Unmanned Aerial Vehicle (UAV)-Based Mapping of Acacia saligna Invasion in the Mediterranean Coast
Abstract
:1. Introduction
2. Materials and Methods
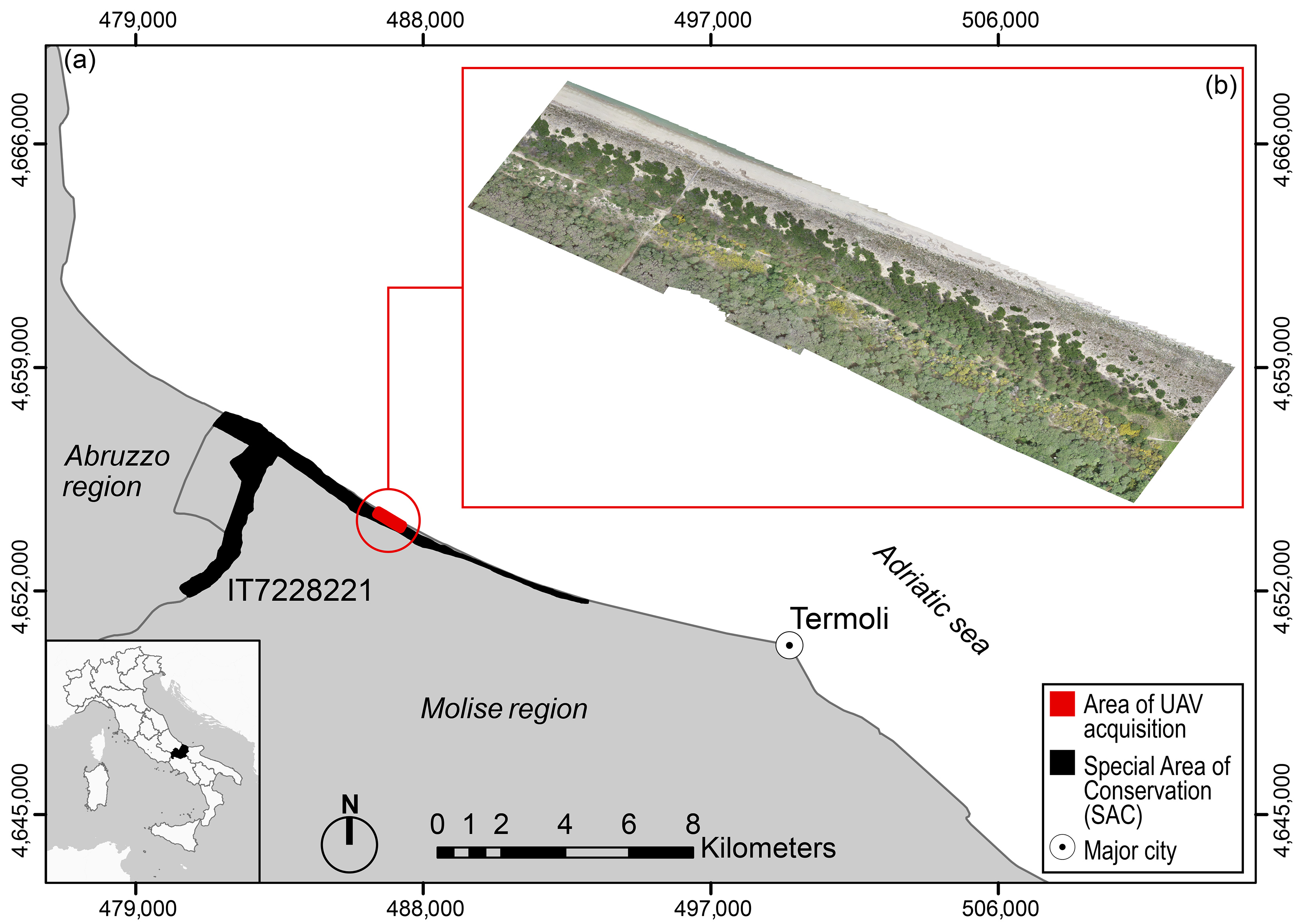
2.1. Study Area and Target Species
2.2. Data Collection and Analysis
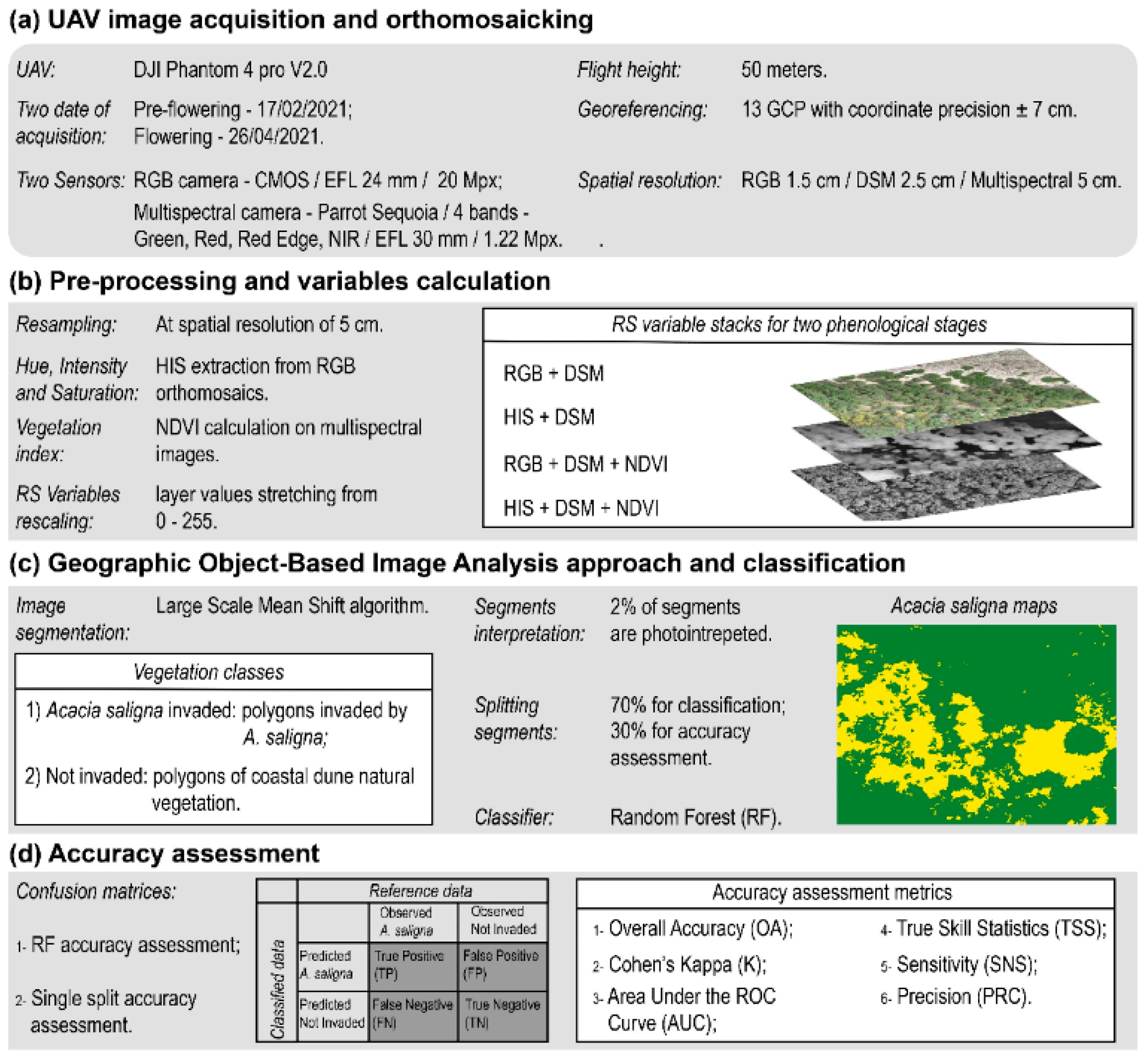
2.2.1. UAV Image Acquisition and Orthomosaicking
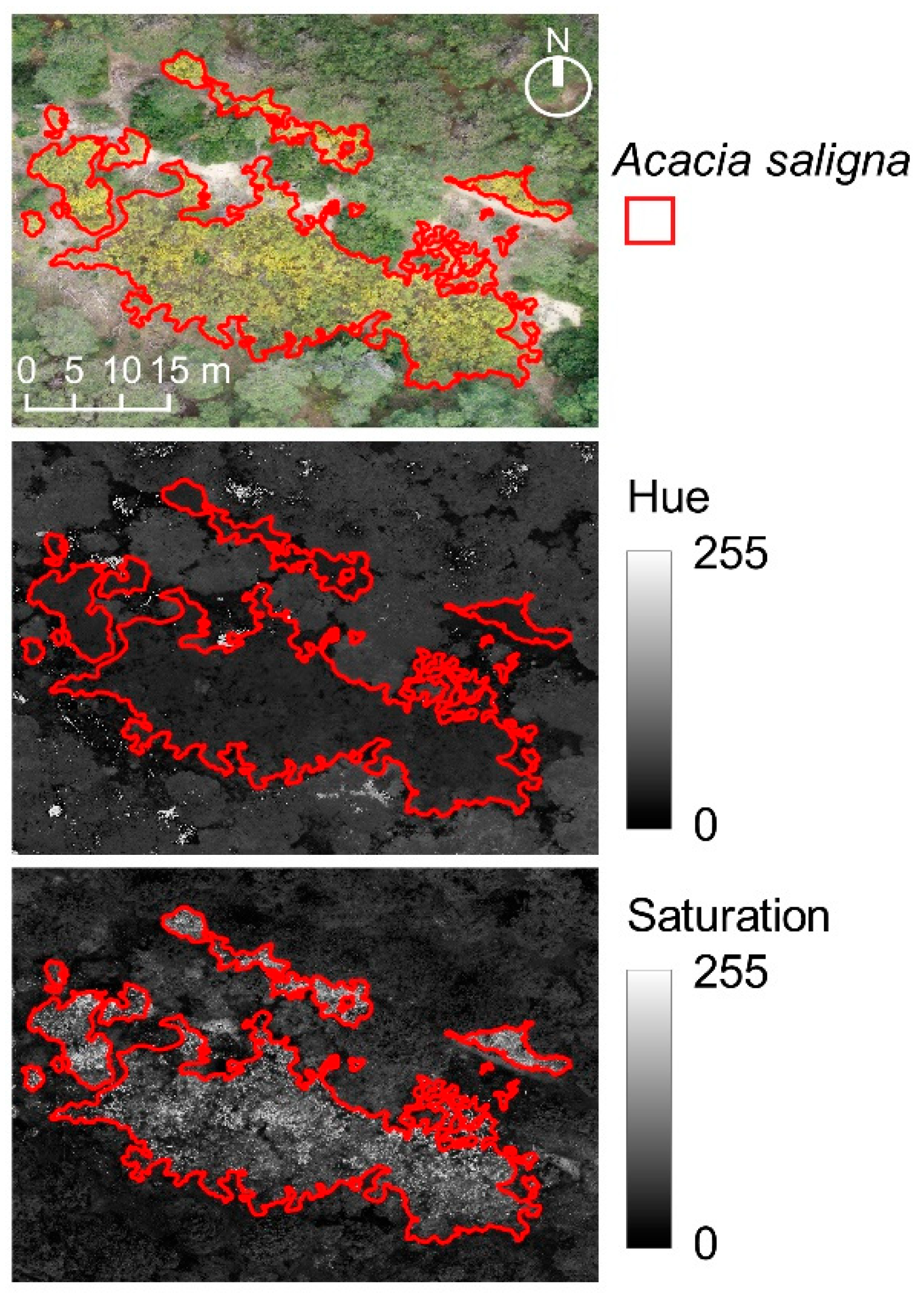
2.2.2. Pre-Processing and Variables Extraction
2.2.3. Geographic Object—Based Image Analysis and Classification
2.2.4. Accuracy Assessment
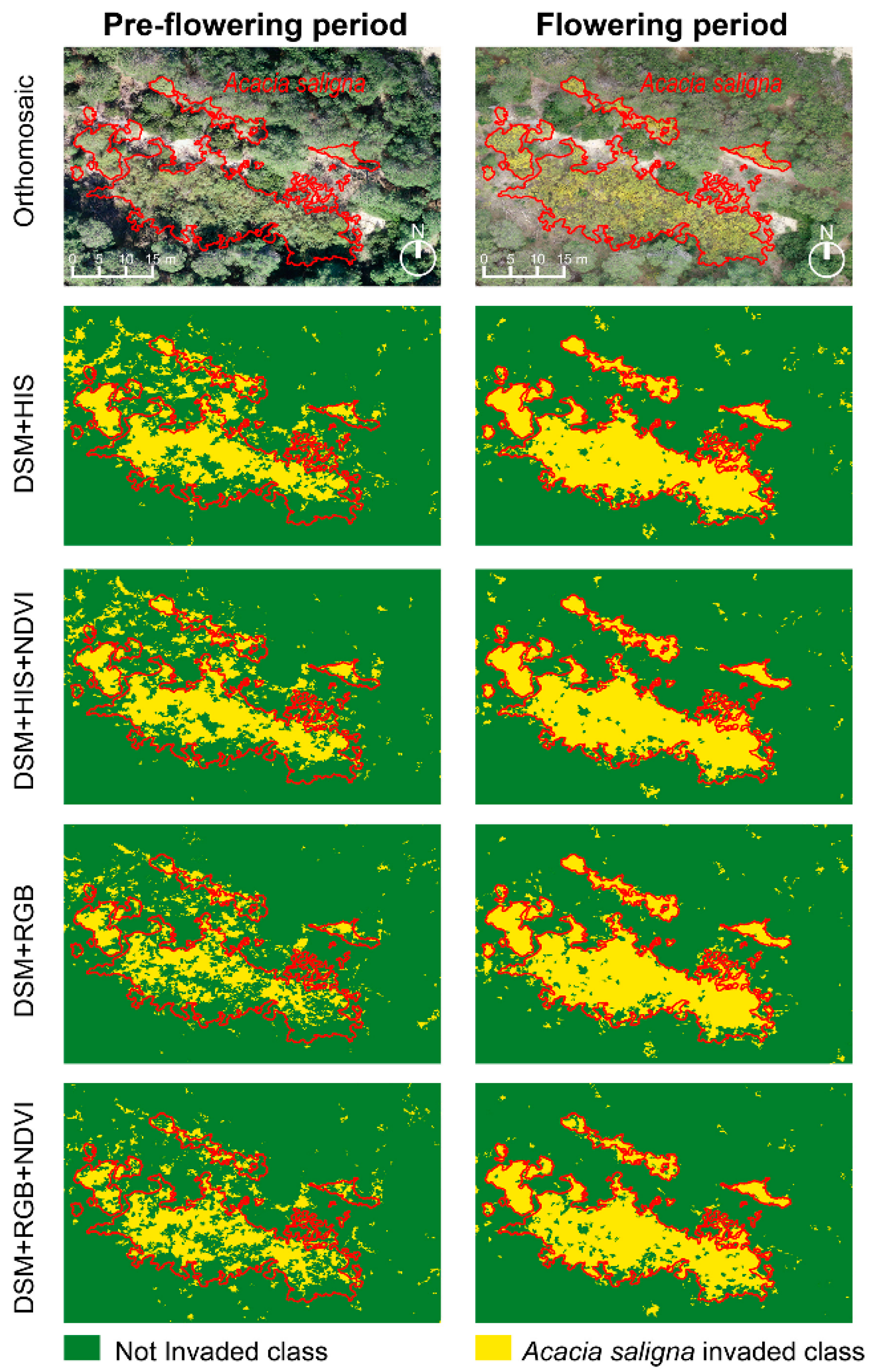
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Hulme, P.E. Trade, transport and trouble: Managing invasive species pathways in an era of globalization. J. Appl. Ecol. 2009, 46, 10–18. [Google Scholar] [CrossRef]
- Pyšek, P.; Richardson, D.M. Invasive species, environmental change and management, and health. Annu. Rev. Environ. Resour. 2010, 35, 25–55. [Google Scholar] [CrossRef] [Green Version]
- Simberloff, D.; Martin, J.L.; Genovesi, P.; Maris, V.; Wardle, D.A.; Aronson, J.; Courchamp, F.; Galil, B.; García-Berthou, E.; Pascal, M.; et al. Impacts of biological invasions: What’s what and the way forward. Trends Ecol. Evol. 2013, 28, 58–66. [Google Scholar] [CrossRef] [Green Version]
- Pyšek, P.; Hulme, P.E.; Simberloff, D.; Bacher, S.; Balckburn, T.M.; Carlton, J.T.; Dawson, W.; Essl, F.; Foxcroft, L.C.; Genovesi, P.; et al. Scientists’ warning on invasive alien species. Biol. Rev. 2020, 95, 1511–1534. [Google Scholar] [CrossRef]
- Lazzaro, L.; Bolpagni, R.; Buffa, G.; Gentili, R.; Lonati, M.; Stinca, A.; Acosta, A.T.R.; Adorni, M.; Aleffi, M.; Allegrezza, M.; et al. Impact of invasive alien plants on native plant communities and Natura 2000 habitats: State of the art, gap analysis and perspectives in Italy. J. Environ. Manag. 2020, 274, 111140. [Google Scholar] [CrossRef]
- Bartz, R.; Kowarik, I. Assessing the environmental impacts of invasive alien plants: A review of assessment approaches. NeoBiota 2019, 43, 69–99. [Google Scholar] [CrossRef] [Green Version]
- Jucker, T.; Carboni, M.; Acosta, A.T.R. Going beyond taxonomic diversity: Deconstructing biodiversity patterns reveals the true cost of iceplant invasion. Divers. Distrib. 2013, 19, 1566–1577. [Google Scholar] [CrossRef]
- Blackburn, T.M.; Essl, F.; Evans, T.; Hulme, P.E.; Jeschke, J.M.; Kühn, I.; Kumschick, S.; Marková, Z.; Mrugała, A.; Nentwig, W.; et al. A unified classification of alien species based on the magnitude of their environmental impacts. PLoS Biol. 2014, 12, e1001850. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loiola, P.P.; de Bello, F.; Chytrý, M.; Götzenberger, L.; Carmona, C.P.; Pyšek, P.; Lososová, Z. Invaders among locals: Alien species decrease phylogenetic and functional diversity while increasing dissimilarity among native community members. J. Ecol. 2018, 106, 2230–2241. [Google Scholar] [CrossRef]
- Tordoni, E.; Petruzzellis, F.; Nardini, A.; Savi, T.; Bacaro, G. Make it simpler: Alien species decrease functional diversity of coastal plant communities. J. Veg. Sci. 2019, 30, 498–509. [Google Scholar] [CrossRef]
- Gutiérrez, J.L. Modification of Habitat Quality by Non-native Species. In Impact of Biological Invasions on Ecosystem Services, 1st ed.; Vilà, M., Hulme, P., Eds.; Invading Nature-Springer Series in Invasion Ecology: Cham, Switzerland, 2017; Volume 12, pp. 33–47. [Google Scholar]
- Bonari, G.; Fantinato, E.; Lazzaro, L.; Sperandii, M.G.; Acosta, A.T.R.; Allegrezza, M.; Assini, S.; Caccianiga, M.; Di Cecco, V.; Frattaroli, A.; et al. Shedding light on typical species: Implications for habitat monitoring. Plant Sociol. 2021, 58, 157–166. [Google Scholar] [CrossRef]
- Rai, P.K.; Singh, J.S. Invasive alien plant species: Their impact on environment, ecosystem services and human health. Ecol. Indic. 2020, 111, 106020. [Google Scholar] [CrossRef]
- Jones, B.A. Tree shade, temperature, and human health: Evidence from invasive species-induced deforestation. Ecol. Econ. 2019, 156, 12–13. [Google Scholar] [CrossRef]
- European Commission. Invasive Alien Species. A European Response, 1st ed.; European Union: Brussels, Belgium, 2014; pp. 1–28. [Google Scholar]
- Branquart, E.; Brundu, G.; Buholzer, S.; Chapman, D.; Ehret, P.; Fried, G.; Starfinger, U.; van Valkenburg, J.; Tanner, R. A prioritization process for invasive alien plant species incorporating the requirements of EU Regulation no. 1143/2014. Bull. OEPP/EPPO Bull. 2016, 46, 603–617. [Google Scholar] [CrossRef] [Green Version]
- Sitzia, T.; Campagnaro, T.; Kowarik, I.; Trentanovi, G. Using forest management to control invasive alien species: Helping implement the new European regulation on invasive alien species. Biol. Invasions 2016, 18, 1–7. [Google Scholar] [CrossRef]
- Lozano, V.; Marzialetti, F.; Carranza, M.L.; Chapman, D.; Branquart, E.; Dološ, K.; Große-Stoltenberg, A.; Fiori, M.; Capece, P.; Brundu, G. Modelling Acacia saligna invasion in a large Mediterranean island using PAB factors: A tool for implementing the European legislation on invasive species. Ecol. Indic. 2020, 116, 106516. [Google Scholar] [CrossRef]
- Vaz, A.S.; Alcaraz-Segura, D.; Campos, J.C.; Vicente, J.R.; Honrado, J.P. Managing plant invasions through the lens of remote sensing: A review of progress and the way forward. Sci. Total Environ. 2018, 642, 1328–1339. [Google Scholar] [CrossRef]
- Müllerová, J.; Pergl, J.; Pyšek, P. Remote sensing as a tool for monitoring plant invasions: Testing the effects of data resolution and image classification approach on the detection of a model plant species Heracleum mantegazzianum (giant hogweed). Int. J. Appl. Earth Obs. 2013, 25, 55–65. [Google Scholar] [CrossRef]
- Niphadkar, M.; Nagendra, H. Remote sensing of invasive plants: Incorporating functional traits into the picture. Int. J. Remote Sens. 2016, 37, 3074–3085. [Google Scholar] [CrossRef]
- Royimani, L.; Mutanga, O.; Odindi, J.; Dube, T.; Matongera, T.N. Advancements in satellite remote sensing for mapping and monitoring of alien invasive plant species (AIPs). Phys. Chem. Earth 2019, 112, 237–245. [Google Scholar] [CrossRef]
- Peerbhay, K.; Mutanga, O.; Ismail, R. The identification and remote detection of alien invasive plants in commercial forests: An overview. S. Afr. J. Geomat. 2016, 5, 49–67. [Google Scholar] [CrossRef] [Green Version]
- Kattenborn, T.; Lopatin, J.; Förster, M.; Braun, A.C.; Fassnacht, F.E. UAV data as alternative to field sampling to map woody invasive species based on combined Sentinel-1 and Sentinel-2 data. Remote Sens. Environ. 2019, 227, 61–73. [Google Scholar] [CrossRef]
- Huang, C.; Asner, G.P. Applications of remote sensing to alien invasive plant studies. Sensors 2009, 9, 4869–4889. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bradley, B.A. Remote detection of invasive plants: A review of spectral, textural and phenological approaches. Biol. Invasions 2014, 16, 1411–1425. [Google Scholar] [CrossRef]
- Pettorelli, N.; Laurance, W.F.; O’Brien, T.G.; Wegmann, M.; Nagendra, H.; Turner, W. Satellite remote sensing for applied ecologists: Opportunities and challenges. J. Appl. Ecol. 2014, 51, 839–848. [Google Scholar] [CrossRef]
- Wang, X.; Wang, L.; Tian, J.; Shi, C. Object-based spectral-phenological features for mapping invasive Spartina alterniflora. Int. J. Appl. Earth Obs. 2021, 101, 102349. [Google Scholar] [CrossRef]
- Nowak, M.M.; Dziób, K.; Bogawski, P. Unmanned aerial vehicles (UAVs) in environmental biology: A review. Eur. J. Ecol. 2018, 4, 56–74. [Google Scholar] [CrossRef]
- Michez, A.; Piégay, H.; Jonathan, L.; Claessens, H.; Lejeune, P. Mapping of riparian invasive species with supervised classification of Unmanned Aerials System (UAS) imagery. Int. J. Appl. Earth Obs. 2016, 44, 88–94. [Google Scholar] [CrossRef]
- Müllerová, J.; Brůna, J.; Bartaloš, T.; Dvořák, P.; Vítková, M.; Pyšek, P. Timing is important: Unmanned aircraft vs. satellite imagery in plant invasion monitoring. Front. Plant Sci. 2017, 8, 887. [Google Scholar] [CrossRef] [Green Version]
- Bolch, E.A.; Santos, M.J.; Ade, C.; Khanna, S.; Basinger, N.T.; Reader, M.O.; Hestir, E.L. Remote detection of invasive alien species. In Remote Sensing of Plant Biodiversity, 1st ed.; Cavender-Bares, J., Gamon, J.A., Townsend, P.A., Eds.; Springer Nature: Cham, Switzerland, 2020; Volume 12, pp. 267–304. [Google Scholar]
- Maslin, B.R. Studies in the genus Acacia-3. The taxonomy of A. salgina (Labill.) H. Wendl. Nuytsia 1974, 1, 332–340. [Google Scholar]
- Thompson, G.D.; Bellstedt, D.U.; Richardson, D.M.; Wilson, J.R.U.; Le Roux, J.J. A tree well travelled: Global genetic structure of the invasive tree Acacia saligna. J. Biogeogr. 2015, 42, 305–314. [Google Scholar] [CrossRef]
- Asefa, G.; Tamir, B. Effects of supplementing different forms of Acacia saligna leaves to grass hay on feed intake and growths of lambs. Trop Sci. 2006, 46, 205–208. [Google Scholar] [CrossRef]
- George, N.; Henry, D.; Yan, G.; Byrne, M. Variability in feed quality between populations of Acacia saligna (Labill.) H. Wendl. (Mimosoideae)—Implications for domestication. Anim. Feed Sci. Technol. 2007, 136, 109–127. [Google Scholar] [CrossRef]
- Avis, A.M. A review of coastal dune stabilization in the Cape province of South Africa. Landsc. Urban Plan. 1989, 18, 55–68. [Google Scholar] [CrossRef]
- Bar, P.; Cohen, O.; Shoshany, M. Invasion rate of the alien species Acacia saligna within coastal sand dune habitats in Israel. Israel J. Plant Sci. 2004, 52, 115–124. [Google Scholar] [CrossRef]
- Donaldson, J.E.; Hui, C.; Richardson, D.M.; Robertson, M.P.; Webber, B.L.; Wilson, J.R.H. Invasion trajectory of alien trees: The role of introduction pathway and planting history. Glob. Change Biol. 2014, 20, 1527–1537. [Google Scholar] [CrossRef]
- Marzialetti, F.; Bazzichetto, M.; Giulio, S.; Acosta, A.T.R.; Stanisci, A.; Malavasi, M.; Carranza, M.L. Modelling Acacia saligna invasion on the Adriatic coastal landscape: An integrative approach using LTER data. Nat. Conserv. Bulg. 2019, 34, 127–144. [Google Scholar] [CrossRef]
- Lehrer, D.; Becker, N.; Bar, P. The economic impact of the invasion of Acacia saligna in Israel. Int. J. Sustain. Dev. World Ecol. 2011, 18, 118–127. [Google Scholar] [CrossRef]
- Del Vecchio, S.; Acosta, A.T.R.; Stanisci, A. The impact of Acacia saligna invasion on Italian coastal dune EC habitats. CR Biol. 2013, 336, 364–369. [Google Scholar] [CrossRef]
- De Sá, N.C.; Castro, P.; Carvalho, S.; Marchante, E.; López-Núñez, F.A.; Marchante, H. Mapping the flowering of an invasive plant using Unmanned Aerial Vehicles: Is there potential for biocontrol monitoring? Front. Plant Sci. 2018, 9, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Paz-Kagan, T.; Silver, M.; Panov, N.; Karnieli, A. Multispectral approach for identifying invasive plant species based on flowering phenology characteristics. Remote Sens. 2019, 11, 953. [Google Scholar] [CrossRef] [Green Version]
- Strydom, M.; Esler, K.J.; Wood, A.R. Acacia saligna seed banks: Sampling methods and dynamics, Western Cape, South Africa. S. Afr. J. Bot. 2012, 79, 140–147. [Google Scholar] [CrossRef]
- Witkowski, E.T.F. Growth of seedlings of the invasive, Acacia saligna and Acacia Cyclops, in relation to soil phosphorus. Aust. J. Ecol. 1994, 19, 290–296. [Google Scholar] [CrossRef]
- Gaol, M.L.; Fox, J.E.D. Reproductive potential of Acacia species in the central wheatbelt: Variation between years. Conserv. Sci. West. Aust. 2002, 4, 147–157. [Google Scholar]
- Darcha, G.; Brhan, A.; Gebrewahid, Y.; Kassa, H.; Mezgebe, K.; Cunningham, P.; Hagazi, N.; Girmay, A.; Birhane, E.; Kelly, R.; et al. Population structure of Acacia saligna in rehabilitated exclosure, northern Ethiopia, Tigray. J. Drylands 2019, 9 (Suppl. 1), 877–884. [Google Scholar]
- Masemola, C.; Cho, M.A.; Ramoelo, A. Sentinel-2 time series based optimal features and time window for mapping invasive Australian native species in KwaZulu Natal, South Africa. Int. J. Appl. Earth Obs. 2020, 93, 102207. [Google Scholar] [CrossRef]
- Paz-Kagan, T.; Caras, T.; Herrmann, I.; Shachak, M.; Karnieli, A. Multiscale mapping of species diversity under changed land use using imaging spectroscopy. Ecol. Appl. 2017, 27, 1466–1484. [Google Scholar] [CrossRef] [PubMed]
- Oldeland, J.; Große-Stoltenberg, A.; Naftal, L.; Strohbach, B.J. The potential of UAV derived image feauters for discriminating Savanaah tree species. In The Roles of Remote Sensing in Nature Conservation. A Pratical Guide and Case Studies, 1st ed.; Díaz-Delgado, R., Lucas, R., Hurford, C., Eds.; Springer Nature: Cham, Switzerland, 2017; pp. 183–202. [Google Scholar]
- Lopatin, J.; Dolos, K.; Kattenborn, T.; Fassnacht, F.E. How canopy shadow affects invasive plant species classification in high spatial resolution remote sensing. Remote Sens. Ecol. Conserv. 2019, 5, 302–317. [Google Scholar] [CrossRef]
- Acosta, A.T.R.; Carranza, M.L.; Izzi, C.F. Are there habitats that contribute best to plant species diversity in coastal dunes? Biodivers. Conserv. 2009, 18, 1087–1098. [Google Scholar] [CrossRef]
- Carranza, M.L.; Acosta, A.T.R.; Stanisci, A.; Pirone, G.; Ciaschetti, G. Ecosystem classification for EU habitat distribution assessment in sandy coastal environments: An application in central Italy. Environ. Monit. Assess. 2008, 140, 99–107. [Google Scholar] [CrossRef]
- Stanisci, A.; Acosta, A.T.R.; Carranza, M.L.; de Chiro, M.; Del Vecchio, S.; Di Martino, L.; Frattaroli, A.R.; Fusco, S.; Izzi, C.F.; Pirone, G.; et al. EU habitats monitoring along the coastal dunes of the LTER sites of Abruzzo and Molise (Italy). Plant Sociol. 2014, 51, 51–56. [Google Scholar] [CrossRef]
- Drius, M.; Malavasi, M.; Acosta, A.T.R.; Ricotta, C.; Carranza, M.L. Boundary-based analysis for the assessment of coastal dune landscape integrity over time. Appl. Geogr. 2013, 45, 41–48. [Google Scholar] [CrossRef]
- Malavasi, M.; Barták, V.; Jucker, T.; Acosta, A.T.R.; Carranza, M.L.; Bazzichetto, M. Strenght in numbers: Combining multi-source remotely sensed data to model plant invasions in coastal dune ecosystems. Remote Sens. 2019, 11, 275. [Google Scholar] [CrossRef] [Green Version]
- Acosta, A.T.R.; Ercole, S.; Stanisci, A.; De Patta Pillar, V.; Blasi, C. Coastal vegetation zonation and dune morphology in some Mediterranean ecosystems. J. Coast. Res. 2007, 23, 1518–1524. [Google Scholar] [CrossRef] [Green Version]
- Marzialetti, F.; Di Febbraro, M.; Malavasi, M.; Giulio, S.; Acosta, A.T.R.; Carranza, M.L. Mapping coastal dune landscape through spectral Rao’s Q temporal diversity. Remote Sens. 2020, 12, 2315. [Google Scholar] [CrossRef]
- Izzi, C.F.; Acosta, A.T.R.; Carranza, M.L.; Ciaschetti, G.; Conti, F.; Di Martino, L.; D’Orazio, G.; Frattaroli, A.; Pirone, G.; Stanisci, A. Sampling the vascular flora in coastal dune ecosystems of Central Italy. Fitosociologia 2007, 44, 129–137. [Google Scholar]
- Turner, D.; Lucieer, A.; Watson, C. An automated technique for generating georectified mosaics from ultra-high resolution Unmanned Aerial Vehicle (UAV) imagery, based on structure from motion (SfM) point clouds. Remote Sens. 2012, 4, 1392–1410. [Google Scholar] [CrossRef] [Green Version]
- Mancini, F.; Dubbini, M.; Gattelli, M.; Stecchi, F.; Fabbri, S.; Gabbianelli, G. Using Unmanned Aerial Vehicles (UAV) for high-resolution reconstruction of topography: The structure from motion approach in coastal environments. Remote Sens. 2013, 5, 6880–6898. [Google Scholar] [CrossRef] [Green Version]
- Moreno, J.F.; Melia, J. An optimum interpolation method applied to the resampling of NOAA AVHRR data. IEEE T. Geosci. Remote 1994, 32, 131–151. [Google Scholar] [CrossRef]
- Laliberte, A.S.; Rango, A. Incorporation of texture, intensity, hue, and saturation for rangeland monitoring with unmanned aircraft imagery 2008. In Proceedings of the International Archives of the Photogrammetry, Remote Sensing, and Spatial Information Sciences, GEOBIA Proceedings, Calgary, AB, Canada, 5–8 August 2008; Hay, G.J., Blaschke, T., Marceau, D., Eds.; ISPRS: Calgary, AB, Canada; Volume XXXVIII-4/C1. [Google Scholar]
- Xie, Y.; Sha, Z.; Yu, M. Remote sensing imagery in vegetation mapping: A review. J. Plant Ecol. 2008, 1, 9–23. [Google Scholar] [CrossRef]
- Ghassemian, H. A review of remote sensing image fusion methods. Inform. Fusion 2016, 32, 75–89. [Google Scholar] [CrossRef]
- GRASS Development Team. Geographic Resources Analysis Support System (GRASS) Software, Version 7.8. Open Source Geospatial Foundation. 2020. Available online: http://grass.osgeo.org (accessed on 24 April 2020).
- Fortner, B.; Meyer, T.E. Number by Colors. A Guide to Using Color to Understand Technical Data, 1st ed.; Springer Science: New York, NY, USA, 1997; pp. 63–84. [Google Scholar]
- Koutsias, N.; Karteris, M.; Chuvieco, E. The use of Intensity-Hue-Saturation transformation of Landsat-5 thematic mapper data for burned land mapping. Photogramm. Eng. Rem. S. 2000, 66, 829–839. [Google Scholar]
- Tu, T.-M.; Lee, Y.-C.; Chang, C.-P.; Huang, P.S. Adjustable intensity-hue-saturation and Brovey transform fusion technique for IKONOS/ QuickBird imagery. Opt. Eng. 2005, 44, 116201. [Google Scholar] [CrossRef] [Green Version]
- Tucker, C.J. Red and photographic infrared linear combinations of monitoring vegetation. Remote Sens. Environ. 1979, 8, 127–150. [Google Scholar] [CrossRef] [Green Version]
- Moore, G.K.; Waltz, F.A. Objective procedure for lineament enhancement and extraction. Photogramm. Eng. Rem. S. 1983, 49, 641–647. [Google Scholar]
- Immitzer, M.; Vuolo, F.; Atzberger, C. First experience with Sentinel-2 data for crop and tree species classifications in Central Europe. Remote Sens. 2016, 8, 166. [Google Scholar] [CrossRef]
- Blaschke, T. Object based image analysis for remote sensing. ISPRS J. Photogramm. 2010, 65, 2–16. [Google Scholar] [CrossRef] [Green Version]
- Blaschke, T.; Hay, G.J.; Kelly, M.; Lang, S.; Hofmann, P.; Addink, E.; Feitosa, R.Q.; van der Meer, F.; van der Werff, H.; van Coillie, F.; et al. Geographic Object-Based Image Analysis—Towards a new paradigm. ISPRS J. Photogramm. 2014, 87, 180–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, B.A.; Ma, L. Image Segmentation and Object-Based Image Analysis for Environmental Monitoring: Recent Areas of Interest, Researchers’ Views on the Future Priorities. Remote Sens. 2020, 12, 1772. [Google Scholar] [CrossRef]
- Chen, G.; Weng, Q.; Hay, G.J.; He, Y. Geographic object-based image analysis (GEOBIA) emerging trends and future opportunities. GISci. Remote Sens. 2018, 55, 159–182. [Google Scholar] [CrossRef]
- Hossain, M.D.; Chen, D. Segmentation for object-based image analysis (OBIA): A review of algorithms and challenges from remote sensing perspective. ISPRS J. Photogramm. 2019, 150, 115–134. [Google Scholar] [CrossRef]
- Michel, J.; Youssefi, D.; Grizonnet, M. Stable Mean-Shift algorithm and its application to the segmentation of arbitrarily large remote sensing images. IEEE Trans. Geosci. Remote 2015, 53, 952–964. [Google Scholar] [CrossRef]
- De Luca, G.; Silva, J.M.N.; Cerasoli, S.; Arújo, J.; Campos, J.; Di Fazio, S.; Modica, G. Object-based land cover classification of Cork Oak Woodlands using UAV imagery and Orfeo Toolbox. Remote Sens. 2019, 11, 1238. [Google Scholar] [CrossRef] [Green Version]
- Comaniciu, D.; Meer, P. Mean shift: A robust approach toward feature space analysis. IEEE Trans. Pattern Anal. 2002, 25, 603–619. [Google Scholar] [CrossRef] [Green Version]
- Millard, K.; Richardson, M. On the importance of training data sample selection in random forest image classification: A case study in Peatland ecosystem mapping. Remote Sens. 2015, 7, 8489–8515. [Google Scholar] [CrossRef] [Green Version]
- Baraldi, A.; Bruzzon, L.; Blonda, P. Quality assessment of classification and cluster maps without ground truth knowledge. IEEE Trans. Geosci. Remote 2005, 43, 857–873. [Google Scholar] [CrossRef]
- Powers, R.P.; Hay, G.J.; Chen, G. How wetland type and area differ through scale: A GEOBIA case study in Alberta’s Boreal Plains. Remote Sens. Environ. 2012, 117, 135–145. [Google Scholar] [CrossRef]
- Shahabi, H.; Jarihani, B.; Piralilou, S.T.; Chittleborough, D.; Avand, M.; Ghorbanzadeh, O. A semi-automated object-base gully networks detection using different machine learning models: A case study of Bowen Catchment, Queensland, Australia. Sensors 2019, 19, 4893. [Google Scholar] [CrossRef] [Green Version]
- Breiman, L. Random Forest. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Cutler, D.R.; Edwards, T.C.; Beard, K.H.; Cutler, A.; Hess, K.T.; Gibson, J.; Lawler, J.J. Random forests for classification in ecology. Ecology 2007, 88, 2783–2792. [Google Scholar] [CrossRef]
- Gislason, P.O.; Benediktsson, J.A.; Sveinsson, J.R. Random Forest for land cover classification. Pattern Recogn. Lett. 2006, 27, 294–300. [Google Scholar] [CrossRef]
- Pal, M. Random forest classifier for remote sensing classification. Int. J. Remote Sens. 2005, 26, 217–222. [Google Scholar] [CrossRef]
- Juel, A.; Groom, G.B.; Svenning, J.-C. Ejrnæs Spatial application of random forest models for fine-scale coastal vegetation classification using object based analysis of aerial orthophoto and DEM data. Int. J. Appl. Earth Obs. 2015, 42, 106–114. [Google Scholar] [CrossRef]
- Wessel, M.; Brandmeier, M.; Tiede, D. Evaluation of different machine learning algorithms for scalable classification of tree types and tree species based on Sentinel-2 data. Remote Sens. 2018, 10, 1419. [Google Scholar] [CrossRef] [Green Version]
- Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 2008, 28, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Routh, D.; Seegmiller, L.; Bettigole, C.; Khun, C.; Oliver, C.D.; Glick, H.B. Improving the reliability of mixture tuned matched filtering remote sensing classification results using supervised learning algorithms and cross-validation. Remote Sens. 2018, 10, 1675. [Google Scholar] [CrossRef] [Green Version]
- Belgiu, M.; Drăgut, L. Random forest in remote sensing: A review of applications and future directions. ISPRS J. Photogramm. 2016, 114, 24–31. [Google Scholar] [CrossRef]
- Grabska, E.; Hostert, P.; Pflugmacher, D.; Ostapowicz, K. Forest stand species mapping using the Sentinel-2 time series. Remote Sens. 2019, 11, 1197. [Google Scholar] [CrossRef] [Green Version]
- Calle, M.L.; Urrea, V. Letter to the editor: Stability of Random Forest importance measures. Brief. Bioinform. 2011, 12, 86–89. [Google Scholar] [CrossRef] [Green Version]
- Mellor, A.; Haywood, A.; Stone, C.; Jones, S. The performance of Random Forest in an operational setting for large area sclerophyll forest classification. Remote Sens. 2013, 5, 2838–2856. [Google Scholar] [CrossRef] [Green Version]
- Lyons, M.B.; Keith, D.A.; Phinn, S.R.; Mason, T.J.; Elith, J. A comparison of resampling methods for remote sensing classification and accuracy assessment. Remote Sens. Environ. 2018, 208, 145–153. [Google Scholar] [CrossRef]
- Ye, S.; Pontius, R.G., Jr.; Rakshit, R. A review of accuracy assessment for object-based image analysis: From per-pixel to per-polygon approaches. ISPRS J. Photogramm. 2018, 14, 137–147. [Google Scholar] [CrossRef]
- Barsi, Á.; Kugler, Z.; László, G.; Abdulmutalib, H.M. Accuracy dimension in Remote Sensing. The International Archives of the Photogrammetry, Remote Sensing and Spatial Information Sciences 2018. Volume XLII-3, pp. 61–67. Available online: https://pdfs.semanticscholar.org/f7d8/b9fece4b5e42a92f8c994d5c198e70fd7118.pdf (accessed on 24 April 2020).
- Congalton, R.G.; Green, K. Assessing the Accuracy of Remotely Sensed Data. Principles and Practices, 2nd ed.; Taylor & Francis Group: Boca Raton, FL, USA, 2009; pp. 1–210. [Google Scholar]
- Landis, J.R.; Koch, G.G. The Measurement of Observer Agreement for Categorical Data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DeLong, E.R.; DeLong, D.M.; Clarke-Pearson, D.L. Comparing the areas under two or more correlated receiver operating characteristic curves: A nonparametric approach. Biometrics 1988, 44, 837–845. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Valverde, A. Insights into the area under the receiver operating characteristic curve (AUC) as a discrimination measure in species distribution modelling. Glob. Ecol. Biogeogr. 2012, 21, 498–507. [Google Scholar] [CrossRef]
- Allouche, O.; Tsoar, A.; Kadmon, R. Assessing the accuracy of species distribution models: Prevalence, kappa and the true skill statistic (TSS). J. Appl. Ecol. 2006, 43, 1223–1232. [Google Scholar] [CrossRef]
- Sofaer, H.R.; Hoeting, J.A.; Jarnevich, C.S. The area under the precision-recall curve as a performance metric for rare binary events. Methods Ecol. Evol. 2019, 10, 565–577. [Google Scholar] [CrossRef]
- Bekkar, M.; Djemaa, H.K.; Alitouche, T.A. Evaluation measures for models assessment over imbalanced data sets. J. Inf. Eng. Appl. 2013, 3, 27–39. [Google Scholar]
- Hanczar, B.; Hua, J.; Sima, C.; Weinstein, J.; Bittner, M.; Dougherty, E. Small-sample precision of ROC-related estimates. Bionformatics 2010, 26, 822–830. [Google Scholar] [CrossRef] [Green Version]
- Vasilakos, C.; Kavroudakis, D.; Georganta, A. Machine learning classification ensemble of multitemporal Sentinel-2 images: The case of a mixed Mediterranean ecosystem. Remote Sens. 2020, 12, 2005. [Google Scholar] [CrossRef]
- Abeysinghe, T.; Milas, A.S.; Arend, K.; Hohman, B.; Reil, P.; Gregory, A.; Vázquez-Ortega, A. Mapping invasive Phragmites australis in the Old Woman Creek estuary using UAV remote sensing and machine learning classifiers. Remote Sens. 2019, 11, 1380. [Google Scholar] [CrossRef] [Green Version]
- Dash, J.P.; Watt, M.S.; Paul, T.S.H.; Morgenroth, J.; Pearse, G.D. Early detection of invasive exotic trees using UAV and manned aircraft multispectral and LiDAR data. Remote Sens. 2019, 11, 1812. [Google Scholar] [CrossRef] [Green Version]
- Shao, G.; Tang, L.; Liao, J. Overselling overall map accuracy misinforms about research reliability. Landsc. Ecol. 2019, 34, 2487–2492. [Google Scholar] [CrossRef] [Green Version]
- Lobo, J.M.; Jiménez-Valverde, A.; Real, R. AUC: A misleading measure of the performance of predictive distribution models. Glob. Ecol. Biogeogr. 2008, 17, 145–151. [Google Scholar] [CrossRef]
- Martins, F.; Alegria, C.; Gil, A. Mapping invasive alien Acacia dealbata Link using ASTER multispectral imagery: A case study in central-eastern of Portugal. For. Syst. 2016, 25, e078. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.; Smith, A.M.; Bourchier, R.S.; Hodge, K.; Ostrander, D. Flowering leafy spurge (Euphorbia esula) detection using unmanned aerial vehicle imagery in biological control sites: Impacts of flight height, flight time and detection method. Weed Technol. 2020, 34, 575–588. [Google Scholar] [CrossRef]
- Hill, D.J.; Tarasoff, C.; Whitworth, G.E.; Baron, J.; Bradshaw, J.L.; Church, J.S. Utility of unmanned aerial vehicles for mapping invasive plant species: A case study on yellow flag iris (Iris pseudacorus L.). Int. J. Remote Sens. 2017, 38, 2083–2105. [Google Scholar] [CrossRef]
- Samiappan, S.; Turnage, G.; Hathcock, L.A.; Moorhead, R. Mapping of invasive phragmites (common reed) in Gulf of Mexico coastal wetlands using multispectral imagery and small unmanned aerial systems. Int. J. Remote Sens. 2017, 38, 2861–2882. [Google Scholar] [CrossRef]
- Millar, M.A.; Byrne, M. Biogeographic origins and reproductive mode of naturalized populations of Acacia saligna. Aust. J. Bot. 2012, 60, 383–395. [Google Scholar] [CrossRef]
- Dorigo, W.; Lucieer, A.; Podobnikar, T.; Čarni, A. Mapping invasive Fallopia japonica by combined spectral, spatial, and temporal analysis of digital orthophotos. Int. J. Appl. Earth Obs. 2012, 19, 185–195. [Google Scholar] [CrossRef]
- Onishi, M.; Ise, T. Explainable identification and mapping of trees using UAV RGB image and deep learning. Sci. Rep. 2021, 11, 903. [Google Scholar] [CrossRef]
- Lehmann, J.R.K.; Prinz, T.; Ziller, S.R.; Thiele, J.; Heringer, G.; Meira-Neto, J.A.A.; Buttschardt, T.K. Open-source processing and analysis of aerial imagery acquired with a low-cost unmanned aerial system to support invasive plant management. Front. Environ. Sci. 2017, 5, 44. [Google Scholar] [CrossRef] [Green Version]
- Müllerová, J.; Bartaloš, T.; Brůna, J.; Dvořak, P.; Vítková, M. Unmanned aircraft in nature conservation: An example from plant invasion. Int. J. Remote Sens. 2017, 38, 2177–2198. [Google Scholar] [CrossRef]
- Baxter, P.W.J.; Hamilton, G. Learning to fly: Integrating spatial ecology with unmanned aerial vehicle surveys. Ecosphere 2018, 9, e02194. [Google Scholar] [CrossRef]
- Alvarez-Taboada, F.; Paredes, C.; Julián-Pelaz, J. Mapping of the invasive species Hakea sericea using Unmanned Aerial Vehicle (UAV) and WorldView-2 imagery and an object-oriented approach. Remote Sens. 2017, 9, 913. [Google Scholar] [CrossRef] [Green Version]


| Acronym | Name | Formula |
|---|---|---|
| OA | Overall Accuracy | |
| K | Cohen’s Kappa | |
| AUC | Area Under the ROC Curve | |
| TSS | True Skill Statistics | |
| SNS | Sensitivity | |
| PRC | Precision |
| Pre—Flowering Period | |||||||
| Raster Stack | AA | OA (%) | K | AUC | TSS | SNS (%) | PRC (%) |
| DSM + HIS | RF | 93.37 ± 0.67 | 0.57 ± 0.06 | 0.90 ± 0.02 | 0.48 ± 0.09 | 52.49 ± 8.82 | 76.55 ± 6.64 |
| SS | 93.61 | 0.62 | 0.77 | 0.54 | 56.18 | 78.12 | |
| DSM + RGB | RF | 91.17 ± 0.80 | 0.38 ± 0.07 | 0.84 ± 0.01 | 0.30 ± 0.09 | 31.09 ± 8.58 | 65.42 ± 7.57 |
| SS | 94.42 | 0.66 | 0.80 | 0.60 | 61.45 | 78.46 | |
| DSM + HIS + NDVI | RF | 93.05 ± 1.06 | 0.55 ± 0.11 | 0.90 ± 0.02 | 0.47 ± 0.12 | 49.01 ± 11.71 | 74.04 ± 6.35 |
| SS | 94.33 | 0.65 | 0.79 | 0.58 | 60 | 78.13 | |
| DSM + RGB + NDVI | RF | 91.20 ± 0.79 | 0.39 ± 0.07 | 0.87 ± 0.02 | 0.31 ± 0.09 | 30.96 ± 9.37 | 69.02 ± 11.21 |
| SS | 94.28 | 0.68 | 0.82 | 0.64 | 66.67 | 76.32 | |
| Flowering Period | |||||||
| Raster Stack | AA | OA (%) | K | AUC | TSS | SNS (%) | PRC (%) |
| DSM + HIS | RF | 94.92 ± 0.60 | 0.73 ± 0.04 | 0.95 ± 0.01 | 0.65 ± 0.07 | 65.77 ± 8.27 | 88.73 ± 4.33 |
| SS | 96.61 | 0.82 | 0.88 | 0.75 | 75.76 | 94.94 | |
| DSM + RGB | RF | 93.35 ± 0.49 | 0.62 ± 0.04 | 0.91 ± 0.01 | 0.51 ± 0.05 | 51.51 ± 5.19 | 88.57 ± 4.52 |
| SS | 95.60 | 0.76 | 0.83 | 0.66 | 66.67 | 95.65 | |
| DSM + HIS + NDVI | RF | 94.92 ± 0.92 | 0.75 ± 0.05 | 0.95 ± 0.01 | 0.67 ± 0.08 | 67.95 ± 8.70 | 88.96 ± 5.42 |
| SS | 95.75 | 0.80 | 0.87 | 0.74 | 74.75 | 94.94 | |
| DSM + RGB + NDVI | RF | 92.56 ± 1.21 | 0.61 ± 0.08 | 0.91 ± 0.02 | 0.51 ± 0.10 | 51.29 ± 10.16 | 89.36 ± 10.76 |
| SS | 95.87 | 0.80 | 0.86 | 0.72 | 72.45 | 94.67 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marzialetti, F.; Frate, L.; De Simone, W.; Frattaroli, A.R.; Acosta, A.T.R.; Carranza, M.L. Unmanned Aerial Vehicle (UAV)-Based Mapping of Acacia saligna Invasion in the Mediterranean Coast. Remote Sens. 2021, 13, 3361. https://doi.org/10.3390/rs13173361
Marzialetti F, Frate L, De Simone W, Frattaroli AR, Acosta ATR, Carranza ML. Unmanned Aerial Vehicle (UAV)-Based Mapping of Acacia saligna Invasion in the Mediterranean Coast. Remote Sensing. 2021; 13(17):3361. https://doi.org/10.3390/rs13173361
Chicago/Turabian StyleMarzialetti, Flavio, Ludovico Frate, Walter De Simone, Anna Rita Frattaroli, Alicia Teresa Rosario Acosta, and Maria Laura Carranza. 2021. "Unmanned Aerial Vehicle (UAV)-Based Mapping of Acacia saligna Invasion in the Mediterranean Coast" Remote Sensing 13, no. 17: 3361. https://doi.org/10.3390/rs13173361
APA StyleMarzialetti, F., Frate, L., De Simone, W., Frattaroli, A. R., Acosta, A. T. R., & Carranza, M. L. (2021). Unmanned Aerial Vehicle (UAV)-Based Mapping of Acacia saligna Invasion in the Mediterranean Coast. Remote Sensing, 13(17), 3361. https://doi.org/10.3390/rs13173361









