Effects of 12 Weeks of Hypertrophy Resistance Exercise Training Combined with Collagen Peptide Supplementation on the Skeletal Muscle Proteome in Recreationally Active Men
Abstract
1. Introduction
2. Materials and Methods
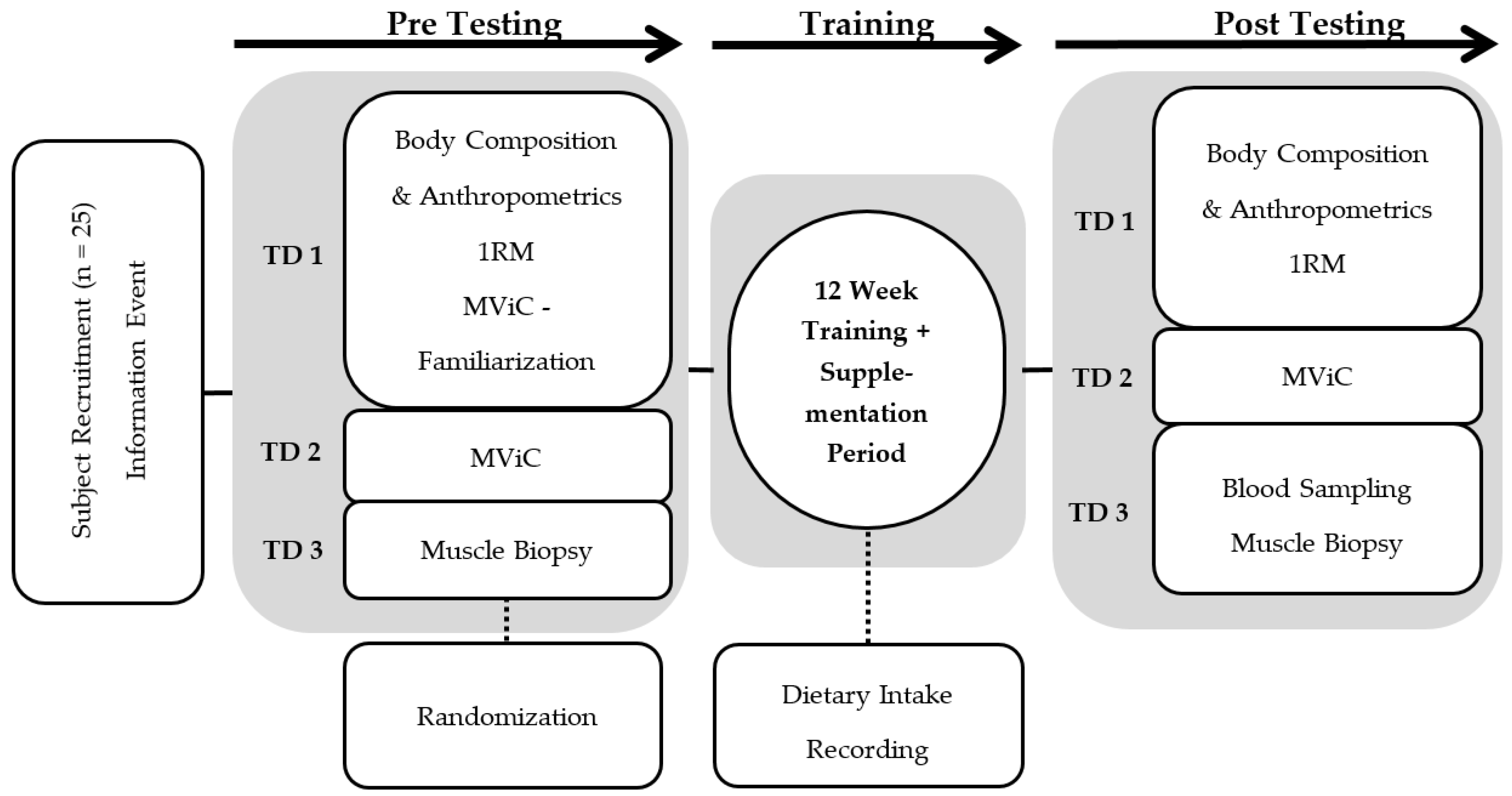
2.1. Experimental Design
2.2. Participants
2.3. Supplementation
2.4. Test Protocols
2.5. Proteomics
2.6. Statistical Analysis
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hawley, J.A.; Burke, L.M.; Phillips, S.M.; Spriet, L.L. Nutritional modulation of training-induced skeletal muscle adaptations. J. Appl. Physiol. 2011, 110, 834–845. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Jentoft, A.J.; Baeyens, J.P.; Bauer, J.M.; Boirie, Y.; Cederholm, T.; Landi, F.; Martin, F.C.; Michel, J.-P.; Rolland, Y.; Schneider, S.M.; et al. Sarcopenia: European consensus on definition and diagnosis: Report of the European Working Group on Sarcopenia in Older People. Age Ageing 2010, 39, 412–423. [Google Scholar] [CrossRef] [PubMed]
- Häkkinen, K.; Alén, M.; Komi, P.V. Changes in isometric force- and relaxation-time, electromyographic and muscle fibre characteristics of human skeletal muscle during strength training and detraining. Acta Physiol. Scand. 1985, 125, 573–585. [Google Scholar] [CrossRef] [PubMed]
- Morton, R.W.; Murphy, K.T.; McKellar, S.R.; Schoenfeld, B.J.; Henselmans, M.; Helms, E.; Aragon, A.A.; Devries, M.C.; Banfield, L.; Krieger, J.W.; et al. A systematic review, meta-analysis and meta-regression of the effect of protein supplementation on resistance training-induced gains in muscle mass and strength in healthy adults. Br. J. Sports Med. 2018, 52, 376–384. [Google Scholar] [CrossRef]
- Hoppeler, H. Molecular networks in skeletal muscle plasticity. J. Exp. Biol. 2016, 219, 205–213. [Google Scholar] [CrossRef]
- Norton, L.E.; Layman, D.K. Leucine regulates translation initiation of protein synthesis in skeletal muscle after exercise. J. Nutr. 2006, 136, 533S–537S. [Google Scholar] [CrossRef] [PubMed]
- Purslow, P.P. The structure and functional significance of variations in the connective tissue within muscle. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2002, 133, 947–966. [Google Scholar] [CrossRef]
- Mackey, A.L.; Magnan, M.; Chazaud, B.; Kjaer, M. Human skeletal muscle fibroblasts stimulate in vitro myogenesis and in vivo muscle regeneration. J. Physiol. 2017, 595, 5115–5127. [Google Scholar] [CrossRef] [PubMed]
- Kjaer, M. Role of extracellular matrix in adaptation of tendon and skeletal muscle to mechanical loading. Physiol. Rev. 2004, 84, 649–698. [Google Scholar] [CrossRef] [PubMed]
- Shaw, G.; Lee-Barthel, A.; Ross, M.L.; Wang, B.; Baar, K. Vitamin C-enriched gelatin supplementation before intermittent activity augments collagen synthesis. Am. J. Clin. Nutr. 2017, 105, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Dressler, P.; Gehring, D.; Zdzieblik, D.; Oesser, S.; Gollhofer, A.; König, D. Improvement of functional ankle properties following supplementation with specific collagen peptides in athletes with chronic ankle instability. J. Bodyw. Mov. Ther. 2018, 22, 858. [Google Scholar] [CrossRef]
- Lugo, J.P.; Saiyed, Z.M.; Lane, N.E. Efficacy and tolerability of an undenatured type II collagen supplement in modulating knee osteoarthritis symptoms: A multicenter randomized, double-blind, placebo-controlled study. Nutr. J. 2016, 15, 14. [Google Scholar] [CrossRef]
- Lugo, J.P.; Saiyed, Z.M.; Lau, F.C.; Molina, J.P.L.; Pakdaman, M.N.; Shamie, A.N.; Udani, J.K. Undenatured type II collagen (UC-II®) for joint support: A randomized, double-blind, placebo-controlled study in healthy volunteers. J. Int. Soc. Sports Nutr. 2013, 10, 48. [Google Scholar] [CrossRef]
- Clark, K.L.; Sebastianelli, W.; Flechsenhar, K.R.; Aukermann, D.F.; Meza, F.; Millard, R.L.; Deitch, J.R.; Sherbondy, P.S.; Albert, A. 24-Week study on the use of collagen hydrolysate as a dietary supplement in athletes with activity-related joint pain. Curr. Med. Res. Opin. 2008, 24, 1485–1496. [Google Scholar] [CrossRef]
- Zdzieblik, D.; Oesser, S.; Gollhofer, A.; Koenig, D. Corrigendum: Improvement of activity-related knee joint discomfort following supplementation of specific collagen peptides. Appl. Physiol. Nutr. Metab. 2017, 42, 1237. [Google Scholar] [CrossRef] [PubMed]
- Bruyère, O.; Zegels, B.; Leonori, L.; Rabenda, V.; Janssen, A.; Bourges, C.; Reginster, J.-Y. Effect of collagen hydrolysate in articular pain: A 6-month randomized, double-blind, placebo controlled study. Complement. Ther. Med. 2012, 20, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Sugihara, F.; Suzuki, K.; Inoue, N.; Venkateswarathirukumara, S. A double-blind, placebo-controlled, randomised, clinical study on the effectiveness of collagen peptide on osteoarthritis. J. Sci. Food Agric. 2015, 95, 702–707. [Google Scholar] [CrossRef]
- Clifford, T.; Ventress, M.; Allerton, D.M.; Stansfield, S.; Tang, J.C.Y.; Fraser, W.D.; Vanhoecke, B.; Prawitt, J.; Stevenson, E. The effects of collagen peptides on muscle damage, inflammation and bone turnover following exercise: A randomized, controlled trial. Amino Acids 2019. [Google Scholar] [CrossRef]
- Zdzieblik, D.; Oesser, S.; Baumstark, M.W.; Gollhofer, A.; Konig, D. Collagen peptide supplementation in combination with resistance training improves body composition and increases muscle strength in elderly sarcopenic men: A randomised controlled trial. Br. J. Nutr. 2015, 114, 1237–1245. [Google Scholar] [CrossRef]
- Padrão, A.I.; Ferreira, R.; Amado, F.; Vitorino, R.; Duarte, J.A. Uncovering the exercise-related proteome signature in skeletal muscle. Proteomics 2016, 16, 816–830. [Google Scholar] [CrossRef]
- Petriz, B.A.; Gomes, C.P.C.; Almeida, J.A.; de Oliveira, G.P.; Ribeiro, F.M.; Pereira, R.W.; Franco, O.L. The Effects of Acute and Chronic Exercise on Skeletal Muscle Proteome. J. Cell. Physiol. 2017, 232, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Tibiana, R.A.; Franco, O.L.; Cunha, G.V.; Sousa, N.M.F.; Sousa Neto, I.V.; Carvalho, M.M.; Almeida, J.A.; Durigan, J.L.Q.; Marqueti, R.C.; Navalta, J.W.; et al. The Effects of Resistance Training Volume on Skeletal Muscle Proteome. Int. J. Exerc. Physiol. 2017, 10, 1051–1066. [Google Scholar]
- Burniston, J.G.; Hoffman, E.P. Proteomic responses of skeletal and cardiac muscle to exercise. Expert Rev. Proteomics 2011, 8, 361–377. [Google Scholar] [CrossRef]
- Kirmse, M.; Oertzen-Hagemann, V.; de Marées, M.; Bloch, W.; Platen, P. Prolonged collagen peptide supplementation and resistance exercise training affects body composition in recreationally active men. Nutrients 2019. submit for publication. [Google Scholar]
- Kraemer, W.J.; Fry, A.C. Strength Testing: Development and Evaluation of Methodology. In Physiological Assessment of Human Fitness; Maud, P., Foster, C., Eds.; Human Kinetics: Champaign, IL, USA, 1995; pp. 115–138. [Google Scholar]
- Oesser, S.; Adam, M.; Babel, W.; Seifert, J. Oral administration of (14)C labeled gelatin hydrolysate leads to an accumulation of radioactivity in cartilage of mice (C57/BL). J. Nutr. 1999, 129, 1891–1895. [Google Scholar] [CrossRef]
- Bergstrom, J. Percutaneous needle biopsy of skeletal muscle in physiological and clinical research. Scand. J. Clin. Lab. Invest. 1975, 35, 609–616. [Google Scholar] [CrossRef]
- Winter, L.; Wittig, I.; Peeva, V.; Eggers, B.; Heidler, J.; Chevessier, F.; Kley, R.A.; Barkovits, K.; Strecker, V.; Berwanger, C.; et al. Mutant desmin substantially perturbs mitochondrial morphology, function and maintenance in skeletal muscle tissue. Acta Neuropathol. 2016, 132, 453–473. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Shinde, S.; Koch, M.-H.; Eisenacher, M.; Galozzi, S.; Lerari, T.; Barkovits, K.; Subedi, P.; Krüger, R.; Kuhlmann, K.; et al. Low-bias phosphopeptide enrichment from scarce samples using plastic antibodies. Sci. Rep. 2015, 5, 11438. [Google Scholar] [CrossRef] [PubMed]
- Apweiler, R.; Bairoch, A.; Wu, C.H.; Barker, W.C.; Boeckmann, B.; Ferro, S.; Gasteiger, E.; Huang, H.; Lopez, R.; Magrane, M.; et al. UniProt: The Universal Protein knowledgebase. Nucleic Acids Res. 2004, 32, D115–D119. [Google Scholar] [CrossRef]
- Reidegeld, K.A.; Eisenacher, M.; Kohl, M.; Chamrad, D.; Körting, G.; Blüggel, M.; Meyer, H.E.; Stephan, C. An easy-to-use Decoy Database Builder software tool, implementing different decoy strategies for false discovery rate calculation in automated MS/MS protein identifications. Proteomics 2008, 8, 1129–1137. [Google Scholar] [CrossRef]
- Uszkoreit, J.; Maerkens, A.; Perez-Riverol, Y.; Meyer, H.E.; Marcus, K.; Stephan, C.; Kohlbacher, O.; Eisenacher, M. PIA: An Intuitive Protein Inference Engine with a Web-Based User Interface. J. Proteome Res. 2015, 14, 2988–2997. [Google Scholar] [CrossRef] [PubMed]
- Uszkoreit, J.; Perez-Riverol, Y.; Eggers, B.; Marcus, K.; Eisenacher, M. Protein inference using PIA workflows and PSI standard file formats. J. Proteome Res. 2018, 18, 741–747. [Google Scholar] [CrossRef]
- Turewicz, M.; Kohl, M.; Ahrens, M.; Mayer, G.; Uszkoreit, J.; Naboulsi, W.; Bracht, T.; Megger, D.A.; Sitek, B.; Marcus, K.; et al. BioInfra.Prot: A comprehensive proteomics workflow including data standardization, protein inference, expression analysis and data publication. J. Biotechnol. 2017, 261, 116–125. [Google Scholar] [CrossRef] [PubMed]
- Burger, T. Gentle Introduction to the Statistical Foundations of False Discovery Rate in Quantitative Proteomics. J. Proteome Res. 2018, 17, 12–22. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Mi, H.; Muruganujan, A.; Casagrande, J.T.; Thomas, P.D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 2013, 8, 1551. [Google Scholar] [CrossRef]
- Supek, F.; Bošnjak, M.; Škunca, N.; Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 2011, 6, e21800. [Google Scholar] [CrossRef]
- Aoi, W.; Sakuma, K. Skeletal muscle: A novel and intriguing characteristics as a secretory organ. Biodiscovery 2013, 7, 2. [Google Scholar] [CrossRef]
- Pedersen, B.K.; Steensberg, A.; Fischer, C.; Keller, C.; Keller, P.; Plomgaard, P.; Febbraio, M.; Saltin, B. Searching for the exercise factor: Is IL-6 a candidate? J. Muscle Res. Cell Moti. 2003, 24, 113–119. [Google Scholar] [CrossRef]
- Yu, M.; Stepto, N.K.; Chibalin, A.V.; Fryer, L.G.D.; Carling, D.; Krook, A.; Hawley, J.A.; Zierath, J.R. Metabolic and mitogenic signal transduction in human skeletal muscle after intense cycling exercise. J. Physiol. 2003, 546, 327–335. [Google Scholar] [CrossRef]
- Yu, M.; Blomstrand, E.; Chibalin, A.V.; Krook, A.; Zierath, J.R. Marathon running increases ERK1/2 and p38 MAP kinase signalling to downstream targets in human skeletal muscle. J. Physiol. 2001, 536, 273–282. [Google Scholar] [CrossRef]
- Egan, B.; Zierath, J.R. Exercise metabolism and the molecular regulation of skeletal muscle adaptation. Cell Metab. 2013, 17, 162–184. [Google Scholar] [CrossRef]
- Bodine, S.C.; Stitt, T.N.; Gonzalez, M.; Kline, W.O.; Stover, G.L.; Bauerlein, R.; Zlotchenko, E.; Scrimgeour, A.; Lawrence, J.C.; Glass, D.J.; et al. Akt/mTOR pathway is a crucial regulator of skeletal muscle hypertrophy and can prevent muscle atrophy in vivo. Nat. Cell Biol. 2001, 3, 1014–1019. [Google Scholar] [CrossRef] [PubMed]
- Peake, J.M.; Della Gatta, P.; Suzuki, K.; Nieman, D.C. Cytokine expression and secretion by skeletal muscle cells: Regulatory mechanisms and exercise effects. Exerc. Immunol. Rev. 2015, 21, 8–25. [Google Scholar]
- Hargreaves, M. Skeletal Muscle Metabolism during Exercise in Humans. Clin. Exp. Pharmacol. Physiol. 2000, 27, 225–228. [Google Scholar] [CrossRef]
- Scheerer, N.; Dehne, N.; Stockmann, C.; Swoboda, S.; Baba, H.A.; Neugebauer, A.; Johnson, R.S.; Fandrey, J. Myeloid hypoxia-inducible factor-1α is essential for skeletal muscle regeneration in mice. J. Immunol. 2013, 191, 407–414. [Google Scholar] [CrossRef] [PubMed]
- Paulsen, G.; Lauritzen, F.; Bayer, M.L.; Kalhovde, J.M.; Ugelstad, I.; Owe, S.G.; Hallén, J.; Bergersen, L.H.; Raastad, T. Subcellular movement and expression of HSP27, alphaB-crystallin, and HSP70 after two bouts of eccentric exercise in humans. J. Appl. Physiol. 2009, 107, 570–582. [Google Scholar] [CrossRef] [PubMed]
- Chereau, D.; Boczkowska, M.; Skwarek-Maruszewska, A.; Fujiwara, I.; Hayes, D.B.; Rebowski, G.; Lappalainen, P.; Pollard, T.D.; Dominguez, R. Leiomodin is an actin filament nucleator in muscle cells. Science (New York, N.Y.) 2008, 320, 239–243. [Google Scholar] [CrossRef]
- Son, H.J.; Kim, H.J.; Kim, C.K. The Effect of Resistance and Endurance Training on Muscle Proteome Expression in Human Skeletal Muscle. Med. Sci. Sports Exerc. 2011, 43, 303. [Google Scholar] [CrossRef]
- Sorichter, S.; Mair, J.; Koller, A.; Gebert, W.; Rama, D.; Calzolari, C.; Artner-Dworzak, E.; Puschendorf, B. Skeletal troponin I as a marker of exercise-induced muscle damage. J. Appl. Physiol. 1997, 83, 1076–1082. [Google Scholar] [CrossRef]
- Carlsson, L.; Yu, J.-G.; Moza, M.; Carpén, O.; Thornell, L.-E. Myotilin: A prominent marker of myofibrillar remodelling. Neurom. Disord. 2007, 17, 61–68. [Google Scholar] [CrossRef]
- Bloch, R.J.; Capetanaki, Y.; O’Neill, A.; Reed, P.; Williams, M.W.; Resneck, W.G.; Porter, N.C.; Ursitti, J.A. Costameres: Repeating structures at the sarcolemma of skeletal muscle. Clin. Orthop. Relat. Res. 2002, S203–S210. [Google Scholar] [CrossRef]
- Kitakaze, T.; Sakamoto, T.; Kitano, T.; Inoue, N.; Sugihara, F.; Harada, N.; Yamaji, R. The collagen derived dipeptide hydroxyprolyl-glycine promotes C2C12 myoblast differentiation and myotube hypertrophy. Biochem. Biophys. Res. Commun. 2016, 478, 1292–1297. [Google Scholar] [CrossRef] [PubMed]
- Frantz, C.; Stewart, K.M.; Weaver, V.M. The extracellular matrix at a glance. J. Cell Sci. 2010, 123, 4195–4200. [Google Scholar] [CrossRef]
- Sorensen, J.R.; Skousen, C.; Holland, A.; Williams, K.; Hyldahl, R.D. Acute extracellular matrix, inflammatory and MAPK response to lengthening contractions in elderly human skeletal muscle. Exp. Gerontol. 2018, 106, 28–38. [Google Scholar] [CrossRef]
- Kroll, T.G.; Peters, B.P.; Hustad, C.M.; Jones, P.A.; Killen, P.D.; Ruddon, R.W. Expression of laminin chains during myogenic differentiation. J. Biol. Chem. 1994, 269, 9270–9277. [Google Scholar]
- Wenstrup, R.J.; Florer, J.B.; Brunskill, E.W.; Bell, S.M.; Chervoneva, I.; Birk, D.E. Type V collagen controls the initiation of collagen fibril assembly. J. Biol. Chem. 2004, 279, 53331–53337. [Google Scholar] [CrossRef]
- Toriello, H.V.; Glover, T.W.; Takahara, K.; Byers, P.H.; Miller, D.E.; Higgins, J.V.; Greenspan, D.S. A translocation interrupts the COL5A1 gene in a patient with Ehlers–Danlos syndrome and hypomelanosis of Ito. Nat. Genet. 1996, 13, 361–365. [Google Scholar] [CrossRef]
- Baghdadi, M.B.; Castel, D.; Machado, L.; Fukada, S.-I.; Birk, D.E.; Relaix, F.; Tajbakhsh, S.; Mourikis, P. Reciprocal signalling by Notch–Collagen V–CALCR retains muscle stem cells in their niche. Nature 2018, 557, 714–718. [Google Scholar] [CrossRef] [PubMed]
- Timmons, J.A. Variability in training-induced skeletal muscle adaptation. J. Appl. Physiol. 2011, 110, 846–853. [Google Scholar] [CrossRef] [PubMed]
- Pakharukova, N.A.; Pastushkova, L.K.; Moshkovskii, S.A.; Larina, I.M. Variability of the healthy human proteome. Biochemi. (Mosc.) Suppl. Ser. B: Biomed. Chem. 2011, 5, 203. [Google Scholar] [CrossRef]
- Seppinen, L.; Pihlajaniemi, T. The multiple functions of collagen XVIII in development and disease. Matrix Biol. 2011, 30, 83–92. [Google Scholar] [CrossRef]
- O’Reilly, M.S.; Boehm, T.; Shing, Y.; Fukai, N.; Vasios, G.; Lane, W.S.; Flynn, E.; Birkhead, J.R.; Olsen, B.R.; Folkman, J. Endostatin: An Endogenous Inhibitor of Angiogenesis and Tumor Growth. Cell 1997, 88, 277–285. [Google Scholar] [CrossRef]
- Svensson, L.; Närlid, I.; Oldberg, Å. Fibromodulin and lumican bind to the same region on collagen type I fibrils. FEBS Lett. 2000, 470, 178–182. [Google Scholar] [CrossRef]
- Hyldahl, R.D.; Nelson, B.; Xin, L.; Welling, T.; Groscost, L.; Hubal, M.J.; Chipkin, S.; Clarkson, P.M.; Parcell, A.C. Extracellular matrix remodeling and its contribution to protective adaptation following lengthening contractions in human muscle. FASEB J. 2015, 29, 2894–2904. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | Collagen Group (n = 12) | Placebo Group (n = 13) | Interaction | Post-Test Difference between Groups | ||
|---|---|---|---|---|---|---|
| Pre | Post | Pre | Post | p Value | p Value | |
| BM (kg) | 81.4 ± 6.6 | 84.4 ± 6.3 * | 77.9 ± 4.1 | 79.4 ± 5.1 * | 0.035 | n.s. (0.039) |
| FM (kg) | 10.3 ± 3.6 | 10.9 ± 4.1 * | 8.4 ± 2.2 | 9.4 ± 2.4 * | n.s. (0.534) | - |
| FFM (kg) | 71.2 ± 5.7 | 73.8 ± 5.3 * | 69.6 ± 4.0 | 70.3 ± 4.3 * | 0.014 | n.s. (0.080) |
| SQ (kg) | 114.3 ± 20.2 | 140.8 ± 24.3 * | 108.7 ± 8.3 | 126.3 ± 13.9 * | n.s. (0.073) | n.s. (0.084) |
| DL (kg) | 131.7 ± 19.6 | 156.3 ± 21.2 * | 128.0 ± 15.6 | 143.9 ± 10.6 * | n.s. (0.234) | - |
| BP (kg) | 80.4 ± 14.0 | 94.4 ± 15.6 * | 84.4 ± 13.9 | 94.2 ± 10.2 * | n.s. (0.099) | n.s. (0.987) |
| R (kg) | 85.0 ± 11.3 | 98.3 ± 12.6 * | 91.2 ± 8.9 | 97.3 ± 8.1 * | 0.025 | n.s. (0.812) |
| MViC (nM) | 294.3 ± 56.3 | 323.6 ± 72.1 * | 260.7 ± 25.0 | 275.2 ± 31.3 * | n.s. (0.066) | n.s. (0.059) |
| Exercise | Collagen Group | Placebo Group | Unpaired t-test |
|---|---|---|---|
| Delta | Delta | p value | |
| BM (kg) | 3.01 ± 2.01 | 1.50 ± 1.29 | 0.035 |
| FM (kg) | 0.58 ± 1.47 | 0.90 ± 1.09 | n.s. (0.534) |
| FFM (kg) | 2.56 ± 2.22 | 0.70 ± 1.14 | 0.014 |
| SQ (kg) | 26.5 ± 13.2 | 17.6 ± 9.9 | n.s. (0.073) |
| DL (kg) | 24.5 ± 19.1 | 15.9 ± 13.9 | n.s. (0.234) |
| BP (kg) | 14.0 ± 6.6 | 9.8 ± 5.5 | n.s. (0.099) |
| R (kg) | 13.3 ± 8.7 | 6.1 ± 5.8 | 0.025 |
| MViC (nM) | 29.4 ± 22.9 | 14.5 ± 14.3 | n.s. (0.066) |
| UniProt | Gene Name | Protein Description | Fold Change (FC) | ||
|---|---|---|---|---|---|
| Collagen | Placebo | ||||
| Only upregulated in COL before 12 weeks | |||||
| - | - | - | |||
| Only upregulated in PLA before 12 weeks | |||||
| P10636 | MAPT | Microtubule-associated protein tau | - | 1.63 | |
| P02144 | MB | Myoglobin | - | 1.71 | |
| P09493 | TPM1 | Tropomyosin alpha-1 chain | - | 1.52 | |
| Pathway | Number of Pathways | Fold Enrichment | FDR |
|---|---|---|---|
| Cell Cycle | 47 | ||
| Signal Transduction | 31 | ||
| PIP3 activates AKT signaling | 3.98 | 3.56 × 10−3 | |
| MAPK family signaling cascades | 5.22 | 7.42 × 10−6 | |
| MAPK6/MAPK4 signaling | 9.65 | 1.47 × 10−5 | |
| MAPK1/MAPK3 signaling | 6.09 | 1.63 × 10−6 | |
| RAF/MAP kinase cascade | 6.22 | 1.37 × 10−6 | |
| Regulation of RAS by GAPs | 12.63 | 2.48 × 10−6 | |
| Immune System | 27 | ||
| MAP kinase activation | 7.57 | 7.58 × 10−3 | |
| Cytokine signaling in immune system | 3.71 | 7.00 × 10−7 | |
| Signaling by Interleukins | 4.25 | 2.51 × 10−6 | |
| Interleukin-1 family signaling | 7.66 | 8.30 × 10−6 | |
| Interleukin-1 signaling | 10.39 | 8.47 × 10−7 | |
| MAP3K8 (TPL2)-dependent MAPK1/3 activation | 17.89 | 9.74 × 10−3 | |
| Interleukin-12 family signaling | 6.70 | 3.23 × 10−2 | |
| Metabolism of Proteins | 23 | ||
| Translation | 3.91 | 1.13 × 10−3 | |
| Eukaryotic translation elongation | 9.14 | 2.11 × 10−5 | |
| Eukaryotic translation initiation | 6.42 | 7.41 × 10−4 | |
| Eukaryotic translation termination | 6.16 | 6.39 × 10−3 | |
| Protein folding | 7.63 | 2.41 × 10−4 | |
| Post-chaperonin tubulin folding pathway | 17.35 | 1.79 × 10−3 | |
| Cooperation of prefoldin and TriC/CCT in actin and tubulin folding | 17.89 | 3.86 × 10−5 | |
| Formation of tubulin folding intermediates by CCT/TriC | 19.09 | 1.90 × 10−4 | |
| Prefoldin mediated transfer of substrate to CCT/TriC | 21.21 | 1.74 × 10−5 | |
| Post-translational protein modification | 2.61 | 1.89 × 10−6 | |
| UCH proteinases | 11.93 | 1.04 × 10−7 | |
| Deubiquitination | 5.08 | 1.00 × 10−5 | |
| Metabolism | 15 | ||
| The citric acid (TCA) cycle and respiratory electron transport | 8.27 | 1.08 × 10−7 | |
| Pyruvate metabolism and citric acid (TCA) cycle | 8.84 | 4.29 × 10−3 | |
| Pyruvate metabolism | 12.72 | 4.76 × 10−3 | |
| Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins | 7.76 | 2.13 × 10−5 | |
| Respiratory electron transport | 7.63 | 2.42 × 10−4 | |
| Metabolism of amino acids and derivatives | 6.8 | 4.19 × 10−11 | |
| Histidine, lysine, phenylalanine, tyrosine, proline and tryptophan catabolism | 10.37 | 2.29 × 10−3 | |
| Glyoxylate metabolism and glycine degradation | 12.31 | 5.25 × 10−3 | |
| Gluconeogenesis | 11.57 | 6.32 × 10−3 | |
| Cellular Response to external stimuli | 11 | ||
| Cellular responses to stress | 6.31 | 1.04 × 10−10 | |
| Cellular response to hypoxia | 12.72 | 7.28 × 10−7 | |
| Oxygen-dependent proline hydroxylation of HIF-1α | 14.46 | 3.37 × 10−7 | |
| Cellular response to heat stress | 13.16 | 5.52 × 10−8 | |
| HSF1 activation | 39.77 | 1.15 × 10−5 | |
| HSF1-dependent transactivation | 31.81 | 9.12 × 10−8 | |
| Attenuation phase | 34.08 | 1.98 × 10−5 | |
| Regulation of HSF1-mediated heat shock response | 7.12 | 9.56 × 10−3 | |
| HSP90 chaperone cycle for steroid hormone receptors (SHR) | 13.63 | 6.36 × 10−6 | |
| Metabolism of mRNA | 11 | ||
| Developmental Biology | 6 | ||
| Gene Expression | 6 | ||
| Muscle Contraction | 3 | ||
| Striated muscle contraction | 23.86 | 8.26 × 10−8 | |
| Smooth muscle contraction | 10.91 | 7.55 × 10−3 | |
| Others | 12 | ||
| CREB phosphorylation through the activation of CaMKII | 19.09 | 5.28 × 10−3 |
| UniProt | Gene Name | Protein Description | Fold Change | FDR |
|---|---|---|---|---|
| Only upregulated in COL after 12 weeks of training | ||||
| P35609 | ACTN2 | Alpha-actinin-2 | 1.57 | 0.006 |
| Q9GZV1 | ANKR2 | Ankyrin repeat domain-containing protein 2 | 1.62 | 0.025 |
| O95817 | BAG3 | BAG family molecular chaperone regulator 3 | 1.87 | 0.004 |
| O00499 | BIN1 | Myc box-dependent-interacting protein 1 | 1.79 | 0.000 |
| P17661; P07197 | DESM | Desmin | 1.70 | 0.002 |
| O00757 | F16P2 | Fructose-1,6-bisphosphatase isozyme 2 | 1.51 | 0.020 |
| Q96A32 | MLRS | Myosin regulatory light chain 2, skeletal muscle isoform | 1.70 | 0.007 |
| P13535 | MYH8 | Myosin-8 | 2.55 | 0.047 |
| P05976; Q15111 | MYL1 | Myosin light chain 1/3, skeletal muscle isoform | 1.61 | 0.009 |
| Q9UBF9 | MYOTI | Myotilin | 1.51 | 0.008 |
| O14974 | MYPT1 | Protein phosphatase 1 regulatory subunit 12A | 1.54 | 0.021 |
| Q53GG5 | PDLI3 | PDZ and LIM domain protein 3 | 1.93 | 0.000 |
| P60900 | PSA6 | Proteasome subunit alpha type-6 | 1.56 | 0.017 |
| Q9UHP9 | SMPX | Small muscular protein | 2.03 | 0.000 |
| A8MU46 | SMTL1 | Smoothelin-like protein 1 | 2.05 | 0.003 |
| P23327 | SRCH | Sarcoplasmic reticulum histidine-rich calcium-binding protein | 1.96 | 0.003 |
| Q9H7C4 | SYNCI | Syncoilin | 1.51 | 0.026 |
| O15061 | SYNEM | Synemin | 1.57 | 0.003 |
| Q8N3V7 | SYNPO | Synaptopodin | 1.82 | 0.016 |
| Q9NZQ9 | TMOD4 | Tropomodulin-4 | 1.92 | 0.002 |
| P02585; P27482 | TNNC2 | Troponin C, skeletal muscle | 1.98 | 0.002 |
| P48788 | TNNI2 | Troponin I, fast skeletal muscle | 1.98 | 0.006 |
| P07951; O75330 | TPM2 | Tropomyosin beta chain | 1.88 | 0.000 |
| P06753 | TPM3 | Tropomyosin alpha-3 chain | 2.07 | 0.004 |
| Q9BYV2 | TRI54 | Tripartite motif-containing protein 54 | 2.15 | 0.033 |
| Only upregulated in PLA group after 12 weeks of training | ||||
| P02511 | CRYAB | Alpha-crystallin B chain | 2.01 | 0.037 |
| Q6P5Q4 | LMOD2 | Leiomodin-2 | 2.80 | 0.014 |
| UniProt | Gene Name | Protein Description | Fold Change (FC) | FDR | |||
|---|---|---|---|---|---|---|---|
| Collagen | Placebo | Collagen | Placebo | ||||
| Upregulated in both groups after 12 weeks | |||||||
| P12110 | CO6A2 | Collagen alpha-2(VI) chain | 2.86 | 3.17 | 0.001 | 0.000 | |
| P02751 | FINC | Fibronectin | 4.46 | 2.24 | 0.007 | 0.021 | |
| P11047 | LAMC1 | Laminin subunit gamma 1 | 1.90 | 1.71 | 0.023 | 0.022 | |
| Only upregulated in COL after 12 weeks | |||||||
| P20908 | CO5A1 | Collagen alpha-1(V) chain | 108.74 | - | 0.018 | - | |
| P39059 | COFA1 | Collagen alpha-1(XV) chain | 1.69 | - | 0.028 | - | |
| P39060 | COIA1 | Collagen alpha-1(XVIII) chain | 2.85 | - | 0.001 | - | |
| Only upregulated in PLA after 12 weeks | |||||||
| P51884 | LUM | Lumican | - | 1.59 | - | 0.001 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oertzen-Hagemann, V.; Kirmse, M.; Eggers, B.; Pfeiffer, K.; Marcus, K.; de Marées, M.; Platen, P. Effects of 12 Weeks of Hypertrophy Resistance Exercise Training Combined with Collagen Peptide Supplementation on the Skeletal Muscle Proteome in Recreationally Active Men. Nutrients 2019, 11, 1072. https://doi.org/10.3390/nu11051072
Oertzen-Hagemann V, Kirmse M, Eggers B, Pfeiffer K, Marcus K, de Marées M, Platen P. Effects of 12 Weeks of Hypertrophy Resistance Exercise Training Combined with Collagen Peptide Supplementation on the Skeletal Muscle Proteome in Recreationally Active Men. Nutrients. 2019; 11(5):1072. https://doi.org/10.3390/nu11051072
Chicago/Turabian StyleOertzen-Hagemann, Vanessa, Marius Kirmse, Britta Eggers, Kathy Pfeiffer, Katrin Marcus, Markus de Marées, and Petra Platen. 2019. "Effects of 12 Weeks of Hypertrophy Resistance Exercise Training Combined with Collagen Peptide Supplementation on the Skeletal Muscle Proteome in Recreationally Active Men" Nutrients 11, no. 5: 1072. https://doi.org/10.3390/nu11051072
APA StyleOertzen-Hagemann, V., Kirmse, M., Eggers, B., Pfeiffer, K., Marcus, K., de Marées, M., & Platen, P. (2019). Effects of 12 Weeks of Hypertrophy Resistance Exercise Training Combined with Collagen Peptide Supplementation on the Skeletal Muscle Proteome in Recreationally Active Men. Nutrients, 11(5), 1072. https://doi.org/10.3390/nu11051072






