Figure 1.
Global view of Mars showing enlarged and approximate locations of the target regions.
Figure 1.
Global view of Mars showing enlarged and approximate locations of the target regions.
Figure 2.
(a) Gypsum spectrum highlighting the 1750 nm absorption feature. The different coloured lines indicate regions where CRISM measures the spectrum in consistent 6.55 nm increments, with breaks marking interruptions in the regular sampling pattern. (b) Zoomed-in view showing the interpolation anchors used to identify the absorption feature. (c) BD1750_2 values for 31 minerals, with only gypsum and alunite showing strong responses, demonstrating the diagnostic role of this spectral product in mineral detection.
Figure 2.
(a) Gypsum spectrum highlighting the 1750 nm absorption feature. The different coloured lines indicate regions where CRISM measures the spectrum in consistent 6.55 nm increments, with breaks marking interruptions in the regular sampling pattern. (b) Zoomed-in view showing the interpolation anchors used to identify the absorption feature. (c) BD1750_2 values for 31 minerals, with only gypsum and alunite showing strong responses, demonstrating the diagnostic role of this spectral product in mineral detection.
Figure 3.
(a) The Best Matching Unit (BMU, red) and its radius of influence (Image A). During training, the weight vectors of neighbouring neurons are updated, with the adjustment strength decreasing with distance from the BMU. Neuron coordinates remain fixed, but their weight vectors move closer together (Image B). (b) The topological arrangement of neurons in a 50 × 50 self-organising map. Together, these panels visualise SOM training and neuron organisation.
Figure 3.
(a) The Best Matching Unit (BMU, red) and its radius of influence (Image A). During training, the weight vectors of neighbouring neurons are updated, with the adjustment strength decreasing with distance from the BMU. Neuron coordinates remain fixed, but their weight vectors move closer together (Image B). (b) The topological arrangement of neurons in a 50 × 50 self-organising map. Together, these panels visualise SOM training and neuron organisation.
Figure 4.
Pixel coordinate system used in this study.
Figure 4.
Pixel coordinate system used in this study.
Figure 5.
This heat map illustrates how summary products serve as input features. Rather than displaying all 29 products for each pixel, it shows a subset of recalculated summary products for selected minerals from the MRO CRISM Type Spectra Library, highlighting spectral variation across mineral types. Each horizontal row can be thought of as an individual pixel from a TER-3 image file.
Figure 5.
This heat map illustrates how summary products serve as input features. Rather than displaying all 29 products for each pixel, it shows a subset of recalculated summary products for selected minerals from the MRO CRISM Type Spectra Library, highlighting spectral variation across mineral types. Each horizontal row can be thought of as an individual pixel from a TER-3 image file.
Figure 7.
Our study uses image data from Viviano et al. [
14] as a source of ground truth for comparison with outputs derived from the SOM model. For ease of comparison, we show a snapshot of the referenced image in the top-right corner of the corresponding panel and provide a larger version in
Appendix A. We highlight the limitations of these visual comparisons in
Section 3.1. Both images are rotated 180 degrees to align with Image D from
Figure A3 and have a resolution of 390 × 640 pixels, with the X-axis oriented vertically to match image rows. (
a) Unprojected map of FRT0000634B with an overlay based on Image D of
Figure A3. Faint pale-pink areas indicate illite, with the clustering box reducing the SOM model’s ability to resolve thin strips of this mineral. (
b) Pixel locations of illite identified by the top five neurons most correlated with the processed MRO CRISM Type Spectra Library. Thin white strips demonstrate the SOM model’s capacity to resolve subtle spectral differences within regions of spectrally similar pixels, as indicated by the surrounding pink shading in (
a).
Figure 7.
Our study uses image data from Viviano et al. [
14] as a source of ground truth for comparison with outputs derived from the SOM model. For ease of comparison, we show a snapshot of the referenced image in the top-right corner of the corresponding panel and provide a larger version in
Appendix A. We highlight the limitations of these visual comparisons in
Section 3.1. Both images are rotated 180 degrees to align with Image D from
Figure A3 and have a resolution of 390 × 640 pixels, with the X-axis oriented vertically to match image rows. (
a) Unprojected map of FRT0000634B with an overlay based on Image D of
Figure A3. Faint pale-pink areas indicate illite, with the clustering box reducing the SOM model’s ability to resolve thin strips of this mineral. (
b) Pixel locations of illite identified by the top five neurons most correlated with the processed MRO CRISM Type Spectra Library. Thin white strips demonstrate the SOM model’s capacity to resolve subtle spectral differences within regions of spectrally similar pixels, as indicated by the surrounding pink shading in (
a).
![Remotesensing 17 03578 g007 Remotesensing 17 03578 g007]()
Figure 8.
Reference to clusters shown in
Figure 7. The SOM grid contains 2500 neurons. In this rotated 3D orientation, neuron [0, 0] is at the left-hand corner, [50, 50] at the right-hand corner, [50, 0] at the top apex, and [0, 50] at the bottom apex. (
a) Neuron weight values for
, with both clusters showing a distinct spike—their separation indicates additional high-dimensional differences. (
b) Neuron weight values for
, where cluster 0 shows elevated absorption, while clusters 1 and 2 are more subdued and not directly visible from this view, leading the SOM to topologically separate them.
Figure 8.
Reference to clusters shown in
Figure 7. The SOM grid contains 2500 neurons. In this rotated 3D orientation, neuron [0, 0] is at the left-hand corner, [50, 50] at the right-hand corner, [50, 0] at the top apex, and [0, 50] at the bottom apex. (
a) Neuron weight values for
, with both clusters showing a distinct spike—their separation indicates additional high-dimensional differences. (
b) Neuron weight values for
, where cluster 0 shows elevated absorption, while clusters 1 and 2 are more subdued and not directly visible from this view, leading the SOM to topologically separate them.
Figure 9.
SOM-driven semi-automated selection of neurons 2412 and 2461–2464, representing the SOM-identified topological cluster containing a pixel verified as illite. White pixels indicate individual pixels clustered by these neurons. The snippet shows the SOM neuron grid with
absorption, highlighting a separation between neuron 2454 and the neighbouring cluster of neurons (2461–2464) with more elevated
absorption. The snippet is positioned over an otherwise uninformative area of the image as per previous images. This image is rotated 180 degrees to align with Image D from
Figure A3 and has a resolution of 390 × 640 pixels, with the x-axis oriented vertically to align with the image rows.
Figure 9.
SOM-driven semi-automated selection of neurons 2412 and 2461–2464, representing the SOM-identified topological cluster containing a pixel verified as illite. White pixels indicate individual pixels clustered by these neurons. The snippet shows the SOM neuron grid with
absorption, highlighting a separation between neuron 2454 and the neighbouring cluster of neurons (2461–2464) with more elevated
absorption. The snippet is positioned over an otherwise uninformative area of the image as per previous images. This image is rotated 180 degrees to align with Image D from
Figure A3 and has a resolution of 390 × 640 pixels, with the x-axis oriented vertically to align with the image rows.
Figure 10.
We use image data from Viviano et al. [
14] as a source of ground truth for comparison with outputs derived from the SOM model. For ease of comparison, a snapshot of the referenced image is shown in the top-right corner of the corresponding panel, with a larger version provided in
Appendix A. Readers are reminded of the limitations of these visual comparisons, as discussed in
Section 3.1. (
a) Overlay based on Image H of
Figure A2, showing cyan areas of low-Ca pyroxene identified by the top five correlated neurons. The maximum pixel correlation in Cluster 0 is 0.87. (
b) Overlay using the top fifty correlated neurons, where white pixels indicate those clustered by the neurons and underlying cyan areas indicate low-Ca pyroxene. Correlation values range from 0.81 to 0.87, with the latter in Cluster 2. Both images are unprojected, rotated 180 degrees to align with Image H from
Figure A2, and have a resolution of 480 × 640 pixels, with the x-axis oriented vertically to match image rows.
Figure 10.
We use image data from Viviano et al. [
14] as a source of ground truth for comparison with outputs derived from the SOM model. For ease of comparison, a snapshot of the referenced image is shown in the top-right corner of the corresponding panel, with a larger version provided in
Appendix A. Readers are reminded of the limitations of these visual comparisons, as discussed in
Section 3.1. (
a) Overlay based on Image H of
Figure A2, showing cyan areas of low-Ca pyroxene identified by the top five correlated neurons. The maximum pixel correlation in Cluster 0 is 0.87. (
b) Overlay using the top fifty correlated neurons, where white pixels indicate those clustered by the neurons and underlying cyan areas indicate low-Ca pyroxene. Correlation values range from 0.81 to 0.87, with the latter in Cluster 2. Both images are unprojected, rotated 180 degrees to align with Image H from
Figure A2, and have a resolution of 480 × 640 pixels, with the x-axis oriented vertically to match image rows.
![Remotesensing 17 03578 g010 Remotesensing 17 03578 g010]()
Figure 11.
Heat map of SOM neurons showing correlation with low-Ca pyroxene in region FRT000064D9 based on comparison with the MRO CRISM Type Spectra Library. Each pixel in this image corresponds to a neuron in the SOM model, starting with neuron 0 at the top-left corner [0, 0] and ending with neuron 2499 at the bottom-right corner [50, 50], comprising a total of 2500 neurons.
Figure 11.
Heat map of SOM neurons showing correlation with low-Ca pyroxene in region FRT000064D9 based on comparison with the MRO CRISM Type Spectra Library. Each pixel in this image corresponds to a neuron in the SOM model, starting with neuron 0 at the top-left corner [0, 0] and ending with neuron 2499 at the bottom-right corner [50, 50], comprising a total of 2500 neurons.
Figure 12.
(a) Number of times each neuron was the BMU, with lower hit counts concentrated in the upper-left quadrant. (b) Heat map of SOM neurons showing correlation with Mg-carbonate based on comparison with the MRO CRISM Type Spectra Library. Neurons correlated with Mg-carbonate cluster in the top-left quadrant, whereas Fe-olivine would otherwise correlate to neurons in the top-right quadrant. Each pixel in these images corresponds to a neuron in the SOM model, numbered from neuron 0 at [0, 0] in the top left to neuron 2499 at [50, 50] in the bottom right, for a total of 2500 neurons.
Figure 12.
(a) Number of times each neuron was the BMU, with lower hit counts concentrated in the upper-left quadrant. (b) Heat map of SOM neurons showing correlation with Mg-carbonate based on comparison with the MRO CRISM Type Spectra Library. Neurons correlated with Mg-carbonate cluster in the top-left quadrant, whereas Fe-olivine would otherwise correlate to neurons in the top-right quadrant. Each pixel in these images corresponds to a neuron in the SOM model, numbered from neuron 0 at [0, 0] in the top left to neuron 2499 at [50, 50] in the bottom right, for a total of 2500 neurons.
Figure 13.
Unprojected map view of FRT00003E12 using the recommended CR2 overlay, highlighting potentially Mg-carbonate-rich areas. Unlike other images, no individual pixels are marked in this overlay. The yellow regions are a direct result of the overlay and represent isolated and distinct mineral clustering in this area. The SOM-delineated regions were generated by selecting the top 15 neurons—those with a correlation with the reference spectra exceeding 0.8, i.e., the highest correlating neurons located in the top-left corner of the neuron heat map.
Figure 13.
Unprojected map view of FRT00003E12 using the recommended CR2 overlay, highlighting potentially Mg-carbonate-rich areas. Unlike other images, no individual pixels are marked in this overlay. The yellow regions are a direct result of the overlay and represent isolated and distinct mineral clustering in this area. The SOM-delineated regions were generated by selecting the top 15 neurons—those with a correlation with the reference spectra exceeding 0.8, i.e., the highest correlating neurons located in the top-left corner of the neuron heat map.
Figure 14.
Mineral mapping, correlation, and hit rate for each neuron in target region FRT0000634B. Each pixel in these images corresponds to a neuron in the SOM model, starting with neuron 0 at the top-left corner [0, 0] and ending with neuron 2499 at the bottom-right corner [50, 50], comprising a total of 2500 neurons. (a) The number of minerals mapped to each neuron. Neurons with multiple mapped minerals suggest a high degree of feature correlation among their mineral groups. (b) Neuron correlation with Al-smectite. (c) The number of times each neuron was identified as the BMU. The SOM model appears to cluster uninformative neurons near the centre of the map, where we observe the highest hit counts. (d) Neuron correlation with talc.
Figure 14.
Mineral mapping, correlation, and hit rate for each neuron in target region FRT0000634B. Each pixel in these images corresponds to a neuron in the SOM model, starting with neuron 0 at the top-left corner [0, 0] and ending with neuron 2499 at the bottom-right corner [50, 50], comprising a total of 2500 neurons. (a) The number of minerals mapped to each neuron. Neurons with multiple mapped minerals suggest a high degree of feature correlation among their mineral groups. (b) Neuron correlation with Al-smectite. (c) The number of times each neuron was identified as the BMU. The SOM model appears to cluster uninformative neurons near the centre of the map, where we observe the highest hit counts. (d) Neuron correlation with talc.
Figure 17.
Orientation note: In panels (b–d), the X-axis corresponds to the vertical axis and the Y-axis to the horizontal axis in panel (a). (a) SOM U-Matrix for target region FRT00007375 with annotated neurons. Each pixel represents a neuron in the 50 × 50 grid, numbered from 0 at the top left [0, 0] to 2499 at the bottom right [50, 50]. (b) R3920 reflectance across the SOM grid. (c) BD2230 reflectance, showing spikes for both neurons 276 and 607. (d) BD2265 reflectance, where only neuron 276 exhibits a strong absorption feature.
Figure 17.
Orientation note: In panels (b–d), the X-axis corresponds to the vertical axis and the Y-axis to the horizontal axis in panel (a). (a) SOM U-Matrix for target region FRT00007375 with annotated neurons. Each pixel represents a neuron in the 50 × 50 grid, numbered from 0 at the top left [0, 0] to 2499 at the bottom right [50, 50]. (b) R3920 reflectance across the SOM grid. (c) BD2230 reflectance, showing spikes for both neurons 276 and 607. (d) BD2265 reflectance, where only neuron 276 exhibits a strong absorption feature.
Figure 18.
SOM outputs for target region FRT00007375. (a) Neuron hit rate, showing how often each neuron was identified as the BMU, with many neurons allocated to unique spectra in the bottom-left corner. (b) Rotated 3D view of the same SOM grid to emphasise BD1300 responses; the dense cluster corresponds to the bottom-left corner of panel (a). (c) Neuron correlation with Fe-olivine, aligning closely with the low-hit-rate cluster in panel (a). (d) Neuron correlation with margarite, showing a more diffuse correlation than for Fe-olivine. Panels (a,c,d) show the SOM 50 × 50 neuron grid.
Figure 18.
SOM outputs for target region FRT00007375. (a) Neuron hit rate, showing how often each neuron was identified as the BMU, with many neurons allocated to unique spectra in the bottom-left corner. (b) Rotated 3D view of the same SOM grid to emphasise BD1300 responses; the dense cluster corresponds to the bottom-left corner of panel (a). (c) Neuron correlation with Fe-olivine, aligning closely with the low-hit-rate cluster in panel (a). (d) Neuron correlation with margarite, showing a more diffuse correlation than for Fe-olivine. Panels (a,c,d) show the SOM 50 × 50 neuron grid.
Figure 19.
Mineral categories associated with the neurons located at each corner of the SOM model for target region FRT00007375. Unlike other corners, sulphate-like spectra are lacking, whereas Fe-silicate and phyllosilicate spectra are more abundant.
Figure 19.
Mineral categories associated with the neurons located at each corner of the SOM model for target region FRT00007375. Unlike other corners, sulphate-like spectra are lacking, whereas Fe-silicate and phyllosilicate spectra are more abundant.
Figure 21.
Unprojected map view of FRT00007375 showing the MAF overlay, intended to highlight Fe-silicate regions (
Figure A1). The overlay provides limited guidance compared to the SOM, which clearly identifies concentrated clusters of spectra potentially corresponding to Fe-oxides and mafic silicates (
Figure 20). Image resolution is 390 × 640 pixels, with the x-axis oriented vertically to match image rows.
Figure 21.
Unprojected map view of FRT00007375 showing the MAF overlay, intended to highlight Fe-silicate regions (
Figure A1). The overlay provides limited guidance compared to the SOM, which clearly identifies concentrated clusters of spectra potentially corresponding to Fe-oxides and mafic silicates (
Figure 20). Image resolution is 390 × 640 pixels, with the x-axis oriented vertically to match image rows.
Figure 23.
Performance trends in SOM clustering with larger SOM configurations. (a) Normalised MWD, calculated by dividing MWD by the number of neurons in each SOM configuration. (b) Gradient of the normalised MWD for SOM series #1 and SOM series #2 across larger SOM configurations, indicating that beyond a 30 × 30 SOM size, the incremental improvement in MWD becomes minimal.
Figure 23.
Performance trends in SOM clustering with larger SOM configurations. (a) Normalised MWD, calculated by dividing MWD by the number of neurons in each SOM configuration. (b) Gradient of the normalised MWD for SOM series #1 and SOM series #2 across larger SOM configurations, indicating that beyond a 30 × 30 SOM size, the incremental improvement in MWD becomes minimal.
Figure 24.
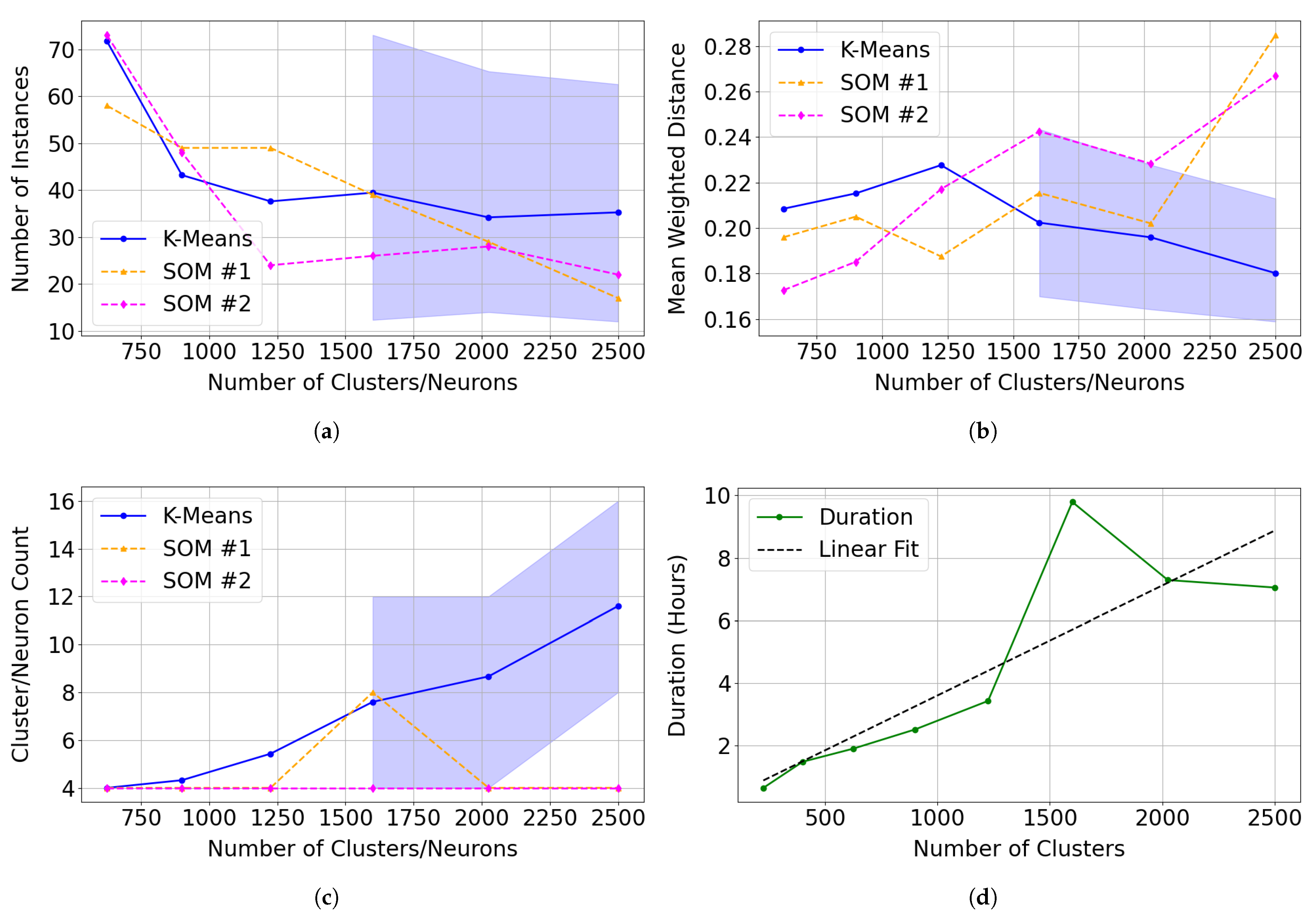
Comparison of clustering performance between 30 independent k-means trials and two SOM model series (SOM #1 and SOM #2) across varying cluster/neuron counts. Mean performance is shown as solid lines. For k-means, the shaded region represents the 5th and 95th percentiles across the 30 trials. Illite was not reliably clustered by k-means at fewer than 1600 clusters. We calculate error metrics using only clusters or neurons with correlation with the summary products derived from the illite spectrum in the MRO CRISM Type Spectra Library above 0.70. (a) Total number of instances assigned to clusters (K-Means) or neurons (SOM) identified as representing illite. (b) Mean weighted distance, indicating the compactness and cohesion of clusters (K-Means) or neurons (SOM). (c) Total number of clusters (K-Means) or neurons (SOM) matching illite across tested model sizes. (d) The execution time for SOM #1 across tested model sizes.
Figure 24.
Comparison of clustering performance between 30 independent k-means trials and two SOM model series (SOM #1 and SOM #2) across varying cluster/neuron counts. Mean performance is shown as solid lines. For k-means, the shaded region represents the 5th and 95th percentiles across the 30 trials. Illite was not reliably clustered by k-means at fewer than 1600 clusters. We calculate error metrics using only clusters or neurons with correlation with the summary products derived from the illite spectrum in the MRO CRISM Type Spectra Library above 0.70. (a) Total number of instances assigned to clusters (K-Means) or neurons (SOM) identified as representing illite. (b) Mean weighted distance, indicating the compactness and cohesion of clusters (K-Means) or neurons (SOM). (c) Total number of clusters (K-Means) or neurons (SOM) matching illite across tested model sizes. (d) The execution time for SOM #1 across tested model sizes.
Table 1.
Target regions and, where applicable, the original [X, Y] pixel coordinates of mineral locations shown in remote sensing and overlay images used in this research. These coordinates correspond to positions prior to any image rotation. For more details, refer to
Section 3.1. * The presence of Fe-olivine, Fe-oxides, or other mafic silicate minerals remains speculative and is not confirmed.
Table 1.
Target regions and, where applicable, the original [X, Y] pixel coordinates of mineral locations shown in remote sensing and overlay images used in this research. These coordinates correspond to positions prior to any image rotation. For more details, refer to
Section 3.1. * The presence of Fe-olivine, Fe-oxides, or other mafic silicate minerals remains speculative and is not confirmed.
| Mineral | Figure | Target Region | Original X | Original Y | Rotated |
|---|
| Illite | Figure 7a | FRT0000634B | NA | NA | YES |
| Illite | Figure 7b | FRT0000634B | NA | NA | YES |
| Low-Ca Pyroxene | Figure 10a | FRT000064D9 | 174 | 528 | YES |
| Low-Ca Pyroxene | Figure 10b | FRT000064D9 | 174 | 528 | YES |
| Mg-carbonate | Figure 13 | FRT00003E12 | 103 | 136 | NO |
| Fe-olivine * | Figure 20a | FRT00007375 | NA | NA | NO |
Table 2.
Left table: Numbers ordered in a 5 × 5 grid from 1 to 25. Right table: Total Euclidean distance of each number from every other number.
Table 2.
Left table: Numbers ordered in a 5 × 5 grid from 1 to 25. Right table: Total Euclidean distance of each number from every other number.
| Left | Right |
|---|
| 1 | 2 | 3 | 4 | 5 | 300 | 277 | 256 | 237 | 220 |
| 6 | 7 | 8 | 9 | 10 | 205 | 192 | 181 | 172 | 165 |
| 11 | 12 | 13 | 14 | 15 | 160 | 157 | 156 | 157 | 160 |
| 16 | 17 | 18 | 19 | 20 | 165 | 172 | 181 | 192 | 205 |
| 21 | 22 | 23 | 24 | 25 | 220 | 237 | 256 | 277 | 300 |
Table 3.
Left table: Randomly initialised numbers from 1 to 25. Right table: SOM ordering after 100,000 training iterations.
Table 3.
Left table: Randomly initialised numbers from 1 to 25. Right table: SOM ordering after 100,000 training iterations.
| Left | Right |
|---|
| 19 | 7 | 14 | 18 | 22 | 24.56 | 21.44 | 18 | 16.3 | 15 |
| 15 | 24 | 5 | 20 | 17 | 23 | 19.41 | 17.56 | 16.53 | 14.52 |
| 8 | 21 | 11 | 9 | 13 | 17 | 16 | 14 | 12.68 | 7.38 |
| 25 | 10 | 23 | 1 | 6 | 13 | 11 | 10 | 6 | 3.41 |
| 4 | 12 | 3 | 2 | 16 | 12 | 10.62 | 9 | 5 | 1.33 |
Table 4.
Left table: Randomly initialised numbers from 1 to 25. Right table: SOM ordering after 100,000 training iterations with training instances biased toward 20 (drawn with 75% probability). Large-scale topological patterns are preserved, but neuron convergence toward 20 dilutes the ability to observe lower-order topological patterns.
Table 4.
Left table: Randomly initialised numbers from 1 to 25. Right table: SOM ordering after 100,000 training iterations with training instances biased toward 20 (drawn with 75% probability). Large-scale topological patterns are preserved, but neuron convergence toward 20 dilutes the ability to observe lower-order topological patterns.
| Left | Right |
|---|
| 12 | 5 | 1 | 7 | 20 | 13 | 11 | 8 | 3.45 | 1.66 |
| 25 | 13 | 11 | 19 | 16 | 15.15 | 13.87 | 7 | 6 | 5 |
| 4 | 24 | 2 | 3 | 15 | 19 | 20 | 15 | 9 | 10 |
| 10 | 18 | 22 | 14 | 8 | 22.56 | 20.08 | 19.82 | 16 | 12 |
| 21 | 23 | 9 | 17 | 6 | 24.45 | 21 | 18 | 17 | 14 |
Table 5.
SOM model execution time (hours and minutes) by number of neurons.
Table 5.
SOM model execution time (hours and minutes) by number of neurons.
| Number of Neurons | Execution Time (Hours and Minutes) |
|---|
| 225 | 00:38 |
| 400 | 01:28 |
| 625 | 01:54 |
| 900 | 02:31 |
| 1225 | 03:25 |
| 1600 | 09:47 |
| 2025 | 07:17 |
| 2500 | 07:03 |
Table 6.
Comparison of normalised mean weighted distance (MWD) and its gradient for SOM series #1 across different neuron counts.
Table 6.
Comparison of normalised mean weighted distance (MWD) and its gradient for SOM series #1 across different neuron counts.
| Neurons | MWD | Norm. MWD | Gradient |
|---|
| 225 | 0.3269 | 1.453 × 10−3 | × 10−6 |
| 400 | 0.3150 | 7.874 × 10−4 | × 10−6 |
| 625 | 0.2916 | 4.665 × 10−4 | × 10−6 |
| 900 | 0.2719 | 3.021 × 10−4 | × 10−7 |
| 1225 | 0.2566 | 2.095 × 10−4 | × 10−7 |
| 1600 | 0.2480 | 1.550 × 10−4 | × 10−7 |
| 2025 | 0.2356 | 1.163 × 10−4 | × 10−8 |
| 2500 | 0.2017 | 8.069 × 10−5 | × 10−8 |
Table 7.
Comparison of normalised mean weighted distance (MWD) and its gradient for SOM series #2 across different neuron counts.
Table 7.
Comparison of normalised mean weighted distance (MWD) and its gradient for SOM series #2 across different neuron counts.
| Neurons | MWD | Norm. MWD | Gradient |
|---|
| 225 | 0.3074 | 1.366 × 10−3 | × 10−6 |
| 400 | 0.2971 | 7.427 × 10−4 | × 10−6 |
| 625 | 0.2763 | 4.421 × 10−4 | × 10−7 |
| 900 | 0.2653 | 2.947 × 10−4 | × 10−7 |
| 1225 | 0.2511 | 2.050 × 10−4 | × 10−7 |
| 1600 | 0.2442 | 1.527 × 10−4 | × 10−7 |
| 2025 | 0.2253 | 1.112 × 10−4 | × 10−8 |
| 2500 | 0.2298 | 9.192 × 10−5 | × 10−8 |