The Evolution, Spread and Global Threat of H6Nx Avian Influenza Viruses
Abstract
1. Introduction
2. History and Emergence of H6Nx Viruses
2.1. History of H6Nx Isolation and Detection
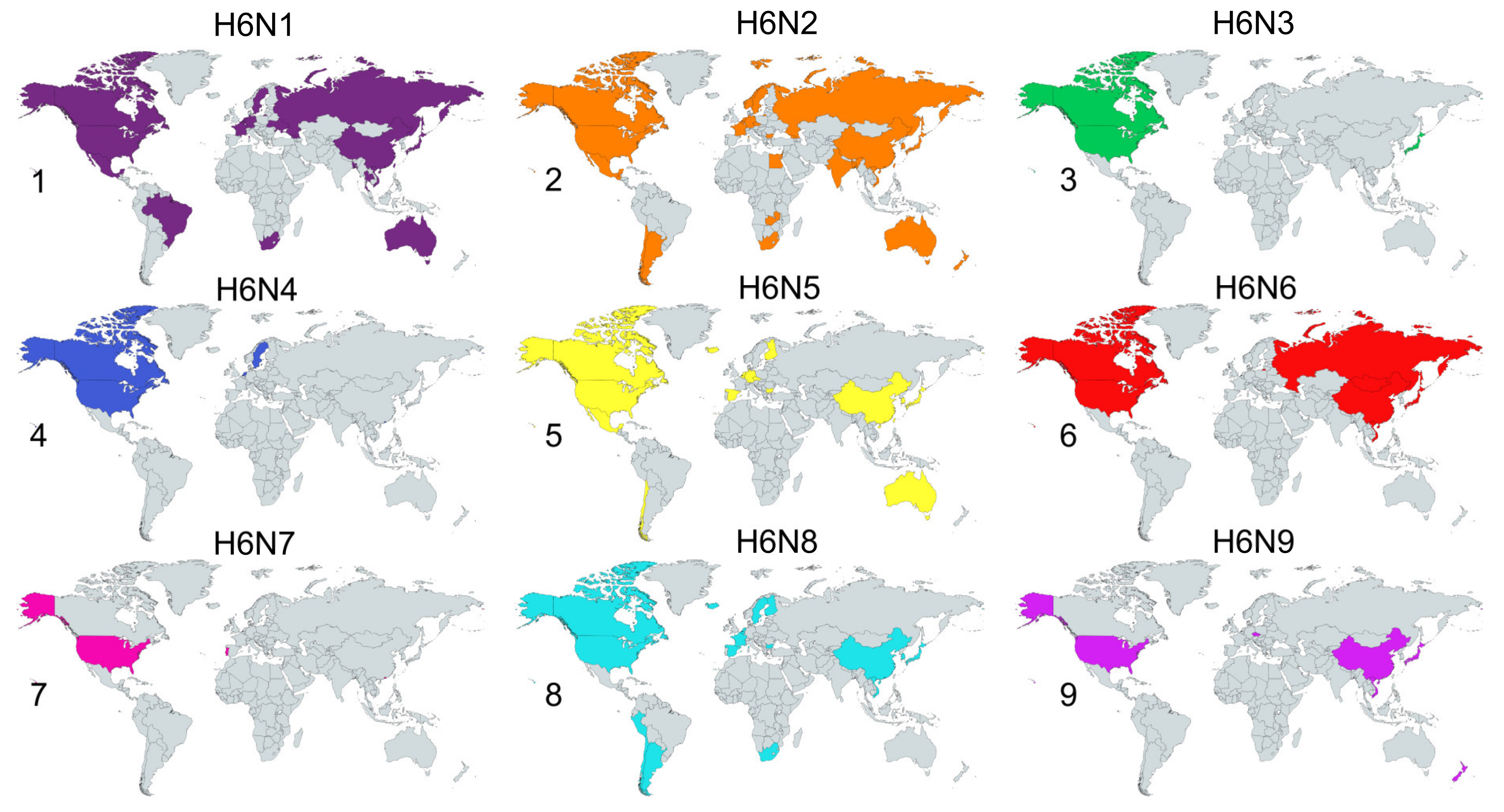
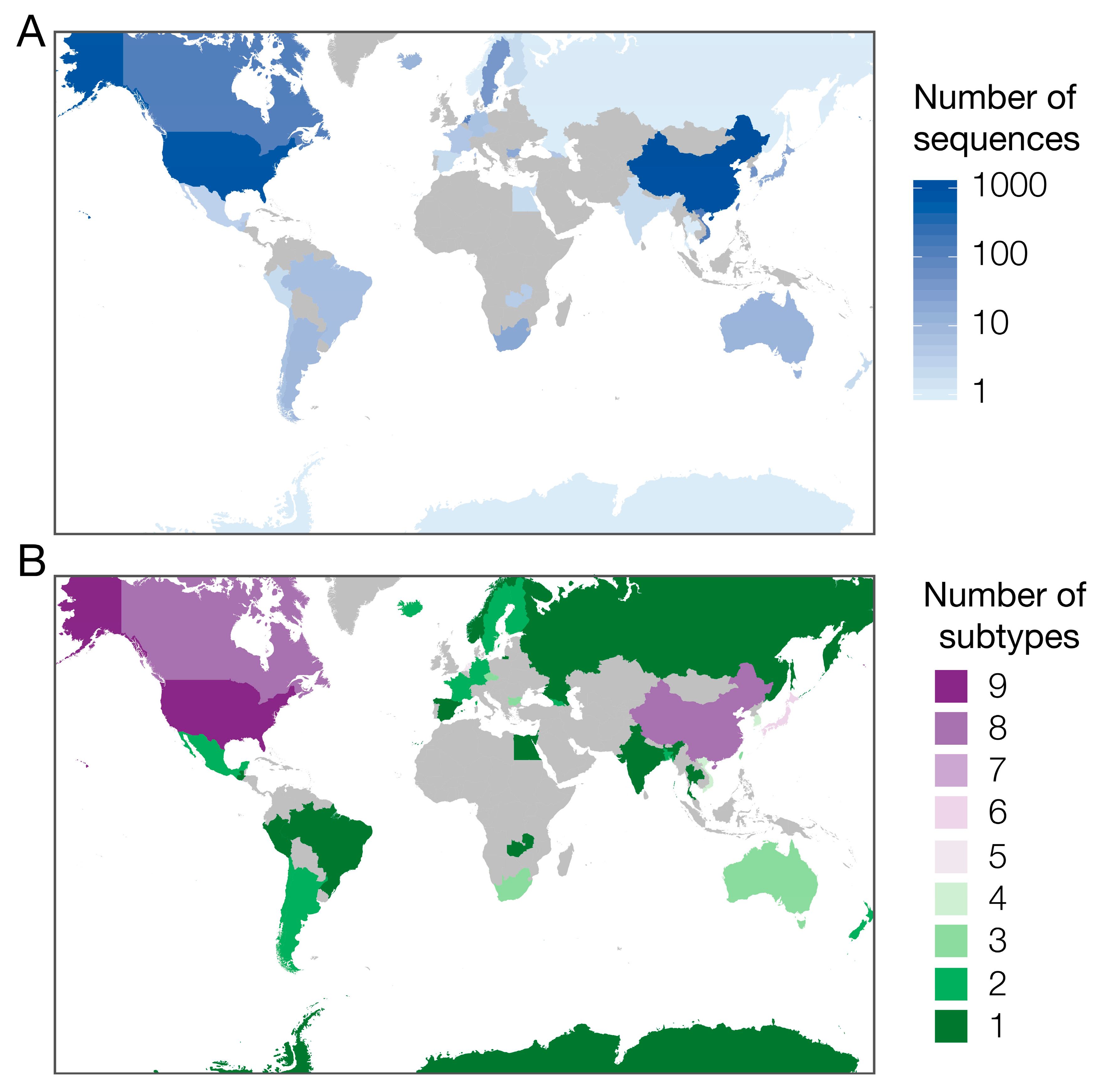
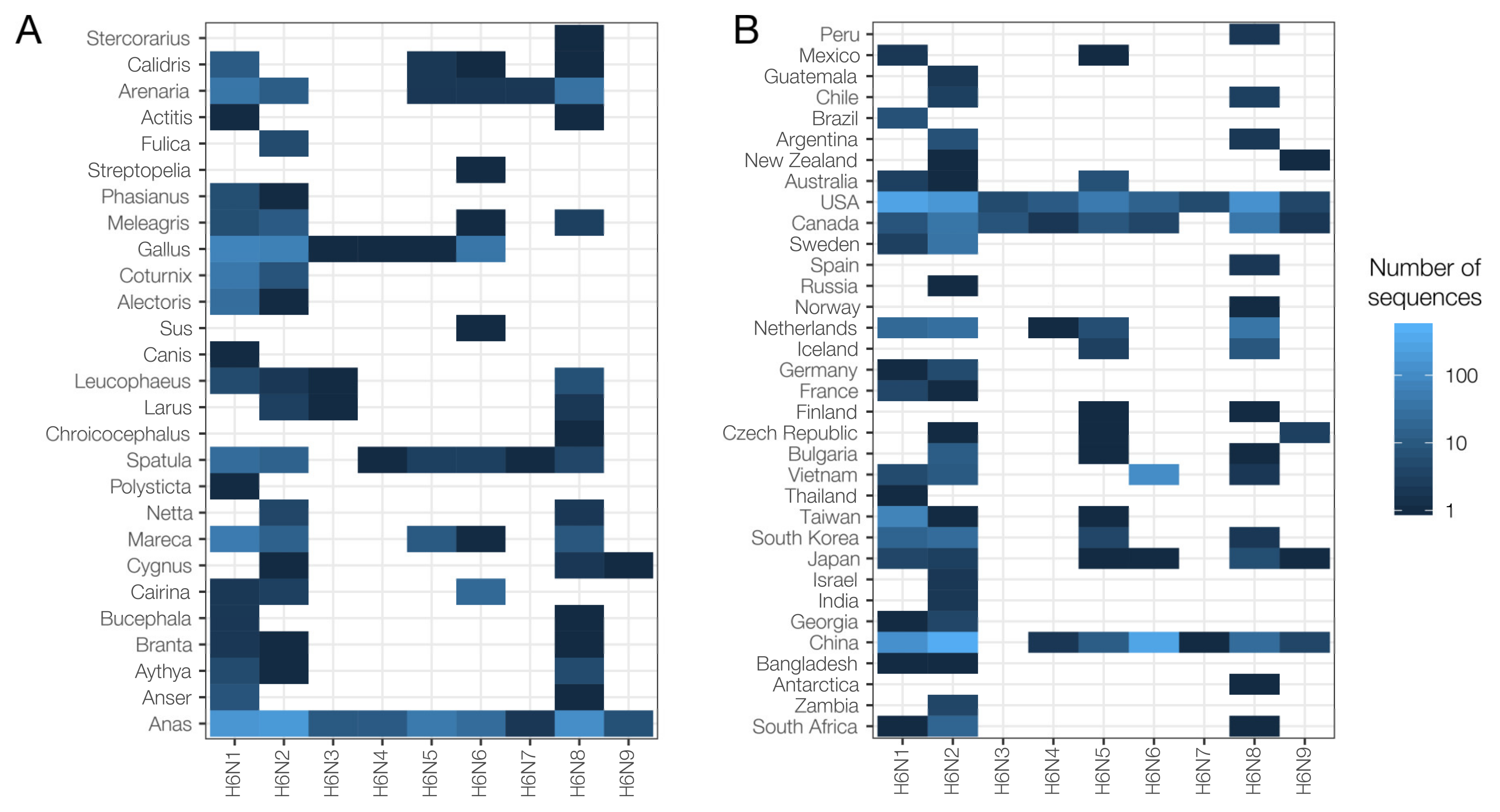
2.2. Species and Geographical Distribution of H6Nx Viruses
2.3. Global Surveillance of H6Nx Viruses
3. Genomic Sequence Availability, Phylogenetics and Nomenclature
3.1. Availability of Sequence Data
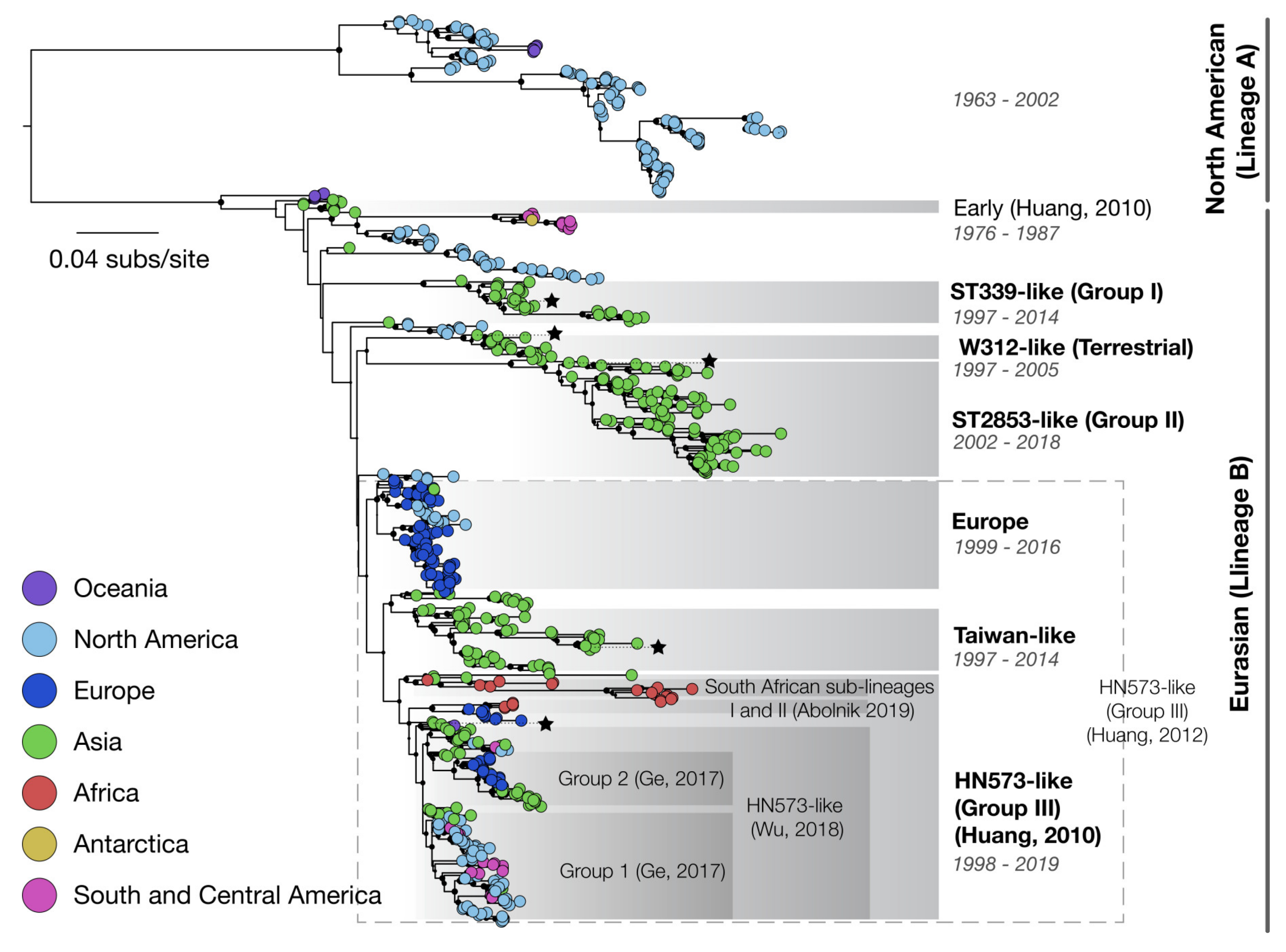
3.2. Phylogenetics
3.3. Nomenclature
3.4. Molecular Epidemiology and Virus Evolution
4. H6Nx Viruses in Avian Species
4.1. Domestic Poultry Infections with H6Nx Viruses
4.2. H6Nx Infections in Avian Species in East and Southeast Asia
4.2.1. Domestic Poultry
4.2.2. Wild Aquatic Birds
4.3. H6Nx Infections in Avian Species in Africa
4.3.1. Domestic Poultry
4.3.2. Wild Aquatic Birds
4.4. H6Nx Infections in Avian Species in the UK and Europe
4.5. H6Nx Infections in Avian Species in the Americas
4.5.1. Domestic Poultry
4.5.2. Wild Aquatic Birds
5. H6Nx Viruses in Mammalian Species and Zoonotic Infection
5.1. Human Infections with H6Nx Viruses
5.2. H6Nx Studies in Laboratory Animals
5.3. Canine Infection with H6Nx Viruses
5.4. H6Nx Virus Infections in Swine
6. Adaptation and Mutations in H6Nx Viruses
6.1. Mammalian Adaptation of H6Nx Viruses
6.1.1. Receptor Preference
6.1.2. HA Mutations
6.1.3. NA Mutations
6.1.4. Internal Gene Mutations
6.2. Poultry Adaptation of H6Nx Viruses
7. Vaccination and Control
8. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wright, P.F.; Neumann, G.; Kawaoka, Y. Orthomyxoviruses. Fields Virol. 2013, 1186–1243. [Google Scholar]
- Arbeitskreis Blut, U.B.B.K. Influenza Virus. Transfus. Med. Hemother. 2009, 36, 32–39. [Google Scholar] [CrossRef]
- Zohari, S.; Gyarmati, P.; Ejdersund, A.; Berglöf, U.; Thorén, P.; Ehrenberg, M.; Czifra, G.; Belák, S.; Waldenström, J.; Olsen, B.; et al. Phylogenetic analysis of the non-structural (NS) gene of influenza A viruses isolated from mallards in Northern Europe in 2005. Virol. J. 2008, 5, 147. [Google Scholar] [CrossRef] [PubMed]
- Zanaty, A.M.; Erfan, A.M.; Mady, W.H.; Amer, F.; Nour, A.A.; Rabie, N.; Samy, M.; Selim, A.A.; Hassan, W.M.M.; Naguib, M.M. Avian influenza virus surveillance in migratory birds in Egypt revealed a novel reassortant H6N2 subtype. Avian Res. 2019, 10. [Google Scholar] [CrossRef]
- Hill, S.C.; Hansen, R.; Watson, S.; Coward, V.; Russell, C.; Cooper, J.; Essen, S.; Everest, H.; Parag, K.V.; Fiddaman, S.; et al. Comparative micro-epidemiology of pathogenic avian influenza virus outbreaks in a wild bird population. Philos. Trans. R. Soc. B Biol. Sci. 2019, 374, 20180259. [Google Scholar] [CrossRef]
- Wu, Y.; Wu, Y.; Tefsen, B.; Shi, Y.; Gao, G.F. Bat-derived influenza-like viruses H17N10 and H18N11. Trends Microbiol. 2014, 22, 183–191. [Google Scholar] [CrossRef]
- Capua, I.; Mutinelli, F.; Pozza, M.D.; Donatelli, I.; Puzelli, S.; Cancellotti, F.M. The 1999–2000 avian influenza (H7N1) epidemic in Italy: Veterinary and human health implications. Acta Trop. 2002, 83, 7–11. [Google Scholar] [CrossRef]
- Berhane, Y.; Hisanaga, T.; Kehler, H.; Neufeld, J.; Manning, L.; Argue, C.; Handel, K.; Hooper-McGrevy, K.; Jonas, M.; Robinson, J.; et al. Highly pathogenic avian influenza virus A (H7N3) in domestic poultry, Saskatchewan, Canada, 2007. Emerg. Infect. Dis. 2009, 15, 1492–1495. [Google Scholar] [CrossRef]
- Fouchier, R.A.; Schneeberger, P.M.; Rozendaal, F.W.; Broekman, J.M.; Kemink, S.A.; Munster, V.; Kuiken, T.; Rimmelzwaan, G.F.; Schutten, M.; Van Doornum, G.J.; et al. Avian influenza A virus (H7N7) associated with human conjunctivitis and a fatal case of acute respiratory distress syndrome. Proc. Natl. Acad. Sci. USA 2004, 101, 1356–1361. [Google Scholar] [CrossRef] [PubMed]
- Franҫa, M.S.; Brown, J.D. Influenza Pathobiology and Pathogenesis in Avian Species; Springer International Publishing: Cham, Switzerland, 2014; pp. 221–242. [Google Scholar]
- Post, J.; de Geus, E.D.; Vervelde, L.; Cornelissen, J.B.W.J.; Rebel, J.M.J. Systemic distribution of different low pathogenic avian influenza (LPAI) viruses in chicken. Virol. J. 2013, 10, 23. [Google Scholar] [CrossRef] [PubMed]
- OIE. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals (Mammals, Birds and Bees), Chapter 3.3.4 Avian Influenza (Infection with Avian Influenza Viruses), Terrestrial Manual, 8th ed.; World Organisation for Animal Health: Paris, France, 2018; Volumes 1–3. [Google Scholar]
- Baron, J.; Tarnow, C.; Mayoli-Nussle, D.; Schilling, E.; Meyer, D.; Hammami, M.; Schwalm, F.; Steinmetzer, T.; Guan, Y.; Garten, W.; et al. Matriptase, HAT, and TMPRSS2 Activate the Hemagglutinin of H9N2 Influenza A Viruses. J. Virol. 2013, 87, 1811–1820. [Google Scholar] [CrossRef]
- Foster, J.E. Viruses as Pathogens: Animal Viruses Affecting Wild and Domesticated Species. In Viruses; Tennant, P., Fermin, G., Foster, J.E., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 189–216. [Google Scholar]
- Authority, E.F.S.; Gonzales, J.L.; Roberts, H.; Smietanka, K.; Baldinelli, F.; Ortiz-Pelaez, A.; Verdonck, F. Assessment of low pathogenic avian influenza virus transmission via raw poultry meat and raw table eggs. Efsa J. 2018, 16, e05431. [Google Scholar] [CrossRef]
- Horman, W.S.J.; Nguyen, T.H.O.; Kedzierska, K.; Bean, A.G.D.; Layton, D.S. The Drivers of Pathology in Zoonotic Avian Influenza: The Interplay Between Host and Pathogen. Front. Immunol. 2018, 9, 1812. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Quan, C.; Xie, Y.; Ke, C.; Nie, Y.; Chen, Q.; Hu, T.; Chen, J.; Wong, G.; Wang, Q.; et al. Continued reassortment of avian H6 influenza viruses from Southern China, 2014–2016. Transbound. Emerg. Dis. 2019, 66, 592–598. [Google Scholar] [CrossRef] [PubMed]
- Munster, V.J.; Baas, C.; Lexmond, P.; Waldenström, J.; Wallensten, A.; Fransson, T.; Rimmelzwaan, G.F.; Beyer, W.E.P.; Schutten, M.; Olsen, B.; et al. Spatial, Temporal, and Species Variation in Prevalence of Influenza A Viruses in Wild Migratory Birds. PLoS Pathog. 2007, 3, e61. [Google Scholar] [CrossRef]
- Lang, G. A Review of Influenza in Canadian Domestic and Wild Birds. Avian Dis. 2003, 47, 21–27. [Google Scholar]
- Wang, G.; Deng, G.; Shi, J.; Luo, W.; Zhang, G.; Zhang, Q.; Liu, L.; Jiang, Y.; Li, C.; Sriwilaijaroen, N.; et al. H6 influenza viruses pose a potential threat to human health. J. Virol. 2014, 88, 3953–3964. [Google Scholar] [CrossRef]
- Ge, Y.; Chai, H.; Fan, Z.; Wang, X.; Yao, Q.; Ma, J.; Chen, S.; Hua, Y.; Deng, G.; Chen, H. New H6 influenza virus reassortment strains isolated from Anser fabalis in Anhui Province, China. Virol. J. 2017, 14. [Google Scholar] [CrossRef]
- Robinson, T.; Pozzi, F. Mapping supply and demand for animal-source foods to 2030. Anim. Prod. Health Work. Pap. 2011, 2, 1–154. [Google Scholar]
- Huang, K.; Bahl, J.; Fan, X.H.; Vijaykrishna, D.; Cheung, C.L.; Webby, R.J.; Webster, R.G.; Chen, H.; Smith, G.J.D.; Peiris, J.S.M.; et al. Establishment of an H6N2 Influenza Virus Lineage in Domestic Ducks in Southern China. J. Virol. 2010, 84, 6978–6986. [Google Scholar] [CrossRef]
- Huang, K.; Zhu, H.; Fan, X.; Wang, J.; Cheung, C.L.; Duan, L.; Hong, W.; Liu, Y.; Li, L.; Smith, D.K.; et al. Establishment and Lineage Replacement of H6 Influenza Viruses in Domestic Ducks in Southern China. J. Virol. 2012, 86, 6075–6083. [Google Scholar] [CrossRef] [PubMed]
- Scotch, M.; Lam, T.T.Y.; Pabilonia, K.L.; Anderson, T.; Baroch, J.; Kohler, D.; DeLiberto, T.J. Diffusion of influenza viruses among migratory birds with a focus on the Southwest United States. Infect. Genet. Evol. 2014, 26, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Hatchette, T.F.; Walker, D.; Johnson, C.; Baker, A.; Pryor, S.P.; Webster, R.G. Influenza A viruses in feral Canadian ducks: Extensive reassortment in nature. J. Gen. Virol. 2004, 85, 2327–2337. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.I.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Posada, D.; Kozlov, A.M.; Stamatakis, A.; Morel, B.; Flouri, T. ModelTest-NG: A New and Scalable Tool for the Selection of DNA and Protein Evolutionary Models. Mol. Biol. Evol. 2019, 37, 291–294. [Google Scholar] [CrossRef]
- World Health Organization/World Organisation for Animal Health/Food; Agriculture Organization H5N1 Evolution Working Group. Revised and updated nomenclature for highly pathogenic avian influenza A (H5N1) viruses. Influenza Other Respir. Viruses 2014, 8, 384–388. [Google Scholar] [CrossRef]
- Rimondi, A.; Xu, K.; Craig, M.I.; Shao, H.; Ferreyra, H.; Rago, M.V.; Romano, M.; Uhart, M.; Sutton, T.; Ferrero, A.; et al. Phylogenetic Analysis of H6 Influenza Viruses Isolated from Rosy-Billed Pochards (Netta peposaca) in Argentina Reveals the Presence of Different HA Gene Clusters. J. Virol. 2011, 85, 13354–13362. [Google Scholar] [CrossRef]
- Wu, H.; Yang, F.; Liu, F.; Lu, R.; Peng, X.; Chen, B.; Yao, H.; Wu, N. Isolation and characterization of novel reassortant H6N1 avian influenza viruses from chickens in Eastern China. Virol. J. 2018, 15, 164. [Google Scholar] [CrossRef]
- Abolnik, C.; Bisschop, S.; Gerdes, T.; Olivier, A.; Horner, R. Outbreaks of avian influenza H6N2 viruses in chickens arose by a reassortment of H6N8 and H9N2 ostrich viruses. Virus Genes 2007, 34, 37–45. [Google Scholar] [CrossRef]
- Abolnik, C.; Strydom, C.; Rauff, D.L.; Wandrag, D.B.R.; Petty, D. Continuing evolution of H6N2 influenza a virus in South African chickens and the implications for diagnosis and control. BMC Vet. Res. 2019, 15. [Google Scholar] [CrossRef]
- Rauff, D.; Strydom, C.; Abolnik, C. Evolutionary consequences of a decade of vaccination against subtype H6N2 influenza. Virology 2016, 498, 226–239. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Pujanauski, L.M.; Davis, A.S.; Schwartzman, L.M.; Chertow, D.S.; Baxter, D.; Scherler, K.; Hartshorn, K.L.; Slemons, R.D.; Walters, K.A.; et al. Contemporary avian influenza A virus subtype H1, H6, H7, H10, and H15 hemagglutinin genes encode a mammalian virulence factor similar to the 1918 pandemic virus H1 hemagglutinin. mBio 2014, 5, e02116. [Google Scholar] [CrossRef]
- Chin, P.S.; Hoffmann, E.; Webby, R.; Webster, R.G.; Guan, Y.; Peiris, M.; Shortridge, K.F. Molecular Evolution of H6 Influenza Viruses from Poultry in Southeastern China: Prevalence of H6N1 Influenza Viruses Possessing Seven A/Hong Kong/156/97 (H5N1)-Like Genes in Poultry. J. Virol. 2002, 76, 507–516. [Google Scholar] [CrossRef] [PubMed]
- Peacock, T.P.; James, J.; Sealy, J.E.; Iqbal, M. A Global Perspective on H9N2 Avian Influenza Virus. Viruses 2019, 11, 620. [Google Scholar] [CrossRef] [PubMed]
- Ozaki, H.; Guan, Y.; Peiris, M.; Webster, R.; Webby, R. Changing patterns of h6 influenza viruses in Hong Kong poultry markets. Influenza Res. Treat. 2011, 2011, 702092. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.-H.; Okamatsu, M.; Matsuno, K.; Hiono, T.; Ogasawara, K.; Nguyen, L.T.; Van Nguyen, L.; Nguyen, T.N.; Nguyen, T.T.; Van Pham, D.; et al. Genetic and antigenic characterization of H5, H6 and H9 avian influenza viruses circulating in live bird markets with intervention in the center part of Vietnam. Vet. Microbiol. 2016, 192, 194–203. [Google Scholar] [CrossRef]
- Sealy, J.E.; Fournie, G.; Trang, P.H.; Dang, N.H.; Sadeyen, J.R.; Thanh, T.L.; Doorn, H.R.; Bryant, J.E.; Iqbal, M. Poultry trading behaviours in Vietnamese live bird markets as risk factors for avian influenza infection in chickens. Transbound. Emerg. Dis. 2019. [Google Scholar] [CrossRef]
- Kinde, H.; Read, D.H.; Daft, B.M.; Hammarlund, M.; Moore, J.; Uzal, F.; Mukai, J.; Woolcock, P. The occurrence of avian influenza A subtype H6N2 in commercial layer flocks in Southern California (2000-02): Clinicopathologic findings. Avian Dis. 2003, 47, 1214–1218. [Google Scholar] [CrossRef]
- Shortridge, K.F. Is China an influenza epicentre? Chin. Med. J. 1997, 110, 637–641. [Google Scholar]
- Shortridge, K.F.; Stuart-Harris, C.H. An influenza epicentre. Lancet 1982, 320, 812–813. [Google Scholar] [CrossRef]
- Cheung, C.L.; Vijaykrishna, D.; Smith, G.J.D.; Fan, X.H.; Zhang, J.X.; Bahl, J.; Duan, L.; Huang, K.; Tai, H.; Wang, J.; et al. Establishment of Influenza A Virus (H6N1) in Minor Poultry Species in Southern China. J. Virol. 2007, 81, 10402–10412. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.-S.; Chang, P.-C.; Shien, J.-H.; Cheng, M.-C.; Chen, C.-L.; Shieh, H.K. Genetic and pathogenic characterization of H6N1 avian influenza viruses isolated in Taiwan between 1972 and 2005. Avian Dis. 2006, 50, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Fournié, G.; Guitian, J.; Desvaux, S.; Mangtani, P.; Ly, S.; Cong, V.C.; San, S.; Dung, D.H.; Holl, D.; Pfeiffer, D.U.; et al. Identifying Live Bird Markets with the Potential to Act as Reservoirs of Avian Influenza A (H5N1) Virus: A Survey in Northern Viet Nam and Cambodia. PLoS ONE 2012, 7, e37986. [Google Scholar] [CrossRef]
- Fournié, G.; Tripodi, A.; Nguyen, T.T.T.; Nguyen, V.T.; Tran, T.T.; Bisson, A.; Pfeiffer, D.U.; Newman, S.H. Investigating poultry trade patterns to guide avian influenza surveillance and control: A case study in Vietnam. Sci. Rep. 2016, 6, 29463. [Google Scholar] [CrossRef]
- Webster, R.G. Wet markets—A continuing source of severe acute respiratory syndrome and influenza? Lancet 2004, 363, 234–236. [Google Scholar] [CrossRef]
- Bi, Y.; Chen, Q.; Wang, Q.; Chen, J.; Jin, T.; Wong, G.; Quan, C.; Liu, J.; Wu, J.; Yin, R.; et al. Genesis, Evolution and Prevalence of H5N6 Avian Influenza Viruses in China. Cell Host Microbe 2016, 20, 810–821. [Google Scholar] [CrossRef]
- Peng, C.; Sun, H.; Li, J.; Hou, G.; Wang, S.; Liu, S.; Zhuang, Q.; Cheng, S.; Chen, J.; Jiang, W. Molecular epidemiological survey and complete genomic phylogenetic analysis of H6 subtype avian influenza viruses in poultry in China from 2011 to 2016. Infect. Genet. Evol. 2018, 65, 91–95. [Google Scholar] [CrossRef]
- Abdelwhab, E.M.; Hassan, M.K.; Abdel-Moneim, A.S.; Naguib, M.M.; Mostafa, A.; Hussein, I.T.M.; Arafa, A.; Erfan, A.M.; Kilany, W.H.; Agour, M.G.; et al. Introduction and enzootic of A/H5N1 in Egypt: Virus evolution, pathogenicity and vaccine efficacy ten years on. Infect. Genet. Evol. 2016, 40, 80–90. [Google Scholar] [CrossRef]
- Simulundu, E.; Ishii, A.; Igarashi, M.; Mweene, A.S.; Suzuki, Y.; Hang Ombe, B.M.; Namangala, B.; Moonga, L.; Manzoor, R.; Ito, K.; et al. Characterization of influenza A viruses isolated from wild waterfowl in Zambia. J. Gen. Virol. 2011, 92, 1416–1427. [Google Scholar] [CrossRef]
- Corrand, L.; Delverdier, M.; Lucas, M.-N.; Croville, G.; Facon, C.; Balloy, D.; Ducatez, M.; Guérin, J.L. A low-pathogenic avian influenza H6N1 outbreak in a turkey flock in France: A comprehensive case report. Avian Pathol. 2012, 41, 569–577. [Google Scholar] [CrossRef]
- Hafez, H.; Prusas, C.; de Jackel, S.C.; Aldehoff, D.; Werner, O. Investigations on avian influenza A in meat turkey flocks in Germany. Arch. Fur. Geflugelkd. 2003, 67, 11–15. [Google Scholar]
- Landman, W.J.M.; Germeraad, E.A.; Kense, M.J. An avian influenza virus H6N1 outbreak in commercial layers: Case report and reproduction of the disease. Avian Pathol. 2019, 48, 98–110. [Google Scholar] [CrossRef] [PubMed]
- Zu Dohna, H.; Li, J.; Cardona, C.J.; Miller, J.; Carpenter, T.E. Invasions by Eurasian Avian Influenza Virus H6 Genes and Replacement of Its North American Clade. Emerg. Infect. Dis. 2009, 15, 1040–1045. [Google Scholar] [CrossRef] [PubMed]
- Benkaroun, J.; Shoham, D.; Kroyer, A.N.K.; Whitney, H.; Lang, A.S. Analysis of influenza A viruses from gulls: An evaluation of inter-regional movements and interactions with other avian and mammalian influenza A viruses. Cogent Biol. 2016, 2. [Google Scholar] [CrossRef]
- Woolcock, P.; Suarez, D.; Kuney, D. Low-pathogenicity avian influenza virus (H6N2) in chickens in California, 2000–2002. Avian Dis. 2003, 47, 872–881. [Google Scholar] [CrossRef]
- Webby, R.J.; Woolcock, P.R.; Krauss, S.L.; Webster, R.G. Reassortment and interspecies transmission of North American H6N2 influenza viruses. Virology 2002, 295, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wille, M.; Dobbin, A.; Walzthöni, N.M.; Robertson, G.J.; Ojkic, D.; Whitney, H.; Lang, A.S. Genetic Structure of Avian Influenza Viruses from Ducks of the Atlantic Flyway of North America. PLoS ONE 2014, 9, e86999. [Google Scholar] [CrossRef]
- He, W.; Mullarkey, C.E.; Duty, J.A.; Moran, T.M.; Palese, P.; Miller, M.S. Broadly Neutralizing Anti-Influenza Virus Antibodies: Enhancement of Neutralizing Potency in Polyclonal Mixtures and IgA Backbones. J. Virol. 2015, 89, 3610–3618. [Google Scholar] [CrossRef]
- Lee, C.-W.; Saif, Y.M. Avian influenza virus. Comp. Immunol. Microbiol. Infect. Dis. 2009, 32, 301–310. [Google Scholar] [CrossRef]
- Imai, M.; Watanabe, T.; Hatta, M.; Das, S.C.; Ozawa, M.; Shinya, K.; Zhong, G.; Hanson, A.; Katsura, H.; Watanabe, S.; et al. Experimental adaptation of an influenza H5 HA confers respiratory droplet transmission to a reassortant H5 HA/H1N1 virus in ferrets. Nature 2012, 486, 420–428. [Google Scholar] [CrossRef]
- Herfst, S.; Schrauwen, E.J.; Linster, M.; Chutinimitkul, S.; de Wit, E.; Munster, V.J.; Sorrell, E.M.; Bestebroer, T.M.; Burke, D.F.; Smith, D.J. Airborne transmission of influenza A/H5N1 virus between ferrets. Science 2012, 336, 1534–1541. [Google Scholar] [CrossRef] [PubMed]
- Peacock, T.P.; Benton, D.J.; Sadeyen, J.-R.; Chang, P.; Sealy, J.E.; Bryant, J.E.; Martin, S.R.; Shelton, H.; McCauley, J.W.; Barclay, W.S. Variability in H9N2 haemagglutinin receptor-binding preference and the pH of fusion. Emerg. Microbes Infect. 2017, 6, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Qi, J.; Bi, Y.; Zhang, W.; Wang, M.; Zhang, B.; Wang, M.; Liu, J.; Yan, J.; Shi, Y. Adaptation of avian influenza A (H6N1) virus from avian to human receptor-binding preference. Embo J. 2015, 34, 1661–1673. [Google Scholar] [CrossRef] [PubMed]
- Zou, S.; Gao, R.; Zhang, Y.; Li, X.; Chen, W.; Bai, T.; Dong, L.; Wang, D.; Shu, Y. Molecular characterization of H6 subtype influenza viruses in southern China from 2009 to 2011. Emerg. Microbes. Infect. 2016, 5, 1–8. [Google Scholar] [CrossRef]
- Myers, K.P.; Setterquist, S.F.; Capuano, A.W.; Gray, G.C. Infection due to 3 avian influenza subtypes in United States veterinarians. Clin. Infect. Dis. 2007, 45, 4–9. [Google Scholar] [CrossRef]
- Quan, C.; Wang, Q.; Zhang, J.; Zhao, M.; Dai, Q.; Huang, T.; Zhang, Z.; Mao, S.; Nie, Y.; Liu, J.; et al. Avian Influenza A Viruses among Occupationally Exposed Populations, China, 2014–2016. Emerg. Infect. Dis. 2019, 25, 2215–2225. [Google Scholar] [CrossRef]
- Su, S.; Qi, W.; Chen, J.; Zhu, W.; Huang, Z.; Xie, J.; Zhang, G. Seroepidemiological evidence of avian influenza A virus transmission to pigs in southern China. J. Clin. Microbiol. 2013, 51, 601–602. [Google Scholar] [CrossRef][Green Version]
- Beare, A.; Webster, R. Replication of avian influenza viruses in humans. Arch. Virol. 1991, 119, 37–42. [Google Scholar] [CrossRef]
- Hoffmann, E.; Stech, J.; Leneva, I.; Krauss, S.; Scholtissek, C.; San Chin, P.; Peiris, M.; Shortridge, K.F.; Webster, R.G. Characterization of the influenza A virus gene pool in avian species in southern China: Was H6N1 a derivative or a precursor of H5N1? J. Virol. 2000, 74, 6309–6315. [Google Scholar] [CrossRef]
- Cheng, K.; Yu, Z.; Chai, H.; Sun, W.; Xin, Y.; Zhang, Q.; Huang, J.; Zhang, K.; Li, X.; Yang, S.; et al. PB2-E627K and PA-T97I substitutions enhance polymerase activity and confer a virulent phenotype to an H6N1 avian influenza virus in mice. Virology 2014, 468, 207–213. [Google Scholar] [CrossRef]
- Ni, F.; Kondrashkina, E.; Wang, Q. Structural and Functional Studies of Influenza Virus A/H6 Hemagglutinin. PLoS ONE 2015, 10, e0134576. [Google Scholar] [CrossRef] [PubMed]
- Gillim-Ross, L.; Santos, C.; Chen, Z.; Aspelund, A.; Yang, C.F.; Ye, D.; Jin, H.; Kemble, G.; Subbarao, K. Avian Influenza H6 Viruses Productively Infect and Cause Illness in Mice and Ferrets. J. Virol. 2008, 82, 10854–10863. [Google Scholar] [CrossRef]
- Belser, J.A.; Katz, J.M.; Tumpey, T.M. The ferret as a model organism to study influenza A virus infection. Dis. Model. Mech. 2011, 4, 575–579. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, C.D.; Subbarao, K. The contribution of animal models to the understanding of the host range and virulence of influenza A viruses. Microbes Infect. 2011, 13, 502–515. [Google Scholar] [CrossRef]
- Jia, N.; Barclay, W.S.; Roberts, K.; Yen, H.L.; Chan, R.W.; Lam, A.K.; Air, G.; Peiris, J.S.; Dell, A.; Nicholls, J.M.; et al. Glycomic characterization of respiratory tract tissues of ferrets: Implications for its use in influenza virus infection studies. J. Biol. Chem. 2014, 289, 28489–28504. [Google Scholar] [CrossRef] [PubMed]
- Long, J.S.; Benfield, C.T.; Barclay, W.S. One-way trip: Influenza virus’ adaptation to gallinaceous poultry may limit its pandemic potential. BioEssays 2015, 37, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention, N.C.f.I.a.R.D.N. Public Health Threat of Highly Pathogenic Asian Avian Influenza A (H5N1) Virus. Available online: https://www.cdc.gov/flu/avianflu/h5n1-threat.htm (accessed on 1 June 2020).
- Sun, H.; Kaplan, B.S.; Guan, M.; Zhang, G.; Ye, J.; Long, L.P.; Blackmon, S.; Yang, C.-K.; Chiang, M.-J.; Xie, H.; et al. Pathogenicity and transmission of a swine influenza A(H6N6) virus. Emerg. Microbes Infect. 2017, 6, e17. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.-T.; Wang, C.-H.; Chueh, L.-L.; Su, B.L.; Wang, L.C. Influenza A(H6N1) Virus in Dogs, Taiwan. Emerg. Infect. Dis. 2015, 21, 2154–2157. [Google Scholar] [CrossRef]
- Yuan, J.; Zhang, L.; Kan, X.; Jiang, L.; Yang, J.; Guo, Z.; Ren, Q. Origin and Molecular Characteristics of a Novel 2013 Avian Influenza A(H6N1) Virus Causing Human Infection in Taiwan. Clin. Infect. Dis. 2013, 57, 1367–1368. [Google Scholar] [CrossRef]
- Tan, L.; Su, S.; Smith, D.K.; He, S.; Zheng, Y.; Shao, Z.; Ma, J.; Zhu, H.; Zhang, G. A Combination of HA and PA Mutations Enhances Virulence in a Mouse-Adapted H6N6 Influenza A Virus. J. Virol. 2014, 88, 14116–14125. [Google Scholar] [CrossRef]
- Kumar, M.; Nagarajan, S.; Murugkar, H.V.; Saikia, B.; Singh, B.; Mishra, A.; Tripathi, S.K.; Agarwal, S.; Shukla, S.; Kulkarni, D.D.; et al. Emergence of novel reassortant H6N2 avian influenza viruses in ducks in India. Infect. Genet. Evol. 2018, 61, 20–23. [Google Scholar] [CrossRef] [PubMed]
- Sealy, J.E.; Yaqub, T.; Peacock, T.P.; Chang, P.; Ermetal, B.; Clements, A.; Sadeyen, J.-R.; Mehboob, A.; Shelton, H.; Bryant, J.E. Association of increased receptor-binding avidity of influenza A (H9N2) viruses with escape from antibody-based immunity and enhanced zoonotic potential. Emerg. Infect. Dis. 2019, 25, 63. [Google Scholar] [CrossRef]
- Shelton, H.; Ayora-Talavera, G.; Ren, J.; Loureiro, S.; Pickles, R.J.; Barclay, W.S.; Jones, I.M. Receptor Binding Profiles of Avian Influenza Virus Hemagglutinin Subtypes on Human Cells as a Predictor of Pandemic Potential. J. Virol. 2011, 85, 1875–1880. [Google Scholar] [CrossRef] [PubMed]
- Tzarum, N.; de Vries, R.P.; Zhu, X.; Yu, W.; McBride, R.; Paulson, J.C.; Wilson, I.A. Structure and Receptor Binding of the Hemagglutinin from a Human H6N1 Influenza Virus. Cell Host Microbe 2015, 17, 369–376. [Google Scholar] [CrossRef]
- De Vries, R.P.; Tzarum, N.; Peng, W.; Thompson, A.J.; Wickramasinghe, I.N.A.; de la Pena, A.T.T.; van Breemen, M.J.; Bouwman, K.M.; Zhu, X.; McBride, R. A single mutation in Taiwanese H6N1 influenza hemagglutinin switches binding to human-type receptors. Embo Mol. Med. 2017, 9, 1314–1325. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.H.; Yang, J.R.; Wu, H.S.; Chang, M.C.; Lin, J.S.; Lin, C.Y.; Liu, Y.L.; Lo, Y.C.; Yang, C.H.; Chuang, J.H.; et al. Human infection with avian influenza A H6N1 virus: An epidemiological analysis. Lancet Respir. Med. 2013, 1, 771–778. [Google Scholar] [CrossRef]
- De Graaf, M.; Fouchier, R.A. Role of receptor binding specificity in influenza A virus transmission and pathogenesis. Embo J. 2014, 33, 823–841. [Google Scholar] [CrossRef] [PubMed]
- Suttie, A.; Deng, Y.-M.; Greenhill, A.R.; Dussart, P.; Horwood, P.F.; Karlsson, E.A. Inventory of molecular markers affecting biological characteristics of avian influenza A viruses. Virus Genes 2019, 55, 739–768. [Google Scholar] [CrossRef]
- Qu, Z.; Ma, S.; Kong, H.; Deng, G.; Shi, J.; Liu, L.; Suzuki, Y.; Chen, H. Identification of a key amino acid in hemagglutinin that increases human-type receptor binding and transmission of an H6N2 avian influenza virus. Microbes Infect. 2017, 19, 655–660. [Google Scholar] [CrossRef]
- Long, J.S.; Giotis, E.S.; Moncorge, O.; Frise, R.; Mistry, B.; James, J.; Morisson, M.; Iqbal, M.; Vignal, A.; Skinner, M.A.; et al. Species difference in ANP32A underlies influenza A virus polymerase host restriction. Nature 2016, 529, 101–104. [Google Scholar] [CrossRef]
- Gao, H.; Liu, J.; Kong, W.; Sun, H.; Pu, J.; Chang, K.C.; Xu, G.; Sun, Y.; Qi, L.; Wang, J. PA-X is a virulence factor in avian H9N2 influenza virus. J. Gen. Virol. 2015, 96, 2587–2594. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Mo, Y.; Wang, X.; Gu, M.; Hu, Z.; Zhong, L.; Wu, Q.; Hao, X.; Hu, S.; Liu, W.; et al. PA-X Decreases the Pathogenicity of Highly Pathogenic H5N1 Influenza A Virus in Avian Species by Inhibiting Virus Replication and Host Response. J. Virol. 2015, 89, 4126. [Google Scholar] [CrossRef] [PubMed]
- DeLuca, D.S.; Keskin, D.B.; Zhang, G.L.; Reinherz, E.L.; Brusic, V. PB1-F2 Finder: Scanning influenza sequences for PB1-F2 encoding RNA segments. BMC Bioinform. 2011, 12, S6. [Google Scholar] [CrossRef] [PubMed]
- Zell, R.; Krumbholz, A.; Eitner, A.; Krieg, R.; Halbhuber, K.-J.; Wutzler, P. Prevalence of PB1-F2 of influenza A viruses. J. Gen. Virol. 2007, 88, 536–546. [Google Scholar] [CrossRef]
- James, J.; Howard, W.; Iqbal, M.; Nair, V.; Barclay, W.; Shelton, H. Influenza A Virus PB1-F2 Protein Prolongs Viral Shedding in Chickens Lengthening the Transmission Window. J. Gen. Virol. 2016, 97. [Google Scholar] [CrossRef]
- James, J.; Smith, N.; Ross, C.; Iqbal, M.; Goodbourn, S.; Digard, P.; Barclay, W.S.; Shelton, H. The cellular localization of avian influenza virus PB1-F2 protein alters the magnitude of IFN2 promoter and NFkappaB-dependent promoter antagonism in chicken cells. J. Gen. Virol. 2019, 100, 414–430. [Google Scholar] [CrossRef]
- Finkelstein, D.B.; Mukatira, S.; Mehta, P.K.; Obenauer, J.C.; Su, X.; Webster, R.G.; Naeve, C.W. Persistent Host Markers in Pandemic and H5N1 Influenza Viruses. J. Virol. 2007, 81, 10292–10299. [Google Scholar] [CrossRef]
- Gambaryan, A.S.; Tuzikov, A.B.; Pazynina, G.V.; Desheva, J.A.; Bovin, N.V.; Matrosovich, M.N.; Klimov, A.I. 6-sulfo sialyl Lewis X is the common receptor determinant recognized by H5, H6, H7 and H9 influenza viruses of terrestrial poultry. Virol. J. 2008, 5, 85. [Google Scholar] [CrossRef]
- Kuchipudi, S.V.; Nelli, R.; White, G.A.; Bain, M.; Chang, K.C.; Dunham, S. Differences in influenza virus receptors in chickens and ducks: Implications for interspecies transmission. J. Mol. Genet. Med. Int. J. Biomed. Res. 2009, 3, 143. [Google Scholar] [CrossRef]
- Costa, T.; Chaves, A.J.; Valle, R.; Darji, A.; van Riel, D.; Kuiken, T.; Majó, N.; Ramis, A. Distribution patterns of influenza virus receptors and viral attachment patterns in the respiratory and intestinal tracts of seven avian species. Vet. Res. 2012, 43, 28. [Google Scholar] [CrossRef]
- Gambaryan, A.; Webster, R.; Matrosovich, M. Differences between influenza virus receptors on target cells of duck and chicken. Arch. Virol. 2002, 147, 1197–1208. [Google Scholar] [CrossRef] [PubMed]
- Wan, H.; Perez, D.R. Quail carry sialic acid receptors compatible with binding of avian and human influenza viruses. Virology 2006, 346, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Banks, J.; Plowright, L. Additional Glycosylation at the Receptor Binding Site of the Hemagglutinin (HA) for H5 and H7 Viruses May Be an Adaptation to Poultry Hosts, but Does It Influence Pathogenicity? Avian Dis. 2003, 47, 942–950. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Carney, P.J.; Chang, J.C.; Villanueva, J.M.; Stevens, J. Structure and receptor binding preferences of recombinant hemagglutinins from avian and human H6 and H10 influenza A virus subtypes. J. Virol. 2015, 89, 4612–4623. [Google Scholar] [CrossRef]
- Bergervoet, S.A.; Germeraad, E.A.; Alders, M.; Roose, M.M.; Engelsma, M.Y.; Heutink, R.; Bouwstra, R.; Fouchier, R.A.M.; Beerens, N. Susceptibility of Chickens to Low Pathogenic Avian Influenza (LPAI) Viruses of Wild Bird- and Poultry-Associated Subtypes. Viruses 2019, 11, 1010. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Santos, C.; Aspelund, A.; Gillim-Ross, L.; Jin, H.; Kemble, G.; Subbarao, K. Evaluation of Live Attenuated Influenza A Virus H6 Vaccines in Mice and Ferrets. J. Virol. 2009, 83, 65. [Google Scholar] [CrossRef]
- Chen, Z.; Zhou, H.; Kim, L.; Jin, H. The Receptor Binding Specificity of the Live Attenuated Influenza H2 and H6 Vaccine Viruses Contributes to Vaccine Immunogenicity and Protection in Ferrets. J. Virol. 2012, 86, 2780–2786. [Google Scholar] [CrossRef][Green Version]
| Country | Period | Subtype | Species | No. of Sequenced Strains | Total |
|---|---|---|---|---|---|
| Antarctica | 2011 | H6N8 | Brown Skua, Chinstrap Penguin | 2 | 2 |
| Argentina | 2007–2011 | H6N2 | Silver Teal, Comb Duck, Rosy-Billed Pochard, Yellow-Billed Pintail | 7 | 9 |
| 2007 | H6N8 | Rosy-Billed Pochard | 2 | ||
| Australia | 1979 | H6N1 | Pacific Duck, Gray Teal | 3 | 8 |
| 1979 | H6N2 | Eurasian Coot | 1 | ||
| 1972–1980 | H6N5 | Pacific Black Duck, Shearwater, | 4 | ||
| Bangladesh | 2015 | H6N1 | Duck (unspecified) | 1 | 2 |
| 2016 | H6N2 | Mallard | 1 | ||
| Belgium | 2008–2010 | H6N1 | European Herring Gull, Turkey | 3 | 21 |
| 2011–2016 | H6N2 | Mute Swan, Eurasian Teal, Mallard | 5 | ||
| 2009 | H6N5 | Mallard | 1 | ||
| 2010–2016 | H6N8 | Mallard, Ruddy turnstone, Tufted Duck, Coot, Duck (unspecified) | 12 | ||
| Brazil | 2012 | H6N1 | White-Rumped Sandpiper | 6 | 6 |
| Bulgaria | 2008–2010 | H6N2 | Mule Duck | 12 | 14 |
| 2010 | H6N5 | Mule Duck | 1 | ||
| 2010 | H6N8 | Mule Duck | 1 | ||
| Canada | 1981–2010 | H6N1 | Mallard, Blue Winged teal, Green-Winged Teal | 10 | 88 |
| 1978–2005 | H6N2 | Mallard, Pintail Duc, Shoveler Duck, Turkey, Blue-Winged Teal, Common Coot | 47 | ||
| 1985–1990 | H6N3 | Mallard, Pintail Duck | 8 | ||
| 1979–1989 | H6N4 | Mallard, Pintail Duck, Blue-Winged Teal | 3 | ||
| 1978–2003 | H6N5 | Mallard, Pintail Duck, Blue-Winged Teal | 12 | ||
| 1982–2010 | H6N6 | Mallard, Pintail Duck, Blue-Winged Teal, Widgeon, American Black Duck | 6 | ||
| 1978–1996 | H6N8 | Mallard, Pintail Duck, Blue-Winged Teal, Redhead, Turkey, Green-Winged Teal | 40 | ||
| Chile | 2016 | H6N2 | Franklin’s gull, Kelp Gull | 3 | 6 |
| 2016–2017 | H6N8 | Upland Goose, Mallard, Kelp Gull | 3 | ||
| China | 1997–2017 | H6N1 | Bean Goose, Chicken, Chukar, Duck (unspecified), guineafowl, Mallard, Partridge, Pheasant, Quail, Shoveler Duck, Common Teal, Turkey | 109 | 937 |
| 2000–2017 | H6N2 | Wild Bird (unspecified), Chicken, Chukar, Duck (unspecified), Environmental sample, Bean Goose, Goose (unspecified), Guineafowl, Mallard, Partridge, Pheasant, Pigeon, quail, Common Teal, Turkey | 462 | ||
| 2003–2017 | H6N5 | Duck (Unspecified), Mallard, Shoveler Duck | 11 | ||
| 2004–2016 | H6N6 | Chicken, Duck (unspecified), Environmental Sample, Goose (unspecified), Mallard, Muscovy Duck, Pigeon, Turtledove, Swine | 321 | ||
| 2000–2016 | H6N8 | Wild Bird (unspecified), Duck (unspecified), Environmental sample, Goose (unspecified), Mallard, Common Teal, Wild Duck | 32 | ||
| 2001–2007 | H6N9 | Duck (unspecified), wild Duck | 2 | ||
| Czech Republic | 2009 | H6N2 | Mallard | 3 | 8 |
| 2007 | H6N5 | Mallard | 2 | ||
| 2009–2010 | H6N9 | Mallard | 3 | ||
| Finland | 2007 | H6N5 | Mallard | 1 | 2 |
| 2007 | H6N8 | Mallard | 1 | ||
| France | 1997–2010 | H6N1 | Turkey, Muscovy Duck | 5 | 22 |
| 2000–2005 | H6N2 | Pekin duck, Turkey, Mallard | 15 | ||
| 2001 | H6N8 | Turkey | 2 | ||
| Georgia | 2015 | H6N1 | Mallard | 1 | 4 |
| 2010–2015 | H6N2 | Mallard, Common Coot, Domestic Duck | 3 | ||
| Germany | 1968–1999 | H6N1 | Turkey, Duck (unspecified) | 2 | 9 |
| 1999–2002 | H6N2 | Turkey | 6 | ||
| 1998 | H6N5 | Turkey | 1 | ||
| Guatemala | 2013 | H6N2 | Blue-Winged Teal | 2 | 2 |
| Hong Kong | 1976–2003 | H6N1 | Chicken, Chukar, Duck (unspecified), Guineafowl, Pheasant, Pigeon, Quail, Common Teal, Waterfowl | 38 | 69 |
| 1977–2002 | H6N2 | Duck (unspecified), Goose (unspecified), Guineafowl, Quail, Silky Chicken | 19 | ||
| 1977–2001 | H6N4 | Chicken, Goose (unspecified), Pheasant, Quail | 5 | ||
| 1976–1980 | H6N5 | Duck (unspecified) | 2 | ||
| 1997 | H6N7 | Goose (unspecified) | 1 | ||
| 1977 | H6N8 | Duck (unspecified) | 1 | ||
| 1977–1997 | H6N9 | Duck (unspecified), Goose (unspecified) | 3 | ||
| Iceland | 2011 | H6N5 | Greylag Goose | 3 | 12 |
| 2011 | H6N8 | Greylag Goose, Pink-footed Goose | 9 | ||
| India | 2014–2015 | H6N2 | Duck (unspecified) | 2 | 2 |
| Israel | 2001 | H6N2 | Wild Bird (unspecified) | 2 | 2 |
| Japan | 2004–2017 | H6N1 | Chicken, Duck (unspecified) | 12 | 44 |
| 2000–2017 | H6N2 | Duck (unspecified) | 15 | ||
| 1980 | H6N3 | Gull (unspecified) | 1 | ||
| 2006–2008 | H6N5 | Duck (unspecified), Wild Bird (unspecified) | 5 | ||
| 2014 | H6N6 | Wild Bird (unspecified) | 1 | ||
| 1996–2017 | H6N8 | Mallard, Wild Bird (unspecified), Whooper Swan, Duck (unspecified) | 8 | ||
| 2001–2008 | H6N9 | Duck (unspecified), Swan (unspecified) | 2 | ||
| Mexico | 2008 | H6N1 | Northern Shoveler, Green-Winged Teal | 2 | 4 |
| 2007 | H6N2 | Mallard | 1 | ||
| 2009 | H6N5 | Green-Winged Teal | 1 | ||
| Mongolia | 2011 | H6N6 | Common Teal | 1 | 1 |
| Netherlands | 1999–2012 | H6N1 | Chicken, Eurasian Teal, Eurasian Wigeon, Greater White-Fronted Goose, Greylag Goose, Mallard, Common Teal | 28 | 128 |
| 2000–2014 | H6N2 | Barnacle Goose, Bewick’s Swan, Gadwall Duck, Eurasian Wigeon, Greater White-Fronted Goose, White-Fronted Goose, Greylag Goose, Mallard, Common Teal | 36 | ||
| 2011 | H6N4 | Mallard | 2 | ||
| 1999–2015 | H6N5 | Greater white-fronted Goose, Mallard, Turkey | 9 | ||
| 2004–2014 | H6N8 | Barnacle Goose, Bewick’s Swan, Black-Headed Gull, Greater White-Fronted Goose, Bean Goose, White-Fronted Goose, Mallard, Common Teal | 53 | ||
| New Zealand | 2005 | H6N2 | Mallard | 1 | 2 |
| 2005 | H6N9 | Mallard | 1 | ||
| Norway | 2005–2010 | H6N2 | Mallard, Teal (unspecified) | 13 | 14 |
| 2009 | H6N8 | Mew Gull | 1 | ||
| Peru | 2010 | H6N8 | Egret, Sandpiper | 2 | 2 |
| Portugal | 2006 | H6N7 | 1 | 1 | |
| Russia | 2006–2009 | H6N1 | Duck (unspecified), Teal (unspecified), Herring Gull | 6 | 9 |
| 2006–2011 | H6N2 | Mallard, Gull (unspecified) | 2 | ||
| 2009 | H6N6 | Teal (unspecified) | 1 | ||
| South Africa | 2009 | H6N1 | Ostrich | 1 | 21 |
| 2002–2019 | H6N2 | Chicken | 18 | ||
| 1998–2007 | H6N8 | Ostrich | 2 | ||
| South Korea | 2003–2017 | H6N1 | Wild Bird (unspecified), Duck (unspecified), Spot-Billed Duck, White Pekin Duck, Bean Goose, Mallard, Shorebird (unspecified) | 19 | 64 |
| 2002–2017 | H6N2 | Wild Bird (unspecified), Duck (unspecified), Spot-Billed Duck, White Pekin Duck, White-Fronted Goose, Mallard | 35 | ||
| 2005–2018 | H6N5 | Wild Bird (unspecified), Duck (unspecified), Bean Goose, Mallard | 5 | ||
| 2005–2014 | H6N8 | Wild Bird (unspecified), Spot-Billed Duck, Mallard | 5 | ||
| Spain | 2007 | H6N5 | Mallard | 1 | 3 |
| 2006 | H6N8 | Duck (unspecified) | 2 | ||
| Sweden | 2002–2005 | H6N1 | Mallard | 4 | 48 |
| 2000–2009 | H6N2 | Guillemot, Mallard, Eurasian Wigeon | 38 | ||
| 2014 | H6N4 | Mallard | 1 | ||
| 2005 | H6N8 | Mallard | 5 | ||
| Taiwan | 1972–2014 | H6N1 | Chicken, Dog, Duck (unspecified), Partridge, Human | 78 | 82 |
| 2004 | H6N2 | Duck (unspecified) | 1 | ||
| 1989–2004 | H6N5 | Chicken, Shorebird (unspecified) | 3 | ||
| Thailand | 2005 | H6N1 | Duck (unspecified) | 1 | 1 |
| Ukraine | 2010 | H6N1 | White-Fronted Goose | 1 | 1 |
| USA | 1976–2018 | H6N1 | American Black Duck, America Green-Winged Teal, American Wigeon, Wild Bird (unspecified), Environmental Sample, Blue-Winged Teal, Canada Goose, Common Goldeneye, Duck (unspecified), Long-Tailed Duck, Gadwall Duck, Eurasian Teal, Glaucous Gull, Cackling Goose, Greater White-Fronted Goose, Greater Scaup, Goose (unspecified), gull (unspecified), Hawk, Laughing Gull, Lesser Scaup, Mallard, Mottled Duck, Northern Pintail, Northern Shoveler Duck, Ring-Necked Duck, Ruddy Turnstone, Sanderling, Sandpiper, Shorebird (unspecified), Snow Goose, Steller’s Eider, Turkey, White-Fronted Goose | 349 | 962 |
| 1986–2015 | H6N2 | American Black Duck, America Green-Winged Teal, American Wigeon, Wild Bird (unspecified), Black Duck, Environmental Sample, Blue-Winged Teal, Chicken, Cinnamon Teal, Duck (unspecified), Gadwall Duck, Eurasian Teal, Greater White-Fronted Goose, Herring Gull, Goose (unspecified), Japanese Quail, Laughing Gull, Mallard, Mottled Duck, Northern Pintail, Northern Shoveler Duck, Pintail, Quail, Redhead, Ruddy Turnstone, Shorebird (unspecified), Turkey | 244 | ||
| 1989–2009 | H6N3 | Chicken, Herring Gull, Laughing Gull, Mallard | 7 | ||
| 1982–2016 | H6N4 | Environmental Sample, Gull (unspecified), Laughing Gull, Mallard, Northern Pintail, Ruddy Turnstone, Sanderling, Shorebird | 32 | ||
| 1975–2015 | H6N5 | American Black Duck, America Green-Winged Teal, American Wigeon, Blue-Winged Teal, Chicken, Duck (unspecified), Environmental Sample, Green-Winged Teal, Mallard, Northern Pintail, Pintail, Red Knot, Ruddy Turnstone, Turkey | 58 | ||
| 1980–2018 | H6N6 | American Wigeon, Blue-Winged Teal, Duck (unspecified), Green-Winged Teal, Mallard, Northern Pintail, Ruddy Turnstone, Sanderling, Turkey | 31 | ||
| 2000–2018 | H6N7 | American Green-Winged Teal, Blue-Winged Teal, Mallard, Ruddy Turnstone | 5 | ||
| 1977–2016 | H6N8 | American Black Duck, America Green-Winged Teal, American Wigeon, Wild Bird (unspecified), Black Duck, Blue-Winged Teal, Bufflehead, Canada Goose, Chicken, Duck (unspecified), Domestic Duck, Wood Duck, Environmental Sample, Eurasian Teal, Ross’s Goose, Herring Gull, Knot, Laughing Gull, Lesser Scaup, Mallard, Northern Pintail, Northern Shoveler, Pintail, Ring-Necked Duck, Ruddy Turnstone, Shorebird (unspecified), Shoveler, Turkey | 198 | ||
| 1975–2014 | H6N9 | Duck (unspecified), Long-Tailed Duck, Green-Winged Teal, Mallard | 4 | ||
| Vietnam | 2013–2018 | H6N1 | Chicken, Duck (unspecified), Muscovy Duck | 6 | 174 |
| 2007–2014 | H6N2 | Duck (unspecified), Environmental Samples, Mallard, Muscovy Duck | 40 | ||
| 2010–2018 | H6N6 | Chicken, Duck (unspecified), Environmental Samples, Mallard, Muscovy Duck | 124 | ||
| 2013–2014 | H6N8 | Duck (unspecified), Mallard | 2 | ||
| 2011 | H6N9 | Duck (unspecified) | 1 | ||
| Zambia | 2008 | H6N2 | Duck (unspecified) | 4 | 4 |
| Protein | Strain | Subtype | Host # | Mutation * | Effect | Ref |
|---|---|---|---|---|---|---|
| HA | A/Taiwan/2/2013 | H6N1 | Human | G225D | Increasing its affinity for the human α2-6 linked sialic acid receptors | [83] |
| S228G | Used in triad to overcome the requirement for a hydrophobic residue and preferentially bind to human-like receptors | [74] | ||||
| N137 | ||||||
| V190D | ||||||
| P186L | Binding preference for the human receptor analog | [17] | ||||
| Q226L | Increased affinity for human-type receptor | [67] | ||||
| V135S | Increased affinity for human-type receptor | |||||
| T136 | Increased affinity for human-type receptor | |||||
| V190D | Increased affinity for human-type receptor | [17] | ||||
| A/swine/Guangdong/K6/2010 | H6N6 | Swine | A222V | Increased human α2-6 linked sialic acid receptor preference | [81] | |
| S228G | Increased human α2-6 linked sialic acid receptor preference | |||||
| A/Eurasian teal/Egypt/P2-29/2017 | H6N2 | Avian | T160A | Increased binding to human-type influenza receptor | [51] | |
| A/swine/Guangdong/K6/2010 [GDK6-MA] | H6N6 | Mouse | L111F | Increasing its affinity for the human α2-6 linked sialic acid receptors | [84] | |
| H156N | Increased plaque size on MDCKs have enhanced early stage viral replication (H156N only) and concurrent occurrence alongside PA-I38M creates a significantly more virulent variant which compensates for lack of PB2-E627K | |||||
| S263R | ||||||
| A/chicken/Guangdong/S1311/2010 | H6N6 | Avian | I55V | Increased affinity for human-type receptor | [85] | |
| S137N | Allows the binding to both human- and avian-like receptors | [67] | ||||
| E190 | ||||||
| G228S | ||||||
| G225D | Increasing its affinity for the human α2-6 linked sialic acid receptors | |||||
| Q226L | Increased affinity for human-type receptor | |||||
| A/environment/Jiangxi/02.05 YGYXG001/2015 | H6N6 | Avian | S228G | Increased affinity for human-type receptor | [17] | |
| PB2 | A/Mallard/San Jiang/275/2007 (MA-P8M3) | H6N1 | Mice | E627K | Enhanced RNA polymerase activity and viral replication which contributes to mammalian adaptation | [73] |
| A/canine/Taiwan/E01/2014 | H6N1 | Dog | E627K | Enhanced RNA polymerase activity and viral replication which contributes to mammalian adaptation | [82] | |
| A/Eurasian teal/Egypt/P2-29/2017 | H6N2 | Avian | V291I | Host specificity shift towards human | [51] | |
| A/duck/Zambia/03/08 | H6N2 | Avian | 475M | Increased affinity for human-type receptor | [52] | |
| A/swine/Guangdong/K6/2010 [GDK6-MA] | H6N6 | Mouse | E627K | Enhanced RNA polymerase activity and viral replication which contributes to mammalian adaptation | [84] | |
| PA | A/Mallard/San Jiang/275/2007 (MA-P8M3) | H6N1 | Mice | T97I | Enhanced RNA polymerase activity and viral replication which contributes to mammalian adaptation | [73] |
| A/swine/Guangdong/K6/2010 [GDK6-MA] | H6N6 | Mouse | I38M | Raised polymerase activity in vitro | [84] | |
| M1 | A/Eurasian teal/Egypt/P2-29/2017 | H6N2 | Avian | N30D | When occurring concurrently, creates increased virulence (as seen in H5N1) | [51] |
| T215A | ||||||
| M2 | A/chicken/Guangdong/S1312/2010 | H6N2 | Avian | S31N | Increased amantadine and rimantadine resistance | [20] |
| A/chicken/Guangdong/S1311/2010 | H6N6 | Avian | S31N | Increased amantadine and rimantadine resistance | [20] | |
| A/duck/India/11CL01/2014 | H6N2 | Avian | V27I | Increased amantadine resistance | [34] | |
| NS1 | A/Eurasian teal/Egypt/P2-29/2017 | H6N2 | Avian | P42S | Increased virulence | [51] |
| N205S | ||||||
| A/wild duck/Shantou/2853/2003 | H6N2 | Avian | D92 | Increased virulence | [17] | |
| PB1-F2 | A/Eurasian teal/Egypt/P2-29/2017 | H6N2 | Avian | N66S | Altered virulence and antivirus response in mice | [51] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Everest, H.; Hill, S.C.; Daines, R.; Sealy, J.E.; James, J.; Hansen, R.; Iqbal, M. The Evolution, Spread and Global Threat of H6Nx Avian Influenza Viruses. Viruses 2020, 12, 673. https://doi.org/10.3390/v12060673
Everest H, Hill SC, Daines R, Sealy JE, James J, Hansen R, Iqbal M. The Evolution, Spread and Global Threat of H6Nx Avian Influenza Viruses. Viruses. 2020; 12(6):673. https://doi.org/10.3390/v12060673
Chicago/Turabian StyleEverest, Holly, Sarah C. Hill, Rebecca Daines, Joshua E. Sealy, Joe James, Rowena Hansen, and Munir Iqbal. 2020. "The Evolution, Spread and Global Threat of H6Nx Avian Influenza Viruses" Viruses 12, no. 6: 673. https://doi.org/10.3390/v12060673
APA StyleEverest, H., Hill, S. C., Daines, R., Sealy, J. E., James, J., Hansen, R., & Iqbal, M. (2020). The Evolution, Spread and Global Threat of H6Nx Avian Influenza Viruses. Viruses, 12(6), 673. https://doi.org/10.3390/v12060673







