Further Evidence for in Utero Transmission of Equine Hepacivirus to Foals
Abstract
1. Introduction
2. Materials and Methods
2.1. Necrospy and Histological Analysis
2.2. Molecular Detection and Characterisation of the EqHV Strains
2.2.1. Nucleic Acid Extraction and Quantitative RT-PCR
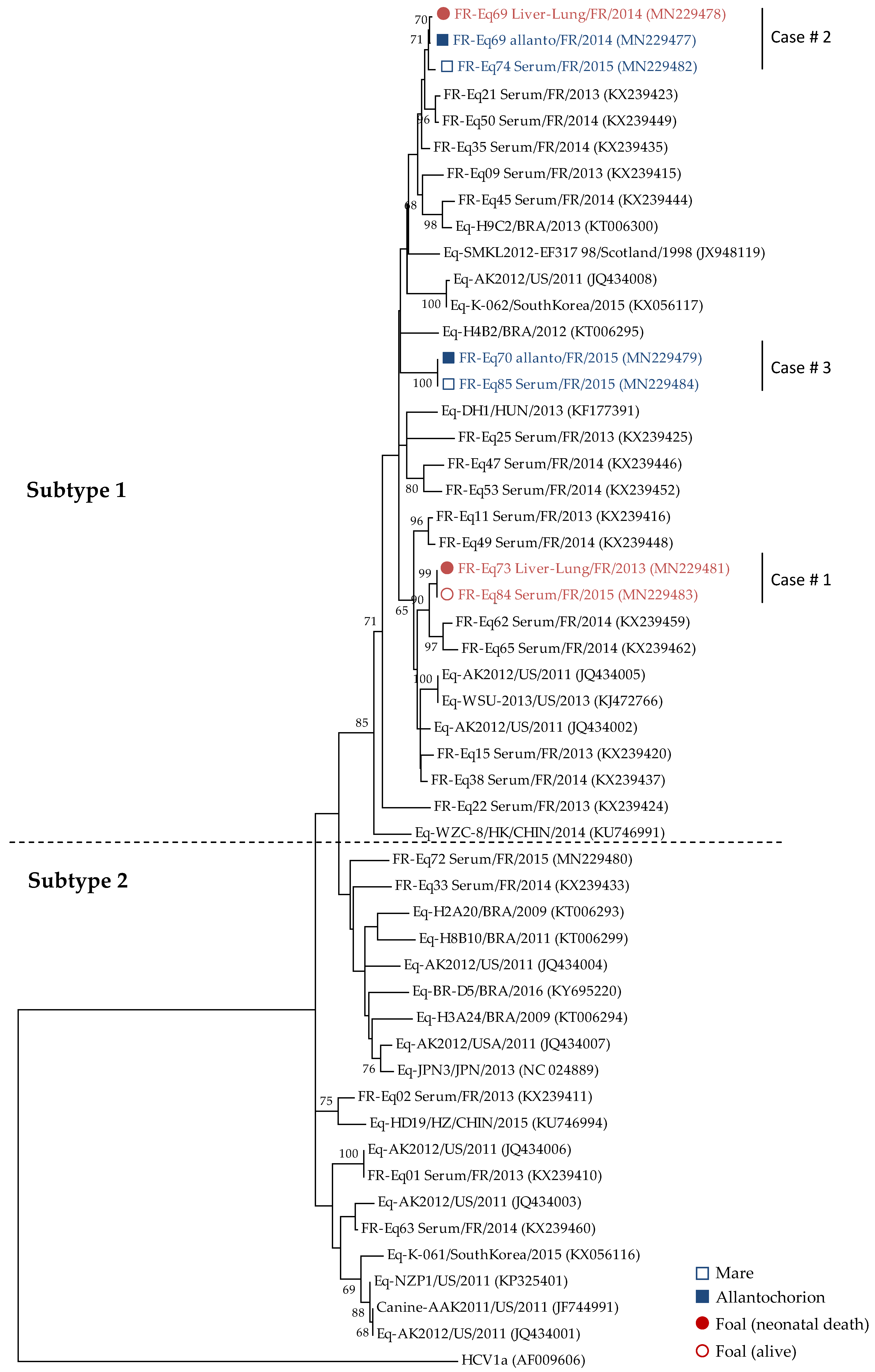
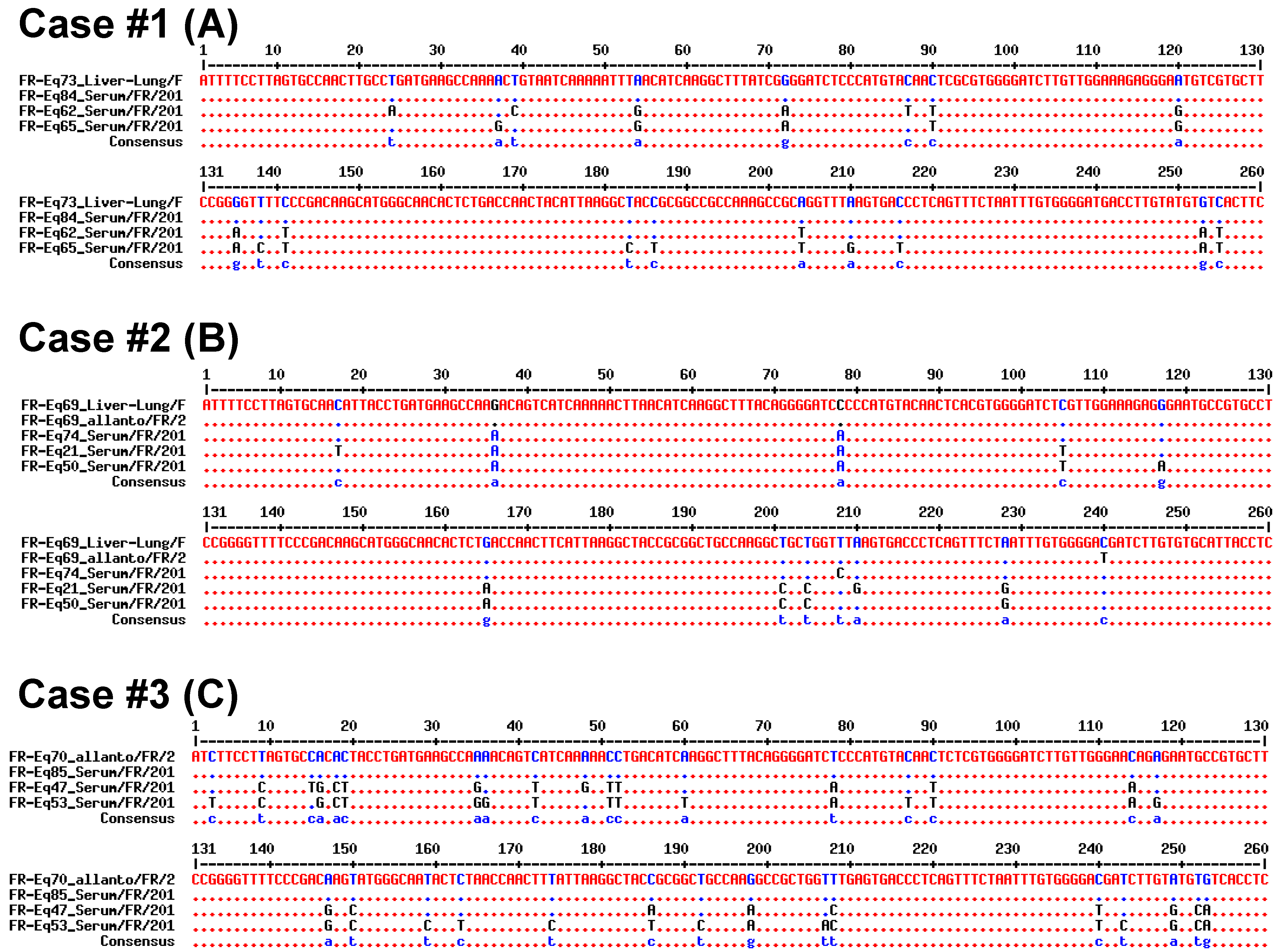
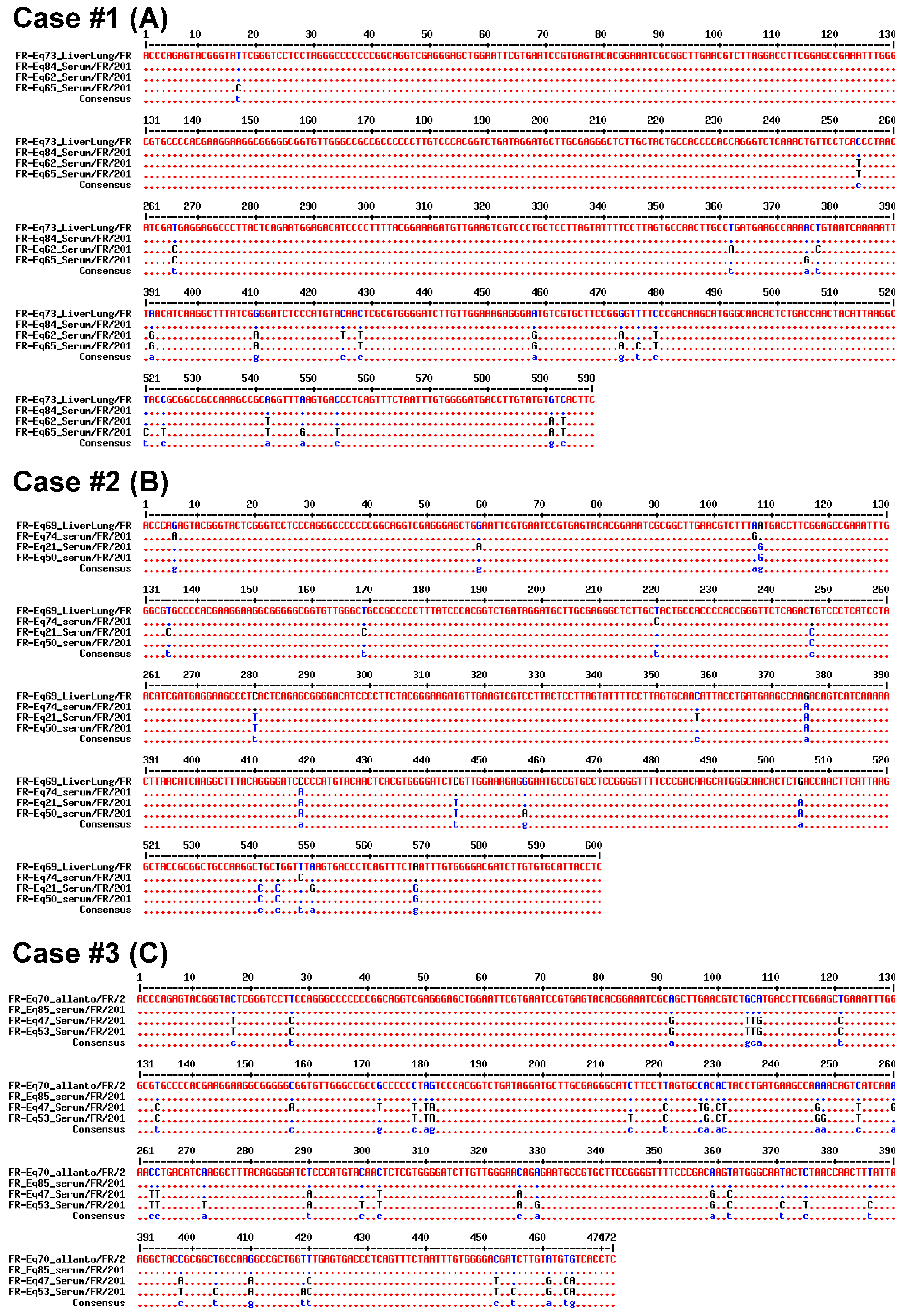
2.2.2. Sequencing and Phylogenic Analysis
3. Results
3.1. Prevalence of EqHV in Perinatal Foal Deaths
3.2. Vertical Transmission of EqHV from Mare to Foal
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Burbelo, P.D.; Dubovi, E.J.; Simmonds, P.; Medina, J.L.; Henriquez, J.A.; Mishra, N.; Wagner, J.; Tokarz, R.; Cullen, J.M.; Iadarola, M.J.; et al. Serology-Enabled Discovery of Genetically Diverse Hepaciviruses in a New Host. J. Virol. 2012, 86, 6171–6178. [Google Scholar] [CrossRef] [PubMed]
- Pronost, S.; Hue, E.; Fortier, C.; Foursin, M.; Fortier, G.; Desbrosse, F.; Rey, F.A.; Pitel, P.-H.; Richard, E.; Saunier, B. Prevalence of Equine Hepacivirus Infections in France and Evidence for Two Viral Subtypes Circulating Worldwide. Transbound. Emerg. Dis. 2016, 64, 1884–1897. [Google Scholar] [CrossRef]
- Tanaka, T.; Kasai, H.; Yamashita, A.; Okuyama-Dobashi, K.; Yasumoto, J.; Maekawa, S.; Enomoto, N.; Okamoto, T.; Matsuura, Y.; Morimatsu, M.; et al. Hallmarks of Hepatitis C Virus in Equine Hepacivirus. J. Virol. 2014, 88, 13352–13366. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gemaque, B.S.; Junior Souza de Souza, A.; do Carmo Pereira Soares, M.; Malheiros, A.P.; Silva, A.L.; Alves, M.M.; Gomes-Gouvêa, M.S.; Pinho, J.R.R.; Ferreira de Figueiredo, H.; Ribeiro, D.B.; et al. Hepacivirus infection in domestic horses, Brazil, 2011–2013. Emerg. Infect. Dis. 2014, 20, 2180–2182. [Google Scholar] [CrossRef] [PubMed]
- Lyons, S.; Kapoor, A.; Sharp, C.; Schneider, B.S.; Wolfe, N.D.; Culshaw, G.; Corcoran, B.; McGorum, B.C.; Simmonds, P. Nonprimate Hepaciviruses in Domestic Horses, United Kingdom. Emerg. Infect. Dis. 2012, 18, 1976–1982. [Google Scholar] [CrossRef]
- Badenhorst, M.; Tegtmeyer, B.; Todt, D.; Guthrie, A.; Feige, K.; Campe, A.; Steinmann, E.; Cavalleri, J.M.V. First detection and frequent occurrence of Equine Hepacivirus in horses on the African continent. Vet. Microbiol. 2018, 223, 51–58. [Google Scholar] [CrossRef]
- Figueiredo, A.S.; Lampe, E.; do Espírito-Santo, M.P.; do Amaral Mello, F.C.; de Almeida, F.Q.; de Lemos, E.R.S.; Godoi, T.L.O.S.; Dimache, L.A.G.; dos Santos, D.R.L.; Villar, L.M. Identification of two phylogenetic lineages of equine hepacivirus and high prevalence in Brazil. Vet. J. 2015, 206, 414–416. [Google Scholar] [CrossRef]
- Matsuu, A.; Hobo, S.; Ando, K.; Sanekata, T.; Sato, F.; Endo, Y.; Amaya, T.; Osaki, T.; Horie, M.; Masatani, T.; et al. Genetic and serological surveillance for non-primate hepacivirus in horses in Japan. Vet. Microbiol. 2015, 179, 219–227. [Google Scholar] [CrossRef]
- Pfaender, S.; Cavalleri, J.M.V.; Walter, S.; Doerrbecker, J.; Campana, B.; Brown, R.J.P.; Burbelo, P.D.; Postel, A.; Hahn, K.; Anggakusuma; et al. Clinical course of infection and viral tissue tropism of hepatitis C virus-like nonprimate hepaciviruses in horses. Hepatology 2015, 61, 447–459. [Google Scholar] [CrossRef]
- Scheel, T.K.H.; Simmonds, P.; Kapoor, A. Surveying the global virome: Identification and characterization of HCV-related animal hepaciviruses. Antivir. Res. 2015, 115, 83–93. [Google Scholar] [CrossRef]
- Pronost, S.; Fortier, C.; Hue, E.; Desbrosse, F.; Foursin, M.; Fortier, G.; Saunier, B.; Pitel, P.-H. Hépacivirus, pégivirus, TDAV: Une nouvelle triade de virus hépatiques chez le cheval? Pratique Vétérinaire Équine 2018, 197, 24–31. [Google Scholar]
- Badenhorst, M.; de Heus, P.; Auer, A.; Rümenapf, T.; Tegtmeyer, B.; Kolodziejek, J.; Nowotny, N.; Steinmann, E.; Cavalleri, J.-M.V. No Evidence of Mosquito Involvement in the Transmission of Equine Hepacivirus (Flaviviridae) in an Epidemiological Survey of Austrian Horses. Viruses 2019, 11, 1014. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Giannitti, F.; Low, J.; Keyes, C.; Ullmann, L.S.; Deng, X.; Aleman, M.; Pesavento, P.A.; Pusterla, N.; Delwart, E. Exploring the virome of diseased horses. J. Gen. Virol. 2015, 96, 2721–2733. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, J.D.; Evanoff, R.; Wilkinson, T.E.; Divers, T.J.; Knowles, D.P.; Mealey, R.H. Experimental transmission of equine hepacivirus in horses as a model for hepatitis C virus. Hepatology 2015, 61, 1533–1546. [Google Scholar] [CrossRef] [PubMed]
- Scheel, T.K.H.; Kapoor, A.; Nishiuchi, E.; Brock, K.V.; Yu, Y.; Andrus, L.; Gu, M.; Renshaw, R.W.; Dubovi, E.J.; McDonough, S.P.; et al. Characterization of nonprimate hepacivirus and construction of a functional molecular clone. Proc. Natl. Acad. Sci. USA 2015, 112, 2192–2197. [Google Scholar] [CrossRef] [PubMed]
- Indolfi, G.; Easterbrook, P.; Dusheiko, G.; El-Sayed, M.H.; Jonas, M.M.; Thorne, C.; Bulterys, M.; Siberry, G.; Walsh, N.; Chang, M.-H.; et al. Hepatitis C virus infection in children and adolescents. Lancet Gastroenterol. Hepatol. 2019, 4, 477–487. [Google Scholar] [CrossRef]
- Benova, L.; Mohamoud, Y.A.; Calvert, C.; Abu-Raddad, L.J. Vertical Transmission of Hepatitis C Virus: Systematic Review and Meta-analysis. Clin. Infect. Dis. 2014, 59, 765–773. [Google Scholar] [CrossRef]
- Tovo, P.-A.; Calitri, C.; Scolfaro, C.; Gabiano, C.; Garazzino, S. Vertically acquired hepatitis C virus infection: Correlates of transmission and disease progression. World J. Gastroenterol. 2016, 22, 1382–1392. [Google Scholar] [CrossRef]
- Gather, T.; Walter, S.; Todt, D.; Pfaender, S.; Brown, R.J.P.; Postel, A.; Becher, P.; Moritz, A.; Hansmann, F.; Baumgaertner, W.; et al. Vertical transmission of hepatitis C virus-like non-primate hepacivirus in horses. J. Gen. Virol. 2016, 97, 2540–2551. [Google Scholar] [CrossRef]
- Rooney, J.R. Autopsy of the Horse. Technique and Interpretation; The Williams & Wilkins Company: Baltimore, MD, USA, 1970. [Google Scholar]
- Laugier, C.; Foucher, N.; Sevin, C.; Leon, A.; Tapprest, J. A 24-Year Retrospective Study of Equine Abortion in Normandy (France). J. Equine Vet. Sci. 2011, 31, 116–123. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef] [PubMed]
- Leigh, J.W.; Bryant, D. Popart: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Bandelt, H.J.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Postel, A.; Cavalleri, J.-M.V.; Pfaender, S.; Walter, S.; Steinmann, E.; Fischer, N.; Feige, K.; Haas, L.; Becher, P. Frequent presence of hepaci and pegiviruses in commercial equine serum pools. Vet. Microbiol. 2016, 182, 8–14. [Google Scholar] [CrossRef]
- Lu, G.; Huang, J.; Yang, Q.; Xu, H.; Wu, P.; Fu, C.; Li, S. Identification and genetic characterization of hepacivirus and pegivirus in commercial equine serum products in China. PLoS ONE 2017, 12, e0189208. [Google Scholar] [CrossRef]
- Pfaender, S.; Walter, S.; Grabski, E.; Todt, D.; Bruening, J.; Romero-Brey, I.; Gather, T.; Brown, R.J.; Hahn, K.; Puff, C. Immune protection against reinfection with nonprimate hepacivirus. Proc. Natl. Acad. Sci. USA 2017, 114, E2430–E2439. [Google Scholar] [CrossRef]
- Altan, E.; Li, Y.; Sabino-Santos, G., Jr.; Sawaswong, V.; Barnum, S.; Pusterla, N.; Deng, X.; Delwart, E. Viruses in Horses with Neurologic and Respiratory Diseases. Viruses 2019, 11, 942. [Google Scholar] [CrossRef]
- Paim, W.P.; Weber, M.N.; Cibulski, S.P.; da Silva, M.S.; Puhl, D.E.; Budaszewski, R.F.; Varela, A.P.M.; Mayer, F.Q.; Canal, C.W. Characterization of the viral genomes present in commercial batches of horse serum obtained by high-throughput sequencing. Biologicals 2019, 61, 1–7. [Google Scholar] [CrossRef]
- Pybus, O.G.; Thézé, J. Hepacivirus cross-species transmission and the origins of the hepatitis C virus. Curr. Opin. Virol. 2016, 16, 1–7. [Google Scholar] [CrossRef]
- Divers, T.J.; Barton, M.H. Chapter 13: Disorders of the Liver. In Equine Internal Medicine, 4th ed.; Reed, S., Baylyd, W., Sellon, D., Saunders, Sellon, D., Eds.; Elsevier: Saint-louis, MO, USA, 2017; pp. 843–887. ISBN 978-0-323-44552-8. [Google Scholar]
- LeBlanc, M.M. Immunologic considerations. In Equine Clinical Neonatalogy; Lea & Febiger: Philadelphia, PA, USA, 1990; pp. 275–294. ISBN 978-0-8121-1184-2. [Google Scholar]
- Pfaender, S.; Walter, S.; Todt, D.; Behrendt, P.; Doerrbecker, J.; Wölk, B.; Engelmann, M.; Gravemann, U.; Seltsam, A.; Steinmann, J.; et al. Assessment of cross-species transmission of hepatitis C virus-related non-primate hepacivirus in a population of humans at high risk of exposure. J. Gen. Virol. 2015, 96, 2636–2642. [Google Scholar] [CrossRef] [PubMed]
- Steininger, C.; Kundi, M.; Jatzko, G.; Kiss, H.; Lischka, A.; Holzmann, H. Increased risk of mother-to-infant transmission of hepatitis C virus by intrapartum infantile exposure to maternal blood. J. Infect. Dis. 2003, 187, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Gould, L.H.; Fikrig, E. West Nile virus: A growing concern? J. Clin. Investig. 2004, 113, 1102–1107. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention (CDC). Intrauterine West Nile virus infection—New York, 2002. MMWR Morb. Mortal. Wkly. Rep. 2002, 51, 1135–1136. [Google Scholar]
- Chaturvedi, U.C.; Mathur, A.; Chandra, A.; Das, S.K.; Tandon, H.O.; Singh, U.K. Transplacental infection with Japanese encephalitis virus. J. Infect. Dis. 1980, 141, 712–715. [Google Scholar] [CrossRef] [PubMed]
- Chye, J.K.; Lim, C.T.; Ng, K.B.; Lim, J.M.; George, R.; Lam, S.K. Vertical transmission of dengue. Clin. Infect. Dis. 1997, 25, 1374–1377. [Google Scholar] [CrossRef]
- Khaiboullina, S.F.; Ribeiro, F.M.; Uppal, T.; Martynova, E.V.; Rizvanov, A.A.; Verma, S.C. Zika Virus Transmission Through Blood Tissue Barriers. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Pronost, S.; Hue, E.; Fortier, C.; Foursin, M.; Fortier, G.; Desbrosse, F.; Rey, F.; Pitel, P.-H.; Saunier, B. Identification of equine hepacivirus infections in France: Facts and Physiopathological insights. J. Equine Vet. Sci. 2016, 39, S22. [Google Scholar] [CrossRef]
| Cases (Year) | Subjects | Sampling | Life Status (/Birth) | Viral Loads | EqHV Strains |
|---|---|---|---|---|---|
| Case #1 (2013 & 2015) | Foal #1 (4 days) | Liver+Lung | Neonatal death (4 days) | 2.4 × 107 copies/g | FR-Eq73_Liver-Lung/FR/2013 |
| Foal #2 (2 months) | Serum | Alive (2 months) | 1.1 × 107 copies/mL | FR-Eq84_Serum/FR/2015 | |
| Case #2 (2014) | Foal (2 days) | Liver+Lung | Neonatal death (2 days) | 2.3 × 107 copies/g | FR-Eq69_Liver-Lung/FR/2014 |
| Mare (5 years) | Allantochorion | n.a. | 1.5 × 104 copies/g | FR-Eq69_Allanto/FR/2014 | |
| Serum | Alive (10 months) | 7.7 × 107 copies/ml | FR-Eq74_Serum/FR/2015 | ||
| Case #3 (2015) | Foal (2 months) | Serum | Alive (50 days) | Negative | n.a. |
| Mare (7 years) | Allantochorion | n.a. | 3.9 × 104 copies/g | FR-Eq70_Allanto/FR/2015 | |
| Serum | Alive | 1.8 × 103 copies/mL | FR-Eq85_Serum/FR/2015 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pronost, S.; Fortier, C.; Marcillaud-Pitel, C.; Tapprest, J.; Foursin, M.; Saunier, B.; Pitel, P.-H.; Paillot, R.; Hue, E.S. Further Evidence for in Utero Transmission of Equine Hepacivirus to Foals. Viruses 2019, 11, 1124. https://doi.org/10.3390/v11121124
Pronost S, Fortier C, Marcillaud-Pitel C, Tapprest J, Foursin M, Saunier B, Pitel P-H, Paillot R, Hue ES. Further Evidence for in Utero Transmission of Equine Hepacivirus to Foals. Viruses. 2019; 11(12):1124. https://doi.org/10.3390/v11121124
Chicago/Turabian StylePronost, Stephane, Christine Fortier, Christel Marcillaud-Pitel, Jackie Tapprest, Marc Foursin, Bertrand Saunier, Pierre-Hugues Pitel, Romain Paillot, and Erika S. Hue. 2019. "Further Evidence for in Utero Transmission of Equine Hepacivirus to Foals" Viruses 11, no. 12: 1124. https://doi.org/10.3390/v11121124
APA StylePronost, S., Fortier, C., Marcillaud-Pitel, C., Tapprest, J., Foursin, M., Saunier, B., Pitel, P.-H., Paillot, R., & Hue, E. S. (2019). Further Evidence for in Utero Transmission of Equine Hepacivirus to Foals. Viruses, 11(12), 1124. https://doi.org/10.3390/v11121124







