A Fusion Peptide in the Spike Protein of MERS Coronavirus
Abstract
1. Introduction
2. Materials and Methods
2.1. Peptide Synthesis
2.2. Giant Unilamellar Vesicles (GUVs)
2.3. GUV Deformation Assay
2.4. Statistical Analysis
2.5. Cell Culture
2.6. Constructs and Mutagenesis
2.7. Syncytium Formation
2.8. Immunofluorescence
3. Results
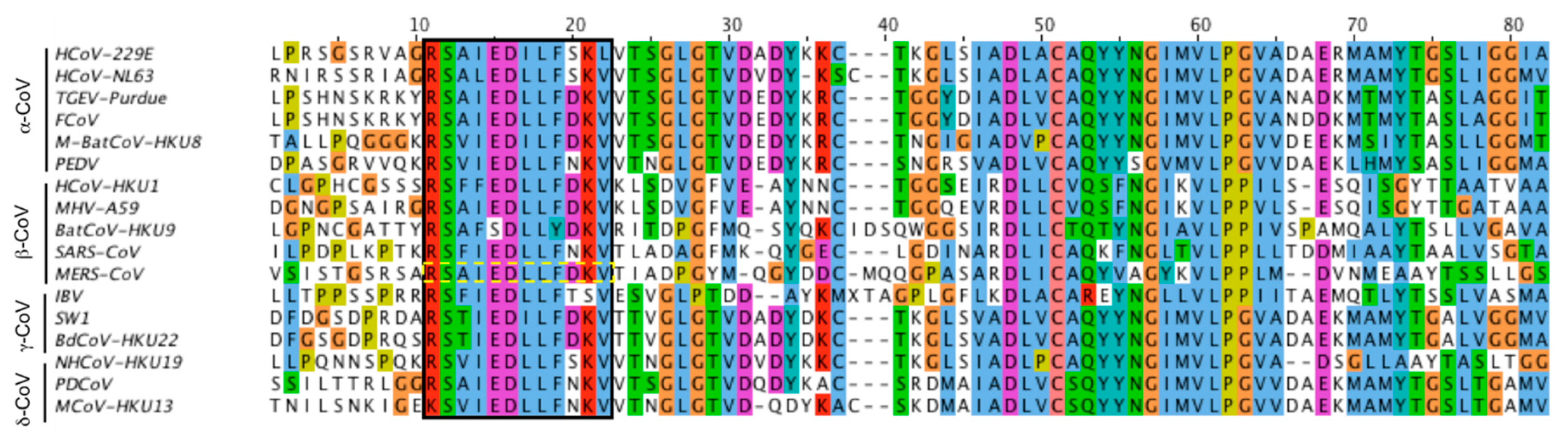
3.1. Identification of the Putative MERS-CoV Fusion Peptide
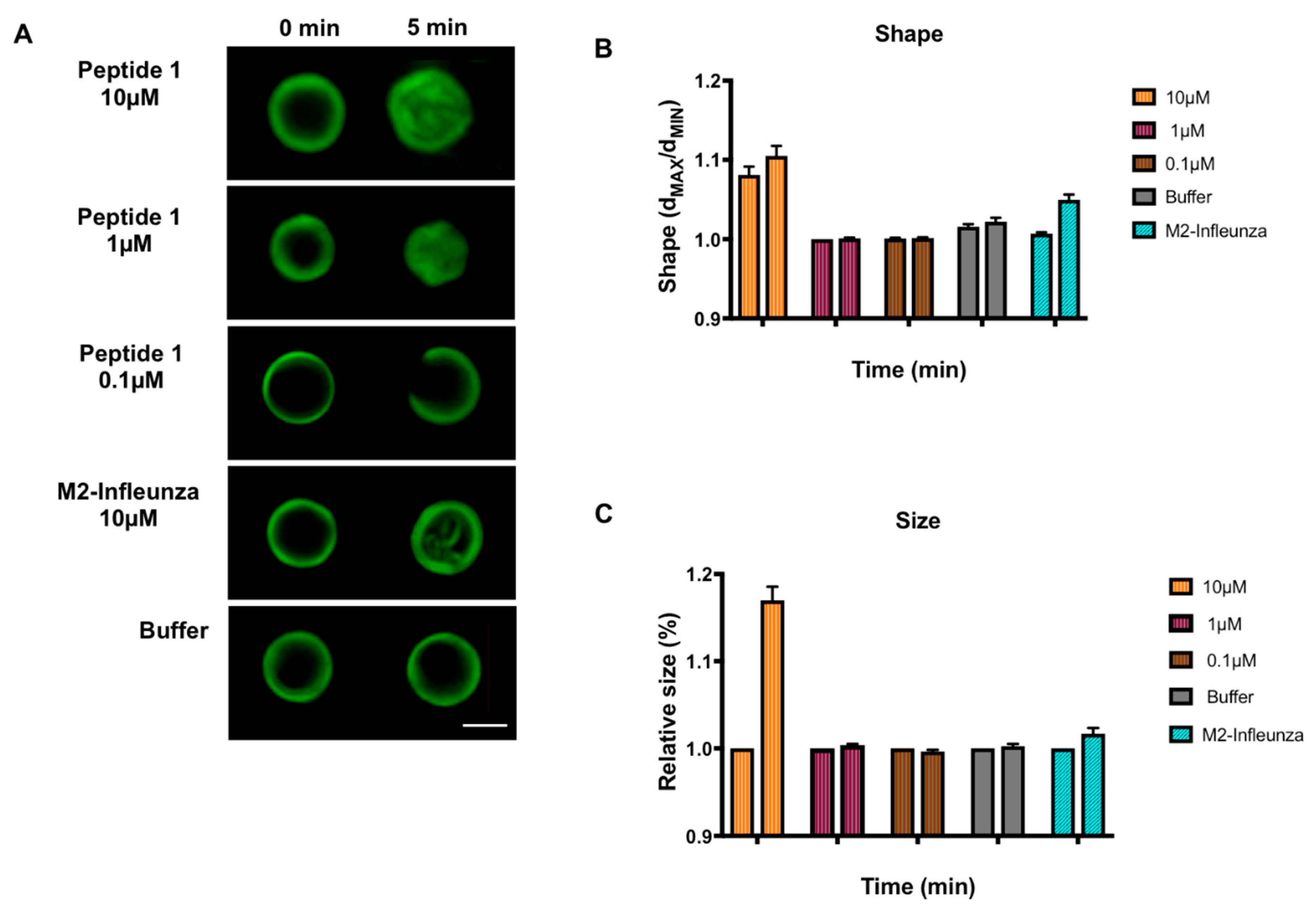
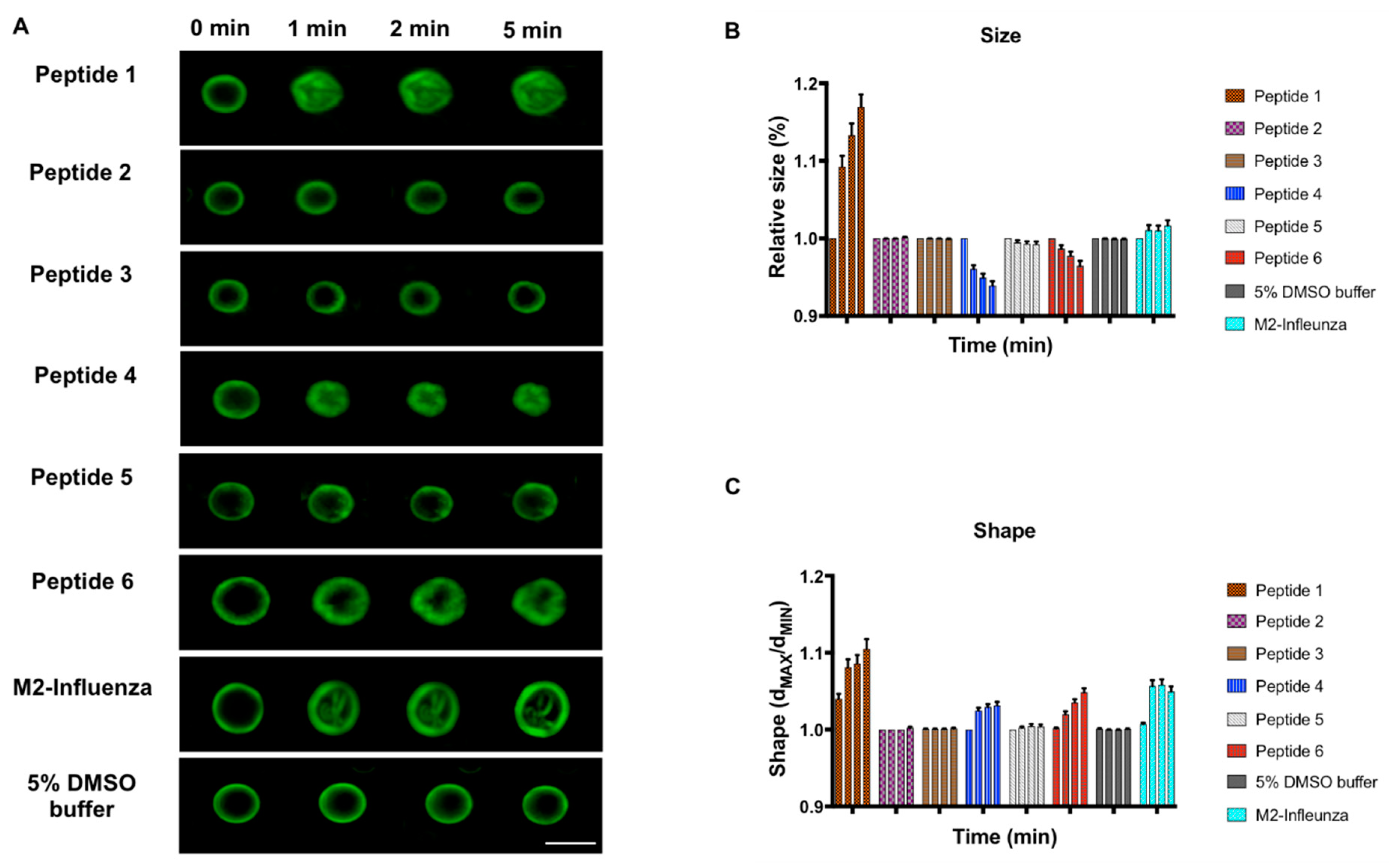
3.2. Effect of MERS-CoV Putative Fusion Peptide on Shape and Size of GUVs
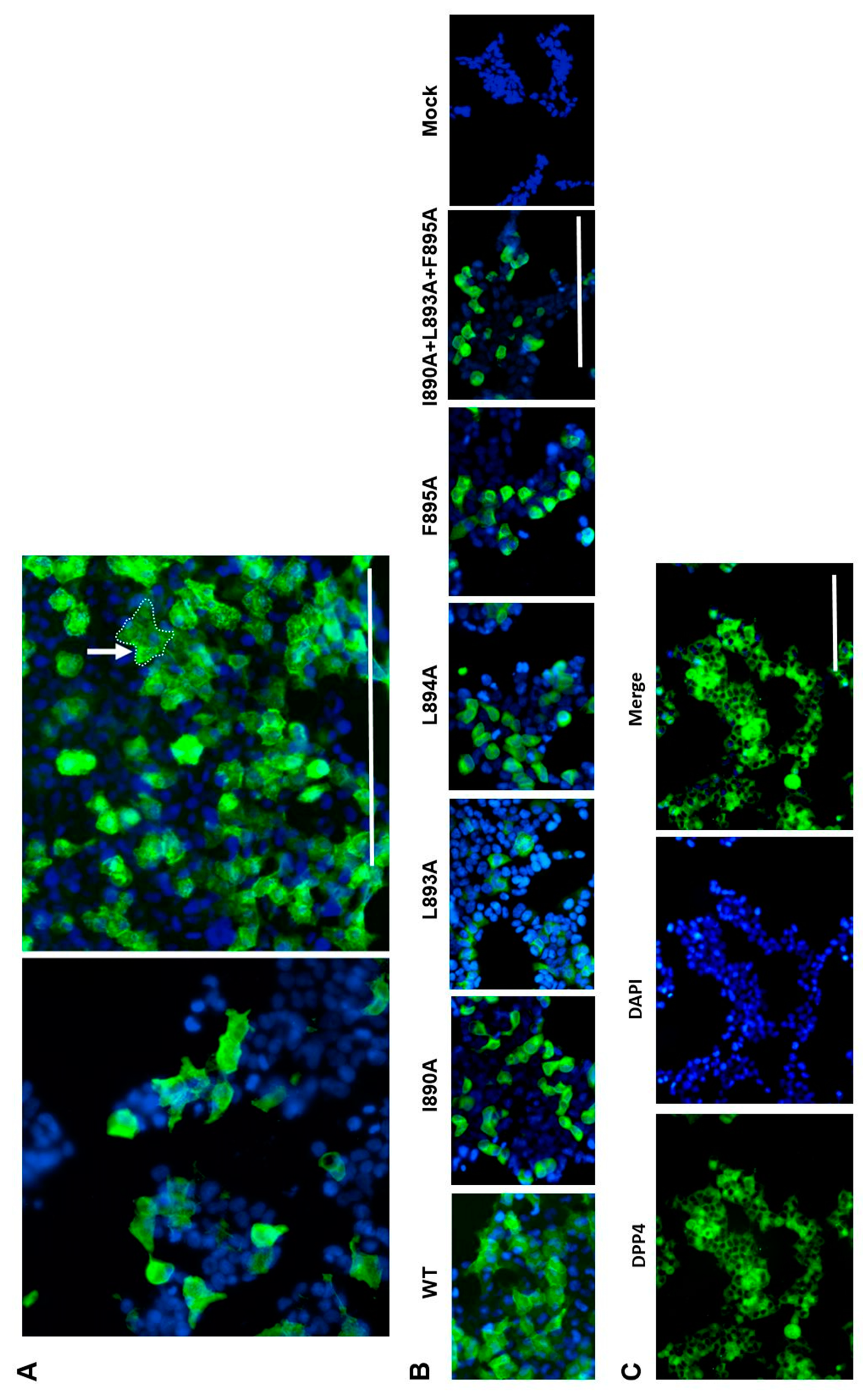
3.3. Role of Identified Peptides in Syncytium Formation
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Coleman, C.M.; Frieman, M.B. Coronaviruses: Important Emerging Human Pathogens. J. Virol. 2014, 88, 5209–5212. [Google Scholar] [CrossRef]
- Drosten, C. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M. Osterhaus AD FR. Isolation of a Novel Coronavirus from a Man with Pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Hu, B.; Ge, X.; Wang, L.F.; Shi, Z. Bat origin of human coronaviruses Coronaviruses: Emerging and re-emerging pathogens in humans and animals Susanna Lau Positive-strand RNA viruses. Virol. J. 2015, 12, 1–10. [Google Scholar]
- Bolles, M.; Donaldson, E.; Baric, R. SARS-CoV and emergent coronaviruses: Viral determinants of interspecies transmission. Curr. Opin. Virol. 2011, 1, 624–634. [Google Scholar] [CrossRef]
- Li, F. Receptor Recognition Mechanisms of Coronaviruses: A Decade of Structural Studies. J. Virol. 2015, 89, 1954–1964. [Google Scholar] [CrossRef]
- Van Boheemen, S.; de Graaf, M.; Lauber, C.; Bestebroer, T.M.; Raj, V.S.; Zaki, A.M.; Osterhaus, A.D.; Haagmans, B.L.; Gorbalenya, A.E.; Snijder, E.J.; et al. Genomic Characterization of Newly Discovered Coronavirus Associated with Acute Respiratory Distress Syndrome in Humans. MBio 2012, 3, e00473-12. [Google Scholar] [CrossRef]
- Raj, V.S.; Mou, H.; Smits, S.L.; Dekkers, D.H.; Müller, M.A.; Dijkman, R.; Muth, D.; Demmers, J.A.; Zaki, A.; Fouchier, R.A.; et al. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 2013, 495, 251–254. [Google Scholar] [CrossRef]
- Li, W.; Hulswit, R.J.; Widjaja, I.; Raj, V.S.; McBride, R.; Peng, W.; Widagdo, W.; Tortorici, M.A.; Van Dieren, B.; Lang, Y.; et al. Identification of sialic acid-binding function for the Middle East respiratory syndrome coronavirus spike glycoprotein. Proc. Natl. Acad. Sci. USA 2017, 114, E8508–E8517. [Google Scholar] [CrossRef]
- Yuan, Y.; Cao, D.; Zhang, Y.; Ma, J.; Qi, J.; Wang, Q.; Lu, G.; Wu, Y.; Yan, J.; Shi, Y.; et al. Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef]
- Shirato, K.; Kawase, M.; Matsuyama, S. Middle East Respiratory Syndrome Coronavirus Infection Mediated by the Transmembrane Serine Protease TMPRSS2. J. Virol. 2013, 87, 12552–12561. [Google Scholar] [CrossRef]
- Du, L.; Yang, Y.; Zhou, Y.; Lu, L.; Li, F.; Jiang, S. MERS-CoV spike protein: A key target for antivirals. Expert Opin. Ther. Targets 2017, 21, 131–143. [Google Scholar] [CrossRef]
- Earp, L.J.; Delos, S.E.; Park, H.E.; White, J.M.; Shirato, K.; Kawase, M. The many mechanisms of viral membrane fusion proteins. Curr. Top. Microbiol. Immunol. 2005, 285, 25–66. [Google Scholar]
- Bosch, B.J.; Bartelink, W.; Rottier, P.J.M. Cathepsin L functionally cleaves the severe acute respiratory syndrome coronavirus class I fusion protein upstream of rather than adjacent to the fusion peptide. J. Virol. 2008, 82, 8887–8890. [Google Scholar] [CrossRef]
- Belouzard, S.; Millet, J.K.; Licitra, B.N.; Whittaker, G.R. Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses 2012, 4, 1011–1033. [Google Scholar] [CrossRef]
- Madu, I.G.; Roth, S.L.; Belouzard, S.; Whittaker, G.R. Characterization of a Highly Conserved Domain within the Severe Acute Respiratory Syndrome Coronavirus Spike Protein S2 Domain with Characteristics of a Viral Fusion Peptide. J. Virol. 2009, 83, 7411–7421. [Google Scholar] [CrossRef]
- Lai, A.L.; Millet, J.K.; Daniel, S.; Freed, J.H.; Whittaker, G.R. The SARS-CoV Fusion Peptide Forms an Extended Bipartite Fusion Platform that Perturbs Membrane Order in a Calcium-Dependent Manner. J. Mol. Biol. 2017, 429, 3875–3892. [Google Scholar] [CrossRef]
- Koynova, R.; Caffrey, M. Phases and phase transitions of the phosphatidylcholines. Biochim. Biophys. Acta Rev. Biomembr. 1998, 1376, 91–145. [Google Scholar] [CrossRef]
- Rossman, J.S.; Jing, X.; Leser, G.P.; Lamb, R.A. Influenza Virus M2 Protein Mediates ESCRT-Independent Membrane Scission. Cell 2010, 142, 902–913. [Google Scholar] [CrossRef]
- Sapay, N.; Guermeur, Y.; Deléage, G. Prediction of amphipathic in-plane membrane anchors in monotopic proteins using a SVM classifier. BMC Bioinform. 2006, 7, 1–11. [Google Scholar] [CrossRef]
- Belouzard, S.; Chu, V.C.; Whittaker, G.R. Activation of the SARS coronavirus spike protein via sequential proteolytic cleavage at two distinct sites. Proc. Natl. Acad. Sci. USA 2009, 106, 5871–5876. [Google Scholar] [CrossRef]
- Millet, J.K.; Goldstein, M.E.; Labitt, R.N.; Hsu, H.L.; Daniel, S.; Whittaker, G.R. A camel-derived MERS-CoV with a variant spike protein cleavage site and distinct fusion activation properties. Emerg. Microbes Infect. 2016, 5, e126–e129. [Google Scholar] [CrossRef]
- Takahashi, T.; Nomura, F.; Yokoyama, Y.; Tanaka-Takiguchi, Y.; Homma, M.; Takiguchi, K. Multiple membrane interactions and versatile vesicle deformations elicited by melittin. Toxins 2013, 5, 637–664. [Google Scholar] [CrossRef]
- Yamada, Y.; Liu, D.X. Proteolytic Activation of the Spike Protein at a Novel RRRR/S Motif Is Implicated in Furin-Dependent Entry, Syncytium Formation, and Infectivity of Coronavirus Infectious Bronchitis Virus in Cultured Cells. J. Virol. 2009, 83, 8744–8758. [Google Scholar] [CrossRef]
- Volz, A.; Kupke, A.; Song, F.; Jany, S.; Fux, R.; Shams-Eldin, H.; Schmidt, J.; Becker, C.; Eickmann, M.; Becker, S.; et al. Protective Efficacy of Recombinant Modified Vaccinia Virus Ankara Delivering Middle East Respiratory Syndrome Coronavirus Spike Glycoprotein. J. Virol. 2015, 89, 8651–8656. [Google Scholar] [CrossRef]
- Jiaming, L.; Yanfeng, Y.; Yao, D.; Yawei, H.; Linlin, B.; Baoying, H.; Jinghua, Y.; Gao, G.F.; Chuan, Q.; Wenjie, T. The recombinant N-terminal domain of spike proteins is a potential vaccine against Middle East respiratory syndrome coronavirus (MERS-CoV) infection. Vaccine 2017, 35, 10–18. [Google Scholar] [CrossRef]
- Channappanavar, R.; Lu, L.; Xia, S.; Du, L.; Meyerholz, D.K.; Perlman, S.; Jiang, S. Protective effect of intranasal regimens containing peptidic middle east respiratory syndrome coronavirus fusion inhibitor against MERS-CoV infection. J. Infect. Dis. 2015, 212, 1894–1903. [Google Scholar] [CrossRef]
- Lee, M.C.S.; Orci, L.; Hamamoto, S.; Futai, E.; Ravazzola, M.; Schekman, R. Sar1p N-terminal helix initiates membrane curvature and completes the fission of a COPII vesicle. Cell 2005, 122, 605–617. [Google Scholar] [CrossRef]
| No. | Designation | Residues | Sequence |
|---|---|---|---|
| 1 | Peptide 1 (wild type) | 884–898 | RSARSAIEDLLFDKV |
| 2 | Peptide 2 | I890A | RSARSAAEDLLFDKV |
| 3 | Peptide 3 | L893A | RSARSAIEDALFDKV |
| 4 | Peptide 4 | L894A | RSARSAIEDLAFDKV |
| 5 | Peptide 5 | F895A | RSARSAIEDLLADKV |
| 6 | Peptide 6 | V898A | RSARSAIEDLLFDKA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alsaadi, E.A.J.; Neuman, B.W.; Jones, I.M. A Fusion Peptide in the Spike Protein of MERS Coronavirus. Viruses 2019, 11, 825. https://doi.org/10.3390/v11090825
Alsaadi EAJ, Neuman BW, Jones IM. A Fusion Peptide in the Spike Protein of MERS Coronavirus. Viruses. 2019; 11(9):825. https://doi.org/10.3390/v11090825
Chicago/Turabian StyleAlsaadi, Entedar A. J., Benjamin W. Neuman, and Ian M. Jones. 2019. "A Fusion Peptide in the Spike Protein of MERS Coronavirus" Viruses 11, no. 9: 825. https://doi.org/10.3390/v11090825
APA StyleAlsaadi, E. A. J., Neuman, B. W., & Jones, I. M. (2019). A Fusion Peptide in the Spike Protein of MERS Coronavirus. Viruses, 11(9), 825. https://doi.org/10.3390/v11090825






