A Vegetarian Diet Is a Major Determinant of Gut Microbiota Composition in Early Pregnancy
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Faecal DNA Extraction
2.3. 16S rRNA Sequencing
2.4. Statistical Analysis
3. Results
3.1. Study Participants
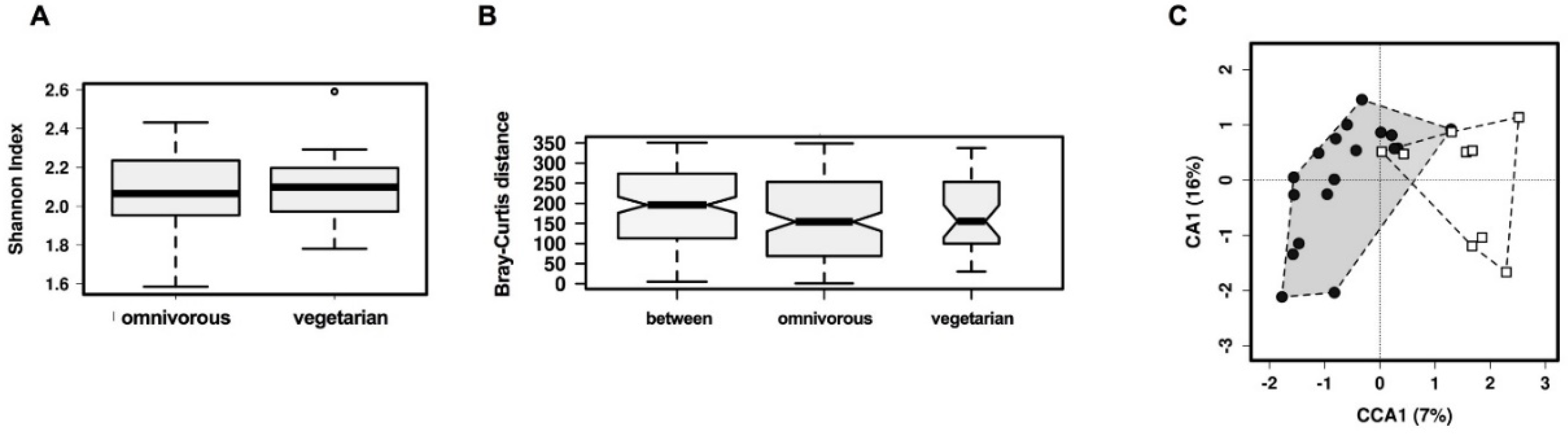
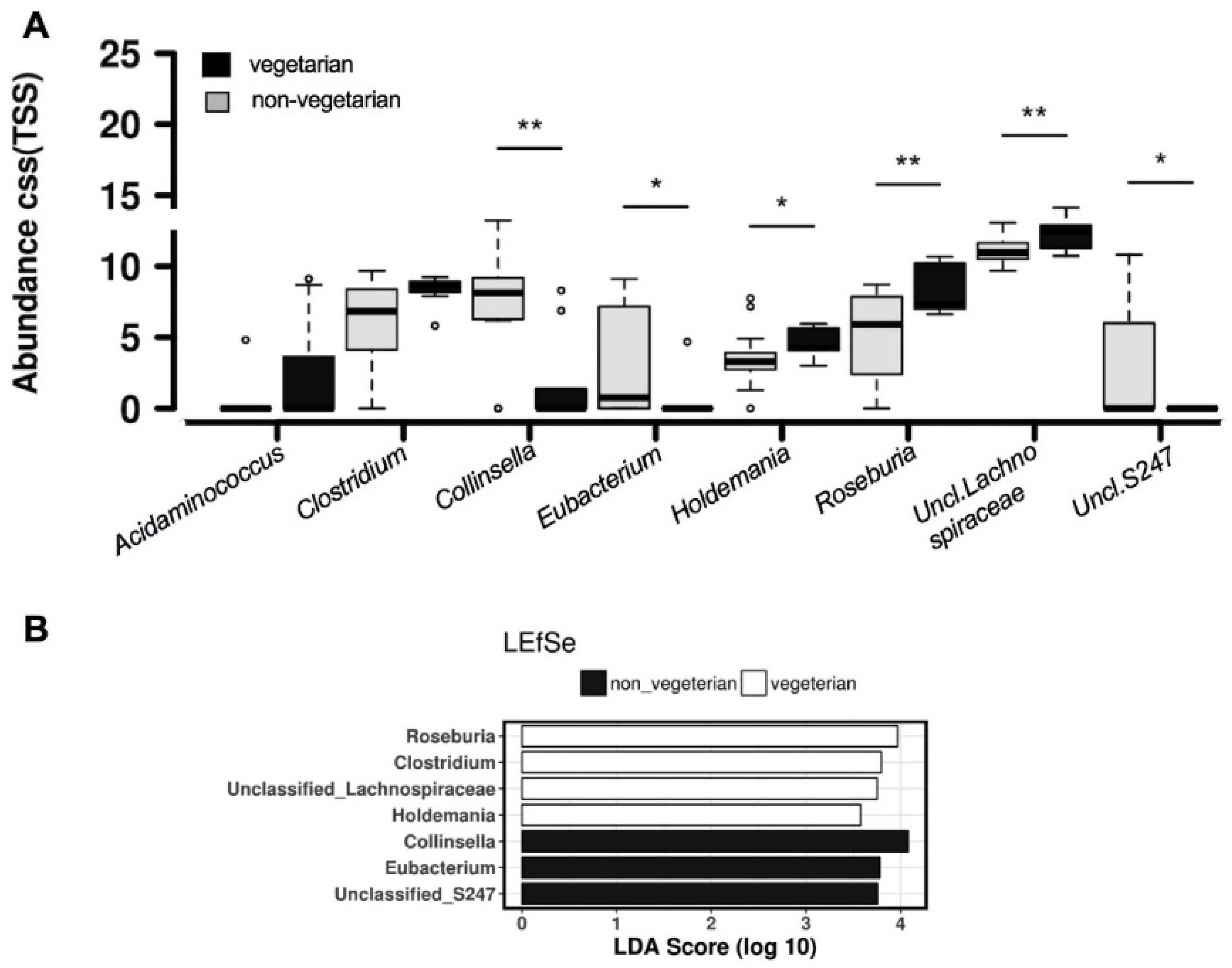
3.2. Comparison of Overall Gut Microbiota Composition
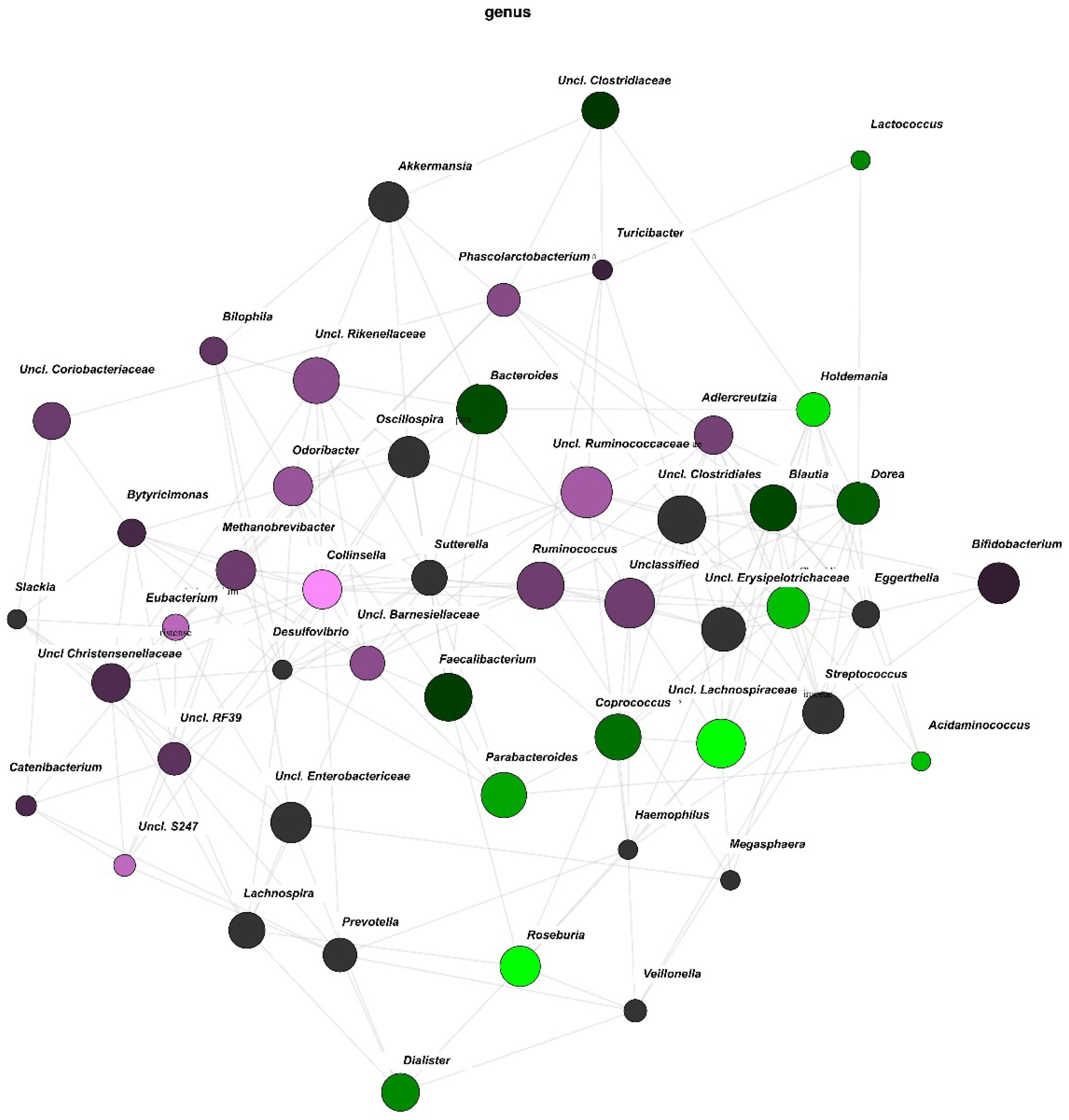
3.3. Network Analysis
3.4. Regression Analyses of Microbiota Profiles with Anthropometric Data
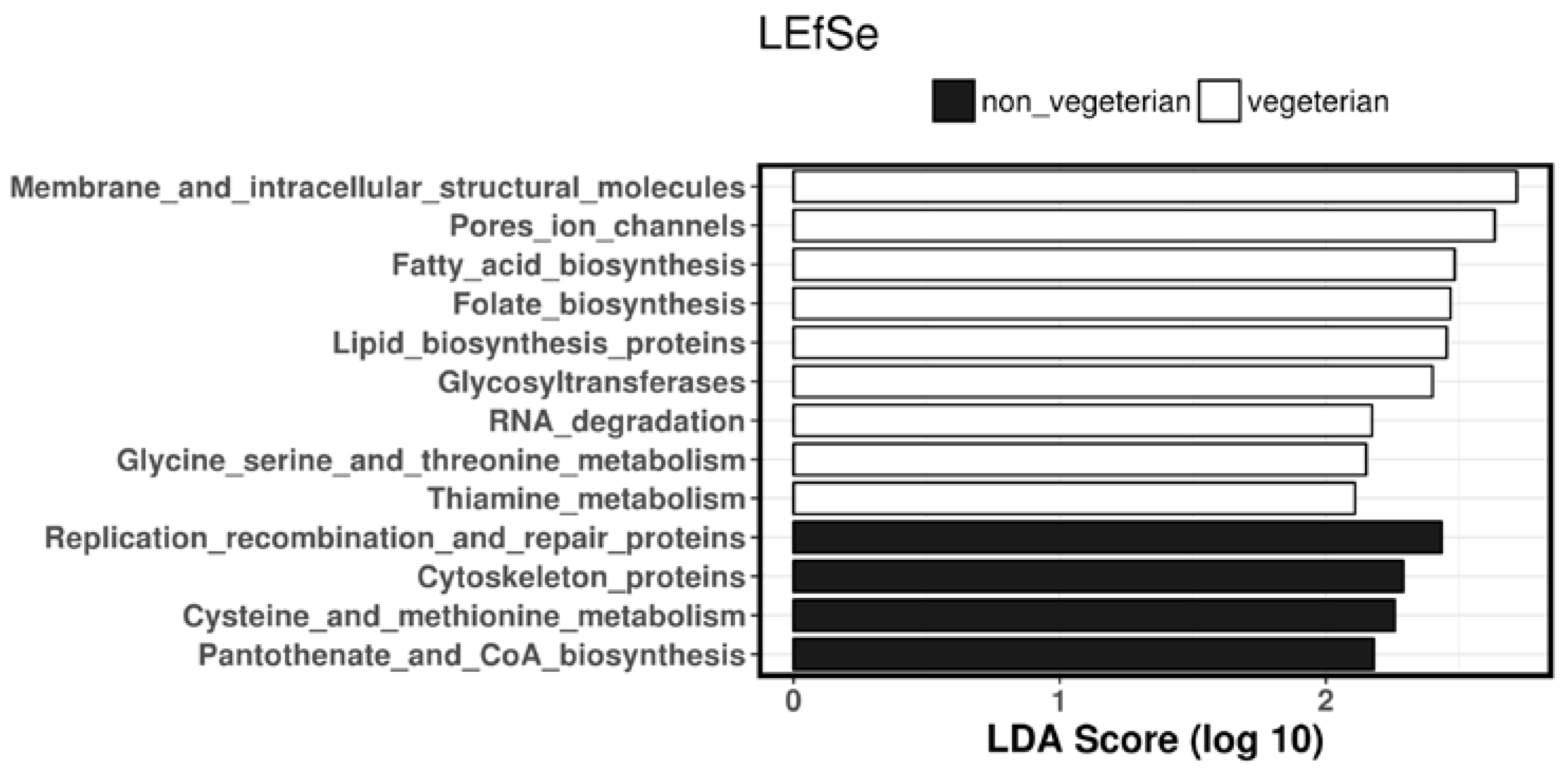
3.5. Predicted Biosynthesis Function Analyses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Losasso, C.; Eckert, E.M.; Mastrorilli, E.; Villiger, J.; Mancin, M.; Patuzzi, I.; Di Cesare, A.; Cibin, V.; Barrucci, F.; Pernthaler, J.; et al. Assessing the Influence of Vegan, Vegetarian and Omnivore Oriented Westernized Dietary Styles on Human Gut Microbiota: A Cross Sectional Study. Front. Microbiol. 2018, 9, 317. [Google Scholar] [CrossRef] [PubMed]
- Franco-de-Moraes, A.C.; de Almeida-Pititto, B.; da Rocha Fernandes, G.; Gomes, E.P.; da Costa Pereira, A.; Ferreira, S.R.G. Worse inflammatory profile in omnivores than in vegetarians associates with the gut microbiota composition. Diabetol. Metab. Syndr. 2017, 9, 62. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Bjorkman, A.; Cai, K.; Liu, G.; Wang, C.; Li, Y.; Xia, H.; Sun, L.; Kristiansen, K.; Wang, J.; et al. Impact of a 3-Months Vegetarian Diet on the Gut Microbiota and Immune Repertoire. Front. Immunol. 2018, 9, 908. [Google Scholar] [CrossRef] [PubMed]
- McMacken, M.; Shah, S. A plant-based diet for the prevention and treatment of type 2 diabetes. J. Geriatr. Cardiol. 2017, 14, 342–354. [Google Scholar] [PubMed]
- Kim, M.S.; Hwang, S.S.; Park, E.J.; Bae, J.W. Strict vegetarian diet improves the risk factors associated with metabolic diseases by modulating gut microbiota and reducing intestinal inflammation. Environ. Microbiol. Rep. 2013, 5, 765–775. [Google Scholar] [CrossRef] [PubMed]
- Koren, O.; Goodrich, J.K.; Cullender, T.C.; Spor, A.; Laitinen, K.; Kling Backhed, H.; Gonzalez, A.; Werner, J.J.; Angenent, L.T.; Knight, R.; et al. Host Remodeling of the Gut Microbiome and Metabolic Changes during Pregnancy. Cell 2012, 150, 470–480. [Google Scholar] [CrossRef] [PubMed]
- Nitert, M.D.; Barrett, H.L.; Foxcroft, K.; Tremellen, A.; Wilkinson, S.; Lingwood, B.; Tobin, J.M.; McSweeney, C.; O’Rourke, P.; McIntyre, H.D.; et al. SPRING: An RCT study of probiotics in the prevention of gestational diabetes mellitus in overweight and obese women. BMC Pregnancy Childbirth 2013, 13, 50. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Arango, L.F.; Barrett, H.L.; McIntyre, H.D.; Callaway, L.K.; Morrison, M.; Dekker Nitert, M.; SPRING Trial Group. Connections Between the Gut Microbiome and Metabolic Hormones in Early Pregnancy in Overweight and Obese Women. Diabetes 2016, 65, 2214–2223. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Morrison, M. Improved extraction of PCR-quality community DNA from digesta and fecal samples. Biotechniques 2004, 36, 808–812. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Zakrzewski, M.; Proietti, C.; Ellis, J.J.; Hasan, S.; Brion, M.-J.; Berger, B.; Krause, L. Calypso: A user-friendly web-server for mining and visualizing microbiome–environment interactions. Bioinformatics 2017, 33, 782–783. [Google Scholar] [CrossRef] [PubMed]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Arango, L.F.; Barrett, H.L.; Wilkinson, S.A.; Callaway, L.K.; McIntyre, H.D.; Morrison, M.; Dekker Nitert, M. Low dietary fiber intake increases Collinsella abundance in the gut microbiota of overweight and obese pregnant women. Gut Microbes 2017, 16, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Ruengsomwong, S.; La-Ongkham, O.; Jiang, J.; Wannissorn, B.; Nakayama, J.; Nitisinprasert, S. Microbial Community of Healthy Thai Vegetarians and Non-Vegetarians, Their Core Gut Microbiota, and Pathogen Risk. J. Microbiol. Biotechnol. 2016, 26, 1723–1735. [Google Scholar] [CrossRef] [PubMed]
- Furet, J.P.; Firmesse, O.; Gourmelon, M.; Bridonneau, C.; Tap, J.; Mondot, S.; Dore, J.; Corthier, G. Comparative assessment of human and farm animal faecal microbiota using real-time quantitative PCR. FEMS Microbiol. Ecol. 2009, 68, 351–362. [Google Scholar] [CrossRef] [PubMed]
- Matijasic, B.B.; Obermajer, T.; Lipoglavsek, L.; Grabnar, I.; Avgustin, G.; Rogelj, I. Association of dietary type with fecal microbiota in vegetarians and omnivores in Slovenia. Eur. J. Nutr. 2014, 53, 1051–1064. [Google Scholar] [CrossRef] [PubMed]
- Roytio, H.; Mokkala, K.; Vahlberg, T.; Laitinen, K. Dietary intake of fat and fibre according to reference values relates to higher gut microbiota richness in overweight pregnant women. Br. J. Nutr. 2017, 118, 343–352. [Google Scholar] [CrossRef] [PubMed]
- Holscher, H.D.; Taylor, A.M.; Swanson, K.S.; Novotny, J.A.; Baer, D.J. Almond Consumption and Processing Affects the Composition of the Gastrointestinal Microbiota of Healthy Adult Men and Women: A Randomized Controlled Trial. Nutrients 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Vanegas, S.M.; Meydani, M.; Barnett, J.B.; Goldin, B.; Kane, A.; Rasmussen, H.; Brown, C.; Vangay, P.; Knights, D.; Jonnalagadda, S.; et al. Substituting whole grains for refined grains in a 6-wk randomized trial has a modest effect on gut microbiota and immune and inflammatory markers of healthy adults. Am. J. Clin. Nutr. 2017, 105, 635–650. [Google Scholar] [CrossRef] [PubMed]
- Smith-Brown, P.; Morrison, M.; Krause, L.; Davies, P.S. Dairy and plant based food intakes are associated with altered faecal microbiota in 2 to 3 year old Australian children. Sci. Rep. 2016, 6, 32385. [Google Scholar] [CrossRef] [PubMed]
- Stanislawski, M.A.; Dabelea, D.; Wagner, B.D.; Sontag, M.K.; Lozupone, C.A.; Eggesbø, M. Pre-pregnancy weight, gestational weight gain, and the gut microbiota of mothers and their infants. Microbiome 2017, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Canani, R.B.; Costanzo, M.D.; Leone, L.; Pedata, M.; Meli, R.; Calignano, A. Potential beneficial effects of butyrate in intestinal and extraintestinal diseases. World J. Gastroenterol. 2011, 17, 1519–1528. [Google Scholar] [CrossRef] [PubMed]
- Haro, C.; Montes-Borrego, M.; Rangel-Zuniga, O.A.; Alcala-Diaz, J.F.; Gomez-Delgado, F.; Perez-Martinez, P.; Delgado-Lista, J.; Quintana-Navarro, G.M.; Tinahones, F.J.; Landa, B.B.; et al. Two Healthy Diets Modulate Gut Microbial Community Improving Insulin Sensitivity in a Human Obese Population. J. Clin. Endocrinol. Metab. 2016, 101, 233–242. [Google Scholar] [CrossRef] [PubMed]
- D’Hoe, K.; Conterno, L.; Fava, F.; Falony, G.; Vieira-Silva, S.; Vermeiren, J.; Tuohy, K.; Raes, J. Prebiotic Wheat Bran Fractions Induce Specific Microbiota Changes. Front. Microbiol. 2018, 9, 31. [Google Scholar] [CrossRef] [PubMed]
- Parsons, J.B.; Rock, C.O. Bacterial lipids: metabolism and membrane homeostasis. Prog. Lipid Res. 2013, 5, 249–276. [Google Scholar] [CrossRef] [PubMed]
- De Weirdt, R.; Hernandez-Sanabria, E.; Fievez, V.; Mees, E.; Geirnaert, A.; Van Herreweghen, F.; Vilchez-Vargas, R.; Van den Abbeele, P.; Jauregui, R.; Pieper Dietmar, H.; et al. Mucosa-associated biohydrogenating microbes protect the simulated colon microbiome from stress associated with high concentrations of poly-unsaturated fat. Environ. Microbiol. 2016, 19, 722–739. [Google Scholar] [CrossRef] [PubMed]
- Watson, H.; Mitra, S.; Croden, F.C.; Taylor, M.; Wood, H.M.; Perry, S.L.; Spencer, J.A.; Quirke, P.; Toogood, G.J.; Lawton, C.L.; et al. A randomised trial of the effect of omega-3 polyunsaturated fatty acid supplements on the human intestinal microbiota. Gut 2017. [Google Scholar] [CrossRef] [PubMed]
- Baer, D.J.; Stote, K.S.; Henderson, T.; Paul, D.R.; Okuma, K.; Tagami, H.; Kanahori, S.; Gordon, D.T.; Rumpler, W.V.; Ukhanova, M.; et al. The Metabolizable Energy of Dietary Resistant Maltodextrin is Variable and Alters Fecal Microbiota Composition in Adult Men. J. Nutr. 2014, 144, 1023–1029. [Google Scholar] [CrossRef] [PubMed]
- Willems, A.; Moore, W.E.; Weiss, N.; Collins, M.D. Phenotypic and phylogenetic characterization of some Eubacterium-like isolates containing a novel type B wall murein from human feces: Description of Holdemania filiformis gen. nov. sp. nov. Int. J. Syst. Bacteriol. 1997, 47, 1201–1204. [Google Scholar] [CrossRef] [PubMed]
| Vegetarian | Omnivorous | p Value | |
|---|---|---|---|
| N | 9 | 18 | ND |
| Maternal age (years) | 33 (29–34) | 34 (32–37) | 0.38 |
| Maternal BMI (kg/m2) * | 28.3 (26.5–35.5) | 28.4 (26.5–35–3) | 0.91 |
| Ethnicity | ND | ||
| Caucasian (%) | 7 (77.8) | 16 (88.8) | |
| Indian (%) | 2 (22.2) | 1 (5.6) | |
| Asian (%) | 0 (0) | 1 (5.6) | |
| Parity $ | 0 (0–2) | 1 (1–2) | ND |
| Systolic blood pressure (mmHg) | 110 (107–118) | 110 (101–112) | 0.46 |
| Diastolic blood pressure (mmHg) | 60 (58–70) | 63 (60–70) | 0.59 |
| Glucose (mmol/L) | 4.5 (4.4–4.6) | 4.3 (4.1–4.4) | 0.04 |
| HbA1c (%) | 4.8 (4.7–5.1) | 4.7 (4.6–5.0) | 0.41 |
| Future GDM (%) | 1 (11.1) | 2 (11.1) | 1 |
| Insulin | 4.7 (3.8–7.4) | 7.3 (4.6–8.3) | 0.38 |
| Total Cholesterol (mmol/L) | 5.4 (4.7–6.1) | 5.3 (4.6–5.9) | 0.71 |
| HDL cholesterol (mmol/L) | 2 (1.7–2.2) | 1.7 (1.6–2.0) | 0.11 |
| LDL cholesterol (mmol/L) | 3 (2.5–3.2) | 3.0 (2.4–3.3) | 0.88 |
| VLDL cholesterol (mmol/L) | 0.6 (0.4–0.7) | 0.6 (0.4–0.8) | 0.75 |
| Triglycerides (mmol/L) | 1.3 (1.0–1.8) | 1.4 (0.8–1.7) | 0.87 |
| Fetal sex (F/M) | 6/3 | 11/7 | 1 |
| Birth weight (g) | 3572 (3193–3992) | 3397 (2976–3978) | 0.52 |
| Birth length (cm) | 51.5 (49.1–53.0) | 50 (49.2–55.3) | 0.82 |
| Vegetarian | Omnivorous | p Value | |
|---|---|---|---|
| Overall Energy intake (kJ/day) | 4988 (3531–6387) | 5528 (5021–6428) | 0.15 |
| Protein (g/day) | 42.5 (32.7–59.2) | 71.1 (61.3–77) | 0.0003 |
| Iron intake (g/day)) | 8.1 | 8.4 | 0.56 |
| Carbohydrate (g/day) | 129.1 (106.5–154.4) | 144.6 (133.5–167) | 0.21 |
| Starch (g/day) | 76.5 (47.5–95.1) | 74.0 (54.1–88.0) | 0.78 |
| Sugars (g/day) | 56.0 (44.2–72.1) | 75.3 (64.2–92.3) | 0.02 |
| Dietary fibre (g/day) | 20.9 (13.9–26.7) | 16.5 (13.5–19.6) | 0.13 |
| Glycaemic Index | 51.5 (49.0–52.1) | 49.1 (46.7–52.6) | 0.40 |
| Saturated fatty acids (g/day) | 13.9 (11.3–23.8) | 21.5 (17.4–26.5) | 0.04 |
| Monounsaturated fatty acids (g/day) | 18.0 (10.6–23.8) | 18.9 (16.4–23.5) | 0.53 |
| Polyunsaturated fatty acids (g/day) | 11.6 (9.1–15.8) | 6.5 (5.0–8.6) | 0.006 |
| α-Linoleic acid (g/day) | 0.97 | 0.63 | 0.16 |
| Linoleic acid (g/day) | 10.4 | 5.7 | 0.008 |
| Arachidonic acid (g/day) | 0.02 | 0.05 | 0.05 |
| EPA (g/day) | 0.003 | 0.078 | 0.002 |
| DHA (g/day) | 0.01 | 0.17 | 0.002 |
| Genus | Rho | p Value | |
|---|---|---|---|
| HbA1c | Ruminococcus | −0.59 | 0.002 |
| Turicibacter | −0.47 | 0.016 | |
| Insulin | Coprococcus | −0.38 | 0.050 |
| Total cholesterol | Uncl. RF39 | −0.40 | 0.036 |
| Ruminococcus | 0.39 | 0.043 | |
| HDL cholesterol | Uncl. Coriobacteriaceae | −0.43 | 0.027 |
| Parabacteroides | 0.38 | 0.050 | |
| LDL cholesterol | Uncl. RF39 | −0.43 | 0.029 |
| VLDL cholesterol | Lachnospira | −0.43 | 0.029 |
| Collinsella | 0.39 | 0.048 | |
| Triglycerides | Lachnospira | −0.40 | 0.038 |
| Dietary Intake (g/day) | |||
| Protein | Adlercreutzia | 0.46 | 0.017 |
| Carbohydrates | Dialister | −0.42 | 0.028 |
| Ruminococcus | 0.41 | 0.034 | |
| Starch | Dialister | −0.47 | 0.013 |
| Uncl. Rikenellaceae | −0.42 | 0.030 | |
| Coprococcus | 0.41 | 0.033 | |
| Uncl. Clostridiaceae | 0.39 | 0.047 | |
| Fibre | Uncl. Lachnospiraceae | 0.67 | 0.0002 |
| Coprococcus | 0.58 | 0.002 | |
| Haemophilus | 0.44 | 0.021 | |
| Roseburia | 0.42 | 0.031 | |
| Clostridium | 0.41 | 0.036 | |
| Holdemania | 0.38 | 0.050 | |
| Glycaemic index | Holdemania | 0.51 | 0.007 |
| Prevotella | −0.48 | 0.011 | |
| Uncl. Costridiaceae | 0.44 | 0.020 | |
| Poly-unsaturated fatty acids | Holdemania | 0.47 | 0.012 |
| Collinsella | −0.46 | 0.017 | |
| Roseburia | 0.41 | 0.033 | |
| Uncl. Rikenellaceae | −0.41 | 0.035 | |
| Saturated fatty acids | Roseburia | −0.40 | 0.038 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barrett, H.L.; Gomez-Arango, L.F.; Wilkinson, S.A.; McIntyre, H.D.; Callaway, L.K.; Morrison, M.; Dekker Nitert, M. A Vegetarian Diet Is a Major Determinant of Gut Microbiota Composition in Early Pregnancy. Nutrients 2018, 10, 890. https://doi.org/10.3390/nu10070890
Barrett HL, Gomez-Arango LF, Wilkinson SA, McIntyre HD, Callaway LK, Morrison M, Dekker Nitert M. A Vegetarian Diet Is a Major Determinant of Gut Microbiota Composition in Early Pregnancy. Nutrients. 2018; 10(7):890. https://doi.org/10.3390/nu10070890
Chicago/Turabian StyleBarrett, Helen L., Luisa F. Gomez-Arango, Shelley A. Wilkinson, H. David McIntyre, Leonie K. Callaway, Mark Morrison, and Marloes Dekker Nitert. 2018. "A Vegetarian Diet Is a Major Determinant of Gut Microbiota Composition in Early Pregnancy" Nutrients 10, no. 7: 890. https://doi.org/10.3390/nu10070890
APA StyleBarrett, H. L., Gomez-Arango, L. F., Wilkinson, S. A., McIntyre, H. D., Callaway, L. K., Morrison, M., & Dekker Nitert, M. (2018). A Vegetarian Diet Is a Major Determinant of Gut Microbiota Composition in Early Pregnancy. Nutrients, 10(7), 890. https://doi.org/10.3390/nu10070890






