Unsupervised Diffusion and Volume Maximization-Based Clustering of Hyperspectral Images
Abstract
1. Introduction
2. Background
2.1. Background on Unsupervised HSI Clustering
2.2. Background on Spectral Graph Theory
Background on Diffusion Geometry
2.3. Background on Spectral Unmixing
2.3.1. Background on the HySime Algorithm
2.3.2. Background on the AVMAX Algorithm
3. Diffusion and Volume Maximization-Based Image Clustering
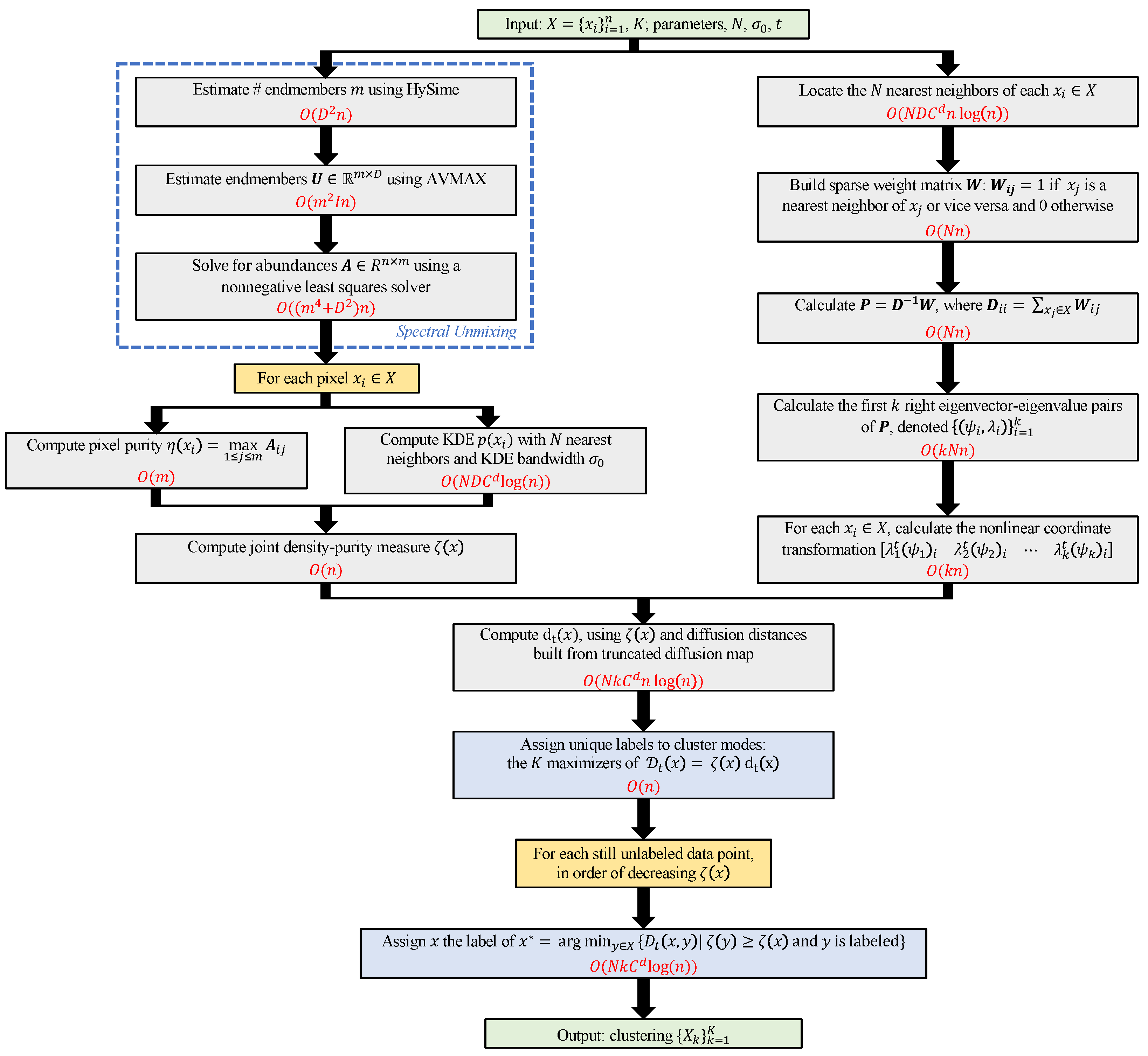
| Algorithm 1: Diffusion and Volume maximization-based Image Clustering |
Input: X (HSI), N (# nearest neighbors), (KDE scale), t (diffusion time), K (# clusters) Output: (clustering)
|
3.1. Computational Complexity
3.2. Comparison with Learning by Unsupervised Nonlinear Diffusion
4. Experiments and Discussion
4.1. Analysis of Benchmark HSI Datasets
- 1.
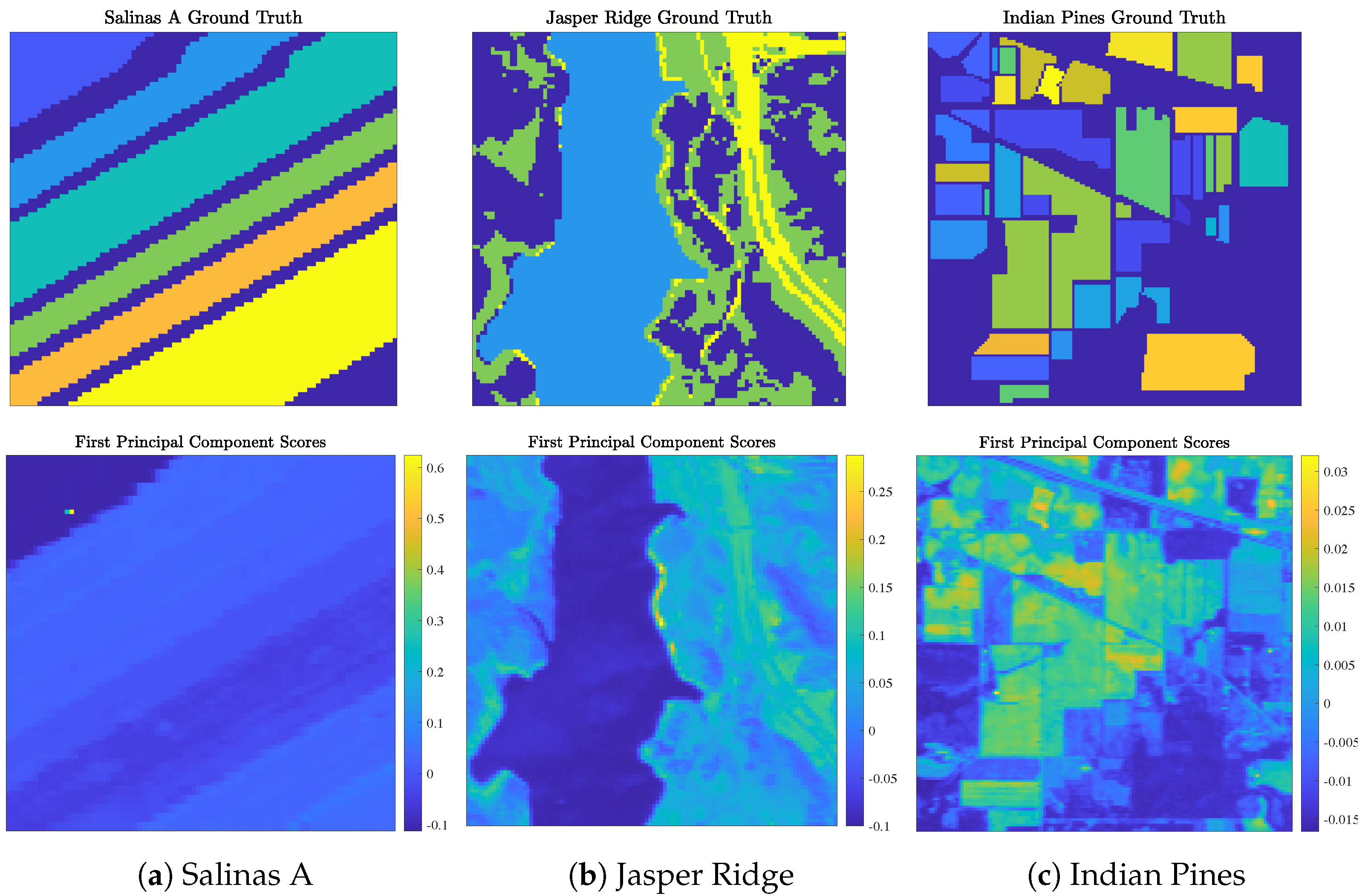
- Salinas A (Figure 5a) was recorded by the Airborne Visible/Infrared Imaging Spectrometer (AVIRIS) sensor over farmland in Salinas Valley, California, USA, in 1998 at a spatial resolution of 1.3 m. Spectral signatures, ranging in recorded wavelength from 380 nm to 2500 nm across 224 spectral bands, were recorded across pixels (). Gaussian noise (with mean 0 and standard deviation ) was added to each pixel to differentiate two pixels with identical spectral signatures. The Salinas A scene contains ground truth classes corresponding to crop types.
- 2.
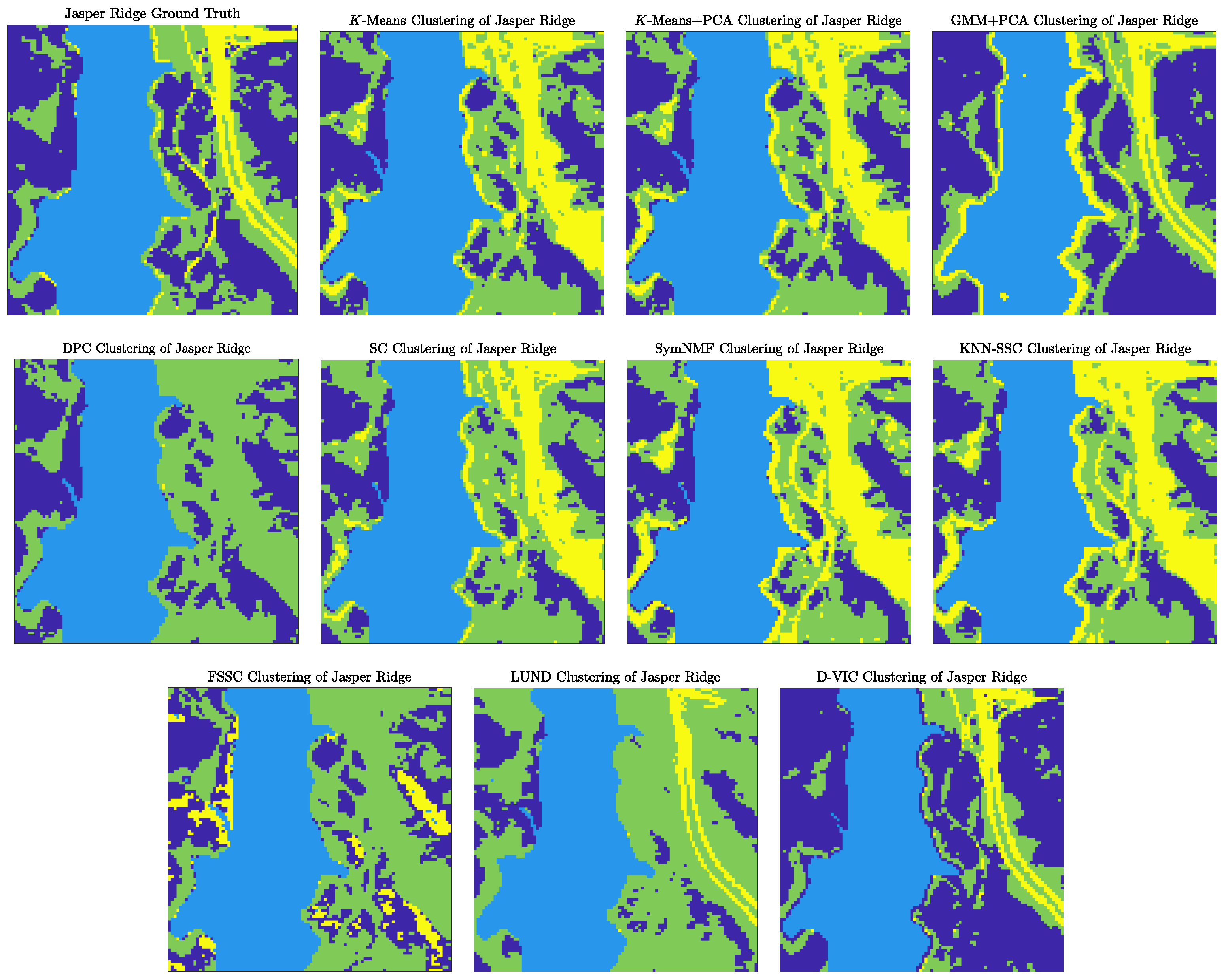
- Jasper Ridge (Figure 5b) was recorded by the AVIRIS sensor over the Jasper Ridge Biological Preserve, California, USA, in 1989 at a spatial resolution of 5 m. Spectral signatures, ranging in recorded wavelength from 380 nm to 2500 nm across 224 spectral bands, were recorded across spatial dimensions of pixels ( 10,000). The Jasper Ridge scene contains ground truth endmembers: road, soil, water, and trees. Ground truth labels were recovered by selecting the material of the highest ground truth abundance for each pixel.
- 3.
- Indian Pines (Figure 5c) was recorded by the AVIRIS sensor over farmland in northwest Indiana, USA, in 1992 at a low spatial resolution of 20 m. Spectral signatures, ranging in recorded wavelength from 400 nm to 2500 nm across 224 spectral bands, were recorded across spatial dimensions of pixels ( 21,025). The Indian Pines scene contains ground truth classes (e.g., crop types and manufactured structures) and many unlabeled pixels.
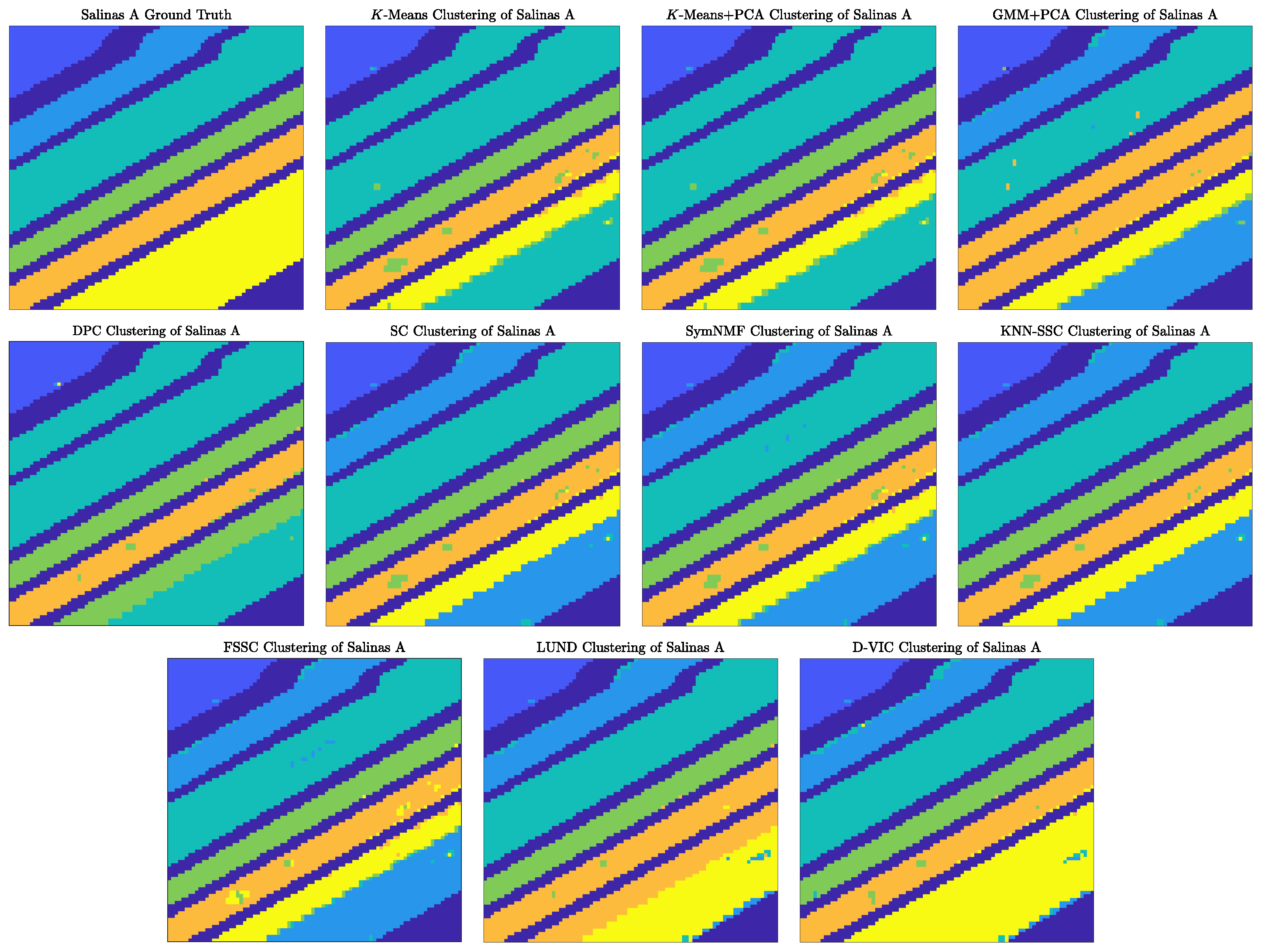
4.1.1. Discussion of Benchmark HSI Experiments
4.1.2. Runtime Analysis
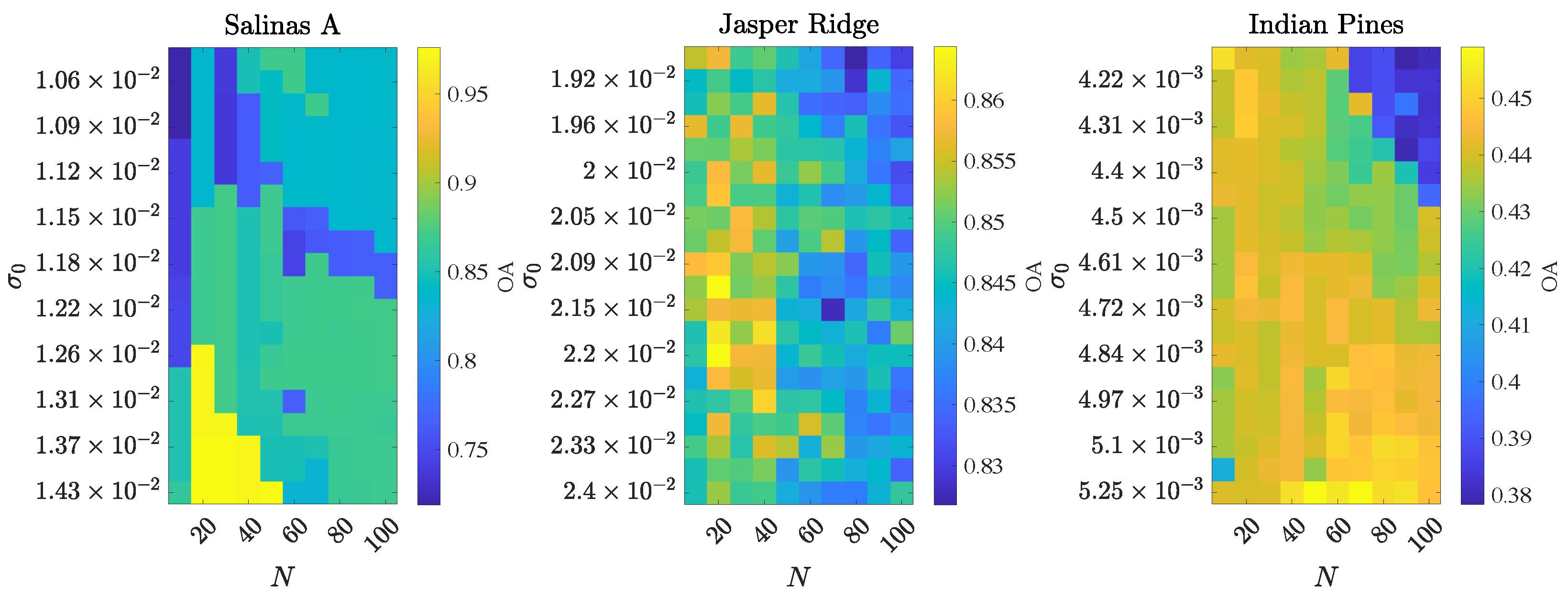
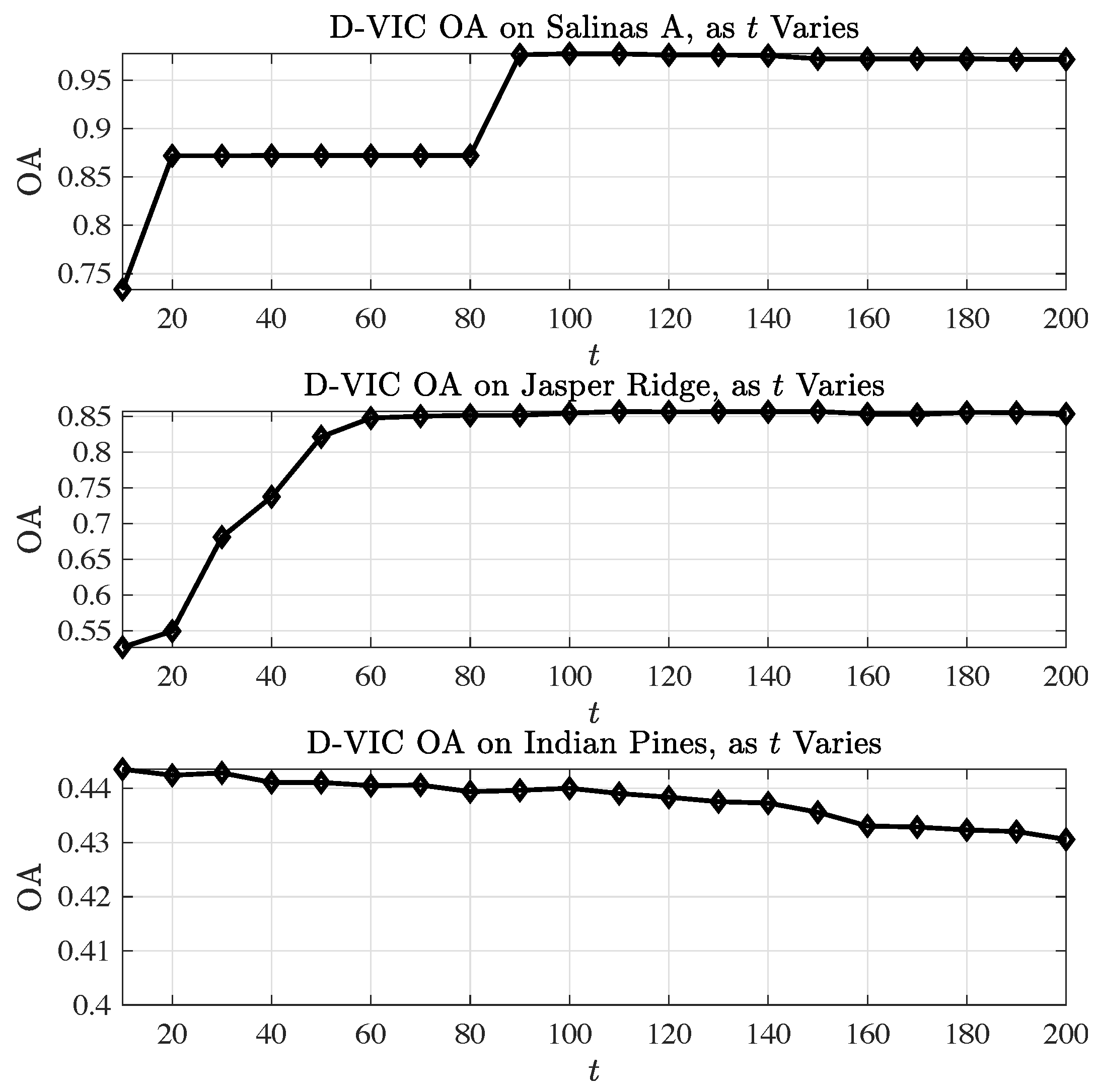
4.1.3. Robustness to Hyperparameter Selection
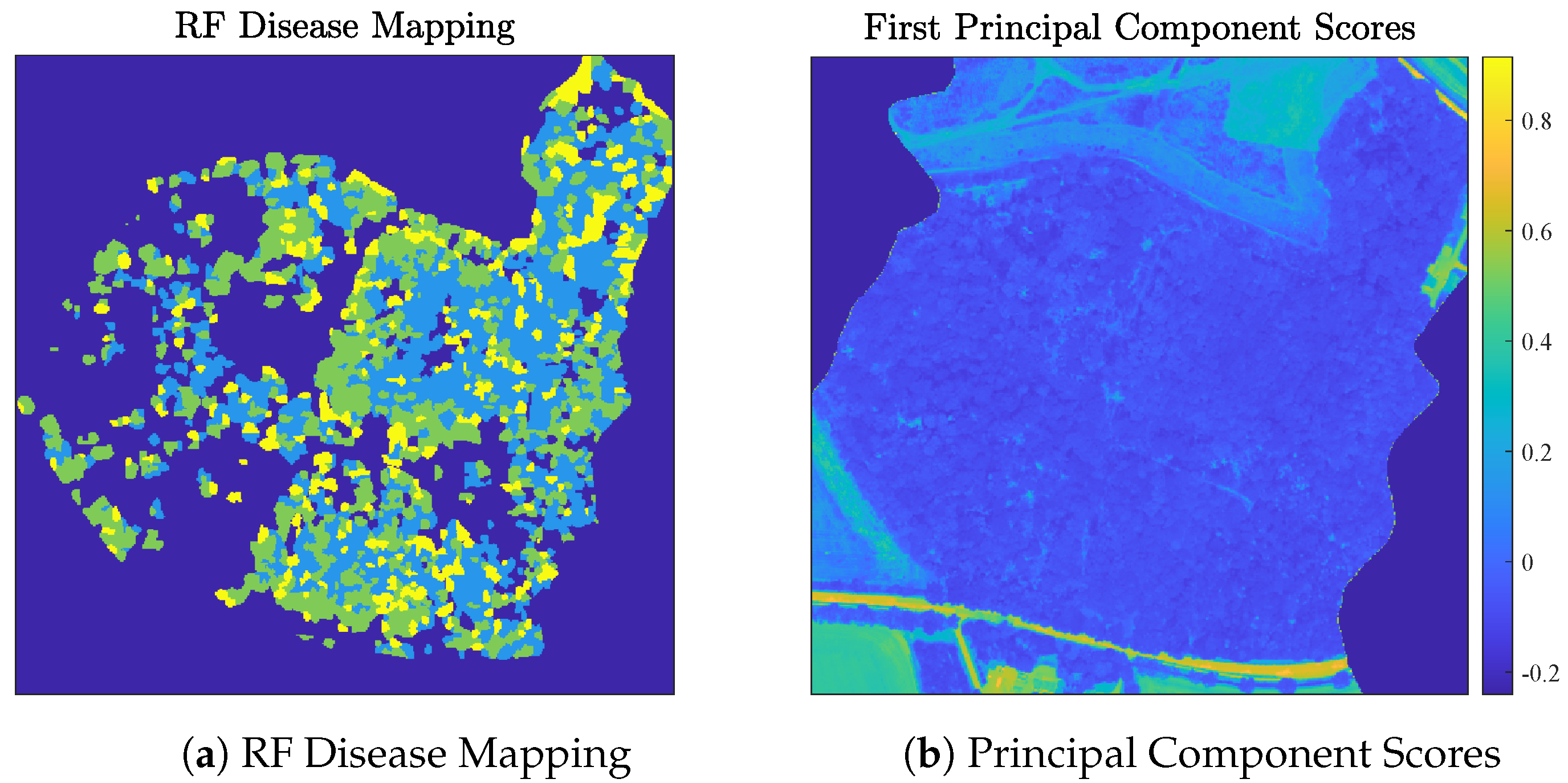
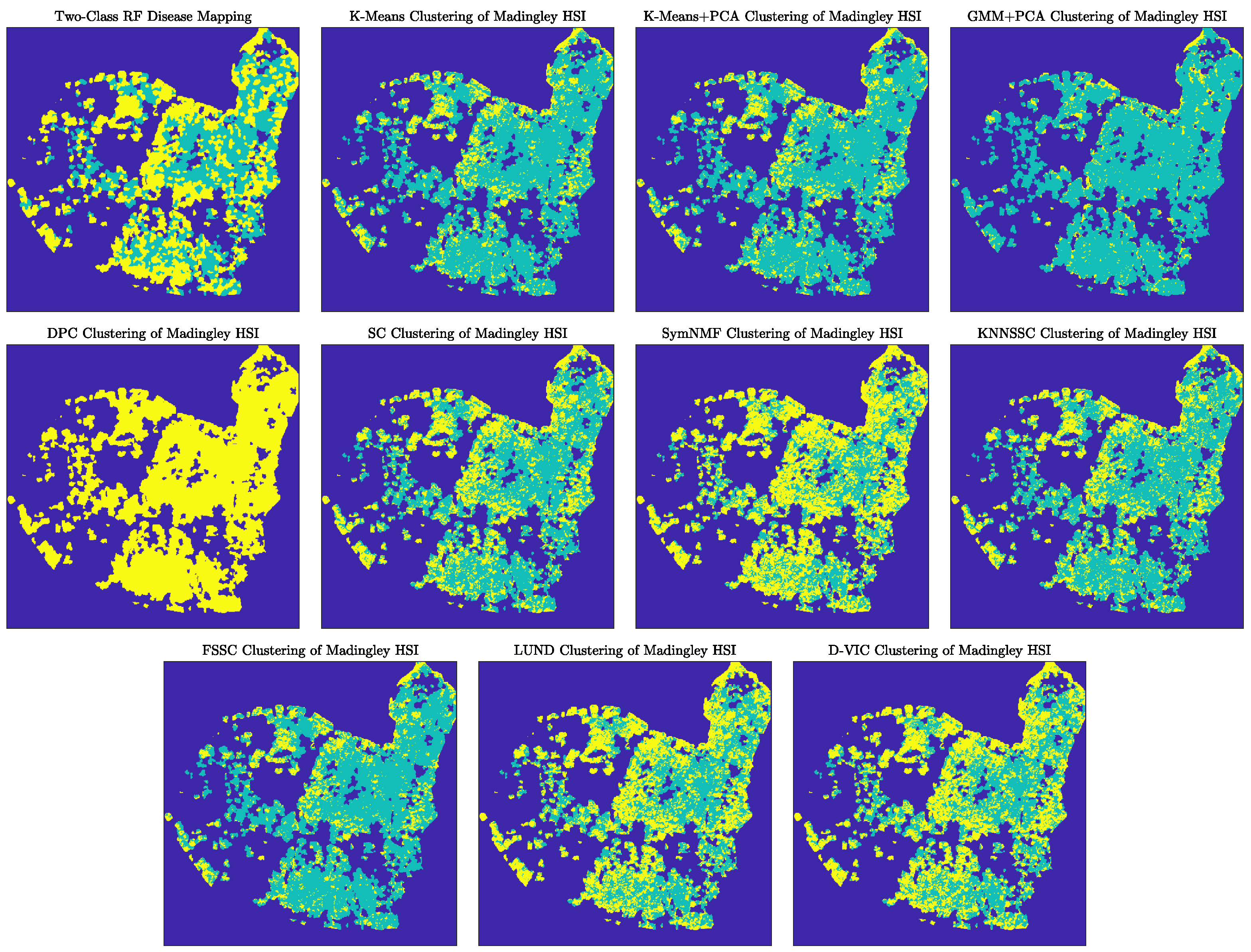
4.2. Analysis of the Madingley HSI Dataset
Discussion of Madingley Experiments
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Hyperparameter Optimization
| Parameter 1 Grid | Parameter 2 Grid | Parameter 3 Grid | |
|---|---|---|---|
| K-Means | — | — | — |
| K-Means + PCA | — | — | — |
| GMM + PCA | — | — | — |
| DPC [135] | — | ||
| SC [41] | — | — | |
| SymNMF [21] | — | — | |
| KNN-SSC [19,20] | — | ||
| FSSC [46] | |||
| LUND [42] | |||
| D-VIC |
References
- Eismann, M.T. Hyperspectral Remote Sensing; SPIE: Philadelphia, PA, USA, 2012. [Google Scholar]
- Ghamisi, P.; Plaza, J.; Chen, Y.; Li, J.; Plaza, A.J. Advanced spectral classifiers for hyperspectral images: A review. IEEE Geosci. Remote Sens. Mag. 2017, 5, 8–32. [Google Scholar] [CrossRef]
- Plaza, A.; Martín, G.; Plaza, J.; Zortea, M.; Sánchez, S. Recent developments in endmember extraction and spectral unmixing. In Optical Remote Sensing: Advances in Signal Processing and Exploitation Techniques; Springer: Berlin/Heidelberg, Germany, 2011; pp. 235–267. [Google Scholar]
- Edelman, G.J.; Gaston, E.; Van Leeuwen, T.G.; Cullen, P.; Aalders, M.C. Hyperspectral imaging for non-contact analysis of forensic traces. Forensic Sci. Int. 2012, 223, 28–39. [Google Scholar] [CrossRef]
- Adam, E.; Mutanga, O.; Rugege, D. Multispectral and hyperspectral remote sensing for identification and mapping of wetland vegetation: A review. Wetl. Ecol. Manag. 2010, 18, 281–296. [Google Scholar] [CrossRef]
- Hirano, A.; Madden, M.; Welch, R. Hyperspectral image data for mapping wetland vegetation. Wetlands 2003, 23, 436–448. [Google Scholar] [CrossRef]
- Clevers, J.G.P.W.; Kooistra, L.; Schaepman, M.E. Estimating canopy water content using hyperspectral remote sensing data. Int. J. Appl. Earth Obs. Geoinf. 2010, 12, 119–125. [Google Scholar]
- Dalponte, M.; Bruzzone, L.; Gianelle, D. Fusion of hyperspectral and LIDAR remote sensing data for classification of complex forest areas. IEEE Trans. Geosci. Remote Sens. 2008, 46, 1416–1427. [Google Scholar] [CrossRef]
- Wang, S.; Guan, K.; Zhang, C.; Lee, D.; Margenot, A.J.; Ge, Y.; Peng, J.; Zhou, W.; Zhou, Q.; Huang, Y. Using soil library hyperspectral reflectance and machine learning to predict soil organic carbon: Assessing potential of airborne and spaceborne optical soil sensing. Remote Sens. Environ. 2022, 271, 112914. [Google Scholar] [CrossRef]
- Jia, J.; Wang, Y.; Chen, J.; Guo, R.; Shu, R.; Wang, J. Status and application of advanced airborne hyperspectral imaging technology: A review. Infr. Phys. Technol. 2020, 104, 103115. [Google Scholar] [CrossRef]
- Price, J.C. Spectral band selection for visible-near infrared remote sensing: Spectral-Spatial resolution tradeoffs. IEEE Trans. Geosci. Remote Sens. 1997, 35, 1277–1285. [Google Scholar] [CrossRef]
- Bioucas-Dias, J.M.; Plaza, A.; Camps-Valls, G.; Scheunders, P.; Nasrabadi, N.; Chanussot, J. Hyperspectral remote sensing data analysis and future challenges. IEEE Geosci. Remote Sens. Mag. 2013, 1, 6–36. [Google Scholar] [CrossRef]
- Laparrcr, V.; Santos-Rodriguez, R. Spatial/spectral information trade-off in hyperspectral images. In Proceedings of the International Geosci Remote Sensing Symposium, Milan, Italy, 26–31 July 2015; IEEE: New York, NY, USA, 2015; pp. 1124–1127. [Google Scholar]
- Miao, L.; Qi, H. Endmember extraction from highly mixed data using minimum volume constrained nonnegative matrix factorization. IEEE Trans. Geosci. Remote Sens. 2007, 45, 765–777. [Google Scholar] [CrossRef]
- Pacheco-Labrador, J.; Migliavacca, M.; Ma, X.; Mahecha, M.; Carvalhais, N.; Weber, U.; Benavides, R.; Bouriaud, O.; Barnoaiea, I.; Coomes, D.A. Challenging the link between functional and spectral diversity with radiative transfer modeling and data. Remote Sens. Environ. 2022, 280, 113170. [Google Scholar] [CrossRef]
- Jia, J.; Wang, Y.; Zhuang, X.; Yao, Y.; Wang, S.; Zhao, D.; Shu, R.; Wang, J. High spatial resolution shortwave infrared imaging technology based on time delay and digital accumulation method. Inf. Phys. Technol. 2017, 81, 305–312. [Google Scholar] [CrossRef]
- Friedman, J.; Hastie, T.; Tibshirani, R. The Elements of Statistical Learning; Springer: Berlin/Heidelberg, Germany, 2001; Volume 1, Springer Series in Statistics. [Google Scholar]
- Murphy, J.M.; Maggioni, M. Unsupervised clustering and active learning of hyperspectral images with nonlinear diffusion. IEEE Trans. Geosci. Remote Sens. 2018, 57, 1829–1845. [Google Scholar] [CrossRef]
- Abdolali, M.; Gillis, N. Beyond linear subspace clustering: A comparative study of nonlinear manifold clustering algorithms. Comput. Sci. Rev. 2021, 42, 100435. [Google Scholar] [CrossRef]
- Zhuang, L.; Wang, J.; Lin, Z.; Yang, A.Y.; Ma, Y.; Yu, N. Locality-preserving low-rank representation for graph construction from nonlinear manifolds. Neurocomputing 2016, 175, 715–722. [Google Scholar] [CrossRef]
- Kuang, D.; Ding, C.; Park, H. Symmetric nonnegative matrix factorization for graph clustering. In Proceedings of the SIAM International Conference Data Min, Anaheim, CA, USA, 26–28 April 2012; SIAM: Philadelphia, PA, USA, 2012; pp. 106–117. [Google Scholar]
- Wang, R.; Nie, N.; Wang, Z.; He, F.; Li, X. Scalable graph-based clustering with nonnegative relaxation for large hyperspectral image. IEEE Trans. Geosci. Remote Sens. 2019, 57, 7352–7364. [Google Scholar] [CrossRef]
- Camps-Valls, G.; Marsheva, T.V.B.; Zhou, D. Semi-supervised graph-based hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 2007, 45, 3044–3054. [Google Scholar] [CrossRef]
- Gao, Y.; Ji, R.; Cui, P.; Dai, Q.; Hua, G. Hyperspectral image classification through bilayer graph-based learning. IEEE Trans. Image Process 2014, 23, 2769–2778. [Google Scholar] [CrossRef]
- Wu, H.; Prasad, S. Semi-supervised deep learning using pseudo labels for hyperspectral image classification. IEEE Trans. Image Process 2017, 27, 1259–1270. [Google Scholar] [CrossRef]
- Yang, X.; Ye, Y.; Li, X.; Lau, R.Y.; Zhang, X.; Huang, X. Hyperspectral image classification with deep learning models. IEEE Trans. Geosci. Remote Sens. 2018, 56, 5408–5423. [Google Scholar] [CrossRef]
- Nalepa, J.; Myller, M.; Imai, Y.; Honda, K.I.; Takeda, T.; Antoniak, M. Unsupervised segmentation of hyperspectral images using 3-D convolutional autoencoders. IEEE Geosci. Remote Sens. Lett. 2020, 17, 1948–1952. [Google Scholar] [CrossRef]
- Gillis, N.; Kuang, D.; Park, H. Hierarchical clustering of hyperspectral images using rank-two nonnegative matrix factorization. IEEE Trans. Geosci. Remote Sens. 2014, 53, 2066–2078. [Google Scholar] [CrossRef]
- Li, K.; Qin, Y.; Ling, Q.; Wang, Y.; Lin, Z.; An, W. Self-supervised deep subspace clustering for hyperspectral images with adaptive self-expressive coefficient matrix initialization. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2021, 14, 3215–3227. [Google Scholar] [CrossRef]
- Sun, J.; Wang, W.; Wei, X.; Fang, L.; Tang, X.; Xu, Y.; Yu, H.; Yao, W. Deep clustering with intraclass distance constraint for hyperspectral images. IEEE Trans. Geosci. Remote Sens. 2020, 59, 4135–4149. [Google Scholar] [CrossRef]
- Zhou, J.; Kwan, C.; Ayhan, B.; Eismann, M.T. A novel cluster kernel RX algorithm for anomaly and change detection using hyperspectral images. IEEE Trans. Geosci. Remote Sens. 2016, 54, 6497–6504. [Google Scholar] [CrossRef]
- Cui, K.; Plemmons, R.J. Unsupervised classification of AVIRIS-NG hyperspectral images. In Proceedings of the Workshop on Hyperspectral Imaging and Signal Processing: Evolution in Remote Sensing, Amsterdam, The Netherlands, 24–26 March 2021; IEEE: New York, NY, USA, 2021; pp. 1–5. [Google Scholar]
- Cui, K.; Li, R.; Polk, S.L.; Murphy, J.M.; Plemmons, R.J.; Chan, R.H. Unsupervised spatial-spectral hyperspectral image reconstruction and clustering with diffusion geometry. In Proceedings of the Workshop Hyperspectral Image Signal Process Evolution in Remote Sensing, Rome, Italy, 13–16 September 2022; IEEE: New York, NY, USA, 2022; pp. 1–5. [Google Scholar]
- Bachmann, C.M.; Ainsworth, T.L.; Fusina, R.A. Exploiting manifold geometry in hyperspectral imagery. IEEE Trans. Geosci. Remote Sens. 2005, 43, 441–454. [Google Scholar] [CrossRef]
- Coifman, R.R.; Lafon, S. Diffusion maps. Appl. Comput. Harm. Anal. 2006, 21, 5–30. [Google Scholar] [CrossRef]
- Baral, H.; Queloz, V.; Hosoya, T. Hymenoscyphus fraxineus, the correct scientific name for the fungus causing ash dieback in Europe. IMA Fungus 2014, 5, 79–80. [Google Scholar] [CrossRef]
- McKinney, L.; Nielsen, L.; Collinge, D.; Thomsen, I.; Hansen, J.; Kjær, E. The ash dieback crisis: Genetic variation in resistance can prove a long-term solution. Plant Pathol. 2014, 63, 485–499. [Google Scholar] [CrossRef]
- Stone, C.; Mohammed, C. Application of remote sensing technologies for assessing planted forests damaged by insect pests and fungal pathogens: A review. Curr. For. Rep. 2017, 3, 75–92. [Google Scholar]
- Waser, L.T.; Küchler, M.; Jütte, K.; Stampfer, T. Evaluating the potential of WorldView-2 data to classify tree species and different levels of ash mortality. Remote Sens. 2014, 6, 4515–4545. [Google Scholar] [CrossRef]
- Chan, A.H.Y.; Barnes, C.; Swinfield, T.; Coomes, D.A. Monitoring ash dieback (Hymenoscyphus fraxineus) in British forests using hyperspectral remote sensing. Remote Sens. Ecol. Conserv. 2021, 7, 306–320. [Google Scholar] [CrossRef]
- Ng, A.Y.; Jordan, M.I.; Weiss, Y. On spectral clustering: Analysis and an algorithm. Adv. Neural Inf. Process Syst. 2002, 14, 849–856. [Google Scholar]
- Maggioni, M.; Murphy, J.M. Learning by unsupervised nonlinear diffusion. J. Mach. Learn. Res. 2019, 20, 1–56. [Google Scholar]
- Cahill, N.D.; Czaja, W.; Messinger, D.W. Schroedinger Eigenmaps with Nondiagonal Potentials for Spatial-Spectral Clustering of Hyperspectral Imagery; SPIE: Philadelphia, PA, USA, 2014; Volume 9088, pp. 27–39. [Google Scholar]
- Theodoridis, S.; Koutroumbas, K. Pattern Recognition; Elsevier: Amsterdam, The Netherlands, 2006. [Google Scholar]
- Zhu, W.; Chayes, V.; Tiard, A.; Sanchez, S.; Dahlberg, D.; Bertozzi, A.L.; Osher, S.; Zosso, D.; Kuang, D. Unsupervised classification in hyperspectral imagery with nonlocal total variation and primal-dual hybrid gradient algorithm. IEEE Trans. Geosci. Remote Sens. 2017, 55, 2786–2798. [Google Scholar] [CrossRef]
- Wang, J.; Ma, Z.; Nie, F.; Li, X. Fast self-supervised clustering with anchor graph. IEEE Trans. Neural Netw. Learn. Syst. 2021, 33, 4199–4212. [Google Scholar] [CrossRef]
- Bandyopadhyay, D.; Mukherjee, S. Tree species classification from hyperspectral data using graph-regularized neural networks. arXiv 2022, arXiv:2208.08675. [Google Scholar]
- Tenenbaum, J.B.; de Silva, V.; Langford, J.C. A global geometric framework for nonlinear dimensionality reduction. Science 2000, 290, 2319–2323. [Google Scholar] [CrossRef]
- Roweis, S.T.; Saul, L.K. Nonlinear dimensionality reduction by locally linear embedding. Science 2000, 290, 2323–2326. [Google Scholar] [CrossRef]
- Belkin, M.; Niyogi, P. Laplacian eigenmaps and spectral techniques for embedding and clustering. Adv. Neural Inf. Process Syst. 2001, 585–591. [Google Scholar]
- Rohe, K.; Chatterjee, S.; Yu, B. Spectral clustering and the high-dimensional stochastic blockmodel. Ann. Stat. 2011, 39, 1878–1915. [Google Scholar] [CrossRef]
- Murphy, J.M.; Polk, S.L. A multiscale environment for learning by diffusion. Appl. Comput. Harm. Anal. 2022, 57, 58–100. [Google Scholar] [CrossRef]
- Nadler, B.; Galun, M. Fundamental limitations of spectral clustering. Adv. Neural Inf. Process Syst. 2007, 19, 1017–1024. [Google Scholar]
- Dilokthanakul, N.; Mediano, P.A.M.; Garnelo, M.; Lee, M.C.H.; Salimbeni, H.; Arulkumaran, K.; Shanahan, M. Deep unsupervised clustering with Gaussian mixture variational autoencoders. arXiv 2016, arXiv:1611.02648. [Google Scholar]
- Min, E.; Guo, X.; Liu, Q.; Zhang, G.; Cui, J.; Long, J. A survey of clustering with deep learning: From the perspective of network architecture. IEEE Access 2018, 6, 39501–39514. [Google Scholar] [CrossRef]
- Tasissa, A.; Nguyen, D.; Murphy, J.M. Deep diffusion processes for active learning of hyperspectral images. In Proceedings of the 2021 IEEE International Geoscience and Remote Sensing Symposium IGARSS, Brussels, Belgium, 11–16 July 2021; IEEE: New York, NY, USA, 2021; pp. 3665–3668. [Google Scholar]
- Nguyen, A.; Yosinski, J.; Clune, J. Deep neural networks are easily fooled: High confidence predictions for unrecognizable images. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 427–436. [Google Scholar]
- Szegedy, C.; Zaremba, W.; Sutskever, I.; Bruna, J.; Erhan, D.; Goodfellow, I.; Fergus, R. Intriguing properties of neural networks. In Proceedings of the International Conference Learn Represent, Banff, AB, Canada, 14–16 April 2014. [Google Scholar]
- Haeffele, B.D.; You, C.; Vidal, R. A Critique of Self-Expressive Deep Subspace Clustering. In Proceedings of the International Conference Learn Represent, Addis Ababa, Ethiopia, 26–30 April 2020. [Google Scholar]
- Polk, S.L.; Murphy, J.M. Multiscale clustering of hyperspectral images through spectral-spatial diffusion geometry. In Proceedings of the International Geoscience and Remote Sensing Symposium IGARSS, Brussels, Belgium, 11–16 July 2021; pp. 4688–4691. [Google Scholar]
- Haghverdi, L.; Büttner, M.; Wolf, F.A.; Buettner, F.; Theis, F.J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 2016, 13, 845–848. [Google Scholar] [CrossRef]
- Van Dijk, D.; Sharma, R.; Nainys, J.; Yim, K.; Kathail, P.; Carr, A.J.; Burdziak, C.; Moon, K.R.; Chaffer, C.L.; Pattabiraman, D. Recovering gene interactions from single-cell data using data diffusion. Cell 2018, 174, 716–729. [Google Scholar] [CrossRef]
- Zhao, Z.; Singer, A. Rotationally invariant image representation for viewing direction classification in cryo-EM. J. Struct. Biol. 2014, 186, 153–166. [Google Scholar] [CrossRef]
- Moon, K.R.; van Dijk, D.; Wang, Z.; Gigante, S.; Burkhardt, D.B.; Chen, W.S.; Yim, K.; van den Elzen, A.; Hirn, M.J.; Coifman, R.R. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 2019, 37, 1482–1492. [Google Scholar] [CrossRef]
- Rohrdanz, M.A.; Zheng, W.; Maggioni, M.; Clementi, C. Determination of reaction coordinates via locally scaled diffusion map. J. Chem. Phys. 2011, 134, 03B624. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Rohrdanz, M.A.; Maggioni, M.; Clementi, C. Polymer reversal rate calculated via locally scaled diffusion map. J. Chem. Phys. 2011, 134, 144109. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Ferguson, A.L. Molecular enhanced sampling with autoencoders: On-the-fly collective variable discovery and accelerated free energy landscape exploration. J. Comput. Chem. 2018, 39, 2079–2102. [Google Scholar] [CrossRef]
- Coifman, R.R.; Lafon, S.; Lee, A.B.; Maggioni, M.; Nadler, B.; Warner, F.; Zucker, S.W. Geometric diffusions as a tool for harmonic analysis and structure definition of data: Diffusion maps. Proc. Natl. Acad. Sci. USA 2005, 102, 7426–7431. [Google Scholar] [CrossRef]
- Nadler, B.; Lafon, S.; Coifman, R.R.; Kevrekidis, I.G. Diffusion maps, spectral clustering and reaction coordinates of dynamical systems. Appl. Comput. Harmon. Anal. 2006, 21, 113–127. [Google Scholar] [CrossRef]
- Chan, T.; Ma, W.; Ambikapathi, A.; Chi, C. A simplex volume maximization framework for hyperspectral endmember extraction. IEEE Trans. Geosci. Remote Sens. 2011, 49, 4177–4193. [Google Scholar] [CrossRef]
- Winter, M.E. N-FINDR: An algorithm for fast autonomous spectral end-member determination in hyperspectral data. In Imaging Spectrometry V; SPIE: Philadelphia, PA, USA, 1999; Volume 3753, pp. 266–275. [Google Scholar]
- Manolakis, D.; Siracusa, C.; Shaw, G. Hyperspectral subpixel target detection using the linear mixing model. IEEE Trans. Geosci. Remote Sens. 2001, 39, 1392–1409. [Google Scholar] [CrossRef]
- Zhao, X.L.; Wang, F.; Huang, T.Z.; Ng, M.K.; Plemmons, R.J. Deblurring and sparse unmixing for hyperspectral images. IEEE Trans. Geosci. Remote Sens. 2013, 51, 4045–4058. [Google Scholar] [CrossRef]
- Berisha, S.; Nagy, J.G.; Plemmons, R.J. Deblurring and sparse unmixing of hyperspectral images using multiple point spread functions. SIAM J. Sci. Comput. 2015, 37, S389–S406. [Google Scholar] [CrossRef]
- Wang, L.; Feng, Y.; Gao, Y.; Wang, Z.; He, M. Compressed sensing reconstruction of hyperspectral images based on spectral unmixing. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2018, 11, 1266–1284. [Google Scholar] [CrossRef]
- Cerra, D.; Müller, R.; Reinartz, P. Noise reduction in hyperspectral images through spectral unmixing. IEEE Geosci. Remote Sens. Lett. 2013, 11, 109–113. [Google Scholar] [CrossRef]
- Rasti, B.; Scheunders, P.; Ghamisi, P.; Licciardi, G.; Chanussot, J. Noise reduction in hyperspectral imagery: Overview and application. Remote Sens. 2018, 10, 482. [Google Scholar] [CrossRef]
- Rasti, B.; Koirala, B.; Scheunders, P.; Ghamisi, P. How hyperspectral image unmixing and denoising can boost each other. Remote Sens. 2020, 12, 1728. [Google Scholar] [CrossRef]
- Ertürk, A.; Güllü, M.K.; Çeşmeci, D.; Gerçek, D.; Ertürk, S. Spatial resolution enhancement of hyperspectral images using unmixing and binary particle swarm optimization. IEEE Geosci. Remote Sens. Lett. 2014, 11, 2100–2104. [Google Scholar] [CrossRef]
- Bendoumi, M.A.; He, M.; Mei, S. Hyperspectral image resolution enhancement using high-resolution multispectral image based on spectral unmixing. IEEE Trans. Geosci. Remote Sens. 2014, 52, 6574–6583. [Google Scholar] [CrossRef]
- Kordi Ghasrodashti, E.; Karami, A.; Heylen, R.; Scheunders, P. Spatial resolution enhancement of hyperspectral images using spectral unmixing and Bayesian sparse representation. Remote Sens. 2017, 9, 541. [Google Scholar] [CrossRef]
- Villa, A.; Chanussot, J.; Benediktsson, J.A.; Jutten, C. Spectral unmixing for the classification of hyperspectral images at a finer spatial resolution. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2010, 5, 521–533. [Google Scholar] [CrossRef]
- Dópido, I.; Villa, A.; Plaza, A.; Gamba, P. A quantitative and comparative assessment of unmixing-based feature extraction techniques for hyperspectral image classification. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2012, 5, 421–435. [Google Scholar] [CrossRef]
- Ertürk, A.; Plaza, A. Informative change detection by unmixing for hyperspectral images. IEEE Geosci. Remote Sens. Lett. 2015, 12, 1252–1256. [Google Scholar] [CrossRef]
- Liu, S.; Bruzzone, L.; Bovolo, F.; Du, P. Unsupervised multitemporal spectral unmixing for detecting multiple changes in hyperspectral images. IEEE Trans. Geosci. Remote Sens. 2016, 54, 2733–2748. [Google Scholar] [CrossRef]
- Camalan, S.; Cui, K.; Pauca, V.P.; Alqahtani, S.; Silman, M.; Chan, R.; Plemmons, R.J.; Dethier, E.N.; Fernandez, L.E.; Lutz, D.A. Change detection of Amazonian alluvial gold mining using deep learning and Sentinel-2 imagery. Remote Sens. 2022, 14, 1746. [Google Scholar] [CrossRef]
- Li, H.; Wu, K.; Xu, Y. An Integrated Change Detection Method Based on Spectral Unmixing and the CNN for Hyperspectral Imagery. Remote Sens. 2022, 14, 2523. [Google Scholar] [CrossRef]
- Qu, Y.; Wang, W.; Guo, R.; Ayhan, B.; Kwan, C.; Vance, S.; Qi, H. Hyperspectral anomaly detection through spectral unmixing and dictionary-based low-rank decomposition. IEEE Trans. Geosci. Remote Sens. 2018, 56, 4391–4405. [Google Scholar] [CrossRef]
- Ma, D.; Yuan, Y.; Wang, Q. Hyperspectral anomaly detection via discriminative feature learning with multiple-dictionary sparse representation. Remote Sens. 2018, 10, 745. [Google Scholar] [CrossRef]
- Somers, B.; Asner, G.P.; Tits, L.; Coppin, P. Endmember variability in spectral mixture analysis: A review. Remote Sens. Environ. 2011, 115, 1603–1616. [Google Scholar] [CrossRef]
- Quintano, C.; Fernández-Manso, A.; Shimabukuro, Y.E.; Pereira, G. Spectral unmixing. Int. J. Remote Sens. 2012, 33, 5307–5340. [Google Scholar] [CrossRef]
- Bioucas-Dias, J.M.; Plaza, A.; Dobigeon, N.; Parente, M.; Du, Q.; Gader, P.; Chanussot, J. Hyperspectral unmixing overview: Geometrical, statistical, and sparse regression-based approaches. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2012, 5, 354–379. [Google Scholar] [CrossRef]
- Heylen, R.; Parente, M.; Gader, P. A review of nonlinear hyperspectral unmixing methods. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 1844–1868. [Google Scholar] [CrossRef]
- Borsoi, R.; Imbiriba, T.; Bermudez, J.C.; Richard, C.; Chanussot, J.; Drumetz, L.; Tourneret, J.Y.; Zare, A.; Jutten, C. Spectral Variability in Hyperspectral Data Unmixing: A Comprehensive Review. IEEE Geosci. Remote Sens. Mag. 2021, 9, 223–270. [Google Scholar] [CrossRef]
- Chang, C.; Wu, C.; Liu, W.; Ouyang, Y. A new growing method for simplex-based endmember extraction algorithm. IEEE Trans. Geosci. Remote Sens. 2006, 44, 2804–2819. [Google Scholar] [CrossRef]
- Neville, R. Automatic endmember extraction from hyperspectral data for mineral exploration. In Proceedings of the Fourth International Airborne Remote Sensing Conference and Exhibition/21st Canadian Symposium on Remote Sensing, Ottawa, ON, Canada, 21–24 June 1999. [Google Scholar]
- Boardman, J.W.; Kruse, F.A.; Green, R.O. Mapping Target Signatures via Partial Unmixing of AVIRIS Data; Technical Report; Jet Propulsion Laboratory: Pasadena, CA, USA, 1995. [Google Scholar]
- Boardman, J.W. Automating spectral unmixing of AVIRIS data using convex geometry concepts. Annu. JPL Airborne Geosci. Workshop 1993, 1, 11–14. [Google Scholar]
- Chan, T.; Chi, C.; Huang, Y.; Ma, W. A convex analysis-based minimum-volume enclosing simplex algorithm for hyperspectral unmixing. IEEE Trans. Signal Process 2009, 57, 4418–4432. [Google Scholar] [CrossRef]
- Nascimento, J.M.; Dias, J.M. Vertex component analysis: A fast algorithm to unmix hyperspectral data. IEEE Trans. Geosci. Remote Sens. 2005, 43, 898–910. [Google Scholar] [CrossRef]
- Clasen, A.; Somers, B.; Pipkins, K.; Tits, L.; Segl, K.; Brell, M.; Kleinschmit, B.; Spengler, D.; Lausch, A.; Förster, M. Spectral unmixing of forest crown components at close range, airborne and simulated Sentinel-2 and EnMAP spectral imaging scale. Remote Sens. 2015, 7, 15361–15387. [Google Scholar] [CrossRef]
- Heylen, R.; Burazerovic, D.; Scheunders, P. Fully constrained least squares spectral unmixing by simplex projection. IEEE Trans. Geosci. Remote Sens. 2011, 49, 4112–4122. [Google Scholar] [CrossRef]
- Hendrix, E.M.; Garcia, I.; Plaza, J.; Martin, G.; Plaza, A. A new minimum-volume enclosing algorithm for endmember identification and abundance estimation in hyperspectral data. IEEE Trans. Geosci. Remote Sens. 2011, 50, 2744–2757. [Google Scholar] [CrossRef]
- Iordache, M.D.; Bioucas-Dias, J.M.; Plaza, A. Sparse unmixing of hyperspectral data. IEEE Trans. Geosci. Remote Sens. 2011, 49, 2014–2039. [Google Scholar] [CrossRef]
- Berman, M.; Kiiveri, H.; Lagerstrom, R.; Ernst, A.; Dunne, R.; Huntington, J.F. ICE: A statistical approach to identifying endmembers in hyperspectral images. IEEE Trans. Signal Process 2004, 42, 2085–2095. [Google Scholar] [CrossRef]
- Zare, A.; Gader, P. Sparsity promoting iterated constrained endmember detection in hyperspectral imagery. IEEE Geosci. Remote Sens. Lett. 2007, 4, 446–450. [Google Scholar] [CrossRef]
- Dobigeon, N.; Moussaoui, S.; Coulon, M.; Tourneret, J.; Hero, A.O. Joint Bayesian endmember extraction and linear unmixing for hyperspectral imagery. IEEE Trans. Signal Process 2009, 57, 4355–4368. [Google Scholar] [CrossRef]
- Moussaoui, S.; Brie, D.; Mohammad-Djafari, A.; Carteret, C. Separation of non-negative mixture of non-negative sources using a Bayesian approach and MCMC sampling. IEEE Trans. Signal Process 2006, 54, 4133–4145. [Google Scholar] [CrossRef]
- Themelis, K.E.; Rontogiannis, A.A.; Koutroumbas, K.D. A novel hierarchical Bayesian approach for sparse semisupervised hyperspectral unmixing. IEEE Trans. Signal Process 2011, 60, 585–599. [Google Scholar] [CrossRef]
- Palsson, B.; Ulfarsson, M.O.; Sveinsson, J.R. Convolutional autoencoder for spectral–spatial hyperspectral unmixing. IEEE Trans. Geosci. Remote Sens. 2020, 59, 535–549. [Google Scholar] [CrossRef]
- Su, Y.; Li, J.; Plaza, A.; Marinoni, A.; Gamba, P.; Chakravortty, S. DAEN: Deep autoencoder networks for hyperspectral unmixing. IEEE Trans. Geosci. Remote Sens. 2019, 57, 4309–4321. [Google Scholar] [CrossRef]
- Palsson, B.; Sigurdsson, J.; Sveinsson, J.R.; Ulfarsson, M.O. Hyperspectral unmixing using a neural network autoencoder. IEEE Access 2018, 6, 25646–25656. [Google Scholar] [CrossRef]
- Qu, Y.; Qi, H. uDAS: An untied denoising autoencoder with sparsity for spectral unmixing. IEEE Trans. Geosci. Remote Sens. 2018, 57, 1698–1712. [Google Scholar] [CrossRef]
- Ozkan, S.; Kaya, B.; Akar, G.B. Endnet: Sparse autoencoder network for endmember extraction and hyperspectral unmixing. IEEE Trans. Geosci. Remote Sens. 2018, 57, 482–496. [Google Scholar] [CrossRef]
- Zhang, X.; Sun, Y.; Zhang, J.; Wu, P.; Jiao, L. Hyperspectral unmixing via deep convolutional neural networks. IEEE Geosci. Remote Sens. Lett. 2018, 15, 1755–1759. [Google Scholar] [CrossRef]
- Su, Y.; Marinoni, A.; Li, J.; Plaza, J.; Gamba, P. Stacked nonnegative sparse autoencoders for robust hyperspectral unmixing. IEEE Geosci. Remote Sens. Lett. 2018, 15, 1427–1431. [Google Scholar] [CrossRef]
- Khajehrayeni, F.; Ghassemian, H. Hyperspectral unmixing using deep convolutional autoencoders in a supervised scenario. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2020, 13, 567–576. [Google Scholar] [CrossRef]
- Feng, X.; Li, H.; Li, J.; Du, Q.; Plaza, A.; Emery, W.J. Hyperspectral unmixing using sparsity-constrained deep nonnegative matrix factorization with total variation. IEEE Trans. Geosci. Remote Sens. 2018, 56, 6245–6257. [Google Scholar] [CrossRef]
- Guilfoyle, K.J.; Althouse, M.L.; Chang, C. A quantitative and comparative analysis of linear and nonlinear spectral mixture models using radial basis function neural networks. IEEE Trans. Geosci. Remote Sens. 2001, 39, 2314–2318. [Google Scholar] [CrossRef]
- Licciardi, G.A.; Del Frate, F. Pixel unmixing in hyperspectral data by means of neural networks. IEEE Trans. Geosci. Remote Sens. 2011, 49, 4163–4172. [Google Scholar] [CrossRef]
- Charles, A.S.; Olshausen, B.A.; Rozell, C.J. Learning sparse codes for hyperspectral imagery. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2011, 5, 963–978. [Google Scholar] [CrossRef]
- Wang, M.; Zhao, M.; Chen, J.; Rahardja, S. Nonlinear unmixing of hyperspectral data via deep autoencoder networks. IEEE Geosci. Remote Sens. Lett. 2019, 16, 1467–1471. [Google Scholar] [CrossRef]
- Yokoya, N.; Chanussot, J.; Iwasaki, A. Nonlinear unmixing of hyperspectral data using semi-nonnegative matrix factorization. IEEE Trans. Geosci. Remote Sens. 2013, 52, 1430–1437. [Google Scholar] [CrossRef]
- Halimi, A.; Altmann, Y.; Dobigeon, N.; Tourneret, J.Y. Nonlinear unmixing of hyperspectral images using a generalized bilinear model. IEEE Trans. Geosci. Remote Sens. 2011, 49, 4153–4162. [Google Scholar] [CrossRef]
- Chen, J.; Richard, C.; Honeine, P. Nonlinear unmixing of hyperspectral data based on a linear-mixture/nonlinear-fluctuation model. IEEE Trans. Signal Process 2012, 61, 480–492. [Google Scholar] [CrossRef]
- Heylen, R.; Scheunders, P. A multilinear mixing model for nonlinear spectral unmixing. IEEE Trans. Geosci. Remote Sens. 2015, 54, 240–251. [Google Scholar] [CrossRef]
- Heylen, R.; Burazerovic, D.; Scheunders, P. Non-linear spectral unmixing by geodesic simplex volume maximization. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2010, 5, 534–542. [Google Scholar] [CrossRef]
- Bioucas-Dias, J.M.; Nascimento, J.M. Hyperspectral subspace identification. IEEE Trans. Geosci. Remote Sens. 2008, 46, 2435–2445. [Google Scholar] [CrossRef]
- Chang, C. A review of virtual dimensionality for hyperspectral imagery. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2018, 11, 1285–1305. [Google Scholar] [CrossRef]
- Chang, C.; Du, Q. Estimation of number of spectrally distinct signal sources in hyperspectral imagery. IEEE Trans. Geosci. Remote Sens. 2004, 42, 608–619. [Google Scholar] [CrossRef]
- Chan, T.H.; Ma, W.K.; Chi, C.Y.; Wang, Y. A convex analysis framework for blind separation of non-negative sources. IEEE Trans. Signal Process 2008, 56, 5120–5134. [Google Scholar] [CrossRef]
- Bro, R.; De Jong, S. A fast non-negativity-constrained least squares algorithm. J. Chem. 1997, 11, 393–401. [Google Scholar] [CrossRef]
- Chen, J.; Richard, C.; Honeine, P. Nonlinear estimation of material abundances in hyperspectral images with ℓ1-norm spatial regularization. IEEE Trans. Geosci. Remote Sens. 2013, 52, 2654–2665. [Google Scholar] [CrossRef]
- Heinz, D.C.; Chang, C.I. Fully constrained least squares linear spectral mixture analysis method for material quantification in hyperspectral imagery. IEEE Trans. Geosci. Remote Sens. 2001, 39, 529–545. [Google Scholar] [CrossRef]
- Rodriguez, A.; Laio, A. Clustering by fast search and find of density peaks. Science 2014, 344, 1492–1496. [Google Scholar] [CrossRef]
- Beygelzimer, A.; Kakade, S.; Langford, J. Cover trees for nearest neighbor. In Proceedings of the International Conference Mach Learn, Orlando, FL, USA, 25–29 June 2006; pp. 97–104. [Google Scholar]
- Shi, J.; Malik, J. Normalized cuts and image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2000, 22, 888–905. [Google Scholar]
- Polk, S.L.; Chan, A.H.Y.; Cui, K.; Plemmons, R.J.; Coomes, D.A.; Murphy, J.M. Unsupervised detection of ash dieback disease (Hymenoscyphus fraxineus) using diffusion-based hyperspectral image clustering. In Proceedings of the International Geoscience and Remote Sensing Symposium, Kuala Lumpur, Malaysia, 17–22 July 2022; pp. 2287–2290. [Google Scholar]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Swinfield, T.; Both, S.; Riutta, T.; Bongalov, B.; Elias, D.; Majalap-Lee, N.; Ostle, N.; Svátek, M.; Kvasnica, J.; Milodowski, D. Imaging spectroscopy reveals the effects of topography and logging on the leaf chemistry of tropical forest canopy trees. Glob Chang. Biol. 2020, 26, 989–1002. [Google Scholar] [CrossRef] [PubMed]
- Kotzagiannidis, M.S.; Schönlieb, C.B. Semi-supervised superpixel-based multi-feature graph learning for hyperspectral image data. IEEE Trans. Geosci. Remote Sens. 2021, 60, 4703612. [Google Scholar] [CrossRef]
- Qin, A.; Shang, Z.; Tian, J.; Wang, Y.; Zhang, T.; Tang, Y.Y. Spectral–spatial graph convolutional networks for semisupervised hyperspectral image classification. IEEE Geosci. Remote Sens. Lett. 2018, 16, 241–245. [Google Scholar] [CrossRef]
- Hong, D.; Gao, L.; Yao, J.; Zhang, B.; Plaza, A.; Chanussot, J. Graph convolutional networks for hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 2020, 59, 5966–5978. [Google Scholar] [CrossRef]
- Sun, H.; Zheng, X.; Lu, X. A supervised segmentation network for hyperspectral image classification. IEEE Trans. Image Process 2021, 30, 2810–2825. [Google Scholar] [CrossRef] [PubMed]
- Kavalerov, I.; Li, W.; Czaja, W.; Chellappa, R. 3-D Fourier scattering transform and classification of hyperspectral images. IEEE Trans. Geosci. Remote Sens. 2020, 59, 10312–10327. [Google Scholar] [CrossRef]
- Murphy, J.M.; Maggioni, M. Spectral–spatial diffusion geometry for hyperspectral image clustering. IEEE Geosci. Remote Sens. Lett. 2019, 17, 1243–1247. [Google Scholar] [CrossRef]
- Murphy, J.M. Spatially regularized active diffusion learning for high-dimensional images. Pattern Recognit. Lett. 2020, 135, 213–220. [Google Scholar] [CrossRef]
- Keys, R. Cubic convolution interpolation for digital image processing. IEEE Trans. Signal Process 1981, 29, 1153–1160. [Google Scholar] [CrossRef]
- Murphy, J.M. Patch-Based Diffusion Learning for Hyperspectral Image Clustering. In Proceedings of the International Geoscience and Remote Sensing Symposium, Waikoloa, HI, USA, 26 September–2 October 2020; IEEE: New York, NY, USA, 2020; pp. 1042–1045. [Google Scholar]
- Fauvel, M.; Tarabalka, Y.; Benediktsson, J.A.; Chanussot, J.; Tilton, J.C. Advances in spectral-spatial classification of hyperspectral images. Proc. IEEE 2012, 101, 652–675. [Google Scholar] [CrossRef]
- Ghamisi, P.; Benediktsson, J.A.; Ulfarsson, M.O. Spectral–spatial classification of hyperspectral images based on hidden Markov random fields. IEEE Trans. Geosci. Remote Sens. 2013, 52, 2565–2574. [Google Scholar] [CrossRef]
- Fang, L.; Li, S.; Duan, W.; Ren, J.; Benediktsson, J.A. Classification of hyperspectral images by exploiting spectral–spatial information of superpixel via multiple kernels. IEEE Trans. Geosci. Remote Sens. 2015, 53, 6663–6674. [Google Scholar] [CrossRef]
- Tarabalka, Y.; Benediktsson, J.A.; Chanussot, J. Spectral–spatial classification of hyperspectral imagery based on partitional clustering techniques. IEEE Trans. Geosci. Remote Sens. 2009, 47, 2973–2987. [Google Scholar] [CrossRef]
- Mohan, A.; Sapiro, G.; Bosch, E. Spatially coherent nonlinear dimensionality reduction and segmentation of hyperspectral images. IEEE Geosci. Remote Sens. Lett. 2007, 4, 206–210. [Google Scholar] [CrossRef]
- Polk, S.L. Diffusion-Based Clustering of High-Dimensional Datasets. Ph.D. Thesis, Tufts University, Medford, MA, USA, 2022. [Google Scholar]
- Maggioni, M.; Murphy, J.M. Learning by active nonlinear diffusion. Found. Data Sci. 2019, 1, 271. [Google Scholar] [CrossRef]
- Gerg, I.; Kun, D. Hyperspectral Toolbox. 2022. Available online: https://github.com/davidkun/HyperSpectralToolbox (accessed on 12 December 2021).



| Dataset | Spatial Resolution | Spectral Range | Spatial Dimensions | Num. Pixels | Num. Spectral Bands | Num. Clusters |
|---|---|---|---|---|---|---|
| Salinas A | 1.3 m | 380–2500 nm | ||||
| Jasper Ridge | 5.0 m | 380–2500 nm | 10,000 | |||
| Indian Pines | 20 m | 400–2500 nm | 21,025 |
| Salinas A | Jasper Ridge | Indian Pines | ||||
|---|---|---|---|---|---|---|
| OA | OA | OA | ||||
| K-Means | 0.764 | 0.703 | 0.784 | 0.703 | 0.383 | 0.315 |
| K-Means + PCA | 0.764 | 0.703 | 0.785 | 0.703 | 0.382 | 0.316 |
| GMM + PCA | 0.611 | 0.512 | 0.789 | 0.701 | 0.364 | 0.292 |
| DPC | 0.629 | 0.529 | 0.809 | 0.727 | 0.410 | 0.271 |
| SC | 0.834 | 0.797 | 0.760 | 0.670 | 0.382 | 0.314 |
| SymNMF | 0.828 | 0.791 | 0.662 | 0.542 | 0.365 | 0.304 |
| KNN-SSC | 0.844 | 0.809 | 0.726 | 0.629 | 0.371 | 0.308 |
| FSSC | 0.830 | 0.793 | 0.780 | 0.691 | 0.396 | 0.281 |
| LUND | 0.887 | 0.860 | 0.815 | 0.737 | 0.404 | 0.312 |
| D-VIC | 0.976 | 0.970 | 0.865 | 0.805 | 0.445 | 0.350 |
| Salinas A | Jasper Ridge | Indian Pines | |
|---|---|---|---|
| K-Means | 0.04 | 0.10 | 1.04 |
| K-Means + PCA | 0.10 | 0.14 | 0.58 |
| GMM + PCA | 0.13 | 0.23 | 2.19 |
| DPC | 3.20 | 6.41 | 25.77 |
| SC | 1.82 | 3.15 | 14.54 |
| SymNMF | 3.50 | 4.42 | 48.29 |
| KNN-SSC | 4.11 | 7.91 | 103.05 |
| FSSC | 13.53 | 30.40 | 130.72 |
| LUND | 2.35 | 4.14 | 14.74 |
| D-VIC | 4.95 | 7.64 | 23.70 |
| KM | KM + PCA | GMM + PCA | DPC | SC | SymNMF | KNN-SSC | FSSC | LUND | D-VIC | |
|---|---|---|---|---|---|---|---|---|---|---|
| OA | 0.570 | 0.570 | 0.477 | 0.555 | 0.595 | 0.630 | 0.651 | 0.608 | 0.648 | 0.645 |
| 0.245 | 0.245 | 0.099 | 0.000 | 0.300 | 0.243 | 0.328 | 0.262 | 0.296 | 0.287 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Polk, S.L.; Cui, K.; Chan, A.H.Y.; Coomes, D.A.; Plemmons, R.J.; Murphy, J.M. Unsupervised Diffusion and Volume Maximization-Based Clustering of Hyperspectral Images. Remote Sens. 2023, 15, 1053. https://doi.org/10.3390/rs15041053
Polk SL, Cui K, Chan AHY, Coomes DA, Plemmons RJ, Murphy JM. Unsupervised Diffusion and Volume Maximization-Based Clustering of Hyperspectral Images. Remote Sensing. 2023; 15(4):1053. https://doi.org/10.3390/rs15041053
Chicago/Turabian StylePolk, Sam L., Kangning Cui, Aland H. Y. Chan, David A. Coomes, Robert J. Plemmons, and James M. Murphy. 2023. "Unsupervised Diffusion and Volume Maximization-Based Clustering of Hyperspectral Images" Remote Sensing 15, no. 4: 1053. https://doi.org/10.3390/rs15041053
APA StylePolk, S. L., Cui, K., Chan, A. H. Y., Coomes, D. A., Plemmons, R. J., & Murphy, J. M. (2023). Unsupervised Diffusion and Volume Maximization-Based Clustering of Hyperspectral Images. Remote Sensing, 15(4), 1053. https://doi.org/10.3390/rs15041053








