Biphasic Packing of DNA and Internal Proteins in Bacteriophage T4 Heads Revealed by Bubblegram Imaging
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of Virions and Recording of Bubblegrams
2.2. Estimation of DNA Content of a T4 Virion in Coaxial Spool Models
2.3. Image Averaging and Difference Images
3. Results and Discussion
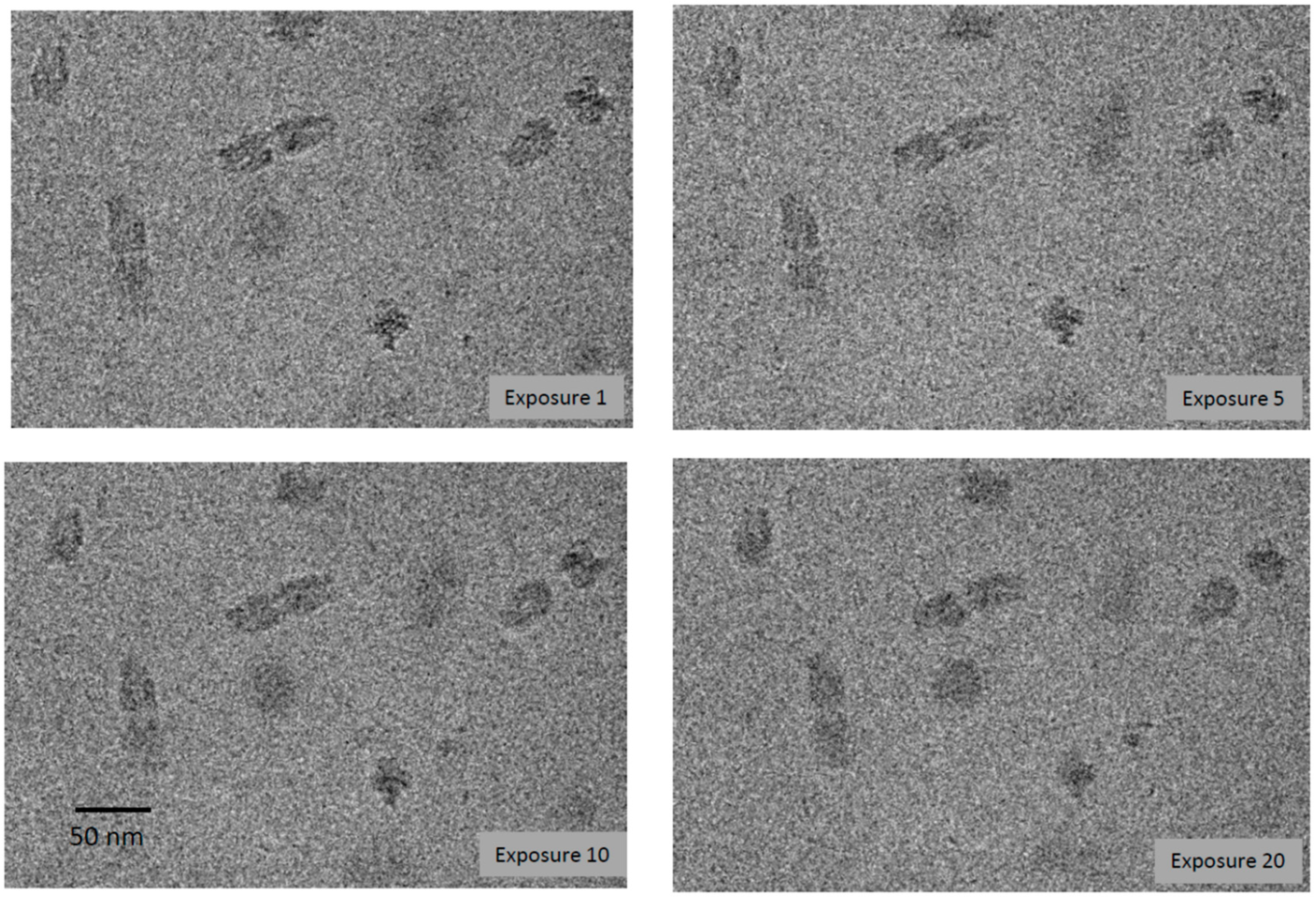
3.1. DNA Complexes Are Bubbling-Reluctant: Origami Do Not Bubble after a Dose of 450 el/Å2
3.2. Bubblegrams of Bacteriophage T4 Heads
3.3. Bubblegram Imaging Supports a Biphasic Model for Wild-Type (Prolate) T4 Heads
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Richards, K.E.; Williams, R.C.; Calendar, R. Mode of DNA packing within bacteriophage heads. J. Mol. Biol. 1973, 78, 255–259. [Google Scholar] [CrossRef]
- Black, L.W.; Showe, M.K.; Steven, A.C. Morphogenesis of the T4 head. In Molecular Biology of Bacteriophage T4; Karam, J., Ed.; American Society for Microbiology: Washington, DC, USA, 1994; pp. 218–258. [Google Scholar]
- Black, L.W. DNA packaging in dsDNA bacteriophages. Annu. Rev. Microbiol. 1989, 43, 267–292. [Google Scholar] [CrossRef] [PubMed]
- Earnshaw, W.C.; King, J.; Harrison, S.C.; Eiserling, F.A. The structural organization of DNA packaged within the heads of T4 wild-type, isometric and giant bacteriophages. Cell 1978, 14, 559–568. [Google Scholar] [CrossRef]
- Harrison, S.C. Packaging of DNA into bacteriophage heads: A model. J. Mol. Biol. 1983, 171, 577–580. [Google Scholar] [CrossRef]
- Black, L.W.; Newcomb, W.W.; Boring, J.W.; Brown, J.C. Ion etching bacteriophage T4: Support for a spiral-fold model of packaged DNA. Proc. Natl. Acad. Sci. USA 1985, 82, 7960–7964. [Google Scholar] [CrossRef]
- Widom, J.; Baldwin, R.L. Tests of spool models for DNA packaging in phage lambda. J. Mol. Biol. 1983, 171, 419–437. [Google Scholar] [CrossRef]
- Cerritelli, M.E.; Cheng, N.; Rosenberg, A.H.; McPherson, C.E.; Booy, F.P.; Steven, A.C. Encapsidated Conformation of Bacteriophage T7 DNA. Cell 1997, 91, 271–280. [Google Scholar] [CrossRef]
- Duda, R.L.; Ross, P.D.; Cheng, N.; Firek, B.A.; Hendrix, R.W.; Conway, J.F.; Steven, A.C. Structure and Energetics of Encapsidated DNA in Bacteriophage HK97 Studied by Scanning Calorimetry and Cryo-electron Microscopy. J. Mol. Biol. 2009, 391, 471–483. [Google Scholar] [CrossRef]
- Chang, J.; Weigele, P.; King, J.; Chiu, W.; Jiang, W. Cryo-EM Asymmetric Reconstruction of Bacteriophage P22 Reveals Organization of its DNA Packaging and Infecting Machinery. Structure 2006, 14, 1073–1082. [Google Scholar] [CrossRef]
- Lander, G.C.; Tang, L.; Casjens, S.R.; Gilcrease, E.B.; Prevelige, P.; Poliakov, A.; Potter, C.S.; Carragher, B.; Johnson, J.E. The Structure of an Infectious P22 Virion Shows the Signal for Headful DNA Packaging. Science 2006, 312, 1791–1795. [Google Scholar] [CrossRef]
- Booy, F.P.; Newcomb, W.W.; Trus, B.L.; Brown, J.C.; Baker, T.S.; Steven, A.C. Liquid-crystalline, phage-like packing of encapsidated DNA in herpes simplex virus. Cell 1991, 64, 1007–1015. [Google Scholar] [CrossRef]
- McElwee, M.; Vijayakrishnan, S.; Rixon, F.; Bhella, D. Structure of the herpes simplex virus portal-vertex. PLoS Biol. 2018, 16, e2006191. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-T.; Jih, J.; Dai, X.H.; Bi, G.-Q.; Zhou, Z.H. Cryo-EM structures of herpes simplex virus type 1 portal vertex and packaged genome. Nature 2019, 570, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Aebi, U.; Bijlenga, R.; Broek, J.v.d.; Broek, R.v.d.; Eiserling, F.; Kellenberger, C.; Kellenberger, E.; Mesyanzhinov, V.; Müller, L.; Showe, M.; et al. The transformation of tau particles into T4 heads. II. Transformations of the surface lattice and related observations on form determination. J. Supramol. Struct. 1974, 2, 253–275. [Google Scholar] [CrossRef]
- Fokine, A.; Chipman, P.R.; Leiman, P.G.; Mesyanzhinov, V.V.; Rao, V.B.; Rossmann, M.G. Molecular architecture of the prolate head of bacteriophage T4. Proc. Natl. Acad. Sci. USA 2004, 101, 6003–6008. [Google Scholar] [CrossRef]
- Doermann, A.H.; Eiserling, F.A.; Boehner, L. Genetic Control of Capsid Length in Bacteriophage T4. I. Isolation and Preliminary Description of Four New Mutants. J. Virol. 1973, 12, 374–385. [Google Scholar] [CrossRef]
- Bijlenga, R.; Aebi, U.; Kellenberger, E. Properties and structure of a gene 24-controlled T4 giant phage. J. Mol. Biol. 1976, 103, 469–498. [Google Scholar] [CrossRef]
- Aebi, U.; Bijlenga, R.K.L.; Heggeler, B.T.; Kistler, J.; Steven, A.C.; Smith, P.R. Comparison of the structural and chemical composition of giant T-even phage heads. J. Supramol. Struct. 1976, 5, 475–495. [Google Scholar] [CrossRef]
- Cheng, N.; Wu, W.; Steven, A.C.; Thomas, J.; Black, L. Bubblegrams reveal the inner body of bacteriophage phiKZ. Microsc. Microanal. 2012, 18, 112. [Google Scholar] [CrossRef]
- Mishyna, M.; Volokh, O.; Danilova, Y.; Gerasimova, N.; Pechnikova, E.; Sokolova, O.S. Effects of radiation damage in studies of protein-DNA complexes by cryo-EM. Micron 2017, 96, 57–64. [Google Scholar] [CrossRef]
- Leapman, R.D.; Sun, S. Cryo-electron energy loss spectroscopy: Observations on vitrified hydrated specimens and radiation damage. Ultramicroscopy 1995, 59, 71–79. [Google Scholar] [CrossRef]
- Bai, X.-C.; Martin, T.G.; Scheres, S.H.W.; Dietz, H. Cryo-EM structure of a 3D DNA-origami object. Proc. Natl. Acad. Sci. USA 2012, 109, 20012–20017. [Google Scholar] [CrossRef] [PubMed]
- Cheng, N.; Wu, W.; Watts, N.R.; Steven, A.C. Exploiting radiation damage to map proteins in nucleoprotein complexes: The internal structure of bacteriophage T7. J. Struct. Biol. 2014, 185, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Leavitt, J.C.; Cheng, N.; Gilcrease, E.B.; Motwani, T.; Teschke, C.M.; Casjens, S.R.; Steven, A.C. Localization of the Houdinisome (Ejection Proteins) inside the Bacteriophage P22 Virion by Bubblegram Imaging. mBio 2016, 7. [Google Scholar] [CrossRef]
- Fontana, J.; Jurado, K.A.; Cheng, N.; Ly, N.L.; Fuchs, J.R.; Gorelick, R.J.; Engelman, A.; Steven, A. Distribution and Redistribution of HIV-1 Nucleocapsid Protein in Immature, Mature, and Integrase-Inhibited Virions: A Role for Integrase in Maturation. J. Virol. 2015, 89, 9765–9780. [Google Scholar] [CrossRef]
- Mullaney, J.M.; Black, L.W. Capsid Targeting Sequence Targets Foreign Proteins into Bacteriophage T4 and Permits Proteolytic Processing. J. Mol. Biol. 1996, 261, 372–385. [Google Scholar] [CrossRef]
- Mullaney, J.M.; Black, L.W. Activity of foreign proteins targeted within the bacteriophage T4 head and prohead: Implications for packaged DNA structure. J. Mol. Biol. 1998, 283, 913–929. [Google Scholar] [CrossRef]
- Cerritelli, M.E.; Trus, B.L.; Smith, C.S.; Cheng, N.; Conway, J.F.; Steven, A.C. A second symmetry mismatch at the portal vertex of bacteriophage T7: 8-fold symmetry in the procapsid core. J. Mol. Biol. 2003, 327, 1–6. [Google Scholar] [CrossRef]
- Mullaney, J.M.; Black, L.W. Bacteriophage T4 Capsid Packaging and Unpackaging of DNA and Proteins. Methods Mol. Biol. 2014, 1108, 69–85. [Google Scholar] [CrossRef]
- Mullaney, J.M.; Thompson, R.B.; Gryczynski, Z.; Black, L.W. Green fluorescent protein as a probe of rotational mobility within bacteriophage T4. J. Virol. Methods 2000, 88, 35–40. [Google Scholar] [CrossRef]
- Yu, T.-Y.; Schaefer, J. REDOR NMR Characterization of DNA Packaging in Bacteriophage T4. J. Mol. Biol. 2008, 382, 1031–1042. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, W.; Cheng, N.; Black, L.W.; Dietz, H.; Steven, A.C. Biphasic Packing of DNA and Internal Proteins in Bacteriophage T4 Heads Revealed by Bubblegram Imaging. Viruses 2020, 12, 1282. https://doi.org/10.3390/v12111282
Wu W, Cheng N, Black LW, Dietz H, Steven AC. Biphasic Packing of DNA and Internal Proteins in Bacteriophage T4 Heads Revealed by Bubblegram Imaging. Viruses. 2020; 12(11):1282. https://doi.org/10.3390/v12111282
Chicago/Turabian StyleWu, Weimin, Naiqian Cheng, Lindsay W. Black, Hendrik Dietz, and Alasdair C. Steven. 2020. "Biphasic Packing of DNA and Internal Proteins in Bacteriophage T4 Heads Revealed by Bubblegram Imaging" Viruses 12, no. 11: 1282. https://doi.org/10.3390/v12111282
APA StyleWu, W., Cheng, N., Black, L. W., Dietz, H., & Steven, A. C. (2020). Biphasic Packing of DNA and Internal Proteins in Bacteriophage T4 Heads Revealed by Bubblegram Imaging. Viruses, 12(11), 1282. https://doi.org/10.3390/v12111282




