The Regulation of Flavivirus Infection by Hijacking Exosome-Mediated Cell–Cell Communication: New Insights on Virus–Host Interactions
Abstract
1. Introduction
2. Isolation Techniques and Heterogeneity in Exosomes Size
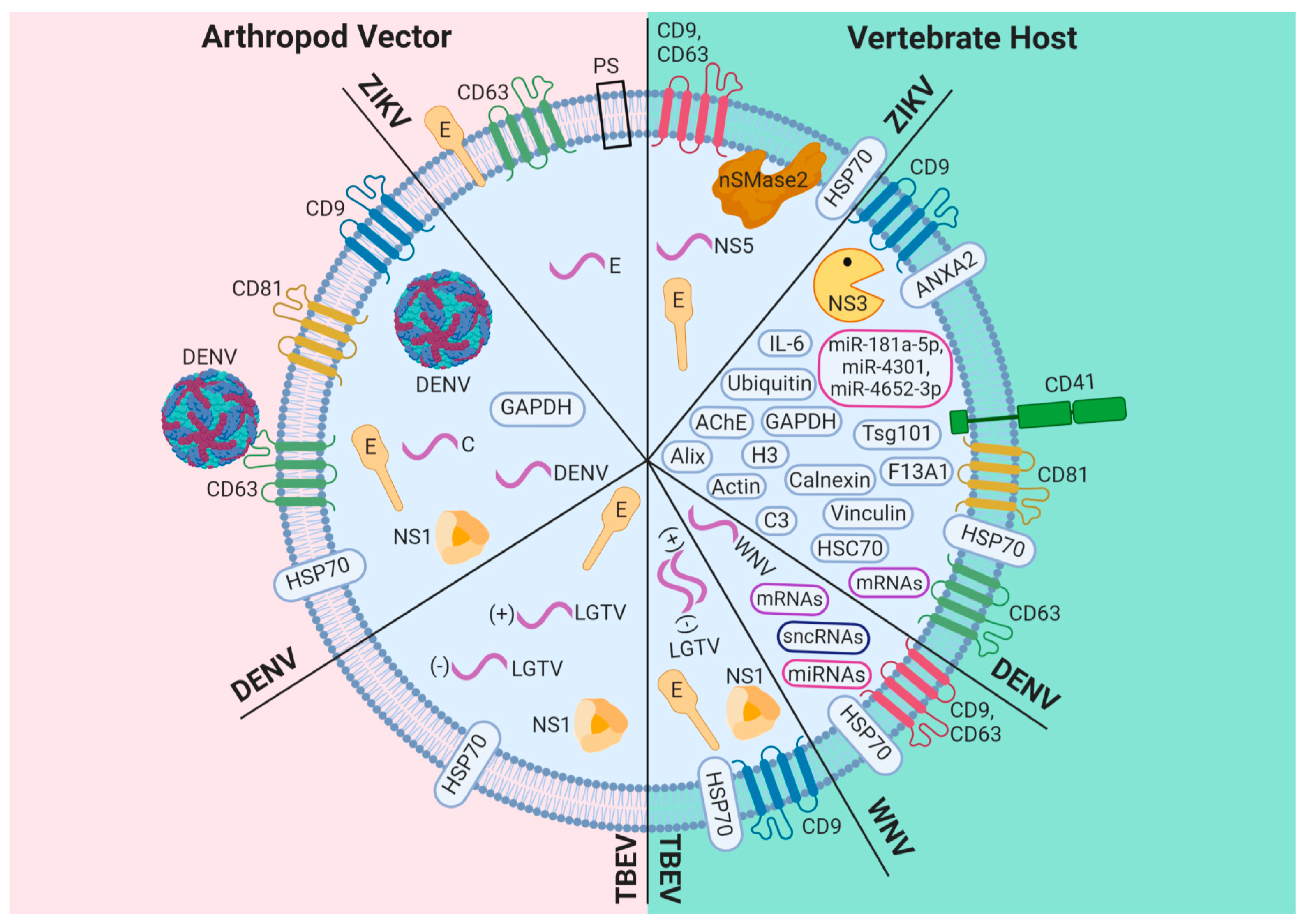
3. Exosomes: A New Mechanism of Viral Dissemination Within and Among Hosts
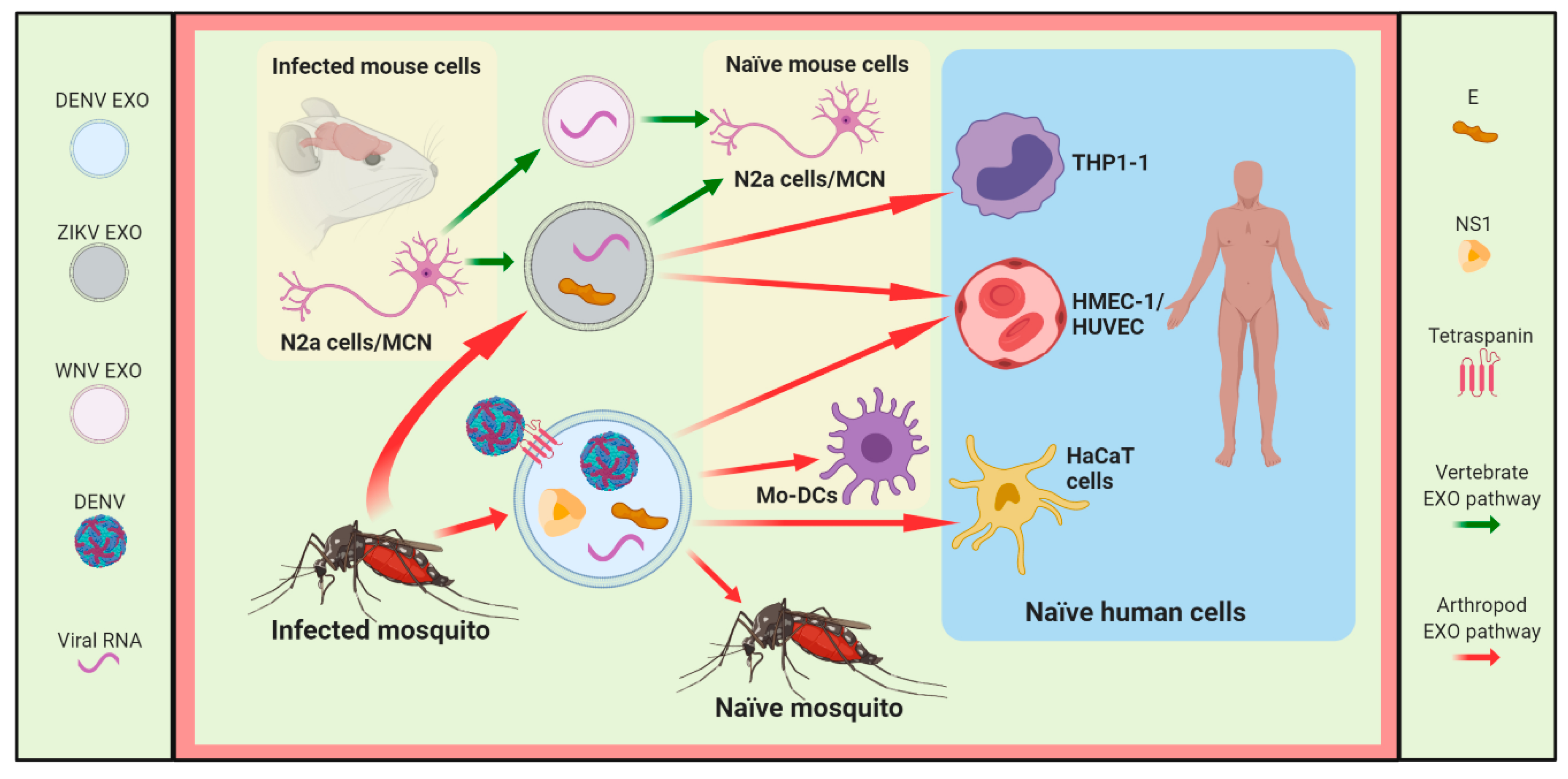
3.1. DENV
3.2. ZIKV
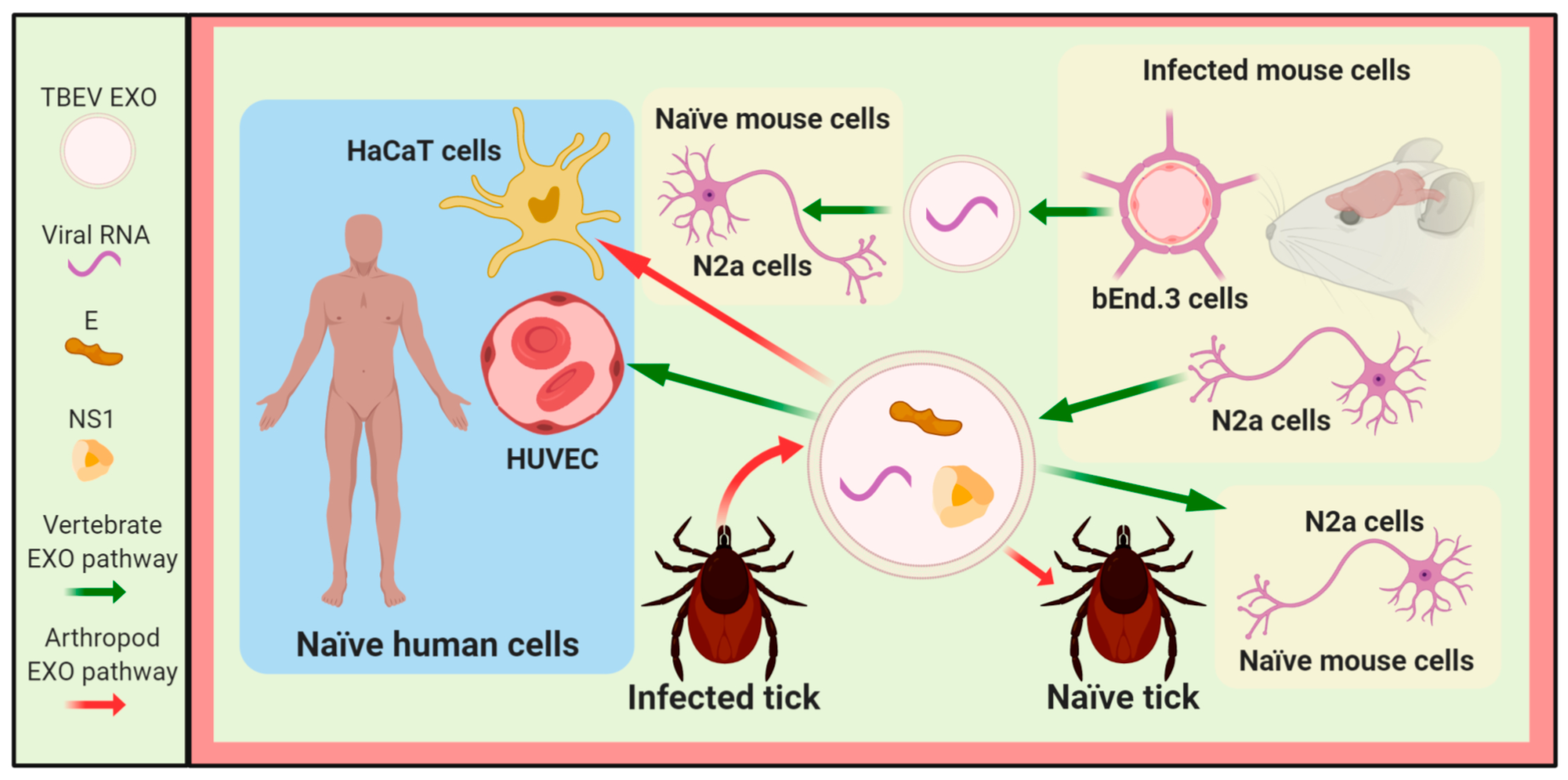
3.3. Other Flaviviruses
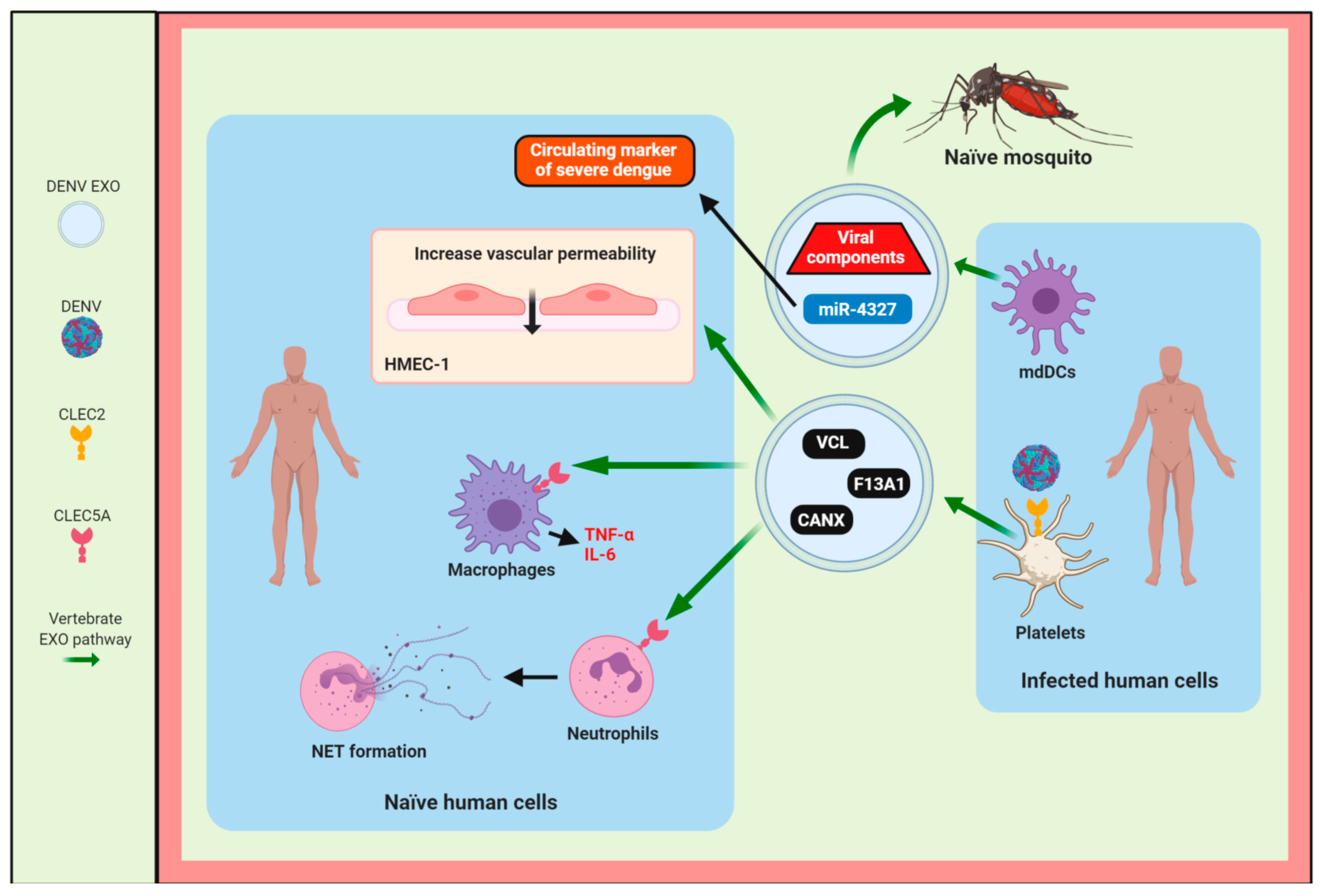
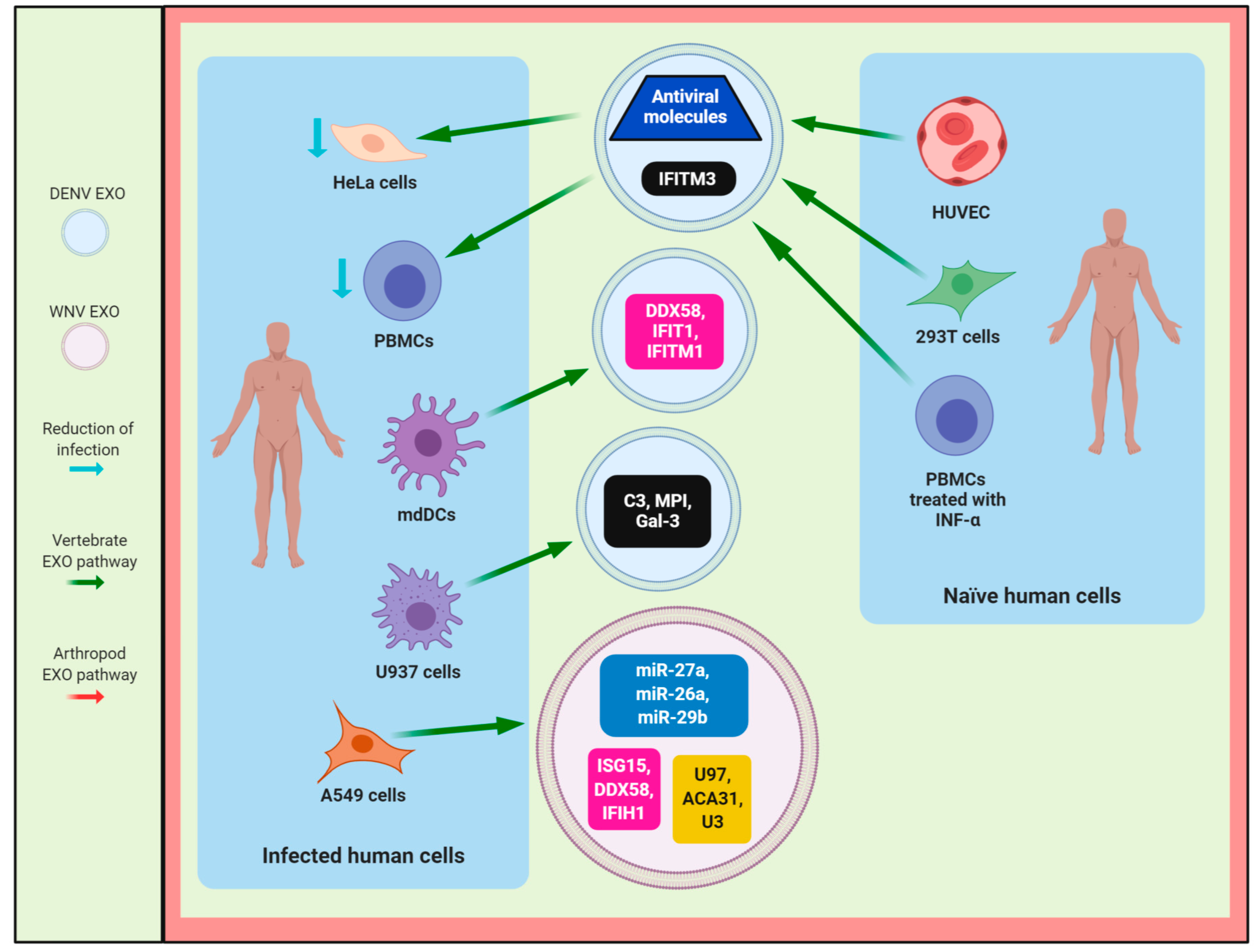
4. The Role of Exosomes in the Host Immune Response During Flavivirus Infection
4.1. DENV
4.2. ZIKV
5. Exosomes as a Transplacental Infection Route of ZIKV: A Hypothesis
6. Concluding Remarks
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Barrows, N.J.; Campos, R.K.; Liao, K.-C.; Prasanth, K.R.; Soto-Acosta, R.; Yeh, S.-C.; Schott-Lerner, G.; Pompon, J.; Sessions, O.M.; Bradrick, S.S.; et al. Biochemistry and Molecular Biology of Flaviviruses. Chem. Rev. 2018, 118, 4448–4482. [Google Scholar] [CrossRef]
- Gould, E.A.; Solomon, T. Pathogenic flaviviruses. Lancet 2008, 371, 500–509. [Google Scholar] [CrossRef]
- Holbrook, M.R. Historical Perspectives on Flavivirus Research. Viruses 2017, 9, 97. [Google Scholar] [CrossRef]
- Mandl, C.W. Steps of the tick-borne encephalitis virus replication cycle that affect neuropathogenesis. Virus Res. 2005, 111, 161–174. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Kuhn, R.J.; Rossmann, M.G. A structural perspective of the flavivirus life cycle. Nat. Rev. Microbiol. 2005, 3, 13–22. [Google Scholar] [CrossRef]
- Mayer, K.A.; Stöckl, J.; Zlabinger, G.J.; Gualdoni, G.A. Hijacking the Supplies: Metabolism as a Novel Facet of Virus-Host Interaction. Front. Immunol. 2019, 10, 1533. [Google Scholar] [CrossRef]
- Thaker, S.K.; Ch’ng, J.; Christofk, H.R. Viral hijacking of cellular metabolism. BMC Biol. 2019, 17, 59. [Google Scholar] [CrossRef]
- Martínez-Rojas, P.P.; Quiroz-García, E.; Monroy-Martínez, V.; Agredano-Moreno, L.T.; Jiménez-García, L.F.; Ruiz-Ordaz, B.H. Participation of Extracellular Vesicles from Zika-Virus-Infected Mosquito Cells in the Modification of Naïve Cells’ Behavior by Mediating Cell-to-Cell Transmission of Viral Elements. Cells 2020, 9, 123. [Google Scholar] [CrossRef]
- Martins, S.d.T.; Kuczera, D.; Lötvall, J.; Bordignon, J.; Alves, L.R. Characterization of Dendritic Cell-Derived Extracellular Vesicles During Dengue Virus Infection. Front Microbiol. 2018, 9, 1792. [Google Scholar] [CrossRef]
- Reyes-Ruiz, J.M.; Osuna-Ramos, J.F.; De Jesús-González, L.A.; Hurtado-Monzón, A.M.; Farfan-Morales, C.N.; Cervantes-Salazar, M.; Bolaños, J.; Cigarroa-Mayorga, O.E.; Martín-Martínez, E.S.; Medina, F.; et al. Isolation and characterization of exosomes released from mosquito cells infected with dengue virus. Virus Res. 2019, 266, 1–14. [Google Scholar] [CrossRef]
- Slonchak, A.; Clarke, B.; Mackenzie, J.; Amarilla, A.A.; Setoh, Y.X.; Khromykh, A.A. West Nile virus infection and interferon alpha treatment alter the spectrum and the levels of coding and noncoding host RNAs secreted in extracellular vesicles. BMC Genom. 2019, 20, 474. [Google Scholar] [CrossRef] [PubMed]
- Sung, P.-S.; Huang, T.-F.; Hsieh, S.-L. Extracellular vesicles from CLEC2-activated platelets enhance dengue virus-induced lethality via CLEC5A/TLR2. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Velandia-Romero, M.L.; Calderón-Peláez, M.A.; Balbás-Tepedino, A.; Márquez-Ortiz, R.A.; Madroñero, L.J.; Prieto, A.B.; Castellanos, J.E. Extracellular vesicles of U937 macrophage cell line infected with DENV-2 induce activation in endothelial cells EA.hy926. PLoS ONE 2020, 15, e0227030. [Google Scholar] [CrossRef]
- Vora, A.; Zhou, W.; Londono-Renteria, B.; Woodson, M.; Sherman, M.B.; Colpitts, T.M.; Neelakanta, G.; Sultana, H. Arthropod EVs mediate dengue virus transmission through interaction with a tetraspanin domain containing glycoprotein Tsp29Fb. Proc. Natl. Acad. Sci. USA 2018, 201720125. [Google Scholar] [CrossRef]
- Zhou, W.; Woodson, M.; Neupane, B.; Bai, F.; Sherman, M.B.; Choi, K.H.; Neelakanta, G.; Sultana, H. Exosomes serve as novel modes of tick-borne flavivirus transmission from arthropod to human cells and facilitates dissemination of viral RNA and proteins to the vertebrate neuronal cells. PLoS Pathog. 2018, 14, e1006764. [Google Scholar] [CrossRef]
- Zhou, W.; Woodson, M.; Sherman, M.B.; Neelakanta, G.; Sultana, H. Exosomes mediate Zika virus transmission through SMPD3 neutral Sphingomyelinase in cortical neurons. Emerg. Microbes Infect. 2019, 8, 307–326. [Google Scholar] [CrossRef]
- Margolis, L.; Sadovsky, Y. The biology of extracellular vesicles: The known unknowns. PLoS Biol. 2019, 17, e3000363. [Google Scholar] [CrossRef]
- Gheinani, A.H.; Vögeli, M.; Baumgartner, U.; Vassella, E.; Draeger, A.; Burkhard, F.C.; Monastyrskaya, K. Improved isolation strategies to increase the yield and purity of human urinary exosomes for biomarker discovery. Sci. Rep. 2018, 8, 3945. [Google Scholar] [CrossRef]
- Regev-Rudzki, N.; Wilson, D.W.; Carvalho, T.G.; Sisquella, X.; Coleman, B.M.; Rug, M.; Bursac, D.; Angrisano, F.; Gee, M.; Hill, A.F.; et al. Cell-Cell Communication between Malaria-Infected Red Blood Cells via Exosome-like Vesicles. Cell 2013, 153, 1120–1133. [Google Scholar] [CrossRef]
- Cocucci, E.; Racchetti, G.; Meldolesi, J. Shedding microvesicles: Artefacts no more. Trends Cell Biol. 2009, 19, 43–51. [Google Scholar] [CrossRef]
- Kalra, H.; Simpson, R.J.; Ji, H.; Aikawa, E.; Altevogt, P.; Askenase, P.; Bond, V.C.; Borràs, F.E.; Breakefield, X.; Budnik, V.; et al. Vesiclepedia: A Compendium for Extracellular Vesicles with Continuous Community Annotation. PLoS Biol. 2012, 10, e1001450. [Google Scholar] [CrossRef] [PubMed]
- Bukong, T.N.; Momen-Heravi, F.; Kodys, K.; Bala, S.; Szabo, G. Exosomes from hepatitis C infected patients transmit HCV infection and contain replication competent viral RNA in complex with Ago2-miR122-HSP90. PLoS Pathog. 2014, 10, e1004424. [Google Scholar] [CrossRef] [PubMed]
- Correa, R.; Caballero, Z.; De León, L.F.; Spadafora, C. Extracellular Vesicles Could Carry an Evolutionary Footprint in Interkingdom Communication. Front. Cell. Infect. Microbiol. 2020, 10, 76. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, M.; Martin-Jaular, L.; Lavieu, G.; Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 2019, 21, 9–17. [Google Scholar] [CrossRef]
- Meckes, D.G. Exosomal Communication Goes Viral. J. Virol. 2015, 89, 5200–5203. [Google Scholar] [CrossRef]
- Ramakrishnaiah, V.; Thumann, C.; Fofana, I.; Habersetzer, F.; Pan, Q.; de Ruiter, P.E.; Willemsen, R.; Demmers, J.A.A.; Stalin Raj, V.; Jenster, G.; et al. Exosome-mediated transmission of hepatitis C virus between human hepatoma Huh7.5 cells. Proc. Natl. Acad. Sci. USA 2013, 110, 13109–13113. [Google Scholar] [CrossRef]
- Skotland, T.; Sandvig, K.; Llorente, A. Lipids in exosomes: Current knowledge and the way forward. Progress Lipid Res. 2017, 66, 30–41. [Google Scholar] [CrossRef]
- Pfrieger, F.W.; Vitale, N. Cholesterol and the journey of extracellular vesicles. J. Lipid Res. 2018, 59, 2255–2261. [Google Scholar] [CrossRef] [PubMed]
- Osuna-Ramos, J.F.; Reyes-Ruiz, J.M.; del Ángel, R.M. The Role of Host Cholesterol During Flavivirus Infection. Front Cell Infect. Microbiol. 2018, 8, 388. [Google Scholar] [CrossRef]
- Zhu, X.; He, Z.; Yuan, J.; Wen, W.; Huang, X.; Hu, Y.; Lin, C.; Pan, J.; Li, R.; Deng, H.; et al. IFITM3-containing exosome as a novel mediator for anti-viral response in dengue virus infection. Cell. Microbiol. 2015, 17, 105–118. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 2006, 30, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Kowal, J.; Arras, G.; Colombo, M.; Jouve, M.; Morath, J.P.; Primdal-Bengtson, B.; Dingli, F.; Loew, D.; Tkach, M.; Théry, C. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl. Acad. Sci. USA 2016, 113, E968–E977. [Google Scholar] [CrossRef] [PubMed]
- Chiang, C.; Chen, C. Toward characterizing extracellular vesicles at a single-particle level. J. Biomed. Sci. 2019, 26, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Skliar, M.; Chernyshev, V.S. Imaging of Extracellular Vesicles by Atomic Force Microscopy. JoVE 2019, 151, e59254. [Google Scholar] [CrossRef] [PubMed]
- Smolarz, M.; Pietrowska, M.; Matysiak, N.; Mielańczyk, Ł.; Widłak, P. Proteome Profiling of Exosomes Purified from a Small Amount of Human Serum: The Problem of Co-Purified Serum Components. Proteomes 2019, 7, 18. [Google Scholar] [CrossRef]
- Turchinovich, A.; Drapkina, O.; Tonevitsky, A. Transcriptome of Extracellular Vesicles: State-of-the-Art. Front. Immunol. 2019, 10, 202. [Google Scholar] [CrossRef]
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L.; et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 2018, 20, 332–343. [Google Scholar] [CrossRef]
- Chang, W.; Wang, J. Exosomes and Their Noncoding RNA Cargo Are Emerging as New Modulators for Diabetes Mellitus. Cells 2019, 8, 853. [Google Scholar] [CrossRef]
- Garikipati, V.N.S.; Shoja-Taheri, F.; Davis, M.E. Kishore Raj Extracellular Vesicles and the Application of System Biology and Computational Modeling in Cardiac Repair. Circ. Res. 2018, 123, 188–204. [Google Scholar] [CrossRef]
- Gho, Y.S.; Lee, C. Emergent properties of extracellular vesicles: A holistic approach to decode the complexity of intercellular communication networks. Mol. BioSyst. 2017, 13, 1291–1296. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367, 6478. [Google Scholar] [CrossRef] [PubMed]
- McKelvey, K.J.; Powell, K.L.; Ashton, A.W.; Morris, J.M.; McCracken, S.A. Exosomes: Mechanisms of Uptake. J. Circ. Biomark. 2015, 4, 7. [Google Scholar] [CrossRef] [PubMed]
- Carpp, L.N.; Galler, R.; Bonaldo, M.C. Interaction between the yellow fever virus nonstructural protein NS3 and the host protein Alix contributes to the release of infectious particles. Microbes Infect. 2011, 13, 85–95. [Google Scholar] [CrossRef]
- Welsch, S.; Miller, S.; Romero-Brey, I.; Merz, A.; Bleck, C.K.E.; Walther, P.; Fuller, S.D.; Antony, C.; Krijnse-Locker, J.; Bartenschlager, R. Composition and Three-Dimensional Architecture of the Dengue Virus Replication and Assembly Sites. Cell Host Microbe 2009, 5, 365–375. [Google Scholar] [CrossRef]
- Tabata, K.; Arimoto, M.; Arakawa, M.; Nara, A.; Saito, K.; Omori, H.; Arai, A.; Ishikawa, T.; Konishi, E.; Suzuki, R.; et al. Unique Requirement for ESCRT Factors in Flavivirus Particle Formation on the Endoplasmic Reticulum. Cell Rep. 2016, 16, 2339–2347. [Google Scholar] [CrossRef] [PubMed]
- Bissig, C.; Gruenberg, J. ALIX and the multivesicular endosome: ALIX in Wonderland. Trends Cell Biol. 2014, 24, 19–25. [Google Scholar] [CrossRef]
- Larios, J.; Mercier, V.; Roux, A.; Gruenberg, J. ALIX- and ESCRT-III–dependent sorting of tetraspanins to exosomes. J. Cell Biol. 2020, 219, 3. [Google Scholar] [CrossRef] [PubMed]
- De Jesús-González, L.A.; Cervantes-Salazar, M.; Reyes-Ruiz, J.M.; Osuna-Ramos, J.F.; Farfán-Morales, C.N.; Palacios-Rápalo, S.N.; Pérez-Olais, J.H.; Cordero-Rivera, C.D.; Hurtado-Monzón, A.M.; Ruíz-Jiménez, F.; et al. The Nuclear Pore Complex: A Target for NS3 Protease of Dengue and Zika Viruses. Viruses 2020, 12, 583. [Google Scholar] [CrossRef] [PubMed]
- Desole, G.; Sinigaglia, A.; Riccetti, S.; Masi, G.; Pacenti, M.; Trevisan, M.; Barzon, L. Modelling Neurotropic Flavivirus Infection in Human Induced Pluripotent Stem Cell-Derived Systems. Int. J. Mol. Sci. 2019, 20, 5404. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Nudelman, G.; Shimoni, Y.; Kumar, M.; Ding, Y.; López, C.; Hayot, F.; Wetmur, J.G.; Sealfon, S.C. Role of Cell-to-Cell Variability in Activating a Positive Feedback Antiviral Response in Human Dendritic Cells. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Hurtado-Monzón, A.M.; Cordero-Rivera, C.D.; Farfan-Morales, C.N.; Osuna-Ramos, J.F.; De Jesús-González, L.A.; Reyes-Ruiz, J.M.; del Ángel, R.M. The role of anti-flavivirus humoral immune response in protection and pathogenesis. Rev. Med Virol. 2020, e2100. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Jordán, J.L.; Fredericksen, B.L. How Flaviviruses Activate and Suppress the Interferon Response. Viruses 2010, 2, 676–691. [Google Scholar] [CrossRef]
- Randall, R.E.; Goodbourn, S. Interferons and viruses: An interplay between induction, signalling, antiviral responses and virus countermeasures. J. Gen. Virol. 2008, 89, 1–47. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, O.; Akira, S. Innate immunity to virus infection. Immunol. Rev. 2009, 227, 75–86. [Google Scholar] [CrossRef]
- Best, S.M. The Many Faces of the Flavivirus NS5 Protein in Antagonism of Type I Interferon Signaling. J. Virol. 2017, 91, e01970-16. [Google Scholar] [CrossRef]
- Best, S.M.; Morris, K.L.; Shannon, J.G.; Robertson, S.J.; Mitzel, D.N.; Park, G.S.; Boer, E.; Wolfinbarger, J.B.; Bloom, M.E. Inhibition of interferon-stimulated JAK-STAT signaling by a tick-borne flavivirus and identification of NS5 as an interferon antagonist. J. Virol. 2005, 79, 12828–12839. [Google Scholar] [CrossRef]
- Robertson, S.J.; Mitzel, D.N.; Taylor, R.T.; Best, S.M.; Bloom, M.E. Tick-borne flaviviruses: Dissecting host immune responses and virus countermeasures. Immunol. Res. 2009, 43, 172–186. [Google Scholar] [CrossRef]
- Brass, A.L.; Huang, I.-C.; Benita, Y.; John, S.P.; Krishnan, M.N.; Feeley, E.M.; Ryan, B.J.; Weyer, J.L.; van der Weyden, L.; Fikrig, E.; et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 2009, 139, 1243–1254. [Google Scholar] [CrossRef]
- Chan, Y.K.; Huang, I.-C.; Farzan, M. IFITM Proteins Restrict Antibody-Dependent Enhancement of Dengue Virus Infection. PLoS ONE 2012, 7, e34508. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Weidner, J.M.; Qing, M.; Pan, X.-B.; Guo, H.; Xu, C.; Zhang, X.; Birk, A.; Chang, J.; Shi, P.-Y.; et al. Identification of five interferon-induced cellular proteins that inhibit west nile virus and dengue virus infections. J. Virol. 2010, 84, 8332–8341. [Google Scholar] [CrossRef] [PubMed]
- Steinman, R.M.; Cohn, Z.A. Identification of a novel cell type in peripheral lymphoid organs of mice. I. Morphology, quantitation, tissue distribution. J. Exp. Med. 1973, 137, 1142–1162. [Google Scholar] [CrossRef]
- Ho, L.J.; Wang, J.J.; Shaio, M.F.; Kao, C.L.; Chang, D.M.; Han, S.W.; Lai, J.H. Infection of human dendritic cells by dengue virus causes cell maturation and cytokine production. J. Immunol. 2001, 166, 1499–1506. [Google Scholar] [CrossRef]
- Mishra, R.; Lata, S.; Ali, A.; Banerjea, A.C. Dengue haemorrhagic fever: A job done via exosomes? Emerg. Microbes Infect. 2019, 8, 1626–1635. [Google Scholar] [CrossRef] [PubMed]
- Holley, C.L.; Topkara, V.K. An introduction to small non-coding RNAs: miRNA and snoRNA. Cardiovasc. Drugs Ther. 2011, 25, 151–159. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Q.; Gui, L.; Cai, Y.; Deng, X.; Li, C.; Guo, Q.; He, X.; Huang, J. Let-7e inhibits TNF-α expression by targeting the methyl transferase EZH2 in DENV2-infected THP-1 cells. J. Cell. Physiol. 2018, 233, 8605–8616. [Google Scholar] [CrossRef]
- Bott, A.; Erdem, N.; Lerrer, S.; Hotz-Wagenblatt, A.; Breunig, C.; Abnaof, K.; Wörner, A.; Wilhelm, H.; Münstermann, E.; Ben-Baruch, A.; et al. miRNA-1246 induces pro-inflammatory responses in mesenchymal stem/stromal cells by regulating PKA and PP2A. Oncotarget 2017, 8, 43897–43914. [Google Scholar] [CrossRef] [PubMed]
- Pham, A.M.; Langlois, R.A.; tenOever, B.R. Replication in Cells of Hematopoietic Origin Is Necessary for Dengue Virus Dissemination. PLoS Pathog. 2012, 8, e1002465. [Google Scholar] [CrossRef]
- Bavia, L.; Mosimann, A.L.P.; Aoki, M.N.; Duarte dos Santos, C.N. A glance at subgenomic flavivirus RNAs and microRNAs in flavivirus infections. Virol. J. 2016, 13, 84. [Google Scholar] [CrossRef]
- Tambyah, P.A.; Ching, C.S.; Sepramaniam, S.; Ali, J.M.; Armugam, A.; Jeyaseelan, K. microRNA expression in blood of dengue patients. Ann. Clin. Biochem. 2016, 53, 466–476. [Google Scholar] [CrossRef]
- Silveira, G.F.; Meyer, F.; Delfraro, A.; Mosimann, A.L.P.; Coluchi, N.; Vasquez, C.; Probst, C.M.; Báfica, A.; Bordignon, J.; Dos Santos, C.N.D. Dengue virus type 3 isolated from a fatal case with visceral complications induces enhanced proinflammatory responses and apoptosis of human dendritic cells. J. Virol. 2011, 85, 5374–5383. [Google Scholar] [CrossRef] [PubMed]
- McCandless, E.E.; Zhang, B.; Diamond, M.S.; Klein, R.S. CXCR4 antagonism increases T cell trafficking in the central nervous system and improves survival from West Nile virus encephalitis. Proc. Natl. Acad. Sci. USA 2008, 105, 11270–11275. [Google Scholar] [CrossRef] [PubMed]
- Arjona, A.; Foellmer, H.G.; Town, T.; Leng, L.; McDonald, C.; Wang, T.; Wong, S.J.; Montgomery, R.R.; Fikrig, E.; Bucala, R. Abrogation of macrophage migration inhibitory factor decreases West Nile virus lethality by limiting viral neuroinvasion. J. Clin. Investig. 2007, 117, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Punyadee, N.; Mairiang, D.; Thiemmeca, S.; Komoltri, C.; Pan-Ngum, W.; Chomanee, N.; Charngkaew, K.; Tangthawornchaikul, N.; Limpitikul, W.; Vasanawathana, S.; et al. Microparticles provide a novel biomarker to predict severe clinical outcomes of dengue virus infection. J. Virol. 2015, 89, 1587–1607. [Google Scholar] [CrossRef]
- Hottz, E.D.; Medeiros-de-Moraes, I.M.; Vieira-de-Abreu, A.; de Assis, E.F.; Vals-de-Souza, R.; Castro-Faria-Neto, H.C.; Weyrich, A.S.; Zimmerman, G.A.; Bozza, F.A.; Bozza, P.T. Platelet activation and apoptosis modulate monocyte inflammatory responses in dengue. J. Immunol. 2014, 193, 1864–1872. [Google Scholar] [CrossRef]
- Wu, M.-F.; Chen, S.-T.; Yang, A.-H.; Lin, W.-W.; Lin, Y.-L.; Chen, N.-J.; Tsai, I.-S.; Li, L.; Hsieh, S.-L. CLEC5A is critical for dengue virus–induced inflammasome activation in human macrophages. Blood 2013, 121, 95–106. [Google Scholar] [CrossRef]
- Wu, M.-F.; Chen, S.-T.; Hsieh, S.-L. Distinct regulation of dengue virus-induced inflammasome activation in human macrophage subsets. J. Biomed. Sci. 2013, 20, 36. [Google Scholar] [CrossRef] [PubMed]
- Gholipour, N.; Ohradanova-Repic, A.; Ahangari, G. A novel report of MiR-4301 induces cell apoptosis by negatively regulating DRD2 expression in human breast cancer cells. J. Cell. Biochem. 2018, 119, 6408–6417. [Google Scholar] [CrossRef] [PubMed]
- Komina, A.; Palkina, N.; Aksenenko, M.; Tsyrenzhapova, S.; Ruksha, T. Antiproliferative and Pro-Apoptotic Effects of MiR-4286 Inhibition in Melanoma Cells. PLoS ONE 2016, 11, e0168229. [Google Scholar] [CrossRef]
- Venkatadri, R.; Muni, T.; Iyer, A.K.V.; Yakisich, J.S.; Azad, N. Role of apoptosis-related miRNAs in resveratrol-induced breast cancer cell death. Cell Death Dis 2016, 7, e2104. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Xu, H. MicroRNA-181a-5p Regulates Inflammatory Response of Macrophages in Sepsis. Open Med. (Wars.) 2019, 14, 899–908. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Yuan, J.; Zhang, F.; Lei, Q.; Zhang, T.; Li, K.; Guo, J.; Hong, Y.; Bu, G.; Lv, X.; et al. MicroRNA-181a-5p and microRNA-181a-3p cooperatively restrict vascular inflammation and atherosclerosis. Cell Death Dis. 2019, 10, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Wang, F.-L.; Wang, H.-B.; Dong, N.; Zhu, X.-M.; Wu, Y.; Wang, Y.-T.; Yao, Y.-M. TNF-α mRNA is negatively regulated by microRNA-181a-5p in maturation of dendritic cells induced by high mobility group box-1 protein. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Burton, G.J.; Fowden, A.L. The placenta: A multifaceted, transient organ. Philos. Trans. R Soc. Lond. B Biol. Sci. 2015, 370, 20140066. [Google Scholar] [CrossRef] [PubMed]
- Jensen, O.E.; Chernyavsky, I.L. Blood Flow and Transport in the Human Placenta. Annu. Rev. Fluid Mech. 2019, 51, 25–47. [Google Scholar] [CrossRef]
- Plitman Mayo, R. Advances in Human Placental Biomechanics. Comput. Struct. Biotechnol. J. 2018, 16, 298–306. [Google Scholar] [CrossRef]
- Delorme-Axford, E.; Sadovsky, Y.; Coyne, C.B. The Placenta as a Barrier to Viral Infections. Annu. Rev. Virol. 2014, 1, 133–146. [Google Scholar] [CrossRef]
- Robbins, J.R.; Bakardjiev, A.I. Pathogens and the placental fortress. Curr. Opin. Microbiol. 2012, 15, 36–43. [Google Scholar] [CrossRef]
- Zeldovich, V.B.; Bakardjiev, A.I. Host Defense and Tolerance: Unique Challenges in the Placenta. PLoS Pathog. 2012, 8, e1002804. [Google Scholar] [CrossRef]
- Costa, H.E.; Gouilly, J.; Mansuy, J.-M.; Chen, Q.; Levy, C.; Cartron, G.; Veas, F.; Al-Daccak, R.; Izopet, J.; Jabrane-Ferrat, N. ZIKA virus reveals broad tissue and cell tropism during the first trimester of pregnancy. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- de Noronha, L.; Zanluca, C.; Burger, M.; Suzukawa, A.A.; Azevedo, M.; Rebutini, P.Z.; Novadzki, I.M.; Tanabe, L.S.; Presibella, M.M.; Duarte dos Santos, C.N. Zika Virus Infection at Different Pregnancy Stages: Anatomopathological Findings, Target Cells and Viral Persistence in Placental Tissues. Front. Microbiol. 2018, 9, 2266. [Google Scholar] [CrossRef]
- Rabelo, K.; Souza, L.J.; Salomão, N.G.; Oliveira, E.R.A.; de Paula Sentinelli, L.; Lacerda, M.S.; Saraquino, P.B.; Rosman, F.C.; Basílio-de-Oliveira, R.; Carvalho, J.J.; et al. Placental Inflammation and Fetal Injury in a Rare Zika Case Associated With Guillain-Barré Syndrome and Abortion. Front. Microbiol. 2018, 9, 1018. [Google Scholar] [CrossRef]
- Aagaard, K.M.; Lahon, A.; Suter, M.A.; Arya, R.P.; Seferovic, M.D.; Vogt, M.B.; Hu, M.; Stossi, F.; Mancini, M.A.; Harris, R.A.; et al. Primary Human Placental Trophoblasts are Permissive for Zika Virus (ZIKV) Replication. Sci. Rep. 2017, 7, 1–14. [Google Scholar] [CrossRef]
- Miner, J.J.; Cao, B.; Govero, J.; Smith, A.M.; Fernandez, E.; Cabrera, O.H.; Garber, C.; Noll, M.; Klein, R.S.; Noguchi, K.K.; et al. Zika Virus Infection during Pregnancy in Mice Causes Placental Damage and Fetal Demise. Cell 2016, 165, 1081–1091. [Google Scholar] [CrossRef]
- Tan, L.; Lacko, L.A.; Zhou, T.; Tomoiaga, D.; Hurtado, R.; Zhang, T.; Sevilla, A.; Zhong, A.; Mason, C.E.; Noggle, S.; et al. Pre- and peri-implantation Zika virus infection impairs fetal development by targeting trophectoderm cells. Nat. Commun. 2019, 10, 1–12. [Google Scholar] [CrossRef]
- Mlakar, J.; Korva, M.; Tul, N.; Popović, M.; Poljšak-Prijatelj, M.; Mraz, J.; Kolenc, M.; Resman Rus, K.; Vesnaver Vipotnik, T.; Fabjan Vodušek, V.; et al. Zika Virus Associated with Microcephaly. N. Engl. J. Med. 2016, 374, 951–958. [Google Scholar] [CrossRef] [PubMed]
- Tabata, T.; Petitt, M.; Puerta-Guardo, H.; Michlmayr, D.; Harris, E.; Pereira, L. Zika Virus Replicates in Proliferating Cells in Explants from First-Trimester Human Placentas, Potential Sites for Dissemination of Infection. J. Infect. Dis. 2018, 217, 1202–1213. [Google Scholar] [CrossRef]
- Delorme-Axford, E.; Bayer, A.; Sadovsky, Y.; Coyne, C.B. Autophagy as a mechanism of antiviral defense at the maternal–fetal interface. Autophagy 2013, 9, 2173–2174. [Google Scholar] [CrossRef] [PubMed]
- Chiramel, A.I.; Best, S.M. Role of autophagy in Zika virus infection and pathogenesis. Virus Res. 2018, 254, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Liang, Q.; Luo, Z.; Zeng, J.; Chen, W.; Foo, S.-S.; Lee, S.-A.; Ge, J.; Wang, S.; Goldman, S.A.; Zlokovic, B.V.; et al. Zika Virus NS4A and NS4B Proteins Deregulate Akt-mTOR Signaling in Human Fetal Neural Stem Cells to Inhibit Neurogenesis and Induce Autophagy. Cell Stem. Cell 2016, 19, 663–671. [Google Scholar] [CrossRef]
- Peng, H.; Liu, B.; Yves, T.D.; He, Y.; Wang, S.; Tang, H.; Ren, H.; Zhao, P.; Qi, Z.; Qin, Z. Zika Virus Induces Autophagy in Human Umbilical Vein Endothelial Cells. Viruses 2018, 10, 259. [Google Scholar] [CrossRef] [PubMed]
- Baixauli, F.; López-Otín, C.; Mittelbrunn, M. Exosomes and Autophagy: Coordinated Mechanisms for the Maintenance of Cellular Fitness. Front. Immunol. 2014, 5, 403. [Google Scholar] [CrossRef] [PubMed]
- Fader, C.M.; Colombo, M.I. Autophagy and multivesicular bodies: Two closely related partners. Cell Death Differ. 2009, 16, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Camfield, R.; Gorski, S.M. The interplay between exosomes and autophagy—partners in crime. J. Cell Sci. 2018, 131, jcs215210. [Google Scholar] [CrossRef]
- Cao, B.; Parnell, L.A.; Diamond, M.S.; Mysorekar, I.U. Inhibition of autophagy limits vertical transmission of Zika virus in pregnant mice. J. Exp. Med. 2017, 214, 2303–2313. [Google Scholar] [CrossRef]
- Gratton, R.; Agrelli, A.; Tricarico, P.M.; Brandão, L.; Crovella, S. Autophagy in Zika Virus Infection: A Possible Therapeutic Target to Counteract Viral Replication. Int. J. Mol. Sci. 2019, 20, 1048. [Google Scholar] [CrossRef]
- Zhang, S.; Yi, C.; Li, C.; Zhang, F.; Peng, J.; Wang, Q.; Liu, X.; Ye, X.; Li, P.; Wu, M.; et al. Chloroquine inhibits endosomal viral RNA release and autophagy-dependent viral replication and effectively prevents maternal to fetal transmission of Zika virus. Antivir. Res. 2019, 169, 104547. [Google Scholar] [CrossRef]
| Flavivirus | Exosomes | Size (nm) | Reference | ||||
|---|---|---|---|---|---|---|---|
| Isolation Methods | Analysis Methods | Arthropod Vector | Vertebrate Host | Uninfected | Infected | ||
| ZIKV | IP (CD63 positive) | NTA | C6/36 cells | 268.9 ± 8.20 107.8 ± 3.10 | 319.3 ± 11.50 125.5 ± 1.60 | [8] | |
| ZIKV | DG | Cryo-EM | Primary culture of C57BL/6 mice cortical neurons | 30-350 [50-200 (↑), 200-350 (↓)] | 30-350 [50-150 (↑), 150-350 (↓)] | [16] | |
| DENV2 | IP (CD9 positive) | TEM | C6/36 cells | 46.77 ± 2.98 | 81.18 ± 4.70 | [10] | |
| DENV2 | IP (CD9 positive) | AFM | C6/36 cells | 55.91 ± 2.71 | 95.07 ± 13.34 | [10] | |
| DENV2 | IP (CD9 positive) | DLS | C6/36 cells | 42.77 ± 2.29 | 97.19 ± 10.50 | [10] | |
| DENV2 | DG | Cryo-EM | C6/36 cells | 30-250 [50-100 (↑), 150-200 (↓)] | 30-250 [50-100 (↑), 100-150 (↓)] | [14] | |
| DENV2 | UC | MET | U937 cells | N/A | ~100 | [13] | |
| DENV2 | UC | NTA and TEM | Human Platelets | 50-150 | 50-150 | [12] | |
| DENV3 | UC | MET | mdDCs | 30-180 | 30-180 | [9] | |
| DENV3 | UC | NTA | mdDCs | <200 [100 (↑)] | <200 [100 (↑)] | [9] | |
| WNV | EP Kit | TEM | Lung cancer A549 cells | ~100 | ~100 | [11] | |
| LGTV (TBEV) | DG | Cryo-EM | Ixodes scapularis ISEG tick cells | 30-250 [50-100 (↑), 150-250 (↓)] | 30-250 [50-100 (↑), 100-250 (↓)] | [15] | |
| LGTV (TBEV) | DG | Cryo-EM | N2a neuronal cells | 30-250 [50-100 (↑), 150-250 (↓)] | 30-250 [50-100 (↑), 150-250 (↓)] | [15] | |
| LGTV (TBEV) | EP Kit | Cryo-EM | N2a neuronal cells | 30-200 | 30-200 | [15] | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reyes-Ruiz, J.M.; Osuna-Ramos, J.F.; De Jesús-González, L.A.; Palacios-Rápalo, S.N.; Cordero-Rivera, C.D.; Farfan-Morales, C.N.; Hurtado-Monzón, A.M.; Gallardo-Flores, C.E.; Alcaraz-Estrada, S.L.; Salas-Benito, J.S.; et al. The Regulation of Flavivirus Infection by Hijacking Exosome-Mediated Cell–Cell Communication: New Insights on Virus–Host Interactions. Viruses 2020, 12, 765. https://doi.org/10.3390/v12070765
Reyes-Ruiz JM, Osuna-Ramos JF, De Jesús-González LA, Palacios-Rápalo SN, Cordero-Rivera CD, Farfan-Morales CN, Hurtado-Monzón AM, Gallardo-Flores CE, Alcaraz-Estrada SL, Salas-Benito JS, et al. The Regulation of Flavivirus Infection by Hijacking Exosome-Mediated Cell–Cell Communication: New Insights on Virus–Host Interactions. Viruses. 2020; 12(7):765. https://doi.org/10.3390/v12070765
Chicago/Turabian StyleReyes-Ruiz, José Manuel, Juan Fidel Osuna-Ramos, Luis Adrián De Jesús-González, Selvin Noé Palacios-Rápalo, Carlos Daniel Cordero-Rivera, Carlos Noe Farfan-Morales, Arianna Mahely Hurtado-Monzón, Carla Elizabeth Gallardo-Flores, Sofía L. Alcaraz-Estrada, Juan Santiago Salas-Benito, and et al. 2020. "The Regulation of Flavivirus Infection by Hijacking Exosome-Mediated Cell–Cell Communication: New Insights on Virus–Host Interactions" Viruses 12, no. 7: 765. https://doi.org/10.3390/v12070765
APA StyleReyes-Ruiz, J. M., Osuna-Ramos, J. F., De Jesús-González, L. A., Palacios-Rápalo, S. N., Cordero-Rivera, C. D., Farfan-Morales, C. N., Hurtado-Monzón, A. M., Gallardo-Flores, C. E., Alcaraz-Estrada, S. L., Salas-Benito, J. S., & del Ángel, R. M. (2020). The Regulation of Flavivirus Infection by Hijacking Exosome-Mediated Cell–Cell Communication: New Insights on Virus–Host Interactions. Viruses, 12(7), 765. https://doi.org/10.3390/v12070765







