High-Resolution Mapping Coastal Wetland Vegetation Using Frequency-Augmented Deep Learning Method
Highlights
- First use of very-high resolution (2 cm) image data for Mapping Coastal Wetland Vegetation: We have used drone imagery and deep learning techniques to complete the task of fine tuned classification of Wetland Vegetation.
- A method for coastal vegetation classification from high resolution imagery is proposed: In this paper, we proposal a augment frequency-domain features network—AFDFNet. Experimental results demonstrate that AFDFNet consistently outperforms existing deep learning models, achieving state-of-the-art performance.
- An Multi-scale Feature Enhancement Module: We designed a multi-scale feature enhancement module to compensate for the misclassification phenomenon caused by the lack of frequency domain features and contextual information in the network.
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset Construction
2.2. AFDFNet Model Development
2.2.1. Overall Framework
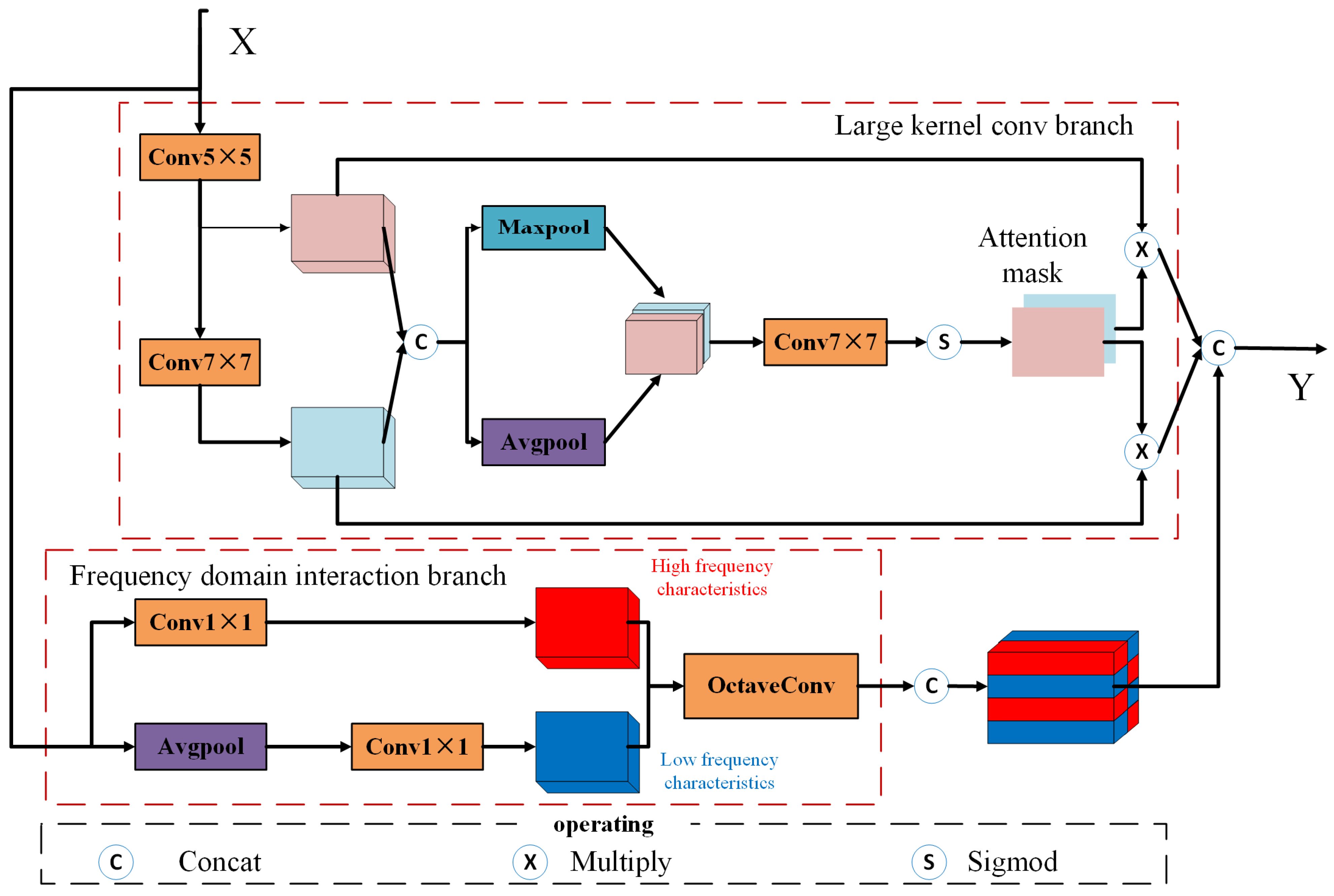
2.2.2. Multi-Scale Feature Enhancement Module (MFEM)
- (1)
- Large-kernel Convolution Branch
- denotes the output of the large-kernel branch,
- is the backbone feature map,
- and represent spatial attention masks generated via average pooling and max pooling, respectively.
- (2)
- Frequency-domain Interaction Branch
- is the output of the frequency-domain interaction branch.
- is the backbone feature map.
- (3)
- Decoder with Cascaded MFEMs
- denotes the decoder features at each scale.
2.2.3. Loss Function
- denotes the prediction generated from the final decoding layer,
- denotes the prediction obtained from deeper decoder features,
- denotes the actual prediction result.
- denotes height and width
- denotes pixel row and column numbers
2.3. Model Evaluation Indicators
2.4. Comparison Method
3. Results
3.1. Coastal Wetland Vegetation Classification Mapping and Analysis
3.2. Comparison and Analysis of Model Results
4. Discussion
4.1. Evaluation of the AFDFNet Based on Ablation Experiments and Parameter Analysis
4.2. Influence of Frequency-Domain Features on the Classification of Typical Coastal Vegetation
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Gedan, K.B.; Kirwan, M.L.; Wolanski, E.; Barbier, E.B.; Silliman, B.R. The Present and Future Role of Coastal Wetland Vegetation in Protecting Shorelines: Answering Recent Challenges to the Paradigm. Clim. Change 2011, 106, 7–29. [Google Scholar] [CrossRef]
- Van De Vijsel, R.C.; Van Belzen, J.; Bouma, T.J.; Van Der Wal, D.; Borsje, B.W.; Temmerman, S.; Cornacchia, L.; Gourgue, O.; Van De Koppel, J. Vegetation Controls on Channel Network Complexity in Coastal Wetlands. Nat. Commun. 2023, 14, 7158. [Google Scholar] [CrossRef] [PubMed]
- Feagin, R.A.; Lozada-Bernard, S.M.; Ravens, T.M.; Möller, I.; Yeager, K.M.; Baird, A.H. Does Vegetation Prevent Wave Erosion of Salt Marsh Edges? Proc. Natl. Acad. Sci. USA 2009, 106, 10109–10113. [Google Scholar] [CrossRef] [PubMed]
- Mcleod, E.; Chmura, G.L.; Bouillon, S.; Salm, R.; Björk, M.; Duarte, C.M.; Lovelock, C.E.; Schlesinger, W.H.; Silliman, B.R. A Blueprint for Blue Carbon: Toward an Improved Understanding of the Role of Vegetated Coastal Habitats in Sequestering CO2. Front. Ecol. Environ. 2011, 9, 552–560. [Google Scholar] [CrossRef]
- Duarte, C.M.; Middelburg, J.J.; Caraco, N. Major Role of Marine Vegetation on the Oceanic Carbon Cycle. Biogeosciences 2005, 2, 1–8. [Google Scholar] [CrossRef]
- Li, S.; Dragicevic, S.; Castro, F.A.; Sester, M.; Winter, S.; Coltekin, A.; Pettit, C.; Jiang, B.; Haworth, J.; Stein, A.; et al. Geospatial Big Data Handling Theory and Methods: A Review and Research Challenges. ISPRS J. Photogramm. Remote Sens. 2016, 115, 119–133. [Google Scholar] [CrossRef]
- Akinaga, T.; Saito, M.; Onodera, S.; Hyodo, F. UAV Visual Imagery-Based Evaluation of Blue Carbon as Seagrass Beds on a Tidal Flat Scale. Remote Sens. Appl. Soc. Environ. 2025, 37, 101430. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, Q.; Sample, J.E.; Hancke, K.; Salberg, A.-B. Coastal habitat mapping with UAV multi-sensor data: An experiment among DCNN-based approaches. ISPRS Ann. Photogramm. Remote Sens. Spat. Inf. Sci. 2022, 3, 439–445. [Google Scholar] [CrossRef]
- James, D.; Collin, A.; Mury, A.; Letard, M. Enhancing UAV coastal mapping using infrared pansharpening. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2021, 43, 257–264. [Google Scholar] [CrossRef]
- Morgan, G.R.; Stevenson, L.; Wang, C.; Avtar, R. UAS Remote Sensing for Coastal Wetland Vegetation Biomass Estimation: A Destructive vs. Non-Destructive Sampling Experiment. Remote Sens. 2025, 17, 2335. [Google Scholar] [CrossRef]
- Morgan, G.R.; Hodgson, M.E.; Wang, C.; Schill, S.R. Unmanned Aerial Remote Sensing of Coastal Vegetation: A Review. Ann. GIS 2022, 28, 385–399. [Google Scholar] [CrossRef]
- Zhang, Z.; Shen, X.; Yan, C.; Li, R.; Li, B. Unveiling Seaward Expansion Pattern in Mangrove Forests Using UAV Remote Sensing and Deep Learning. Ecol. Indic. 2025, 178, 114054. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, Y.; Ca, J.; Qin, Q.; Feng, Y.; Yan, J. Semantic Segmentation Network for Mangrove Tree Species Based on UAV Remote Sensing Images. Sci. Rep. 2024, 14, 29860. [Google Scholar] [CrossRef] [PubMed]
- Ke, L.; Lu, Y.; Tan, Q.; Zhao, Y.; Wang, Q. Precise Mapping of Coastal Wetlands Using Time-Series Remote Sensing Images and Deep Learning Model. Front. For. Glob. Change 2024, 7, 1409985. [Google Scholar] [CrossRef]
- Cruz, C.; McGuinness, K.; Perrin, P.M.; O’Connell, J.; Martin, J.R.; Connolly, J. Improving the Mapping of Coastal Invasive Species Using UAV Imagery and Deep Learning. Int. J. Remote Sens. 2023, 44, 5713–5735. [Google Scholar] [CrossRef]
- Yuan, S.; Liang, X.; Lin, T.; Chen, S.; Liu, R.; Wang, J.; Zhang, H.; Gong, P. A Comprehensive Review of Remote Sensing in Wetland Classification and Mapping. arXiv 2025, arXiv:2504.10842. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, L.; Zhao, T.; Chen, X.; Lin, S.; Wang, J.; Mi, J.; Liu, W. GWL_FCS30: A Global 30 m Wetland Map with a Fine Classification System Using Multi-Sourced and Time-Series Remote Sensing Imagery in 2020. Earth Syst. Sci. Data 2023, 15, 265–293. [Google Scholar] [CrossRef]
- Wang, X.; Xiao, X.; Zou, Z.; Hou, L.; Qin, Y.; Dong, J.; Doughty, R.B.; Chen, B.; Zhang, X.; Chen, Y.; et al. Mapping Coastal Wetlands of China Using Time Series Landsat Images in 2018 and Google Earth Engine. ISPRS J. Photogramm. Remote Sens. 2020, 163, 312–326. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, L.; Zhao, T.; Wang, J.; Liu, W.; Chen, X. Global Annual Wetland Dataset at 30 m with a Fine Classification System from 2000 to 2022. Sci. Data 2024, 11, 310. [Google Scholar] [CrossRef]
- Peng, K.; Jiang, W.; Hou, P.; Wu, Z.; Cui, T. Detailed Wetland-Type Classification Using Landsat-8 Time-Series Images: A Pixel- and Object-Based Algorithm with Knowledge (POK). GIScience Remote Sens. 2024, 61, 2293525. [Google Scholar] [CrossRef]
- Klemas, V. Remote Sensing of Coastal Wetland Biomass: An Overview. J. Coast. Res. 2013, 290, 1016–1028. [Google Scholar] [CrossRef]
- Doughty, C.L.; Cavanaugh, K.C. Mapping Coastal Wetland Biomass from High Resolution Unmanned Aerial Vehicle (UAV) Imagery. Remote Sens. 2019, 11, 540. [Google Scholar] [CrossRef]
- Du, Y.; Wang, J.; Liu, Z.; Yu, H.; Li, Z.; Cheng, H. Evaluation on Spaceborne Multispectral Images, Airborne Hyperspectral, and LiDAR Data for Extracting Spatial Distribution and Estimating Aboveground Biomass of Wetland Vegetation Suaeda salsa. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2019, 12, 200–209. [Google Scholar] [CrossRef]
- Cai, L.; Tang, D.; Levy, G.; Liu, D. Remote Sensing of the Impacts of Construction in Coastal Waters on Suspended Particulate Matter Concentration—the Case of the Yangtze River Delta, China. Int. J. Remote Sens. 2016, 37, 2132–2147. [Google Scholar] [CrossRef]
- Ouyang, Z.-T.; Gao, Y.; Xie, X.; Guo, H.-Q.; Zhang, T.-T.; Zhao, B. Spectral Discrimination of the Invasive Plant Spartina Alterniflora at Multiple Phenological Stages in a Saltmarsh Wetland. PLoS ONE 2013, 8, e67315. [Google Scholar] [CrossRef]
- Wang, R.; Su, Y.; Sun, X.; Wang, M.; Feng, M. Rapid and Automated Mapping Method of Spartina Alterniflora Combines Tidal Imagery and Phenological Characteristics. Environ. Monit. Assess. 2025, 197, 1136. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; IEEE: Las Vegas, NV, USA, 2016; pp. 770–778. [Google Scholar]
- Gao, N.; Du, X.; Yang, M.; Zhao, X.; Gao, E.; Yang, Y. Extraction of Suaeda Salsa from UAV Imagery Assisted by Adaptive Capture of Contextual Information. Remote Sens. 2025, 17, 2022. [Google Scholar] [CrossRef]
- Gao, F.; Fu, M.; Cao, J.; Dong, J.; Du, Q. Adaptive Frequency Enhancement Network for Remote Sensing Image Semantic Segmentation. IEEE Trans. Geosci. Remote Sens. 2025, 63, 5619415. [Google Scholar] [CrossRef]
- Chen, Y.; Fan, H.; Xu, B.; Yan, Z.; Kalantidis, Y.; Rohrbach, M.; Yan, S.; Feng, J. Drop an Octave: Reducing Spatial Redundancy in Convolutional Neural Networks with Octave Convolution. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), Seoul, Republic of Korea, 27 October–2 November 2019. [Google Scholar]
- Huang, Y.; Wang, J.; Wu, P.; Duan, Z.; Li, X.; Tang, J. Impacts of Spartina Alterniflora Invasion on Coastal Carbon Cycling Within a Native Phragmites Australis-Dominated Wetland. Agric. For. Meteorol. 2025, 363, 110405. [Google Scholar] [CrossRef]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, USA, 2016; ISBN 978-0-262-33737-3. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation; Springer international publishing: Cham, Switzerland, 2015. [Google Scholar]
- Lin, T.-Y.; Dollar, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature Pyramid Networks for Object Detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Chen, J.; Lu, Y.; Yu, Q.; Luo, X.; Adeli, E.; Wang, Y.; Lu, L.; Yuille, A.L.; Zhou, Y. TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation. arXiv 2021, arXiv:2102.04306. [Google Scholar] [CrossRef]
- Xie, E.; Wang, W.; Yu, Z.; Anandkumar, A.; Alvarez, J.M.; Luo, P. SegFormer: Simple and Efficient Design for Semantic Segmentation with Transformers. Adv. Neural Inf. Process. Syst. 2021, 34, 12077–12090. [Google Scholar]
- Zhou, L.; Zhang, C.; Wu, M. D-LinkNet: LinkNet with Pretrained Encoder and Dilated Convolution for High Resolution Satellite Imagery Road Extraction. In Proceedings of the 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), Lake City, UT, USA, 18–22 June 2018; IEEE: Salt Lake City, UT, USA, 2018; pp. 192–1924. [Google Scholar]
- Zheng, J.; Shao, A.; Yan, Y.; Wu, J.; Zhang, M. Remote Sensing Semantic Segmentation via Boundary Supervision-Aided Multiscale Channelwise Cross Attention Network. IEEE Trans. Geosci. Remote Sens. 2023, 61, 1–14. [Google Scholar] [CrossRef]
- Han, K.; Wang, Y.; Chen, H.; Chen, X.; Guo, J.; Liu, Z.; Tang, Y.; Xiao, A.; Xu, C.; Xu, Y.; et al. A Survey on Vision Transformer. IEEE Trans. Pattern Anal. Mach. Intell. 2023, 45, 87–110. [Google Scholar] [CrossRef]
- Minaee, S.; Boykov, Y.Y.; Porikli, F.; Plaza, A.J.; Kehtarnavaz, N.; Terzopoulos, D. Image Segmentation Using Deep Learning: A Survey. IEEE Trans. Pattern Anal. Mach. Intell. 2021, 44, 3523–3542. [Google Scholar] [CrossRef] [PubMed]
- Ke, Y.; Han, Y.; Cui, L.; Sun, P.; Min, Y.; Wang, Z.; Zhuo, Z.; Zhou, Q.; Yin, X.; Zhou, D. Suaeda Salsa Spectral Index for Suaeda Salsa Mapping and Fractional Cover Estimation in Intertidal Wetlands. ISPRS J. Photogramm. Remote Sens. 2024, 207, 104–121. [Google Scholar] [CrossRef]
- Harris, J.M.; Broussard, W.P.; Nelson, J.A. Evaluating Coastal Wetland Restoration Using Drones and High-Resolution Imagery. Estuaries Coasts 2024, 47, 1359–1375. [Google Scholar] [CrossRef]
- Yang, Y.; Yuan, G.; Li, J. SFFNet: A Wavelet-Based Spatial and Frequency Domain Fusion Network for Remote Sensing Segmentation. IEEE Trans. Geosci. Remote Sens. 2024, 62, 3000617. [Google Scholar] [CrossRef]
- Yang, M.; Qin, J.; Wang, X.; Gu, Y. Research on the Wetland Vegetation Classification Method Based on Cross-Satellite Hyperspectral Images. JMSE 2025, 13, 801. [Google Scholar] [CrossRef]
- Liu, G.; Liu, C.; Wu, X.; Li, Y.; Zhang, X.; Xu, J. Optimization of Remote-Sensing Image-Segmentation Decoder Based on Multi-Dilation and Large-Kernel Convolution. Remote Sens. 2024, 16, 2851. [Google Scholar] [CrossRef]
- Wang, X.; Shu, L.; Han, R.; Yang, F.; Gordon, T.; Wang, X.; Xu, H. A Survey of Farmland Boundary Extraction Technology Based on Remote Sensing Images. Electronics 2023, 12, 1156. [Google Scholar] [CrossRef]









| Formula | ||
|---|---|---|
| Accuracy | (8) | |
| Kappa | (9) | |
| () | (10) | |
| (11) | ||
| Recall | (12) | |
| Miou | (13) | |
| Accuracy | Kappa | Miou | Recall | |
|---|---|---|---|---|
| unet | 0.9032 | 0.7511 | 0.5422 | 0.6032 |
| AFDFNet | 0.9485 | 0.8274 | 0.6922 | 0.7371 |
| transunet | 0.9392 | 0.8566 | 0.6733 | 0.7066 |
| D-linknet | 0.8791 | 0.6894 | 0.4876 | 0.5431 |
| deeplabv3 | 0.8922 | 0.6334 | 0.61 | 0.6822 |
| MCCA | 0.9086 | 0.7689 | 0.6499 | 0.7147 |
| segformer | 0.8527 | 0.6367 | 0.5368 | 0.6007 |
| Accuracy | Kappa | mIoU | Recall | |
|---|---|---|---|---|
| baseline | 0.9073 | 0.8578 | 0.6092 | 0.6322 |
| Baseline + a | 0.9351 | 0.7963 | 0.6331 | 0.7066 |
| Baseline + b | 0.9366 | 0.7752 | 0.6564 | 0.6891 |
| Baseline + a + b | 0.9431 | 0.8522 | 0.6884 | 0.7268 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Gao, N.; Du, X.; Xu, P.; Gao, E.; Yang, Y. High-Resolution Mapping Coastal Wetland Vegetation Using Frequency-Augmented Deep Learning Method. Remote Sens. 2026, 18, 247. https://doi.org/10.3390/rs18020247
Gao N, Du X, Xu P, Gao E, Yang Y. High-Resolution Mapping Coastal Wetland Vegetation Using Frequency-Augmented Deep Learning Method. Remote Sensing. 2026; 18(2):247. https://doi.org/10.3390/rs18020247
Chicago/Turabian StyleGao, Ning, Xinyuan Du, Peng Xu, Erding Gao, and Yixin Yang. 2026. "High-Resolution Mapping Coastal Wetland Vegetation Using Frequency-Augmented Deep Learning Method" Remote Sensing 18, no. 2: 247. https://doi.org/10.3390/rs18020247
APA StyleGao, N., Du, X., Xu, P., Gao, E., & Yang, Y. (2026). High-Resolution Mapping Coastal Wetland Vegetation Using Frequency-Augmented Deep Learning Method. Remote Sensing, 18(2), 247. https://doi.org/10.3390/rs18020247





