Application of Super-Resolution and Advanced Quantitative Microscopy to the Spatio-Temporal Analysis of Influenza Virus Replication
Abstract
1. Introduction
2. Overview of Super-Resolution Microscopy and Novel Imaging Methods to Study Influenza Virus Replication
3. Understanding Virus Structure
4. Understanding Viral Entry and Trafficking
5. Understanding Fusion and Uncoating
6. Understanding Assembly, Budding and Release
7. Understanding Viral Restriction by the Host Cell and Supporting the Development of Antiviral Strategies
8. Limitations of SRM Approaches
9. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Influenza (Seasonal). Available online: https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal) (accessed on 22 June 2020).
- Houser, K.; Subbarao, K. Influenza Vaccines: Challenges and Solutions. Cell Host Microbe 2015, 17, 295–300. [Google Scholar] [CrossRef]
- Mostafa, A.; Abdelwhab, E.M.; Mettenleiter, T.C.; Pleschka, S. Zoonotic Potential of Influenza A Viruses: A Comprehensive Overview. Viruses 2018, 10, 497. [Google Scholar] [CrossRef]
- World Health Organization. Ten threats to global health in 2019. 2019. Available online: https://www.who.int/news-room/feature-stories/ten-threats-to-global-health-in-2019 (accessed on 29 June 2020).
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef]
- Wagner, R.; Matrosovich, M.; Klenk, H.-D. Functional balance between haemagglutinin and neuraminidase in influenza virus infections. Rev. Med. Virol. 2002, 12, 159–166. [Google Scholar] [CrossRef]
- Gamblin, S.; Skehel, J.J. Influenza Hemagglutinin and Neuraminidase Membrane Glycoproteins. J. Biol. Chem. 2010, 285, 28403–28409. [Google Scholar] [CrossRef]
- Badham, M.D.; Rossman, J.S. Filamentous Influenza Viruses. Curr. Clin. Microbiol. Rep. 2016, 3, 155–161. [Google Scholar] [CrossRef]
- Dadonaite, B.; Vijayakrishnan, S.; Fodor, E.; Bhella, D.; Hutchinson, E.C. Filamentous influenza viruses. J. Gen. Virol. 2016, 97, 1755–1764. [Google Scholar] [CrossRef]
- Calder, L.J.; Wasilewski, S.; Berriman, J.A.; Rosenthal, P.B. Structural organization of a filamentous influenza A virus. Proc. Natl. Acad. Sci. USA 2010, 107, 10685–10690. [Google Scholar] [CrossRef]
- Benton, D.J.; Gamblin, S.J.; Rosenthal, P.B.; Skehel, J.J. Structural transitions in influenza haemagglutinin at membrane fusion pH. Nat. Cell Biol. 2020, 583, 150–153. [Google Scholar] [CrossRef]
- Wasilewski, S.; Calder, L.J.; Grant, T.; Rosenthal, P.B. Distribution of surface glycoproteins on influenza A virus determined by electron cryotomography. Vaccine 2012, 30, 7368–7373. [Google Scholar] [CrossRef]
- Huang, B.; Babcock, H.; Zhuang, X. Breaking the Diffraction Barrier: Super-Resolution Imaging of Cells. Cell 2010, 143, 1047–1058. [Google Scholar] [CrossRef]
- Lelek, M.; Di Nunzio, F.; Henriques, R.; Charneau, P.; Arhel, N.J.; Zimmer, C. Superresolution imaging of HIV in infected cells with FlAsH-PALM. Proc. Natl. Acad. Sci. USA 2012, 109, 8564–8569. [Google Scholar] [CrossRef]
- Müller, B.; Heilemann, M. Shedding new light on viruses: Super-resolution microscopy for studying human immunodeficiency virus. Trends Microbiol. 2013, 21, 522–533. [Google Scholar] [CrossRef]
- Chojnacki, J.; Eggeling, C. Super-resolution fluorescence microscopy studies of human immunodeficiency virus. Retrovirology 2018, 15, 1–16. [Google Scholar] [CrossRef]
- Chojnacki, J.; Staudt, T.; Glass, B.; Bingen, P.; Engelhardt, J.; Anders, M.; Schneider, J.; Müller, B.; Hell, S.W.; Kräusslich, H.-G. Maturation-Dependent HIV-1 Surface Protein Redistribution Revealed by Fluorescence Nanoscopy. Science 2012, 338, 524–528. [Google Scholar] [CrossRef]
- Van Engelenburg, S.B.; Shtengel, G.; Sengupta, P.; Waki, K.; Jarnik, M.; Ablan, S.D.; Freed, E.O.; Hess, H.F.; Lippincott-Schwartz, J. Distribution of ESCRT Machinery at HIV Assembly Sites Reveals Virus Scaffolding of ESCRT Subunits. Science 2014, 343, 653–656. [Google Scholar] [CrossRef]
- Kim, D.; Deerinck, T.J.; Sigal, Y.M.; Babcock, H.; Ellisman, M.H.; Zhuang, X. Correlative Stochastic Optical Reconstruction Microscopy and Electron Microscopy. PLoS ONE 2015, 10, e0124581. [Google Scholar] [CrossRef]
- Buttler, C.A.; Pezeshkian, N.; Fernandez, M.V.; Aaron, J.; Norman, S.; Freed, E.O.; Van Engelenburg, S.B. Single molecule fate of HIV-1 envelope reveals late-stage viral lattice incorporation. Nat. Commun. 2018, 9, 1–15. [Google Scholar] [CrossRef]
- Laine, R.F.; Goodfellow, G.; Young, L.J.; Travers, J.; Carroll, D.; Dibben, O.; Bright, H.; Kaminski, C.F. Structured illumination microscopy combined with machine learning enables the high throughput analysis and classification of virus structure. eLife 2018, 7. [Google Scholar] [CrossRef]
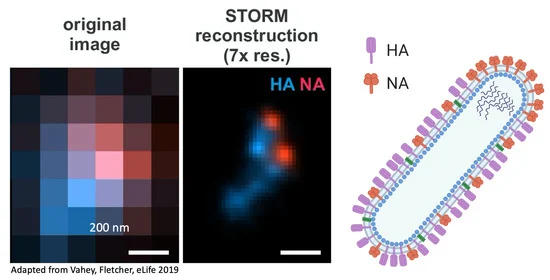
- Vahey, M.D.; Fletcher, D.A. Low-Fidelity Assembly of Influenza A Virus Promotes Escape from Host Cells. Cell 2019, 176, 281–294.e19. [Google Scholar] [CrossRef]
- Burdick, R.C.; Li, C.; Munshi, M.; Rawson, J.M.O.; Nagashima, K.; Hu, W.-S.; Pathak, V.K. HIV-1 uncoats in the nucleus near sites of integration. Proc. Natl. Acad. Sci. USA 2020, 117, 5486–5493. [Google Scholar] [CrossRef]
- Bhagwat, A.R.; Le Sage, V.; Nturibi, E.; Kulej, K.; Jones, J.E.; Guo, M.; Kim, E.T.; Garcia, B.A.; Weitzman, M.D.; Shroff, H.; et al. Quantitative live cell imaging reveals influenza virus manipulation of Rab11A transport through reduced dynein association. Nat. Commun. 2020, 11, 1–14. [Google Scholar] [CrossRef]
- Laine, R.F.; Albecka, A.; Van De Linde, S.; Rees, E.J.; Crump, C.M.; Kaminski, C.F. Structural analysis of herpes simplex virus by optical super-resolution imaging. Nat. Commun. 2015, 6, 5980. [Google Scholar] [CrossRef]
- Sieben, C.; Sezgin, E.; Eggeling, C.; Manley, S. Influenza A viruses use multivalent sialic acid clusters for cell binding and receptor activation. PLoS Pathog. 2020, 16, e1008656. [Google Scholar] [CrossRef]
- Hell, S.W.; Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: Stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 1994, 19, 780–782. [Google Scholar] [CrossRef]
- Baharom, F.; Thomas, O.S.; Lepzien, R.; Mellman, I.; Chalouni, C.; Smed-Sörensen, A. Visualization of early influenza A virus trafficking in human dendritic cells using STED microscopy. PLoS ONE 2017, 12, e0177920. [Google Scholar] [CrossRef]
- Gustafsson, M.G.L. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. SHORT COMMUNICATION. J. Microsc. 2000, 198, 82–87. [Google Scholar] [CrossRef]
- Banerjee, I.; Miyake, Y.; Nobs, S.P.; Schneider, C.; Horvath, P.; Kopf, M.; Matthias, P.; Helenius, A.; Yamauchi, Y. Influenza A virus uses the aggresome processing machinery for host cell entry. Science 2014, 346, 473–477. [Google Scholar] [CrossRef]
- Gray, R.D.M.; Albrecht, D.; Beerli, C.; Huttunen, M.; Cohen, G.H.; White, I.J.; Burden, J.J.; Henriques, R.; Mercer, J. Nanoscale polarization of the entry fusion complex of vaccinia virus drives efficient fusion. Nat. Microbiol. 2019, 4, 1636–1644. [Google Scholar] [CrossRef]
- Betzig, E.; Patterson, G.H.; Sougrat, R.; Lindwasser, O.W.; Olenych, S.; Bonifacino, J.S.; Davidson, M.W.; Lippincott-Schwartz, J.; Hess, H.F. Imaging Intracellular Fluorescent Proteins at Nanometer Resolution. Science 2006, 313, 1642–1645. [Google Scholar] [CrossRef]
- Hess, S.T.; Girirajan, T.P.K.; Mason, M.D. Ultra-High Resolution Imaging by Fluorescence Photoactivation Localization Microscopy. Biophys. J. 2006, 91, 4258–4272. [Google Scholar] [CrossRef]
- Rust, M.J.; Bates, M.; Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 2006, 3, 793–796. [Google Scholar] [CrossRef]
- Heilemann, M.; Van De Linde, S.; Schüttpelz, M.; Kasper, R.; Seefeldt, B.; Mukherjee, A.; Tinnefeld, P.; Sauer, M. Subdiffraction-Resolution Fluorescence Imaging with Conventional Fluorescent Probes. Angew. Chem. Int. Ed. 2008, 47, 6172–6176. [Google Scholar] [CrossRef]
- Sharonov, A.; Hochstrasser, R.M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl. Acad. Sci. USA 2006, 103, 18911–18916. [Google Scholar] [CrossRef]
- Fölling, J.; Bossi, M.; Bock, H.; Medda, R.; Wurm, C.A.; Hein, B.; Jakobs, S.; Eggeling, C.; Hell, S.W. Fluorescence nanoscopy by ground-state depletion and single-molecule return. Nat. Methods 2008, 5, 943–945. [Google Scholar] [CrossRef]
- Manley, S.; Gillette, J.M.; Patterson, G.H.; Shroff, H.; Hess, H.F.; Betzig, E.; Lippincott-Schwartz, J. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 2008, 5, 155–157. [Google Scholar] [CrossRef]
- Vahey, M.D.; Fletcher, D.A. Influenza A virus surface proteins are organized to help penetrate host mucus. eLife 2019, 8. [Google Scholar] [CrossRef]
- Chozinski, T.J.; Halpern, A.R.; Okawa, H.; Kim, H.-J.; Tremel, G.J.; Wong, R.O.; Vaughan, J.C. Expansion microscopy with conventional antibodies and fluorescent proteins. Nat. Methods 2016, 13, 485–488. [Google Scholar] [CrossRef]
- Takeda, M.; Leser, G.P.; Russell, C.J.; Lamb, R.A. Influenza virus hemagglutinin concentrates in lipid raft microdomains for efficient viral fusion. Proc. Natl. Acad. Sci. USA 2003, 100, 14610–14617. [Google Scholar] [CrossRef]
- Simons, K.; Ikonen, E. Functional rafts in cell membranes. Nature 1997, 387, 569–572. [Google Scholar] [CrossRef]
- Barman, S.; Nayak, D.P. Analysis of the Transmembrane Domain of Influenza Virus Neuraminidase, a Type II Transmembrane Glycoprotein, for Apical Sorting and Raft Association. J. Virol. 2000, 74, 6538–6545. [Google Scholar] [CrossRef] [PubMed]
- Hess, S.T.; Gould, T.J.; Gudheti, M.V.; Maas, S.A.; Mills, K.D.; Zimmerberg, J. Dynamic clustered distribution of hemagglutinin resolved at 40 nm in living cell membranes discriminates between raft theories. Proc. Natl. Acad. Sci. USA 2007, 104, 17370–17375. [Google Scholar] [CrossRef] [PubMed]
- Steinhauer, C.; Forthmann, C.; Vogelsang, J.; Tinnefeld, P. Superresolution Microscopy on the Basis of Engineered Dark States. J. Am. Chem. Soc. 2008, 130, 16840–16841. [Google Scholar] [CrossRef] [PubMed]
- Itano, M.S.; Steinhauer, C.; Schmied, J.J.; Forthmann, C.; Liu, P.; Neumann, A.K.; Thompson, N.L.; Tinnefeld, P.; Jacobson, K. Super-Resolution Imaging of C-Type Lectin and Influenza Hemagglutinin Nanodomains on Plasma Membranes Using Blink Microscopy. Biophys. J. 2012, 102, 1534–1542. [Google Scholar] [CrossRef]
- Chen, F.; Tillberg, P.W.; Boyden, E.S. Expansion microscopy. Science 2015, 347, 543–548. [Google Scholar] [CrossRef]
- Chang, J.-B.; Chen, F.; Yoon, Y.-G.; Jung, E.E.; Babcock, H.; Kang, J.S.; Asano, S.; Suk, H.-J.; Pak, N.; Tillberg, P.W.; et al. Iterative expansion microscopy. Nat. Methods 2017, 14, 593–599. [Google Scholar] [CrossRef]
- Gambarotto, D.; Zwettler, F.U.; Le Guennec, M.; Schmidt-Cernohorska, M.; Fortun, D.; Borgers, S.; Heine, J.; Schloetel, J.-G.; Reuss, M.; Unser, M.; et al. Imaging cellular ultrastructures using expansion microscopy (U-ExM). Nat. Methods 2018, 16, 71–74. [Google Scholar] [CrossRef]
- Mascheroni, L.; Scherer, K.M.; Manton, J.D.; Ward, E.; Dibben, O.; Kaminski, C.F. Combining sample expansion and light sheet microscopy for the volumetric imaging of virus-infected cells with optical super-resolution. bioRxiv 2020. [Google Scholar] [CrossRef]
- Aho, V.; Mäntylä, E.; Ekman, A.; Hakanen, S.; Mattola, S.; Chen, J.-H.; Weinhardt, V.; Ruokolainen, V.; Sodeik, B.; Larabell, C.; et al. Quantitative Microscopy Reveals Stepwise Alteration of Chromatin Structure during Herpesvirus Infection. Viruses 2019, 11, 935. [Google Scholar] [CrossRef]
- Gao, R.; Yu, C.C.J.; Gao, L.; Piatkevich, K.D.; Neve, R.L.; Upadhyayula, S.; Boyden, E.S. A highly homogeneous expansion microscopy polymer composed of tetrahedron-like monomers. bioRxiv 2019, 1, 814111. [Google Scholar]
- Gambarotto, D.; Hamel, V.; Guichard, P. Ultrastructure expansion microscopy (U-ExM). In Methods in Cell Biology; Elsevier Inc.: Amsterdam, The Netherlands, 2020; pp. 1–25. [Google Scholar]
- Power, R.M.; Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat. Methods 2017, 14, 360–373. [Google Scholar] [CrossRef] [PubMed]
- Griffin, B.A. Specific Covalent Labeling of Recombinant Protein Molecules Inside Live Cells. Science 1998, 281, 269–272. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Lin, A.J.; Golan, D.E.; Walsh, C.T. Site-specific protein labeling by Sfp phosphopantetheinyl transferase. Nat. Protoc. 2006, 1, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-W.; Ting, A.Y. Transglutaminase-Catalyzed Site-Specific Conjugation of Small-Molecule Probes to Proteins in Vitro and on the Surface of Living Cells. J. Am. Chem. Soc. 2006, 128, 4542–4543. [Google Scholar] [CrossRef] [PubMed]
- Theile, C.S.; Witte, M.D.; Blom, A.E.M.; Kundrat, L.; Ploegh, H.L.; Guimaraes, C.P. Site-specific N-terminal labeling of proteins using sortase-mediated reactions. Nat. Protoc. 2013, 8, 1800–1807. [Google Scholar] [CrossRef]
- Lakadamyali, M.; Rust, M.J.; Babcock, H.P.; Zhuang, X. Visualizing infection of individual influenza viruses. Proc. Natl. Acad. Sci. USA 2003, 100, 9280–9285. [Google Scholar] [CrossRef]
- Brandenburg, B.; Zhuang, X. Virus trafficking—learning from single-virus tracking. Nat. Rev. Genet. 2007, 5, 197–208. [Google Scholar] [CrossRef]
- Liu, S.-L.; Wang, Z.-G.; Xie, H.; Liu, A.-A.; Lamb, D.C.; Pang, D.-W. Single-Virus Tracking: From Imaging Methodologies to Virological Applications. Chem. Rev. 2020, 120, 1936–1979. [Google Scholar] [CrossRef]
- Liu, S.-L.; Zhang, Z.-L.; Tian, Z.-Q.; Zhao, H.-S.; Liu, H.; Sun, E.-Z.; Xiao, G.F.; Zhang, W.; Wang, H.-Z.; Pang, D.-W. Effectively and Efficiently Dissecting the Infection of Influenza Virus by Quantum-Dot-Based Single-Particle Tracking. ACS Nano 2011, 6, 141–150. [Google Scholar] [CrossRef]
- Liu, S.-L.; Zhang, L.-J.; Wang, Z.-G.; Zhang, Z.-L.; Wu, Q.-M.; Sun, E.-Z.; Shi, Y.-B.; Pang, D.-W. Globally Visualizing the Microtubule-Dependent Transport Behaviors of Influenza Virus in Live Cells. Anal. Chem. 2014, 86, 3902–3908. [Google Scholar] [CrossRef]
- Qin, C.; Li, W.; Li, Q.; Yin, W.; Zhang, X.; Zhang, Z.; Zhang, X.-E.; Zhang, M. Real-time dissection of dynamic uncoating of individual influenza viruses. Proc. Natl. Acad. Sci. USA 2019, 116, 2577–2582. [Google Scholar] [CrossRef] [PubMed]
- Weiss, S. Fluorescence Spectroscopy of Single Biomolecules. Science 1999, 283, 1676–1683. [Google Scholar] [CrossRef] [PubMed]
- Das, D.K.; Govindan, R.; Nikić-Spiegel, I.; Krammer, F.; Lemke, E.A.; Munro, J.B. Direct Visualization of the Conformational Dynamics of Single Influenza Hemagglutinin Trimers. Cell 2018, 174, 926–937.e12. [Google Scholar] [CrossRef] [PubMed]
- Femino, A.M. Visualization of Single RNA Transcripts in Situ. Science 1998, 280, 585–590. [Google Scholar] [CrossRef] [PubMed]
- Chou, Y.-Y.; Vafabakhsh, R.; Doğanay, S.; Gao, Q.; Ha, T.; Palese, P. One influenza virus particle packages eight unique viral RNAs as shown by FISH analysis. Proc. Natl. Acad. Sci. USA 2012, 109, 9101–9106. [Google Scholar] [CrossRef] [PubMed]
- Chou, Y.-Y.; Heaton, N.S.; Gao, Q.; Palese, P.; Singer, R.H.; Lionnet, T. Colocalization of Different Influenza Viral RNA Segments in the Cytoplasm before Viral Budding as Shown by Single-molecule Sensitivity FISH Analysis. PLoS Pathog. 2013, 9, e1003358. [Google Scholar] [CrossRef]
- Von Chamier, L.; Laine, R.F.; Henriques, R. Artificial intelligence for microscopy: What you should know. Biochem. Soc. Trans. 2019, 47, 1029–1040. [Google Scholar] [CrossRef]
- Belthangady, C.; Royer, L.A. Applications, promises, and pitfalls of deep learning for fluorescence image reconstruction. Nat. Methods 2019, 16, 1215–1225. [Google Scholar] [CrossRef]
- Schelker, M.; Mair, C.M.; Jolmes, F.; Welke, R.-W.; Klipp, E.; Herrmann, A.; Flöttmann, M.; Sieben, C. Viral RNA Degradation and Diffusion Act as a Bottleneck for the Influenza A Virus Infection Efficiency. PLoS Comput. Biol. 2016, 12, e1005075. [Google Scholar] [CrossRef]
- Schwille, P.; Korlach, J.; Webb, W.W. Fluorescence correlation spectroscopy with single-molecule sensitivity on cell and model membranes. Cytometry 1999, 36, 176–182. [Google Scholar] [CrossRef]
- Chojnacki, J.; Waithe, D.; Carravilla, P.; Huarte, N.; Galiani, S.; Enderlein, J.; Eggeling, C. Envelope glycoprotein mobility on HIV-1 particles depends on the virus maturation state. Nat. Commun. 2017, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Haralampiev, I.; Prisner, S.; Nitzan, M.; Schade, M.; Jolmes, F.; Schreiber, M.; Loidolt-Krüger, M.; Jongen, K.; Chamiolo, J.; Nilson, N.; et al. Selective flexible packaging pathways of the segmented genome of influenza A virus. Nat. Commun. 2020, 11, 1–13. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Sun, E.; Bujny, M.V.; Kim, D.; Davidson, M.W.; Zhuang, X. Dual Function of CD81 in Influenza Virus Uncoating and Budding. PLoS Pathog. 2013, 9, e1003701. [Google Scholar] [CrossRef] [PubMed]
- Sieczkarski, S.B.; Whittaker, G.R. Characterization of the host cell entry of filamentous influenza virus. Arch. Virol. 2005, 150, 1783–1796. [Google Scholar] [CrossRef] [PubMed]
- De Vries, E.; Tscherne, D.M.; Wienholts, M.J.; Cobos-Jiménez, V.; Scholte, F.; García-Sastre, A.; Rottier, P.J.M.; De Haan, C.A.M. Dissection of the Influenza A Virus Endocytic Routes Reveals Macropinocytosis as an Alternative Entry Pathway. PLoS Pathog. 2011, 7, e1001329. [Google Scholar] [CrossRef] [PubMed]
- De Conto, F.; Covan, S.; Arcangeletti, M.C.; Orlandini, G.; Gatti, R.; Dettori, G.; Chezzi, C. Differential infectious entry of human influenza A/NWS/33 virus (H1N1) in mammalian kidney cells. Virus Res. 2011, 155, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Edinger, T.O.; Pohl, M.O.; Stertz, S. Entry of influenza A virus: Host factors and antiviral targets. J. Gen. Virol. 2014, 95, 263–277. [Google Scholar] [CrossRef]
- Karakus, U.; Pohl, M.O.; Stertz, S. Breaking the Convention: Sialoglycan Variants, Coreceptors, and Alternative Receptors for Influenza A Virus Entry. J. Virol. 2019, 94, 01357-19. [Google Scholar] [CrossRef]
- Eierhoff, T.; Hrincius, E.R.; Rescher, U.; Ludwig, S.; Ehrhardt, C. The Epidermal Growth Factor Receptor (EGFR) Promotes Uptake of Influenza A Viruses (IAV) into Host Cells. PLoS Pathog. 2010, 6, e1001099. [Google Scholar] [CrossRef]
- Stauffer, S.; Feng, Y.; Nebioglu, F.; Heilig, R.; Picotti, P.; Helenius, A. Stepwise Priming by Acidic pH and a High K+ Concentration Is Required for Efficient Uncoating of Influenza A Virus Cores after Penetration. J. Virol. 2014, 88, 13029–13046. [Google Scholar] [CrossRef]
- Li, S.; Sieben, C.; Ludwig, K.; Höfer, C.T.; Chiantia, S.; Herrmann, A.; Eghiaian, F.; Schaap, I.A. pH-Controlled Two-Step Uncoating of Influenza Virus. Biophys. J. 2014, 106, 1447–1456. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhuang, X. Epsin 1 is a cargo-specific adaptor for the clathrin-mediated endocytosis of the influenza virus. Proc. Natl. Acad. Sci. USA 2008, 105, 11790–11795. [Google Scholar] [CrossRef] [PubMed]
- Floyd, D.L.; Ragains, J.R.; Skehel, J.J.; Harrison, S.C.; Van Oijen, A.M. Single-particle kinetics of influenza virus membrane fusion. Proc. Natl. Acad. Sci. USA 2008, 105, 15382–15387. [Google Scholar] [CrossRef] [PubMed]
- Sakai, T.; Ohuchi, M.; Imai, M.; Mizuno, T.; Kawasaki, K.; Kuroda, K.; Yamashina, S. Dual Wavelength Imaging Allows Analysis of Membrane Fusion of Influenza Virus inside Cells. J. Virol. 2006, 80, 2013–2018. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Huh, H.; Lee, S.-H.; Huang, F. Three-Dimensional Single-Molecule Localization Microscopy in Whole-Cell and Tissue Specimens. Annu. Rev. Biomed. Eng. 2020, 22, 155–184. [Google Scholar] [CrossRef]
- Miyake, Y.; Keusch, J.J.; Decamps, L.; Ho-Xuan, H.; Iketani, S.; Gut, H.; Kutay, U.; Helenius, A.; Yamauchi, Y. Influenza virus uses transportin 1 for vRNP debundling during cell entry. Nat. Microbiol. 2019, 4, 578–586. [Google Scholar] [CrossRef]
- Gao, Q.; Chou, Y.-Y.; Doğanay, S.; Vafabakhsh, R.; Ha, T.; Palese, P. The Influenza A Virus PB2, PA, NP, and M Segments Play a Pivotal Role during Genome Packaging. J. Virol. 2012, 86, 7043–7051. [Google Scholar] [CrossRef]
- Nakatsu, S.; Sagara, H.; Sakai-Tagawa, Y.; Sugaya, N.; Noda, T.; Kawaoka, Y. Complete and Incomplete Genome Packaging of Influenza A and B Viruses. mBio 2016, 7, e01248-16. [Google Scholar] [CrossRef]
- Noda, T.; Murakami, S.; Nakatsu, S.; Imai, H.; Muramoto, Y.; Shindo, K.; Sagara, H.; Kawaoka, Y. Importance of the 1+7 configuration of ribonucleoprotein complexes for influenza A virus genome packaging. Nat. Commun. 2018, 9, 1–10. [Google Scholar] [CrossRef]
- Gavazzi, C.; Yver, M.; Isel, C.; Smyth, R.P.; Rosa-Calatrava, M.; Lina, B.; Moulès, V.; Marquet, R. A functional sequence-specific interaction between influenza A virus genomic RNA segments. Proc. Natl. Acad. Sci. USA 2013, 110, 16604–16609. [Google Scholar] [CrossRef]
- Essere, B.; Yver, M.; Gavazzi, C.; Terrier, O.; Isel, C.; Fournier, E.; Giroux, F.; Textoris, J.; Julien, T.; Socratous, C.; et al. Critical role of segment-specific packaging signals in genetic reassortment of influenza A viruses. Proc. Natl. Acad. Sci. USA 2013, 110, E3840–E3848. [Google Scholar] [CrossRef]
- Inagaki, A.; Goto, H.; Kakugawa, S.; Ozawa, M.; Kawaoka, Y. Competitive Incorporation of Homologous Gene Segments of Influenza A Virus into Virions. J. Virol. 2012, 86, 10200–10202. [Google Scholar] [CrossRef] [PubMed]
- Lakdawala, S.S.; Wu, Y.; Wawrzusin, P.; Kabat, J.; Broadbent, A.J.; Lamirande, E.W.; Fodor, E.; Altan-Bonnet, N.; Shroff, H.; Subbarao, K. Influenza A Virus Assembly Intermediates Fuse in the Cytoplasm. PLoS Pathog. 2014, 10, e1003971. [Google Scholar] [CrossRef]
- Eisfeld, A.J.; Kawakami, E.; Watanabe, T.; Neumann, G.; Kawaoka, Y. RAB11A Is Essential for Transport of the Influenza Virus Genome to the Plasma Membrane. J. Virol. 2011, 85, 6117–6126. [Google Scholar] [CrossRef] [PubMed]
- Vale-Costa, S.; Alenquer, M.; Sousa, A.L.; Kellen, B.; Ramalho, J.; Tranfield, E.M.; Amorim, M.J. Influenza A virus ribonucleoproteins modulate host recycling by competing with Rab11 effectors. J. Cell Sci. 2016, 129, 1697–1710. [Google Scholar] [CrossRef] [PubMed]
- Cabantous, S.; Terwilliger, T.C.; Waldo, G.S. Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein. Nat. Biotechnol. 2005, 23, 102–107. [Google Scholar] [CrossRef] [PubMed]
- Avilov, S.; Magnus, J.; Cusack, S.; Naffakh, N. Time-Resolved Visualisation of Nearly-Native Influenza A Virus Progeny Ribonucleoproteins and Their Individual Components in Live Infected Cells. PLoS ONE 2016, 11, e0149986. [Google Scholar] [CrossRef]
- Chen, B.J.; Leser, G.P.; Morita, E.; Lamb, R.A. Influenza Virus Hemagglutinin and Neuraminidase, but Not the Matrix Protein, Are Required for Assembly and Budding of Plasmid-Derived Virus-Like Particles. J. Virol. 2007, 81, 7111–7123. [Google Scholar] [CrossRef]
- Rossman, J.S.; Jing, X.; Leser, G.P.; Balannik, V.; Pinto, L.H.; Lamb, R.A. Influenza Virus M2 Ion Channel Protein Is Necessary for Filamentous Virion Formation. J. Virol. 2010, 84, 5078–5088. [Google Scholar] [CrossRef]
- Rossman, J.S.; Jing, X.; Leser, G.P.; Lamb, R.A. Influenza Virus M2 Protein Mediates ESCRT-Independent Membrane Scission. Cell 2010, 142, 902–913. [Google Scholar] [CrossRef]
- Gudheti, M.V.; Curthoys, N.M.; Gould, T.J.; Kim, D.; Gunewardene, M.S.; Gabor, K.A.; Gosse, J.A.; Kim, C.H.; Zimmerberg, J.; Hess, S.T. Actin Mediates the Nanoscale Membrane Organization of the Clustered Membrane Protein Influenza Hemagglutinin. Biophys. J. 2013, 104, 2182–2192. [Google Scholar] [CrossRef] [PubMed]
- Curthoys, N.M.; Mlodzianoski, M.J.; Parent, M.; Butler, M.B.; Raut, P.; Wallace, J.; Lilieholm, J.; Mehmood, K.; Maginnis, M.S.; Waters, H.; et al. Influenza Hemagglutinin Modulates Phosphatidylinositol 4,5-Bisphosphate Membrane Clustering. Biophys. J. 2019, 116, 893–909. [Google Scholar] [CrossRef] [PubMed]
- Kolpe, A.; Arista-Romero, M.; Schepens, B.; Pujals, S.; Saelens, X.; Albertazzi, L. Super-resolution microscopy reveals significant impact of M2e-specific monoclonal antibodies on influenza A virus filament formation at the host cell surface. Sci. Rep. 2019, 9, 4450. [Google Scholar] [CrossRef] [PubMed]
- Pileri, P.; Uematsu, Y.; Campagnoli, S.; Galli, G.; Falugi, F.; Petracca, R.; Weiner, A.J.; Houghton, M.; Rosa, D.; Grandi, G.; et al. Binding of Hepatitis C Virus to CD81. Science 1998, 282, 938–941. [Google Scholar] [CrossRef]
- Muranyi, W.; Malkusch, S.; Müller, B.; Heilemann, M.; Kräusslich, H.-G. Super-Resolution Microscopy Reveals Specific Recruitment of HIV-1 Envelope Proteins to Viral Assembly Sites Dependent on the Envelope C-Terminal Tail. PLoS Pathog. 2013, 9, e1003198. [Google Scholar] [CrossRef]
- Pezeshkian, N.; Groves, N.S.; Van Engelenburg, S.B. Single-molecule imaging of HIV-1 envelope glycoprotein dynamics and Gag lattice association exposes determinants responsible for virus incorporation. Proc. Natl. Acad. Sci. USA 2019, 116, 25269–25277. [Google Scholar] [CrossRef]
- Feeley, E.M.; Sims, J.S.; John, S.P.; Chin, C.R.; Pertel, T.; Chen, L.-M.; Gaiha, G.D.; Ryan, B.J.; Donis, R.O.; Elledge, S.J.; et al. IFITM3 Inhibits Influenza A Virus Infection by Preventing Cytosolic Entry. PLoS Pathog. 2011, 7, e1002337. [Google Scholar] [CrossRef]
- Spence, J.S.; He, R.; Hoffmann, H.-H.; Das, T.; Thinon, E.; Rice, C.M.; Peng, T.; Chandran, K.; Hang, H.C. IFITM3 directly engages and shuttles incoming virus particles to lysosomes. Nat. Chem. Biol. 2019, 15, 259–268. [Google Scholar] [CrossRef]
- Kummer, S.; Avinoam, O.; Kräusslich, H.-G. IFITM3 Clusters on Virus Containing Endosomes and Lysosomes Early in the Influenza A Infection of Human Airway Epithelial Cells. Viruses 2019, 11, 548. [Google Scholar] [CrossRef]
- Pereira, P.M.; Gustafsson, N.; Marsh, M.; Mhlanga, M.M.; Henriques, R. Super-beacons: Open-source probes with spontaneous tuneable blinking compatible with live-cell super-resolution microscopy. Traffic 2020, 21, 375–385. [Google Scholar] [CrossRef]
- Liedmann, S.; Hrincius, E.R.; Guy, C.S.; Anhlan, D.; Dierkes, R.; Carter, R.R.; Wu, G.; Staeheli, P.; Green, D.R.; Wolff, T.; et al. Viral suppressors of the RIG-I-mediated interferon response are pre-packaged in influenza virions. Nat. Commun. 2014, 5, 5645. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, I.; Yamauchi, Y.; Helenius, A.; Horvath, P. High-Content Analysis of Sequential Events during the Early Phase of Influenza A Virus Infection. PLoS ONE 2013, 8, e68450. [Google Scholar] [CrossRef] [PubMed]
- Harke, B.; Keller, J.; Ullal, C.K.; Westphal, V.; Schönle, A.; Hell, S.W. Resolution scaling in STED microscopy. Opt. Express 2008, 16, 4154–4162. [Google Scholar] [CrossRef] [PubMed]
- Wäldchen, S.; Lehmann, J.; Klein, T.; Van De Linde, S.; Sauer, M. Light-induced cell damage in live-cell super-resolution microscopy. Sci. Rep. 2015, 5, srep15348. [Google Scholar] [CrossRef]
- Tosheva, K.L.; Yuan, Y.; Pereira, P.; Culley, S.; Henriques, R. Between life and death: Strategies to reduce phototoxicity in super-resolution microscopy. J. Phys. D Appl. Phys. 2020, 53, 163001. [Google Scholar] [CrossRef]
- Chen, B.-C.; Legant, W.R.; Wang, K.; Shao, L.; Milkie, D.E.; Davidson, M.W.; Janetopoulos, C.; Wu, X.S.; Hammer, J.A.; Liu, Z.; et al. Lattice light-sheet microscopy: Imaging molecules to embryos at high spatiotemporal resolution. Science 2014, 346, 1257998. [Google Scholar] [CrossRef]
- Wang, L.; Tran, M.; D’Este, E.; Roberti, J.; Koch, B.; Xue, L.; Johnsson, K. A general strategy to develop cell permeable and fluorogenic probes for multicolour nanoscopy. Nat. Chem. 2020, 12, 165–172. [Google Scholar] [CrossRef]
- Wang, L.; Frei, M.S.; Salim, A.; Johnsson, K. Small-Molecule Fluorescent Probes for Live-Cell Super-Resolution Microscopy. J. Am. Chem. Soc. 2019, 141, 2770–2781. [Google Scholar] [CrossRef]
- Jradi, F.M.; Lavis, L.D. Chemistry of Photosensitive Fluorophores for Single-Molecule Localization Microscopy. ACS Chem. Biol. 2019, 14, 1077–1090. [Google Scholar] [CrossRef]
- Zheng, Q.; Ayala, A.X.; Chung, I.; Weigel, A.V.; Ranjan, A.; Falco, N.; Grimm, J.B.; Tkachuk, A.N.; Wu, C.; Lippincott-Schwartz, J.; et al. Rational Design of Fluorogenic and Spontaneously Blinking Labels for Super-Resolution Imaging. ACS Cent. Sci. 2019, 5, 1602–1613. [Google Scholar] [CrossRef]
- Von Chamier, L.; Laine, R.F.; Jukkala, J.; Spahn, C.; Krentzel, D.; Nehme, E.; Lerche, M.; Hernández-Pérez, S.; Mattila, P.K.; Karinou, E.; et al. ZeroCostDL4Mic: An open platform to use Deep-Learning in Microscopy. bioRxiv 2020. [Google Scholar] [CrossRef]
- Gray, R.D.M.; Beerli, C.; Pereira, P.M.; Scherer, K.M.; Samolej, J.; Bleck, C.K.E.; Mercer, J.; Henriques, R. VirusMapper: Open-source nanoscale mapping of viral architecture through super-resolution microscopy. Sci. Rep. 2016, 6, 29132. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Touizer, E.; Sieben, C.; Henriques, R.; Marsh, M.; Laine, R.F. Application of Super-Resolution and Advanced Quantitative Microscopy to the Spatio-Temporal Analysis of Influenza Virus Replication. Viruses 2021, 13, 233. https://doi.org/10.3390/v13020233
Touizer E, Sieben C, Henriques R, Marsh M, Laine RF. Application of Super-Resolution and Advanced Quantitative Microscopy to the Spatio-Temporal Analysis of Influenza Virus Replication. Viruses. 2021; 13(2):233. https://doi.org/10.3390/v13020233
Chicago/Turabian StyleTouizer, Emma, Christian Sieben, Ricardo Henriques, Mark Marsh, and Romain F. Laine. 2021. "Application of Super-Resolution and Advanced Quantitative Microscopy to the Spatio-Temporal Analysis of Influenza Virus Replication" Viruses 13, no. 2: 233. https://doi.org/10.3390/v13020233
APA StyleTouizer, E., Sieben, C., Henriques, R., Marsh, M., & Laine, R. F. (2021). Application of Super-Resolution and Advanced Quantitative Microscopy to the Spatio-Temporal Analysis of Influenza Virus Replication. Viruses, 13(2), 233. https://doi.org/10.3390/v13020233