A Proposed Methodology to Analyze Plant Growth and Movement from Phenomics Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. Image Acquisition
2.3. Image Analysis
2.4. Statistical Analysis
3. Results
3.1. Acquisition of Coordinates and Calibration
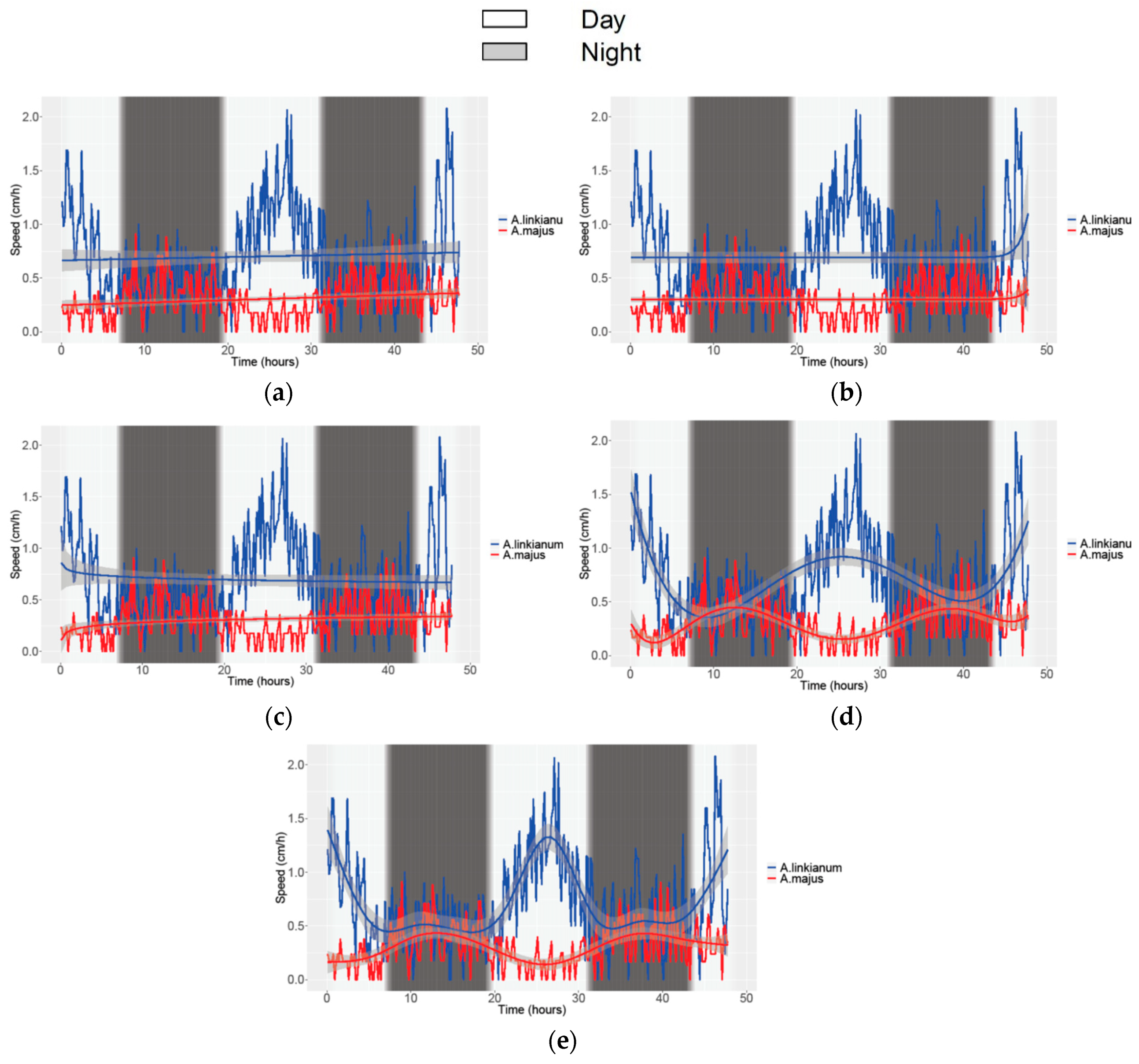
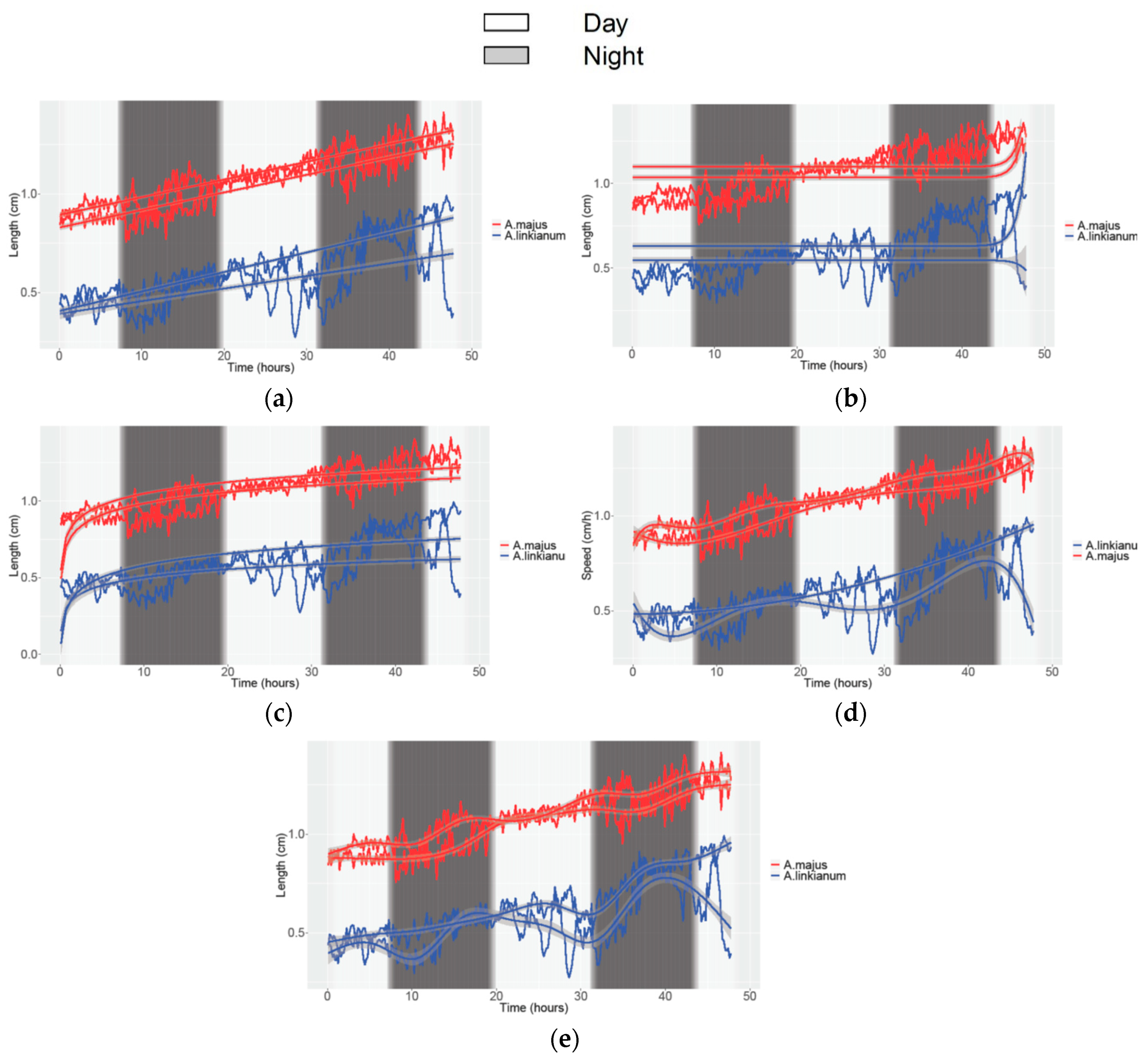
3.2. Obtaining Data of Growth and Movement in Antirrhinum
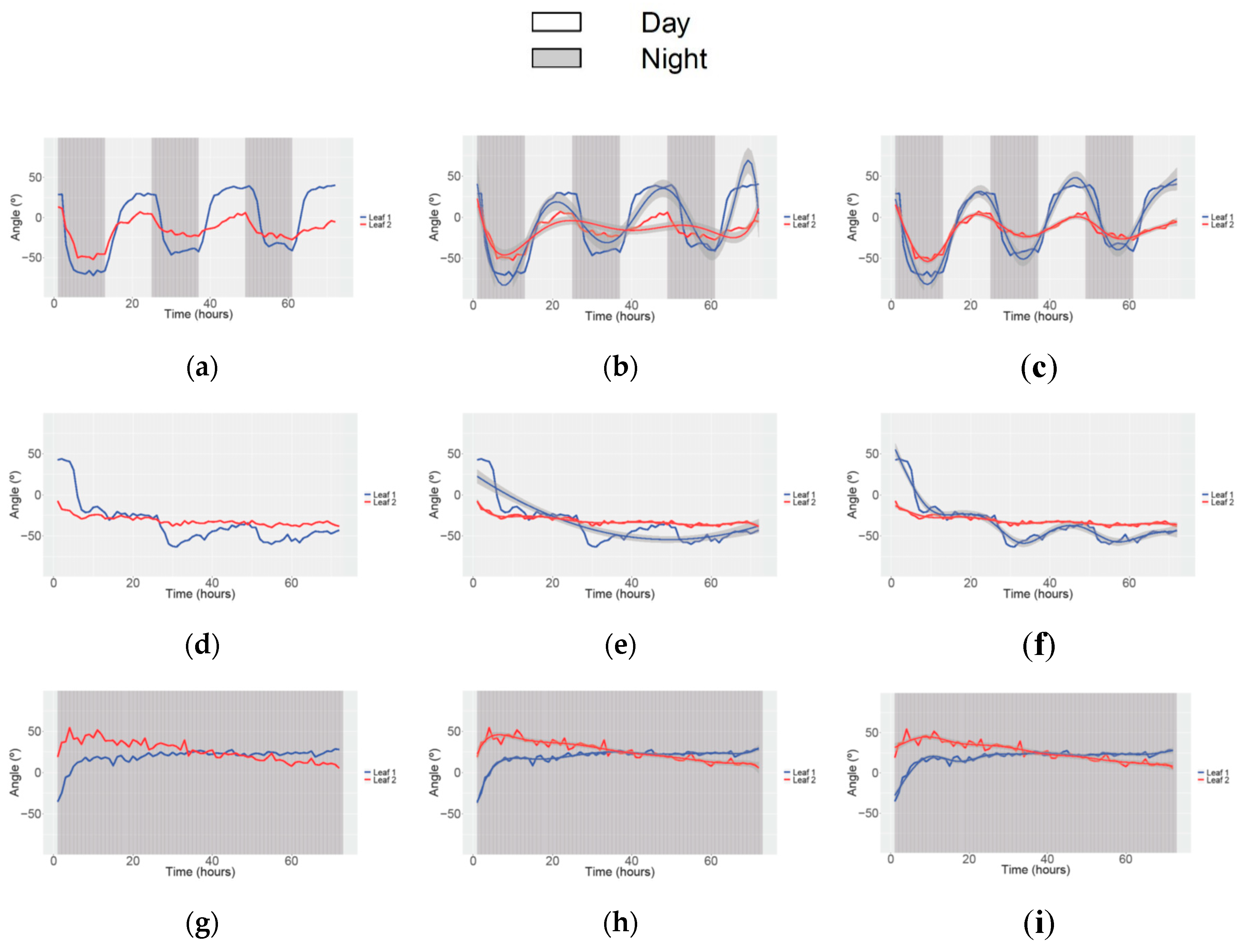
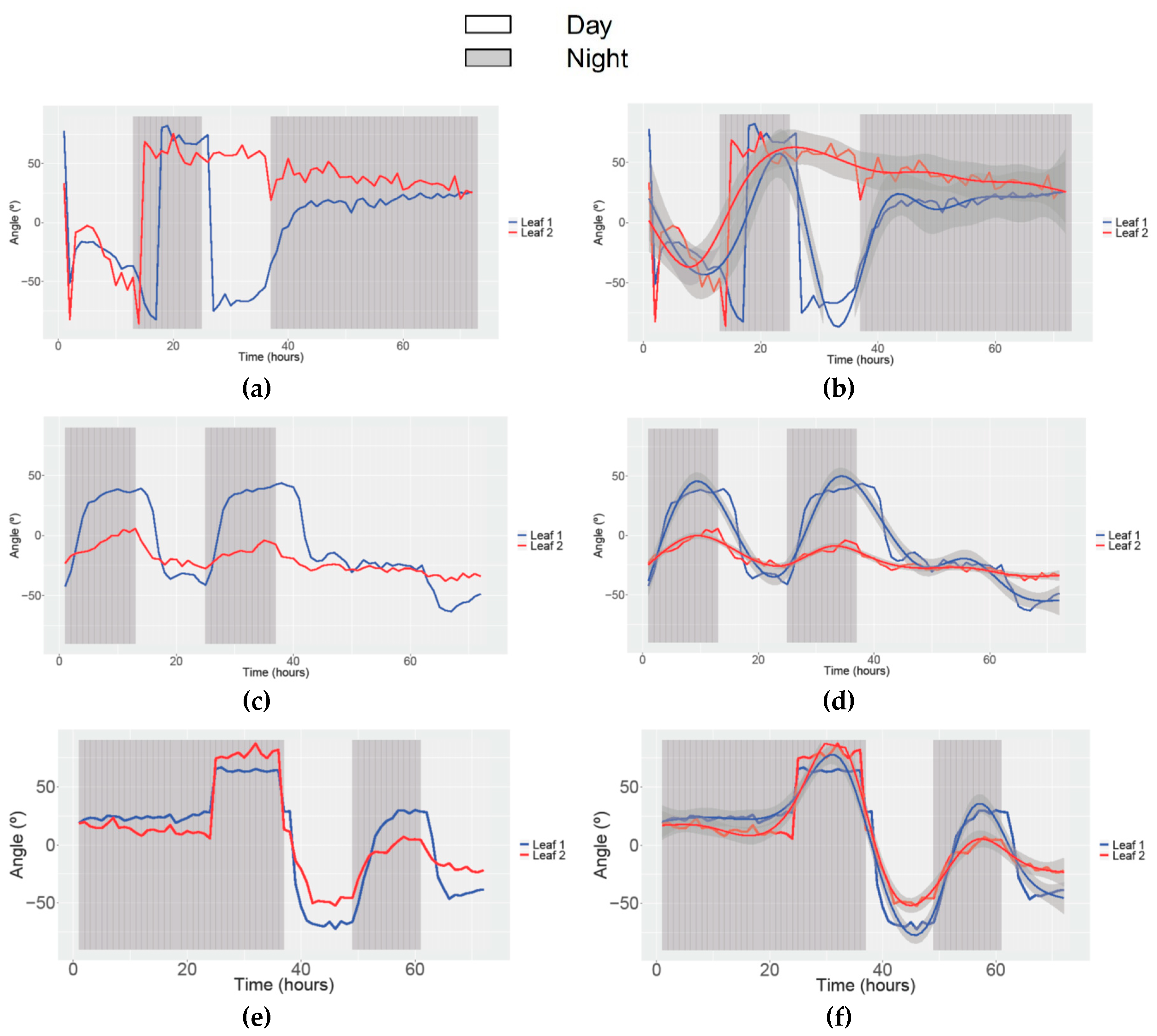
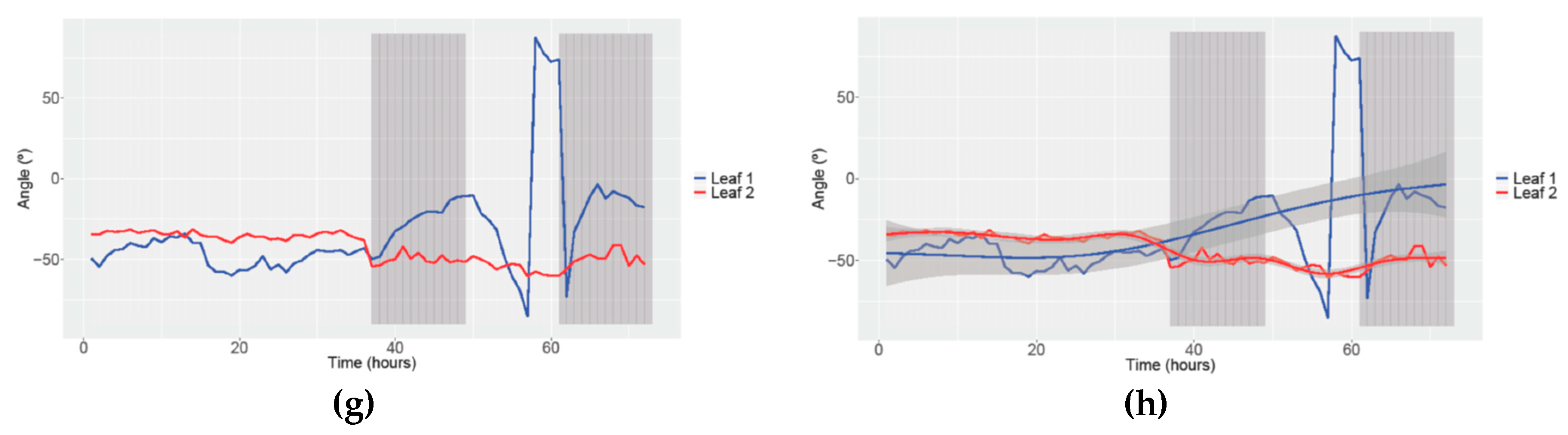
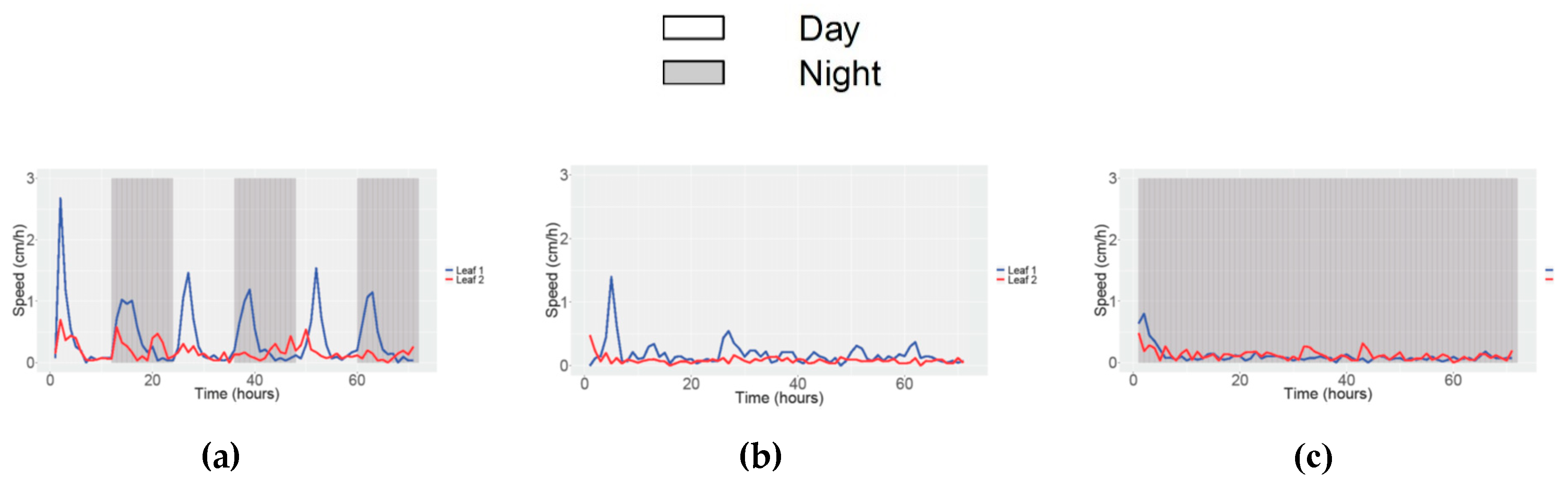
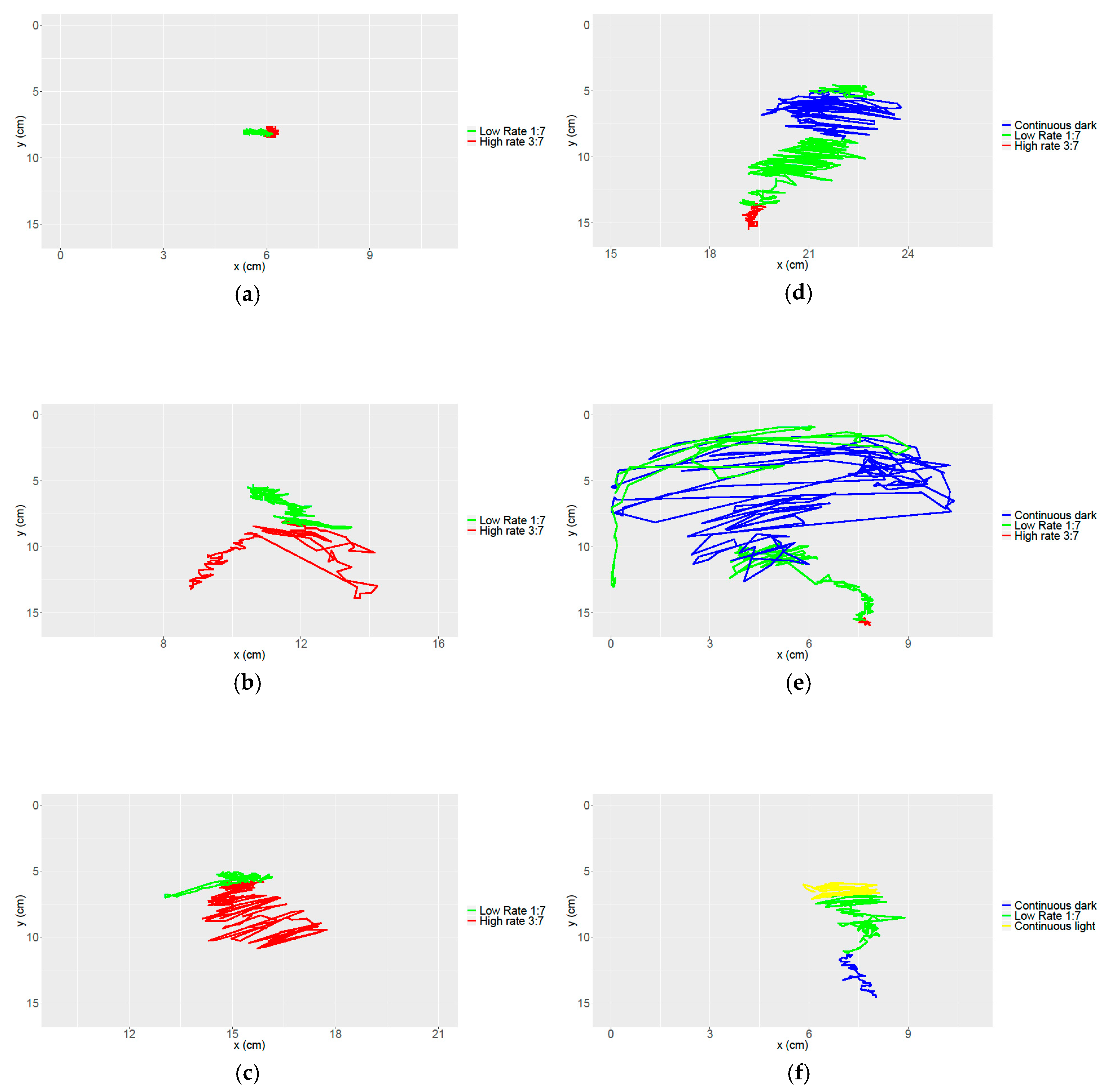
3.3. Obtaining Data of Movement and Speed in Petunia
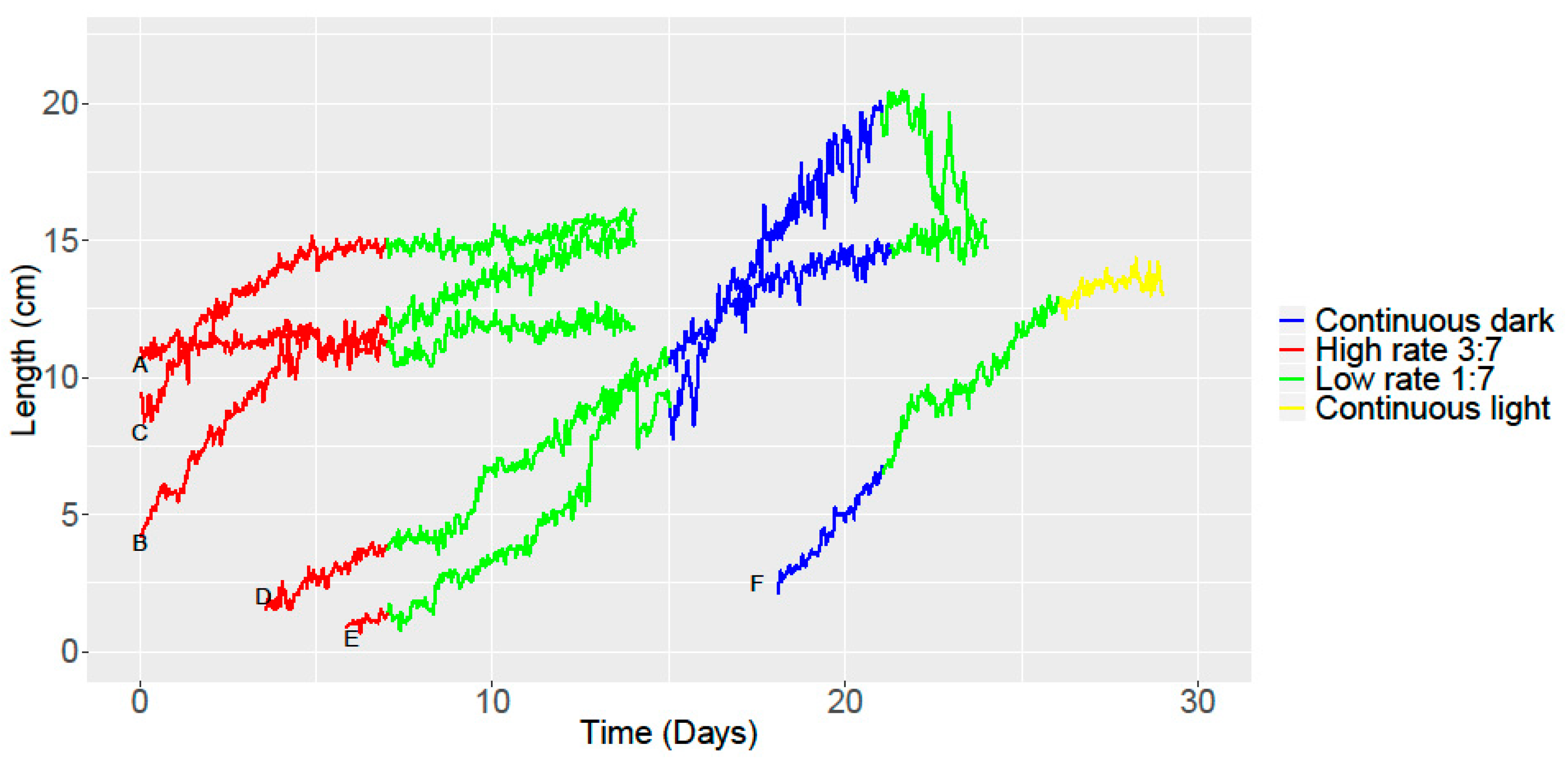
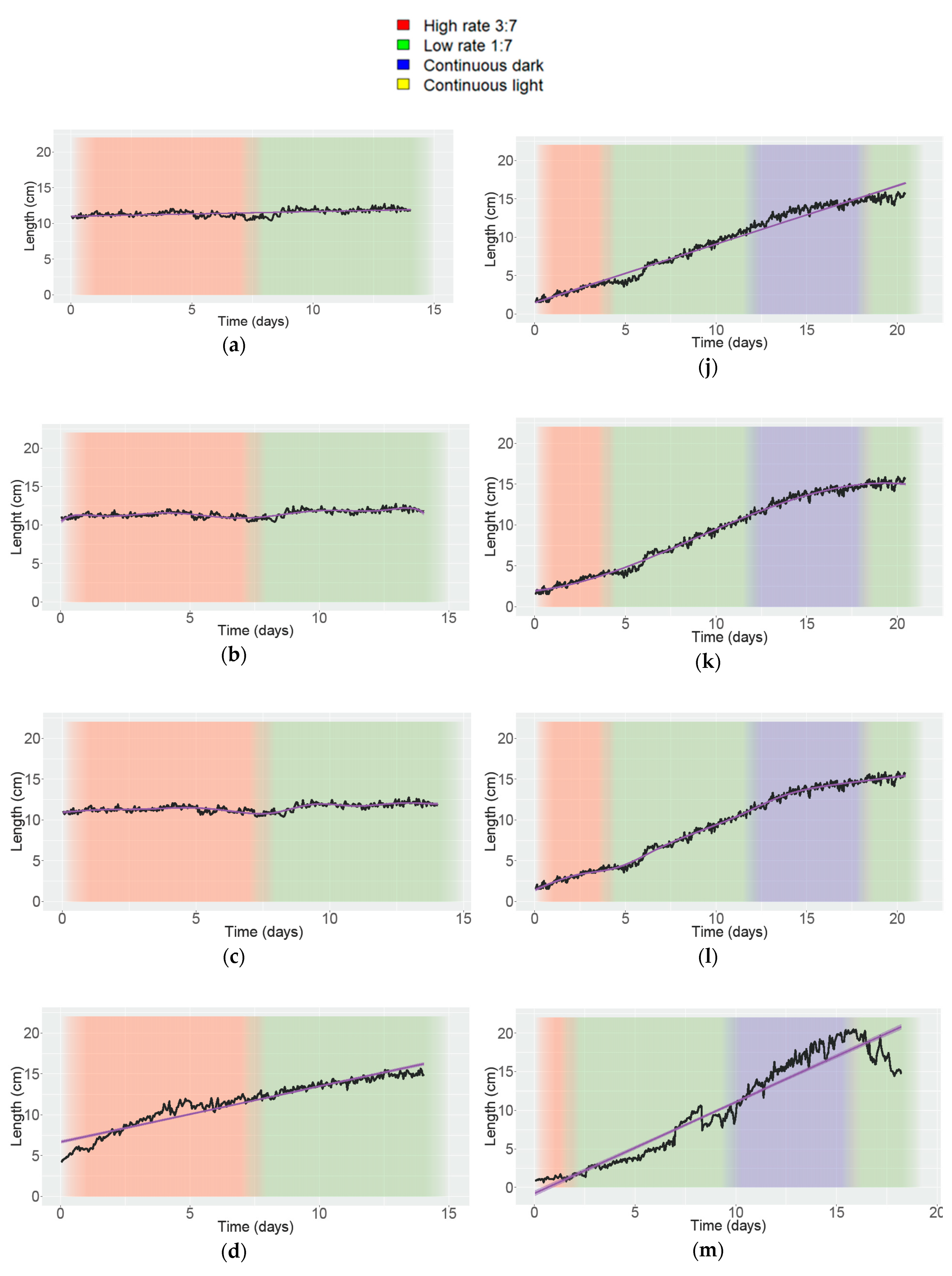
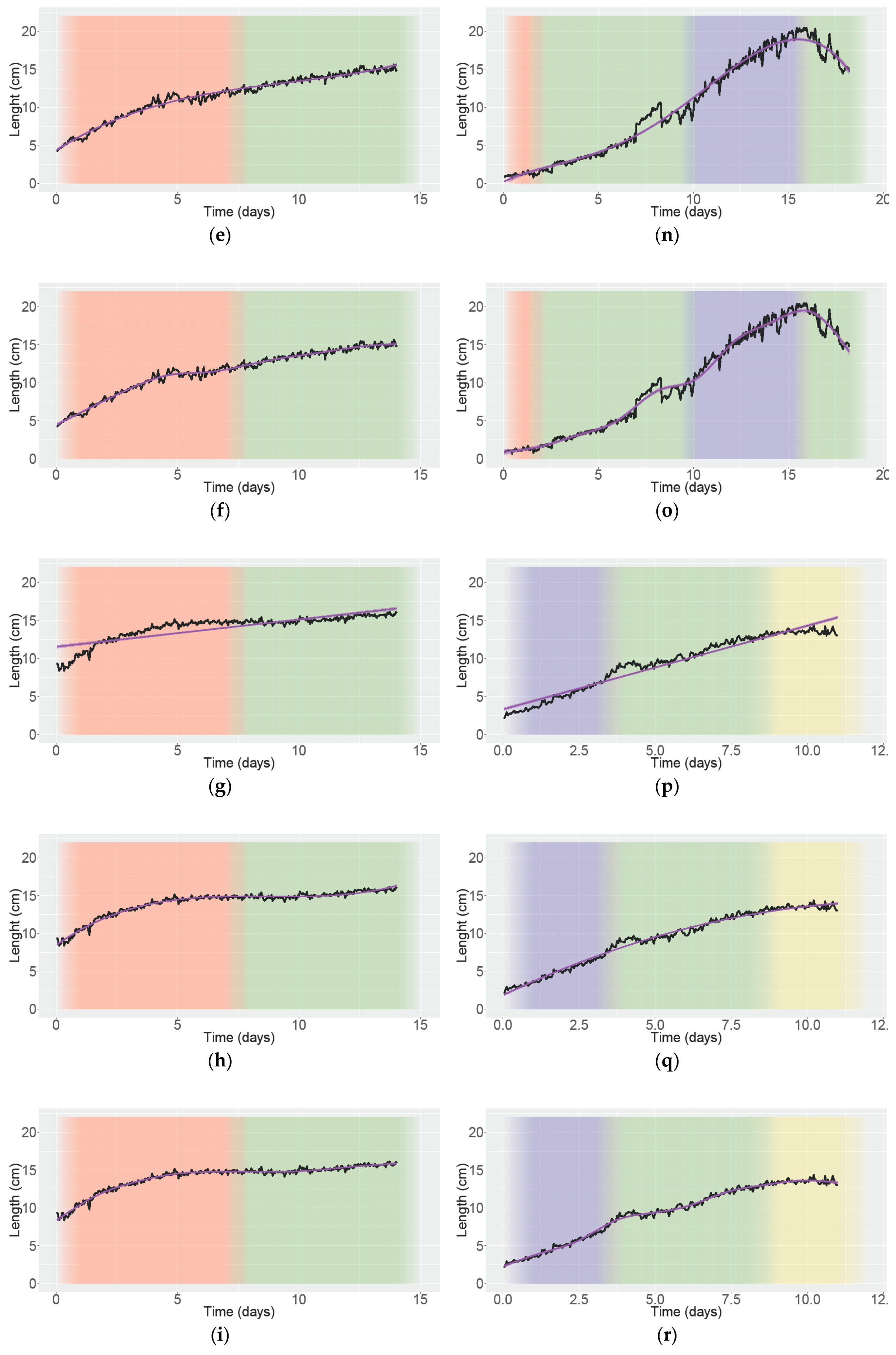
3.4. Analyzing Growth and Movement in Strawberry
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Darwin, C. On the movements and habits of climbing plants. J. Linn. Soc. Lond. 1865, 9, 1–118. [Google Scholar] [CrossRef]
- Schaffner, J.H. Observations on the nutation of Helianthus annuus. Bot. Gaz. 1898, 25, 395–403. [Google Scholar] [CrossRef]
- Barkai, N.; Leibler, S. Circadian clocks limited by noise. Nature 2000, 403, 267–268. [Google Scholar] [CrossRef] [PubMed]
- Hotta, C.T.; Gardner, M.J.; Hubbard, K.E.; Baek, S.J.; Dalchau, N.; Suhita, D.; Dodd, A.N.; Webb, A.A.R. Modulation of environmental responses of plants by circadian clocks. Plant Cell Environ. 2007, 30, 333–349. [Google Scholar] [CrossRef] [PubMed]
- Shim, J.S.; Imaizumi, T. Circadian clock and photoperiodic response in arabidopsis: From seasonal flowering to redox homeostasis. Biochemistry 2014, 54, 157–170. [Google Scholar] [CrossRef]
- Hecht, V.; Knowles, C.L.; Schoor, J.K.V.; Liew, L.C.; Jones, S.E.; Lambert, M.J.M.; Weller, J.L.; Oa, G.H.W. Pea LATE BLOOMER1 is a GIGANTEA ortholog with roles in photoperiodic flowering, deetiolation, and transcriptional regulation of circadian clock gene homologs. Plant Physiol. 2007, 144, 648–661. [Google Scholar] [CrossRef]
- Annunziata, M.G.; Apelt, F.; Carillo, P.; Krause, U.; Feil, R.; Koehl, K.; Lunn, J.E.; Stitt, M. Response of Arabidopsis primary metabolism and circadian clock to low night temperature in a natural light environment. J. Exp. Bot. 2018, 69, 4881–4895. [Google Scholar] [CrossRef]
- Gould, P.D.; Locke, J.C.W.; Larue, C.; Southern, M.M.; Davis, S.J.; Hanano, S.; Moyle, R.; Milich, R.; Putterill, J.; Millar, A.J.; et al. The molecular basis of temperature compensation in the Arabidopsis circadian clock. Plant Cell 2006, 18, 1177–1187. [Google Scholar] [CrossRef]
- McClung, C.R. Plant circadian rhythms. Plant Cell 2006, 18, 792–803. [Google Scholar] [CrossRef]
- Terry, M.I.; Pérez-Sanz, F.; Díaz-Galián, M.V.; Pérez de los Cobos, F.; Navarro, P.J.; Egea-Cortines, M.; Weiss, J. The Petunia CHANEL gene is a ZEITLUPE ortholog coordinating growth and scent profiles. Cells 2019, 8, 343. [Google Scholar] [CrossRef]
- Sun, X.; Cahill, J.; Van Hautegem, T.; Feys, K.; Whipple, C.; Novák, O.; Delbare, S.; Versteele, C.; Demuynck, K.; De Block, J. Altered expression of maize PLASTOCHRON1 enhances biomass and seed yield by extending cell division duration. Nat. Commun. 2017, 8, 14752. [Google Scholar] [CrossRef] [PubMed]
- Beemster, G.T.S.; De Vusser, K.; De Tavernier, E.; De Bock, K.; Inzé, D. Variation in growth rate between Arabidopsis ecotypes is correlated with cell division and A-type cyclin-dependent kinase activity. Plant Physiol. 2002, 129, 854–864. [Google Scholar] [CrossRef]
- Paine, C.E.T.; Marthews, T.R.; Vogt, D.R.; Purves, D.; Rees, M.; Hector, A.; Turnbull, L.A. How to fit nonlinear plant growth models and calculate growth rates: An update for ecologists. Methods Ecol. Evol. 2012, 3, 245–256. [Google Scholar] [CrossRef]
- Biskup, B.; Scharr, H.; Fischbach, A.; Wiese-Klinkenberg, A.; Schurr, U.; Walter, A. Diel growth cycle of isolated leaf discs analyzed with a novel, high-throughput three-dimensional imaging method is identical to that of intact leaves. Plant Physiol. 2009, 149, 1452–1461. [Google Scholar] [CrossRef] [PubMed]
- Nusinow, D.A.; Helfer, A.; Hamilton, E.E.; King, J.J.; Imaizumi, T.; Schultz, T.F.; Farré, E.M.; Kay, S.A.; Farre, E.M. The ELF4-ELF3-LUX complex links the circadian clock to diurnal control of hypocotyl growth. Nature 2011, 475, 398. [Google Scholar] [CrossRef]
- Yoshihara, T.; Iino, M. Circumnutation of rice coleoptiles: Effects of light and gravity. Plant Cell Physiol. 2003, 44, S16. [Google Scholar]
- Takagi, S.; Takano, T.; Wakamatsu, R. Circumnutation of azuki bean epicotyls. Plant Cell Physiol. 2003, 44, S15. [Google Scholar]
- Johnsson, A.; Solheim, B.G.B.; Iversen, T.-H. Gravity amplifies and microgravity decreases circumnutations in Arabidopsis thaliana stems: Results from a space experiment. New Phytol. 2009, 182, 621–629. [Google Scholar] [CrossRef]
- Hennessey, T.L.; Field, C.B. Evidence of multiple circadian oscillators in bean-plants. J. Biol. Rhythm. 1992, 7, 105–113. [Google Scholar] [CrossRef]
- Hoshizaki, T.; Hamner, K.C. Circadian leaf movements: Persistence in bean plants grown in continuous high-intensity light. Science 1964, 144, 1240–1241. [Google Scholar] [CrossRef]
- Müller, N.A.; Wijnen, C.L.; Srinivasan, A.; Ryngajllo, M.; Ofner, I.; Lin, T.; Ranjan, A.; West, D.; Maloof, J.N.; Sinha, N.R.; et al. Domestication selected for deceleration of the circadian clock in cultivated tomato. Nat. Genet. 2016, 48, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Bollig, I. Different circadian rhythms regulate photoperiodic flowering response and leaf movement in Pharbitis nil (L.) Choisy. Planta 1977, 135, 137–142. [Google Scholar] [CrossRef] [PubMed]
- Dornbusch, T.; Michaud, O.; Xenarios, I.; Fankhauser, C. Differentially phased leaf growth and movements in Arabidopsis depend on coordinated circadian and light regulation. Plant Cell 2014, 26, 3911–3921. [Google Scholar] [CrossRef] [PubMed]
- Egea-Cortines, M.; Ruíz-Ramón, F.; Weiss, J.; Ruiz-Ramon, F.; Weiss, J. Circadian regulation of horticultural traits: Integration of environmental signals in plants. In Horticultural Reviews; Janick, J., Ed.; Wiley: Hoboken, NJ, USA, 2013; Volume 41, pp. 1–46. ISBN 978-1-118-70741-8. [Google Scholar]
- Egea-Cortines, M.; Doonan, J. Editorial: Phenomics. Front. Plant Sci. 2018, 9, 678. [Google Scholar] [CrossRef]
- Behmann, J.; Mahlein, A.-K.; Rumpf, T.; Römer, C.; Plümer, L. A review of advanced machine learning methods for the detection of biotic stress in precision crop protection. Precis. Agric. 2015, 16, 239–260. [Google Scholar] [CrossRef]
- Deligiannidis, L.; Arabnia, H.R. Emerging Trends in Image Processing, Computer Vision and Pattern Recognition; Morgan Kaufmann: Burlington, MA, USA, 2014. [Google Scholar]
- Li, L.; Zhang, Q.; Huang, D. A review of imaging techniques for plant phenotyping. Sensors 2014, 14, 20078–20111. [Google Scholar] [CrossRef]
- Perez-Sanz, F.; Navarro, P.J.; Egea-Cortines, M. Plant phenomics: An overview of image acquisition technologies and image data analysis algorithms. GigaScience 2017, 6, 1–18. [Google Scholar] [CrossRef]
- Furbank, R.T. Plant phenomics: From gene to form and function. Funct. Plant Biol. 2009, 36, 5–6. [Google Scholar]
- Fiorani, F.; Rascher, U.; Jahnke, S.; Schurr, U. Imaging plants dynamics in heterogenic environments. Curr. Opin. Biotechnol. 2012, 23, 227–235. [Google Scholar] [CrossRef]
- Millar, A.J.; Carre, I.A.; Strayer, C.A.; Chua, N.-H.; Kay, S.A. Circadian clock mutants in Arabidopsis identified by luciferase imaging. Science 1995, 267, 1161–1163. [Google Scholar] [CrossRef]
- Xiong, L.; Lee, B.; Ishitani, M.; Lee, H.; Zhang, C.; Zhu, J.-K. FIERY1 encoding an inositol polyphosphate 1-phosphatase is a negative regulator of abscisic acid and stress signaling in Arabidopsis. Genes Dev. 2001, 15, 1971–1984. [Google Scholar] [CrossRef] [PubMed]
- Terry, M.I.; Pérez-Sanz, F.; Navarro, P.J.; Weiss, J.; Egea-Cortines, M. The Snapdragon LATE ELONGATED HYPOCOTYL plays a dual role in activating floral growth and scent emission. Cells 2019, 8, 920. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Hernández, V.; Hermans, B.; Weiss, J.; Egea-Cortines, M. Genetic analysis of natural variation in antirrhinum scent profiles identifies benzoic acid carboxymethyl transferase as the major locus controlling methyl benzoate synthesis. Front. Plant Sci. 2017, 8, 27. [Google Scholar] [CrossRef] [PubMed]
- Koes, R. Evolution and development of virtual inflorescences. Trends Plant Sci. 2008, 13, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Bombarely, A.; Moser, M.; Amrad, A.; Bao, M.; Bapaume, L.; Barry, C.S.; Bliek, M.; Boersma, M.R.; Borghi, L.; Bruggmann, R. Insight into the evolution of the Solanaceae from the parental genomes of Petunia hybrida. Nat. Plants 2016, 2, 16074. [Google Scholar] [CrossRef]
- Shulaev, V.; Sargent, D.J.; Crowhurst, R.N.; Mockler, T.C.; Folkerts, O.; Delcher, A.L.; Jaiswal, P.; Mockaitis, K.; Liston, A.; Mane, S.P. The genome of woodland strawberry (Fragaria vesca). Nat. Genet. 2011, 43, 109. [Google Scholar] [CrossRef]
- Schwarz-Sommer, Z.; Gübitz, T.; Weiss, J.; Gómez-di-Marco, P.; Delgado-Benarroch, L.; Hudson, A.; Egea-Cortines, M. A molecular recombination map of Antirrhinum majus. BMC Plant Biol. 2010, 10, 275. [Google Scholar] [CrossRef]
- Mitchell, A.Z.; Hanson, M.R.; Skvirsky, R.C.; Ausubel, F.M. Anther culture of Petunia: Genotypes with high frequency of callus, root, or plantlet formation. Z. Für Pflanzenphysiol. 1980, 100, 131–145. [Google Scholar] [CrossRef]
- Available online: https://www.ntesistemas.es/ (accessed on 26 November 2016).
- Navarro, P.; Fernández, C.; Weiss, J.; Egea-Cortines, M. Development of a configurable growth chamber with a computer vision system to study circadian rhythm in plants. Sensors 2012, 12, 15356–15375. [Google Scholar] [CrossRef]
- Valverde, F.; Mouradov, A.; Soppe, W.; Ravenscroft, D.; Samach, A.; Coupland, G. Photoreceptor regulation of CONSTANS protein in photoperiodic flowering. Science 2004, 303, 1003–1006. [Google Scholar] [CrossRef]
- Wu, G.; Anafi, R.C.; Hughes, M.E.; Kornacker, K.; Hogenesch, J.B. MetaCycle: An integrated R package to evaluate periodicity in large scale data. Bioinformatics 2016, 32, 3351–3353. [Google Scholar] [CrossRef] [PubMed]
- Hughes, M.E.; Hogenesch, J.B.; Kornacker, K. JTK_CYCLE: An efficient nonparametric algorithm for detecting rhythmic components in genome-scale data sets. J. Biol. Rhythm. 2010, 25, 372–380. [Google Scholar] [CrossRef] [PubMed]
- Kodinariya, T.M.; Makwana, P.R. Review on determining number of cluster in K-Means clustering. Int. J. Adv. Res. Comput. Sci. Manag. Stud. 2013, 1, 90–95. [Google Scholar]
- Leister, D.; Varotto, C.; Pesaresi, P.; Niwergall, A.; Salamini, F. Large-scale evaluation of plant growth in Arabidopsis thaliana by non-invasive image analysis. Plant Physiol. Biochem. 1999, 37, 671–678. [Google Scholar] [CrossRef]
- Golzarian, M.R.; Frick, R.A.; Rajendran, K.; Berger, B.; Roy, S.; Tester, M.; Lun, D.S.; Tackenberg, O.; Niklas, K.; Enquist, B.; et al. Accurate inference of shoot biomass from high-throughput images of cereal plants. Plant Methods 2011, 7, 2. [Google Scholar] [CrossRef]
- Chen, D.; Neumann, K.; Friedel, S.; Kilian, B.; Chen, M.; Altmann, T.; Klukas, C. Dissecting the phenotypic components of crop plant growth and drought responses based on high-throughput image analysis. Plant Cell 2014, 26, 4636–4655. [Google Scholar] [CrossRef]
- Fung-Uceda, J.; Lee, K.; Seo, P.J.; Polyn, S.; De Veylder, L.; Mas, P. The circadian clock sets the time of DNA replication licensing to regulate growth in Arabidopsis. Dev. Cell 2018, 45, 101–113. [Google Scholar] [CrossRef]
- Nieto, C.; López-Salmerón, V.; Davière, J.-M.; Prat, S. ELF3-PIF4 interaction regulates plant growth independently of the evening complex. Curr. Biol. 2015, 25, 187–193. [Google Scholar] [CrossRef]
- Harmer, S.L.; Brooks, C.J. Growth-mediated plant movements: Hidden in plain sight. Curr. Opin. Plant Biol. 2018, 41, 89–94. [Google Scholar] [CrossRef]
- Stolarz, M. Circumnutation as a visible plant action and reaction. Plant Signal. Behav. 2009, 4, 380–387. [Google Scholar] [CrossRef]
- Buda, A.; Zawadzki, T.; Krupa, M.; Stolarz, M.; Okulski, W. Daily and infradian rhythms of circumnutation intensity in Helianthus annuus. Physiol. Plant. 2003, 119, 582–589. [Google Scholar] [CrossRef]
- Britz, S.; Galston, A. Physiology of movements in the stems of seedling Pisum sativum L. cv Alaska III. Phototropism in relation to gravitropism, nutation, and growth. Plant Physiol. 1983, 71, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Yoshihara, T.; Iino, M. Circumnutation of rice coleoptiles: Its relationships with gravitropism and absence in lazy mutants. Plant Cell Environ. 2006, 29, 778–792. [Google Scholar] [CrossRef] [PubMed]
- Gaiser, J.C.; Lomax, T.L. The altered gravitropic response of the lazy-2 mutant of tomato is phytochrome regulated. Plant Physiol. 1993, 102, 339–344. [Google Scholar] [CrossRef] [PubMed]
- Yoshihara, T.; Spalding, E.P.; Iino, M. AtLAZY1 is a signaling component required for gravitropism of the Arabidopsis thaliana inflorescence. Plant J. 2013, 74, 267–279. [Google Scholar] [CrossRef] [PubMed]
- Steppe, K.; Sterck, F.; Deslauriers, A. Diel growth dynamics in tree stems: Linking anatomy and ecophysiology. Trends Plant Sci. 2015, 20, 335–343. [Google Scholar] [CrossRef]
- Matos, D.A.; Cole, B.J.; Whitney, I.P.; MacKinnon, K.J.-M.; Kay, S.A.; Hazen, S.P. Daily changes in temperature, not the circadian clock, regulate growth rate in Brachypodium distachyon. PLoS ONE 2014, 9, e100072. [Google Scholar] [CrossRef]
- Charzewska, A.; Zawadzki, T. Circadian modulation of circumnutation length, period, and shape in Helianthus annuus. J. Plant Growth Regul. 2006, 25, 324–331. [Google Scholar] [CrossRef]
- Huang, H.; Yoo, C.Y.; Bindbeutel, R.; Goldsworthy, J.; Tielking, A.; Alvarez, S.; Naldrett, M.J.; Evans, B.S.; Chen, M.; Nusinow, D.A. PCH1 integrates circadian and light-signaling pathways to control photoperiod-responsive growth in Arabidopsis. eLife 2016, 5, e13292. [Google Scholar] [CrossRef]
- Terry, M.I.; Carrera-Alesina, M.; Weiss, J.; Egea-Cortines, M. Transcriptional structure of Petunia clock in leaves and petals. Genes 2019, 10, 860. [Google Scholar] [CrossRef]
- Niinuma, K.; Fujiwara, S.; Oda, A.; Tajima, T.; Calvino, M.; Ohkoshi, Y.; Yoshida, R.; Nakamichi, N.; Mizuno, T.; Kamada, H.; et al. Roles of a circadian clock in the photoperiodic flowering and organ-movements in Arabidopsis. Plant Cell Physiol. 2006, 47, S1. [Google Scholar]
- Bastien, R.; Meroz, Y. The kinematics of plant nutation reveals a simple relation between curvature and the orientation of differential growth. PLoS Comput. Biol. 2016, 12, e1005238. [Google Scholar] [CrossRef] [PubMed]
- Watson, A.; Ghosh, S.; Williams, M.J.; Cuddy, W.S.; Simmonds, J.; Rey, M.-D.; Hatta, M.A.M.; Hinchliffe, A.; Steed, A.; Reynolds, D.; et al. Speed breeding is a powerful tool to accelerate crop research and breeding. Nat. Plants 2018, 4, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Watson, A.; Gonzalez-Navarro, O.E.; Ramirez-Gonzalez, R.H.; Yanes, L.; Mendoza-Suárez, M.; Simmonds, J.; Wells, R.; Rayner, T.; Green, P.; et al. Speed breeding in growth chambers and glasshouses for crop breeding and model plant research. Nat. Protoc. 2018, 13, 2944–2963. [Google Scholar] [CrossRef] [PubMed]
- Morrow, R.C. LED lighting in horticulture. HortScience 2008, 43, 1947–1950. [Google Scholar] [CrossRef]
- Choi, H.G.; Moon, B.Y.; Kang, N.J. Effects of LED light on the production of strawberry during cultivation in a plastic greenhouse and in a growth chamber. Sci. Hortic. 2015, 189, 22–31. [Google Scholar] [CrossRef]
- Kang, W.H.; Park, J.S.; Park, K.S.; Son, J.E. Leaf photosynthetic rate, growth, and morphology of lettuce under different fractions of red, blue, and green light from light-emitting diodes (LEDs). Hortic. Environ. Biotechnol. 2016, 57, 573–579. [Google Scholar] [CrossRef]

| Experiment | #1 | #2 | #3 |
|---|---|---|---|
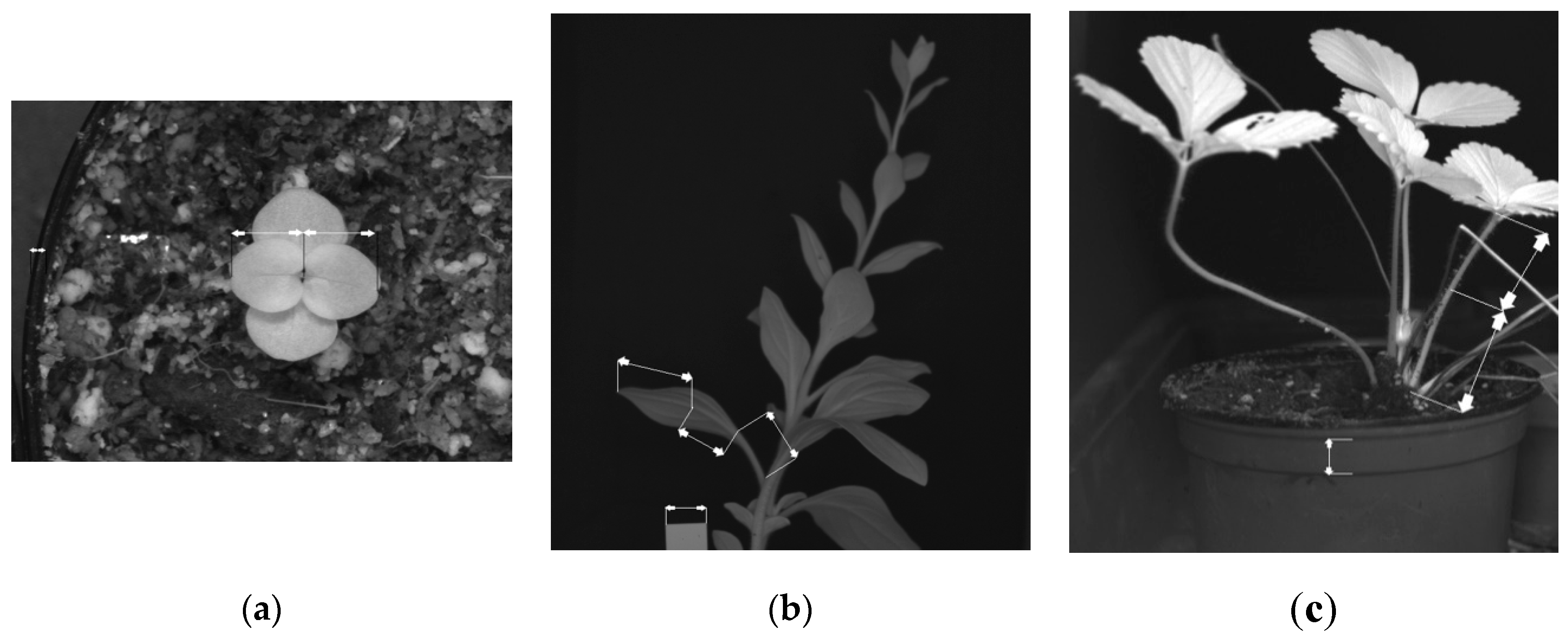
| Species studied | A. linkianum and A. majus | Petunia x hybrida W115 Mitchell | Fragaria x ananassa cv. Fortuna |
| Photoperiod | 12:12 LD | 12:12 LD 24:0 LD 0:24 LD | 12:12 LD 24:0 LD 0:24 LD |
| Thermoperiod | 23:18 °C LD | 23:18 °C LD | 23:8 °C LD |
| Duration of study | 10 days (48 h studied) | 12.5 days | 30 days |
| Growth stage | Seedling | Plant | Plant |
| Images/hour studied | 6 | 1 | 1 |
| Total number of images analyzed | 288 | 300 | 720 |
| Organ studied | Leaf and whole seedling | Leaf + Petiole | Petiole |
| Top/Side view | Top | Side | Side |
| N° of organs studied | 2 per species | 2 | 6 |
| Parameters studied | Movement and Growth | Angle and Movement speed | Movement and Growth |
| Study | Movement Speed | Growth | ||||||||||
| Species | A. linkianum | A. majus | A. linkianum | A. majus | ||||||||
| N° of parameters | 4 | 6 | 5 | 4 | ||||||||
| 2 | 8 | |||||||||||
| Petunia x hybrida | ||||||||||||
| Study | Angle | Speed movement | ||||||||||
| Condition | 12/12 h LD | 24 h Light | 24 h Dark | 12/12 h LD | 24 h Light | 24 h Dark | ||||||
| N° of parameters (Leaf 1 and Leaf 2) | 8 | 6 | 7 | 8 | 2 | 7 | 5 | 4 | 7 | 5 | 5 | 6 |
| Condition | 12/12 h LD-24 h Dark | 24 h Dark-12/12 h LD | 12/12 h LD-24 h Light | 24 h Light-12/12 h LD | ||||||||
| (Leaf 1 and Leaf 2) | 6 | 5 | 7 | 7 | 8 | 1 | 4 | 3 | ||||
| Fragaria ananassa | ||||||||||||
| Stem # | #1 | #2 | #3 | #4 | #5 | #6 | ||||||
| N° of parameters | 8 | 3 | 3 | 3 | 4 | 2 | ||||||
| Trend Line | Adjusted R2/R2 | |||||
|---|---|---|---|---|---|---|
| Movement Speed | Growth | |||||
| A. linkianum | A. majus | A. linkianum | A. majus | |||
| Linear | 0.00/0.00 | 0.03/0.03 | 0.38/0.39 | 0.81/0.81 | 0.83/0.83 | 0.87/0.87 |
| Exponential | 0.01/0.01 | 0.00/0.00 | 0.00/0.00 | 0.15/0.16 | 0.09/0.09 | 0.11/0.11 |
| Logarithmic | 0.00/0.01 | 0.03/0.04 | 0.27/0.28 | 0.52/0.52 | 0.60/0.60 | 0.64/0.64 |
| Polynomial | 0.27/0.28 | 0.27/0.29 | 0.56/0.57 | 0.86/0.86 | 0.87/0.87 | 0.88/0.89 |
| GAM | 0.43/0.44 | 0.28/0.29 | 0.68/0.69 | 0.91/0.91 | 0.89/0.89 | 0.90/0.90 |
| Trend Line | Adjusted R2/R2 | |||||
|---|---|---|---|---|---|---|
| 12/12 h Night/Day | 24 h Day | 24 h Night | ||||
| Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | |
| Linear | 0.18/0.19 | 0.03/0.04 | 0.50/0.51 | 0.62/0.62 | 0.35/0.36 | 0.81/0.81 |
| Exponential | 0.03/0.05 | 0.00/0.01 | 0.00/0.00 | 0.02/0.03 | 0.01/0.02 | 0.07/0.08 |
| Logarithmic | 0.12/0.13 | 0.01/0.03 | 0.79/0.80 | 0.83/0.83 | 0.72/0.72 | 0.52/0.53 |
| Polynomial | 0.80/0.82 | 0.40/0.45 | 0.76/0.77 | 0.89/0.90 | 0.95/0.95 | 0.88/0.89 |
| GAM | 0.92/0.93 | 0.94/0.94 | 0.94/0.95 | 0.86/0.88 | 0.92/0.93 | 0.86/0.88 |
| Trend Line | Adjusted R2/R2 | |||||||
|---|---|---|---|---|---|---|---|---|
| 12/12 h DL 24 h DD | 24 h DD 12/12 h DL | 12/12 h DL 24 h LL | 24 h LL 12/12 h DL | |||||
| Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | |
| Linear | 0.07/0.09 | 0.12/0.13 | 0.22/0.23 | 0.18/0.19 | 0.30/0.31 | 0.57/0.57 | 0.22/0.23 | 0.66/0.67 |
| Exponential | 0.00/0.01 | 0.00/0.00 | 0.02/0.04 | 0.00/0.02 | 0.04/0.05 | 0.03/0.05 | 0.00/0.01 | 0.01/0.02 |
| Logarithmic | 0.02/0.04 | 0.24/0.25 | 0.12/0.13 | 0.07/0.09 | 0.11/0.13 | 0.35/0.36 | 0.12/0.13 | 0.50/0.51 |
| Polynomial | 0.23/0.29 | 0.55/0.58 | 0.77/0.79 | 0.74/0.77 | 0.86/0.87 | 0.57/0.57 | 0.27/0.31 | 0.78/0.79 |
| GAM | 0.60/0.65 | 0.62/0.66 | 0.94/0.94 | 0.93/0.94 | 0.92/0.93 | 0.90/0.91 | 0.26/0.29 | 0.87/0.89 |
| BH.Q | ADJ.P | Period(h) | LAG(phase)(h) | Amplitude | ||
|---|---|---|---|---|---|---|
| 12/12 h L/D | Leaf 1 | 4.4363e-18 | 4.4363e-18 | 24 | 21 | 49.8351 |
| Leaf 2 | 1.3976e-20 | 6.9878e-21 | 25 | 21 | 18.2425 | |
| Continuous light | Leaf 1 | 1 | 1 | 25 | 14 | 1.6507 |
| Leaf 2 | 1 | 1 | 26 | 8.5 | 5.4527 | |
| Continuous dark | Leaf 1 | 0.3894 | 0.1947 | 21 | 3 | 9.7137 |
| Leaf 2 | 1 | 1 | 20 | 6 | 2.1682 |
| Trend Line | Adjusted R2/R2 | |||||
|---|---|---|---|---|---|---|
| 12/12 h Night/Day | 24 h Day | 24 h Night | ||||
| Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | Leaf 1 | Leaf 2 | |
| Linear | 0.01/0.03 | 0.06/0.07 | 0.05/0.07 | 0.06/0.07 | 0.05/0.07 | 0.06/0.07 |
| Exponential | 0.00/0.02 | 0.00/0.00 | 0.00/0.01 | 0.00/0.00 | 0.0.0/0.01 | 0.00/0.00 |
| Logarithmic | 0.04/0.05 | 0.09/0.11 | 0.05/0.06 | 0.21/0.22 | 0.05/0.06 | 0.21/0.22 |
| Polynomial | 0.05/0.12 | 0.16/0.23 | 0.21/0.23 | 0.46/0.50 | 0.02/0.09 | 0.37/0.40 |
| GAM | 0.01/0.02 | 0.23/0.30 | 0.05/0.07 | 0.40/0.46 | 0.05/0.07 | 0.40/0.46 |
| BH.Q | ADJ.P | Period(h) | LAG(phase)(h) | Amplitude | ||
|---|---|---|---|---|---|---|
| 12/12 h L/D | Leaf 1 | 1.1192e-14 | 1.1192e-14 | 12 | 2.5 | 0.3297 |
| Leaf 2 | 0.0461 | 0.0461 | 26 | 21 | 0.0627 | |
| 24 h light | Leaf 1 | 0.1415 | 0.2830 | 27 | 5.5 | 0.0524 |
| Leaf 2 | 0.6317 | 0..6317 | 26 | 4 | 0.0194 | |
| 24 h dark | Leaf 1 | 1 | 1 | 20 | 4 | 0.0308 |
| Leaf 2 | 1 | 1 | 24 | 7 | 0.0214 |
| Trend Line | Adjusted R2/R2 | |||||
|---|---|---|---|---|---|---|
| A | B | C | D | E | F | |
| Linear | 0.32/0.32 | 0.92/0.92 | 0.70/0.70 | 0.97/0.97 | 0.93/0.93 | 0.85/0.85 |
| Exponential | 0.19/0.19 | 0.23/0.23 | 0.14/0.14 | 0.18/0.18 | 0.13/0.13 | 0.27/0.28 |
| Logarithmic | 0.20/0.20 | 0.91/0.91 | 0.92/0.92 | 0.77/0.77 | 0.69/0.69 | 0.85/0.85 |
| Polynomial | 0.66/0.67 | 0.98/0.98 | 0.97/0.97 | 0.99/0.99 | 0.98/0.98 | 0.99/0.99 |
| GAM | 0.61/0.62 | 0.99/0.99 | 0.98/0.98 | 0.99/0.99 | 0.99/0.99 | 0.99/0.99 |
| A | B | C | D | E | F | |
|---|---|---|---|---|---|---|
| Area Under the Curve (AUC) (cm × h) | 160.48 | 160.48 | 196.91 | 189.76 | 182.58 | 102.85 |
| Maximum slope | 0.86 | 1.60 | 2.19 | 1.15 | 2.77 | 2.39 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Díaz-Galián, M.V.; Perez-Sanz, F.; Sanchez-Pagán, J.D.; Weiss, J.; Egea-Cortines, M.; Navarro, P.J. A Proposed Methodology to Analyze Plant Growth and Movement from Phenomics Data. Remote Sens. 2019, 11, 2839. https://doi.org/10.3390/rs11232839
Díaz-Galián MV, Perez-Sanz F, Sanchez-Pagán JD, Weiss J, Egea-Cortines M, Navarro PJ. A Proposed Methodology to Analyze Plant Growth and Movement from Phenomics Data. Remote Sensing. 2019; 11(23):2839. https://doi.org/10.3390/rs11232839
Chicago/Turabian StyleDíaz-Galián, María Victoria, Fernando Perez-Sanz, Jose David Sanchez-Pagán, Julia Weiss, Marcos Egea-Cortines, and Pedro J. Navarro. 2019. "A Proposed Methodology to Analyze Plant Growth and Movement from Phenomics Data" Remote Sensing 11, no. 23: 2839. https://doi.org/10.3390/rs11232839
APA StyleDíaz-Galián, M. V., Perez-Sanz, F., Sanchez-Pagán, J. D., Weiss, J., Egea-Cortines, M., & Navarro, P. J. (2019). A Proposed Methodology to Analyze Plant Growth and Movement from Phenomics Data. Remote Sensing, 11(23), 2839. https://doi.org/10.3390/rs11232839







