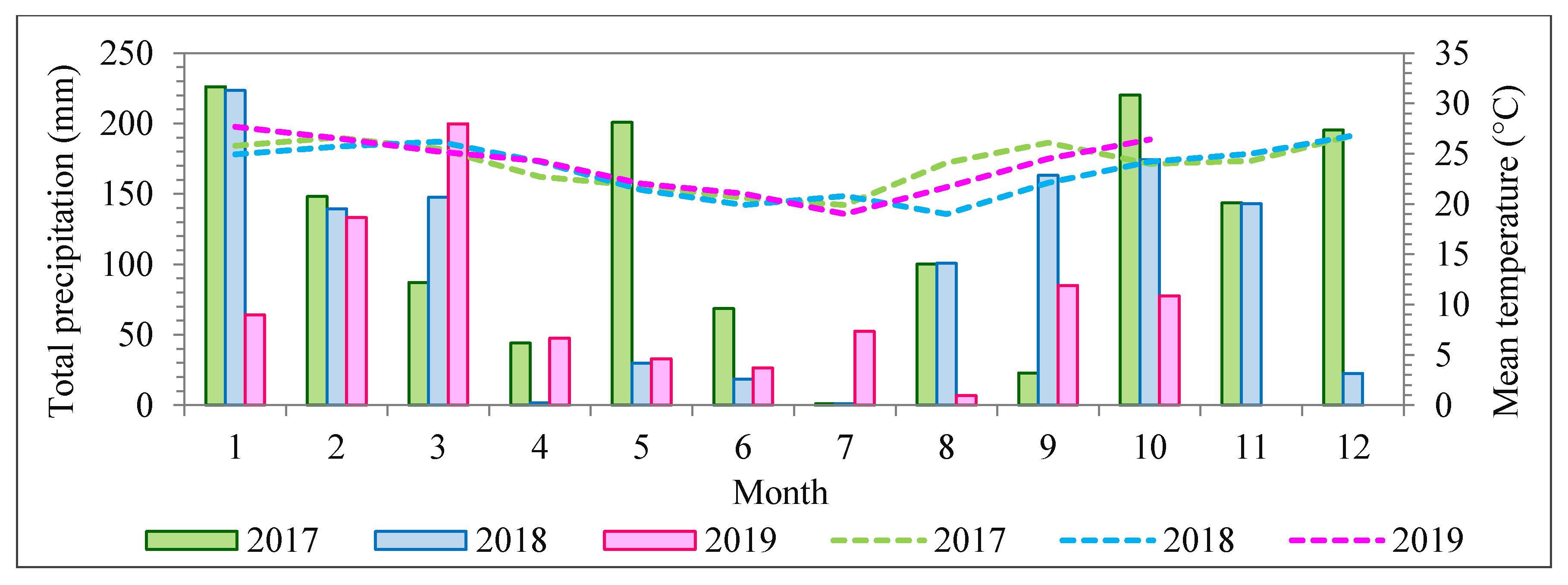2.2. Reference Data
More than 25 tree species with a diameter at breast height (DBH) greater than 3.8 cm were detected during fieldwork [
38]. Tree species were in different successional stages, with the northernmost part of the study area containing trees in the initial stage of succession and the southernmost in a more advanced stage [
39]. Moreover, we located 90 trees of eight species that emerged from the canopy (
Table 1 and
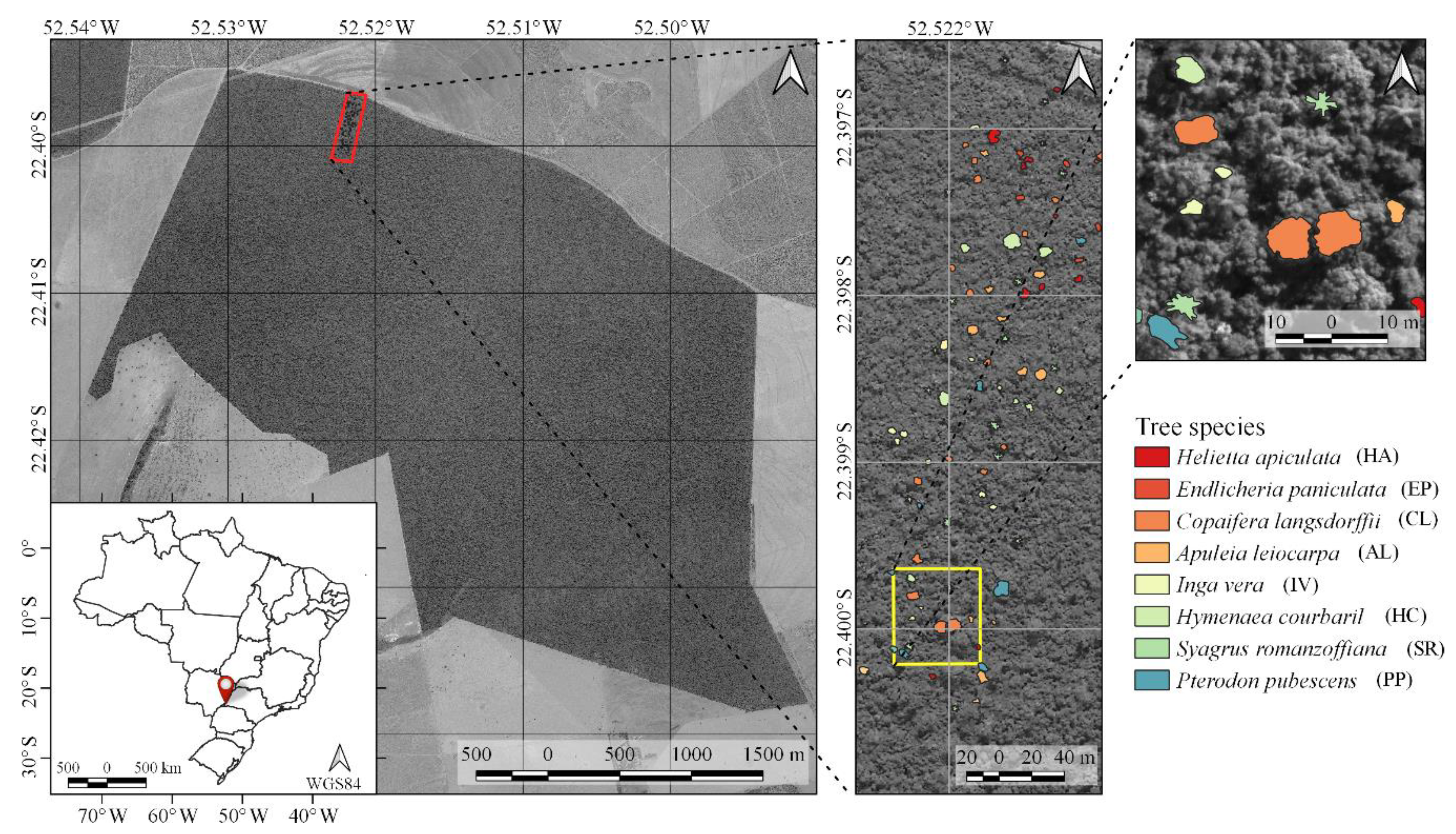
Table 2). Their crowns were manually delineated through visual interpretation of RGB image composites of each dataset (R: 628.73 nm; G: 550.39 nm; B: 506.22 nm).
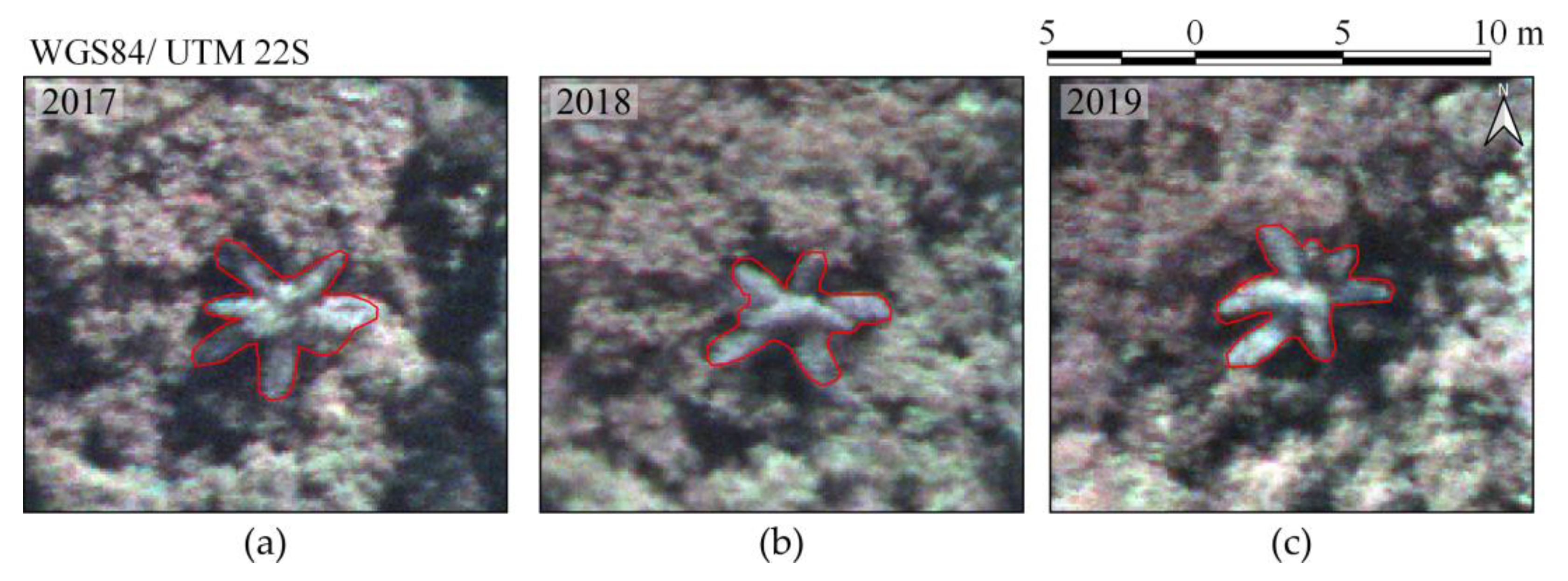
Figure 1 shows examples of delineated individual tree crown (ITC) polygons in the 2017 dataset, and
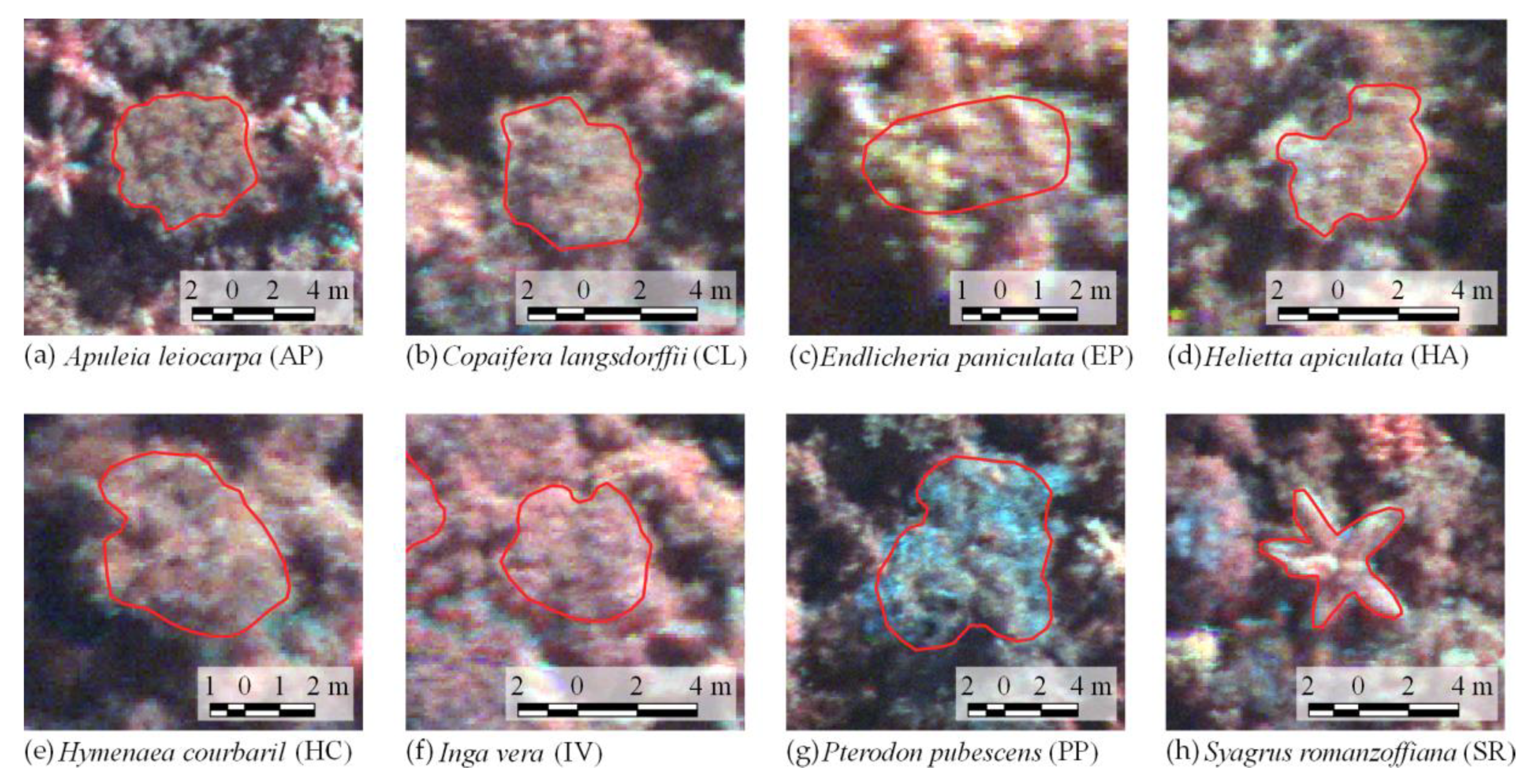
Figure 3 shows canopy examples of each tree species in the mosaic of images acquired during the 2017 flight campaign. These tree species were chosen because they are important for characterizing the successional stage of the forest, e.g.,
Syagrus romanzoffiana (SR), which can be associated with the faunal composition [
40]. It is important to note that smaller tree species were excluded from analysis because the lianas that cover and mix among individuals negatively affect the classification accuracy. From now on, tree species will be called by their abbreviations from
Table 1 and
Table 2.
The number of samples was low for some species because of challenges in acquiring reference data. First, our study area comprised different successional stages; thus, the species composition varied over the area. Second, we used tree samples that emerged from the canopy. In tropical forests, the tree heights can be modeled by the DBH, which can be related to the number of trees per hectare [
43,
44,
45]. According to Lima et al. [
44] and d’Oliveira et al. [
45], the relationship between the DBH and the frequency of trees in tropical forests has an “inverted J shape”, because the number of trees per hectare decreases substantially as the DBH increases, so the number of taller trees also decreases.
2.3. Remote Sensing Data
Hyperspectral images were acquired with a 2D-format camera based on the tunable Fabry–Pérot Interferometer (FPI) from Senop Ltd, model DT-0011 [
46,
47,
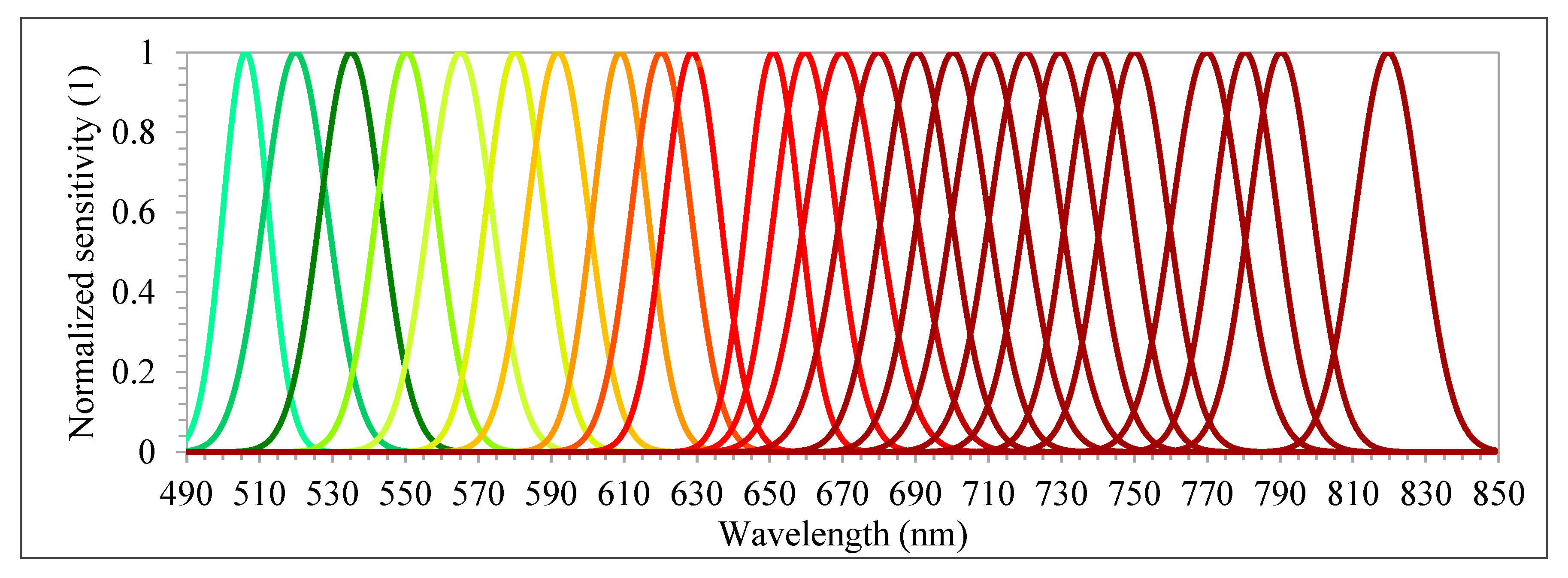
48]. The camera has two sensors, both of which have 1017 × 648 pixels with a pixel size of 5.5 µm. The total weight of the camera is around 700 g with its accessories, which include an irradiance sensor and a Global Positioning System (GPS) receiver. Spectral bands can be selected from the visible (VIS) to near-infrared (NIR) region (500–900 nm), which are acquired sequentially, i.e., the air gap of the FPI moves to acquire the different spectral bands of the same image. The spectral range of the first and second sensors are 647–900 nm and 500–635 nm, respectively. A total of 25 spectral bands were chosen, with the Full-Width at Half Maximum (FWHM) varying from 12.44 to 20.45 nm (
Table 3 and
Figure 4). For this spectral setting, each image cube needs 0.779 s to be acquired. The exposure time was set to 5 ms, and the image blocks were divided into two flight strips, ensuring more than 70% and 50% forward and side overlaps, respectively.
The FPI camera was mounted onboard the UX4 UAV, which is a rotary-wing quadcopter developed by the company Nuvem UAV. The UX4 UAV is almost 90 cm in diameter and 30 cm in height without counting the GPS antenna, which is approximately 15 cm. It is controlled by a PixHawk autopilot. The energy source for the UAV system and its sensors is one six-cell battery of 22 volts and one smaller three-cell battery of 11 volts, which allow the UAV to fly for up to 30 min, depending on payload, battery, and weather conditions. A flight speed of 4 m/s was used to limit the maximum gap between the first and last band of the hyperspectral imager to 3.1 m in a single cube.
During the field campaigns, three radiometric reference targets were placed in the area to enable reflectance calibration. Flight campaigns were performed over the study area (
Figure 1) on 1 July 2017, 16 June 2018, and 13 July 2019, with an above-ground flight height of approximately 160 m and flight speed of 4 m/s. The flight height was selected so that a GSD of 10 cm was obtained. This ensured a good representation of tree crowns that were predominantly over 3 m in diameter.
Table 4 provides more details about the flight time of each campaign and the mean zenith and azimuth angles of the Sun during the image acquisitions.
Images were geometrically and radiometrically processed to obtain hyperspectral image orthomosaics. First, the images were radiometrically corrected from the dark current and nonuniformity of sensors using a dark image acquired before each flight and laboratory parameters [
47,
49].
The geometric processing was performed using the Agisoft PhotoScan software (version 1.3) (Agisoft LLC, St. Petersburg, Russia). In the orientation process, for each year, the exterior orientation parameters (EOPs) of four reference bands (band 3: 550.39 nm; band 8: 609.00 nm; band 14: 679.84 nm; and band 22: 769.89 nm) were estimated in the same Agisoft PhotoScan project in order to reduce misregistration between the datasets. The EOPs of the other bands were calculated using the method developed in [
49,
50]. Positions from the camera GPS were used as initial values and refined using a bundle block adjustment (BBA) and ground control points (GCPs). The number of GCPs varied between datasets, with 3, 3, and 4 used in 2017, 2018, and 2019, respectively. GCPs were placed outside the forest since it was not possible to see the ground from imagery acquired over the forested area. Initially, the base station was defined near the study area, and the global navigation satellite system (GNSS) observations from GCPs were collected and processed in differential mode.
A self-calibrating bundle adjustment was used to estimate the interior orientation parameters (IOPs) of each sensor and for each year of the dataset. After initial image alignment, parameter estimation was optimized with automatic outlier removal using a gradual selection of tie points based on reconstruction uncertainty and reprojection error, together with the manual removal of points. The final products of this step were the calibrated IOPs, EOPs, sparse and dense point clouds, and digital surface model (DSM) of the area with a GSD of 10 cm. These were used in the following radiometric block adjustment and mosaic generation.
Radiometric adjustment processing aims to correct the digital number (DN) of pixels of images from the bidirectional reflectance distribution function (BRDF) effects and differences caused by the different geometries of acquisition due to the UAV and Sun movements. Thus, nonuniformities among images were compensated for by the radBA software, developed at the Finnish Geospatial Research Institute (FGI) [
49,
50]. Equation (1) shows the model used in the software to extract the reflectance value from the DN of each pixel.
where
is the digital number of pixel
in image
;
is the corresponding reflectance factor with respect to the zenithal angle
of the incident and reflected light,
and
, respectively, and with the relative azimuthal angle
, where
refers to the reflected azimuthal angle, and
denotes the incident azimuthal angle;
is the relative correction factor of illumination differences with respect to the reference image; and
and
are the empirical line parameters for the linear transformation between reflectance and DNs.
According to a previous study by Miyoshi et al. [
47], for the study area, the best initial relative correction factor (
) was one (1), with a standard deviation equal to 0.05. It is worth noting that an exception was necessary for the dataset from 2018 because of higher density differences in cloud covering. The 2017 and 2019 flights were carried out in almost blue-sky conditions, with slight differences compensated for by the radiometric block adjustment. The radiometric block adjustment was performed in two steps for the 2018 dataset. First, an initial radiometric block adjustment was performed using initial values of
= 1. In sequence, the final values of
were used as the initial values for the second radiometric block adjustment. Then, the reflectance factor values were estimated using the empirical line method [
51]. The empirical line parameters (
and
) were estimated from the linear relationship between the DN values of three radiometric reference targets with a mean reflectance of 4%, 11%, and 37%. Radiometric reference targets were 90 cm × 90 cm and composed of light-gray, gray, and black synthetic material. Thus, the mosaics of hyperspectral images for each dataset representing the reflectance factor values were obtained.
Additionally, point cloud data from airborne laser scanning (ALS) were provided by the company Fototerra. ALS data were acquired in November 2017 using a Riegl LMS-Q680i laser scanner (RIEGL, Horn, Austria) onboard a manned aircraft at a flight height of 400 m, which resulted in an average density of 8.4 points/m
2. The canopy height model (CHM) was obtained by extracting the digital terrain model (DTM) from the DSM. The processing was performed using the LAStools software (Martin Isenburg, LAStools—efficient tools for LiDAR processing) [
52]. First, the lasnoise tool was applied to withdraw possible noises in the point cloud. Then, the CHM was extracted using the lasheight tool to obtain tree heights in the study area.
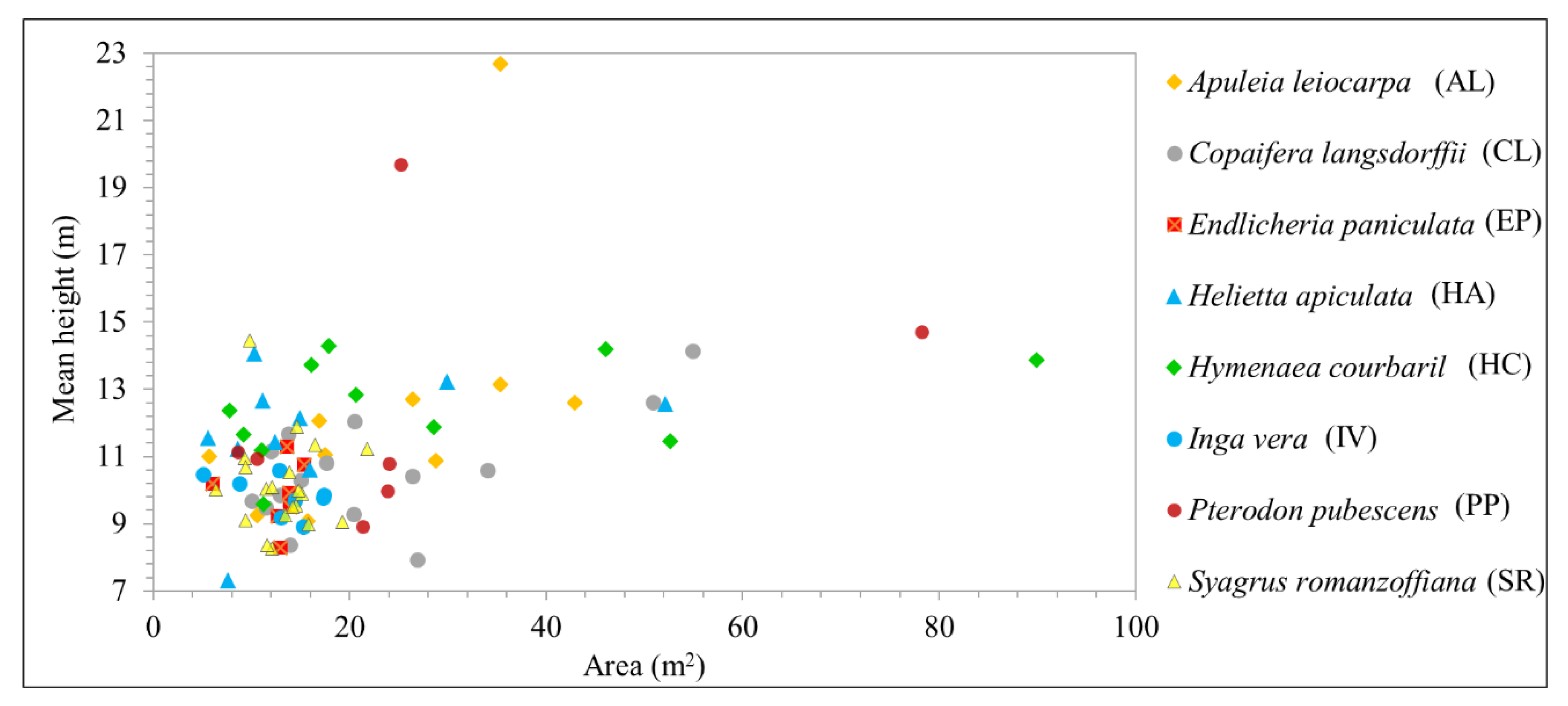
Figure 5 shows the mean height of each tree sample recognized in the field; most of the observed samples fell within a similar height range. Additionally, taller trees were found in the more developed successional stage of the area. Trees of the same species varied in age and were found in regions of different successional stages. For example, PP trees had crown areas of around 25 m
2 and mean heights of 10–20 m. Similarly, HC samples had a mean height of almost 14 m, with tree crown areas ranging from 16 to 90 m
2. It is important to highlight that the ALS data were not used in the classification step, since the objective of this research was to evaluate hyperspectral multitemporal data to improve tree species identification.
2.5. Tree Species Identification with RF
The RF method was used for tree species identification using only the spectral information, since the objective was to verify whether the use of temporal information could improve tree species detection. This method is based on multiple decision trees, and the class is determined by the most popular vote [
6]. Decision trees are composed of different samples, which are drawn with replacement, i.e., one sample can belong to more than one tree [
54]. The RF has been successfully applied for image classification when working with high-dimensional data [
8,
13], and it is less sensitive to feature selection [
54]. The RF was applied using the default parameters of the Weka software version 3.8.3 (The University of Waikato, Hamilton, New Zealand) [
55].
The classification process was carried out five times with four different datasets: (i) the 2017 spectral information (D17); (ii) the 2018 spectral information (D18); (iii) the 2019 spectral information (D19); (iv) the combination of the 2017, 2018, and 2019 spectral information (Dall). For the D17, D18, and D19 datasets, we used the normalized pixel values to extract the spectral features, which are referred to as cases D17_MeanNorm, D18_MeanNorm, and D19_MeanNorm, respectively. Additionally, in the case of the combined dataset (item (iv) in the previously described datasets), the classification was performed using both the normalized and nominal values, referred to as Dall_MeanNorm and Dall_Mean, respectively.
Table 5 summarizes the number of features used in each case.
The number of samples of different species is relatively low and also unbalanced. The leave-one-out cross-validation (LOOCV) method was used to circumvent this problem. LOOCV is a particular case of k-fold cross-validation, where k is equal to the total number of samples of the dataset [
13,
56]. The classification model is trained k times, followed by testing with one subset and training with the remaining subsets. In each iteration, the model is trained using k − 1 samples and tested with the remaining sample. The final accuracy values are obtained by averaging the accuracy values of each iteration [
56]. LOOCV has been successfully applied in tree species classification studies with a small sample size (e.g., less than 10 samples per class [
14]) or an unbalanced number of samples per class [
13].
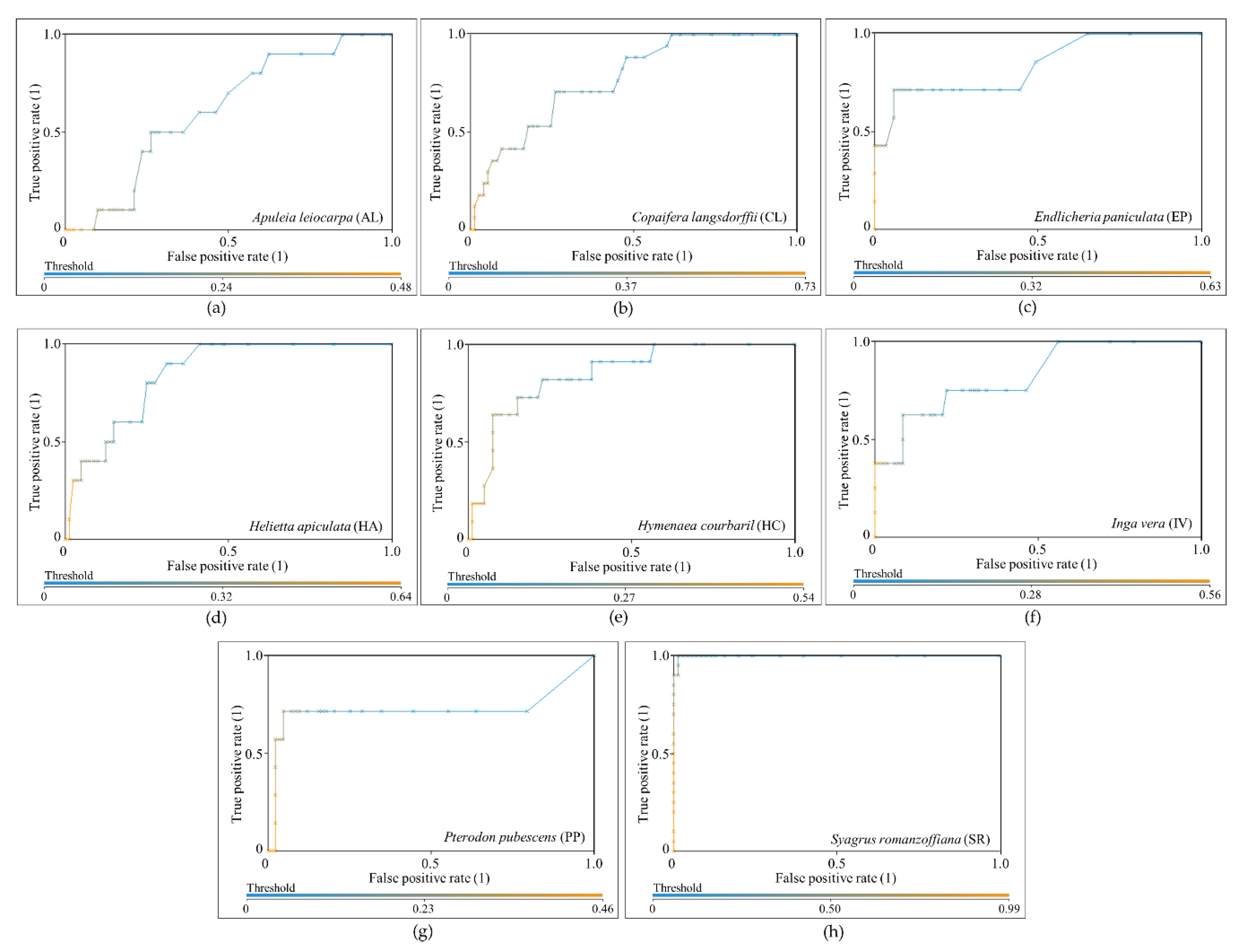
The results were evaluated through the area under the receiver operating characteristic curve, known as AUC (area under the curve) ROC (receiver operating characteristics) or AUCROC [
57,
58,
59]. ROC is the relationship between the false positive rate (FPR), or “1-specificity”, and the true positive rate (TPR), or sensitivity, and it is useful when working with unbalanced classes because it is independent of the class distribution [
59,
60]. When using classifiers such as RF that provide probabilities or scores, thresholds can be applied to acquire different points in the ROC space to form an ROC curve [
60]. AUCROC is the area under the ROC curve, and represents the probability of the classification model correctly classifying a random sample in a specific class. AUCROC varies from 0 to 1 for each class, where a value of 0.5 indicates that the specific classification model is no better than a random assignment, and a value of 1 represents perfect discrimination of a class from the remaining ones [
59]. To the best classification (i.e., the one with highest value of average AUCROC value), the overall accuracy (i.e., the percentage of correctly classified instances of the total number of samples), the user accuracy, and the producer accuracy [
61] were calculated as well.