Assessment of Ionomic, Phenolic and Flavonoid Compounds for a Sustainable Management of Xylella fastidiosa in Morocco
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples Collection
2.2. Polymerase Chain Reaction
2.3. Extract Preparation for Assessment of Total Phenolic and Flavonoid Content
2.4. Total Phenolic Content
2.5. Total Flavonoid Content
2.6. Determination of Leaf Ionome and Soil Parameters
2.7. Statistical Analyses
3. Results
3.1. Polymerase Chain Reaction
3.2. Determination of Leaf Ionome
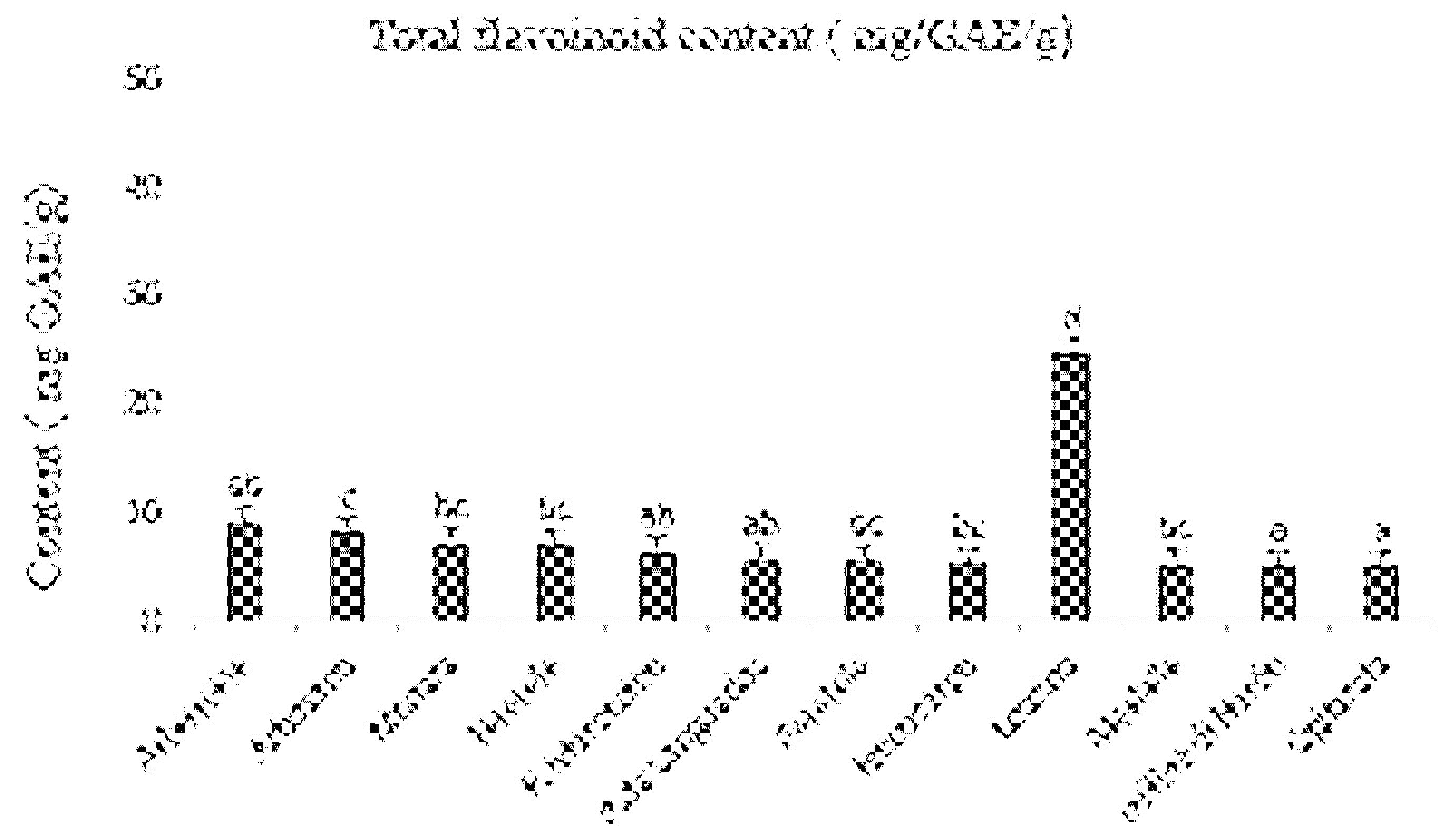
3.3. Determination of Total Phenolic and Flavonoid Content
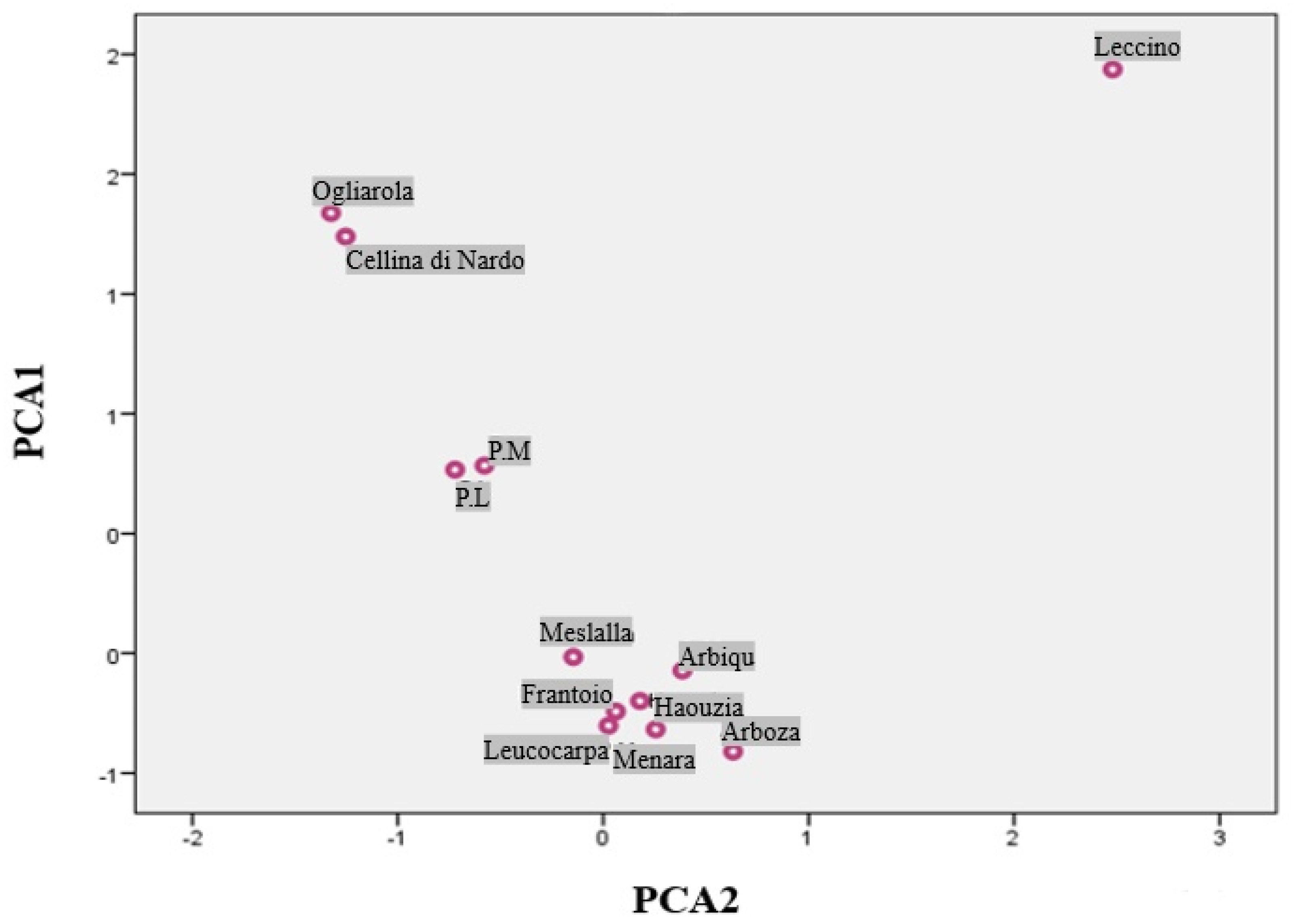
3.4. Principal Component Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Bajoub, A.; Medina-Rodríguez, S.; Olmo-García, L.; Ajal, E.A.; Monasterio, R.P.; Hanine, H.; Fernández-Gutiérrez, A.; Carrasco-Pancorbo, A. In-Depth Two-Year Study of Phenolic Profile Variability among Olive Oils from Autochthonous and Mediterranean Varieties in Morocco, as Revealed by a LC-MS Chemometric Profiling Approach. Int. J. Mol. Sci. 2017, 18, 52. [Google Scholar] [CrossRef]
- FAOSTAT. Food and Agriculture Organization Statistical Data-Base. 2019. Available online: http://faostat.fao.org/default.aspx (accessed on 5 May 2021).
- Bouymajane, A.; El Majdoub, Y.O.; Cacciola, F.; Russo, M.; Salafia, F.; Trozzi, A.; Filali, F.R.; Dugo, P.; Mondello, L. Characterization of Phenolic Compounds, Vitamin E and Fatty Acids from Monovarietal Virgin Olive Oils of “Picholine marocaine” Cultivar. Molecules 2020, 25, 5428. [Google Scholar] [CrossRef] [PubMed]
- Barbara, H.; Terral, J.F.; Ater, M. Première Caractérisa-tion Pomologique des Variétés Locales De L’olivier (Olea Europaea, L.) Des Oliveraies Tradition-nelles des Agroécosystèmes des Montagnes Du Nord-Ouest Du Maroc. Eur. Sci. J. ESJ 2020, 16, 556–575. [Google Scholar]
- Chliyeh, M.; Touhami, A.O.; Filali-Maltouf, A.; El Modafar, C.; Moukhli, A.; Oukabli, A.; Benkirane, R.; Douira, A. Phytophthora Palmivora: A New Pathogen of Olive Trees in Morocco. Atlas J. Biol. 2013, 2, 130–135. [Google Scholar] [CrossRef]
- Sicard, A.; Zeilinger, A.R.; Vanhove, M.; Schartel, T.E.; Beal, D.J.; Daugherty, M.P.; Almeida, R.P. Xylella fastidiosa: Insights into an Emerging Plant Pathogen. Annu. Rev. Phytopathol. 2018, 56, 181–202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, S.M.; Navas-Cortés, J.A.; Bullock, J.M.; Boscia, D.; Chapman, D.S. Estimating the Epidemiology of Emerging Xylella fastidiosa Outbreaks in Ol-ives. Plant Pathol. 2020, 69, 1403–1413. [Google Scholar] [CrossRef]
- Afechtal, M.; Vicent, A.; Saponari, M.; D’Onghia, A.M. Pest risk analysis on Xylella fastidiosa in Morocco. J. Plant Prot. Res. 2018, 58. [Google Scholar]
- Frem, M.; Chapman, D.; Fucilli, V.; Choueiri, E.; El Moujabber, M.; La Notte, P.; Nigro, F. Xylella fastidiosa invasion of new countries in Europe, the Middle East and North Africa: Ranking the potential exposure scenarios. NeoBiota 2020, 59, 77–97. [Google Scholar] [CrossRef]
- Afechtal, M.; Aitfriha, A.; Bibi, I. A Preliminary Survey on the Presence of Xylella fastidiosa in Olive, Citrus and Grapevine Groves in Morocco. Rev. Maroc. Sci. Agron. Vétérinaires 2017, 6, 6–9. [Google Scholar]
- Martelli, G.P. The current status of the quick decline syndrome of olive in southern Italy. Phytoparasitica 2016, 44, 1–10. [Google Scholar] [CrossRef]
- Oepp, B.; Eppo, B. PM 7/24 (4) Xylella fastidiosa. EPPO Bull. 2019, 49, 175–227. [Google Scholar]
- D’Attoma, G.; Morelli, M.; Saldarelli, P.; Saponari, M.; Giampetruzzi, A.; Boscia, D.; Savino, V.N.; De La Fuente, L.; Cobine, P.A. Ionomic Differences between Susceptible and Resistant Olive Cultivars Infected by Xylella fastidiosa in the Outbreak Area of Salento, Italy. Pathogens 2019, 8, 272. [Google Scholar] [CrossRef] [Green Version]
- Del Coco, L.; Migoni, D.; Girelli, C.R.; Angilè, F.; Scortichini, M.; Fanizzi, F.P. Soil and Leaf Ionome Heterogeneity in Xylella fastidiosa Subsp. Pauca-Infected, Non-Infected and Treated Olive Groves in Apulia, Italy. Plants 2020, 9, 760. [Google Scholar] [CrossRef]
- Cruz, L.F.; Cobine, P.A.; De La Fuente, L. Calcium Increases Xy-lella fastidiosa Surface Attachment, Biofilm Formation, and Twitching Motility. Appl. Environ. Microbiol. 2012, 78, 1321–1331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raffini, F.; Bertorelle, G.; Biello, R.; D’Urso, G.; Russo, D.; Bosso, L. From Nucleotides to Satellite Imagery: Approaches to Identify and Manage the Invasive Pathogen Xylella fastidiosa and Its Insect Vectors in Europe. Sustainability 2020, 12, 4508. [Google Scholar] [CrossRef]
- Hood, M.I.; Skaar, E.P. Nutritional immunity: Transition metals at the pathogen–host interface. Nat. Rev. Genet. 2012, 10, 525–537. [Google Scholar] [CrossRef] [PubMed]
- Baxter, I.; Dilkes, B.P. Elemental Profiles Reflect Plant Adaptations to the Environment. Science 2012, 336, 1661–1663. [Google Scholar] [CrossRef] [Green Version]
- Oliver, J.; Sefick, S.A.; Parker, J.K.; Arnold, T.; Cobine, P.A.; De La Fuente, L.; Oliver, J.; Sefick, S.A.; Parker, J.K.; Arnold, T.; et al. Ionome Changes in Xylella fastidiosa–Infected Nicotiana tabacum Correlate with Virulence and Discriminate Between Subspecies of Bacterial Isolates. Mol. Plant-Microbe Interact. 2014, 27, 1048–1058. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Navarrete, F.; De La Fuente, L. Zinc Detoxification Is Required for Full Virulence and Modification of the Host Leaf Ionome by Xylella fastidiosa. Mol. Plant-Microbe Interact. 2015, 28, 497–507. [Google Scholar] [CrossRef] [Green Version]
- Cobine, P.A.; Cruz, L.F.; Navarrete, F.; Duncan, D.; Tygart, M.; De La Fuente, L. Xylella fastidiosa Differentially Accumulates Mineral Elements in Biofilm and Planktonic Cells. PLoS ONE 2013, 8, e5493610. [Google Scholar] [CrossRef] [PubMed]
- Andersen, P.C.; French, W.J. Biophysical Characteristics of Peach-Trees Infected with Phony Peach Disease. Physiol. Mol. Plant Pathol. 1987, 31, 25–40. [Google Scholar] [CrossRef]
- De La Fuente, L.; Parker, J.K.; Oliver, J.E.; Granger, S.; Brannen, P.M.; van Santen, E.; Cobine, P.A. The bacterial pathogen Xylella fastidiosa affects the leaf ionome of plant hosts during infection. PLoS ONE 2013, 8, e62945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodrigues, C.M.; De Souza, A.A.; Takita, M.A.; Kishi, L.T.; Machado, M.A. RNA-Seq analysis of Citrus reticulata in the early stages of Xylella fastidiosa infection reveals auxin-related genes as a defense response. BMC Genom. 2013, 14, 676. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vergine, M.; Nicolì, F.; Sabella, E.; Aprile, A.; De Bellis, L.; Luvisi, A. Secondary Metabolites in Xylella fastidiosa—Plant Interaction. Pathogens 2020, 9, 675. [Google Scholar] [CrossRef] [PubMed]
- Djelouah, K.; Frasheri, D.; Valentini, F.; D’Onghia, A.M.; Digiaro, M. Direct tissue blot immunoassay for detection of Xylella fastidiosa in olive trees. Phytopathol. Mediterr. 2020, 53, 559–564. [Google Scholar]
- Minsavage, G.V.; Thompson, C.M.; Hopkins, D.L.; Leite, R.M.V.B.C.; Stall, R.E. Development of a polymerase chain reaction protocol for detection of Xylella fastidiosa in plant tissue. Phytopathology 1994, 84, 446–461. [Google Scholar] [CrossRef]
- Sanders, T.H.; McMichael, R.W.; Hendrix, K.W. Occurrence of resveratrol in edible peanuts. J. Agric. Food Chem. 2020, 48, 1243–1246. [Google Scholar] [CrossRef]
- Xie, L.; Bolling, B.W. Characterisation of stilbenes in California almonds (Prunus dulcis) by UHPLC–MS. Food Chem. 2014, 148, 300–306. [Google Scholar] [CrossRef]
- Waterhouse, A.L. Determination of total phenolics. Curr. Protoc. Food Anal. Chem. 2002, 6, 111–118. [Google Scholar]
- Barreira, J.C.; Ferreira, I.C.; Oliveira, M.; Pereira, J.A. Antioxidant activities of the extracts from chestnut fower leaf skins and fruit. Food Chem. 2018, 107, 1106–1113. [Google Scholar] [CrossRef]
- European and Mediterranean Plant Protection Organization (EPPO). Diagnostic protocols for regulated pests. In Xylella fastidiosa; Bulletin OEPP/EPPO Bulletin: Paris, France, 2004; pp. 187–192. Available online: https://onlinelibrary.wiley.com/doi/epdf/10.1111/j.1365-2338.2004.00718.x (accessed on 5 May 2021).
- Firrao, G.; Bazzi, C. Specific identification of Xylella fastidiosa using the polymerase chain reaction. Phytopathol. Mediterr. 1994, 33, 90–92. [Google Scholar]
- Harper, S.J.; Ward, L.I.; Clover, G.R.G. Development of LAMP and Real-Time PCR Methods for the Rapid Detection of Xylella fastidiosa for Quarantine and Field Applications. Phytopathology 2010, 100, 1282–1288. [Google Scholar] [CrossRef]
- Kailis, S.; Harris, D. Fertiliser Requirements and Monitoring Nutritional Status; Landlinks Press: Collingwood, Australia, 2007. [Google Scholar]
- Pagan, I.; Garcia-Arenal, F. Tolerance to Plant Pathogens: Theory and Experimental Evidence. Int. J. Mol. Sci. 2018, 19, 810. [Google Scholar] [CrossRef] [Green Version]
- El Chami, D.; Galli, F. An assessment of seaweed extracts: Innovation for sustainable agriculture. Agronomy 2020, 10, 1433. [Google Scholar] [CrossRef]
- Saponari, M.; Giampetruzzi, A.; Loconsole, G.; Boscia, D.; Saldarelli, P. Xylella fastidiosa in Olive in Apulia:Where We Stand. Phytopathology 2019, 109, 175–186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Torres, M.A.; Jones, J.D.G.; Dangl, J.L. Reactive oxygen species signaling in response to pathogens. Plant Physiol. 2006, 141, 373–378. [Google Scholar] [CrossRef] [Green Version]
- Caspi, V.; Droppa, M.; Horvath, G.; Malkin, S.; Marder, J.B.; Raskin, V.I. The effect of copper on chlorophyll organization during greening of barley leaves. Photosynth. Res. 1999, 62, 165–174. [Google Scholar] [CrossRef]
- Eide, D.J. The oxidative stress of zinc deficiency. Metallomics 2011, 3, 1124–1129. [Google Scholar] [CrossRef]
- Cabot, C.; Martos, S.; Llugany, M.; Gallego, B.; Toirà, R.; Poschenrieder, C. A role for zinc in plant defense against pathogens and herbivores. Front. Plant Sci. 2019, 10, 1171. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Hong, J.K.; Lee, S.C.; Sohn, K.H.; Jung, H.W.; Hwang, B.K. CAZFP1, CYS2/HiS(2)-type zinc-finger transcription factor gene functions as a pathogen-induced early-defense gene in Capsicum annuum. Plant Mol. Biol. 2004, 55, 883–904. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Rai, A.K.; Kanwar, S.S.; Sharma, T.R. Comparative analysis of zinc finger proteins involved in plant disease resistance. PLoS ONE 2012, 7, 8. [Google Scholar] [CrossRef] [Green Version]
- Poschenrieder, C.; Tolrà, R.; Barceló, J. Can metals defend plants against biotic stress? Trends Plant Sci. 2006, 11, 288–295. [Google Scholar] [CrossRef]
- Fones, H.N.; Preston, G.M. Reactive oxygen and oxidative stress tolerance in plant pathogenic Pseudomonas. Fems Microbiol. Letters 2012, 327, 1–8. [Google Scholar]
- Scortichini, M. Preliminary results on field trial to control Xylella fastidiosa on olive trees in Puglia. Options Méditerranéennes 2017, 121, 77–78. [Google Scholar]
- Sabella, E.; Luvisi, A.; Aprile, A.; Negro, C.; Vergine, M.; Nicolì, F.; Miceli, A.; De Bellis, L. Xylella fastidiosa induces differential expression of lignification related-genes and lignin accumulation in tolerant olive trees cv. Leccino. J. Plant Physiol. 2018, 220, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Leite, B.; Ishida, M.; Alves, E.; Carrer, H.; Pascholati, S.; Kitajima, E. Genomics and X-ray microanalysis indicate that Ca2+ and thiols mediate the aggregation and adhesion of Xylella fastidiosa. Braz. J. Med. Biol. Res. 2002, 35, 645–650. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gutha, L.R.; Casassa, L.F.; Harbertson, J.F.; Naidu, R.A. Modulation of flavonoid biosynthetic pathway genes and anthocyanins due to virus infection in grapevine (Vitis vinifera L.) leaves. BMC Plant Biol. 2010, 10, 187. [Google Scholar] [CrossRef] [Green Version]
- El Aabidine, A.Z.; Baissac, Y.; Moukhli, A.; Jay-Allemand, C.; Khadari, B.; El Modafar, C. Resistance of olivetree to Spilocaea oleagina is mediated by the synthesis of phenolic compounds. Int. J. Agric. Biol. 2010, 12, 61–67. [Google Scholar]
- Rusjan, D.; Veberič, R.; Mikulič-Petkovšek, M. The response of phenolic compounds in grapes of the variety ‘Chardonnay’ (Vitis vinifera L.) to the infection by phytoplasma Bois noir. Eur. J. Plant Pathol. 2012, 133, 965–974. [Google Scholar] [CrossRef]
- Petridis, A.; Therios, I.; Samouris, G.; Tananaki, C. Salinity-induced changes in phenolic compounds in leaves and roots of four olive cultivars (Olea europaea L.) and their relationship to antioxidant activity. Environ. Exp. Bot. 2012, 79, 37–43. [Google Scholar] [CrossRef]
- Raman, T.; Muthukathan, G. Field Suppression of Fusarium Wilt Disease in Banana by the Combined Application of Native Endophytic and Rhizospheric Bacterial Isolates Possessing Multiple Functions. Phytopathol. Mediterr. 2015, 54, 241–252. [Google Scholar] [CrossRef]
- Wallis, C.M.; Wallingford, A.K.; Chen, J. Effects of cultivar, phenology, and Xylella fastidiosa infection on grapevine Xylem Sap and tissue phenolic content. Physiol. Mol. Plant Pathol. 2013, 84, 28–35. [Google Scholar] [CrossRef]
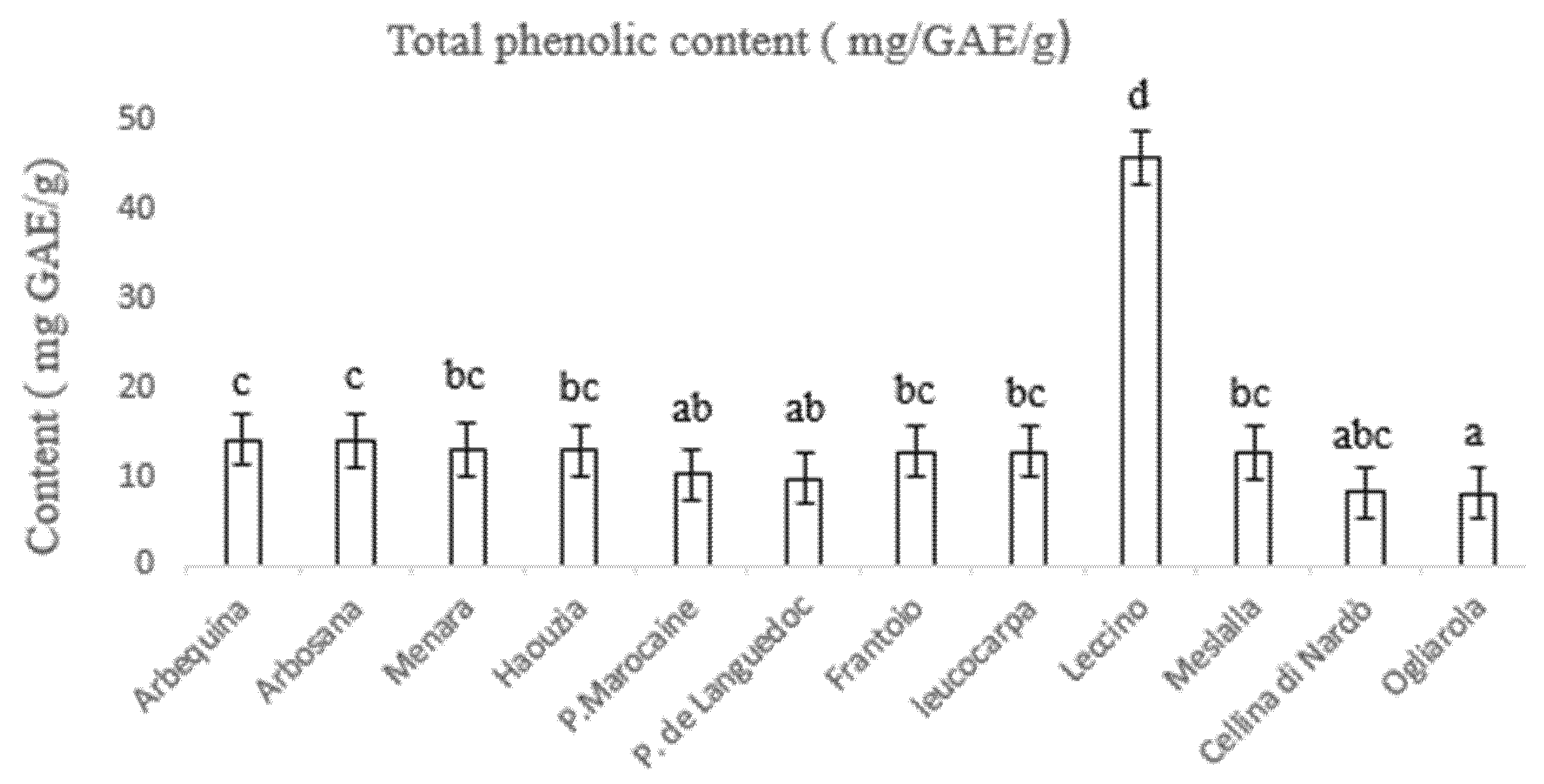
| Ca | Mg | P | Cu | Mn | Na | Zn | Fe | ||
| Reference * | mg/kg or g/kg | 10–14 | 1–1.6 | 1–1.3 | - | 20–36 | <200 | 4–9 | 90–124 |
| Arbiquina | NA | 13.20 | 2.04 | 0.82 | 17.35 | 31.18 | 33.30 | 7.80 | 67.05 |
| Arbozana | NA | 12.52 | 2.09 | 0.93 | 18.50 | 35.38 | 28.60 | 10.23 | 67.60 |
| Menara | NA | 13.35 | 2.01 | 0.72 | 17.03 | 31.18 | 35.75 | 7.86 | 69.46 |
| Haouzia | NA | 14.32 | 1.83 | 0.70 | 17.02 | 29.46 | 35.16 | 7.66 | 70.32 |
| Picholine Marocaine | NA | 19.50 | 0.98 | 0.49 | 10.40 | 23.40 | 41.35 | 5.13 | 80.36 |
| Picholine Languedoc | NA | 20.50 | 0.85 | 0.33 | 9.97 | 21.33 | 41.61 | 4.57 | 80.87 |
| Frantoio | NA | 14.45 | 1.75 | 0.68 | 15.25 | 28.82 | 35.05 | 7.25 | 72.34 |
| Leucocarpa | NA | 14.51 | 1.65 | 0.62 | 15.15 | 28.21 | 34.24 | 7.06 | 72.36 |
| Leccino | NA | 7.02 | 3.62 | 2.51 | 23.82 | 42.63 | 20.21 | 15.61 | 41.3 |
| Meslalla | NA | 15.30 | 1.50 | 0.53 | 14.93 | 26.38 | 40.31 | 7.01 | 73.65 |
| Cellina di Nardò | NA | 27.15 | 0.16 | 0.12 | 7.88 | 18.59 | 48.76 | 4.80 | 93.45 |
| Ogliarola | NA | 27.31 | 0.12 | 0.13 | 7.12 | 17.99 | 49.40 | 4.31 | 95.45 |
| Soil Depth | Sand | Silt | Clay | pH | EC | OM | K2O | P2O5 | CaCo3 |
|---|---|---|---|---|---|---|---|---|---|
| Cm | % | % | % | mS/cm | % | mg·kg−1 | mg·kg−1 | % | |
| 0–35 | 46.8 ± 0.4 | 10.20 | 43.00 | 6.50 | 0.10 | 2.50 | 458.80 | 73.30 | 2.70 |
| 35–70 | 46.10 | 16.10 | 37.60 | 7.80 | 0.10 | 1.60 | 222.50 | 15.10 | 3.10 |
| Mg | Cu | Mn | Na | Zn | |
|---|---|---|---|---|---|
| Soil analysis (mg·kg−1 Fine Fraction) | 237.6 | 19 | 25.3 | 2109 | 0.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
El Handi, K.; Hafidi, M.; Habbadi, K.; El Moujabber, M.; Ouzine, M.; Benbouazza, A.; Sabri, M.; Achbani, E.H. Assessment of Ionomic, Phenolic and Flavonoid Compounds for a Sustainable Management of Xylella fastidiosa in Morocco. Sustainability 2021, 13, 7818. https://doi.org/10.3390/su13147818
El Handi K, Hafidi M, Habbadi K, El Moujabber M, Ouzine M, Benbouazza A, Sabri M, Achbani EH. Assessment of Ionomic, Phenolic and Flavonoid Compounds for a Sustainable Management of Xylella fastidiosa in Morocco. Sustainability. 2021; 13(14):7818. https://doi.org/10.3390/su13147818
Chicago/Turabian StyleEl Handi, Kaoutar, Majida Hafidi, Khaoula Habbadi, Maroun El Moujabber, Mohamed Ouzine, Abdellatif Benbouazza, Miloud Sabri, and El Hassan Achbani. 2021. "Assessment of Ionomic, Phenolic and Flavonoid Compounds for a Sustainable Management of Xylella fastidiosa in Morocco" Sustainability 13, no. 14: 7818. https://doi.org/10.3390/su13147818
APA StyleEl Handi, K., Hafidi, M., Habbadi, K., El Moujabber, M., Ouzine, M., Benbouazza, A., Sabri, M., & Achbani, E. H. (2021). Assessment of Ionomic, Phenolic and Flavonoid Compounds for a Sustainable Management of Xylella fastidiosa in Morocco. Sustainability, 13(14), 7818. https://doi.org/10.3390/su13147818







