The MicroBioDiverSar Project: Exploring the Microbial Biodiversity in Ex Situ Collections of Sardinia
Abstract
1. Introduction
2. Database Design and Implementation
2.1. The Database Architecture
2.2. The Database Information and Schema
2.3. The Features of the Applications: On-Line Integrated Query
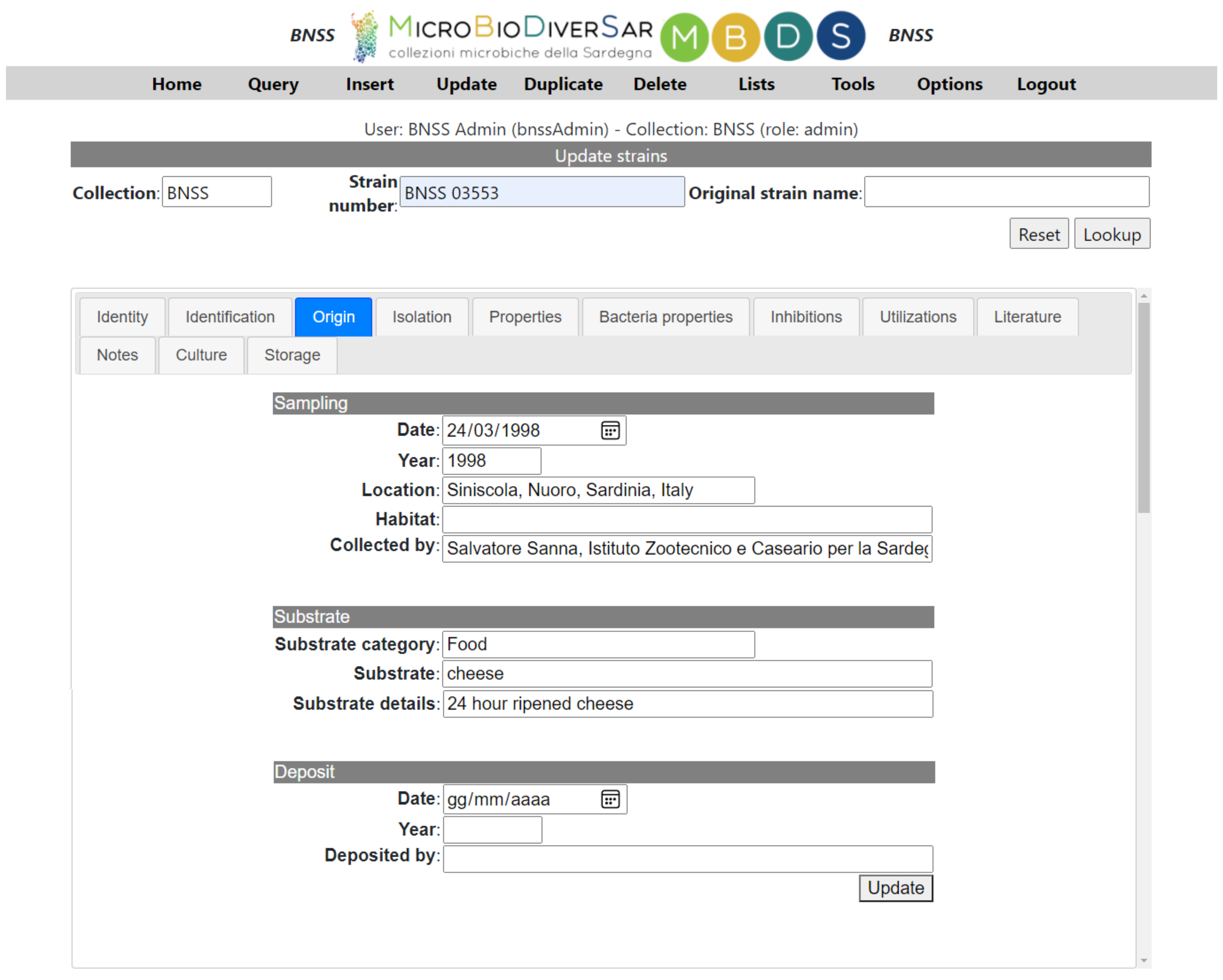
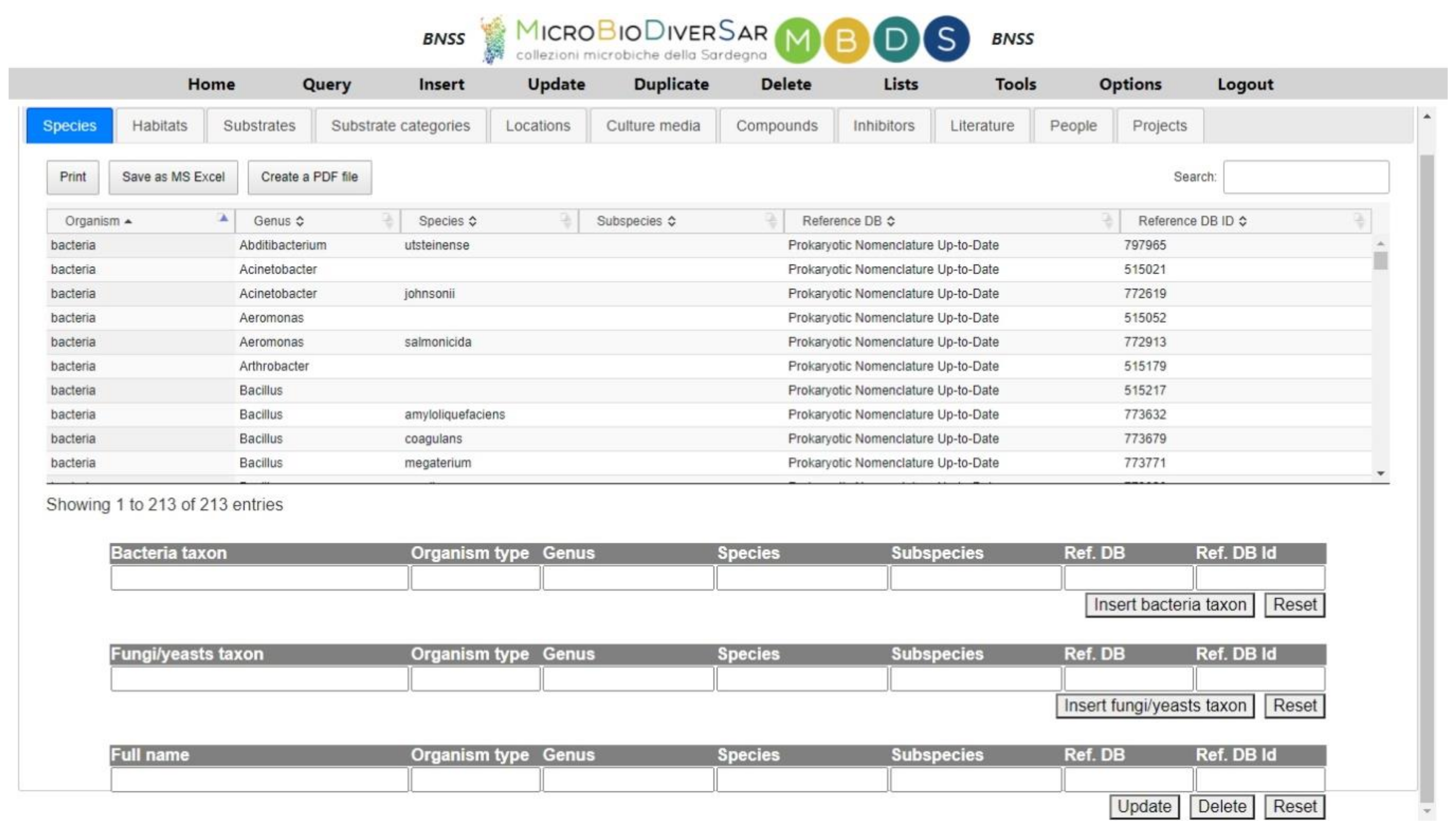
2.4. The Features of the Applications: Local Database Management
3. The MBDS Microbial Collection
3.1. BNSS Agris Sardegna Collection
3.2. Uniss Collection
3.3. Unica Collection
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Food and Agriculture Organization of the United Nations (FAO). The State of the World’s Biodiversity for Food and Agriculture; FAO Commission on Genetic Resources for Food and Agriculture Assessments: Rome, Italy, 2019; Available online: http://www.fao.org/3/CA3129EN/CA3129EN.pdf (accessed on 24 June 2021).
- Food and Agriculture Organization of the United Nations (FAO). Agricultural Biodiversity, Multifunctional Character of Agriculture and Land Conference, Background Paper 1; FAO: Rome, Italy, 1999. [Google Scholar]
- Ministero delle Politiche Agricole, Alimentari e Forestali. Linee Guida per la Conservazione e la Caratterizzazione Della Biodiversità Vegetale, Animale e Microbica di Interesse per L’agricoltura; Ministero delle Politiche Agricole, Alimentari e Forestali: Rome, Italy, 2013. [Google Scholar]
- Barragán-Ocaña, A.; Silva-Borjas, P.; Olmos-Peña, S.; Polanco-Olguín, M. Biotechnology and Bioprocesses: Their Contribution to Sustainability. Processes 2020, 8, 436. [Google Scholar] [CrossRef]
- McCluskey, K. A Review of Living Collections with Special Emphasis on Sustainability and Its Impact on Research across Multiple Disciplines. Biopreser. Biobank. 2017, 15, 20–30. [Google Scholar] [CrossRef]
- De Vero, L.; Boniotti, M.B.; Budroni, M.; Buzzini, P.; Cassanelli, S.; Comunian, R.; Gullo, M.; Logrieco, A.F.; Mannazzu, I.; Musumeci, R.; et al. Preservation, Characterization and Exploitation of Microbial Biodiversity: The Perspective of the Italian Network of Culture Collections. Microorganisms 2019, 7, 685. [Google Scholar] [CrossRef]
- OECD 2007. Best Practice Guidelines for Biological Resource Centres. Available online: https://www.oecd.org/sti/emerging-tech/38777417.pdf (accessed on 18 June 2021).
- CABRI. Guidelines for Catalogue Production. Available online: http://www.cabri.org/guidelines/catalogue/CPcover.html (accessed on 18 June 2021).
- Romano, P.; Kracht, M.; Manniello, M.A.; Stegehuis, G.; Fritze, D. The role of informatics in the coordinated management of biological resources collections. Appl. Bioinform. 2005, 4, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Bottazzi, V.; Ledda, A. Microbiologia del formaggio pecorino “romano”. Ann. Microbiol. 1967, 17, 41–53. [Google Scholar]
- Ledda, A. Microbiologia del formaggio Pecorino “Romano”. Nota II: Caratteri fisiologici di lattobacilli termofili isolati da scotta-fermento. Sci. Tec. Latt. Casearia 1969, 20, 305–314. [Google Scholar]
- Ward, L.J.; Timmins, M.J. Differentiation of Lactobacillus casei, Lactobacillus paracasei and Lactobacillus rhamnosus by polymerase chain reaction. Lett. Appl. Microbiol. 1999, 29, 90–92. [Google Scholar] [CrossRef]
- Cremonesi, P.; Vanoni, L.; Morandi, S.; Silvetti, T.; Castiglioni, B.; Brasca, M. Development of a pentaplex PCR assay for the simultaneous detection of Streptococcus thermophilus, Lactobacillus delbrueckii subsp. bulgaricus, L. delbrueckii subsp. lactis, L. helveticus, L. fermentum in whey starter for Grana Padano cheese. Int. J. Food Microbiol. 2011, 146, 207–211. [Google Scholar] [CrossRef]
- Torriani, S.; Felis, G.E.; Dellaglio, F. Differentiation of Lactobacillus plantarum, L. pentosus, and L. paraplantarum by recA gene sequence analysis and multiplex PCR assay with recA gene-derived primers. Appl. Environ. Microbiol. 2001, 67, 3450–3454. [Google Scholar] [CrossRef]
- Hoppe-Seyler, T.S.; Jaeger, B.; Bockelmann, W.; Noordman, W.H.; Geis, A.; Heller, K.J. Molecular Identification and Differentiation of Staphylococcus Species and Strains of Cheese Origin. Syst. Appl. Microbiol. 2004, 27, 211–218. [Google Scholar] [CrossRef]
- Gevers, D.; Huys, G.; Swings, J. Applicability of rep-PCR fingerprinting for identification of Lactobacillus species. FEMS Microbiol. Lett. 2001, 205, 31–36. [Google Scholar] [CrossRef]
- Gosiewski, T.; Brzychczy-Wloch, M. The use of PFGE method in genotyping of selected bacteria species of the Lactobacillus genus. Methods Mol. Biol. 2015, 1301, 225–240. [Google Scholar] [CrossRef]
- Graves, L.M.; Swaminathan, B. PulseNet standardized protocol for subtyping Listeria monocytogenes by macrorestriction and pulsed-field gel electrophoresis. Int. J. Food Microbiol. 2001, 65, 55–62. [Google Scholar] [CrossRef]
- Tosi, L.; Berruti, G.; Danielsen, M.; Wind, A.; Huys, G.; Morelli, L. Susceptibility of Streptococcus thermophilus to antibiotics. Antonie Van Leeuwenhoek 2007, 92, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Scintu, M.F.; Mannu, L.; Mulargia, A.F.; Comunian, R.; Daga, E.; Paba, A.; Galistu, G. Microbiological Characteristics of Ewe’s Milk and Pecorino Romano PDO Cheese. The Challenge to Sheep and Goats Milk Sectors. Spec. Issue Int. Dairy Fed. 2007. Available online: http://store.fil-idf.org/wp-content/uploads/2016/12/0801Part4.pdf (accessed on 24 June 2021).
- Ledda, A.; Scintu, M.F.; Pirisi, A.; Sanna, S.; Mannu, L. Caratterizzazione tecnologica di ceppi di Lattococchi e di Enterococchi per la produzione di formaggio pecorino Fiore Sardo. Sci. Tec. Latt. Casearia 1994, 45, 443–456. [Google Scholar]
- Mannu, L.; Comunian, R.; Daga, E.; Paba, A.; Demuro, P.P.; Scintu, M.F. Isolamento e caratterizzazione della microflora naturale colonizzante il formaggio Fiore Sardo (DOP). Sci. Tec. Latt. Casearia 2006, 57, 445–454. [Google Scholar]
- Mannu, L.; Comunian, R.; Scintu, M.F. Mesophilic lactobacilli in Fiore Sardo cheese: PCR-identification and evolution during cheese ripening. Int. Dairy J. 2000, 10, 383–389. [Google Scholar] [CrossRef]
- Mannu, L.; Riu, G.; Comunian, R.; Fozzi, M.C.; Scintu, M.F. A preliminary study of lactic acid bacteria in whey starter culture and industrial Pecorino Sardo ewes’ milk cheese: PCR-identification and evolution during ripening. Int. Dairy J. 2002, 12, 17–26. [Google Scholar] [CrossRef]
- Mannu, L.; Paba, A.; Pes, M.; Scintu, M.F. Genotypic and phenotypic heterogeneity among lactococci isolated from traditional Pecorino Sardo cheese. J. Appl. Microbiol. 2000, 89, 191–197. [Google Scholar] [CrossRef]
- Arrizza, S.; Ledda, A.; Sarra, P.G.; Dellaglio, F. Identification of Lactic Acid Bacteria in Gioddu. Sci. Tec. Latt. Casearia 1983, 34, 87–102. [Google Scholar]
- Daga, E.; Mannu, L.; Porcu, S.; Comunian, R.; Paba, A.; Scintu, M.F. Home-made dry sausages produced in Sardinia: An investigation on the microflora. Ital. J. Food Sci. 2007, 19, 297. [Google Scholar]
- Comunian, R.; Ferrocino, I.; Paba, A.; Daga, E.; Campus, M.; Di Salvo, R.; Cauli, E.; Piras, F.; Zurru, R.; Cocolin, L. Evolution of microbiota during spontaneous and inoculated Tonda di Cagliari table olives fermentation and impact on sensory characteristics. LWT 2017, 84, 64–72. [Google Scholar] [CrossRef]
- Paba, A.; Chessa, L.; Daga, E.; Campus, M.; Bulla, M.; Angioni, A.; Sedda, P.; Comunian, R. Do Best-Selected Strains Perform Table Olive Fermentation Better than Undefined Biodiverse Starters? A Comparative Study. Foods 2020, 9, 135. [Google Scholar] [CrossRef]
- Chessa, L.; Paba, A.; Daga, E.; Dupré, I.; Comunian, R. Biodiversity and Safety Assessment of Half-Century Preserved Natural Starter Cultures for Pecorino Romano PDO Cheese. Microorganisms 2021, 9, 1363. [Google Scholar] [CrossRef]
- Chessa, L.; Paba, A.; Daga, E.; Comunian, R. Effect of growth media on natural starter culture composition and performance evaluated with a polyphasic approach. Int. J. Dairy Technol. 2019, 72, 152–158. [Google Scholar] [CrossRef]
- Floris, R.; Scanu, G.; Fois, N.; Rizzo, C.; Malavenda, R.; Spanò, N.; Lo Giudice, A. Intestinal bacterial flora of Mediterranean gilthead sea bream (Sparus aurata Linnaeus) as a novel source of natural surface active compounds. Aquac. Res. 2018, 49, 1262–1273. [Google Scholar] [CrossRef]
- Comunian, R.; Daga, E.; Dupré, I.; Paba, A.; Devirgiliis, C.; Piccioni, V.; Perozzi, G.; Zonenschain, D.; Rebecchi, A.; Morelli, L.; et al. Susceptibility to tetracycline and erythromycin of Lactobacillus paracasei strains isolated from traditional Italian fermented foods. Int. J. Food Microbiol. 2010, 138, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Comunian, R. Identification and Safety Assessment of Enterococchi Isolated from a Sardinian Ewe’s Raw Milk Pdo Cheese (Fiore Sardo). Ph.D. Dissertation, University of Sassari, Sassari, Italy, 2010. [Google Scholar]
- Mannu, L.; Paba, A.; Daga, E.; Comunian, R.; Zanetti, S.; Duprè, I.; Sechi, L.A. Comparison of the incidence of virulence determinants and antibiotic resistance between Enterococcus faecium strains of dairy, animal and clinical origin. Int. J. Food Microbiol. 2003, 88, 291–304. [Google Scholar] [CrossRef]
- Coi, A.L.; Bigey, F.; Mallet, S.; Marsit, S.; Zara, G.; Gladieux, P.; Galeote, V.; Budroni, M.; Dequin, S.; Legras, J.L. Genomic signatures of adaptation to wine biological ageing conditions in biofilm-forming flor yeasts. Mol. Ecol. 2017, 26, 2150–2166. [Google Scholar] [CrossRef]
- Zara, G.; Zara, S.; Pinna, C.; Marceddu, S.; Budroni, M. FLO11 gene length and transcriptional level affect biofilm-forming ability of wild flor strains of Saccharomyces cerevisiae. Microbiology 2009, 155, 3838–3846. [Google Scholar] [CrossRef]
- Mannazzu, I.; Clementi, F.; Ciani, M. Strategies and criteria for the isolation and selection of autochthonous starters. In Biodiversity and Biotechnology of Wine Yeasts; Research Signpost: Trivandrum, India, 2002; pp. 19–34. [Google Scholar]
- Mannazzu, I.; Domizio, P.; Carboni, G.; Zara, S.; Zara, G.; Comitini, F.; Budroni, M.; Ciani, M. Yeast killer toxins: From ecological significance to application. Crit. Rev. Biotechnol. 2019, 39, 603–617. [Google Scholar] [CrossRef]
- Legras, J.-L.; Moreno-Garcia, J.; Zara, S.; Zara, G.; Garcia-Martinez, T.; Mauricio, J.C.; Mannazzu, I.; Coi, A.L.; Bou Zeidan, M.; Dequin, S.; et al. Flor Yeast: New Perspectives Beyond Wine Aging. Front. Microbiol. 2016, 7, 503. [Google Scholar] [CrossRef]
- Zara, G.; Goffrini, P.; Lodi, T.; Zara, S.; Mannazzu, I.; Budroni, M. FLO11 expression and lipid biosynthesis are required for air-liquid biofilm formation in a Saccharomyces cerevisiae flor strain. FEMS Yeast Res. 2012, 12, 864–866. [Google Scholar] [CrossRef] [PubMed]
- Assandri, D.; Pampuro, N.; Zara, G.; Cavallo, E.; Budroni, M. Suitability of Composting Process for the Disposal and Valorization of Brewer’s Spent Grain. Agriculture 2021, 11, 2. [Google Scholar]
- Bianco, A.; Budroni, M.; Zara, S.; Mannazzu, I.; Fancello, F.; Zara, G. The role of microorganisms on biotransformation of brewers’ spent grain. Appl. Microbiol. Biotechnol. 2020, 104, 8661–8678. [Google Scholar] [CrossRef] [PubMed]
- Santona, M.; Sanna, M.L.; Multineddu, C.; Fancello, F.; de la Fuente, S.A.; Dettori, S.; Zara, S. Microbial biodiversity of Sardinian oleic ecosystems. Food Microbiol. 2018, 70, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Bianco, A.; Fancello, F.; Balmas, V.; Dettori, M.; Motroni, A.; Zara, G.; Budroni, M. Microbial communities and malt quality of durum wheat used in brewing. J. Inst. Brew. 2019, 125, 222–229. [Google Scholar] [CrossRef]
- Bianco, A.; Fancello, F.; Balmas, V.; Zara, G.; Dettori, M.; Budroni, M. The microbiome of Sardinian barley and malt. J. Inst. Brew. 2018, 124, 344–351. [Google Scholar] [CrossRef]
- Fancello, F.; Multineddu, C.; Santona, M.; Deiana, P.; Zara, G.; Mannazzu, I.; Budroni, M.; Dettori, S.; Zara, S. Bacterial Biodiversity of Extra Virgin Olive Oils and Their Potential Biotechnological Exploitation. Microorganisms 2020, 8, 97. [Google Scholar] [CrossRef]
- Marongiu, A.; Zara, G.; Legras, J.L.; Del Caro, A.; Mascia, I.; Fadda, C.; Budroni, M. Novel starters for old processes: Use of Saccharomyces cerevisiae strains isolated from artisanal sourdough for craft beer production at a brewery scale. J. Ind. Microbiol. Biotechnol. 2015, 42, 85–92. [Google Scholar] [CrossRef]
- Aru, V.; Pisano, M.B.; Savorani, F.; Engelsen, S.B.; Cosentino, S.; Cesare Marincola, F. Metabolomics analysis of shucked mussels’ freshness. Food Chem. 2016, 205, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Cosentino, S.; Fadda, M.E.; Deplano, M.; Mulargia, A.F.; Palmas, F. Yeasts associated with Sardinian ewe’s dairy products. Int. J. Food Microbiol. 2001, 69, 53–58. [Google Scholar] [CrossRef]
- Cosentino, S.; Matza, O.; Fadda, M.E.; Palmas, F. Hygienic and microbiological quality of starters and coagulants used in the production of sheep’s milk cheese. Ann. Ig. Med. Prev. Comun. 1989, 1, 1521–1528. [Google Scholar]
- Cosentino, S.; Tuberoso, C.; Meloni, V.; Cherchi, A.; Mulargia, A.F.; Porcu, M.; Palmas, F. Valorizzazione dei mieli tipici sardi: Aspetti microbiologici, botanici e fisico-chimici. Riv. Sci. Aliment. 1994, 23, 199–207. [Google Scholar]
- Palmas, F.; Carta, A.; Cosentino, S.; Fadda, M.E.; Giliberto, G.; Mulargia, A.F. Nuova tecnologia per la produzione di Ricotta ovina: Caratteristiche e valutazione della conservabilità. Riv. Sci. Aliment. 1994, 4, 467–472. [Google Scholar]
- Palmas, F.; Cosentino, S.; Fadda, M.E.; Deplano, M.; Mascia, V. Microbial characteristics of Pecorino processed cheese spreads. Lait 1999, 79, 607–613. [Google Scholar] [CrossRef]
- Pisano, M.B.; Deplano, M.; Fadda, M.E.; Cosentino, S. Microbiota of Sardinian Goat’s Milk and Preliminary Characterization of Prevalent LAB Species for Starter or Adjunct Cultures Development. BioMed Res. Int. 2019, 6131404. [Google Scholar] [CrossRef]
- Pisano, M.B.; Fadda, M.E.; Deplano, M.; Corda, A.; Cosentino, S. Microbiological and chemical characterization of Fiore Sardo, a traditional Sardinian cheese made from ewe’s milk. Int. J. Dairy Technol. 2006, 59, 171–179. [Google Scholar] [CrossRef]
- Pisano, M.B.; Scano, P.; Murgia, A.; Cosentino, S.; Caboni, P. Metabolomics and microbiological profile of Italian mozzarella cheese produced with buffalo and cow milk. Food Chem. 2016, 192, 618–624. [Google Scholar] [CrossRef]
- Palmas, F.; Fadda, M.E.; Sanseverino, P. Correlations between environmental contamination and microbiological quality of several milk-cheese products. Nuovi Ann. Ig. Microb. 1985, 36, 351–359. [Google Scholar]
- Muyzer, G.; de Waal, E.C.; Uitterlinden, A.G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E.; Nirenberg, H.I. Molecular Systematics and Phylogeography of the Gibberella fujikuroi Species Complex. Mycologia 1998, 90, 465–493. [Google Scholar] [CrossRef]
- Quere, F.; Deschamps, A.; Urdaci, M.C. DNA probe and PCR-specific reaction for Lactobacillus plantarum. J. Appl. Microbiol. 1997, 82, 783–790. [Google Scholar] [CrossRef] [PubMed]
- Young, J.P.; Downer, H.L.; Eardly, B.D. Phylogeny of the phototrophic rhizobium strain BTAi1 by polymerase chain reaction-based sequencing of a 16S rRNA gene segment. J. Bacteriol. 1991, 173, 2271–2277. [Google Scholar] [CrossRef]
- Saad, N.; Delattre, C.; Urdaci, M.; Schmitter, J.M.; Bressollier, P. An overview of the last advances in probiotic and prebiotic field. LWT Food Sci. Technol. 2013, 50, 1–16. [Google Scholar] [CrossRef]
- Cosentino, S.; Pisano, M.B.; Corda, A.; Fadda, M.E.; Piras, C. Genotypic and technological characterization of enterococci isolated from artisanal Fiore Sardo cheese. J. Dairy Res. 2004, 71, 444–450. [Google Scholar] [CrossRef] [PubMed]
- Fadda, M.E.; Mossa, V.; Deplano, M.; Pisano, M.B.; Cosentino, S. In vitro screening of Kluyveromyces strains isolated from Fiore Sardo cheese for potential use as probiotics. LWT 2017, 75, 100–106. [Google Scholar] [CrossRef]
- Fadda, M.E.; Mossa, V.; Pisano, M.B.; Deplano, M.; Cosentino, S. Occurrence and characterization of yeasts isolated from artisanal Fiore Sardo cheese. Int. J. Food Microbiol. 2004, 95, 51–59. [Google Scholar] [CrossRef]
- Fadda, M.E.; Viale, S.; Deplano, M.; Pisano, M.B.; Cosentino, S. Characterization of yeast population and molecular fingerprinting of Candida zeylanoides isolated from goat’s milk collected in Sardinia. Int. J. Food Microbiol. 2010, 136, 376–380. [Google Scholar] [CrossRef] [PubMed]
- Pisano, M.B.; Casula, M.; Corda, A.; Fadda, M.E.; Deplano, M.; Cosentino, S. In vitro probiotic characteristics of Lactobacillus strains isolated from Fiore Sardo cheese. Ital. J. Food Sci. 2008, 20, 505–516. [Google Scholar]
- Pisano, M.B.; Fadda, M.E.; Melis, R.; Ciusa, M.L.; Viale, S.; Deplano, M.; Cosentino, S. Molecular identification of bacteriocins produced by Lactococcus lactis dairy strains and their technological and genotypic characterization. Food Control. 2015, 51, 1–8. [Google Scholar] [CrossRef]
- Pisano, M.B.; Patrignani, F.; Cosentino, S.; Guerzoni, M.E.; Franz, C.M.A.P.; Holzapfel, W.H. Diversity and functional properties of Lactobacillus plantarum-group strains isolated from Italian cheese products. Dairy Sci. Technol. 2011, 91, 65–76. [Google Scholar] [CrossRef][Green Version]
- Pisano, M.B.; Viale, S.; Conti, S.; Fadda, M.E.; Deplano, M.; Melis, M.P.; Deiana, M.; Cosentino, S. Preliminary evaluation of probiotic properties of Lactobacillus strains isolated from Sardinian dairy products. BioMed Res. Int. 2014, 2014, 286390. [Google Scholar] [CrossRef]
- Piras, C.; Cesare Marincola, F.; Savorani, F.; Engelsen, S.B.; Cosentino, S.; Viale, S.; Pisano, M.B. A NMR metabolomics study of the ripening process of the Fiore Sardo cheese produced with autochthonous adjunct cultures. Food Chem. 2013, 141, 2137–2147. [Google Scholar] [CrossRef]
- Pisano, M.B.; Casula, M.; Serci, V.; Corda, A.; Deplano, M.; Fadda, M.E.; Cosentino, S. Characterization of goats’ milk cheeses manufactured with the addition of adjunct cultures. The Challenge to Sheep and Goat Milk Sectors. Spec. Issue Int. Dairy Fed. 2008, 801, 263–265. [Google Scholar]
- Pisano, M.B.; Elisabetta Fadda, M.; Deplano, M.; Corda, A.; Casula, M.; Cosentino, S. Characterization of Fiore Sardo cheese manufactured with the addition of autochthonous cultures. J. Dairy Res. 2007, 74, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Bukvicki, D.; Siroli, L.; D’Alessandro, M.; Cosentino, S.; Fliss, I.; Ben Said, L.; Hassan, H.; Lanciotti, R.; Patrignani, F. Unravelling the Potential of Lactococcus lactis Strains to Be Used in Cheesemaking Production as Biocontrol Agents. Foods 2020, 9, 1815. [Google Scholar] [CrossRef]
- Cosentino, S.; Fadda, M.E.; Deplano, M.; Melis, R.; Pomata, R.; Pisano, M.B. Antilisterial activity of nisin-like bacteriocin-producing Lactococcus lactis subsp. lactis isolated from traditional Sardinian dairy products. J. Biomed. Biotechnol. 2012, 2012, 376428. [Google Scholar] [CrossRef] [PubMed]
- Cosentino, S.; Viale, S.; Deplano, M.; Fadda, M.E.; Pisano, M.B. Application of Autochthonous Lactobacillus Strains as Biopreservatives to Control Fungal Spoilage in Caciotta Cheese. BioMed Res. Int. 2018, 2018, 3915615. [Google Scholar] [CrossRef] [PubMed]
- Rincón-León, F. Functional Foods. In Encyclopedia of Food Sciences and Nutrition, 2nd ed.; Caballero, B., Finglas, P., Toldra, F., Eds.; Academic Press: Cambridge, MA, USA, 2003. [Google Scholar]

| Entity | Contents | In Relation with |
|---|---|---|
| Strains | Basic information on strains, mainly related to the MIRRI-IS dataset | All, but Chemical compounds |
| Strain properties | Extended information on strains, mainly related to their chemo-physical properties | Strains, Chemical compounds |
| Strain management | Extended information related to the local management of strains, such as lab book notes and position of vials | Strains |
| Strain culture | Extended information on culture management, such as number of available vials in different storage conditions | Strains |
| Sequences | Information on known sequences of the strains | Strains |
| Species | Scientific species names, derived from authoritative sources (Mycobank, Prokaryotic Nomenclature Up-To-Date) | Strains |
| Literature | Information on bibliographic references for publications relatives to strains | Strains |
| People | Information on people who collected, isolated, identified, and deposited the strains | Strains |
| Projects | Information on projects for which strains were collected and identified | Strains |
| Geographic locations | Information on places where samples were collected | Strains |
| Habitats | Information on strain habitats | Strains |
| Substrates | Information on substrates | Strains |
| Substrate categories | Information on substrate categories (useful for queries) | Strains |
| Growth media | Information on available growth media | Strains |
| Chemical compounds | Information on chemicals and their interactions with strains as by-products, inhibitors, etc. | Strain properties |
| User Type | Query Strains | Insert Strain | Update Strain | Delete Strain | View Lists | Update Lists | Backup Restore | Manage Users | Upload Data |
|---|---|---|---|---|---|---|---|---|---|
| Admin | √ | √ | √ | √ | √ | √ | √ | √ | √ |
| Curator | √ | √ | √ | √ | √ | √ | X | X | X |
| Basic | √ | X | X | X | √ | X | X | X | X |
| Species | Isolates (No.) | Origin | |
|---|---|---|---|
| Enterococcus faecium (Orla-Jensen 1919) Schleifer and Kilpper-Bälz 1984 | 489 | dairy, meat, and animal | Fiore Sardo PDO/home made Pecorino Sardo PDO/Natural starter culture/Pecorino Romano PDO/Casu axedu (fresh goat’s cheese)/Traditional dry sausage/ovine feaces |
| Lactiplantibacillus plantarum (Orla-Jensen 1919) Zheng et al. 2020 | 301 | dairy and animal | Fiore Sardo PDO/Casu axedu/Pecorino Romano PDO/ovine faeces |
| Lactococcus lactis (Lister 1873) Schleifer et al. 1986 | 139 | dairy | Fiore Sardo PDO/homemade Pecorino Sardo PDO/Casu axedu (fresh goat’s cheese) |
| Streptococcus thermophilus Orla-Jensen 1919 (Approved Lists 1980) | 133 | dairy | Natural starter culture (scotta-innesto)/Pecorino Romano PDO/Casu axedu/fermented milk (Gioddu)/Industrial Pecorino Sardo PDO |
| Lacticaseibacillus paracasei (Collins et al. 1989) Zheng et al. 2020 | 111 | dairy and meat | Fiore Sardo PDO/Industrial Pecorino Sardo PDO/Pecorino Romano PDO/Traditional dry sausage |
| Lactobacillus delbrueckii subsp. lactis (Orla-Jensen 1919) Weiss et al. 1984 | 95 | dairy | Natural starter culture (scotta-innesto) |
| Enterococcus faecalis (Andrewes and Horder 1906) Schleifer and Kilpper-Bälz 1984 | 92 | dairy | Fiore Sardo PDO/ Pecorino Romano PDO |
| Lactobacillus delbrueckii (Leichmann 1896) Beijerinck 1901 (Approved Lists 1980) | 12 | dairy and animal | Natural starter culture (scotta-innesto)/Industrial Pecorino sardo PDO/Pecorino Romano PDO/ovine feaces |
| Streptococcus gallolyticus subsp. macedonicus (Tsakalidou et al. 1998) Schlegel et al. 2003 | 11 | dairy | Pecorino Romano PDO |
| Limosilactobacillus reuteri (Kandler et al. 1982) Zheng et al. 2020 | 10 | dairy and meat | Natural starter culture (scotta-innesto)/Pecorino Romano PDO/Traditional dry sausage |
| Enterococcus durans (ex Sherman and Wing 1937) Collins et al. 1984 | 10 | dairy | Fiore Sardo PDO |
| Enterococcus hirae Farrow and Collins 1985 | 9 | dairy | Fiore Sardo PDO |
| Lacticaseibacillus rhamnosus (Hansen 1968) Zheng et al. 2020 | 8 | dairy | Fiore Sardo PDO/Industrial Pecorino Sardo PDO |
| Levilactobacillus brevis (Orla-Jensen 1919) Zheng et al. 2020 | 8 | dairy and meat | Fiore Sardo PDO/Traditional dry sausage |
| Lactobacillus helveticus (Orla-Jensen 1919) Bergey et al. 1925 (Approved Lists 1980) | 5 | dairy and animal | Pecorino Romano PDO/ovine faeces |
| Lactobacillus delbrueckii subsp. bulgaricus (Orla-Jensen 1919) Weiss et al. 1984 | 4 | dairy | Fermented milk (Gioddu)/natural starter culture (scotta-innesto) |
| Pediococcus pentosaceus Mees 1934 (Approved Lists 1980) | 4 | dairy | Fiore Sardo PDO |
| Lacticaseibacillus casei (Orla-Jensen 1916) Zheng et al. 2020 | 2 | dairy | Industrial Pecorino Sardo PDO |
| Loigolactobacillus coryniformis (Abo-Elnaga and Kandler 1965) Zheng et al. 2020 | 2 | dairy | Fiore Sardo PDO |
| Lactiplantibacillus paraplantarum (Curk et al. 1996) Zheng et al. 2020 | 1 | dairy | Fiore Sardo PDO |
| Lentilactobacillus hilgardii (Douglas and Cruess 1936) Zheng et al. 2020 | 1 | dairy | Fiore Sardo PDO |
| Lentilactobacillus parabuchneri (Farrow et al. 1989) Zheng et al. 2020 | 1 | dairy | Fiore Sardo PDO |
| Limosilactobacillus fermentum (Beijerinck 1901) Zheng et al. 2020 | 1 | dairy | Pecorino Romano PDO |
| Enterococcus gallinarum (Bridge and Sneath 1982) Collins et al. 1984 | 1 | dairy | Fiore Sardo PDO |
| Latilactobacillus curvatus (Troili-Petersson 1903) Zheng et al. 2020 | 54 | meat and dairy | Traditional dry sausage/Fiore Sardo PDO |
| Staphylococcus pasteuri Chesneau et al. 1993 | 10 | meat and dairy | Traditional dry sausage/Pecorino Romano PDO |
| Staphylococcus warneri Kloos and Schleifer 1975 (Approved Lists 1980) | 5 | meat and dairy | Traditional dry sausage/Pecorino Romano PDO |
| Latilactobacillus sakei (Katagiri et al. 1934) Zheng et al. 2020 | 108 | meat | Traditional dry sausage |
| Staphylococcus equorum Schleifer et al. 1985 | 40 | meat | Traditional dry sausage |
| Staphylococcus xylosus Schleifer and Kloos 1975 (Approved Lists 1980) | 17 | meat | Traditional dry sausage |
| Staphylococcus succinus Lambert et al. 1998 | 10 | meat | Traditional dry sausage |
| Staphylococcus epidermidis (Winslow and Winslow 1908) Evans 1916 (Approved Lists 1980) | 7 | meat | Traditional dry sausage |
| Leuconostoc mesenteroides (Tsenkovskii 1878) van Tieghem 1878 (Approved Lists 1980) | 3 | meat | Traditional dry sausage |
| Staphylococcus saprophyticus (Fairbrother 1940) Shaw et al. 1951 (Approved Lists 1980) | 3 | meat | Traditional dry sausage |
| Staphylococcus vitulinus corrig. Webster et al. 1994 | 2 | meat | Traditional dry sausage |
| Leuconostoc carnosum Shaw and Harding 1989 | 1 | meat | Traditional dry sausage |
| Leuconostoc citreum Farrow et al. 1989 | 1 | meat | Traditional dry sausage |
| Sphingomonas paucimobilis (Holmes et al. 1977) Yabuuchi et al. 1990 | 6 | fish | Fish gut (Sparus aurata) |
| Psychrobacter maritimus Romanenko et al. 2004 | 3 | fish | Fish gut (Sparus aurata) |
| Pseudomonas brenneri Baïda et al. 2002 | 2 | fish | Fish gut (Sparus aurata) |
| Pseudomonas fluorescens Migula 1895 (Approved Lists 1980) | 2 | fish | Fish gut (Sparus aurata) |
| Aeromonas salmonicida (Lehmann and Neumann 1896) Griffin et al. 1953 (Approved Lists 1980) | 1 | fish | Fish gut (Sparus aurata) |
| Yersinia bercovieri Wauters et al. 1988 | 1 | fish | Fish gut (Sparus aurata) |
| Erwinia persicina corrig. Hao et al. 1990 | 1 | fish | Fish gut (Sparus aurata) |
| Lactiplantibacillus pentosus (Zanoni et al. 1987) Zheng et al. 2020 | 29 | plant | Naturally fermented table olives (Tonda di Cagliari) |
| Bacteria | |||
|---|---|---|---|
| Genera | Species | Isolates (No.) | |
| Arthrobacter | spp. | Conn and Dimmick 1947 | 2 |
| Bacillus | amiloliquefaciens | (ex Fukumoto 1943) Priest et al. 1987 | 10 |
| coagulans | Hammer 1915 | 1 | |
| megaterium | de Bary 1884 | 1 | |
| pumilus | Meyer and Gottheil 1901 | 4 | |
| subtilis | (Ehrenberg 1835) Cohn 1872 | 5 | |
| velezensis | Ruiz-García et al. 2005 | 1 | |
| Bifidobacterium | pseudolongum | Mitsuoka 1969 | 1 |
| Brevibacillus | spp. | Shida et al. 1996 | 4 |
| agrii | (Nakamura 1993) Shida et al. 1996 | 9 | |
| invocatus | Logan et al. 2002 | 6 | |
| parabrevis | (Takagi et al. 1993) Shida et al. 1996 | 3 | |
| Cutibacterium | acnes | (Gilchrist 1900) Scholz and Kilian 2016 | 1 |
| Enterococcus | faecium | (Orla-Jensen 1919) Schleifer and Kilpper-Bälz 1984 | 2 |
| hirae | Farrow and Collins 1985 | 2 | |
| Frigobacterium | spp. | Frigoribacterium Kämpfer et al. 2000 | 1 |
| Kocuria | rhizophila | Kovács et al. 1999 | 4 |
| Lacticaseibacillus | rhamnosus | (Hansen 1968) Zheng et al. 2020 | 1 |
| Limosilactobacillus | mucosae | (Roos et al. 2000) Zheng et al. 2020 | 1 |
| Leuconostoc | citreum | Farrow et al. 1989 | 1 |
| Lysinibacillus | spp. | Ahmed et al. 2007 | 1 |
| Microbacterium | aerolatum | Zlamala et al. 2002 | 4 |
| Micrococcus | spp. | Cohn 1872 | 3 |
| Pantoea | spp. | Gavini et al. 1989 | 3 |
| Propionibacterium | acnes | (Gilchrist 1900) Douglas and Gunter 1946 | 1 |
| Staphylococcus | arlettae | Schleifer et al. 1985 | 3 |
| capitis | Kloos and Schleifer 1975 | 1 | |
| epidermidis | (Winslow and Winslow 1908) Evans 1916 | 2 | |
| hominis | Kloos and Schleifer 1975 | 3 | |
| pasteuri | Chesneau et al. 1993 | 3 | |
| warneri | Kloos and Schleifer 1975 | 3 | |
| Streptococcus | equinus | Andrewes and Horder 1906 | 1 |
| spp. | Rosenbach 1884 | 1 | |
| Weissella | cibaria | Björkroth et al. 2002 | 1 |
| Yeasts | |||
| Genera | Species | Isolates (No.) | |
| Aureobasidium | pullulans | (De Bary) G. Arnaud ex Cif., Ribaldi & Corte, 1957 | 3 |
| Candida | adriatrica | Čadež, Cardinali, Ciafardini & G. Péter ,2012 | 6 |
| dendronema | Van der Walt, Klift & D.B. Scott | 1 | |
| diddensiae | (Phaff, Mrak, & O.B. Williams) Fell & S.A. Mey, 1967 | 1 | |
| guilliermondii | (Castell.) Langeron & Guerra, 1938 | 1 | |
| molendinolei | Čadež, Turchetti & G. Péter , 2012 | 8 | |
| temnochilae | S.O. Suh, N.H. Nguyen & M. Blackw., 2005 | 2 | |
| wickerhamii | S.A. Mey. & Yarrow, 1978 | 1 | |
| Clavispora | lusitaniae | Rodr. Mir. and Antonie van Leeuwenhoek, 1979 | 1 |
| Cryptococcus | carnescens | (Verona & Luchetti) M. Takash, 2003 | 1 |
| magnus | (Lodder & Kreger) Baptist & Kurtzman, 1977 | 1 | |
| victoriae | M.J. Montes, Belloch, Galiana, M.D. García, C. Andrés, S. Ferrer, Torr.-Rodr. & J. Guinea, 1999 | 1 | |
| Hanseniaspora | uvarum | (Niehaus) Shehata, Mrak & Phaff ex M.T. Sm., 1984 | 4 |
| Hansenula | anomala | (E.C. Hansen) Syd. & P. Syd, 1919 | 5 |
| californica | (Lodder) Wick, 1951 | 1 | |
| fabiani | Wick, 1965 | 1 | |
| Kluyveromyces | africanus | Van der Walt, Antonie van Leeuwenhoek, 1956 | 1 |
| bulgaricus | (Santa María) Van der Walt, 1971 | 2 | |
| fragilis | (A. Jörg.) Van der Walt, 1971 | 1 | |
| lactis | (Stell.-Dekk.) Van der Wal, 1965 | 1 | |
| veronae | (Lodder & Kreger) Van der Walt, 1971 | 7 | |
| Lodderomyces | elongisporus | (Recca & Mrak) Van der Walt, 1971 | 1 |
| Metschnikowia | pulcherrima | Pitt & Mill, 1968 | 13 |
| Nakazawaea | anatomiae | Kurtzman & Robnett, 2014 | 2 |
| Pichia | farinosa | (Lindner) Guillierm, 1904 | 2 |
| guilliermondii | Wick, 1966 | 1 | |
| kudriavzevii | Boidin, Pignal & Besson, 1965 | 1 | |
| manshurica | Saito, 1914 | 1 | |
| membranifaciens | Hansen, 1904 | 2 | |
| mexicana | M. Miranda, Holzschu, Phaff & Starmer, 1982 | 1 | |
| nakazawae | Kodama, 1975 | 1 | |
| Pirula | salina | (P.A. Dangeard) Printz, 1927 | 1 |
| Rhodosporidium | babjevae | Golubev, 1993 199 | 1 |
| Rhodotorula | glutinis | (Fresen.) F.C. Harrison, 1928 | 2 |
| Saccharomyces | aceti | Santa María, 1958 | 3 |
| bailii | Lindner, 1895 | 3 | |
| bayanus | Sacc, 1895 | 14 | |
| capensis | Van der Walt & Tscheuschner, 1956 | 1 | |
| cerevisiae | Meyen ex E.C. Hansen, Medd. Carlsberg, 1883 | 16 | |
| chevalieri | Reess, 1870 | 10 | |
| ellipsoideus | Guillierm, 1914 | 36 | |
| exiguus | Reess ex E.C. Hansen, 1888 | 1 | |
| fructuum | Lodder & Kreger-van Rij, 1952 | 1 | |
| italicus | Castelli, 1939 | 13 | |
| kluyveri | Phaff, M.W. Mill. & Shifrine, 1956 | 4 | |
| prostoserdovii | Kudryavtsev, 1960 | 1 | |
| Fungi | |||
| Genera | Species | Isolates (No.) | |
| Fusarium | avenaceum | Sacc, 1886 | 3 |
| cortaderiae | O’Donnell, T. Aoki, Kistler & Geiser, 2004 | 1 | |
| crookwellense | L.W. Burgess, P.E. Nelson & Toussoun, 1982 | 2 | |
| culmorum | Sacc, 1895 | 37 | |
| fujikuroi | Nirenberg, 1976 | 1 | |
| globosum | Rheeder, Marasas & P.E. Nelson, 1996 | 2 | |
| graminearum | Schwabe, 1839 | 3 | |
| meridionale | Aoki, 2004 | 1 | |
| oxysporum | Schltdl, 1824 | 3 | |
| phyllophilum | Nirenberg & O’Donnell, 1998 | 1 | |
| pseudograminearum | O’Donnell & T. Aoki, 1999 | 1 | |
| solani | Sacc, 1881 | 2 | |
| F. incarnatum | equiseti complex | Sacc, 1886 | 2 |
| Bacteria | ||
|---|---|---|
| Genera | Species | Isolates (No.) |
| Enterococcus | spp. | 120 |
| durans (ex Sherman and Wing 1937) Collins et al. 1984 | 11 | |
| faecalis (Andrewes and Horder 1906) Schleifer and Kilpper-Balz 1984 | 5 | |
| faecium (Orla-Jensen 1919) Schleifer and Kilpper-Balz 1984 | 41 | |
| Lactobacillus | spp. | 110 |
| helveticus (Orla-Jensen 1919) Bergey et al. 1925 | 1 | |
| Lentilactobacillus | buchneri (Henneberg 1903) Zheng et al. 2020 | 2 |
| Lacticaseibacillus | casei (Orla-Jensen 1916) Zheng et al. 2020 | 4 |
| rhamnosus (Hansen 1968) Zheng et al. 2020 | 1 | |
| paracasei (Collins et al. 1989) Zheng et al. 2020 | 26 | |
| Limosilactobacillus | fermentum (Beijerinck 1901) Zheng et al. 2020 | 3 |
| Levilactobacillus | brevis (Beijerinck 1901) Zheng et al. 2020 | 11 |
| Lactiplantibacillus | paraplantarum (Curk et al. 1996) Zheng et al. 2020 | 1 |
| plantarum (Orla-Jensen 1919) Zheng et al. 2020 | 36 | |
| Latilactobacillus | sakei (Katagiri et al. 1934) Zheng et al. 2020 | 9 |
| Lactococcus | spp. | 25 |
| lactis (Lister 1873) Schleifer et al. 1986 | 53 | |
| cremoris (Orla-Jensen 1919) Li et al. 2021 | 2 | |
| raffinolactis (Orla-Jensen and Hansen 1932) Schleifer et al. 1988 | 1 | |
| lactis subsp. lactis bv. diacetylactis (NCBI:txid44688) | 1 | |
| Leuconostoc | spp. | 15 |
| mesenteroides (Tsenkovskii 1878) van Tieghem 1878 | 5 | |
| Pediococcus | spp. | 4 |
| acidilactici Lindner 1887 (App. Lists 1980) emend. Jud. Commission 1996 | 4 | |
| pentosaceus Mees 1934 | 2 | |
| Streptococcus | thermophilus (Orla-Jensen 1919) Schleifer et al. 1995 | 7 |
| macedonicus Tsakalidou et al. 1998 | 1 | |
| Yeasts | ||
| Genera | Species | Isolates (No.) |
| Candida | albicans (C.P. Robin) Berkhout, De schimmelgeslachten Monilia, Oidium, Oospora en Torula: 44 (1923) | 2 |
| atlantica (Siepmann) S.A. Mey. & Simione, Mycotaxon 66: 100 (1998) | 1 | |
| catenulata Diddens & Lodder, Die anaskosporogenen Hefen, II Hälfte: 486 (1942) | 2 | |
| inconspicua (Lodder & Kreger) S.A. Mey. & Yarrow, International Journal of Systematic Bacteriology 28 (4): 612 (1978) | 2 | |
| krusei (Castell.) Berkhout, De schimmelgeslachten Monilia, Oidium, Oospora en Torula: 44 (1923) | 2 | |
| parapsilosis (Ashford) Langeron & Talice, Annales de Parasitologie Humaine Comparée 10: 54 (1932) | 14 | |
| sake (Saito & Oda) van Uden & H.R. Buckley, Mycotaxon 17: 298 (1983) | 1 | |
| utilis (Henneberg) Lodder & Kreger, The Yeasts: a taxonomic study: 546 (1952) | 2 | |
| zeylanoides (Castell.) Langeron & Guerra, Annales de Parasitologie Humaine Comparée 16 (5): 501 (1938) | 10 | |
| Cryptococcus | curvatus (Diddens & Lodder) Golubev, Mikologiya i Fitopatologiya 15 (6): 467 (1981) | 2 |
| uniguttulatus (Zach) Phaff & Fell, The Yeasts: a taxonomic study: 1140 (1970) | 3 | |
| Cutaneotrichosporon | guehoae (Middelhoven, Scorzetti & Fell) Yurkov, Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Studies in Mycology 81: 140 (2015) | 3 |
| Cystobasidium | slooffiae (E.K. Novák & Vörös-Felkai) Yurkov, Kachalkin, H.M. Daniel, M. Groenew., Libkind, V. de Garcia, Zalar, Gouliam., Boekhout & Begerow, Antonie van Leeuwenhoek 107 (1): 180 (2014) | 3 |
| Debaryomyces | hansenii (Zopf) Lodder & Kreger, The Yeasts: a taxonomic study: 280 (1952) | 36 |
| Filobasidium | uniguttulatum Kwon-Chung, International Journal of Systematic Bacteriology 27: 293 (1977) | 1 |
| Geotrichum | candidum Link, Magazin der Gesellschaft Naturforschenden Freunde Berlin 3 (1): 17, t. 1:26 (1809) | 3 |
| Hannaella | oryzae (Nakase & M. Suzuki) F.Y. Bai & Q.M. Wang, FEMS Yeast Research 8 (5): 805 (2008) | 1 |
| Kluyveromyces | lactis (Stell.-Dekk.) Van der Walt, Antonie van Leeuwenhoek 31: 347 (1965) | 4 |
| marxianus (E.C. Hansen) Van der Walt, Antonie van Leeuwenhoek 31: 347 (1965) | 3 | |
| Pichia | kudriavzevii Boidin, Pignal & Besson, Bulletin de la Société Mycologique de France 81: 589 (1965) | 1 |
| Rhodotorula | mucilaginosa (A. Jörg.) F.C. Harrison, Transactions of the Royal Society of Canada. Section 5, Biological Sciences 22: 191 (1928) | 6 |
| Saccharomyces | cerevisiae (Desm.) Meyen, Archiv für Naturgeschichte 4 (2): 100 (1838) | 1 |
| Trichosporon | aquatile L.R. Hedrick & P.D. Dupont, Antonie van Leeuwenhoek 34: 474-482 (1968) | 3 |
| cutaneum (Beurm., Gougerot & Vaucher bis) M. Ota, Annales de Parasitologie Humaine Comparée 4: 12 (1926) | 3 | |
| gracile (Weigmann & A. Wolff) E. Guého & M.T. Sm., Antonie van Leeuwenhoek 61 (4): 307 (1992) | 1 | |
| jirovecii Frágner, Ceská Mykologie 23 (3): 160 (1969) | 1 | |
| lactis Lopandic, Sugita, Middelhoven, Herzberg & Prillinger, Trichosporon caseorum sp. nov. and Trichosporon lactis sp. nov., two basidiomycetous yeasts isolated from cheeses: 99-116 (2004) | 6 | |
| mucoides E. Guého & M.T. Sm., Antonie van Leeuwenhoek 61 (4): 312 (1992) | 2 | |
| Wickerhamiella | pararugosa (Nakase, Komag. & Fukaz.) C. Vega & Lachance, FEMS Yeast Research 17 (5): fox054, 8 (2017) | 1 |
| Yarrowia | deformans (Zach) M. Groenew. & M.T. Sm., Antonie van Leeuwenhoek 103 (5): 1025 (2013) | 7 |
| lipolytica (Wick., Kurtzman & Herman) Van der Walt & Arx, Antonie van Leeuwenhoek 46 (6): 519 (1980) | 27 |
| Characterization | Bacteria | Yeasts |
|---|---|---|
| Taxonomic- | Morphology | Morphology |
| Biochemical | Gram positive | Fermentation of GLU-GAL-LAC |
| Catalase negative | Assimilation of LAC-LAT-CIT | |
| Gas production from glucose | Urease activity | |
| NH3 production from arginine | Growth at 10°, 15°, 45 °C | |
| Esculin hydrolysis | ||
| Growth at 10°, 15°, 40°, 45 °C | ||
| Tolerance to 6,5/10% NaCl | ||
| Growth at pH 3,9/9,6 | ||
| Carbohydrates fermentation profile | ||
| Molecular | Species-specific PCR | Species-specific PCR |
| PCR-RAPD | PCR-RFLP | |
| Sequencing of 16S rDNA | Sequencing of D1-D2 26S rDNA | |
| Technological | Acidifying activity | Proteolytic activity |
| Proteolytic activity | Lipolytic activity | |
| Lipolytic activity | Growth atl 6, 10, 15, 20% NaCl | |
| Fermentation of citrate | ||
| Milk coagulation | ||
| API ZYM | ||
| Functional | Survival at pH 2.0/2.5 | Survival at pH 2.0/2.5 |
| Simulated stomach-duodenum passage | Simulated stomach-duodenum passage | |
| Autoaggregation ability | Autoaggregation abillity | |
| Hydrophobicity | Hydrophobicity | |
| Adhesion properties | Adhesion properties | |
| Antimicrobial activity in vitro e in situ | Killer activity | |
| Bile salts deconjugation | Bile salts deconjugation | |
| Antibiotics resistance | ||
| Cholesterol assimilation |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daga, E.; Budroni, M.; Multineddu, C.; Cosentino, S.; Deplano, M.; Romano, P.; Comunian, R. The MicroBioDiverSar Project: Exploring the Microbial Biodiversity in Ex Situ Collections of Sardinia. Sustainability 2021, 13, 8494. https://doi.org/10.3390/su13158494
Daga E, Budroni M, Multineddu C, Cosentino S, Deplano M, Romano P, Comunian R. The MicroBioDiverSar Project: Exploring the Microbial Biodiversity in Ex Situ Collections of Sardinia. Sustainability. 2021; 13(15):8494. https://doi.org/10.3390/su13158494
Chicago/Turabian StyleDaga, Elisabetta, Marilena Budroni, Chiara Multineddu, Sofia Cosentino, Maura Deplano, Paolo Romano, and Roberta Comunian. 2021. "The MicroBioDiverSar Project: Exploring the Microbial Biodiversity in Ex Situ Collections of Sardinia" Sustainability 13, no. 15: 8494. https://doi.org/10.3390/su13158494
APA StyleDaga, E., Budroni, M., Multineddu, C., Cosentino, S., Deplano, M., Romano, P., & Comunian, R. (2021). The MicroBioDiverSar Project: Exploring the Microbial Biodiversity in Ex Situ Collections of Sardinia. Sustainability, 13(15), 8494. https://doi.org/10.3390/su13158494








