Detecting Drivers and Predicting Spatial Distribution of Soil Organic Carbon in an Arid Region Using Machine Learning
Highlights
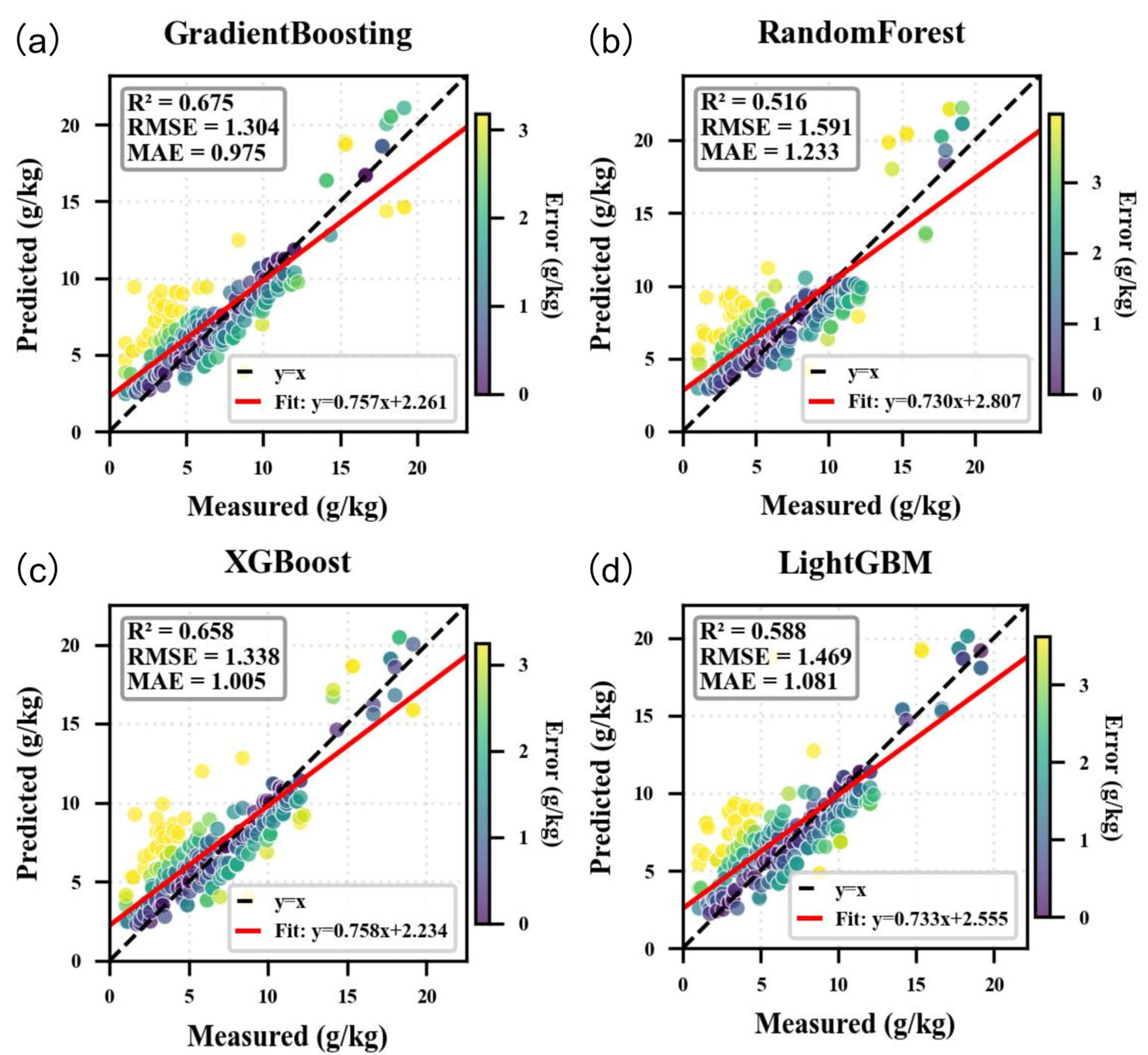
- In the arid Akesai region, the Gradient Boosting model (R2 = 0.675, RMSE = 1.304 g kg−1) outperformed other machine learning algorithms in predicting SOC, with vegetation-type factors (NDVI_PC1) and clay content identified as the most influential positive drivers of SOC accumulation.
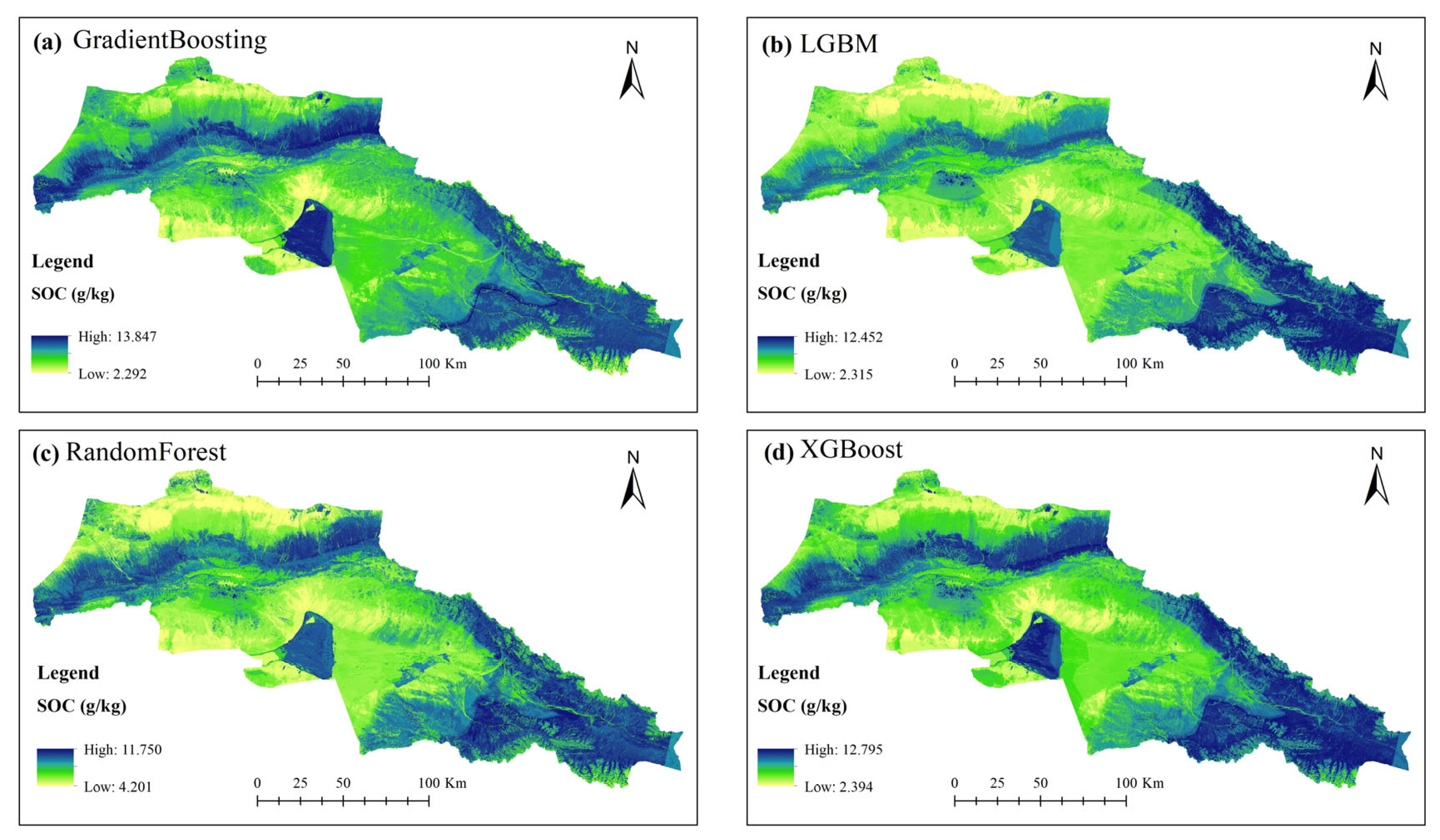
- The spatial distribution of SOC exhibits clear heterogeneity, with higher concentrations in mountainous and valley areas, which is primarily governed by the synergistic interplay of vegetation dynamics, soil texture, and topographic features.
- Methodologically, this study demonstrates that combining ensemble machine learning with interpretable SHAP analysis provides a robust and transparent framework for quantifying the contribution of multiple environmental factors to SOC variability in data-scarce arid regions.
- Practically speaking, the identified key drivers and high-resolution spatial patterns provide a scientific basis for targeted land management, ecological restoration, and more accurate carbon stock assessment in arid ecosystems.
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area
2.2. Soil Sampling and RS Data Acquisition and Preprocessing
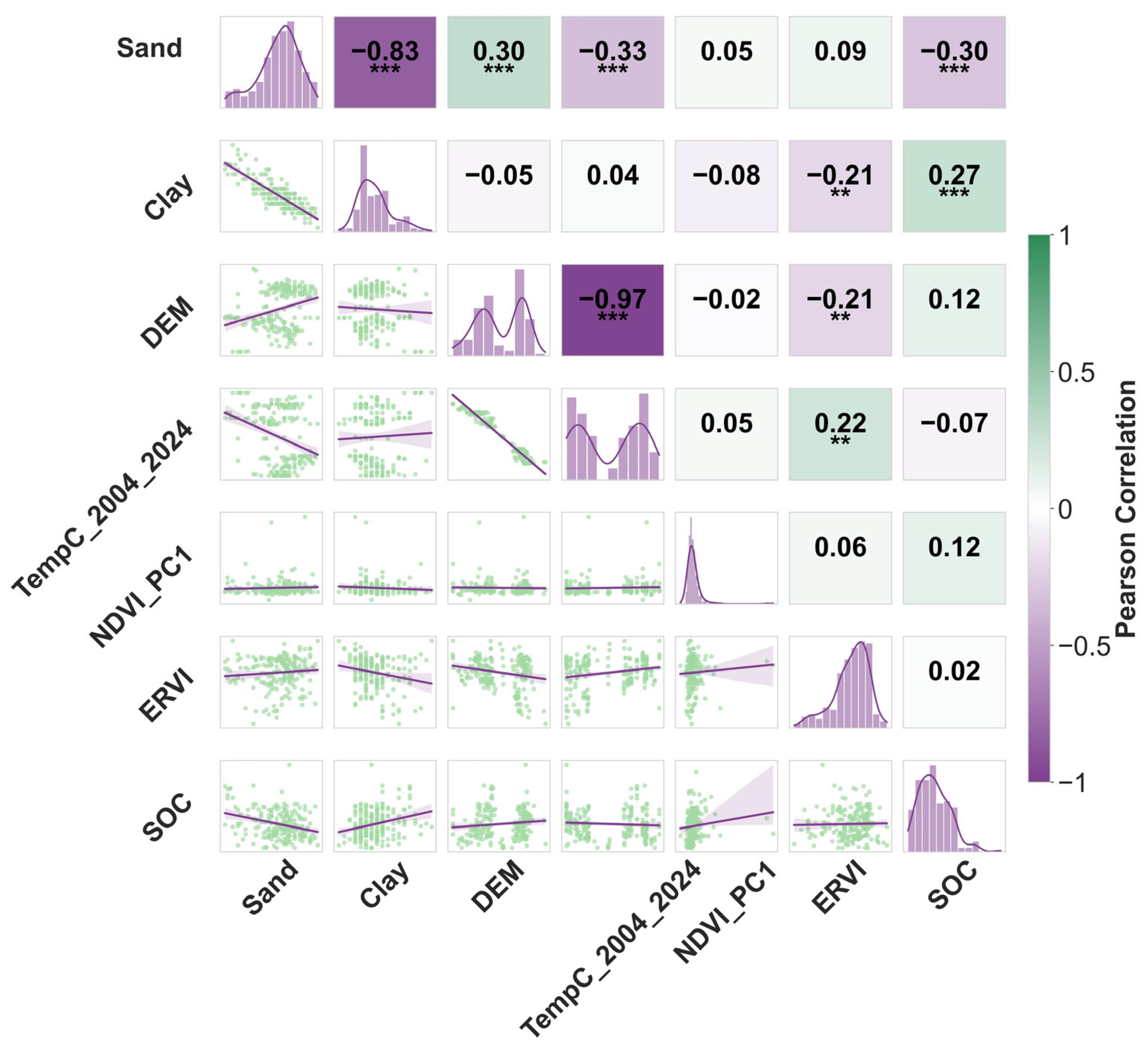
2.3. Variable Selection
2.4. Modeling Framework
2.4.1. Gradient Boosting Trees (GBT)
2.4.2. Random Forest (RF)
2.4.3. eXtreme Gradient Boosting (XGBoost)
2.4.4. Light Gradient Boosting Machine (LGBM)
2.5. Model Interpretability Using SHAP Analysis
2.6. Model Evaluation
3. Results
3.1. Descriptive Statistics of Soil Organic Carbon
3.2. Comparison of Model Accuracy
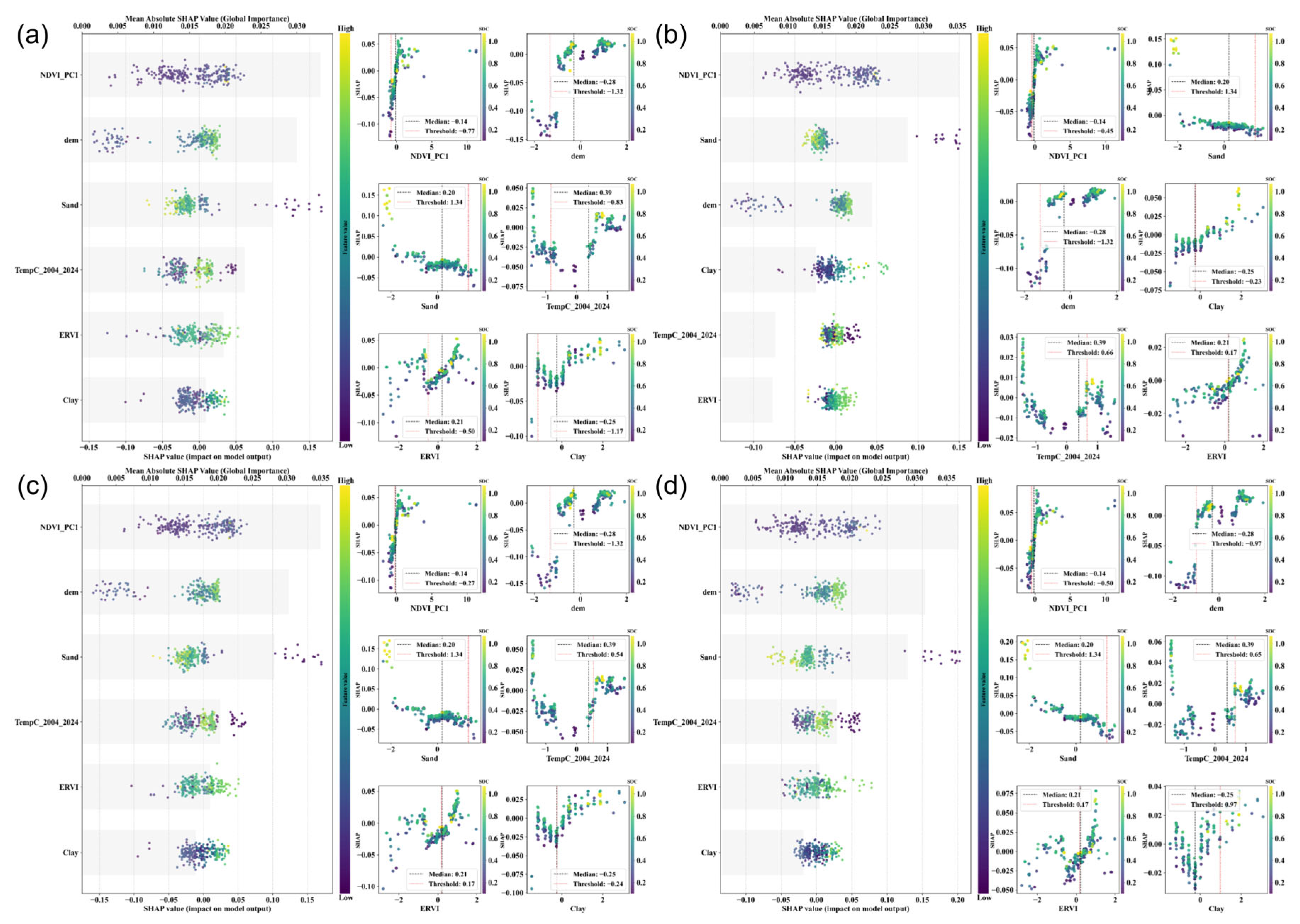
3.3. Feature Importance
3.4. Soil Organic Carbon Mapping
4. Discussion
4.1. Comparing the Effects of Different Characteristics on Soil Organic Carbon Mapping
4.2. Uncertainty Analysis of the Current Study
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Appendix A
References
- Wang, X.; Li, L.; Liu, H.; Song, K.; Wang, L.; Meng, X. Prediction of soil organic matter using VNIR spectral parameters extracted from shape characteristics. Soil Tillage Res. 2022, 216, 105241. [Google Scholar] [CrossRef]
- Bradford, M.A.; Wieder, W.R.; Bonan, G.B.; Fierer, N.; Raymond, P.A.; Crowther, T.W. Managing uncertainty in soil carbon feedbacks to climate change. Nat. Clim. Change 2016, 6, 751–758. [Google Scholar] [CrossRef]
- Wang, X.; Wang, L.; Li, S.; Wang, Z.; Zheng, M.; Song, K. Remote estimates of soil organic carbon using multi-temporal synthetic images and the probability hybrid model. Geoderma 2022, 425, 116066. [Google Scholar] [CrossRef]
- Iucn, D.J.; Gudka, M.; Laban, P.; Metternicht, G.; Alexander, S.; Hannam, I. Land Degradation Neutrality: Implications and Opportunities for Conservation; Technical Brief 2nd Edition; IUCN: Nairobi, Kenya, 2015; 19p. [Google Scholar]
- Lamichhane, S.; Kumar, L.; Wilson, B. Digital soil mapping algorithms and covariates for soil organic carbon mapping and their implications: A review. Geoderma 2019, 352, 395–413. [Google Scholar] [CrossRef]
- Bruun, T.B.; Elberling, B.; de Neergaard, A.; Magid, J. Organic Carbon Dynamics in Different Soil Types After Conversion of Forest to Agriculture. Land Degrad. Dev. 2013, 26, 272–283. [Google Scholar] [CrossRef]
- Saia, S.; Benítez, E.; GarcíA-Garrido, J.M.; Settanni, L.; Amato, G.; Giambalvo, D. The effect of arbuscular mycorrhizal fungi on total plant nitrogen uptake and nitrogen recovery from soil organic material. J. Agric. Sci. 2013, 152, 370–378. [Google Scholar] [CrossRef]
- Cui, Z.A.; Zhang, R.; Wang, W.; Peng, Z.; Wu, Y.; Zhao, Z.; Li, M.; Cong, Y.; Zhang, S.; Li, Z.; et al. Unveiling critical drivers of soil salinity prediction accuracy in remote sensing: A global meta-analysis. Plant Soil 2025, 516, 33–65. [Google Scholar] [CrossRef]
- Zhang, J.; Ge, X.; Hou, X.; Han, L.; Zhang, Z.; Feng, W.; Zhou, Z.; Luo, X. Strategies for Soil Salinity Mapping Using Remote Sensing and Machine Learning in the Yellow River Delta. Remote Sens. 2025, 17, 2619. [Google Scholar] [CrossRef]
- Dai, F.; Zhou, Q.; Lv, Z.; Wang, X.; Liu, G. Spatial prediction of soil organic matter content integrating artificial neural network and ordinary kriging in Tibetan Plateau. Ecol. Indic. 2014, 45, 184–194. [Google Scholar] [CrossRef]
- Liu, S.; An, N.; Yang, J.; Dong, S.; Wang, C.; Yin, Y. Prediction of soil organic matter variability associated with different land use types in mountainous landscape in southwestern Yunnan province, China. Catena 2015, 133, 137–144. [Google Scholar] [CrossRef]
- Mishra, U.; Drewniak, B.; Jastrow, J.D.; Matamala, R.M.; Vitharana, U.W.A. Spatial representation of organic carbon and active-layer thickness of high latitude soils in CMIP5 earth system models. Geoderma 2017, 300, 55–63. [Google Scholar] [CrossRef]
- Zeng, C.; Yang, L.; Zhu, A.X.; Rossiter, D.G.; Liu, J.; Liu, J.; Qin, C.; Wang, D. Mapping soil organic matter concentration at different scales using a mixed geographically weighted regression method. Geoderma 2016, 281, 69–82. [Google Scholar] [CrossRef]
- Mirzaee, S.; Ghorbani-Dashtaki, S.; Mohammadi, J.; Asadi, H.; Asadzadeh, F. Spatial variability of soil organic matter using remote sensing data. Catena 2016, 145, 118–127. [Google Scholar] [CrossRef]
- Hoffmann, U.; Hoffmann, T.; Jurasinski, G.; Glatzel, S.; Kuhn, N.J. Assessing the spatial variability of soil organic carbon stocks in an alpine setting (Grindelwald, Swiss Alps). Geoderma 2014, 232–234, 270–283. [Google Scholar] [CrossRef]
- Piccini, C.; Marchetti, A.; Francaviglia, R. Estimation of soil organic matter by geostatistical methods: Use of auxiliary information in agricultural and environmental assessment. Ecol. Indic. 2014, 36, 301–314. [Google Scholar] [CrossRef]
- Ge, X.; Ding, J.; Teng, D.; Wang, J.; Huo, T.; Jin, X.; Wang, J.; He, B.; Han, L. Updated soil salinity with fine spatial resolution and high accuracy: The synergy of Sentinel-2 MSI, environmental covariates and hybrid machine learning approaches. Catena 2022, 212, 106054. [Google Scholar] [CrossRef]
- Han, L.; Liu, D.; Cheng, G.; Zhang, G.; Wang, L. Spatial distribution and genesis of salt on the saline playa at Qehan Lake, Inner Mongolia, China. Catena 2019, 177, 22–30. [Google Scholar] [CrossRef]
- Han, L.; Ding, J.; Zhang, J.; Chen, P.; Wang, J.; Wang, Y.; Wang, J.; Ge, X.; Zhang, Z. Precipitation events determine the spatiotemporal distribution of playa surface salinity in arid regions: Evidence from satellite data fused via the enhanced spatial and temporal adaptive reflectance fusion model. Catena 2021, 206, 105546. [Google Scholar] [CrossRef]
- Jandl, R.; Rodeghiero, M.; Martinez, C.; Cotrufo, M.F.; Bampa, F.; van Wesemael, B.; Harrison, R.B.; Guerrini, I.A.; Richter, D.D., Jr.; Rustad, L.; et al. Current status, uncertainty and future needs in soil organic carbon monitoring. Sci. Total Environ. 2014, 468-469, 376–383. [Google Scholar] [CrossRef]
- Viaud, V.; Angers, D.A.; Walter, C. Toward Landscape-Scale Modeling of Soil Organic Matter Dynamics in Agroecosystems. Soil Sci. Soc. Am. J. 2010, 74, 1847–1860. [Google Scholar] [CrossRef]
- Aksoy, E.; Yigini, Y.; Montanarella, L. Combining Soil Databases for Topsoil Organic Carbon Mapping in Europe. PLoS ONE 2016, 11, e0152098. [Google Scholar] [CrossRef] [PubMed]
- Rial, M.; Martinez Cortizas, A.; Rodriguez-Lado, L. Understanding the spatial distribution of factors controlling topsoil organic carbon content in European soils. Sci. Total Environ. 2017, 609, 1411–1422. [Google Scholar] [CrossRef]
- Schillaci, C.; Acutis, M.; Lombardo, L.; Lipani, A.; Fantappie, M.; Marker, M.; Saia, S. Spatio-temporal topsoil organic carbon mapping of a semi-arid Mediterranean region: The role of land use, soil texture, topographic indices and the influence of remote sensing data to modelling. Sci. Total Environ. 2017, 601–602, 821–832. [Google Scholar] [CrossRef] [PubMed]
- Yigini, Y.; Panagos, P. Assessment of soil organic carbon stocks under future climate and land cover changes in Europe. Sci. Total Environ. 2016, 557-558, 838–850. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhao, Y.; Taylor, J.; Gaulton, R.; Jin, X.; Song, X.; Li, Z.; Meng, Y.; Chen, P.; Feng, H.; et al. Comparison and transferability of thermal, temporal and phenological-based in-season predictions of above-ground biomass in wheat crops from proximal crop reflectance data. Remote Sens. Environ. 2022, 273, 112967. [Google Scholar] [CrossRef]
- Li, T.; Cui, L.; Kuhnert, M.; McLaren, T.I.; Pandey, R.; Liu, H.; Wang, W.; Xu, Z.; Xia, A.; Dalal, R.C.; et al. A comprehensive review of soil organic carbon estimates: Integrating remote sensing and machine learning technologies. J. Soils Sediments 2024, 24, 3556–3571. [Google Scholar] [CrossRef]
- Rukhovich, D.; Koroleva, P.; Rukhovich, A.; Komissarov, M. A detailed mapping of soil organic matter content in arable land based on the multitemporal soil line coefficients and neural network filtering of big remote sensing data. Geoderma 2024, 447, 116941. [Google Scholar] [CrossRef]
- Dong, C.; Meng, X.; Ruan, W.; Cui, J.; Zhang, X.; Liu, H. An innoval hyperspectral prediction model for soil organic matter in croplands of the Northeast China Mollisols Region. Soil Tillage Res. 2025, 253, 106666. [Google Scholar] [CrossRef]
- Li, X.; Ding, J.; Liu, J.; Ge, X.; Zhang, J. Digital Mapping of Soil Organic Carbon Using Sentinel Series Data: A Case Study of the Ebinur Lake Watershed in Xinjiang. Remote Sens. 2021, 13, 769. [Google Scholar] [CrossRef]
- Wang, X.; Li, S.; Wang, L.; Zheng, M.; Wang, Z.; Song, K. Effects of cropland reclamation on soil organic carbon in China’s black soil region over the past 35 years. Glob. Chang. Biol. 2023, 29, 5460–5477. [Google Scholar] [CrossRef]
- Wang, X.; Song, K.; Wang, Z.; Li, S.; Shang, Y.; Liu, G. Effects of land conversion to cropland on soil organic carbon in montane soils of Northeast China from 1985 to 2020. Catena 2024, 235, 107691. [Google Scholar] [CrossRef]
- Wang, X.; Song, K.; Wang, Z.; Li, S.; Zheng, M.; Wen, Z.; Liu, G. Are topsoil spectra or soil-environmental factors better indicators for discrimination of soil classes? Catena 2022, 218, 106580. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, S.; Adhikari, K.; Wang, Q.; Sui, Y.; Xin, G. Effect of cultivation history on soil organic carbon status of arable land in northeastern China. Geoderma 2019, 342, 55–64. [Google Scholar] [CrossRef]
- Zhang, Z.; Ding, J.; Zhu, C.; Wang, J.; Ge, X.; Li, X.; Han, L.; Chen, X.; Wang, J. Historical and future variation of soil organic carbon in China. Geoderma 2023, 436, 116557. [Google Scholar] [CrossRef]
- Zhou, Z.; Zhang, Z.; Wang, M.; Wang, K.; Ai, J.; Gill, A.; Temirbayeva, K.; Zhu, C. Impact of future climate warming on soil organic carbon in China based on process-based models. Clim. Smart Agric. 2025, 2, 100086. [Google Scholar] [CrossRef]
- Zhang, Z.; Ding, J.; Li, L.; Cao, J.; Wang, K.; Zhu, C.; Ge, X.; Wang, J.; Yang, C.; Li, F.; et al. The impact of extreme climate on soil organic carbon in China. Geogr. Sustain. 2025, 6, 100356. [Google Scholar] [CrossRef]
- Zhang, Z.; Ding, J.; Wang, J.; Ge, X. Prediction of soil organic matter in northwestern China using fractional-order derivative spectroscopy and modified normalized difference indices. Catena 2020, 185, 104257. [Google Scholar] [CrossRef]
- Zhang, Z.; Ding, J.; Zhu, C.; Wang, J.; Ma, G.; Ge, X.; Li, Z.; Han, L. Strategies for the efficient estimation of soil organic matter in salt-affected soils through Vis-NIR spectroscopy: Optimal band combination algorithm and spectral degradation. Geoderma 2021, 382, 114729. [Google Scholar] [CrossRef]
- Guo, B.; Yang, X.; Yang, M.; Sun, D.; Zhu, W.; Zhu, D.; Wang, J. Mapping soil salinity using a combination of vegetation index time series and single-temporal remote sensing images in the Yellow River Delta, China. Catena 2023, 231, 107313. [Google Scholar] [CrossRef]
- Liu, H.; Guo, B.; Yang, X.; Zhao, J.; Li, M.; Huo, Y.; Wang, J. High spatiotemporal resolution vegetation index time series can facilitate enhanced remote sensing monitoring of soil salinization. Plant Soil 2024, 510, 305–327. [Google Scholar] [CrossRef]
- Xing, H.; Niu, J.; Feng, Y.; Hou, D.; Wang, Y.; Wang, Z. A coastal wetlands mapping approach of Yellow River Delta with a hierarchical classification and optimal feature selection framework. Catena 2023, 223, 106897. [Google Scholar] [CrossRef]
- Zhao, J.; Wang, Z.; Zhang, Q.; Niu, Y.; Lu, Z.; Zhao, Z. A novel feature selection criterion for wetland mapping using GF-3 and Sentinel-2 Data. Ecol. Indic. 2025, 171, 113146. [Google Scholar] [CrossRef]
- Miao, J.; Niu, L. A Survey on Feature Selection. Procedia Comput. Sci. 2016, 91, 919–926. [Google Scholar] [CrossRef]
- Bentéjac, C.; Csörgő, A.; Martínez-Muñoz, G. A comparative analysis of gradient boosting algorithms. Artif. Intell. Rev. 2020, 54, 1937–1967. [Google Scholar] [CrossRef]
- Li, X.; Jia, H.; Wang, L. Remote Sensing Monitoring of Drought in Southwest China Using Random Forest and eXtreme Gradient Boosting Methods. Remote Sens. 2023, 15, 4840. [Google Scholar] [CrossRef]
- Cutler, A.; Cutler, D.R.; Stevens, J.R. Random Forests. In Ensemble Machine Learning; Springer: Berlin/Heidelberg, Germany, 2012; pp. 157–175. [Google Scholar]
- Belgiu, M.; Drăguţ, L. Random forest in remote sensing: A review of applications and future directions. ISPRS J. Photogramm. Remote Sens. 2016, 114, 24–31. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. XGBoost. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Liu, Z.; Jiang, P.; De Bock, K.W.; Wang, J.; Zhang, L.; Niu, X. Extreme gradient boosting trees with efficient Bayesian optimization for profit-driven customer churn prediction. Technol. Forecast. Soc. Change 2024, 198, 122945. [Google Scholar] [CrossRef]
- Yan, J.; Xu, Y.; Cheng, Q.; Jiang, S.; Wang, Q.; Xiao, Y.; Ma, C.; Yan, J.; Wang, X. LightGBM: Accelerated genomically designed crop breeding through ensemble learning. Genome Biol. 2021, 22, 271. [Google Scholar] [CrossRef]
- Fan, J.; Ma, X.; Wu, L.; Zhang, F.; Yu, X.; Zeng, W. Light Gradient Boosting Machine: An efficient soft computing model for estimating daily reference evapotranspiration with local and external meteorological data. Agric. Water Manag. 2019, 225, 105758. [Google Scholar] [CrossRef]
- Li, X.; Wang, H.; Qin, S.; Lin, L.; Wang, X.; Cornelis, W. Evaluating ensemble learning in developing pedotransfer functions to predict soil hydraulic properties. J. Hydrol. 2024, 640, 131658. [Google Scholar] [CrossRef]
- Antwarg, L.; Miller, R.M.; Shapira, B.; Rokach, L. Explaining anomalies detected by autoencoders using Shapley Additive Explanations. Expert Syst. Appl. 2021, 186, 115736. [Google Scholar] [CrossRef]
- Al-Najjar, H.A.H.; Pradhan, B.; Beydoun, G.; Sarkar, R.; Park, H.-J.; Alamri, A. A novel method using explainable artificial intelligence (XAI)-based Shapley Additive Explanations for spatial landslide prediction using Time-Series SAR dataset. Gondwana Res. 2023, 123, 107–124. [Google Scholar] [CrossRef]
- Wang, T.; Kang, F.; Cheng, X.; Han, H.; Bai, Y.; Ma, J. Spatial variability of organic carbon and total nitrogen in the soils of a subalpine forested catchment at Mt. Taiyue, China. Catena 2017, 155, 41–52. [Google Scholar] [CrossRef]
- Swetha, R.K.; Dasgupta, S.; Chakraborty, S.; Li, B.; Weindorf, D.C.; Mancini, M.; Silva, S.H.G.; Ribeiro, B.T.; Curi, N.; Ray, D.P. Using Nix color sensor and Munsell soil color variables to classify contrasting soil types and predict soil organic carbon in Eastern India. Comput. Electron. Agric. 2022, 199, 107192. [Google Scholar] [CrossRef]
- Yang, R.-M.; Zhang, G.-L.; Liu, F.; Lu, Y.-Y.; Yang, F.; Yang, F.; Yang, M.; Zhao, Y.-G.; Li, D.-C. Comparison of boosted regression tree and random forest models for mapping topsoil organic carbon concentration in an alpine ecosystem. Ecol. Indic. 2016, 60, 870–878. [Google Scholar] [CrossRef]
- Wan, Q.; Zhu, G.; Guo, H.; Zhang, Y.; Pan, H.; Yong, L.; Ma, H. Influence of Vegetation Coverage and Climate Environment on Soil Organic Carbon in the Qilian Mountains. Sci. Rep. 2019, 9, 17623. [Google Scholar] [CrossRef]
- Hounkpatin, K.O.L.; Stendahl, J.; Lundblad, M.; Karltun, E. Predicting the spatial distribution of soil organic carbon stock in Swedish forests using a group of covariates and site-specific data. Soil 2021, 7, 377–398. [Google Scholar] [CrossRef]
- He, X.; Yang, L.; Li, A.; Zhang, L.; Shen, F.; Cai, Y.; Zhou, C. Soil organic carbon prediction using phenological parameters and remote sensing variables generated from Sentinel-2 images. Catena 2021, 205, 105442. [Google Scholar] [CrossRef]
- Basile-Doelsch, I.; Balesdent, J.; Pellerin, S. Reviews and syntheses: The mechanisms underlying carbon storage in soil. Biogeosciences 2020, 17, 5223–5242. [Google Scholar] [CrossRef]
- Sun, D.; Qiu, X.; Feng, J.; Ru, J.; Song, J.; Wan, S. Forest types control the contribution of litter and roots to labile and persistent soil organic carbon. Biogeochemistry 2024, 167, 1609–1617. [Google Scholar] [CrossRef]
- Sastre, B.; Antón-Iruela, O.; Moreno-Delafuente, A.; Navas, M.J.; Marques, M.J.; González-Canales, J.; Martín-Sanz, J.P.; Ramos, R.; García-Díaz, A.; Bienes, R. Groundcovers Improve Soil Properties in Woody Crops Under Semiarid Climate. Agriculture 2024, 14, 2288. [Google Scholar] [CrossRef]
- Plaza-Bonilla, D.; Arrúe, J.L.; Cantero-Martínez, C.; Fanlo, R.; Iglesias, A.; Álvaro-Fuentes, J. Carbon management in dryland agricultural systems. A review. Agron. Sustain. Dev. 2015, 35, 1319–1334. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, S.; Meng, X.; Wang, L.; Yu, Y.; Zhang, Y.; Zhang, G.; Zhang, S.; Dai, X.; Ruan, W.; et al. Climatic and topographic controls on soil organic carbon distribution across continents. Catena 2025, 260, 109435. [Google Scholar] [CrossRef]
- Zhu, G.; Zhou, L.; He, X.; Wei, P.; Lin, D.; Qian, S.; Zhao, L.; Luo, M.; Yin, X.; Zeng, L.; et al. Effects of Elevation Gradient on Soil Carbon and Nitrogen in a Typical Karst Region of Chongqing, Southwest China. J. Geophys. Res. Biogeosci. 2022, 127, e2021JG006742. [Google Scholar] [CrossRef]
- Schapel, A.; Marschner, P.; Churchman, J. Clay amount and distribution influence organic carbon content in sand with subsoil clay addition. Soil Tillage Res. 2018, 184, 253–260. [Google Scholar] [CrossRef]
- Nguyen-Sy, T. Optimized hybrid XGBoost-CatBoost model for enhanced prediction of concrete strength and reliability analysis using Monte Carlo simulations. Appl. Soft Comput. 2024, 167, 112490. [Google Scholar] [CrossRef]
- Roberts, D.R.; Bahn, V.; Ciuti, S.; Boyce, M.S.; Elith, J.; Guillera-Arroita, G.; Hauenstein, S.; Lahoz-Monfort, J.J.; Schröder, B.; Thuiller, W.; et al. Cross-validation strategies for data with temporal, spatial, hierarchical, or phylogenetic structure. Ecography 2017, 40, 913–929. [Google Scholar] [CrossRef]
- Ma, L.; Liu, Y.; Zhang, X.; Ye, Y.; Yin, G.; Johnson, B.A. Deep learning in remote sensing applications: A meta-analysis and review. ISPRS J. Photogramm. Remote Sens. 2019, 152, 166–177. [Google Scholar] [CrossRef]




| Category | Predictors | Abbreviation |
|---|---|---|
| Vegetation spectral indices | Difference Vegetation Index | SR |
| Enhanced Normalized Difference Vegetation Index | ENDVI | |
| Enhanced Ratio Vegetation Index | ERVI | |
| Enhanced Vegetation Index | EVI | |
| Green Difference Vegetation Index | GDVI | |
| Green Ratio Vegetation Index, | GRVI | |
| Modified Soil Adjusted Vegetation Index | MSAVI | |
| Normalized Difference Vegetation Index | NDVI | |
| Soil Adjusted Vegetation Index | SAVI | |
| Normalized Difference Water Index | NDWI | |
| Simple Ratio Index | SR | |
| Salinity indices | Salinity index I–V | SI1, SI2, SI3, SI4, SI5 |
| Soil properties | – | Clay, Sand |
| Spectral band data | Sentinel-2 bands | B1, B11, B12, B2, B3, B4, B6, B7, B8, B8A, B9 |
| Climate data | Mean precipitation and temperature from 2004 to 2024 | Precip_2004_2024, TempC_2004_2024 |
| Topographic data | DEM | |
| Vegetation-type factors | Principal components of NDVI time-series Maximum, minimum, and mean of NDVI time-series | NDVI_PC1, NDVI_PC2, NDVI_max, NDVI_mean, NDVI_min |
| Dataset | Mean | SD | Skewness | Kurtosis | CV | Min | Median | Max |
|---|---|---|---|---|---|---|---|---|
| SOC (g/kg) | 4.728 | 2.208 | 0.784 | 1.021 | 0.467 | 1.000 | 4.500 | 14.100 |
| Sand (%) | 69.816 | 7.984 | −0.676 | 0.093 | 0.114 | 49.000 | 71.000 | 85.000 |
| Clay (%) | 7.609 | 3.068 | 0.923 | 0.705 | 0.403 | 1.000 | 7.000 | 18.00 |
| NDVI_PC1 | −0.028 | 1.325 | 6.009 | 46.115 | −47.741 | −1.977 | −0.290 | 11.518 |
| DEM (m) | 3114.908 | 708.531 | −0.207 | −1.221 | 0.227 | 1647.000 | 2930.000 | 4509.000 |
| ERVI | 2.553 | 0.294 | −0.879 | 0.591 | 0.115 | 1.594 | 2.614 | 3.152 |
| TempC_2004_2024 (°C) | 0.035 | 6.243 | −0.038 | −1.634 | 180.175 | −8.501 | 2.782 | 9.739 |
| Model | Train R2 | Train RMSE | Train MAE | Train RPIQ | Test R2 | Test RMSE | Test MAE | Test RPIQ |
|---|---|---|---|---|---|---|---|---|
| Gradient Boosting | 0.790 | 1.040 | 0.809 | 3.558 | 0.675 | 1.304 | 0.975 | 2.837 |
| Random Forest | 0.614 | 1.410 | 1.120 | 2.624 | 0.516 | 1.591 | 1.233 | 2.326 |
| XGBoost | 0.761 | 1.111 | 0.847 | 3.331 | 0.658 | 1.338 | 1.005 | 2.765 |
| LightGBM | 0.684 | 1.276 | 0.972 | 2.900 | 0.588 | 1.469 | 1.081 | 2.519 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Chen, G.; Ge, X.; Zhang, Z.; Han, L. Detecting Drivers and Predicting Spatial Distribution of Soil Organic Carbon in an Arid Region Using Machine Learning. Remote Sens. 2026, 18, 535. https://doi.org/10.3390/rs18040535
Chen G, Ge X, Zhang Z, Han L. Detecting Drivers and Predicting Spatial Distribution of Soil Organic Carbon in an Arid Region Using Machine Learning. Remote Sensing. 2026; 18(4):535. https://doi.org/10.3390/rs18040535
Chicago/Turabian StyleChen, Guiren, Xianghe Ge, Zipeng Zhang, and Lijing Han. 2026. "Detecting Drivers and Predicting Spatial Distribution of Soil Organic Carbon in an Arid Region Using Machine Learning" Remote Sensing 18, no. 4: 535. https://doi.org/10.3390/rs18040535
APA StyleChen, G., Ge, X., Zhang, Z., & Han, L. (2026). Detecting Drivers and Predicting Spatial Distribution of Soil Organic Carbon in an Arid Region Using Machine Learning. Remote Sensing, 18(4), 535. https://doi.org/10.3390/rs18040535







