Detection of Two Highly Diverse Peribunyaviruses in Mosquitoes from Palenque, Mexico
Abstract
1. Introduction
2. Materials and Methods
2.1. Mosquito Sampling and RT-PCR Screening
2.2. Genome Sequencing
2.3. Genome and Phylogenetic Analyses
2.4. Cell Culture and Virus Replication Analyses
2.5. Nucleotide Sequence Accession Numbers
3. Results
3.1. Identification of Orthobunyaviruses and Herbeviruses
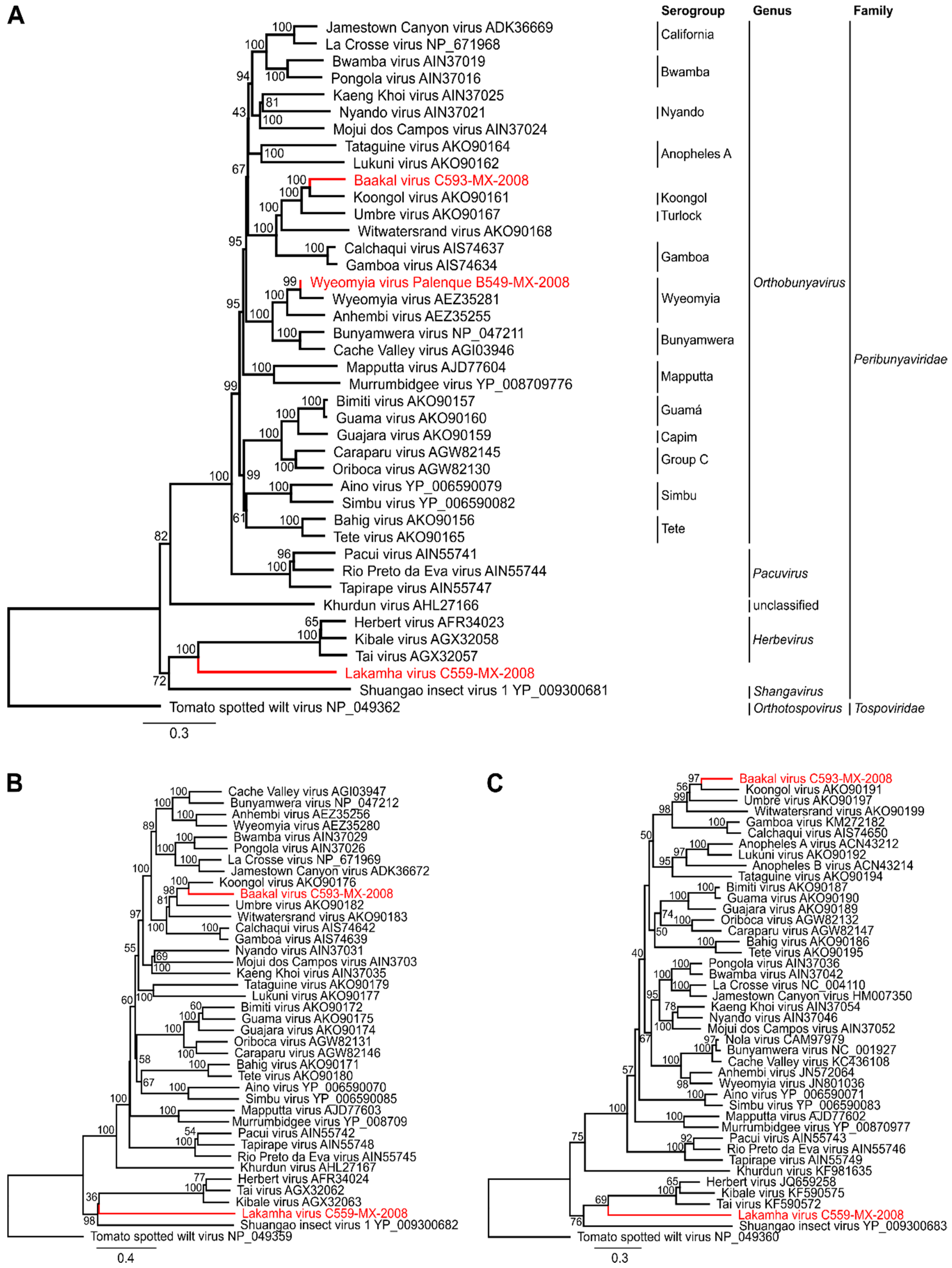
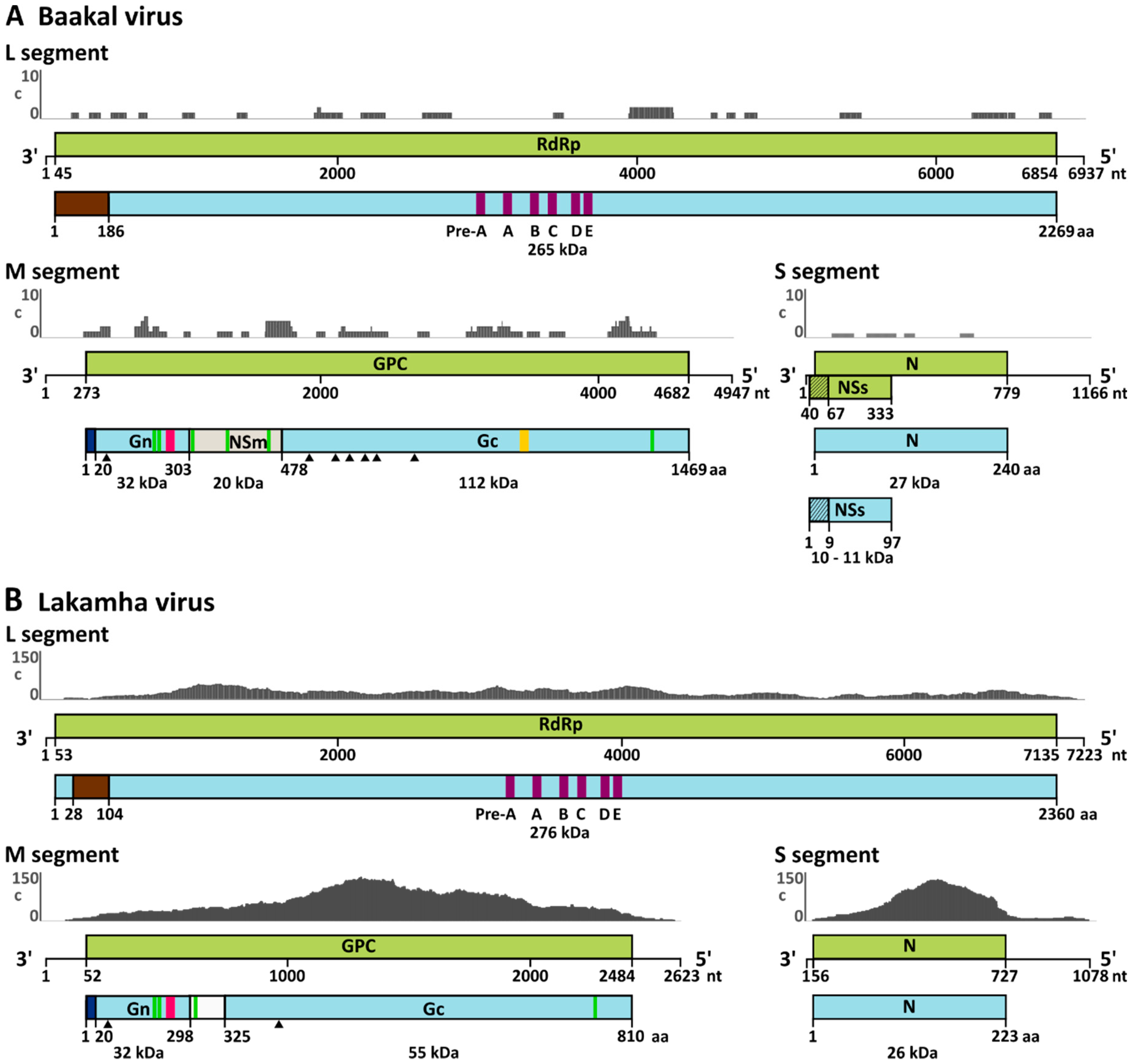
3.2. Full Genome Sequencing and Phylogenetic Relationship
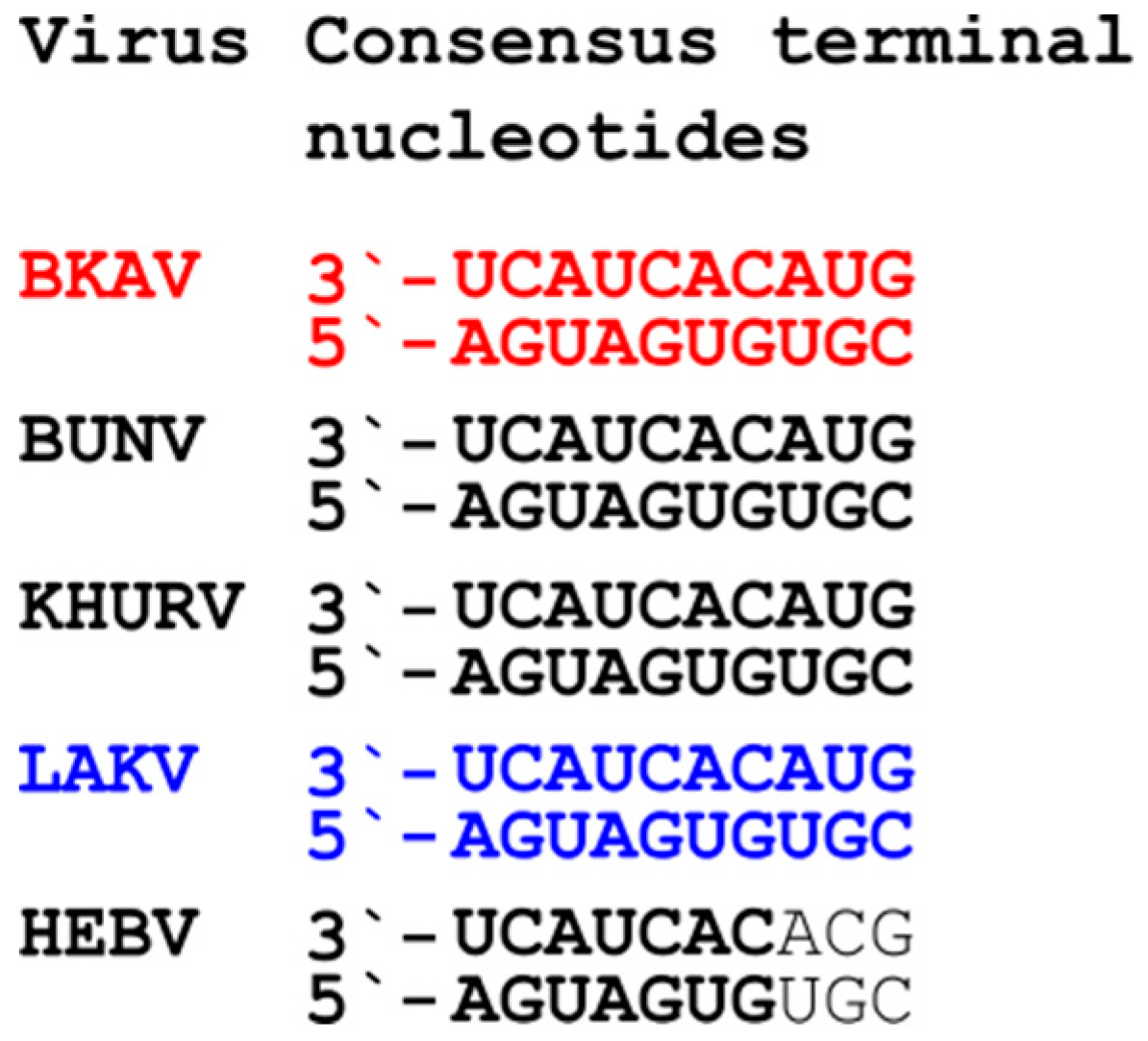
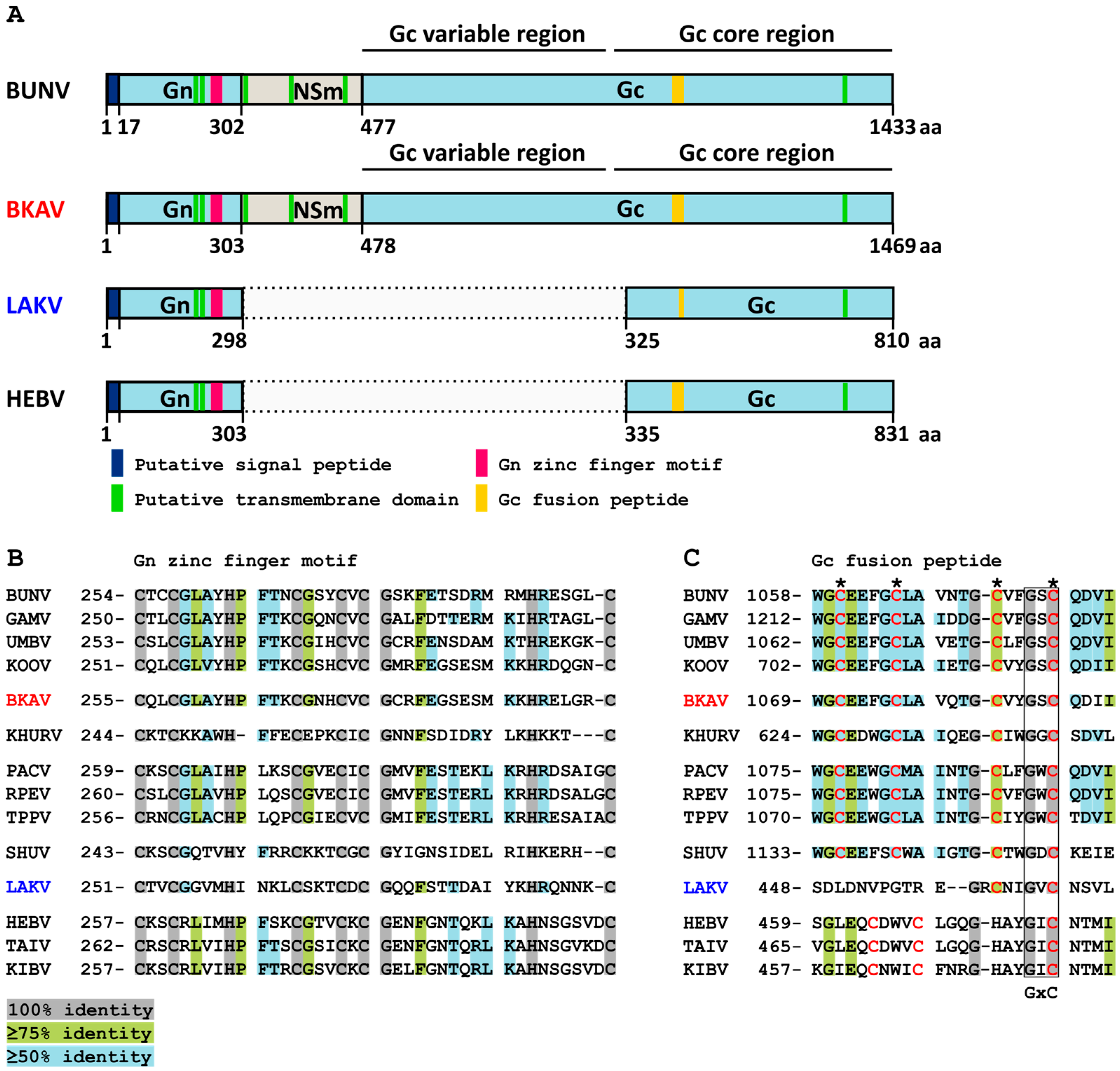
3.3. Genome Analyses
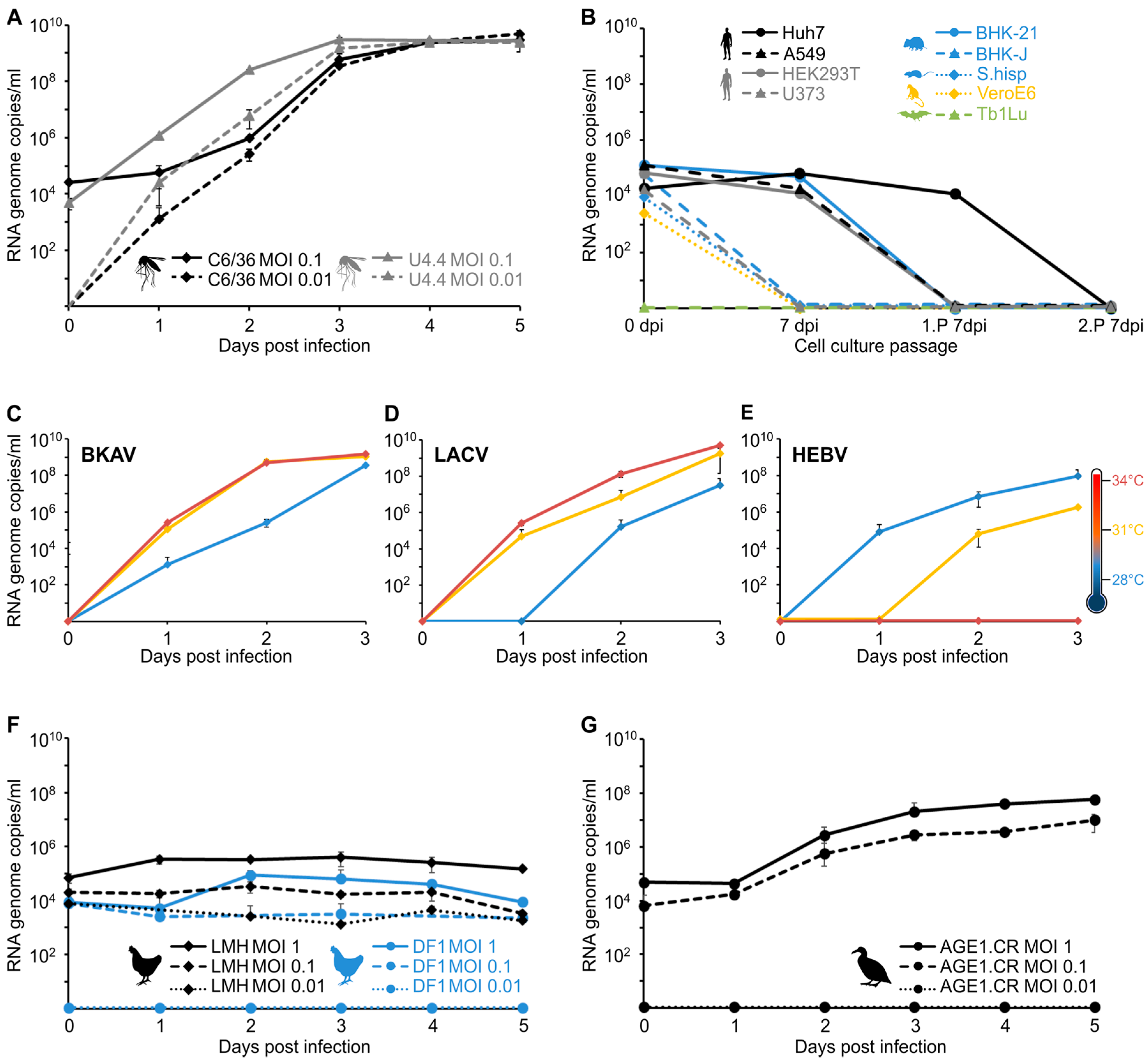
3.4. Virus Isolation and Growth Analyses in Insect and Vertebrate Cells
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Abudurexiti, A.; Adkins, S.; Alioto, D.; Alkhovsky, S.V.; Avšič-Županc, T.; Ballinger, M.J.; Bente, D.A.; Beer, M.; Bergeron, É.; Blair, C.D.; et al. Taxonomy of the order Bunyavirales: Update 2019. Arch. Virol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Dutuze, M.F.; Nzayirambaho, M.; Mores, C.N.; Christofferson, R.C. A Review of Bunyamwera, Batai, and Ngari Viruses: Understudied Orthobunyaviruses With Potential One Health Implications. Front. Vet. Sci. 2018, 5, 69. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.M. Orthobunyaviruses: Recent genetic and structural insights. Nat. Rev. Microbiol. 2014, 12, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Blitvich, B.J.; Saiyasombat, R.; Travassos da Rosa, A.; Tesh, R.B.; Calisher, C.H.; Garcia-Rejon, J.E.; Farfán-Ale, J.A.; Loroño, R.E.; Bates, A.; Loroño-Pino, M.A. Orthobunyaviruses, a Common Cause of Infection of Livestock in the Yucatan Peninsula of Mexico. Am. J. Trop. Med. Hyg. 2012, 87, 1132–1139. [Google Scholar] [CrossRef] [PubMed]
- Romero-Alvarez, D.; Escobar, L.E. Oropouche fever, an emergent disease from the Americas. Microbes Infect. 2018, 20, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Kohl, A.; Dunn, E.F.; Lowen, A.C.; Elliott, R.M. Complementarity, sequence and structural elements within the 3′ and 5′ non-coding regions of the Bunyamwera orthobunyavirus S segment determine promoter strength. J. Gen. Virol. 2004, 85, 3269–3278. [Google Scholar] [CrossRef]
- Schmaljohn, C.S.; Nichol, S.T. Bunyaviridae. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; Volume 2, pp. 1741–1789. [Google Scholar]
- Jonkers, A.H.; Spence, L.; Downs, W.G.; Aitken, T.H.; Tikasingh, E.S. Arbovirus studies in bBush Bush forest, Trinidad, W.I., September 1959-December 1964. V. Virus isolations. Am. J. Trop. Med. Hyg. 1968, 17, 276–284. [Google Scholar] [CrossRef]
- Rodrigues, D.S.; Medeiros, D.B.; Rodrigues, S.G.; Martins, L.C.; de Lima, C.P.; de Oliveira, L.F.; de Vasconcelos, J.M.; Da Silva, D.E.; Cardoso, J.F.; da Silva, S.P.; et al. Pacui Virus, Rio Preto da Eva Virus, and Tapirape Virus, Three Distinct Viruses within the Family Bunyaviridae. Genome Announc. 2014, 2, e00923-14. [Google Scholar] [CrossRef]
- Marklewitz, M.; Zirkel, F.; Rwego, I.B.; Heidemann, H.; Trippner, P.; Kurth, A.; Kallies, R.; Briese, T.; Lipkin, W.I.; Drosten, C.; et al. Discovery of a unique novel clade of mosquito-associated bunyaviruses. J. Virol. 2013, 87, 12850–12865. [Google Scholar] [CrossRef]
- Junglen, S.; Marklewitz, M.; Zirkel, F.; Wollny, R.; Meyer, B.; Heidemann, H.; Metzger, S.; Annan, A.; Dei, D.; Leendertz, F.H.; et al. No Evidence of Gouleako and Herbert Virus Infections in Pigs, Cote d’Ivoire and Ghana. Emerg. Infect. Dis. 2015, 21, 2190–2193. [Google Scholar] [CrossRef]
- Marklewitz, M.; Zirkel, F.; Kurth, A.; Drosten, C.; Junglen, S. Evolutionary and phenotypic analysis of live virus isolates suggests arthropod origin of a pathogenic RNA virus family. Proc. Natl. Acad. Sci. USA 2015, 112, 7536–7541. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.J.; Lefkowitz, E.J.; King, A.M.Q.; Harrach, B.; Harrison, R.L.; Knowles, N.J.; Kropinski, A.M.; Krupovic, M.; Kuhn, J.H.; Mushegian, A.R.; et al. Changes to taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2017). Arch. Virol. 2017, 162, 2505–2538. [Google Scholar] [CrossRef] [PubMed]
- Li, C.X.; Shi, M.; Tian, J.H.; Lin, X.D.; Kang, Y.J.; Chen, L.J.; Qin, X.C.; Xu, J.; Holmes, E.C.; Zhang, Y.Z. Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. eLife 2015, 4, e05378. [Google Scholar] [CrossRef] [PubMed]
- Al’kovskhovskii, S.V.; Shchetinin, A.M.; L’Vov D, K.; Shchelkanov, M.; Deriabin, P.G.; L’Vov D, N.; Samokhvalov, E.I.; Gitel’man, A.K.; Botikov, A.G. [The Khurdun virus (KHURV): A new representative of the orthobunyavirus (Bunyaviridae)]. Vopr. Virusol. 2013, 58, 10–13. [Google Scholar] [PubMed]
- Galkina, I.V.; L’Vov L, N.; Gromashevskii, V.L.; Moskvina, T.M. [Khurdun virus, a presumably new RNA-containing virus associated with coots (Fulica atra), isolated in the Volga river delta]. Vopr. Virusol. 2005, 50, 29–31. [Google Scholar] [PubMed]
- Esposito, D.L.A.; Fonseca, B. Will Mayaro virus be responsible for the next outbreak of an arthropod-borne virus in Brazil? Braz. J. Infect. Dis. 2017, 21, 540–544. [Google Scholar] [CrossRef] [PubMed]
- Mavian, C.; Rife, B.D.; Dollar, J.J.; Cella, E.; Ciccozzi, M.; Prosperi, M.C.F.; Lednicky, J.; Morris, J.G.; Capua, I.; Salemi, M. Emergence of recombinant Mayaro virus strains from the Amazon basin. Sci. Rep. 2017, 7, 8718. [Google Scholar] [CrossRef] [PubMed]
- Mackay, I.M.; Arden, K.E. Mayaro virus: A forest virus primed for a trip to the city? Microbes Infect. 2016, 18, 724–734. [Google Scholar] [CrossRef] [PubMed]
- Romero-Alvarez, D.; Escobar, L.E. Vegetation loss and the 2016 Oropouche fever outbreak in Peru. Mem. Inst. Oswaldo Cruz 2017, 112, 292–298. [Google Scholar] [CrossRef]
- Sakkas, H.; Bozidis, P.; Franks, A.; Papadopoulou, C. Oropouche Fever: A Review. Viruses 2018, 10, 175. [Google Scholar] [CrossRef]
- Travassos da Rosa, J.F.; de Souza, W.M.; Pinheiro, F.P.; Figueiredo, M.L.; Cardoso, J.F.; Acrani, G.O.; Nunes, M.R.T. Oropouche Virus: Clinical, Epidemiological, and Molecular Aspects of a Neglected Orthobunyavirus. Am. J. Trop. Med. Hyg. 2017, 96, 1019–1030. [Google Scholar] [PubMed]
- Hontz, R.D.; Guevara, C.; Halsey, E.S.; Silvas, J.; Santiago, F.W.; Widen, S.G.; Wood, T.G.; Casanova, W.; Vasilakis, N.; Watts, D.M.; et al. Itaya virus, a Novel Orthobunyavirus Associated with Human Febrile Illness, Peru. Emerg. Infect. Dis. 2015, 21, 781–788. [Google Scholar] [CrossRef] [PubMed]
- Kopp, A.; Gillespie, T.R.; Hobelsberger, D.; Estrada, A.; Harper, J.M.; Miller, R.A.; Eckerle, I.; Muller, M.A.; Podsiadlowski, L.; Leendertz, F.H.; et al. Provenance and geographic spread of St. Louis encephalitis virus. mBio 2013, 4, e00322-13. [Google Scholar] [CrossRef] [PubMed]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar] [PubMed]
- Jordan, I.; Vos, A.; Beilfuss, S.; Neubert, A.; Breul, S.; Sandig, V. An avian cell line designed for production of highly attenuated viruses. Vaccine 2009, 27, 748–756. [Google Scholar] [CrossRef]
- Himly, M.; Foster, D.N.; Bottoli, I.; Iacovoni, J.S.; Vogt, P.K. The DF-1 chicken fibroblast cell line: Transformation induced by diverse oncogenes and cell death resulting from infection by avian leukosis viruses. Virology 1998, 248, 295–304. [Google Scholar] [CrossRef] [PubMed]
- Kawaguchi, T.; Nomura, K.; Hirayama, Y.; Kitagawa, T. Establishment and characterization of a chicken hepatocellular carcinoma cell line, LMH. Cancer Res. 1987, 47, 4460–4464. [Google Scholar]
- Calisher, C.H.; Lazuick, J.S.; Justines, G.; Francy, D.B.; Monath, T.P.; Gutierrez, V.E.; Sabattini, M.S.; Bowen, G.S.; Jakob, W.L. Viruses Isolated from Aedeomyia Squamipennis Mosquitoes Collected in Panama, Ecuador, and Argentina: Establishment of the Gamboa Serogroup. Am. J. Trop. Med. Hyg. 1981, 30, 219–223. [Google Scholar] [CrossRef]
- Shchetinin, A.; Lvov, D.; Deriabin, P.; Botikov, A.; Gitelman, A.; Kuhn, J.; Alkhovsky, S. Genetic and Phylogenetic Characterization of Tataguine and Witwatersrand Viruses and Other Orthobunyaviruses of the Anopheles A, Capim, Guamá, Koongol, Mapputta, Tete, and Turlock Serogroups. Viruses 2015, 7, 5987–6008. [Google Scholar] [CrossRef]
- Watanabe, C.; Uchida, Y.; Ito, H.; Ito, T.; Saito, T. Host immune-related gene responses against highly pathogenic avian influenza virus infection in vitro differ among chicken cell lines established from different organs. Vet. Immunol. Immunopathol. 2011, 144, 187–199. [Google Scholar] [CrossRef]
- Bachmann, M.; Breitwieser, T.; Lipps, C.; Wirth, D.; Jordan, I.; Reichl, U.; Frensing, T. Impaired antiviral response of adenovirus-transformed cell lines supports virus replication. J. Gen. Virol. 2016, 97, 293–298. [Google Scholar] [CrossRef] [PubMed]
- Barber, M.R.; Aldridge, J.R., Jr.; Webster, R.G.; Magor, K.E. Association of RIG-I with innate immunity of ducks to influenza. Proc. Natl. Acad. Sci. USA 2010, 107, 5913–5918. [Google Scholar] [CrossRef] [PubMed]
- Giard, D.J.; Aaronson, S.A.; Todaro, G.J.; Arnstein, P.; Kersey, J.H.; Dosik, H.; Parks, W.P. In vitro cultivation of human tumors: Establishment of cell lines derived from a series of solid tumors. J. Natl. Cancer Inst. 1973, 51, 1417–1423. [Google Scholar] [CrossRef] [PubMed]
- Lieber, M.; Smith, B.; Szakal, A.; Nelson-Rees, W.; Todaro, G. A continuous tumor-cell line from a human lung carcinoma with properties of type II alveolar epithelial cells. Int. J. Cancer 1976, 17, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Day, J.F.; Curtis, G.A. When it rains, they soar—And that makes Culex nigripalpus a dangerous mosquito. Am. Entomol. 1994, 40, 162–167. [Google Scholar] [CrossRef]
- Doherty, R.L.; Wetters, E.J.; Gorman, B.M.; Whitehead, R.H. Arbovirus infection in Western Queensland: Serological studies, 1963–1969. Trans. R. Soc. Trop. Med. Hyg. 1970, 64, 740–747. [Google Scholar] [CrossRef]
- Doherty, R.L.; Whitehead, R.H.; Wetters, E.J.; Gorman, B.M.; Carley, J.G. A survey of antibody to 10 arboviruses (Koongol group, Mapputta group and ungrouped) isolated in Queensland. Trans. R. Soc. Trop. Med. Hyg. 1970, 64, 748–753. [Google Scholar] [CrossRef]
- Sanderson, C.J. A serologic survey of Queensland cattle for evidence of arbovirus infections. Am. J. Trop. Med. Hyg. 1969, 18, 433–439. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.M. Catalogue of Arthropod-Borne Viruses of the World. Am. J. Public Health Nations Health. 1968, 58, 2177. [Google Scholar]
- Karabatsos, N. International Catalogue of Arboviruses, Including Certain other Viruses of Vertebrates, 3rd ed.; American Society of Tropical Medicine and Hygiene for The Subcommittee on Information Exchange of the American Committee on Arthropod-borne Viruses: San Antonio, TX, USA, 1985. [Google Scholar]
- Whitehouse, C.A.; Kuhn, J.H.; Wada, J.; Ergunay, K. Family Bunyaviridae. In Global Virology I—Identifying and Investigating Viral Diseases; Shapshak, P., Sinnott, J., Somboonwit, C., Kuhn, J.H., Eds.; Springer: New York, NY, USA, 2015; pp. 199–246. [Google Scholar]
- Calisher, C.H.; Lazuick, J.S.; Sudia, W.D. Brus Laguna Virus, a Gamboa Bunyavirus from Aedeomyia Squamipennis Collected in Honduras. Am. J. Trop. Med. Hyg. 1988, 39, 406–408. [Google Scholar] [CrossRef]
- Eastwood, G.; Loaiza, J.R.; Pongsiri, M.J.; Sanjur, O.I.; Pecor, J.E.; Auguste, A.J.; Kramer, L.D. Enzootic Arbovirus Surveillance in Forest Habitat and Phylogenetic Characterization of Novel Isolates of Gamboa Virus in Panama. Am. J. Trop. Med. Hyg. 2016. [Google Scholar] [CrossRef] [PubMed]
- Chiang, J.O.; Marciel de Souza, W.; Teixeira Nunes, M.R.; Acrani, G.O.; Paes de Andrade Travassos da Rosa, A.; Mesquita de Freitas, N.; Patroca da Silva, S.; Dorta da Silva, P.H.; Watanabe de Sousa, A.; Rodrigues, S.G.; et al. Characterization of the Gamboa Virus Serogroup (Orthobunyavirus Genus, Peribunyaviridae Family). Am. J. Trop. Med. Hyg. 2018, 98, 1502–1511. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.M. Nucleotide sequence analysis of the small (S) RNA segment of Bunyamwera virus, the prototype of the family Bunyaviridae. J. Gen. Virol. 1989, 70, 1281–1285. [Google Scholar] [CrossRef] [PubMed]
- Lambert, A.J.; Lanciotti, R.S. Molecular characterization of medically important viruses of the genus Orthobunyavirus. J. Gen. Virol. 2008, 89, 2580–2585. [Google Scholar] [CrossRef] [PubMed]
- Tilston-Lunel, N.L.; Acrani, G.O.; Randall, R.E.; Elliott, R.M. Generation of Recombinant Oropouche Viruses Lacking the Nonstructural Protein NSm or NSs. J. Virol. 2015, 90, 2616–2627. [Google Scholar] [CrossRef] [PubMed]
- Pollitt, E.; Zhao, J.; Muscat, P.; Elliott, R.M. Characterization of Maguari orthobunyavirus mutants suggests the nonstructural protein NSm is not essential for growth in tissue culture. Virology 2006, 348, 224–232. [Google Scholar] [CrossRef]
- Hart, T.J.; Kohl, A.; Elliott, R.M. Role of the NSs protein in the zoonotic capacity of Orthobunyaviruses. Zoonoses Public Health 2009, 56, 285–296. [Google Scholar] [CrossRef]
- Szemiel, A.M.; Failloux, A.B.; Elliott, R.M. Role of Bunyamwera Orthobunyavirus NSs protein in infection of mosquito cells. Plos. Negl. Trop. Dis. 2012, 6, e1823. [Google Scholar] [CrossRef]
- de Souza, W.M.; Acrani, G.O.; Romeiro, M.F.; Reis, O., Jr.; Tolardo, A.L.; da Silva, S.P.; de Almeida Medeiros, D.B.; Varela, M.; Nunes, M.R.; Figueiredo, L.T. Molecular characterization of Capim and Enseada orthobunyaviruses. Infect. Genet. Evol. J. Mol. Epidemiol. Evol. Genet. Infect. Dis. 2016, 40, 47–53. [Google Scholar] [CrossRef]
- Firth, A.E.; Brierley, I. Non-canonical translation in RNA viruses. J. Gen. Virol. 2012, 93, 1385–1409. [Google Scholar] [CrossRef]
- Panicker, I.S.; Browning, G.F.; Markham, P.F. The Effect of an Alternate Start Codon on Heterologous Expression of a PhoA Fusion Protein in Mycoplasma gallisepticum. PLoS ONE 2015, 10, e0127911. [Google Scholar] [CrossRef] [PubMed]
- Barr, J.N.; Elliott, R.M.; Dunn, E.F.; Wertz, G.W. Segment-specific terminal sequences of Bunyamwera bunyavirus regulate genome replication. Virology 2003, 311, 326–338. [Google Scholar] [CrossRef]
- Barr, J.N.; Wertz, G.W. Role of the conserved nucleotide mismatch within 3′- and 5′-terminal regions of Bunyamwera virus in signaling transcription. J. Virol. 2005, 79, 3586–3594. [Google Scholar] [CrossRef] [PubMed]
- Mazel-Sanchez, B.; Elliott, R.M. Evolution of the Bunyamwera virus polymerase to accommodate deletions within genomic untranslated region sequences. J. Virol. 2015, 89, 3957–3964. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lowen, A.C.; Elliott, R.M. Mutational analyses of the nonconserved sequences in the Bunyamwera Orthobunyavirus S segment untranslated regions. J. Virol. 2005, 79, 12861–12870. [Google Scholar] [CrossRef]
- Chowdhary, R.; Street, C.; Travassos da Rosa, A.; Nunes, M.R.; Tee, K.K.; Hutchison, S.K.; Vasconcelos, P.F.; Tesh, R.B.; Lipkin, W.I.; Briese, T. Genetic characterization of the Wyeomyia group of orthobunyaviruses and their phylogenetic relationships. J. Gen. Virol. 2012, 93, 1023–1034. [Google Scholar] [CrossRef] [PubMed]
- Turell, M.J.; O’Guinn, M.L.; Jones, J.W.; Sardelis, M.R.; Dohm, D.J.; Watts, D.M.; Fernandez, R.; Travassos da Rosa, A.; Guzman, H.; Tesh, R.; et al. Isolation of viruses from mosquitoes (Diptera: Culicidae) collected in the Amazon Basin region of Peru. J. Med. Entomol. 2005, 42, 891–898. [Google Scholar] [CrossRef]
- Aitken, T.H.G.; Spence, L.; Jonkers, A.H.; Anderson, C.R. Wyeomyia-Virus Isolations in Trinidad, West Indies. Am. J. Trop. Med. Hyg. 1968, 17, 886–888. [Google Scholar] [CrossRef]
- de Souza Lopes, O.; de Abreu Sacchetta, L.; Fonseca, I.E.; Lacerda, J.P. Bertioga (Guama group) and Anhembi (Bunyamwera group), two new arboviruses isolated in Sao Paulo, Brazil. Am. J. Trop. Med. Hyg. 1975, 24, 131–134. [Google Scholar] [CrossRef]
- Srihongse, S.; Johnson, C.M. Wyeomyia Subgroup of Arbovirus: Isolation from Man. Science 1965, 149, 863–864. [Google Scholar] [CrossRef]
- Hacker, J.K.; Hardy, J.L. Adsorptive endocytosis of California encephalitis virus into mosquito and mammalian cells: A role for G1. Virology 1997, 235, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Plassmeyer, M.L.; Soldan, S.S.; Stachelek, K.M.; Roth, S.M.; Martin-Garcia, J.; Gonzalez-Scarano, F. Mutagenesis of the La Crosse Virus glycoprotein supports a role for Gc (1066–1087) as the fusion peptide. Virology 2007, 358, 273–282. [Google Scholar] [CrossRef] [PubMed]
- Hellert, J.; Aebischer, A.; Wernike, K.; Haouz, A.; Brocchi, E.; Reiche, S.; Guardado-Calvo, P.; Beer, M.; Rey, F.A. Orthobunyavirus spike architecture and recognition by neutralizing antibodies. Nat. Commun. 2019, 10, 879. [Google Scholar] [CrossRef] [PubMed]

| Virus | Strain | Mosquito Species (Similarity COI in %) | Collection Site |
|---|---|---|---|
| Baakal virus | C593-MX-2008 | Culex nigripalpus (99.67%) | Rainforest |
| Wyeomyia virus Palenque | B549-MX-2008 | Culicinae sp., Tribe Sabethini | Edge |
| Lakamha virus | C559-MX-2008 | Wyeomyia complosa (98.17%) | Edge |
| Virus | Number of Reads/Sample | Segment Size (nt) | Genome Coverage (nt) | Coverage (%) | |
|---|---|---|---|---|---|
| Baakal virus | Total | 1979 | |||
| Virus specific | 59 | ||||
| L segment | 21 | 6937 | 1986 | 28.6 | |
| M segment | 34 | 4947 | 2322 | 46.9 | |
| S segment | 4 | 1166 | 301 | 25.8 | |
| Lakamha virus | Total | 1,146,244 | |||
| Virus specific | 1289 | ||||
| L segment | 547 | 7223 | 7082 | 98.0 | |
| M segment | 557 | 2623 | 2521 | 96.1 | |
| S segment | 185 | 1078 | 1053 | 97.7 |
| Orthobunyavirus | Pacuvirus | Uncl. | Shanga-virus | Uncl. | Herbevirus | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BKAV | KOOV | UMBV | WITV | CALV | GAMV | BUNV | CVV | WYOV | JCV | LACV | PACV | RPEV | TPPV | KHURV | SHUV | LAKV | HEBV | TAIV | KIBV | |
| BKAV | 62.82 | 52.74 | 49.19 | 40.76 | 43.28 | 33.19 | 33.62 | 35.74 | 40.51 | 39.24 | 25.70 | 26.51 | 26.21 | 18.60 | 18.82 | 15.94 | 14.68 | 16.33 | 14.68 | |
| KOOV | 75.46 | 54.66 | 46.34 | 43.16 | 44.02 | 31.91 | 33.62 | 33.62 | 38.14 | 39.83 | 26.05 | 27.73 | 28.99 | 18.60 | 18.15 | 13.55 | 15.08 | 17.13 | 16.67 | |
| UMBV | 68.27 | 70.35 | 44.35 | 43.04 | 43.88 | 35.59 | 36.44 | 41.53 | 37.55 | 39.24 | 25.83 | 25.83 | 25.42 | 19.67 | 18.37 | 15.42 | 14.96 | 17.00 | 16.93 | |
| WITV | 57.92 | 58.69 | 58.83 | 35.37 | 37.40 | 34.01 | 34.01 | 34.82 | 33.47 | 35.08 | 25.60 | 28.40 | 28.40 | 18.11 | 18.15 | 14.12 | 14.45 | 14.89 | 14.07 | |
| CALV | 59.72 | 59.89 | 59.73 | 58.14 | 85.29 | 34.89 | 35.32 | 34.89 | 35.86 | 37.13 | 26.03 | 25.21 | 24.38 | 20.25 | 16.49 | 17.53 | 15.87 | 17.13 | 18.25 | |
| GAMV | 59.63 | 60.09 | 58.54 | 57.22 | 93.56 | 35.74 | 34.89 | 35.74 | 35.86 | 35.86 | 26.45 | 25.21 | 25.62 | 19.42 | 16.84 | 16.33 | 14.29 | 16.33 | 17.06 | |
| BUNV | 47.62 | 47.27 | 47.01 | 46.78 | 49.49 | 49.05 | 90.56 | 63.52 | 44.44 | 43.16 | 32.63 | 29.66 | 30.93 | 18.67 | 15.66 | 17.60 | 18.33 | 19.20 | 16.73 | |
| CVV | 47.18 | 47.80 | 47.14 | 46.65 | 49.71 | 49.27 | 82.09 | 63.95 | 43.59 | 43.59 | 32.20 | 29.24 | 31.78 | 20.33 | 17.08 | 18.00 | 17.13 | 18.80 | 16.33 | |
| WYOV | 48.28 | 49.08 | 47.45 | 47.57 | 49.62 | 49.19 | 67.83 | 67.20 | 47.86 | 48.72 | 31.78 | 28.81 | 29.66 | 19.50 | 18.51 | 18.40 | 17.93 | 17.20 | 16.33 | |
| JCV | 50.48 | 50.92 | 51.51 | 50.50 | 54.44 | 54.71 | 55.08 | 54.46 | 54.99 | 82.13 | 27.31 | 28.99 | 25.63 | 18.60 | 17.67 | 17.13 | 16.27 | 17.13 | 14.29 | |
| LACV | 51.41 | 51.54 | 51.55 | 51.34 | 54.84 | 54.57 | 55.87 | 55.34 | 55.08 | 83.47 | 29.83 | 28.99 | 26.89 | 18.18 | 18.73 | 15.54 | 15.87 | 15.14 | 15.48 | |
| PACV | 39.96 | 40.14 | 40.22 | 40.18 | 42.28 | 41.86 | 42.80 | 43.02 | 43.67 | 43.53 | 43.31 | 64.63 | 55.10 | 17.70 | 15.59 | 17.46 | 16.60 | 15.08 | 15.42 | |
| RPEV | 40.18 | 40.63 | 40.26 | 40.48 | 41.48 | 40.81 | 43.68 | 43.59 | 44.15 | 43.36 | 43.75 | 69.68 | 55.92 | 16.46 | 15.59 | 17.06 | 18.18 | 16.27 | 14.62 | |
| TPPV | 39.29 | 40.26 | 40.38 | 40.97 | 42.26 | 42.07 | 43.50 | 43.50 | 43.48 | 44.26 | 44.35 | 69.06 | 68.09 | 14.88 | 15.99 | 17.93 | 17.46 | 15.94 | 17.46 | |
| KHURV | 29.13 | 29.73 | 29.04 | 28.50 | 29.28 | 28.96 | 30.90 | 30.78 | 31.22 | 30.20 | 30.33 | 30.95 | 31.30 | 31.04 | 12.54 | 17.36 | 16.60 | 15.04 | 16.60 | |
| SHUV | 20.63 | 21.16 | 21.21 | 21.38 | 21.19 | 21.22 | 20.58 | 20.66 | 20.88 | 20.74 | 21.13 | 20.41 | 20.12 | 19.90 | 21.53 | 14.91 | 15.05 | 13.31 | 13.62 | |
| LAKV | 23.69 | 24.57 | 24.44 | 23.98 | 25.44 | 25.64 | 25.74 | 25.70 | 25.65 | 25.52 | 26.37 | 25.78 | 25.87 | 25.48 | 28.36 | 21.12 | 23.43 | 22.27 | 23.43 | |
| HEBV | 24.57 | 24.51 | 23.93 | 23.74 | 24.52 | 24.41 | 24.91 | 24.83 | 24.75 | 23.97 | 24.44 | 24.10 | 24.10 | 24.49 | 27.03 | 20.20 | 30.83 | 66.22 | 72.57 | |
| TAIV | 24.31 | 23.97 | 24.02 | 23.75 | 24.33 | 24.18 | 25.04 | 25.52 | 25.08 | 24.10 | 24.57 | 23.91 | 23.87 | 24.46 | 27.63 | 20.16 | 30.85 | 79.31 | 60.89 | |
| KIBV | 24.43 | 24.21 | 24.10 | 24.06 | 24.57 | 24.38 | 24.68 | 25.36 | 24.88 | 23.90 | 24.30 | 23.91 | 24.35 | 24.74 | 27.12 | 19.77 | 30.45 | 80.02 | 78.61 | ✕ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kopp, A.; Hübner, A.; Zirkel, F.; Hobelsberger, D.; Estrada, A.; Jordan, I.; Gillespie, T.R.; Drosten, C.; Junglen, S. Detection of Two Highly Diverse Peribunyaviruses in Mosquitoes from Palenque, Mexico. Viruses 2019, 11, 832. https://doi.org/10.3390/v11090832
Kopp A, Hübner A, Zirkel F, Hobelsberger D, Estrada A, Jordan I, Gillespie TR, Drosten C, Junglen S. Detection of Two Highly Diverse Peribunyaviruses in Mosquitoes from Palenque, Mexico. Viruses. 2019; 11(9):832. https://doi.org/10.3390/v11090832
Chicago/Turabian StyleKopp, Anne, Alexandra Hübner, Florian Zirkel, Daniel Hobelsberger, Alejandro Estrada, Ingo Jordan, Thomas R. Gillespie, Christian Drosten, and Sandra Junglen. 2019. "Detection of Two Highly Diverse Peribunyaviruses in Mosquitoes from Palenque, Mexico" Viruses 11, no. 9: 832. https://doi.org/10.3390/v11090832
APA StyleKopp, A., Hübner, A., Zirkel, F., Hobelsberger, D., Estrada, A., Jordan, I., Gillespie, T. R., Drosten, C., & Junglen, S. (2019). Detection of Two Highly Diverse Peribunyaviruses in Mosquitoes from Palenque, Mexico. Viruses, 11(9), 832. https://doi.org/10.3390/v11090832





