Parvovirus B19 Uncoating Occurs in the Cytoplasm without Capsid Disassembly and It Is Facilitated by Depletion of Capsid-Associated Divalent Cations
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Antibodies and Chemicals
2.3. Quantitative PCR
2.4. Iodixanol Density Gradient Ultracentrifugation
2.5. Transmission Electron Microscopy
2.6. Nuclease Assay
2.7. Isolation of Cytoplasmic and Nuclear Fractions from Infected Cells
2.8. Immunoprecipitation
2.9. Infectivity Assay
2.10. Complementary-Strand Synthesis
3. Results
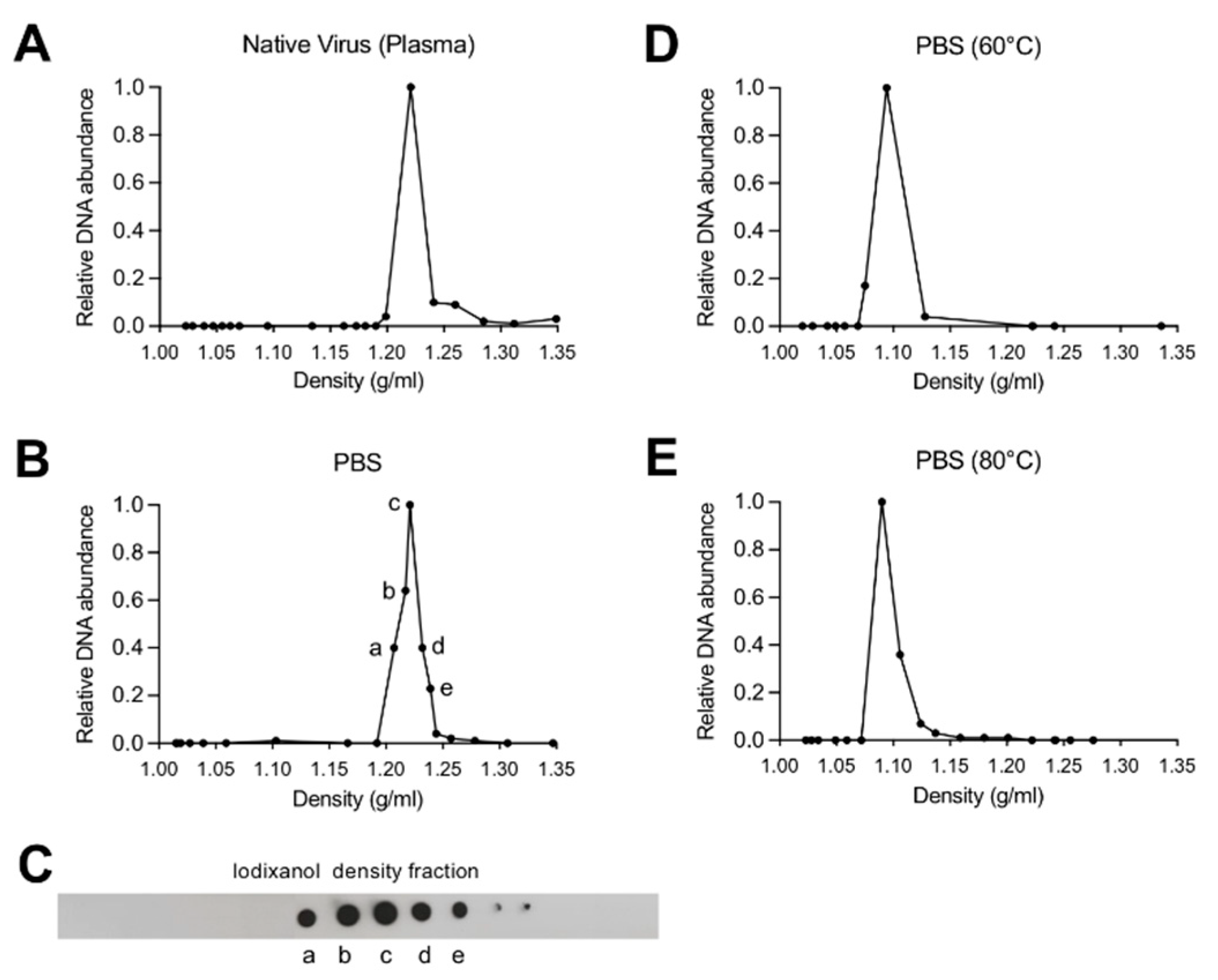
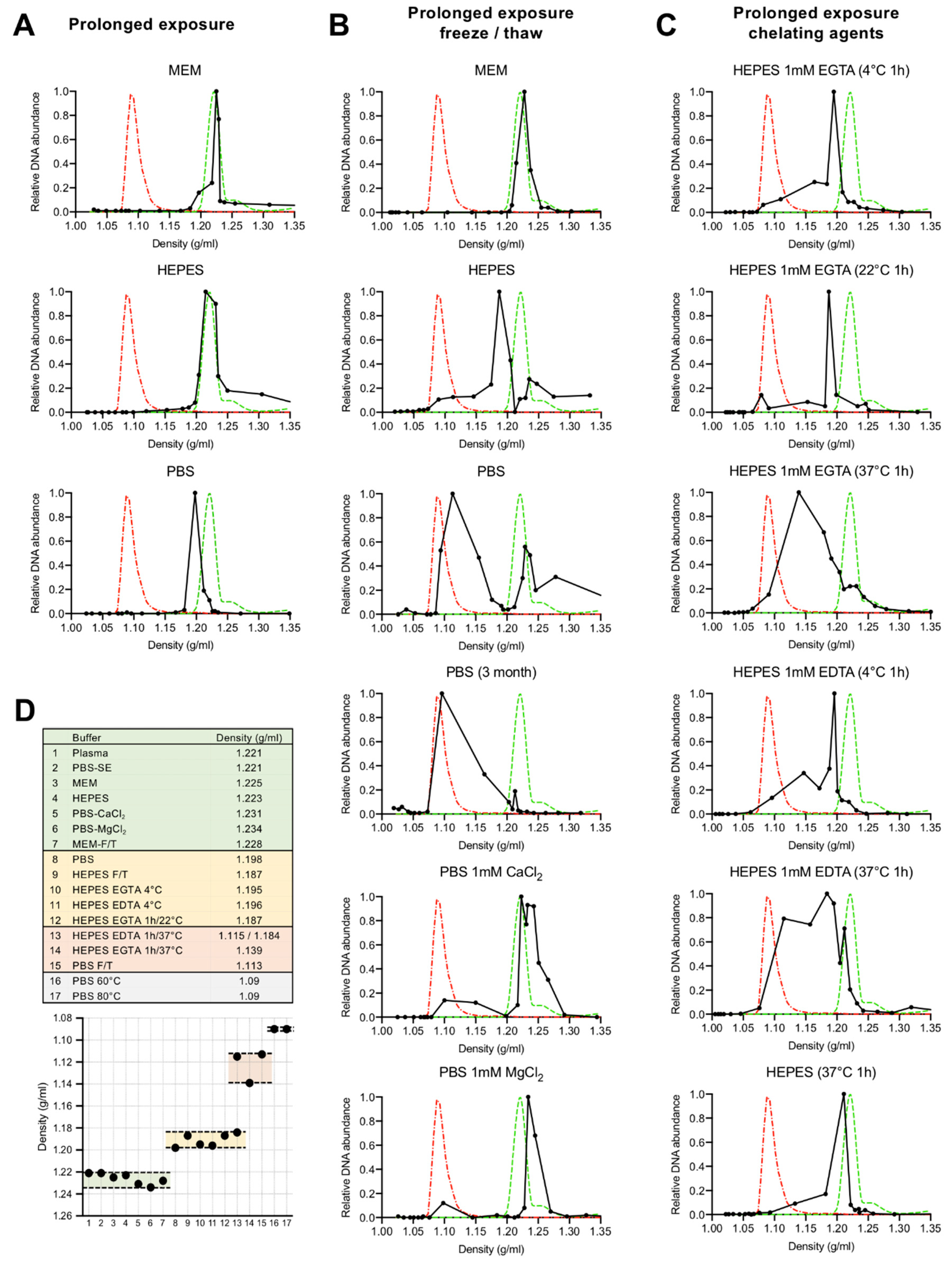
3.1. Depletion of Capsid-Associated Divalent Cations Destabilizes the B19V Capsid
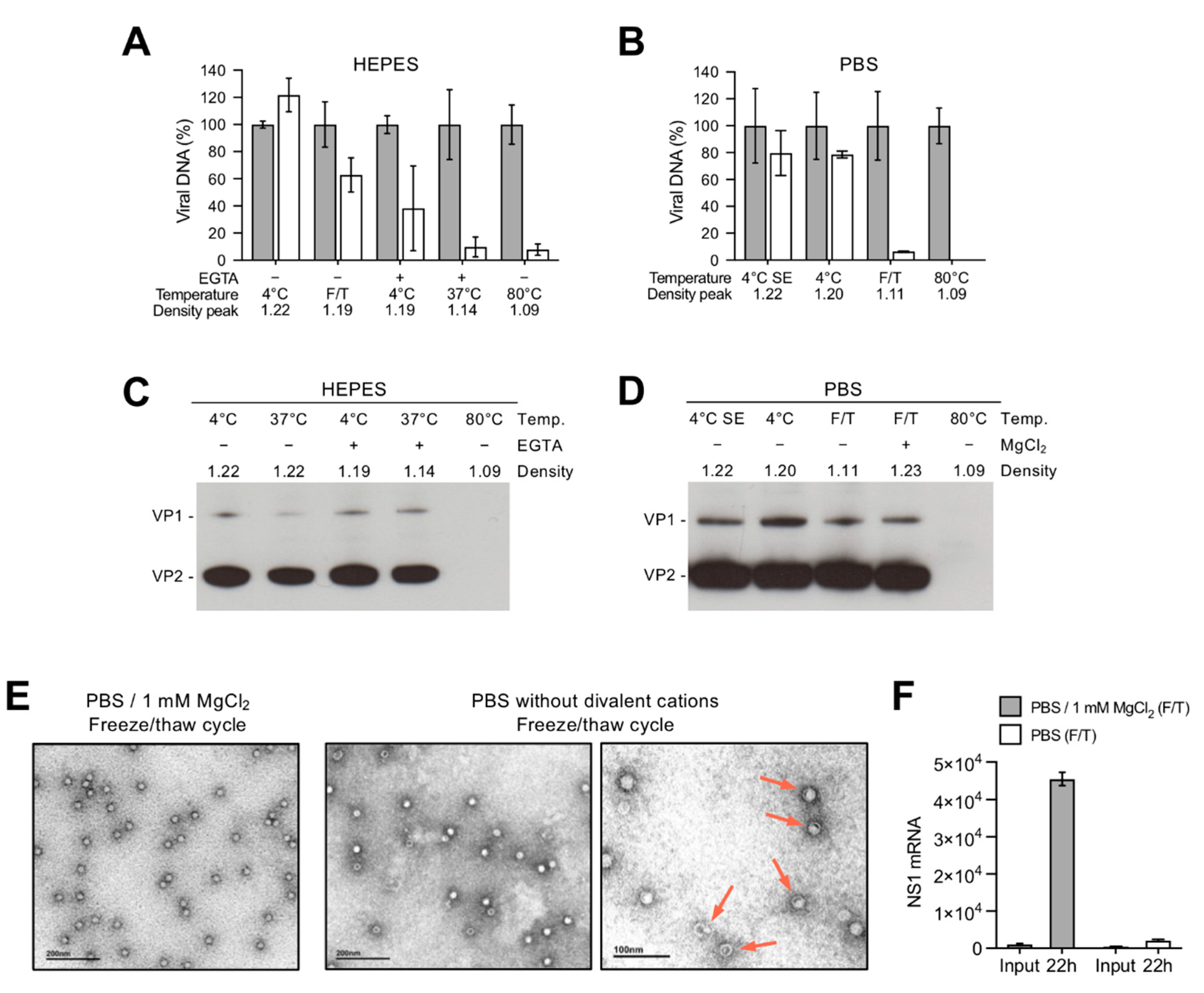
3.2. Depletion of Divalent Cations Facilitates B19V Uncoating without Capsid Disassembly
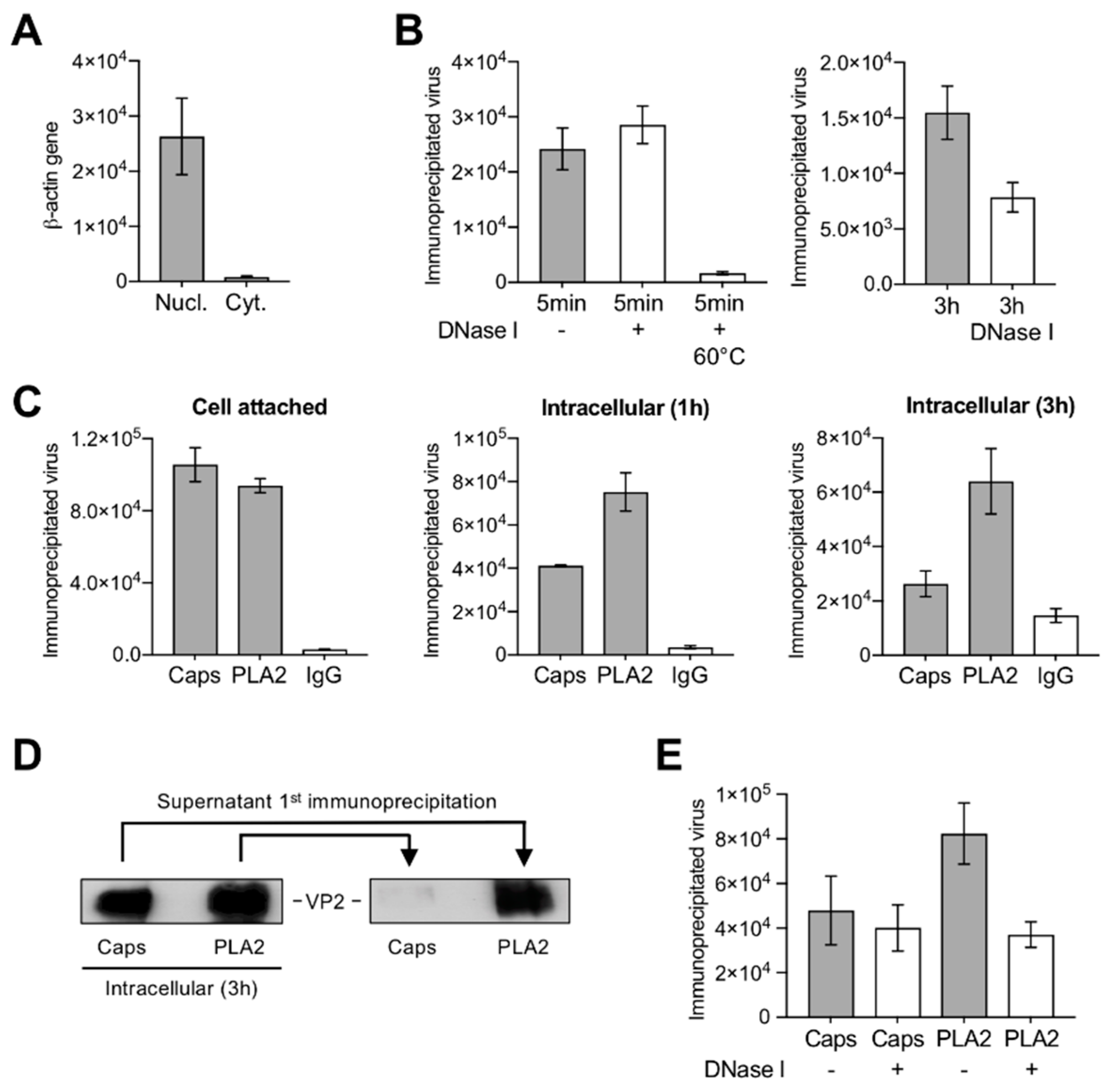
3.3. A Proportion of Incoming Capsids Uncoat in the Cytoplasm without Capsid Disassembly
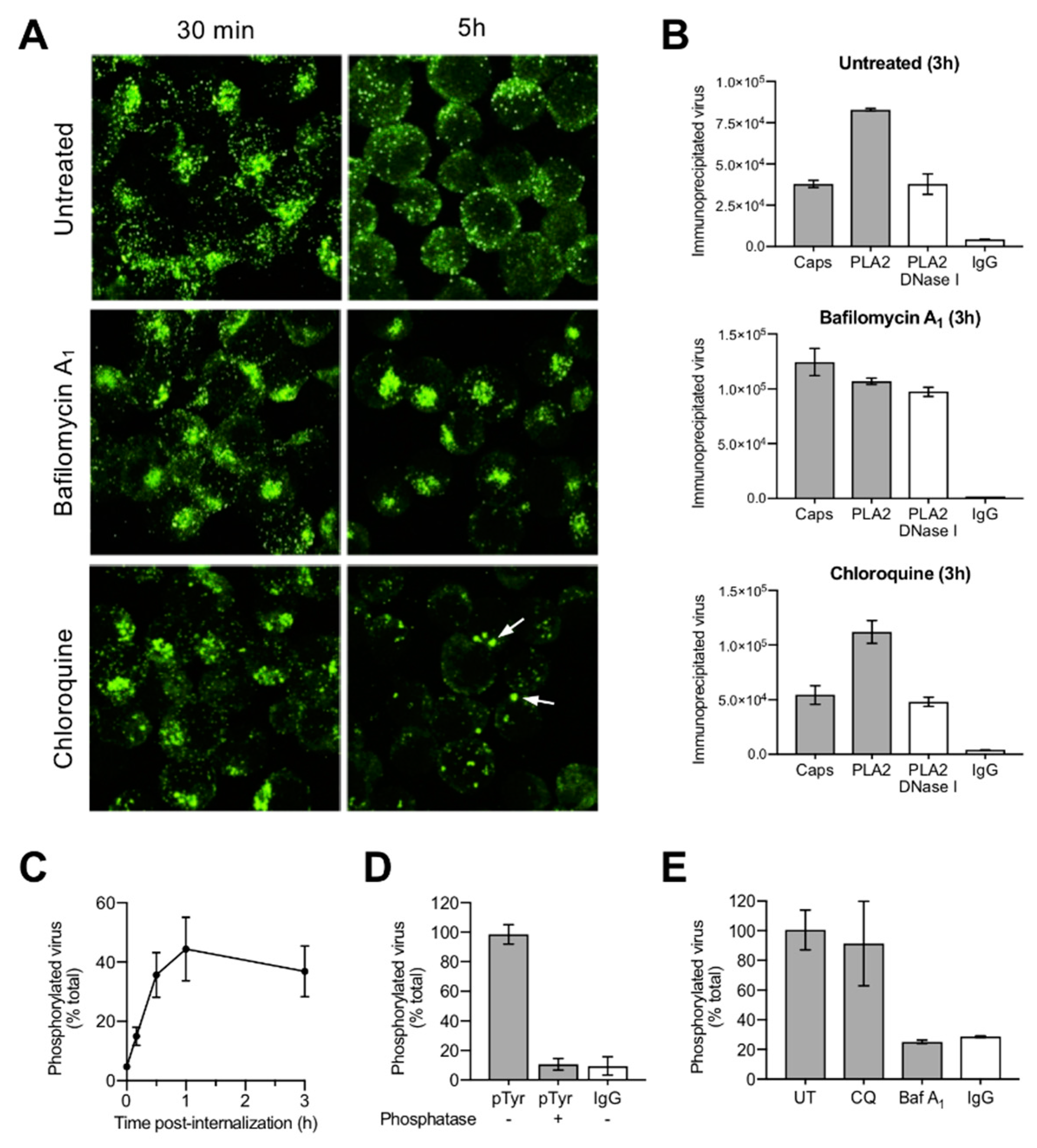
3.4. Uncoating Occurs after Endosomal Escape
3.5. Incoming Capsids Become Reactive to Phosphospecific Antibodies Following Endosomal Escape
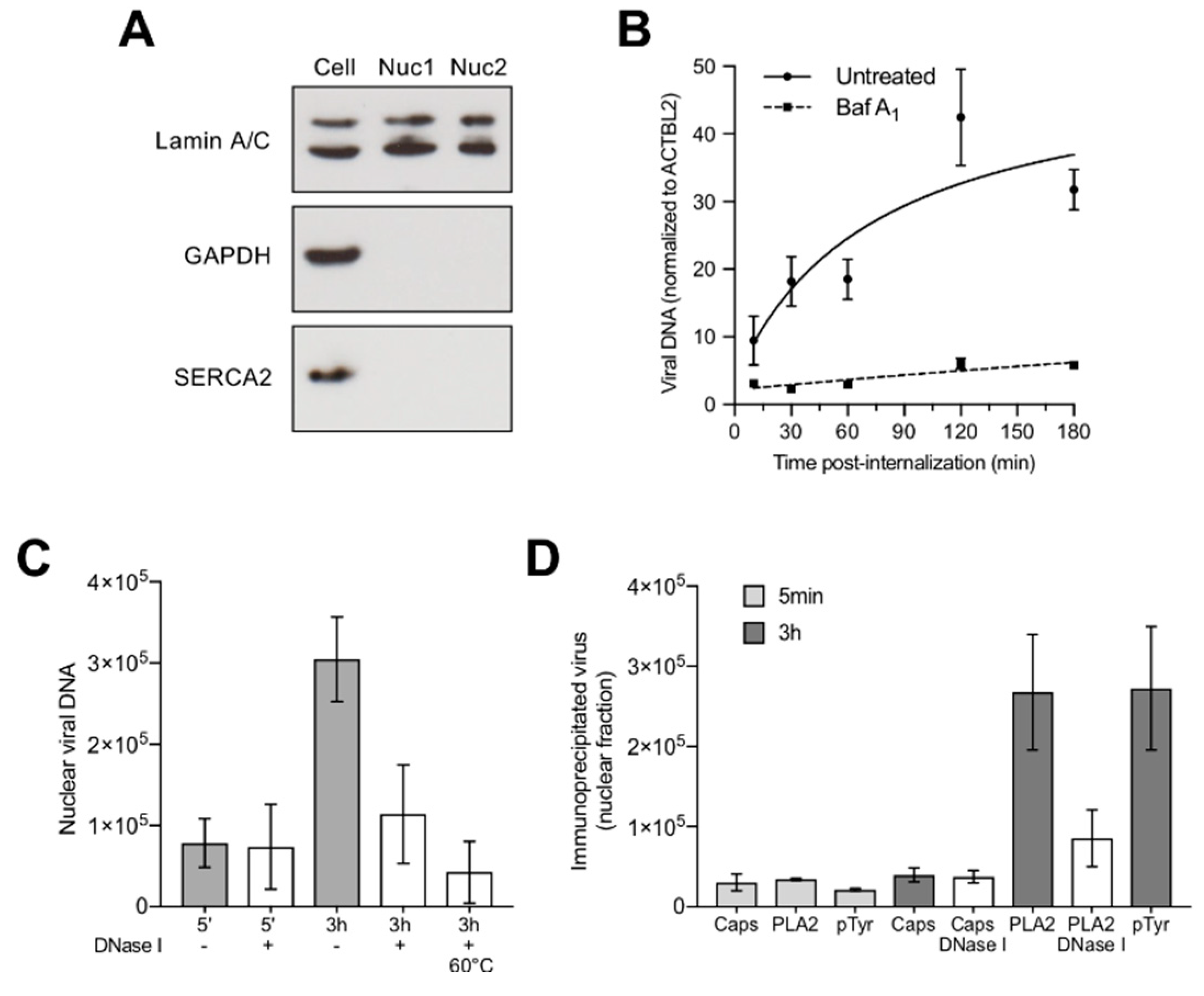
3.6. Capsids Phosphorylated and with Accessible Genomes Accumulated Progressively in the Nuclear Fraction
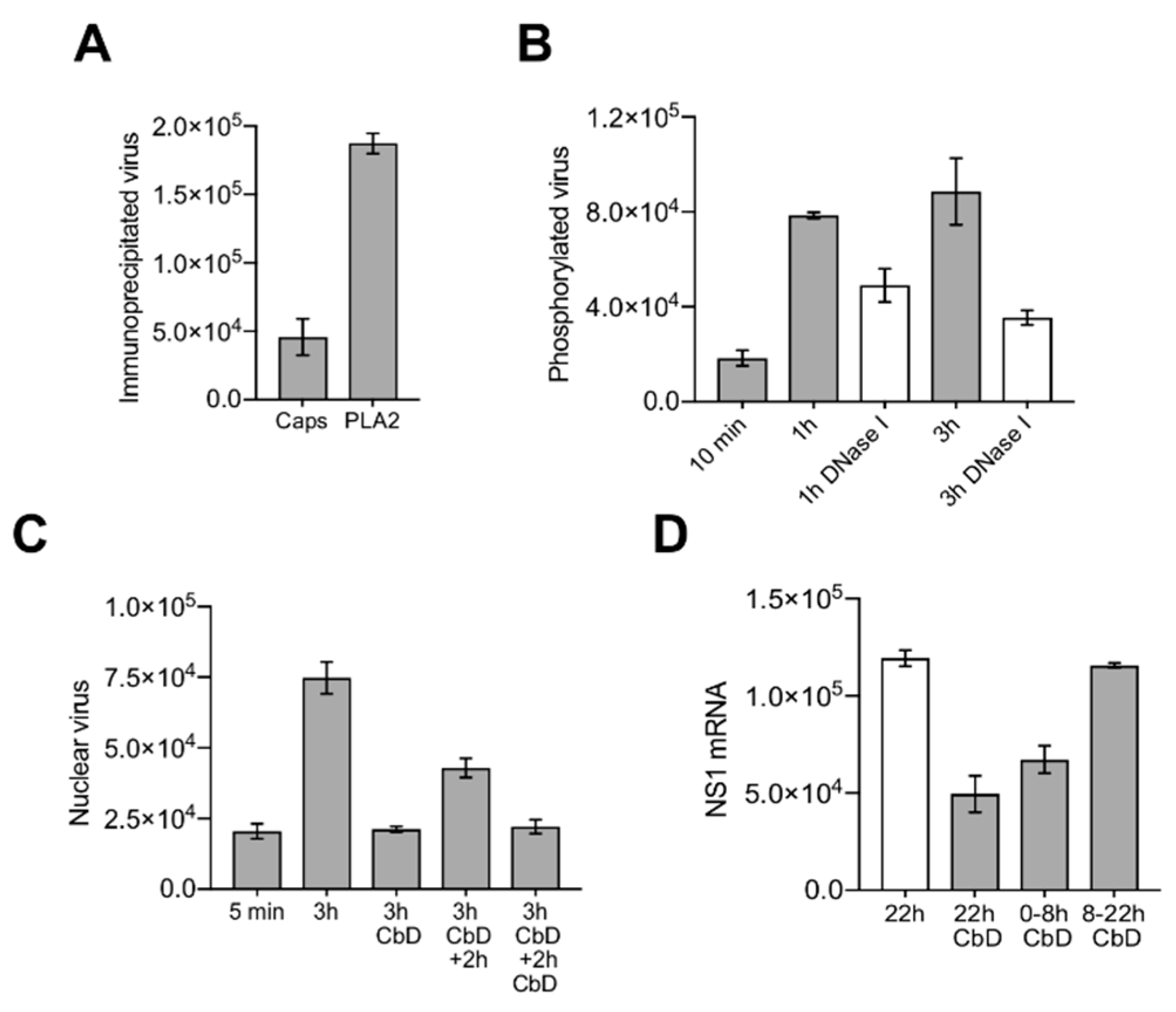
3.7. Nuclear Targeting of B19V Is Mediated by the Microtubule-Dependent, Minus-End-Directed Motor Dynein
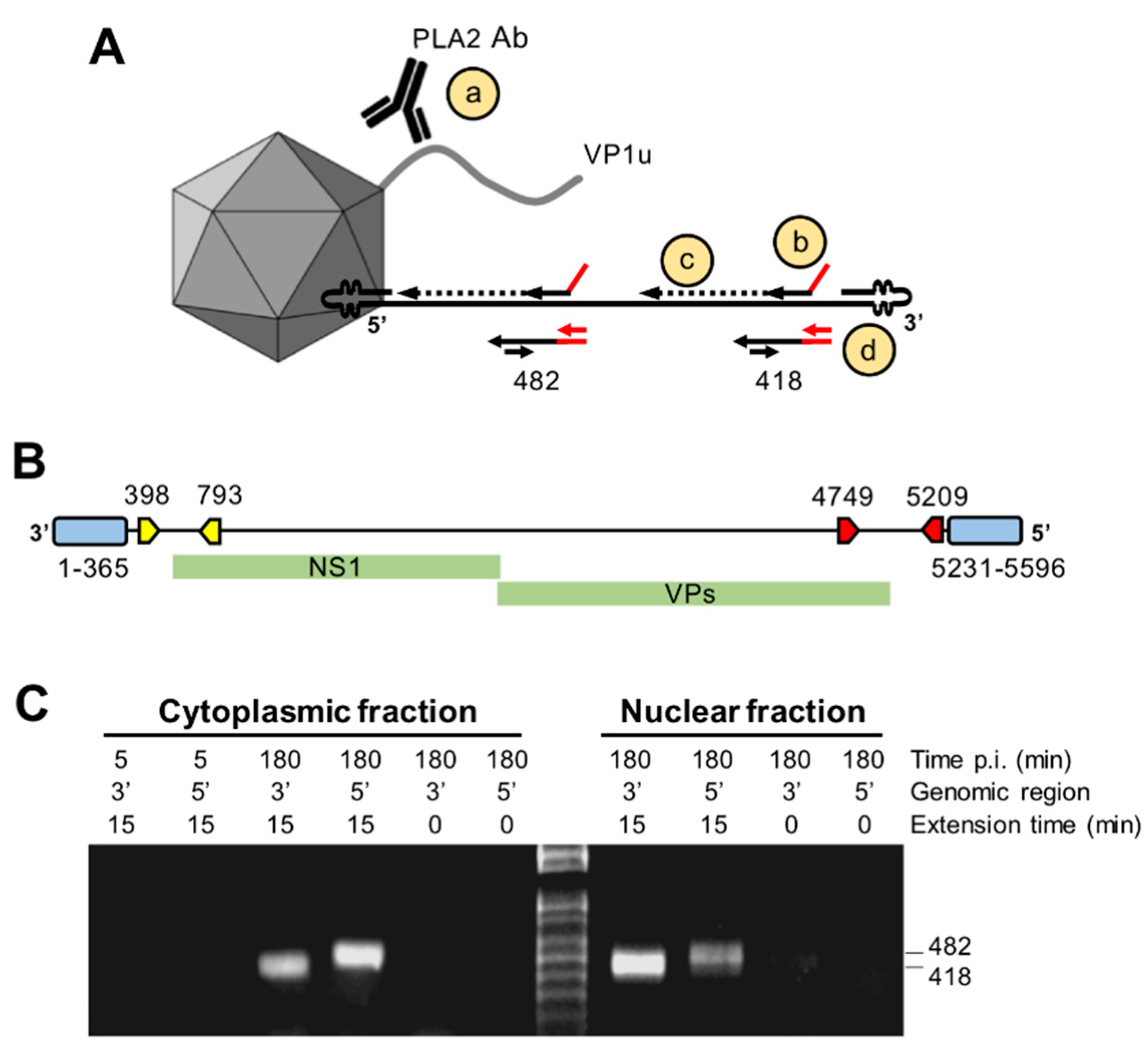
3.8. Capsids Immunoprecipitated from Cytoplasmic and from Nuclear Fractions Support Complementary-Strand DNA Synthesis
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Young, N.S.; Brown, K.E. Parvovirus B19. N. Engl. J. Med. 2004, 350, 586–597. [Google Scholar] [CrossRef] [PubMed]
- Servey, J.T.; Reamy, B.V.; Hodge, J. Clinical presentations of parvovirus B19 infection. Am. Fam. Physician 2007, 75, 373–376. [Google Scholar] [PubMed]
- Cotmore, S.F.; Mckie, V.C.; Anderson, L.J.; Astell, C.R.; Tattersall1, P.; Tattersall, P. Identification of the Major Structural and Nonstructural Proteins Encoded by Human Parvovirus B19 and Mapping of Their Genes by Procaryotic Expression of Isolated Genomic Fragments. J. Virol. 1986, 60, 548–557. [Google Scholar] [PubMed]
- Kilcher, S.; Mercer, J. DNA virus uncoating. Virology 2015, 479–480, 578–590. [Google Scholar] [CrossRef] [PubMed]
- Suomalainen, M.; Greber, U.F. Uncoating of non-enveloped viruses. Curr. Opin. Virol. 2013, 3, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Ravindran, M.S.; Tsai, B. Viruses Utilize Cellular Cues in Distinct Combination to Undergo Systematic Priming and Uncoating. PLoS Pathog. 2016, 12, e1005467. [Google Scholar] [CrossRef]
- Yamauchi, Y.; Greber, U.F. Principles of Virus Uncoating: Cues and the Snooker Ball. Traffic 2016, 17, 569–592. [Google Scholar] [CrossRef]
- Ros, C.; Bayat, N.; Wolfisberg, R.; Almendral, J.M. Protoparvovirus Cell Entry. Viruses 2017, 9, 313. [Google Scholar] [CrossRef]
- Quattrocchi, S.; Ruprecht, N.; Bönsch, C.; Bieli, S.; Zürcher, C.; Boller, K.; Kempf, C.; Ros, C. Characterization of the early steps of human parvovirus B19 infection. J. Virol. 2012, 86, 9274–9284. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Tattersall, P. Parvoviral Host Range and Cell Entry Mechanisms. Adv. Virus Res. 2007, 70, 183–232. [Google Scholar]
- Harbison, C.E.; Chiorini, J.A.; Parrish, C.R. The parvovirus capsid odyssey: From the cell surface to the nucleus. Trends Microbiol. 2008, 16, 208–214. [Google Scholar] [CrossRef]
- Parrish, C.R. Structures and Functions of Parvovirus Capsids and the Process of Cell Infection; Springer: Berlin/Heidelberg, Germany, 2010; pp. 149–176. [Google Scholar]
- Bönsch, C.; Kempf, C.; Ros, C. Interaction of parvovirus B19 with human erythrocytes alters virus structure and cell membrane integrity. J. Virol. 2008, 82, 11784–11791. [Google Scholar] [CrossRef]
- Bönsch, C.; Zuercher, C.; Lieby, P.; Kempf, C.; Ros, C. The globoside receptor triggers structural changes in the B19 virus capsid that facilitate virus internalization. J. Virol. 2010, 84, 11737–11746. [Google Scholar] [CrossRef]
- Leisi, R.; Ruprecht, N.; Kempf, C.; Ros, C. Parvovirus B19 uptake is a highly selective process controlled by VP1u, a novel determinant of viral tropism. J. Virol. 2013, 87, 13161–13167. [Google Scholar] [CrossRef]
- Leisi, R.; Von Nordheim, M.; Ros, C.; Kempf, C. The VP1u Receptor Restricts Parvovirus B19 Uptake to Permissive Erythroid Cells. Viruses 2016, 8, 265. [Google Scholar] [CrossRef]
- Zhang, W.; Tung, C.-H. Lysosome Enlargement Enhanced Photochemotherapy Using a Multifunctional Nanogel. ACS Appl. Mater. Interfaces 2018, 10, 4343–4348. [Google Scholar] [CrossRef]
- Suikkanen, S.; Aaltonen, T.; Nevalainen, M.; Välilehto, O.; Lindholm, L.; Vuento, M.; Vihinen-Ranta, M. Exploitation of microtubule cytoskeleton and dynein during parvoviral traffic toward the nucleus. J. Virol. 2003, 77, 10270–10279. [Google Scholar] [CrossRef]
- Kelkar, S.; De, B.P.; Gao, G.; Wilson, J.M.; Crystal, R.G.; Leopold, P.L. A common mechanism for cytoplasmic dynein-dependent microtubule binding shared among adeno-associated virus and adenovirus serotypes. J. Virol. 2006, 80, 7781–7785. [Google Scholar] [CrossRef]
- Hirosue, S.; Senn, K.; Clément, N.; Nonnenmacher, M.; Gigout, L.; Linden, R.M.; Weber, T. Effect of inhibition of dynein function and microtubule-altering drugs on AAV2 transduction. Virology 2007, 367, 10–18. [Google Scholar] [CrossRef]
- Lyi, S.M.; Tan, M.J.A.; Parrish, C.R. Parvovirus particles and movement in the cellular cytoplasm and effects of the cytoskeleton. Virology 2014, 456–457, 342–352. [Google Scholar] [CrossRef]
- Vihinen-Ranta, M.; Kakkola, L.; Kalela, A.; Vilja, P.; Vuento, M. Characterization of a Nuclear Localization Signal of Canine Parvovirus Capsid Proteins. Eur. J. Biochem. 1997, 250, 389–394. [Google Scholar] [CrossRef]
- Sonntag, F.; Bleker, S.; Leuchs, B.; Fischer, R.; Kleinschmidt, J.A. Adeno-Associated Virus Type 2 Capsids with Externalized VP1/VP2 Trafficking Domains Are Generated prior to Passage through the Cytoplasm and Are Maintained until Uncoating Occurs in the Nucleus. J. Virol. 2006, 80, 11040. [Google Scholar] [CrossRef]
- Johnson, J.S.; Li, C.; DiPrimio, N.; Weinberg, M.S.; McCown, T.J.; Samulski, R.J. Mutagenesis of adeno-associated virus type 2 capsid protein VP1 uncovers new roles for basic amino acids in trafficking and cell-specific transduction. J. Virol. 2010, 84, 8888–8902. [Google Scholar] [CrossRef]
- Popa-Wagner, R.; Porwal, M.; Kann, M.; Reuss, M.; Weimer, M.; Florin, L.; Kleinschmidt, J.A. Impact of VP1-Specific Protein Sequence Motifs on Adeno-Associated Virus Type 2 Intracellular Trafficking and Nuclear Entry. J. Virol. 2012, 86, 9163. [Google Scholar] [CrossRef]
- Nicolson, S.C.; Samulski, R.J. Recombinant adeno-associated virus utilizes host cell nuclear import machinery to enter the nucleus. J. Virol. 2014, 88, 4132–4144. [Google Scholar] [CrossRef]
- Cohen, S.; Marr, A.K.; Garcin, P.; Panté, N. Nuclear envelope disruption involving host caspases plays a role in the parvovirus replication cycle. J. Virol. 2011, 85, 4863–4874. [Google Scholar] [CrossRef]
- Fay, N.; Panté, N. Old foes, new understandings: Nuclear entry of small non-enveloped DNA viruses. Curr. Opin. Virol. 2015, 12, 59–65. [Google Scholar] [CrossRef]
- Lux, K.; Goerlitz, N.; Schlemminger, S.; Perabo, L.; Goldnau, D.; Endell, J.; Leike, K.; Kofler, D.M.; Finke, S.; Hallek, M.; et al. Green fluorescent protein-tagged adeno-associated virus particles allow the study of cytosolic and nuclear trafficking. J. Virol. 2005, 79, 11776–11787. [Google Scholar] [CrossRef]
- Farr, G.A.; Tattersall, P. A conserved leucine that constricts the pore through the capsid fivefold cylinder plays a central role in parvoviral infection. Virology 2004, 323, 243–256. [Google Scholar] [CrossRef]
- Bleker, S.; Sonntag, F.; Kleinschmidt, J.A. Mutational analysis of narrow pores at the fivefold symmetry axes of adeno-associated virus type 2 capsids reveals a dual role in genome packaging and activation of phospholipase A2 activity. J. Virol. 2005, 79, 2528–2540. [Google Scholar] [CrossRef]
- Farr, G.A.; Cotmore, S.F.; Tattersall, P. VP2 cleavage and the leucine ring at the base of the fivefold cylinder control pH-dependent externalization of both the VP1 N terminus and the genome of minute virus of mice. J. Virol. 2006, 80, 161–171. [Google Scholar] [CrossRef]
- Plevka, P.; Hafenstein, S.; Li, L.; D’Abrgamo, A.; Cotmore, S.F.; Rossmann, M.G.; Tattersall, P.; Tattersall, P. Structure of a packaging-defective mutant of minute virus of mice indicates that the genome is packaged via a pore at a 5-fold axis. J. Virol. 2011, 85, 4822–4827. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Tattersall, P. Parvoviruses: Small Does Not Mean Simple. Annu. Rev. Virol. 2014, 1, 517–537. [Google Scholar] [CrossRef]
- Kaufmann, B.; Simpson, A.A.; Rossmann, M.G. The structure of human parvovirus B19. Proc. Natl. Acad. Sci. USA 2004, 101, 11628–11633. [Google Scholar] [CrossRef]
- Cotmore, S.F.; D’Abramo, A.M.; Ticknor, C.M.; Tattersall, P. Controlled Conformational Transitions in the MVM Virion Expose the VP1 N-Terminus and Viral Genome without Particle Disassembly. Virology 1999, 254, 169–181. [Google Scholar] [CrossRef]
- Turnbull, A.E.; Skulimowski, A.; Smythe, J.A.; Alexander, I.E. Adeno-Associated Virus Vectors Show Variable Dependence on Divalent Cations for Thermostability: Implications for Purification and Handling. Hum. Gene Ther. 2000, 11, 629–635. [Google Scholar] [CrossRef]
- Ros, C.; Baltzer, C.; Mani, B.; Kempf, C. Parvovirus uncoating in vitro reveals a mechanism of DNA release without capsid disassembly and striking differences in encapsidated DNA stability. Virology 2006, 345, 137–147. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Tattersall, P. Mutations at the base of the icosahedral five-fold cylinders of minute virus of mice induce 3’-to-5’ genome uncoating and critically impair entry functions. J. Virol. 2012, 86, 69–80. [Google Scholar] [CrossRef]
- Horowitz, E.D.; Rahman, K.S.; Bower, B.D.; Dismuke, D.J.; Falvo, M.R.; Griffith, J.D.; Harvey, S.C.; Asokan, A. Biophysical and ultrastructural characterization of adeno-associated virus capsid uncoating and genome release. J. Virol. 2013, 87, 2994–3002. [Google Scholar] [CrossRef]
- Bernaud, J.; Rossi, A.; Fis, A.; Gardette, L.; Aillot, L.; Büning, H.; Castelnovo, M.; Salvetti, A.; Faivre-Moskalenko, C. Characterization of AAV vector particle stability at the single-capsid level. J. Biol. Phys. 2018, 44, 181–194. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Hafenstein, S.; Tattersall, P. Depletion of virion-associated divalent cations induces parvovirus minute virus of mice to eject its genome in a 3’-to-5’ direction from an otherwise intact viral particle. J. Virol. 2010, 84, 1945–1956. [Google Scholar] [CrossRef]
- Sun, Y.; Klose, T.; Liu, Y.; Modrow, S.; Rossmann, M.G. Structure of parvovirus B19 decorated by Fabs from a human antibody. J. Virol. 2019. [Google Scholar] [CrossRef]
- Gigler, A.; Dorsch, S.; Hemauer, A.; Williams, C.; Kim, S.; Young, N.S.; Zolla-Pazner, S.; Wolf, H.; Gorny, M.K.; Modrow, S. Generation of neutralizing human monoclonal antibodies against parvovirus B19 proteins. J. Virol. 1999, 73, 1974–1979. [Google Scholar]
- Zhong, L.; Li, B.; Jayandharan, G.; Mah, C.S.; Govindasamy, L.; Agbandje-McKenna, M.; Herzog, R.W.; Weigel-Van Aken, K.A.; Hobbs, J.A.; Zolotukhin, S.; et al. Tyrosine-phosphorylation of AAV2 vectors and its consequences on viral intracellular trafficking and transgene expression. Virology 2008, 381, 194–202. [Google Scholar] [CrossRef]
- Roberts, A.J.; Kon, T.; Knight, P.J.; Sutoh, K.; Burgess, S.A. Functions and mechanics of dynein motor proteins. Nat. Rev. Mol. Cell Biol. 2013, 14, 713–726. [Google Scholar] [CrossRef]
- Firestone, A.J.; Weinger, J.S.; Maldonado, M.; Barlan, K.; Langston, L.D.; O’Donnell, M.; Gelfand, V.I.; Kapoor, T.M.; Chen, J.K. Small-molecule inhibitors of the AAA+ ATPase motor cytoplasmic dynein. Nature 2012, 484, 125–129. [Google Scholar] [CrossRef]
- Sainath, R.; Gallo, G. The Dynein Inhibitor Ciliobrevin D Inhibits the Bi-directional Transport of Organelles along Sensory Axons and Impairs NGF- Mediated Regulation of Growth Cones and Axon Branches. Dev. Neurobiol. 2015, 75, 757–777. [Google Scholar] [CrossRef]
- Tsao, J.; Chapman, M.S.; Agbandje, M.; Keller, W.; Smith, K.; Wu, H.; Luo, M.; Smith, T.J.; Rossmann, M.G.; Compans, R.W.; et al. The three-dimensional structure of canine parvovirus and its functional implications. Science 1991, 251, 1456–1464. [Google Scholar] [CrossRef]
- Chapman, M.S.; Rossmann, M.G. Structure, Sequence, and Function Correlations among Parvoviruses. Virology 1993, 194, 491–508. [Google Scholar] [CrossRef]
- Llamas-Saiz, A.L.; Agbandje-McKenna, M.; Wikoff, W.R.; Bratton, J.; Tattersall, P.; Rossmann, M.G. IUCr Structure Determination of Minute Virus of Mice. Acta Crystallogr. Sect. D Biol. Crystallogr. 1997, 53, 93–102. [Google Scholar] [CrossRef]
- Agbandje-McKenna, M.; Llamas-Saiz, A.L.; Wang, F.; Tattersall, P.; Rossmann, M.G. Functional implications of the structure of the murine parvovirus, minute virus of mice. Structure 1998, 6, 1369–1381. [Google Scholar] [CrossRef]
- Kronenberg, S.; Böttcher, B.; von der Lieth, C.W.; Bleker, S.; Kleinschmidt, J.A. A conformational change in the adeno-associated virus type 2 capsid leads to the exposure of hidden VP1 N termini. J. Virol. 2005, 79, 5296–5303. [Google Scholar] [CrossRef]
- Castellanos, M.; Pérez, R.; Rodríguez-Huete, A.; Grueso, E.; Almendral, J.M.; Mateu, M.G. A slender tract of glycine residues is required for translocation of the VP2 protein N-terminal domain through the parvovirus MVM capsid channel to initiate infection. Biochem. J. 2013, 455, 87–94. [Google Scholar] [CrossRef]
- Kaufmann, B.; Chipman, P.R.; Kostyuchenko, V.A.; Modrow, S.; Rossmann, M.G. Visualization of the externalized VP2 N termini of infectious human parvovirus B19. J. Virol. 2008, 82, 7306–7312. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Tattersall, P. Genome packaging sense is controlled by the efficiency of the nick site in the right-end replication origin of parvoviruses minute virus of mice and LuIII. J. Virol. 2005, 79, 2287–2300. [Google Scholar] [CrossRef]
- King, J.A.; Dubielzig, R.; Grimm, D.; Kleinschmidt, J.A. DNA helicase-mediated packaging of adeno-associated virus type 2 genomes into preformed capsids. EMBO J. 2001, 20, 3282–3291. [Google Scholar] [CrossRef]
- Ivanovska, I.; Wuite, G.; Jönsson, B.; Evilevitch, A. Internal DNA pressure modifies stability of WT phage. Proc. Natl. Acad. Sci. USA 2007, 104, 9603–9608. [Google Scholar] [CrossRef]
- Bauer, D.W.; Li, D.; Huffman, J.; Homa, F.L.; Wilson, K.; Leavitt, J.C.; Casjens, S.R.; Baines, J.; Evilevitch, A. Exploring the Balance between DNA Pressure and Capsid Stability in Herpesviruses and Phages. J. Virol. 2015, 89, 9288–9298. [Google Scholar] [CrossRef]
- Shirley, J.A.; Beards, G.M.; Thouless, M.E.; Flewett, T.H. The influence of divalent cations on the stability of human rotavirus. Arch. Virol. 1981, 67, 1–9. [Google Scholar] [CrossRef]
- Sherman, M.B.; Guenther, R.H.; Tama, F.; Sit, T.L.; Brooks, C.L.; Mikhailov, A.M.; Orlova, E.V.; Baker, T.S.; Lommel, S.A. Removal of divalent cations induces structural transitions in red clover necrotic mosaic virus, revealing a potential mechanism for RNA release. J. Virol. 2006, 80, 10395–10406. [Google Scholar] [CrossRef]
- Plevka, P.; Kazaks, A.; Voronkova, T.; Kotelovica, S.; Dishlers, A.; Liljas, L.; Tars, K. The structure of bacteriophage phiCb5 reveals a role of the RNA genome and metal ions in particle stability and assembly. J. Mol. Biol. 2009, 391, 635–647. [Google Scholar] [CrossRef]
- Llauró, A.; Coppari, E.; Imperatori, F.; Bizzarri, A.R.; Castón, J.R.; Santi, L.; Cannistraro, S.; de Pablo, P.J. Calcium ions modulate the mechanics of tomato bushy stunt virus. Biophys. J. 2015, 109, 390–397. [Google Scholar] [CrossRef]
- Kawano, M.; Xing, L.; Tsukamoto, H.; Inoue, T.; Handa, H.; Cheng, R.H. Calcium Bridge Triggers Capsid Disassembly in the Cell Entry Process of Simian Virus 40. J. Biol. Chem. 2009, 284, 34703–34712. [Google Scholar] [CrossRef]
- Hu, X.; Dong, Q.; Yang, J.; Zhang, Y. Recognizing metal and acid radical ion-binding sites by integrating ab initio modeling with template-based transferals. Bioinformatics 2016, 32, 3260–3269. [Google Scholar] [CrossRef]
- Clapham, D.E. Calcium Signaling. Cell 2007, 131, 1047–1058. [Google Scholar] [CrossRef]
- Vihinen-Ranta, M.; Wang, D.; Weichert, W.S.; Parrish, C.R. The VP1 N-terminal sequence of canine parvovirus affects nuclear transport of capsids and efficient cell infection. J. Virol. 2002, 76, 1884–1891. [Google Scholar] [CrossRef]
- Ros, C.; Kempf, C. The ubiquitin–proteasome machinery is essential for nuclear translocation of incoming minute virus of mice. Virology 2004, 324, 350–360. [Google Scholar] [CrossRef]
- Mani, B.; Baltzer, C.; Valle, N.; Almendral, J.M.; Kempf, C.; Ros, C. Low pH-dependent endosomal processing of the incoming parvovirus minute virus of mice virion leads to externalization of the VP1 N-terminal sequence (N-VP1), N-VP2 cleavage, and uncoating of the full-length genome. J. Virol. 2006, 80, 1015–1024. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Caliaro, O.; Marti, A.; Ruprecht, N.; Leisi, R.; Subramanian, S.; Hafenstein, S.; Ros, C. Parvovirus B19 Uncoating Occurs in the Cytoplasm without Capsid Disassembly and It Is Facilitated by Depletion of Capsid-Associated Divalent Cations. Viruses 2019, 11, 430. https://doi.org/10.3390/v11050430
Caliaro O, Marti A, Ruprecht N, Leisi R, Subramanian S, Hafenstein S, Ros C. Parvovirus B19 Uncoating Occurs in the Cytoplasm without Capsid Disassembly and It Is Facilitated by Depletion of Capsid-Associated Divalent Cations. Viruses. 2019; 11(5):430. https://doi.org/10.3390/v11050430
Chicago/Turabian StyleCaliaro, Oliver, Andrea Marti, Nico Ruprecht, Remo Leisi, Suriyasri Subramanian, Susan Hafenstein, and Carlos Ros. 2019. "Parvovirus B19 Uncoating Occurs in the Cytoplasm without Capsid Disassembly and It Is Facilitated by Depletion of Capsid-Associated Divalent Cations" Viruses 11, no. 5: 430. https://doi.org/10.3390/v11050430
APA StyleCaliaro, O., Marti, A., Ruprecht, N., Leisi, R., Subramanian, S., Hafenstein, S., & Ros, C. (2019). Parvovirus B19 Uncoating Occurs in the Cytoplasm without Capsid Disassembly and It Is Facilitated by Depletion of Capsid-Associated Divalent Cations. Viruses, 11(5), 430. https://doi.org/10.3390/v11050430




