11, 94. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Bailo, B.; Toguri, C.; Eny, K.M.; El-Sohemy, A. Genetic variation in taste and its influence on food selection. OMICS 2009, 13, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Bachmanov, A.A.; Bosak, N.P.; Lin, C.; Matsumoto, I.; Ohmoto, M.; Reed, D.R.; Nelson, T.M. Genetics of taste receptors. Curr. Pharm. Des. 2014, 20, 2669–2683. [Google Scholar] [CrossRef]
- Chamoun, E.; Mutch, D.M.; Allen-Vercoe, E.; Buchholz, A.C.; Duncan, A.M.; Spriet, L.L.; Haines, J.; Ma, D.W.L.; Guelph Family Health Study. A review of the associations between single nucleotide polymorphisms in taste receptors, eating behaviors, and health. Crit. Rev. Food Sci. Nutr. 2018, 58, 194–207. [Google Scholar] [CrossRef]
- Hoppu, U.; Laitinen, K.; Jaakkola, J.; Sandell, M. The hTAS2R38 genotype is associated with sugar and candy consumption in preschool boys. J. Hum. Nutr. Diet 2015, 28, 45–51. [Google Scholar] [CrossRef]
- Chandrashekar, J.; Hoon, M.A.; Ryba, N.J.; Zuker, C.S. The receptors and cells for mammalian taste. Nature 2006, 444, 288–294. [Google Scholar] [CrossRef]
- Fushan, A.A.; Simons, C.T.; Slack, J.P.; Drayna, D. Association between common variation in genes encoding sweet taste signaling components and human sucrose perception. Chem. Senses 2010, 35, 579–592. [Google Scholar] [CrossRef] [PubMed]
- Klip, A.; McGraw, T.E.; James, D.E. 30 sweet years of GLUT4. J. Biol. Chem. 2019. [Google Scholar] [CrossRef] [PubMed]
- Ledda, M.; Kutalik, Z.; Souza Destito, M.C.; Souza, M.M.; Cirillo, C.A.; Zamboni, A.; Martin, N.; Morya, E.; Sameshima, K.; Beckmann, J.S.; et al. GWAS of human bitter taste perception identifies new loci and reveals additional complexity of bitter taste genetics. Hum. Mol. Genet. 2014, 23, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Zhong, V.W.; Kuang, A.; Danning, R.D.; Kraft, P.; van Dam, R.M.; Chasman, D.I.; Cornelis, M.C. A genome-wide association study of bitter and sweet beverage consumption. Hum. Mol. Genet. 2019. [Google Scholar] [CrossRef]
- Hwang, L.D.; Lin, C.; Gharahkhani, P.; Cuellar-Partida, G.; Ong, J.S.; An, J.; Gordon, S.D.; Zhu, G.; MacGregor, S.; Lawlor, D.A.; et al. New insight into human sweet taste: A genome-wide association study of the perception and intake of sweet substances. Am. J. Clin. Nutr. 2019, 109, 1724–1737. [Google Scholar] [CrossRef] [PubMed]
- Chapple, I.L.; Bouchard, P.; Cagetti, M.G.; Campus, G.; Carra, M.C.; Cocco, F.; Nibali, L.; Hujoel, P.; Laine, M.L.; Lingstrom, P.; et al. Interaction of lifestyle, behaviour or systemic diseases with dental caries and periodontal diseases: Consensus report of group 2 of the joint EFP/ORCA workshop on the boundaries between caries and periodontal diseases. J. Clin. Periodontol. 2017, 44, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, N.; Nyvad, B. The role of bacteria in the caries process: Ecological perspectives. J. Dent. Res. 2011, 90, 294–303. [Google Scholar] [CrossRef] [PubMed]
- National Institute of Environmental Health Sciences-SNPinfo Web Server. Available online: https://snpinfo.niehs.nih.gov/snpinfo/snptag.html (accessed on 28 June 2019).
- Figshare. Available online: https://doi.org/10.6084/m9.figshare.8001719.v1 (accessed on 28 June 2019).
- Ashi, H.; Campus, G.; Bertéus Forslund, H.; Hafiz, W.; Ahmed, N.; Lingström, P. The Influence of Sweet Taste Perception on Dietary Intake in Relation to Dental Caries and BMI in Saudi Arabian Schoolchildren. Int. J. Dent. 2017, 2017, 4262053. [Google Scholar] [CrossRef]
- Overberg, J.; Hummel, T.; Krude, H.; Wiegand, S. Differences in taste sensitivity between obese and non-obese children and adolescents. Arch. Dis. Child 2012, 97, 1048–1052. [Google Scholar] [CrossRef]
- Mennella, J.A.; Pepino, M.Y.; Reed, D.R. Genetic and environmental determinants of bitter perception and sweet preferences. Pediatrics 2005, 115, 216–222. [Google Scholar] [CrossRef]
- Swedish Food Composition Database. Available online: www.livsmedelsverket.se/en/food-and-content/naringsamnen (accessed on 28 June 2019).
- Johansson, I.; Hallmans, G.; Wikman, A.; Biessy, C.; Riboli, E.; Kaaks, R. Validation and calibration of food-frequency questionnaire measurements in the Northern Sweden Health and Disease cohort. Public Health Nutr. 2002, 5, 487–496. [Google Scholar] [CrossRef] [PubMed]
- Johansson, I.; Van Guelpen, B.; Hultdin, J.; Johansson, M.; Hallmans, G.; Stattin, P. Validity of food frequency questionnaire estimated intakes of folate and other B vitamins in a region without folic acid fortification. Eur. J. Clin. Nutr. 2010, 64, 905–913. [Google Scholar] [CrossRef] [PubMed]
- Klingberg, S.; Winkvist, A.; Hallmans, G.; Johansson, I. Evaluation of plant sterol intake estimated with the Northern Sweden FFQ. Public Health Nutr. 2013, 16, 460–467. [Google Scholar] [CrossRef] [PubMed]
- Wennberg, M.; Vessby, B.; Johansson, I. Evaluation of relative intake of fatty acids according to the Northern Sweden FFQ with fatty acid levels in erythrocyte membranes as biomarkers. Public Health Nutr. 2009, 12, 1477–1484. [Google Scholar] [CrossRef] [PubMed]
- Gabriel, S.B.; Schaffner, S.F.; Nguyen, H.; Moore, J.M.; Roy, J.; Blumenstiel, B.; Higgins, J.; DeFelice, M.; Lochner, A.; Faggart, M.; et al. The structure of haplotype blocks in the human genome. Science 2002, 296, 2225–2229. [Google Scholar] [CrossRef] [PubMed]
- Willett, W.C.; Howe, G.R.; Kushi, L.H. Adjustment for total energy intake in epidemiologic studies. Am. J. Clin. Nutr. 1997, 65, 1220S–1231S. [Google Scholar] [CrossRef]
- Smith, S.R.; Johnson, S.T.; Oldman, S.M.; Duffy, V.B. Pediatric Adapted Liking Survey: A Novel, Feasible and Reliable Dietary Screening in Clinical Practice. Caries Res. 2019, 53, 153–159. [Google Scholar] [CrossRef]
- Misaka, T. Molecular mechanisms of the action of miraculin, a taste-modifying protein. Semin. Cell Dev. Biol. 2013, 24, 222–225. [Google Scholar] [CrossRef]
- Sanematsu, K.; Kitagawa, M.; Yoshida, R.; Nirasawa, S.; Shigemura, N.; Ninomiya, Y. Intracellular acidification is required for full activation of the sweet taste receptor by miraculin. Sci. Rep. 2016, 6, 22807. [Google Scholar] [CrossRef]
- Dias, A.G.; Eny, K.M.; Cockburn, M.; Chiu, W.; Nielsen, D.E.; Duizer, L.; El-Sohemy, A. Variation in the TAS1R2 Gene, Sweet Taste Perception and Intake of Sugars. J. Nutr. Nutr. 2015, 8, 81–90. [Google Scholar] [CrossRef]
- Eny, K.M.; Wolever, T.M.; Corey, P.N.; El-Sohemy, A. Genetic variation in TAS1R2 (Ile191Val) is associated with consumption of sugars in overweight and obese individuals in 2 distinct populations. Am. J. Clin. Nutr. 2010, 92, 1501–1510. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, S.K.; McKinnon, P.J.; Margolskee, R.F. Gustducin is a taste-cell-specific G protein closely related to the transducins. Nature 1992, 357, 563–569. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.T.; Gannon, K.S.; Margolskee, R.F. Transduction of bitter and sweet taste by gustducin. Nature 1996, 381, 796–800. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Yasumatsu, K.; Varadarajan, V.; Yamada, A.; Lem, J.; Ninomiya, Y.; Margolskee, R.F.; Damak, S. Umami taste responses are mediated by alpha-transducin and alpha-gustducin. J. Neurosci. 2004, 24, 7674–7680. [Google Scholar] [CrossRef]
- Izakovicova Holla, L.; Borilova Linhartova, P.; Lucanova, S.; Kastovsky, J.; Musilova, K.; Bartosova, M.; Kukletova, M.; Kukla, L.; Dusek, L. GLUT2 and TAS1R2 Polymorphisms and Susceptibility to Dental Caries. Caries Res. 2015, 49, 417–424. [Google Scholar] [CrossRef] [PubMed]
- Haznedaroğlu, E.; Koldemir-Gündüz, M.; Bakır-Coşkun, N.; Bozkuş, H.M.; Çağatay, P.; Süsleyici-Duman, B.; Menteş, A. Association of sweet taste receptor gene polymorphisms with dental caries experience in school children. Caries Res. 2015, 49, 275–281. [Google Scholar] [CrossRef] [PubMed]
- Robino, A.; Bevilacqua, L.; Pirastu, N.; Situlin, R.; Di Lenarda, R.; Gasparini, P.; Navarra, C.O. Polymorphisms in sweet taste genes (TAS1R2 and GLUT2), sweet liking, and dental caries prevalence in an adult Italian population. Genes Nutr. 2015, 10, 485. [Google Scholar] [CrossRef]
- Kulkarni, G.V.; Chng, T.; Eny, K.M.; Nielsen, D.; Wessman, C.; El-Sohemy, A. Association of GLUT2 and TAS1R2 genotypes with risk for dental caries. Caries Res. 2013, 47, 219–225. [Google Scholar] [CrossRef]
- Laukkanen, O.; Lindström, J.; Eriksson, J.; Valle, T.T.; Hämäläinen, H.; Ilanne-Parikka, P.; Keinänen-Kiukaanniemi, S.; Tuomilehto, J.; Uusitupa, M.; Laakso, M.; et al. Polymorphisms in the SLC2A2 (GLUT2) gene are associated with the conversion from impaired glucose tolerance to type 2 diabetes: The Finnish Diabetes Prevention Study. Diabetes 2005, 54, 2256–2260. [Google Scholar] [CrossRef]
- Willer, C.J.; Bonnycastle, L.L.; Conneely, K.N.; Duren, W.L.; Jackson, A.U.; Scott, L.J.; Narisu, N.; Chines, P.S.; Skol, A.; Stringham, H.M.; et al. Screening of 134 single nucleotide polymorphisms (SNPs) previously associated with type 2 diabetes replicates association with 12 SNPs in nine genes. Diabetes 2007, 56, 256–264. [Google Scholar] [CrossRef]
- Igl, W.; Johansson, A.; Wilson, J.F.; Wild, S.H.; Polasek, O.; Hayward, C.; Vitart, V.; Hastie, N.; Rudan, P.; Gnewuch, C.; et al. Modeling of environmental effects in genome-wide association studies identifies SLC2A2 and HP as novel loci influencing serum cholesterol levels. PLoS Genet. 2010, 6, e1000798. [Google Scholar] [CrossRef] [PubMed]
- Borglykke, A.; Grarup, N.; Sparsø, T.; Linneberg, A.; Fenger, M.; Jeppesen, J.; Hansen, T.; Pedersen, O.; Jørgensen, T. Genetic variant SLC2A2 [corrected] Is associated with risk of cardiovascular disease—Assessing the individual and cumulative effect of 46 type 2 diabetes related genetic variants. PLoS ONE 2012, 7, e50418. [Google Scholar] [CrossRef] [PubMed]
- Meyer, T.E.; Boerwinkle, E.; Morrison, A.C.; Volcik, K.A.; Sanderson, M.; Coker, A.L.; Pankow, J.S.; Folsom, A.R. Diabetes genes and prostate cancer in the Atherosclerosis Risk in Communities study. Cancer Epidemiol. Biomark. Prev. 2010, 19, 558–565. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Toyono, T.; Seta, Y.; Kataoka, S.; Oda, M.; Toyoshima, K. Differential expression of the glucose transporters in mouse gustatory papillae. Cell Tissue Res. 2011, 345, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Cetik, S.; Hupkens, E.; Malaisse, W.J.; Sener, A.; Popescu, I.R. Expression and localization of glucose transporters in rodent submandibular salivary glands. Cell Physiol. Biochem. 2014, 33, 1149–1161. [Google Scholar] [CrossRef] [PubMed]
- Jurysta, C.; Nicaise, C.; Cetik, S.; Louchami, K.; Malaisse, W.J.; Sener, A. Glucose transport by acinar cells in rat parotid glands. Cell Physiol. Biochem. 2012, 29, 325–330. [Google Scholar] [CrossRef]
- Jurysta, C.; Louchami, K.; Malaisse, W.J.; Sener, A. Uptake and efflux of 3-O-methyl-d-glucose in rat parotid cells. Biomed. Rep. 2013, 1, 638–640. [Google Scholar] [CrossRef][Green Version]
- Mascarenhas, P.; Fatela, B.; Barahona, I. Effect of diabetes mellitus type 2 on salivary glucose--a systematic review and meta-analysis of observational studies. PLoS ONE 2014, 9, e101706. [Google Scholar] [CrossRef]
- Goodson, J.M.; Hartman, M.L.; Shi, P.; Hasturk, H.; Yaskell, T.; Vargas, J.; Song, X.; Cugini, M.; Barake, R.; Alsmadi, O.; et al. The salivary microbiome is altered in the presence of a high salivary glucose concentration. PLoS ONE 2017, 12, e0170437. [Google Scholar] [CrossRef]
- Malodobra-Mazur, M.; Bednarska-Chabowska, D.; Olewinski, R.; Chmielecki, Z.; Adamiec, R.; Dobosz, T. Single nucleotide polymorphisms in 5′-UTR of the SLC2A4 gene regulate solute carrier family 2 member 4 gene expression in visceral adipose tissue. Gene 2016, 576, 499–504. [Google Scholar] [CrossRef]
| Factors | All Participants (n = 127) | Preferred Taste (PT)sweet Groups | p-Value | |
|---|---|---|---|---|
| Low (n = 58) | High (n = 69) | |||
| Male, Female, % | 47.2, 52.8 | 37.9, 62.1 | 55.1, 44.9 | 0.054 |
| Body Mass Index 1, kg/m2 | 23.0 (22.4, 23.6) | 22.3 (21.4, 23.3) | 23.6 (22.8, 24.3) | 0.039 |
| Smoking yes, % | 4.7 | 3.4 | 5.8 | 0.687 |
| Swedish snus yes, % | 8.7 | 3.4 | 13.0 | 0.055 |
| Diet intake 1 | ||||
| Energy, kcal/day | 1748 (1624, 1871) | 1666 (1482, 1848) | 1820 (1650, 1990) | 0.226 |
| Carbohydrate, E% | 40.6 (39.3, 42.8) | 40.9 (39.0. 42,8) | 40.3 (38.5, 42.0) | 0.626 |
| Protein, E% | 14.0 (13.4, 14.5) | 13.7 (12.9, 14.5) | 14.2 (13.4, 14.9) | 0.445 |
| Fat, E% | 44.1 (42.8, 45.5) | 44.3 (42.1, 45.9) | 44.0 (42.1, 45.9) | 0.808 |
| Sucrose, E% | 6.0 (5.6, 6.4) | 5.9 (5.4, 6.5) | 6.1 (5.6, 6.6) | 0.652 |
| Saliva flow 1, mL/min | 1.5 (1.4, 1.6) | 1.3 (1.1, 1.5) | 1.7 (1.5, 1.8) | 0.030 |
| Number of teeth | 27.5 (27.2, 27.7) | 27.5 (27.2, 27.7) | 27.5 (27.2, 27.7) | 0.609 |
| Caries status | ||||
| Caries affected 2, % | 56.7 | 60.3 | 53.6 | 0.446 |
| DeFS 1 | 4.4 (3.2, 5.6) | 3.6 (2.4, 4.8) | 5.0 (3.0, 7.0) | 0.238 |
| Pheno-Type | Gene | rs-Number | Minor/Major | Genotype Group 1 vs. 2 | Freq. of Geno-Type Group 1 | Test Group Mean (95% CI) | p-Value | |
|---|---|---|---|---|---|---|---|---|
| Group 1 | Group 2 | |||||||
| TTsweet | GNAT3 | rs17260734 | T/A | TT + TA vs. AA | 69.4% | 3.1 (2.9, 3.2) | 2.6 (2.4, 2.8) | 0.0007 |
| GNAT3 | rs7792845 | T/C | TT + TC vs. CC | 57.3% | 3.1 (2.9, 3.3) | 2.7 (2.5, 2.8) | 0.0002 | |
| SLC2A4 | rs2654185 | A/C | AA + AC vs. CC | 49.6% | 3.1 (2.9, 3.4) | 2.7 (2.6, 2.9) | 0.0016 | |
| SLC2A4 | rs5415 | T/C | TT + TC vs. CC | 43.5% | 3.2 (3.0, 3.4) | 2.7 (2.6, 2.8) | 0.0004 | |
| PTsweet | TAS1R1 | rs4908923 | G/A | GG + GA vs. AA | 18.1% | 4.4 (4.1, 4.8) | 5.5 (5.0, 6.0) | 0.0010 |
| TAS1R2 | rs9988418 | T/C | TT + TC vs. CC | 3.9% | 2.6 (1.3, 3.8) | 4.7 (4.4, 5.0) | 0.0010 | |
| TAS1R2 | rs28652778 | T/C | TT vs. CT + CC | 5.6% | 5.8 (5.1, 6.5) | 4.6 (4.3, 4.9) | 0.0013 | |
| GNAT3 | rs7792845 | T/C | TT vs. CT + CC | 14.5% | 5.7 (5.0, 6.3) | 4.5 (4.1, 4.8) | 0.0011 | |
| TTsour | TAS1R2 | rs12035074 | C/G | CC + CG vs. GG | 44.4% | 3.0 (2.8, 3.2) | 3.4 (3.3, 3.6) | 0.003 |
| GNAT3 | rs6947745 | T/C | TT vs. TG + GG | 5.5% | 3.0 (3.0, 3.1) | 3.3 (3.2, 3.5) | 0.0004 | |
| PTsour | TAS1R1 | rs4908932 | T/G | TT vs. TG + GG | 5.6% | 5.2 (4.8, 5.5) | 4.4 (4.1, 4.7) | 0.0002 |
| TAS1R2 | rs12035074 | C/G | CC + CG vs. GG | 44.4% | 3.9 (3.5, 4.3) | 4.8 (4.5, 5.2) | 0.0003 | |
| TAS1R2 | rs35874116 | C/T | CC vs. CT + TT | 8.7% | 5.4 (4.7, 6.1) | 4.3 (4.0, 4.6) | 0.0018 | |
| TTbitter | TAS2R50 | rs2218820 | T/C | TT vs. TC + CC | 21.3% | 2.6 (2.4, 2.9) | 2.2 (2.1, 2.3) | 0.002 |
| Sweet | TAS1R2 | rs9701796 | G/C | GG + GC vs. CC | 38.4% | 0.9 (0.8, 1.0) | 1.3 (1.1, 1.5) | 0.003 |
| food | SLC2A4 | rs2654185 | A/C | AA vs. CA + CC | 8.0% | 0.7 (0.4, 1,0) | 1.2 (1.0, 1.3) | 0.002 |
| intake | SLC2A4 | rs5418 | G/A | GG + GA vs. AA | 11.2% | 0.7 (0.5, 0.9) | 1.2 (1.0, 1.3) | 4.8 × 10−5 |
| TAS1R2 | rs28374389 | C/T | CC < CT < TT | 7.1% | - | - | 0.003 | |
| TTsweet | PTsweet | TTbitter | PTbitter | TTsour | PTsour | |
|---|---|---|---|---|---|---|
| Food taste clusters | ||||||
| Food preference for sweet foods | 0.130.173 | 0.36<0.001 | ||||
| Intake of sugary foods | 0.180.044 | 0.220.014 | −0.240.008 | |||
| Food preference for bitter foods | 0.140.153 | |||||
| Intake of bitter foods | 0.150.092 | |||||
| Food preference for sour foods | −0.220.019 | −0.100.301 | 0.170.060 | |||
| Intake of sour foods | −0.270.003 | |||||
| Foods items | ||||||
| Intake of sweets | 0.200.025 | −0.190.037 | ||||
| Intake of juice | 0.210.022 | |||||
| Food preference for ice cream | 0.190.039 | |||||
| Food preference for sweet rolls/rusk | 0.240.010 | |||||
| Food preference for raisins | 0.240.008 | |||||
| Intake of syrups | 0.220.012 | −0.230.010 | ||||
| Food preference for non-fermented milk | 0.260.004 | |||||
| Intake of non-fermented milk | 0.210.020 | |||||
| Intake of pancake | −0.180.048 | |||||
| Food preference for Brussels sprouts | 0.190.044 | |||||
| Intake of filter brewed coffee | 0.150.088 | |||||
| Intake of boiled coffee | 0.180.042 | |||||
| Intake of beer | 0.200.026 | 0.180.046 | ||||
| Food preference for juice | −0.230.012 | |||||
| Intake of cookies/cakes | 0.210.018 | |||||
| Intake of apples, pears, peaches | −0.290.001 | |||||
| Food preference for lemon | 0.200.027 | |||||
| Nutrients | ||||||
| Disaccharides, g/day | 0.240.008 | |||||
| Sucrose, g/day | 0.170.056 | −0.220.014 | ||||
| Ascorbic acid, mg/day | −0.210.018 |
| Gene | Rs-Number | Minor/Major | Genotype Group 1 vs. 2 | Freq. of Geno-type Group 1 | DeFS (Group Mean (95% CI)) | p-Value | |
|---|---|---|---|---|---|---|---|
| Group 1 | Group 2 | ||||||
| GNAT3 | rs6962693 | G/T | GG + GT vs. TT | 15.7% | 1.2 (0.5, 2.8) | 4.5 (3.5, 5.8) | 2.0 × 10−3 |
| TAS1R1 | rs4908932 | T/G | TT + TG vs. GG | 33.1% | 6.4 (4.8, 8.6) | 1.9 (1.0, 3.6) | 1.7 × 10−5 |
| TAS1R2 | rs6685177 | A/G | AA vs. AG + GG | 7.1% | 8.8 (6.2, 12.4) | 3.1 (2.0, 4.8) | 4.5 × 10−5 |
| rs28374389 | C/T | CC vs. CT + TT | 7.1% | 8.8 (6.2, 12.4) | 3.1 (2.0, 4.8) | 4.5 × 10−5 | |
| rs28410948 | C/T | CC vs. CT + TT | 10.2% | 8.0 (5.9, 11.0) | 2.9 (1.7, 4.8) | 2.0 × 10−4 | |
| SLC2A2 | rs1996220 | G/A | GG + GA vs. AA | 26.0% | 7.3 (5.4, 9.7) | 2.1 (1.2, 3.7) | 2.0 × 10−6 |
| rs5400 | A/G | AA + AG vs. GG | 26.0% | 7.3 (5.4, 9.7) | 2.1 (1.2, 3.7) | 2.0 × 10−6 | |
| rs11917504 | T/A | TT + TA vs. AA | 23.6% | 7.8 (5.8, 10.4) | 2.2 (1.3, 3.7) | 2.3 × 10−7 | |
| SLC2A4 | rs5415 | T/C | TT vs. TC + CC | 6.6% | 8.8 (4.9, 15.9) | 3.2 (2.3, 4.5) | 3.4 × 10−3 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eriksson, L.; Esberg, A.; Haworth, S.; Holgerson, P.L.; Johansson, I. Allelic Variation in Taste Genes Is Associated with Taste and Diet Preferences and Dental Caries. Nutrients 2019, 11, 1491. https://doi.org/10.3390/nu11071491
Eriksson L, Esberg A, Haworth S, Holgerson PL, Johansson I. Allelic Variation in Taste Genes Is Associated with Taste and Diet Preferences and Dental Caries. Nutrients. 2019; 11(7):1491. https://doi.org/10.3390/nu11071491
Chicago/Turabian StyleEriksson, Linda, Anders Esberg, Simon Haworth, Pernilla Lif Holgerson, and Ingegerd Johansson. 2019. "Allelic Variation in Taste Genes Is Associated with Taste and Diet Preferences and Dental Caries" Nutrients 11, no. 7: 1491. https://doi.org/10.3390/nu11071491
APA StyleEriksson, L., Esberg, A., Haworth, S., Holgerson, P. L., & Johansson, I. (2019). Allelic Variation in Taste Genes Is Associated with Taste and Diet Preferences and Dental Caries. Nutrients, 11(7), 1491. https://doi.org/10.3390/nu11071491





