Model Selection in Atmospheric Remote Sensing with an Application to Aerosol Retrieval from DSCOVR/EPIC, Part 1: Theory
Abstract
1. Introduction
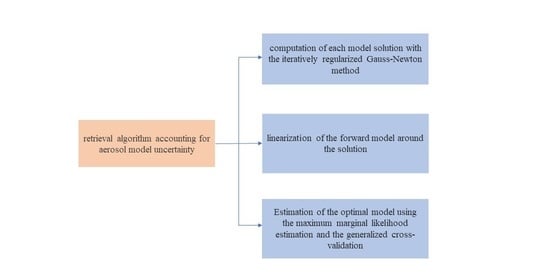
- an iteratively regularized Gauss–Newton method for computing the solution of each model and estimating the ratio of the data error variance and the a priori state variance;
- a linearization of the forward model around the solution for integrating the likelihood density over the state vector;
- parameter choice methods from linear regularization theory for estimating the optimal model and the data error variance.
2. Data Model
- as a Gaussian random vector with zero mean and covariance matrix:and
- as a Gaussian random vector with zero mean and covariance matrix:
- In [3], the covariance matrix was estimated by empirically exploring a set of residuals of model fits to the measurements. Essentially, is assumed to be in the form:where , , and l (representing the non-spectral diagonal variance, the spectral variance, and the correlation length, respectively) are computed by means of an empirical semivariogram, i.e., the variances of the residual differences are calculated for each wavelength pair with the distance , and a theoretical Gaussian variogram model is fitted to these empirical semivariogram values. As compared to Equation (6), in our analysis, we use the diagonal matrix approximation with .
- In principle, the scaling matrix depends on the model error variance through the weighting factor . However, for , we usually have for all . In this case, it follows that , together with and , are close to the identity matrix. This result does not mean that in our model, does not play any role; is included in , which is the subject of an estimation process.
3. Bayesian Approach
3.1. Bayesian Parameter Estimation
3.2. Bayesian Model Selection
- Problem 1.
- Problem 2.
- From Equations (15) and (17), we see that the solution estimates and are expressed in terms of relative evidence , which in turn, according to Equation (14), is expressed in terms of the marginal likelihood density . In view of Equation (8), the computation of the marginal likelihood density requires the knowledge of the likelihood density , and therefore of Equation (10), of the data error variance .
- Problem 3.
- The dependency of the likelihood density on the nonlinear forward model does not allow an analytical integration in Equation (8).
4. Iteratively Regularized Gauss–Newton Method
- is an estimate for the optimal regularization parameter, and
- is the minimizer of the Tikhonov function with regularization parameter , .
- the computation of the regularized solution depends only on the initial value and the ratio q of the geometric sequence which determine the rate of convergence, and
- the regularization parameter at the solution is an estimate for the optimal regularization parameter, and so, for the ratio of the data error variance and the a priori state variance.
5. Parameter Estimation and Model Selection
- the a posteriori density can be expressed as a Gaussian distribution:where:is the a posteriori covariance matrix, and
- as shown in Appendix A, the marginal likelihood density can be computed analytically; the result is:where is the influence matrix at the solution .
- in the framework of maximum marginal likelihood estimation, the data error variance can be estimated by:and the model with the smallest value of the marginal likelihood function, defined by:is optimal.
- in the framework of generalized cross-validation, the data error variance can be estimated by:and the model with the smallest value of the generalized cross-validation function, defined by:is optimal.
- the relative evidence of an approach based on marginal likelihood and the computation of the data error variance in the framework of the Maximum Marginal Likelihood Estimation (MLMMLE) by:
- the relative evidence of an approach based on the marginal likelihood and computation of the data error variance in the framework of the Generalized Cross-Validation (MLGCV) by:
- the relative evidence of an approach based on Maximum Marginal Likelihood Estimation (MMLE) by:
- the relative evidence of an approach based on Generalized Cross-Validation (GCV) by:
6. Algorithm Description
- compute the scaling matrix by means of Equation (5) and the scaled data vector ;
- given the current iterate at step k, compute the forward model , the Jacobian matrix , and the scaled quantities and ;
- compute the linearized data vector:
- compute the singular value decomposition of the quotient matrix , where with for is a diagonal matrix containing the singular values in decreasing order and and are orthogonal matrices containing the left and right singular column vectors and , respectively;
- if , choose , where and are the largest and the smallest singular values, respectively; otherwise, set ;
- compute the minimizer of the Tikhonov function for the linearized equation,and the new iterate, ;
- compute the nonlinear residual vector at ,and the residual ;
- compute the condition number , the scalar quantities:and the normalized covariance matrix:where ;
- if , recompute by means of a step-length algorithm such that ; if the residual cannot be reduced, set . and go to Step 12;
- compute the relative decrease in the residual ;
- if , go to Step 1; otherwise set , and go to Step 12;
- determine such that , for all ;
- since , , , and , compute the estimates:the marginal likelihood:where X stands for the character strings MLMMLE and MLGCV when the character variable Y takes the values L and G, respectively, and the covariance matrix,
- Another estimate for the data error variance is derived in Appendix C; this is given by:
- If a model is far from reality, it is natural to assume that the model parameter errors are large or, equivalently, that the data error variance is large. This observation suggests that, in a deterministic setting, we may define the relative evidence as:where the character variable Y takes the values R, L, or G. In this case, for , the model with the smallest value of the squared residual is considered to be optimal.
7. Application to the EPIC Instrument
- compute the radiances for the moderately absorbing aerosol model and the true values and , ;
- add the measurement noise ,where , , and:which is the maximum signal-to-noise ratio according to the EPIC camera specifications and
- in view of the approximation:identify , , and:implying:
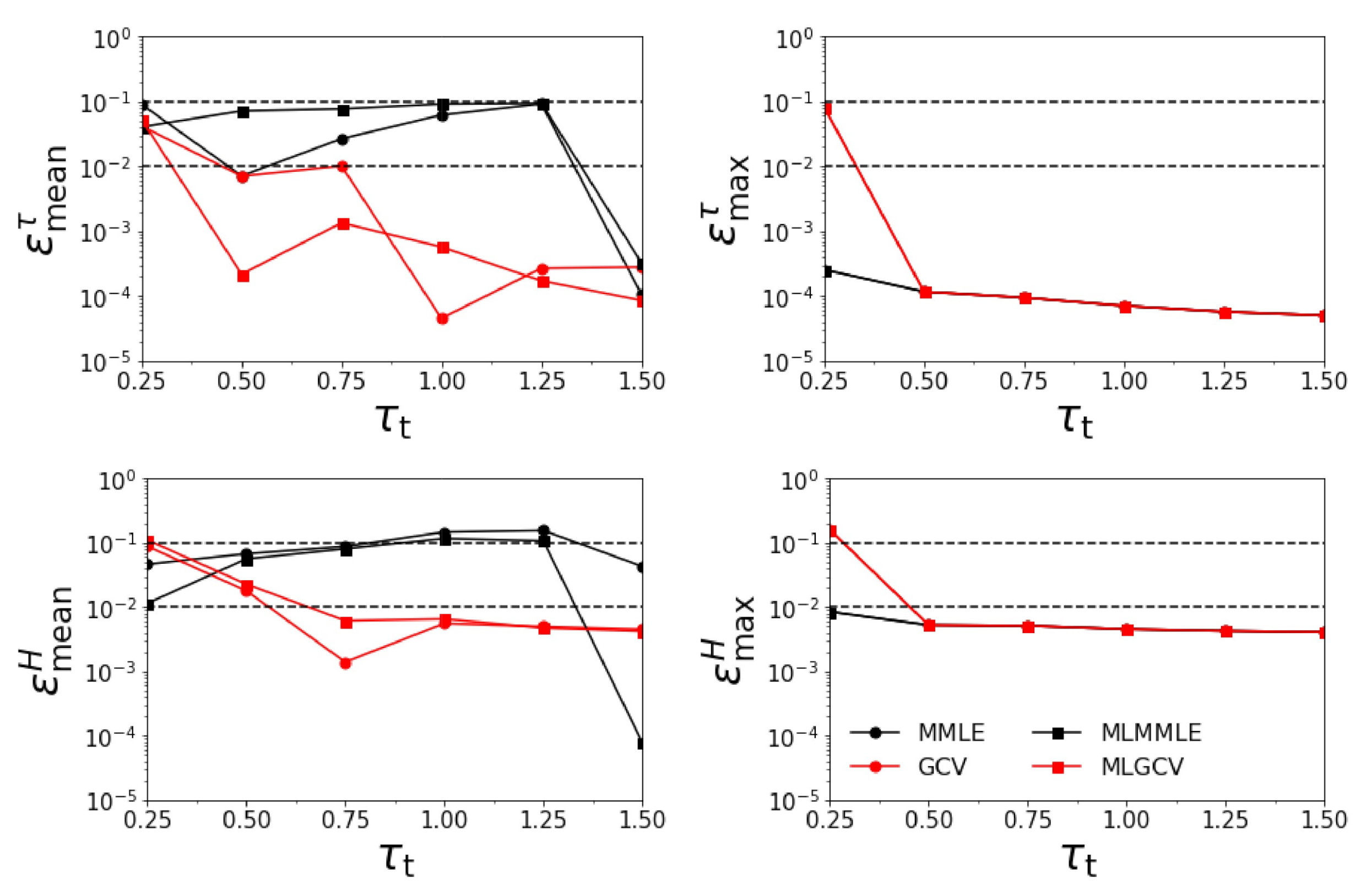
- Test Example 1
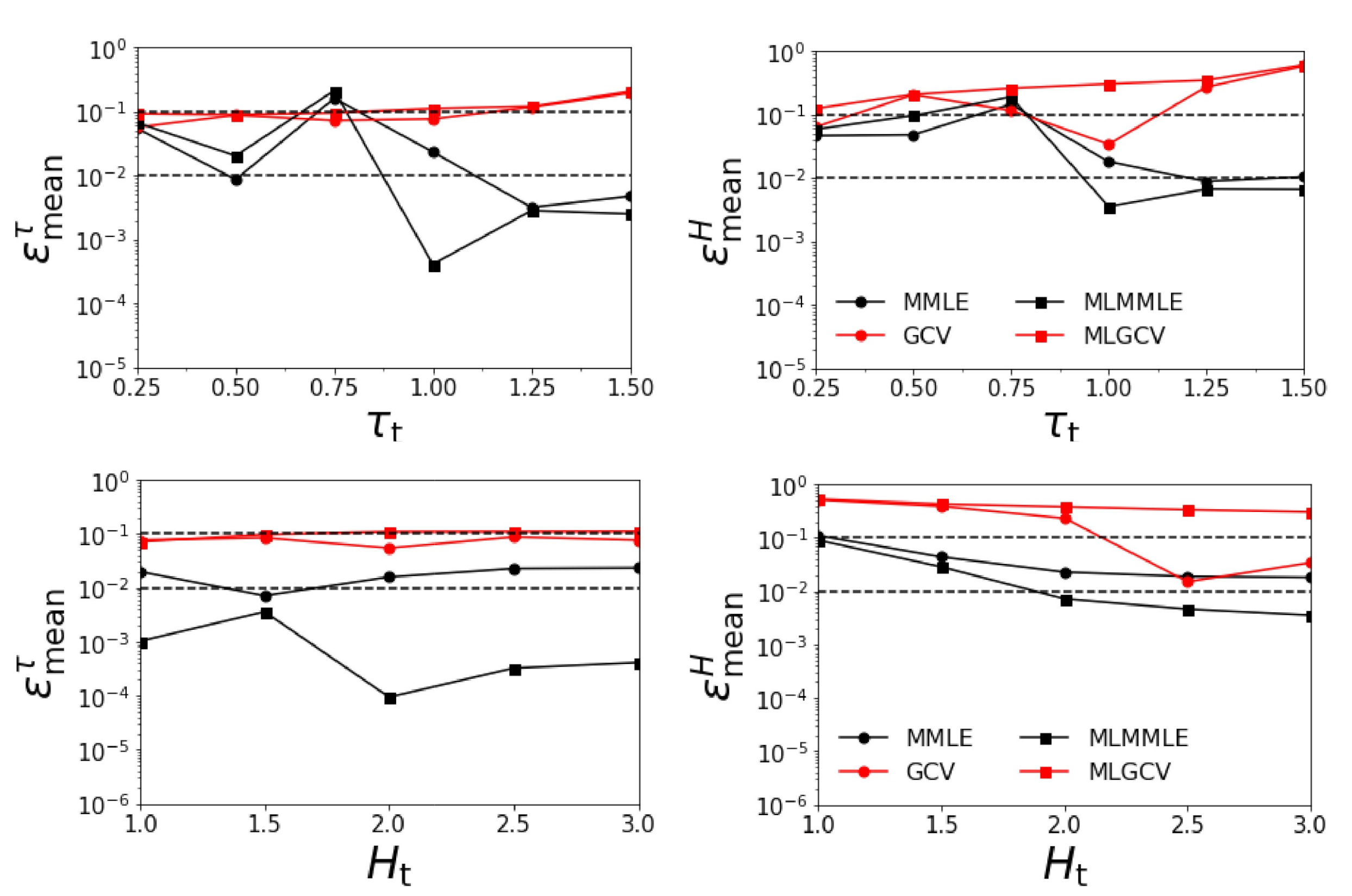
- the relative errors and corresponding to the generalized cross-validation (GCV and MLGCV) are in general smaller than those corresponding to the maximum marginal likelihood estimation (MMLE and MLMMLE).
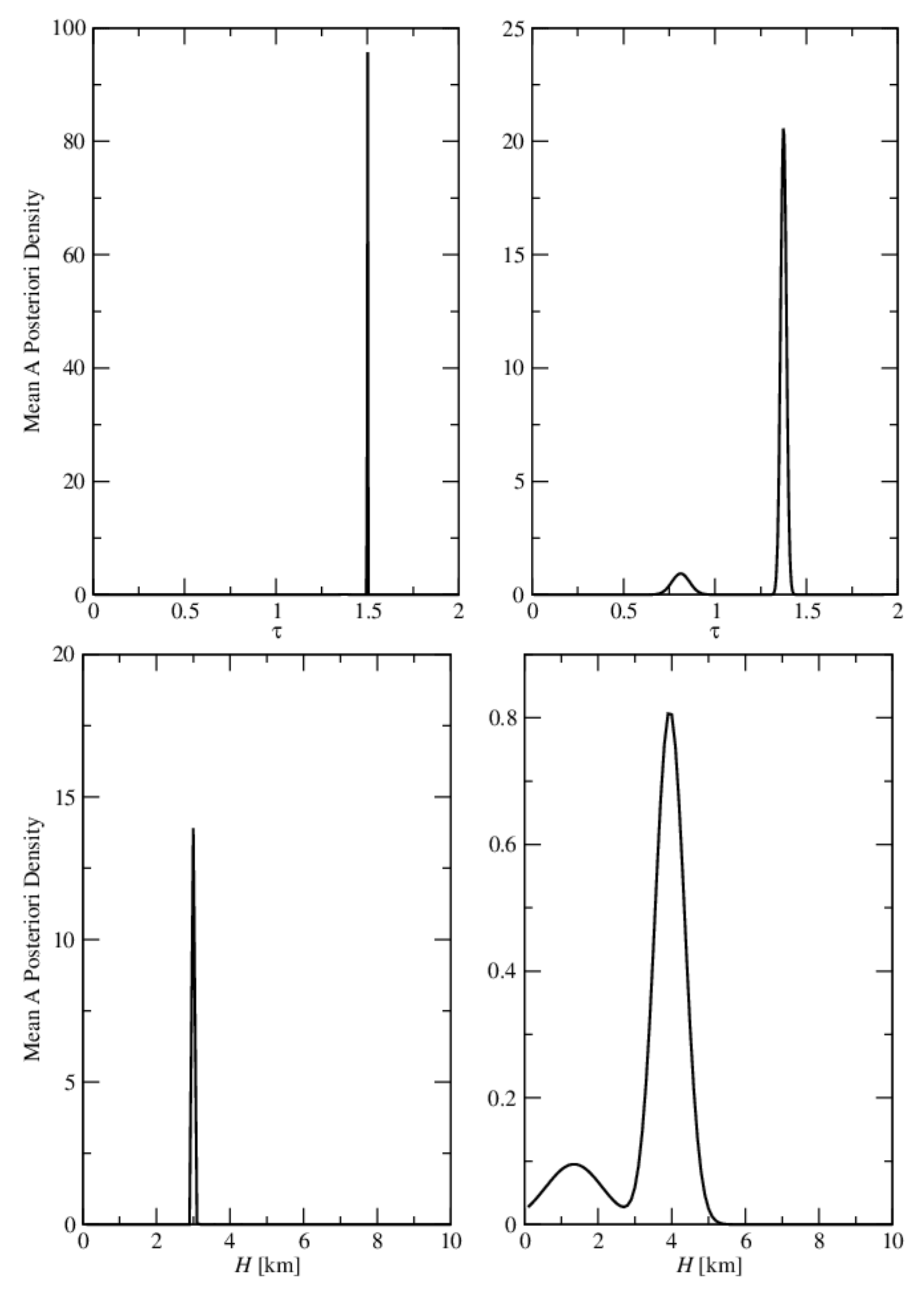
- the relative errors and are very small, and so, we deduce that the algorithm recognizes the exact aerosol model;
- the best method is GCV, in which case the average relative errors are , , , and ;
- the aerosol optical thickness is better retrieved than the aerosol layer height.
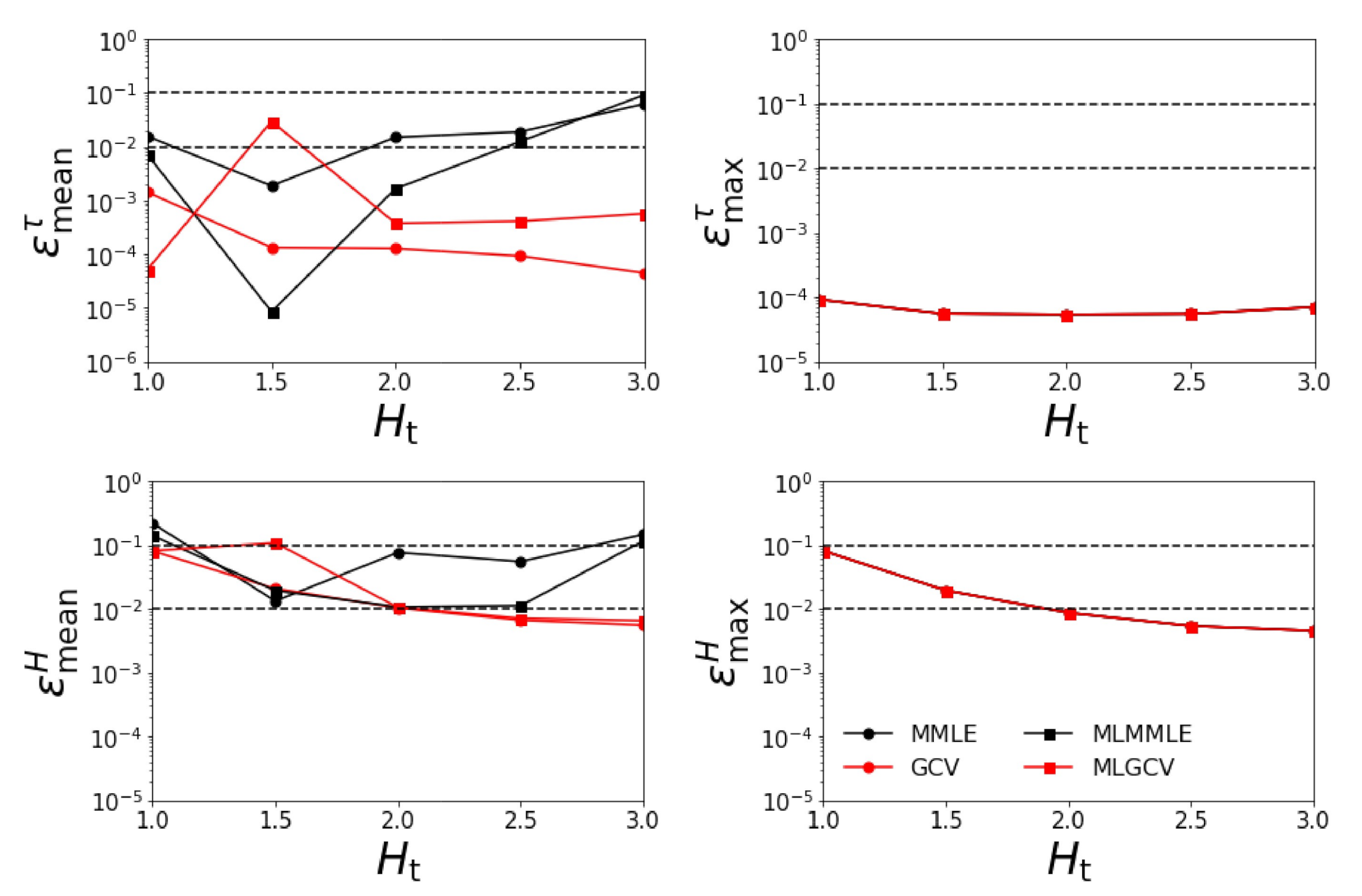
- Test Example 2
- the relative errors and corresponding to the generalized cross-validation (GCV and MLGCV) are still smaller than those corresponding to the maximum marginal likelihood estimation (MMLE and MLMMLE).
- the relative errors and are extremely large, and so, we infer that the maximum solution estimate is completely unrealistic;
- as before, the best method is GCV characterized by the average relative errors , , , and ;
- the relative errors are significantly larger than those corresponding to the case when all four aerosol models are taken into account.
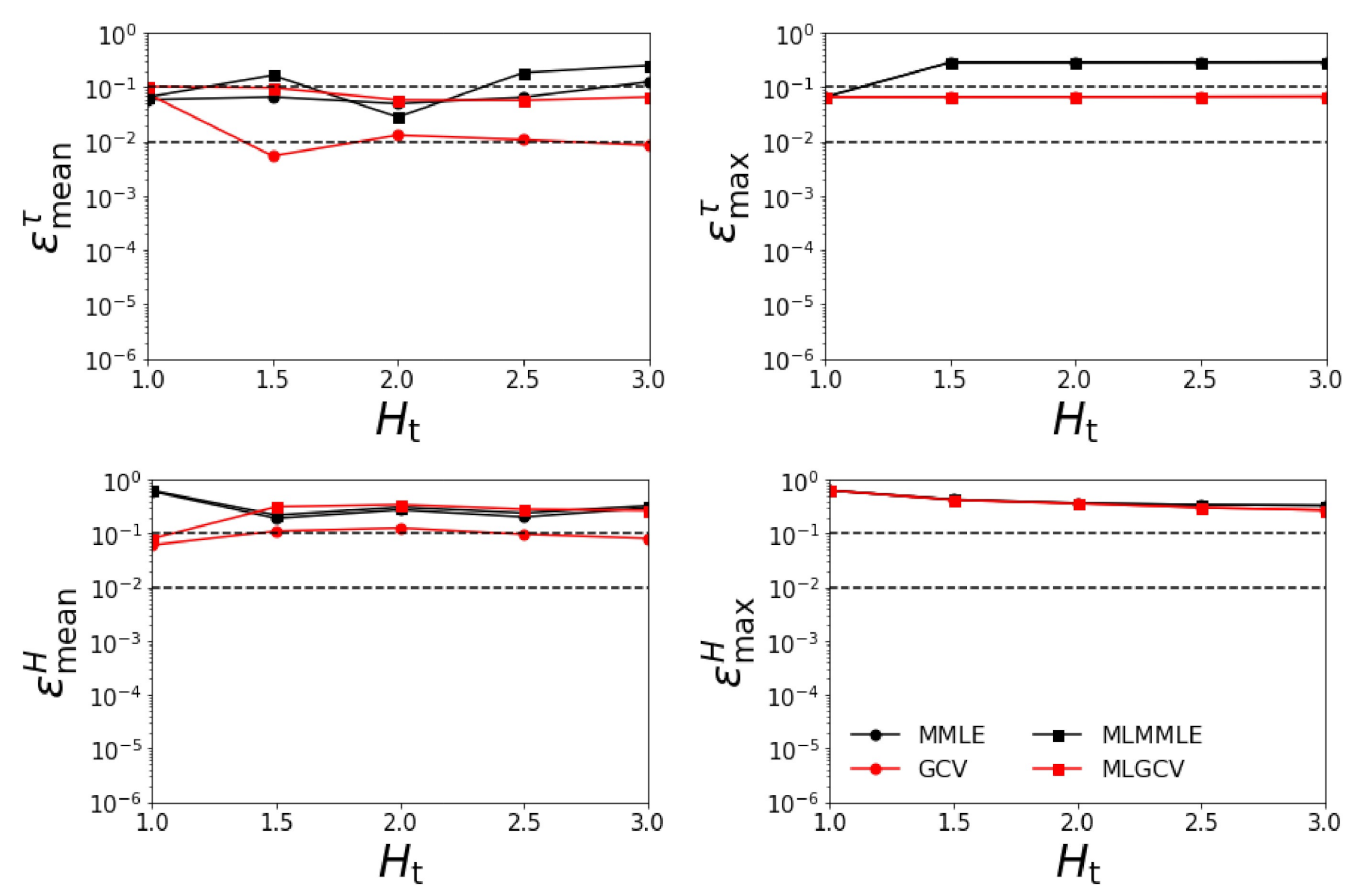
- Test Example 3
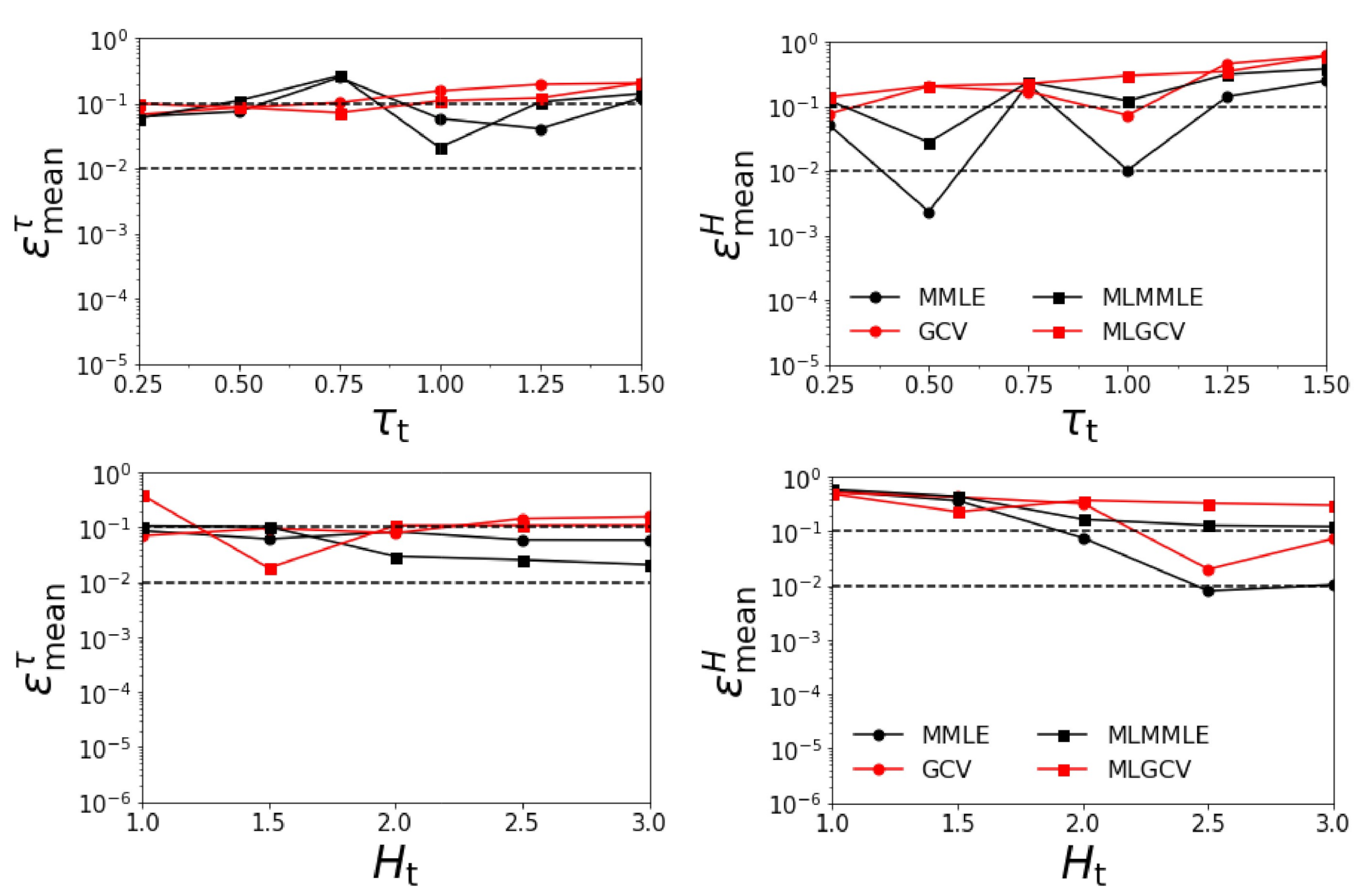
- the relative errors and corresponding to generalized cross-validation (GCV and MLGCV) are in general larger than those corresponding to maximum marginal likelihood estimation (MMLE and MLMMLE);
- the best methods are MMLE and MLMMLE; the average relative errors given by MLMMLE are , , , and ;
- the relative errors are significantly larger than those obtained in the first two test examples (when the surface albedo is known exactly).
- as in the previous scenario, the relative errors and corresponding to generalized cross-validation (GCV and MLGCV) are in general larger than those corresponding to maximum marginal likelihood estimation (MMLE and MLMMLE).
- the best methods are MMLE and MLMMLE; the average relative errors delivered by MMLE are = 0.101, = 0.113, = 0.070, and = 0.204;
- the relative errors are the largest among all the test examples.
8. Conclusions
- an application of the prewhitening technique in order to transform the data model into a scaled model with white noise;
- a deterministic regularization method, i.e., the iteratively regularized Gauss–Newton method, in order to compute the regularized solution (equivalent to the maximum a posteriori estimate of the solution in a Bayesian framework) and to determine the optimal value of the regularization parameter (equivalent to the ratio of the data error and a priori state variances in a Bayesian framework);
- a linearization of the forward model around the solution in order to transform the nonlinear data model into a linear model and, in turn, facilitate an analytical integration of the likelihood density over the state vector;
- an extension of maximum marginal likelihood estimation and generalized cross-validation to model selection and data error variance estimation.
- the two parameter choice methods used (maximum marginal likelihood estimation and generalized cross-validation) and
- the two settings in which the relative evidence is treated (stochastic and deterministic).
- The differences between the results corresponding to the stochastic and deterministic interpretations of the relative evidence are not significant.
- If the surface albedo is assumed to be known, generalized cross-validation is superior to maximum marginal likelihood estimation; if the surface albedo is included in the retrieval, the contrary is true.
- The errors in the aerosol optical thickness retrieval are smaller than those in the aerosol layer height retrieval. In the most realistic situation, when the exact aerosol model and surface albedo are unknown, the average relative errors in the retrieved aerosol optical thickness are about 10%, while the corresponding errors in the aerosol layer height are about 20%.
- The maximum solution estimate is completely unrealistic when both the aerosol model and surface albedo are unknown.
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
Appendix B
- Generalized cross-validation
- Maximum marginal likelihood estimation
Appendix C
References
- Levy, R.C.; Remer, L.A.; Dubovik, O. Global aerosol optical properties and application to Moderate Resolution Imaging Spectroradiometer aerosol retrieval over land. J. Geophys. Res. Atmos. 2007, 112. [Google Scholar] [CrossRef]
- Hoeting, J.A.; Madigan, D.; Raftery, A.E.; Volinsky, C.T. Bayesian Model averaging: A tutorial. Stat. Sci. 1999, 14, 382–401. [Google Scholar]
- Määttä, A.; Laine, M.; Tamminen, J.; Veefkind, J. Quantification of uncertainty in aerosol optical thickness retrieval arising from aerosol microphysical model and other sources, applied to Ozone Monitoring Instrument (OMI) measurements. Atmos. Meas. Tech. 2014, 7, 1185–1199. [Google Scholar] [CrossRef]
- Kauppi, A.; Kolmonen, P.; Laine, M.; Tamminen, J. Aerosol-type retrieval and uncertainty quantification from OMI data. Atmos. Meas. Tech. 2017, 10, 4079–4098. [Google Scholar] [CrossRef]
- Doicu, A.; Trautmann, T.; Schreier, F. Numerical Regularization for Atmospheric Inverse Problems; Springer: Berlin/Heidelberg, Germany, 2010. [Google Scholar] [CrossRef]
- Tarantola, A. Inverse Problem Theory and Methods for Model Parameter Estimation; Society for Industrial and Applied Mathematics (SIAM): Philadelphia, PA, USA, 2005. [Google Scholar] [CrossRef]
- Patterson, H.; Thompson, R. Recovery of inter-block information when block sizes are unequal. Biometrika 1971, 58, 545–554. [Google Scholar] [CrossRef]
- Smyth, G.K.; Verbyla, A.P. A conditional likelihood approach to residual maximum likelihood estimation in generalized linear models. J. R. Stat. Soc. Ser. Methodol. 1996, 58, 565–572. [Google Scholar] [CrossRef]
- Stuart, A.; Ord, J.; Arnols, S. Kendall’s Advanced Theory of Statistics. Volume 2A: Classical Inference and the Linear Model; Oxford University Press Inc.: Oxford, UK, 1999. [Google Scholar]
- Wahba, G. Practical approximate solutions to linear operator equations when the data are noisy. SIAM J. Numer. Anal. 1977, 14, 651–667. [Google Scholar] [CrossRef]
- Wahba, G. Spline Models for Observational Data; Society for Industrial and Applied Mathematics (SIAM): Philadelphia, PA, USA, 1990. [Google Scholar] [CrossRef]
- Marshak, A.; Herman, J.; Szabo, A.; Blank, K.; Carn, S.; Cede, A.; Geogdzhayev, I.; Huang, D.; Huang, L.K.; Knyazikhin, Y.; et al. Earth Observations from DSCOVR EPIC Instrument. Bull. Am. Meteorol. Soc. 2018, 99, 1829–1850. [Google Scholar] [CrossRef] [PubMed]
- Geogdzhayev, I.V.; Marshak, A. Calibration of the DSCOVR EPIC visible and NIR channels using MODIS Terra and Aqua data and EPIC lunar observations. Atmos. Meas. Tech. 2018, 11, 359–368. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Wang, J.; Wang, Y.; Zeng, J.; Torres, O.; Reid, J.; Miller, S.; Martins, J.; Remer, L. Detecting layer height of smoke aerosols over vegetated land and water surfaces via oxygen absorption bands: Hourly results from EPIC/DSCOVR in deep space. Atmos. Meas. Tech. 2019, 12, 3269–3288. [Google Scholar] [CrossRef]
- García, V.M.; Sasi, S.; Efremenko, D.S.; Doicu, A.; Loyola, D. Radiative transfer models for retrieval of cloud parameters from EPIC/DSCOVR measurements. J. Quant. Spectrosc. Radiat. Transf. 2018, 213, 228–240. [Google Scholar] [CrossRef]
- García, V.M.; Sasi, S.; Efremenko, D.S.; Doicu, A.; Loyola, D. Linearized radiative transfer models for retrieval of cloud parameters from EPIC/DSCOVR measurements. J. Quant. Spectrosc. Radiat. Transf. 2018, 213, 241–251. [Google Scholar] [CrossRef]
- Holben, B.; Eck, T.; Slutsker, I.; Tanré, D.; Buis, J.; Setzer, A.; Vermote, E.; Reagan, J.; Kaufman, Y.; Nakajima, T.; et al. AERONET—A federated instrument network and data archive for aerosol characterization. Remote Sens. Environ. 1998, 66, 1–16. [Google Scholar] [CrossRef]

| Model | ||||
|---|---|---|---|---|
| Method | Average Relative Error | |||
|---|---|---|---|---|
| MMLE | 0.046 | 0.091 | 0.027 | 0.102 |
| MLMMLE | 0.009 | 0.062 | 0.022 | 0.059 |
| GCV | 0.009 | 0.020 | 0.001 | 0.024 |
| MLGCV | 0.009 | 0.025 | 0.006 | 0.042 |
| Method | Average Relative Error | |||
|---|---|---|---|---|
| MMLE | 0.095 | 0.206 | 0.073 | 0.318 |
| MLMMLE | 0.180 | 0.210 | 0.139 | 0.348 |
| GCV | 0.060 | 0.111 | 0.022 | 0.096 |
| MLGCV | 0.059 | 0.200 | 0.076 | 0.259 |
| Method | Average Relative Error | |||
|---|---|---|---|---|
| MMLE | 0.042 | 0.046 | 0.018 | 0.042 |
| MLMMLE | 0.051 | 0.060 | 0.011 | 0.027 |
| GCV | 0.101 | 0.213 | 0.076 | 0.237 |
| MLGCV | 0.119 | 0.311 | 0.100 | 0.398 |
| Method | Average Relative Error | |||
|---|---|---|---|---|
| MMLE | 0.101 | 0.113 | 0.070 | 0.204 |
| MLMMLE | 0.117 | 0.203 | 0.057 | 0.291 |
| GCV | 0.136 | 0.269 | 0.109 | 0.274 |
| MLGCV | 0.116 | 0.307 | 0.147 | 0.346 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sasi, S.; Natraj, V.; Molina García, V.; Efremenko, D.S.; Loyola, D.; Doicu, A. Model Selection in Atmospheric Remote Sensing with an Application to Aerosol Retrieval from DSCOVR/EPIC, Part 1: Theory. Remote Sens. 2020, 12, 3724. https://doi.org/10.3390/rs12223724
Sasi S, Natraj V, Molina García V, Efremenko DS, Loyola D, Doicu A. Model Selection in Atmospheric Remote Sensing with an Application to Aerosol Retrieval from DSCOVR/EPIC, Part 1: Theory. Remote Sensing. 2020; 12(22):3724. https://doi.org/10.3390/rs12223724
Chicago/Turabian StyleSasi, Sruthy, Vijay Natraj, Víctor Molina García, Dmitry S. Efremenko, Diego Loyola, and Adrian Doicu. 2020. "Model Selection in Atmospheric Remote Sensing with an Application to Aerosol Retrieval from DSCOVR/EPIC, Part 1: Theory" Remote Sensing 12, no. 22: 3724. https://doi.org/10.3390/rs12223724
APA StyleSasi, S., Natraj, V., Molina García, V., Efremenko, D. S., Loyola, D., & Doicu, A. (2020). Model Selection in Atmospheric Remote Sensing with an Application to Aerosol Retrieval from DSCOVR/EPIC, Part 1: Theory. Remote Sensing, 12(22), 3724. https://doi.org/10.3390/rs12223724