RNASeq Analysis of Aedes albopictus Mosquito Midguts after Chikungunya Virus Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Chikungunya Virus
2.2. Aedes Mosquito Rearing, Infection and RNA Extraction
2.3. qPCR
2.4. RNASeq and Viral Genome Sequencing
2.5. Differential Gene Expression and Gene Ontology Analysis
3. Results
3.1. Whole Genome Sequencing of CHIKV
3.2. RNASeq
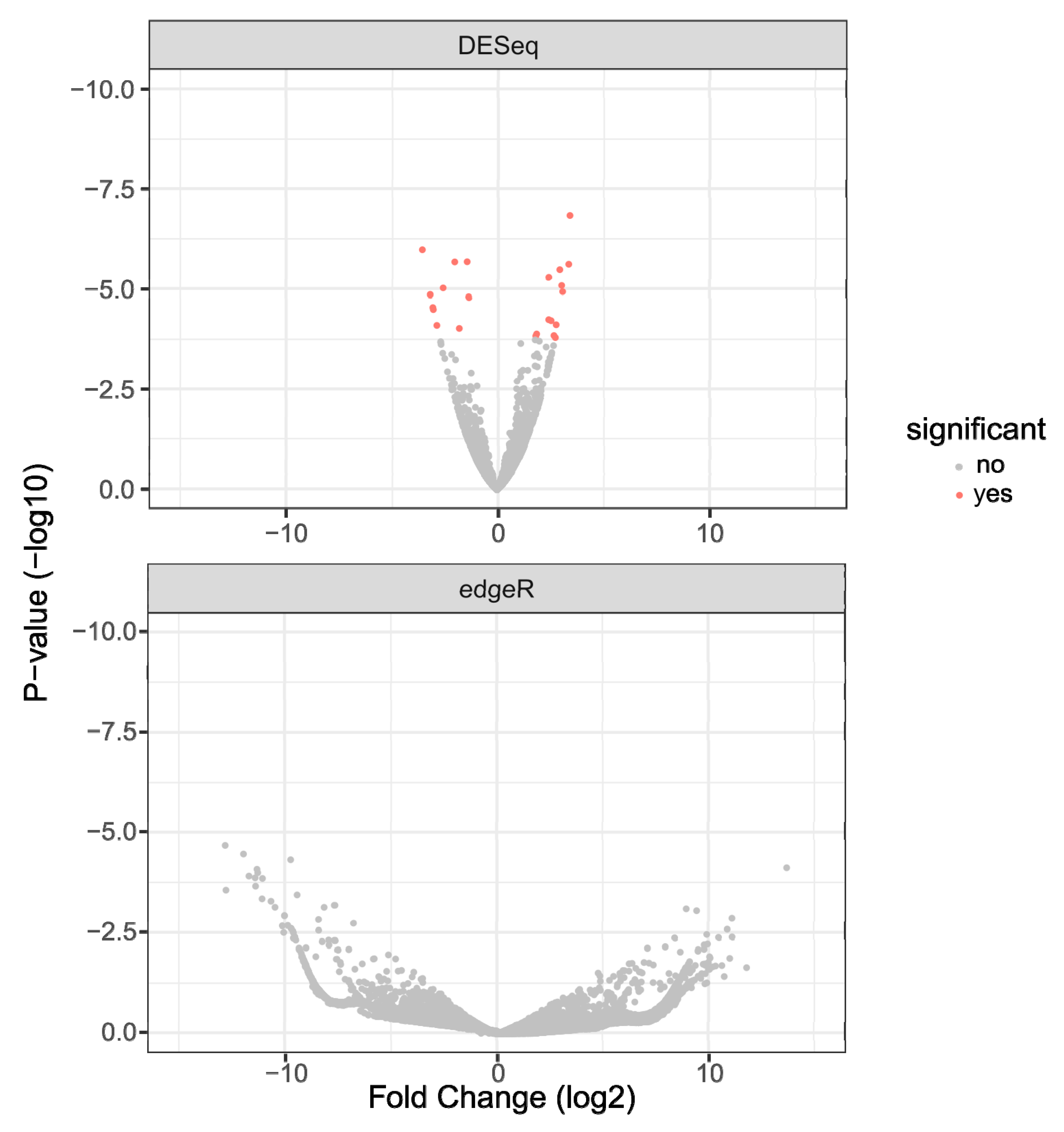
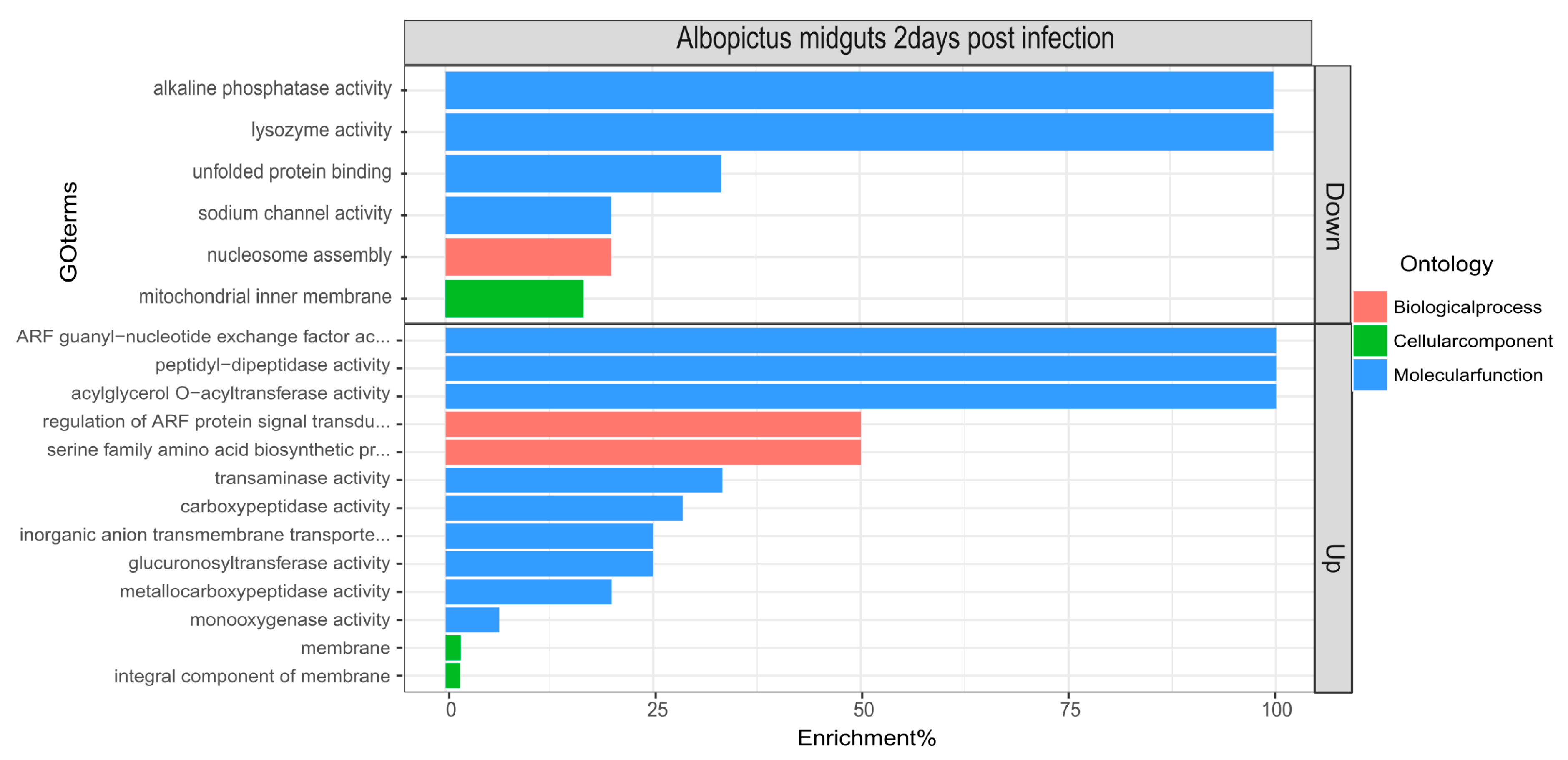
3.3. Differential Expression and TopGO Analysis
3.4. RNASeq Data Validation on qRT-PCR
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504. [Google Scholar] [CrossRef]
- Lumsden, W.H. An epidemic of virus disease in Southern Province, Tanganyika Territory, in 1952-53. II. General description and epidemiology. Trans. R. Soc. Trop. Med. Hyg. 1955, 49, 33–57. [Google Scholar] [CrossRef]
- Robinson, M.C. An epidemic of virus disease in Southern Province, Tanganyika Territory, in 1952-53. I. Clinical features. Trans. R. Soc. Trop. Med. Hyg. 1955, 49, 28–32. [Google Scholar] [CrossRef]
- Kamath, S.; Das, A.K.; Parikh, F.S. Chikungunya. J. Assoc. Physicians India 2006, 54, 725–726. [Google Scholar] [PubMed]
- Higgs, S.; Vanlandingham, D. Chikungunya virus and its mosquito vectors. Vector Borne Zoonotic Dis. 2015, 15, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Simon, F.; Javelle, E.; Oliver, M.; Leparc-Goffart, I.; Marimoutou, C. Chikungunya Virus Infection. Curr. Infect. Dis. Rep. 2011, 13, 218–228. [Google Scholar] [CrossRef] [PubMed]
- Reiter, P.; Fontenille, D.; Paupy, C. Aedes albopictus as an epidemic vector of chikungunya virus: Another emerging problem? Lancet Infect. Dis. 2006, 6, 463–464. [Google Scholar] [CrossRef]
- Tsetsarkin, K.A.; Chen, R.; Weaver, S.C. Interspecies transmission and chikungunya virus emergence. Curr. Opin. Virol. 2016, 16, 143–150. [Google Scholar] [CrossRef]
- Waldock, J.; Chandra, N.L.; Lelieveld, J.; Proestos, Y.; Michael, E.; Christophides, G.; Parham, P.E. The role of environmental variables on Aedes albopictus biology and chikungunya epidemiology. Pathog. Glob. Health 2013, 107, 224–241. [Google Scholar] [CrossRef]
- Rezza, G.; Nicoletti, L.; Angelini, R.; Romi, R.; Finarelli, A.C.; Panning, M.; Cordioli, P.; Fortuna, C.; Boros, S.; Magurano, F.; et al. Infection with chikungunya virus in Italy: An outbreak in a temperate region. Lancet 2007, 370, 1840–1846. [Google Scholar] [CrossRef]
- Enserink, M. Chikungunya: No Longer a Third World Disease. Science 2007, 318, 1860. [Google Scholar] [CrossRef] [PubMed]
- Jupatanakul, N.; Sim, S.; Dimopoulos, G. The insect microbiome modulates vector competence for arboviruses. Viruses 2014, 6, 4294–4313. [Google Scholar] [CrossRef] [PubMed]
- Arias-Goeta, C.; Mousson, L.; Rougeon, F.; Failloux, A.-B. Dissemination and Transmission of the E1-226V Variant of Chikungunya Virus in Aedes albopictus Are Controlled at the Midgut Barrier Level. PLoS ONE 2013, 8, e57548. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Kantor, A.M.; Lin, J.; Passarelli, A.L.; Clem, R.J.; Franz, A.W.E. Infection pattern and transmission potential of chikungunya virus in two New World laboratory-adapted Aedes aegypti strains. Sci. Rep. 2016, 6, 24729. [Google Scholar] [CrossRef] [PubMed]
- Dubrulle, M.; Mousson, L.; Moutailler, S.; Vazeille, M.; Failloux, A.-B. Chikungunya Virus and Aedes Mosquitoes: Saliva Is Infectious as soon as Two Days after Oral Infection. PLoS ONE 2009, 4, e5895. [Google Scholar] [CrossRef] [PubMed]
- Franz, A.W.E.; Kantor, A.M.; Passarelli, A.L.; Clem, R.J. Tissue Barriers to Arbovirus Infection in Mosquitoes. Viruses 2015, 7, 3741–3767. [Google Scholar] [CrossRef] [PubMed]
- Kuno, G.; Chang, G.-J.J. Biological Transmission of Arboviruses: Reexamination of and New Insights into Components, Mechanisms, and Unique Traits as Well as Their Evolutionary Trends. Clin. Microbiol. Rev. 2005, 18, 608–637. [Google Scholar] [CrossRef] [PubMed]
- Chompoosri, J.; Thavara, U.; Tawatsin, A.; Boonserm, R.; Phumee, A.; Sangkitporn, S.; Siriyasatien, P. Vertical transmission of Indian Ocean Lineage of chikungunya virus in Aedes aegypti and Aedes albopictus mosquitoes. Parasites Vectors 2016, 9, 227. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Behura, S.K.; Franz, A.W.E. The midgut transcriptome of Aedes aegypti fed with saline or protein meals containing chikungunya virus reveals genes potentially involved in viral midgut escape. BMC Genom. 2017, 18, 382. [Google Scholar] [CrossRef] [PubMed]
- Etebari, K.; Hegde, S.; Saldaña, M.A.; Widen, S.G.; Wood, T.G.; Asgari, S.; Hughes, G.L. Global Transcriptome Analysis of Aedes aegypti Mosquitoes in Response to Zika Virus Infection. mSphere 2017, 2, e00456-00417. [Google Scholar] [CrossRef] [PubMed]
- Shrinet, J.; Srivastava, P.; Sunil, S. Transcriptome analysis of Aedes aegypti in response to mono-infections and co-infections of dengue virus-2 and chikungunya virus. Biochem. Biophys. Res. Commun. 2017, 492, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Bonizzoni, M.; Dunn, W.A.; Campbell, C.L.; Olson, K.E.; Marinotti, O.; James, A.A. Complex Modulation of the Aedes aegypti Transcriptome in Response to Dengue Virus Infection. PLoS ONE 2012, 7, e50512. [Google Scholar] [CrossRef] [PubMed]
- Fragkoudis, R.; Chi, Y.; Siu, R.W.; Barry, G.; Attarzadeh-Yazdi, G.; Merits, A.; Nash, A.A.; Fazakerley, J.K.; Kohl, A. Semliki Forest virus strongly reduces mosquito host defence signaling. Insect Mol. Biol. 2008, 17, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhou, Y.; Wu, J.; Zheng, P.; Li, Y.; Zheng, X.; Puthiyakunnon, S.; Tu, Z.; Chen, X.G. The expression profile of Aedes albopictus miRNAs is altered by dengue virus serotype-2 infection. Cell Biosci. 2015, 5, 16. [Google Scholar] [CrossRef] [PubMed]
- Pyke, A.T. Partial E1 gene Sequence of Mauritius 2006 isolate 06113879. Unpublished work. 2008. [Google Scholar]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357. [Google Scholar] [CrossRef]
- Lawson, D.; Arensburger, P.; Atkinson, P.; Besansky, N.J.; Bruggner, R.V.; Butler, R.; Campbell, K.S.; Christophides, G.K.; Christley, S.; Dialynas, E.; et al. VectorBase: A data resource for invertebrate vector genomics. Nucleic Acids Res. 2009, 37, D583–D587. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Afgan, E.; Sloggett, C.; Goonasekera, N.; Makunin, I.; Benson, D.; Crowe, M.; Gladman, S.; Kowsar, Y.; Pheasant, M.; Horst, R.; et al. Genomics Virtual Laboratory: A Practical Bioinformatics Workbench for the Cloud. PLoS ONE 2015, 10, e0140829. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644. [Google Scholar] [CrossRef]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-Seq: Reference generation and analysis with Trinity. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Boratyn, G.M.; Schäffer, A.A.; Agarwala, R.; Altschul, S.F.; Lipman, D.J.; Madden, T.L. Domain enhanced lookup time accelerated BLAST. Biol. Direct 2012, 7, 12. [Google Scholar] [CrossRef] [PubMed]
- Alexa, A.; Rahnenfuhrer, J. topGO: Enrichment Analysis for Gene Ontology, R package Version 2.33.0; 2016. Available online: https://bioconductor.org/packages/release/bioc/html/topGO.html (accessed on 6 May 2018).
- Smedley, D.; Haider, S.; Durinck, S.; Pandini, L.; Provero, P.; Allen, J.; Arnaiz, O.; Awedh, M.H.; Baldock, R.; Barbiera, G.; et al. The BioMart community portal: An innovative alternative to large, centralized data repositories. Nucleic Acids Res. 2015, 43, W589–W598. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, M.; Arias-Goeta, C.; Martin, E.; O’Hara, Z.; Lulla, A.; Mousson, L.; Rainey, S.M.; Misbah, S.; Schnettler, E.; Donald, C.L.; et al. Characterization of Aedes aegypti innate-immune pathways that limit Chikungunya virus replication. PLoS Negl. Trop. Dis. 2014, 8, e2994. [Google Scholar] [CrossRef] [PubMed]
- Colpitts, T.M.; Cox, J.; Vanlandingham, D.L.; Feitosa, F.M.; Cheng, G.; Kurscheid, S.; Wang, P.; Krishnan, M.N.; Higgs, S.; Fikrig, E. Alterations in the Aedes aegypti Transcriptome during Infection with West Nile, Dengue and Yellow Fever Viruses. PLoS Pathog. 2011, 7, e1002189. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.; Koch, J.; Rakheja, D.; Pattnaik, A.K.; Brugarolas, J.; Dozmorov, I.; Levine, B.; Wakeland, E.K.; Lee-Kirsch, M.A.; Yan, N. Trex1 regulates lysosomal biogenesis and interferon-independent activation of antiviral genes. Nat. Immunol. 2013, 14, 61–71. [Google Scholar] [CrossRef]
- Isoe, J.; Zamora, J.; Miesfeld, R.L. Molecular Analysis of the Aedes aegypti Carboxypeptidase Gene Family. Insect Biochem. Mol. Biol. 2009, 39, 68–73. [Google Scholar] [CrossRef]
- Johnston, C.; Jiang, W.; Chu, T.; Levine, B. Identification of Genes Involved in the Host Response to Neurovirulent Alphavirus Infection. J. Virol. 2001, 75, 10431–10445. [Google Scholar] [CrossRef]
- Merkling, S.H.; van Rij, R.P. Beyond RNAi: Antiviral defense strategies in Drosophila and mosquito. J. Insect Physiol. 2013, 59, 159–170. [Google Scholar] [CrossRef]
- Swevers, L.; Liu, J.; Smagghe, G. Defense Mechanisms against Viral Infection in Drosophila: RNAi and Non-RNAi. Viruses 2018, 10, 230. [Google Scholar] [CrossRef] [PubMed]
- Vance, J.E.; Karten, B. Niemann-Pick C disease and mobilization of lysosomal cholesterol by cyclodextrin. J. Lipid Res. 2014, 55, 1609–1621. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.H.; Twu, Y.C.; Wang, C.K.; Lin, F.Z.; Lee, C.Y.; Liao, Y.J. Niemann-Pick Type C2 Protein Regulates Free Cholesterol Accumulation and Influences Hepatic Stellate Cell Proliferation and Mitochondrial Respiration Function. Int. J. Mol. Sci. 2018, 19, 1678. [Google Scholar] [CrossRef] [PubMed]
- Jupatanakul, N.; Sim, S.; Dimopoulos, G. Aedes aegypti ML and Niemann-Pick type C family members are agonists of dengue virus infection. Dev. Comp. Immunol. 2014, 43, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.S.; Davidson, C.; Kuehne, A.I.; Bakken, R.; Braigen, S.Z.; Gunn, K.E.; Whelan, S.P.; Brummelkamp, T.R.; Twenhafel, N.A.; Chandran, K.; et al. Niemann-pick C1 is essential for ebolavirus replication and pathogenesis in vivo. mBio 2015, 6, e00565-00515. [Google Scholar] [CrossRef] [PubMed]
- Infante, R.E.; Wang, M.L.; Radhakrishnan, A.; Kwon, H.J.; Brown, M.S.; Goldstein, J.L. NPC2 facilitates bidirectional transfer of cholesterol between NPC1 and lipid bilayers, a step in cholesterol egress from lysosomes. Proc. Natl. Acad. Sci. 2008, 105, 15287–15292. [Google Scholar] [CrossRef]
- Amini-Bavil-Olyaee, S.; Choi, Y.J.; Lee, J.H.; Shi, M.; Huang, I.C.; Farzan, M.; Jung, J.U. The antiviral effector IFITM3 disrupts intracellular cholesterol homeostasis to block viral entry. Cell Host Microbe 2013, 13, 452–464. [Google Scholar] [CrossRef] [PubMed]
- Wichit, S.; Hamel, R.; Bernard, E.; Talignani, L.; Diop, F.; Ferraris, P.; Liegeois, F.; Ekchariyawat, P.; Luplertlop, N.; Surasombatpattana, P.; et al. Imipramine Inhibits Chikungunya Virus Replication in Human Skin Fibroblasts through Interference with Intracellular Cholesterol Trafficking. Sci. Rep. 2017, 7, 3145. [Google Scholar] [CrossRef] [PubMed]
- Cirimotich, C.M.; Scott, J.C.; Phillips, A.T.; Geiss, B.J.; Olson, K.E. Suppression of RNA interference increases alphavirus replication and virus-associated mortality in Aedes aegypti mosquitoes. BMC Microbiol. 2009, 9, 49. [Google Scholar] [CrossRef] [PubMed]
- Saleh, M.C.; Tassetto, M.; van Rij, R.P.; Goic, B.; Gausson, V.; Berry, B.; Jacquier, C.; Antoniewski, C.; Andino, R. Antiviral immunity in Drosophila requires systemic RNA interference spread. Nature 2009, 458, 346–350. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, Y.; Zhou, R.; Zhao, J.; Zhang, Y.; Yi, D.; Li, Q.; Zhou, J.; Guo, F.; Liang, C.; et al. Host Long Noncoding RNA lncRNA-PAAN Regulates the Replication of Influenza A Virus. Viruses 2018, 10, 330. [Google Scholar] [CrossRef] [PubMed]
- Etebari, K.; Asad, S.; Zhang, G.; Asgari, S. Identification of Aedes aegypti Long Intergenic Non-coding RNAs and Their Association with Wolbachia and Dengue Virus Infection. PLoS Negl. Trop. Dis. 2016, 10, e0005069. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.J.; Jiang, S.C.; Wei, H.X.; Deng, S.Q.; He, C.; Peng, H.J. The Differential Expression and Possible Function of Long Noncoding RNAs in Liver Cells Infected by Dengue Virus. Am. J. Trop. Med. Hyg. 2017, 97, 1904–1912. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Ding, C. Roles of LncRNAs in Viral Infections. Front. Cell. Infect. Microbiol. 2017, 7, 205. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.; Wang, T.; Tang, Q.; Li, G.; Wu, P.; Chen, K. Long Non-coding RNAs: Regulators of Viral Infection and the Interferon Antiviral Response. Front. Microbiol. 2018, 9, 1621. [Google Scholar] [CrossRef] [PubMed]
- Du, M.; Yuan, L.; Tan, X.; Huang, D.; Wang, X.; Zheng, Z.; Mao, X.; Li, X.; Yang, L.; Huang, K.; et al. The LPS-inducible lncRNA Mirt2 is a negative regulator of inflammation. Nat. Commun. 2017, 8, 2049. [Google Scholar] [CrossRef] [PubMed]
- Nikhil, S.; Singh, S. Implications of non-coding RNAs in viral infections. Rev. Med. Virol. 2016, 26, 356–368. [Google Scholar] [CrossRef]
- Valadkhan, S.; Gunawardane, L.S. lncRNA-mediated regulation of the interferon response. Virus Res. 2016, 212, 127–136. [Google Scholar] [CrossRef]
| Total Reads (millions) | % Mapped to RefSeq Genome | % of CHIKV Reads | |
|---|---|---|---|
| Infected MG 1 | 170.65 | 60.83% | 0.01% |
| Infected MG 2 | 37.13 | 60.46% | 0.02% |
| Infected MG 3 | 38.02 | 65.28% | 0.10% |
| Control MG1 | 44.77 | 61.53% | 0.00% |
| Control MG2 | 44.72 | 63.15% | 0.00% |
| Genes | Aedes albopictus | Aedes aegypti | |
|---|---|---|---|
| LogFC (RNAseq) | Expression Fold Change (qRT-PCR) | Expression Fold Change (qRT-PCR) | |
| ARF GTPase-activating protein GIT2 | −8.7 | 0.76(↓) | 39.74(↑) |
| NPC2 homolog | 6.29 | 5.35(↑) | 11.63(↑) |
| Mucin-22/FLO-11-like | −8.61 | 0.16(↓) | 7.34(↑) |
| Translocon-associated protein subunit delta | 8.82 | 1.88(↑) | 40.18(↑) |
| ATP-dependent RNA helicase dbp2 | −9.07 | 6.53(↑) | 26.22(↑) |
| Uncharacterized gene coding for Sina and RING_Ubox domains containing protein | 8.84 | 1.51(↑) | 41.07(↑) |
| E3 ubiquitin-protein ligase MARCH6 | −9.04 | 1.03(↑) | 28.65(↑) |
| Ankyrin repeat domain-containing protein 44 | −5.56 | 0.5(↓) | No homologue/Not tested |
| PREDICTED: Aedes albopictus uncharacterized LOC109424229 (LOC109424229), ncRNA | −11.22 | 2.14(↑) | No homologue/Not tested |
| PREDICTED: Aedes albopictus uncharacterized LOC109622934 (LOC109622934), ncRNA | −7.13 | 0.06(↓) | No homologue/Not tested |
| PREDICTED: Aedes albopictus uncharacterized LOC109423409 (LOC109423409), ncRNA | −9.07 | 0.73(↓) | No homologue/Not tested |
| PREDICTED: Aedes albopictus uncharacterized LOC109414360 (LOC109414360), ncRNA | −9.53 | 1.02(↑) | No homologue/Not tested |
| PREDICTED: Aedes albopictus uncharacterized LOC109424229 (LOC109424229), ncRNA | −8.98 | 0.04(↓) | No homologue/Not tested |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vedururu, R.k.; Neave, M.J.; Tachedjian, M.; Klein, M.J.; Gorry, P.R.; Duchemin, J.-B.; Paradkar, P.N. RNASeq Analysis of Aedes albopictus Mosquito Midguts after Chikungunya Virus Infection. Viruses 2019, 11, 513. https://doi.org/10.3390/v11060513
Vedururu Rk, Neave MJ, Tachedjian M, Klein MJ, Gorry PR, Duchemin J-B, Paradkar PN. RNASeq Analysis of Aedes albopictus Mosquito Midguts after Chikungunya Virus Infection. Viruses. 2019; 11(6):513. https://doi.org/10.3390/v11060513
Chicago/Turabian StyleVedururu, Ravi kiran, Matthew J. Neave, Mary Tachedjian, Melissa J. Klein, Paul R. Gorry, Jean-Bernard Duchemin, and Prasad N. Paradkar. 2019. "RNASeq Analysis of Aedes albopictus Mosquito Midguts after Chikungunya Virus Infection" Viruses 11, no. 6: 513. https://doi.org/10.3390/v11060513
APA StyleVedururu, R. k., Neave, M. J., Tachedjian, M., Klein, M. J., Gorry, P. R., Duchemin, J.-B., & Paradkar, P. N. (2019). RNASeq Analysis of Aedes albopictus Mosquito Midguts after Chikungunya Virus Infection. Viruses, 11(6), 513. https://doi.org/10.3390/v11060513






