PCV2 Regulates Cellular Inflammatory Responses through Dysregulating Cellular miRNA-mRNA Networks
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Virus
2.2. Cell Culture and Treatment
2.3. RNA Isolation, Sequencing, and Data Analysis
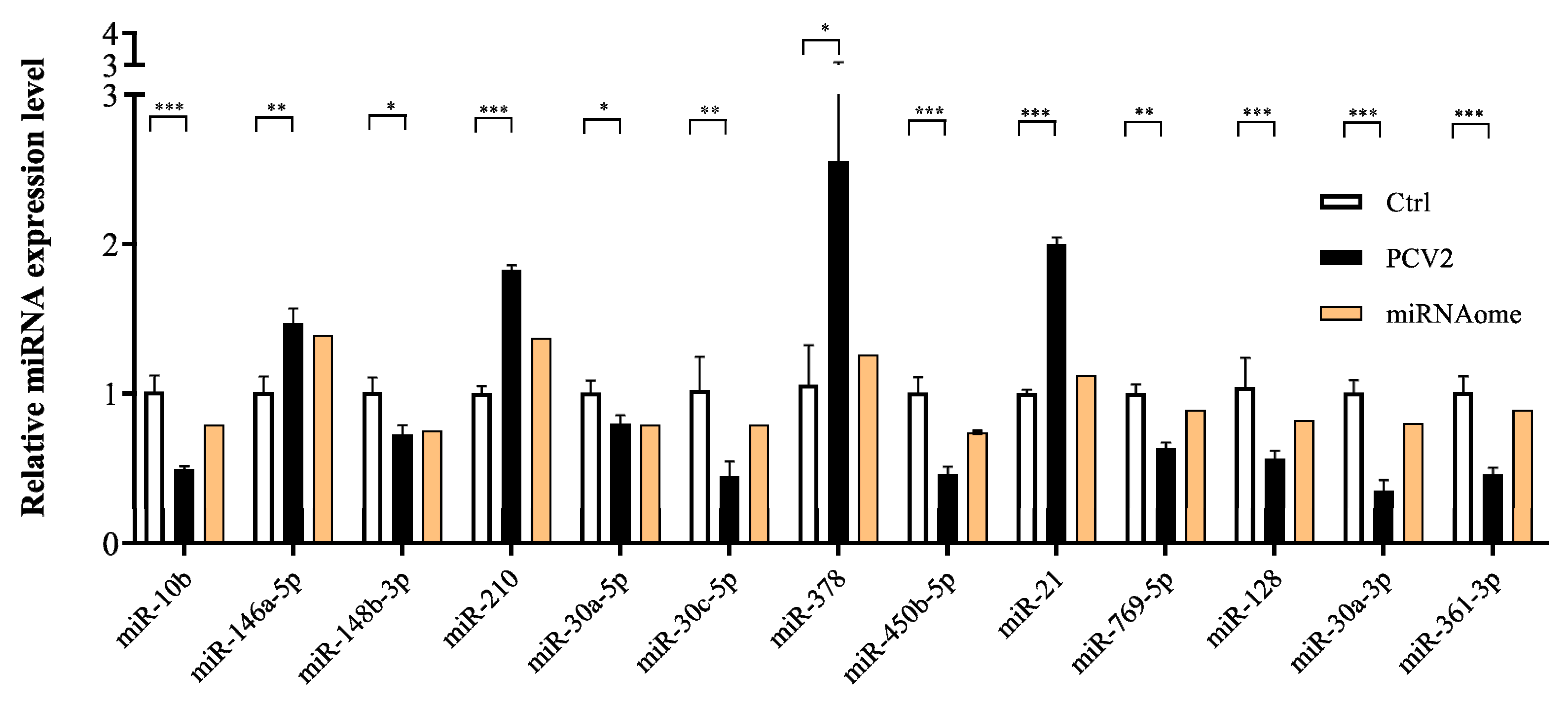
2.4. Validation of DE miRNAs/mRNA with the RT-qPCR Method
2.5. MiRNAs Target Prediction and Functional Enrichment Analysis
2.6. Construction of the miRNA-mRNA Network
3. Results
3.1. PCV2 Replication Kinetics in PK-15 Cells
3.2. Small RNA Sequencing Analysis
3.3. PCV2 Encoded miRNA Capability Evaluation
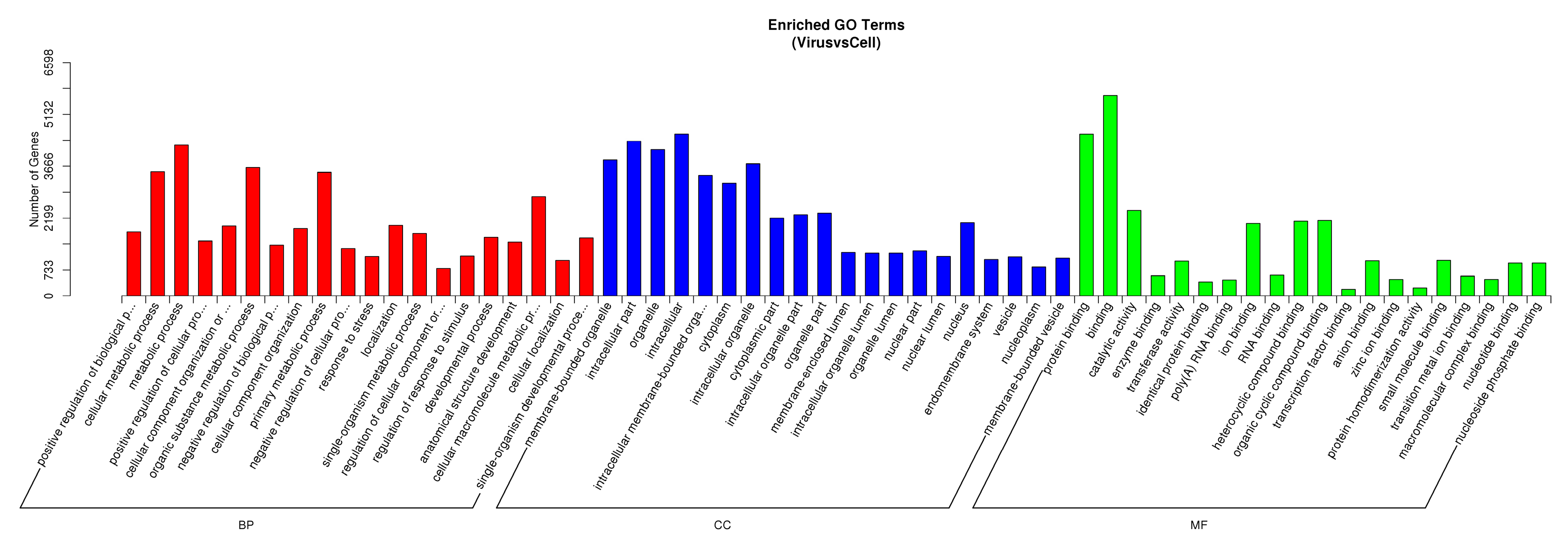
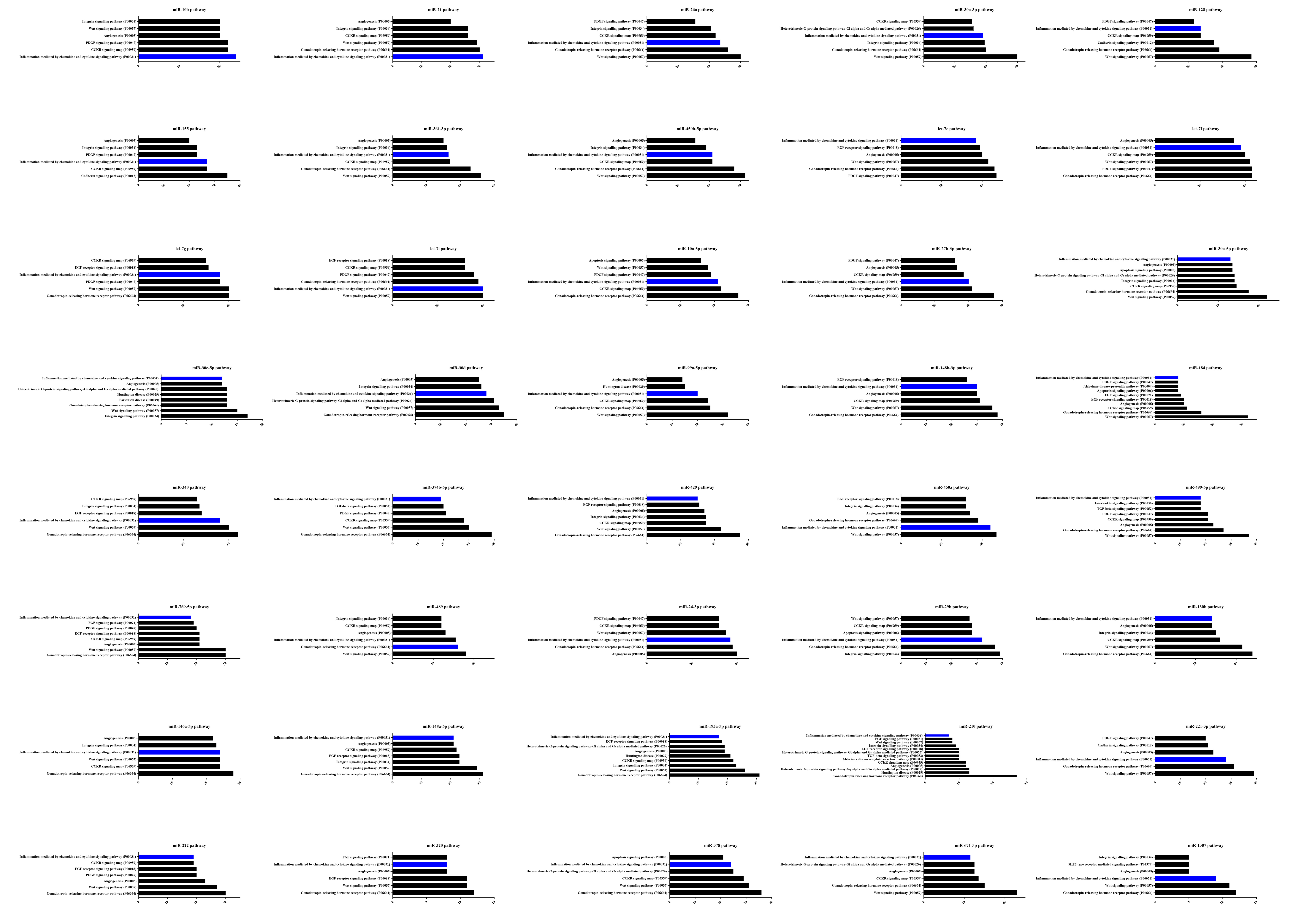
3.4. DE miRNA Target Genes Prediction and Functional Analysis
3.5. Construction of the miRNA-mRNA Network
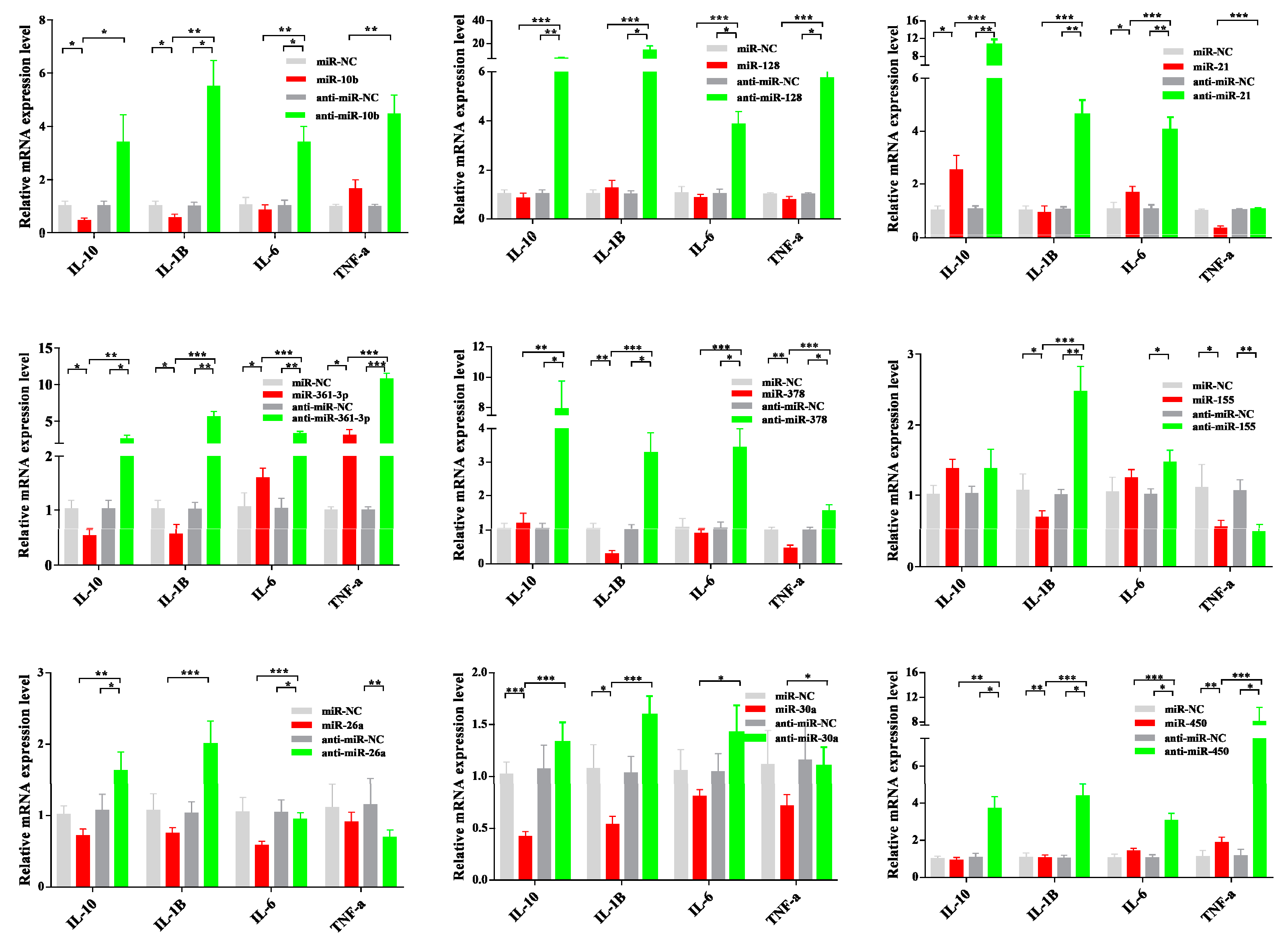
3.6. Dysregulated miRNA Expression Influences PCV2-Triggered Cellular Inflammatory Response
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ellis, J. Porcine circovirus: A historical perspective. Vet. Pathol. 2014, 51, 315–327. [Google Scholar] [CrossRef]
- Meng, X. Porcine Circovirus Type 2 (PCV2): Pathogenesis and Interaction with the Immune System. Annu. Rev. Anim. Biosci. 2013, 1, 43–66. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Wang, D.; Wang, J.; Zhu, S.; She, R.; Ren, X.; Tian, J.; Quan, R.; Hou, L.; Li, Z.; et al. Induction of porcine dermatitis and nephropathy syndrome in piglets by infection with porcine circovirus type 3. J. Virol. 2018, 93. [Google Scholar] [CrossRef] [PubMed]
- Darwich, L.; Segal, J. Pathogenesis of postweaning multisystemic wasting syndrome caused by Porcine circovirus 2: An immune riddle Brief Review. Arch. Virol. 2004, 149, 857–874. [Google Scholar] [CrossRef] [PubMed]
- Opriessnig, T.; Meng, X.; Halbur, P.G. Porcine circovirus type 2—Associated disease: Update on current terminology, clinical manifestations, pathogenesis, diagnosis and intervention strategies. J. Vet. Diagn. Investig. 2007, 615, 591–615. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Liu, S.; Wang, Y.; Deng, F.; Yan, W.; Yang, K.; Chen, H.; He, Q.; Charreyre, C.; Audoneet, J. Transcription analysis of the porcine alveolar macrophage response to porcine circovirus type 2. BMC Genom. 2013, 14, 1. [Google Scholar] [CrossRef]
- Shivdasani, R.A. Review in translational hematology MicroRNAs: Regulators of gene expression and cell differentiation. Blood 2015, 108, 3646–3654. [Google Scholar] [CrossRef]
- Song, G.; Sharma, A.D.; Roll, G.R.; Ng, R.; Lee, A.Y.; Blelloch, R.H.; Frandsen, N.M.; Willenbring, H. MicroRNAs Control Hepatocyte Proliferation During Liver Regeneration. Hepatology 2010, 51, 1735–1743. [Google Scholar] [CrossRef]
- Suh, Y.S.; Bhat, S.; Hong, S.; Shin, M.; Bahk, S.; Cho, K.S.; Kim, S.; Lee, K.; Kim, Y.; Jones, W.D.; et al. Genome-wide microRNA screening reveals that the evolutionary conserved miR-9a regulates body growth by targeting sNPFR1/NPYR. Nat. Commun. 2015, 6, 1–11. [Google Scholar] [CrossRef]
- Gomez, I.G.; Chau, B.N.; Duffield, J.S.; Gomez, I.G.; Mackenna, D.A.; Johnson, B.G.; Kaimal, V.; Roach, A.M.; Ren, S.; Nakagawa, N.; et al. Anti-microRNA-21 oligonucleotides prevent Alport nephropathy progression by stimulating metabolic pathways Find the latest version: Anti-microRNA-21 oligonucleotides prevent Alport nephropathy progression by stimulating metabolic pathways. J. Clin. Investig. 2015, 125, 141–156. [Google Scholar] [CrossRef]
- Meydan, C.; Shenhar-Tsarfaty, S.; Soreq, H. MicroRNA Regulators of Anxiety and Metabolic Disorders. Trends Mol. Med. 2016, 22, 798–812. [Google Scholar] [CrossRef] [PubMed]
- Cheng, A.M.; Byrom, M.W.; Shelton, J.; Ford, L.P. Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res. 2005, 33, 1290–1297. [Google Scholar] [CrossRef] [PubMed]
- Skalsky, R.L.; Cullen, B.R. Viruses, microRNAs, and Host Interactions Rebecca. Annu. Rev. Microbiol. 2013, 64, 123–141. [Google Scholar] [CrossRef] [PubMed]
- Núñez-Hernández, F.; Pérez, L.J.; Vera, G.; Córdoba, S.; Segalés, J.; Sánchez, A.; Núñez, J.I. Evaluation of the capability of the PCV2 genome to encode miRNAs: Lack of viral miRNA expression in an experimental infection. Vet. Res. 2015, 46, 1–5. [Google Scholar] [CrossRef]
- Núñez-Hernández, F.; Pérez, L.J.; Muñoz, M.; Vera, G.; Tomás, A.; Egea, R.; Córdoba, S.; Segalés, J.; Sánchez, A.; Núñez, J.I. Identification of microRNAs in PCV2 subclinically infected pigs by high throughput sequencing. Vet. Res. 2015, 46, 1–7. [Google Scholar] [CrossRef]
- Hong, J.-S.; Kim, N.-H.; Choi, C.-Y.; Lee, J.-S.; Na, D.; Chun, T.; Lee, Y.S. Changes in cellular microRNA expression induced by porcine circovirus type 2-encoded proteins. Vet. Res. 2015, 46, 1–14. [Google Scholar] [CrossRef][Green Version]
- Zhang, P.; Wang, L.; Li, Y.; Jiang, P.; Wang, Y.; Wang, P.; Kang, L.; Wang, Y. Identification and characterization of microRNA in the lung tissue of pigs with different susceptibilities to PCV2 infection. Vet. Res. 2018, 49, 1–13. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Gao, D.; Middleton, R.; Rasko, J.E.J.; Ritchie, W. miREval 2. 0: A web tool for simple microRNA prediction in genome sequences. Bioinformatics 2013, 29, 3225–3226. [Google Scholar] [CrossRef]
- Li, S.C.; Shiau, C.K.; Lin, W.C. Vir-Mir db: Prediction of viral microRNA candidate hairpins. Nucleic Acids Res. 2008, 36, 184–189. [Google Scholar] [CrossRef]
- Friedlander, M.R.; Mackowiak, S.D.; Li, N.; Chen, W.; Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012, 40, 37–52. [Google Scholar] [CrossRef]
- Zhou, L.; Chen, J.; Li, Z.; Li, X.; Hu, X.; Huang, Y.; Zhao, X.; Liang, C.; Wang, Y.; Sun, L.; et al. Integrated Profiling of MicroRNAs and mRNAs: MicroRNAs Located on Xq27. 3 Associate with Clear Cell Renal Cell Carcinoma. PLoS ONE 2010, 5, e15224. [Google Scholar] [CrossRef] [PubMed]
- Xiao, F.; Zuo, Z.; Cai, G.; Kang, S.; Gao, X.; Li, T. miRecords: An integrated resource for microRNA-target interactions. Nucleic Acids Res. 2009, 37, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Huang, X.; Muruganujan, A.; Tang, H.; Mills, C.; Kang, D.; Thomas, P.D. PANTHER version 11: Expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 2017, 45, D183–D189. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Haasnoot, J.; Berkhout, B. RNAi and Cellular miRNAs in Infections by Mammalian Viruses. Methods Mol. Biol. 2011, 721, 23–41. [Google Scholar]
- Shrinet, J.; Jain, S.; Jain, J.; Bhatnagar, R.K.; Sunil, S. Next Generation Sequencing Reveals Regulation of Distinct Aedes microRNAs during Chikungunya Virus Development. PLoS ONE 2014, 8, e2616. [Google Scholar] [CrossRef]
- Gkirtzou, K.; Tsamardinos, I.; Tsakalides, P.; Poirazi, P. MatureBayes: A Probabilistic Algorithm for Identifying the Mature miRNA within Novel Precursors. PLoS ONE 2010, 5, e11843. [Google Scholar] [CrossRef]
- Xuan, P.; Guo, M.; Huang, Y.; Li, W.; Huang, Y. MaturePred: Efficient identification of microRNAs within novel plant pre-miRNAs. PLoS ONE 2011, 6, e27422. [Google Scholar] [CrossRef]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.; Zhang, M.; Li, W.; Wang, Y.; Liu, Y.; He, Q. Proteomic analysis of porcine alveolar macrophages infected with porcine circovirus type 2. J. Proteomics 2012, 75, 3258–3269. [Google Scholar] [CrossRef] [PubMed]
- Gaudet, A.D.; Fonken, L.K.; Watkins, L.R.; Nelson, R.J.; Popovich, P.G. MicroRNAs: Roles in Regulating Neuroinflammation. Neuroscientist 2017, 24, 221–245. [Google Scholar] [CrossRef] [PubMed]
- Sheedy, F.J. Turning 21: Induction of miR-21 as a key switch in the inflammatory response. Front. Immunol. 2015, 6, 1–9. [Google Scholar] [CrossRef]
- Lin, Z.; Ge, J.; Wang, Z.; Ren, J.; Wang, X.; Xiong, H.; Gao, J. Let-7e modulates the inflammatory response in vascular endothelial cells through ceRNA crosstalk. NSci Rep. 2017, 7, 42498. [Google Scholar] [CrossRef]
- Hussain, N.; Zhu, W.; Jiang, C.; Xu, J.; Wu, X. Down-regulation of miR-10a-5p in synoviocytes contributes to TBX5-controlled joint inflammation. J. Cell Mol. Med. 2017, 22, 241–250. [Google Scholar] [CrossRef]
- Jiang, C.; Yu, H.; Sun, Q.; Zhu, W.; Xu, J.; Gao, N.; Zhang, R.; Liu, L.; Wu, X.; Yang, X.; et al. Extracellular microRNA-21 and microRNA- 26a increase in body fluids from rats with antigen induced pulmonary inflammation and children with recurrent wheezing. BMC Pulm. Med. 2016, 16, 50. [Google Scholar] [CrossRef]
- Liang, S.; Song, Z.; Wu, Y.; Gao, Y. MicroRNA-27b Modulates Inflammatory Response and Apoptosis during Mycobacterium tuberculosis Infection. J. Immunol. 2018, 200, 3506–3518. [Google Scholar] [CrossRef]
- Koh, E.H.; Chernis, N.; Saha, P.K.; Xiao, L.; Bader, D.A.; Zhu, B.; Rajapakshe, K.; Hamilton, M.P.; Liu, X.; Perera, D.; et al. miR-30a remodels subcutaneous adipose tissue inflammation to improve insulin sensitivity in obesity. Diabetes 2018, 67, 2541–2553. [Google Scholar] [CrossRef]
- Fang, X.; Sun, D.; Wang, Z.; Yu, Z.; Liu, W. MiR-30a Positively Regulates the Inflammatory Response of Microglia in Experimental Autoimmune Encephalomyelitis. Neurosci. Bull. 2017, 33, 603–615. [Google Scholar] [CrossRef]
- Radu, C.; Simioni, P.; Garlaschelli, K.; Baragetti, A.; Catapano, A.L.; Iori, E.; Fadini, G.P.; Avogaro, A. miR-30c-5p regulates macrophage-mediated inflammation and pro-atherosclerosis pathways. Cardiovasc. Res. 2017, 113, 1627–1638. [Google Scholar]
- Bao, M.; Li, J.; Luo, H.; Tang, L.; Lv, Q.; Li, G.; Zhou, H. NF- κB-Regulated miR-99a Modulates Endothelial Cell Inflammation. Mediat. Inflamm. 2016, 2016, 53081. [Google Scholar] [CrossRef] [PubMed]
- Ong, C.; Yung, L.L.; Dheen, S.T. miR-128 regulates genes associated with inflammation and fibrosis of rat kidney cells in vitro. Anat. Rec. 2018, 301, 913–921. [Google Scholar]
- Xiao, J.; Tang, J.; Chen, Q.; Tang, D.; Liu, M.; Luo, M.; Wang, Y.; Wang, J.; Zhao, Z.; Tang, C.; et al. miR-429 regulates alveolar macrophage inflammatory cytokine production and is involved in LPS-induced acute lung injury. Biochem. J. 2015, 471, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Ma, J.; Li, T.; Han, X. MiR-29b aggravates lipopolysaccharide-induced endothelial cells in flammatory damage by regulation of NF- κ B and JNK signaling pathways. Biomed. Pharmacother. 2018, 99, 451–461. [Google Scholar] [CrossRef]
- Zhang, M.; Zhou, Z.; Wang, J.; Li, S. MiR-130b promotes obesity associated adipose tissue inflammation and insulin resistance in diabetes mice through alleviating M2 macrophage polarization via repression of PPAR- γ. Immunol. Lett. 2016, 180, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Zhang, X.; Li, F.; Yuan, K.; Li, M.; Zhang, J.; Li, B. International Immunopharmacology MiR-130b attenuates vascular in fl ammation via negatively regulating tumor progression locus 2 (Tpl2) expression. Int. Immunopharmacol. 2017, 51, 9–16. [Google Scholar] [CrossRef]
- Zhang, T.; Hu, J.; Wang, X.; Zhao, X.; Li, Z.; Niu, J.; Steer, C.J.; Zheng, G.; Song, G. MicroRNA-378 Promotes Hepatic Inflammation and Fibrosis via Modulation of the NF κB - TNFα Pathway. J. Hepatol. 2018, 70, 87–96. [Google Scholar] [CrossRef]






© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, C.; Sun, Y.; Li, J.; Jiang, C.; Zeng, W.; Zhang, H.; Fan, S.; He, Q. PCV2 Regulates Cellular Inflammatory Responses through Dysregulating Cellular miRNA-mRNA Networks. Viruses 2019, 11, 1055. https://doi.org/10.3390/v11111055
Li C, Sun Y, Li J, Jiang C, Zeng W, Zhang H, Fan S, He Q. PCV2 Regulates Cellular Inflammatory Responses through Dysregulating Cellular miRNA-mRNA Networks. Viruses. 2019; 11(11):1055. https://doi.org/10.3390/v11111055
Chicago/Turabian StyleLi, Chang, Yumei Sun, Jing Li, Changsheng Jiang, Wei Zeng, Hao Zhang, Shengxian Fan, and Qigai He. 2019. "PCV2 Regulates Cellular Inflammatory Responses through Dysregulating Cellular miRNA-mRNA Networks" Viruses 11, no. 11: 1055. https://doi.org/10.3390/v11111055
APA StyleLi, C., Sun, Y., Li, J., Jiang, C., Zeng, W., Zhang, H., Fan, S., & He, Q. (2019). PCV2 Regulates Cellular Inflammatory Responses through Dysregulating Cellular miRNA-mRNA Networks. Viruses, 11(11), 1055. https://doi.org/10.3390/v11111055



