Improved Immune Responses Against Zika Virus After Sequential Dengue and Zika Virus Infection in Humans
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Human Blood Samples
2.3. Viruses and Cell Lines
2.4. HLA Typing
2.5. Serology
2.6. RT-PCR Assays for Detection of DENV and ZIKV
2.7. Viral Sequences
2.8. Peptides
2.9. Ex Vivo IFN-γ ELISPOT Assay
2.10. Immunogenicity and HLA Restrictions Prediction
2.11. Statistics
3. Results
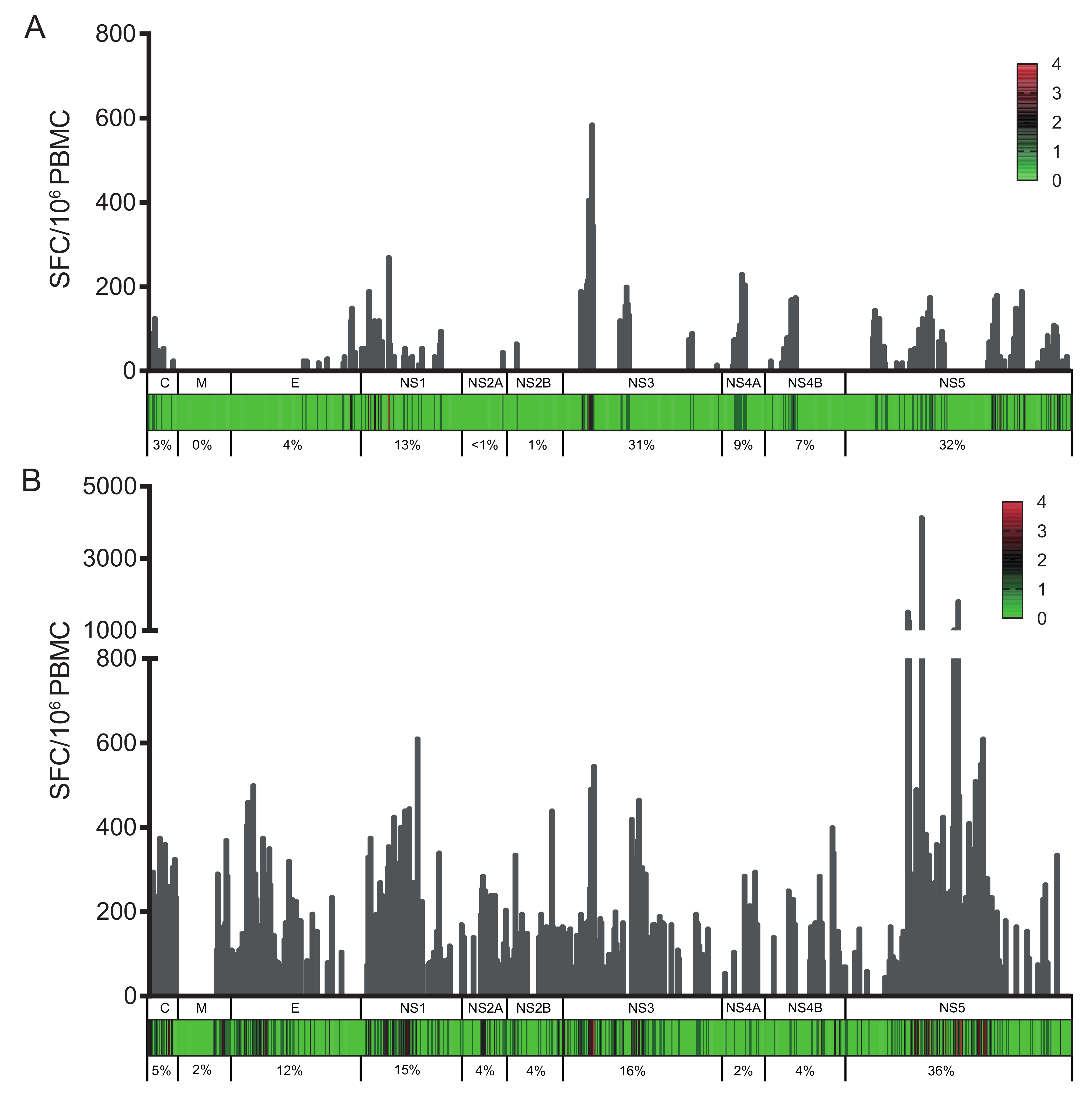
3.1. Identification of Immunodominant Regions of the ZIKV Proteome
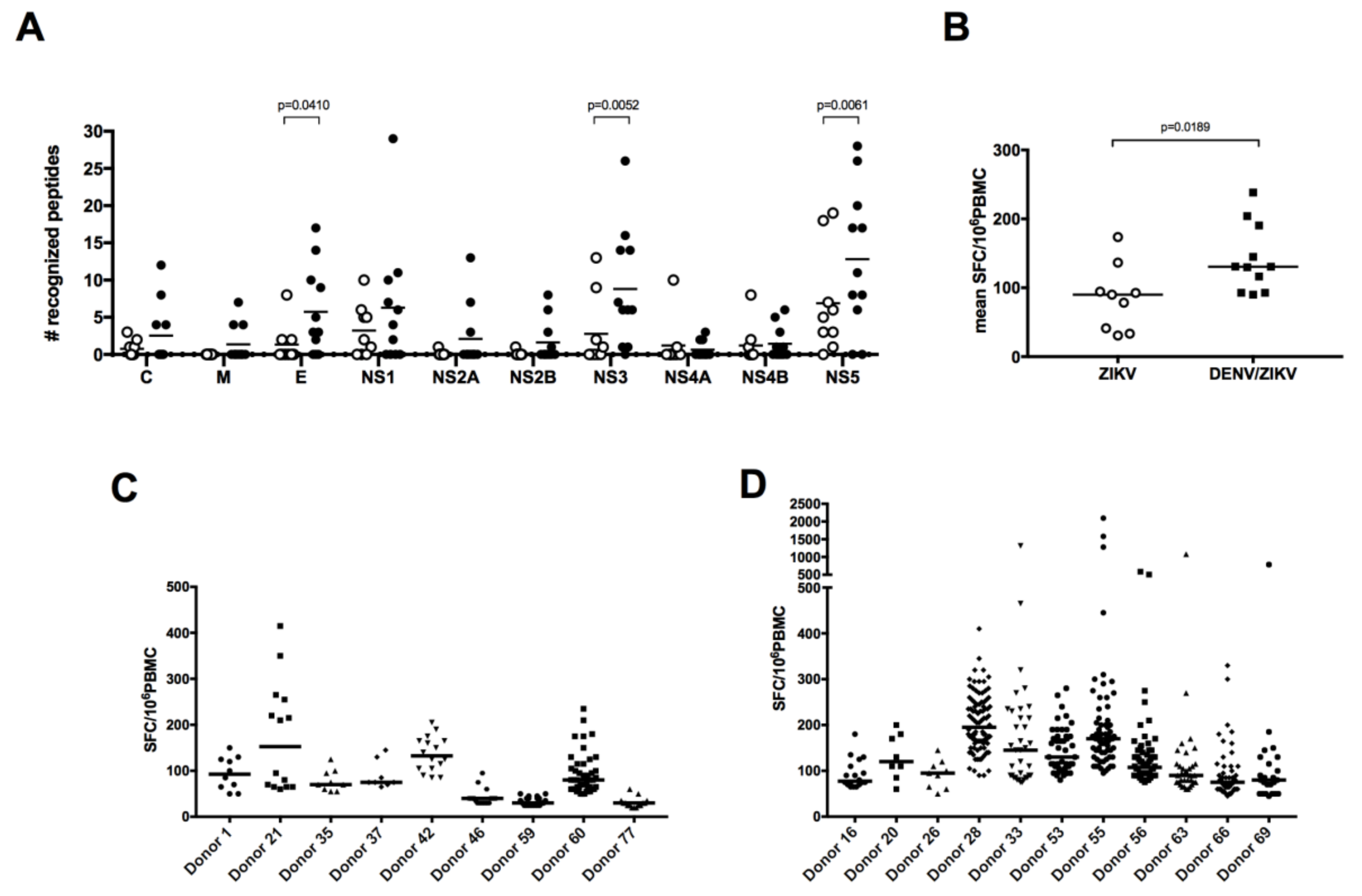
3.2. Broader Responses with a Higher Magnitude in DENV/ZIKV-Immune Donors
3.3. DENV/ZIKV-Cross-Reactive T Cells Mainly Target the NS5 Protein
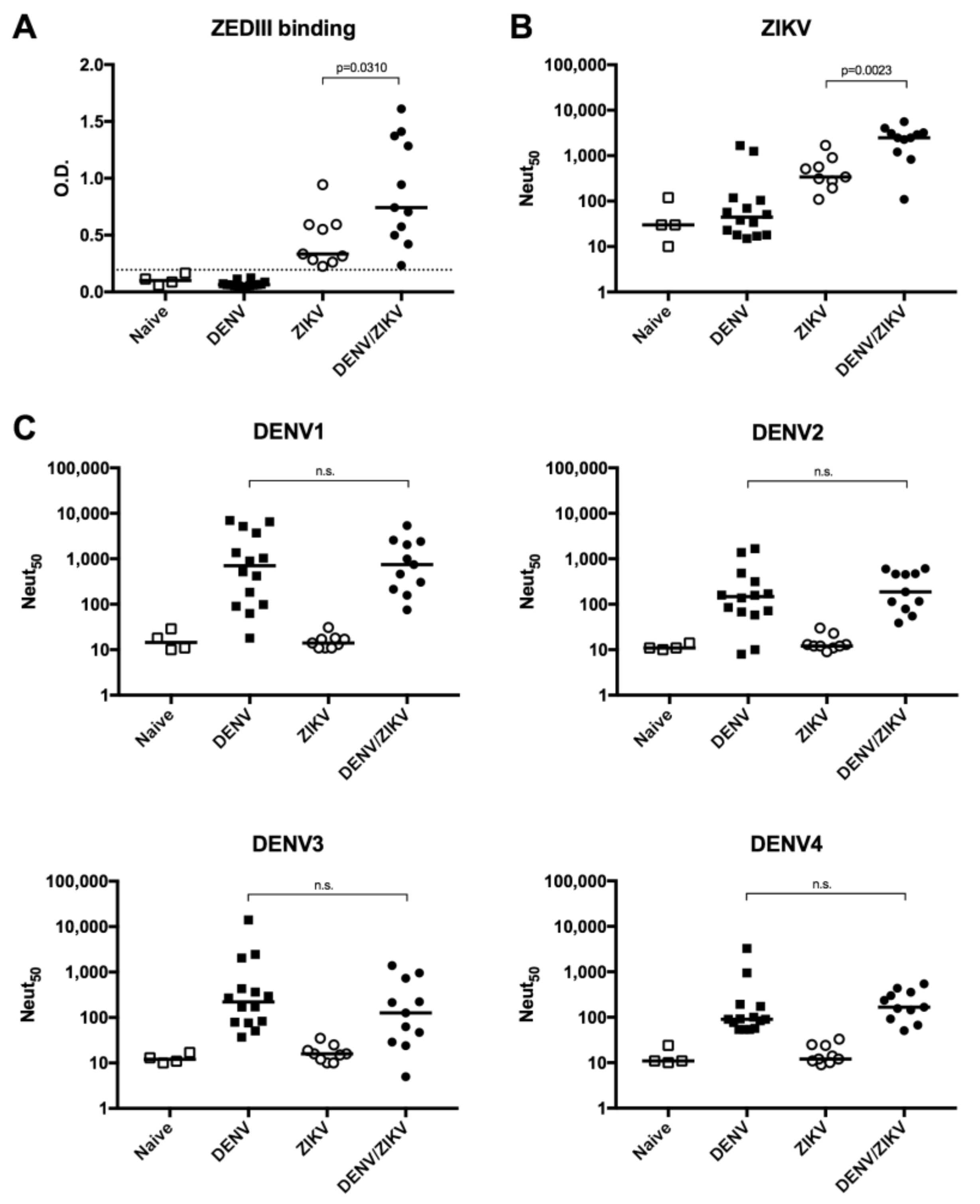
3.4. Increased Neutralizing Antibody Titer against ZIKV in DENV/ZIKV-Immune Donors
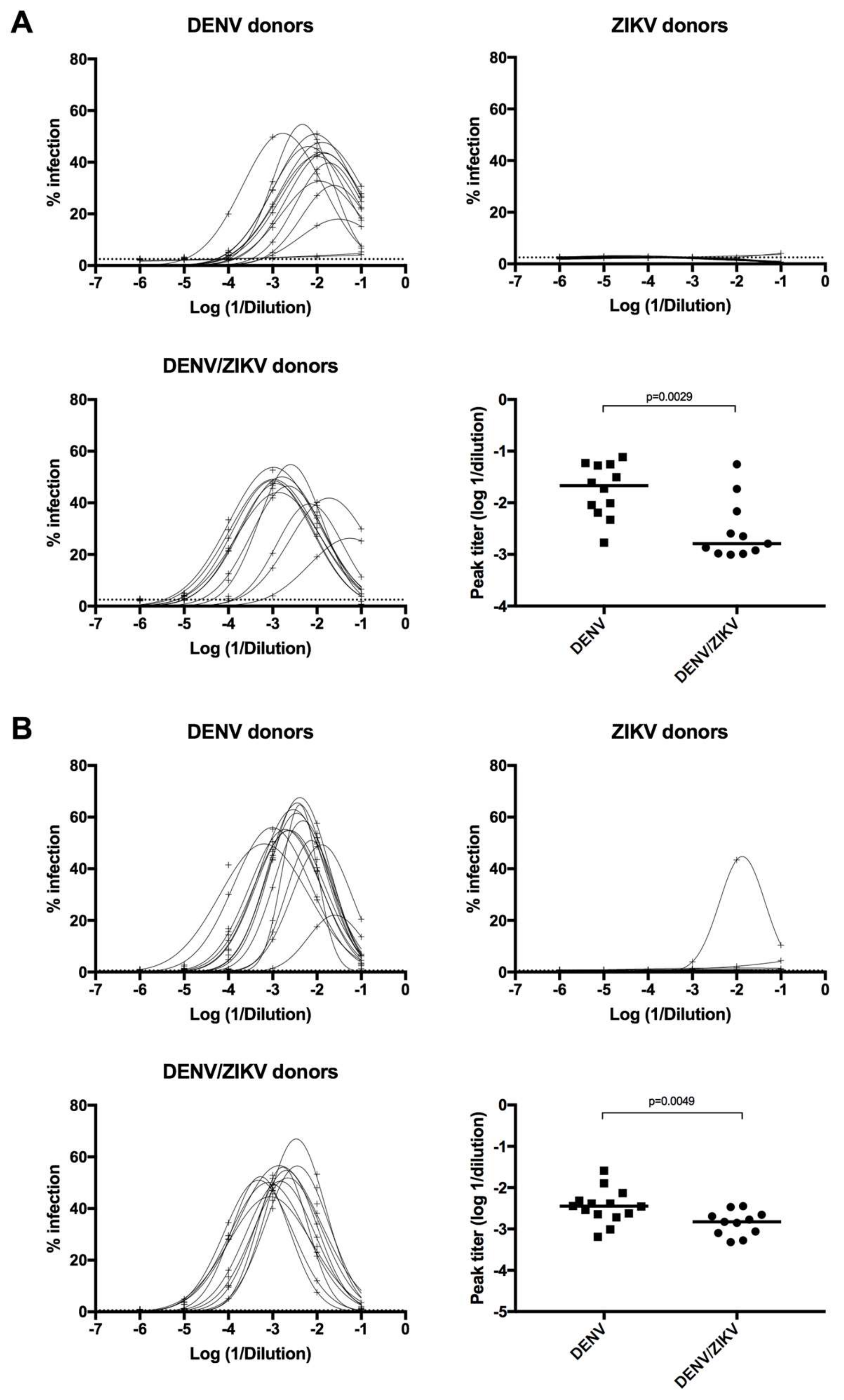
3.5. Decreased Enhancing Potential on ZIKV Infection in DENV/ZIKV-Immune Donors
3.6. High Titer of Neutralizing Antibodies with Low Enhancing Activity are Associated with Strong T-Cell Responses
4. Discussion
5. Patents
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kuno, G.; Chang, G.J.; Tsuchiya, K.R.; Karabatsos, N.; Cropp, C.B. Phylogeny of the genus Flavivirus. J. Virol. 1998, 72, 73–83. [Google Scholar] [PubMed]
- Dick, G.W.; Kitchen, S.F.; Haddow, A.J. Zika virus. I. Isolations and serological specificity. Trans. R Soc. Trop. Med. Hyg. 1952, 46, 509–520. [Google Scholar] [CrossRef]
- Duffy, M.R.; Chen, T.H.; Hancock, W.T.; Powers, A.M.; Kool, J.L.; Lanciotti, R.S.; Pretrick, M.; Marfel, M.; Holzbauer, S.; Dubray, C.; et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. N. Engl. J. Med. 2009, 360, 2536–3543. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Roche, C.; Teissier, A.; Robin, E.; Berry, A.L.; Mallet, H.P.; Sall, A.A.; Musso, D. Zika virus, French polynesia, South pacific, 2013. Emerg. Infect. Dis. 2014, 20, 1085–1086. [Google Scholar] [CrossRef] [PubMed]
- Campos, G.S.; Bandeira, A.C.; Sardi, S.I. Zika Virus Outbreak, Bahia, Brazil. Emerg. Infect. Dis. 2015, 21, 1885–1886. [Google Scholar] [CrossRef] [PubMed]
- Dupont-Rouzeyrol, M.; O’Connor, O.; Calvez, E.; Daures, M.; John, M.; Grangeon, J.P.; Gourinat, A.C. Co-infection with Zika and dengue viruses in 2 patients, New Caledonia, 2014. Emerg. Infect. Dis. 2015, 21, 381–382. [Google Scholar] [CrossRef] [PubMed]
- Zanluca, C.; Melo, V.C.; Mosimann, A.L.; Santos, G.I.; Santos, C.N.; Luz, K. First report of autochthonous transmission of Zika virus in Brazil. Mem. Inst. Oswaldo Cruz. 2015, 110, 569–572. [Google Scholar] [CrossRef] [PubMed]
- Pacheco, O.; Beltran, M.; Nelson, C.A.; Valencia, D.; Tolosa, N.; Farr, S.L.; Padilla, A.V.; Tong, V.T.; Cuevas, E.L.; Espinosa-Bode, A.; et al. Zika virus disease in Colombia-preliminary report. N. Engl. J. Med. 2016. [Google Scholar] [CrossRef] [PubMed]
- Metsky, H.C.; Matranga, C.B.; Wohl, S.; Schaffner, S.F.; Freije, C.A.; Winnicki, S.M.; West, K.; Qu, J.; Baniecki, M.L.; Gladden-Young, A.; et al. Zika virus evolution and spread in the Americas. Nature 2017, 546, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Oehler, E.; Watrin, L.; Larre, P.; Leparc-Goffart, I.; Lastere, S.; Valour, F.; Baudouin, L.; Mallet, H.; Musso, D.; Ghawche, F. Zika virus infection complicated by Guillain-Barre syndrome--case report, French Polynesia, December 2013. Eurosurveillance 2014, 19, 20720. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Blake, A.; Mons, S.; Lastere, S.; Roche, C.; Vanhomwegen, J.; Dub, T.; Baudouin, L.; Teissier, A.; Larre, P.; et al. Guillain-Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: A case-control study. Lancet. 2016, 387, 1531–1539. [Google Scholar] [CrossRef]
- Cauchemez, S.; Besnard, M.; Bompard, P.; Dub, T.; Guillemette-Artur, P.; Eyrolle-Guignot, D.; Salje, H.; Van Kerkhove, M.D.; Abadie, V.; Garel, C.; et al. Association between Zika virus and microcephaly in French Polynesia, 2013-15: A retrospective study. Lancet. 2016, 387, 2125–2132. [Google Scholar] [CrossRef]
- Soares de Araujo, J.S.; Regis, C.T.; Gomes, R.G.; Tavares, T.R.; Rocha Dos Santos, C.; Assuncao, P.M.; Nobrega, R.V.; Pinto, D.F.; Bezerra, B.V.; Mattos, S.D. Microcephaly in north-east Brazil: A retrospective study on neonates born between 2012 and 2015. Bull. World Health Organ. 2016, 94, 835–840. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, J.Y.; Du, S.Y.; Shan, C.; Nie, K.X.; Zhang, R.; Li, X.F.; Zhang, R.; Wang, T.; Qin, C.F.; et al. Evolutionary enhancement of Zika virus infectivity in Aedes aegypti mosquitoes. Nature 2017, 545, 482–486. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.; Luo, H.; Shan, C.; Muruato, A.E.; Nunes, B.T.D.; Medeiros, D.B.A.; Zou, J.; Xie, X.; Giraldo, M.I.; Vasconcelos, P.F.C.; et al. An evolutionary NS1 mutation enhances Zika virus evasion of host interferon induction. Nat. Commun. 2018, 9, 414. [Google Scholar] [CrossRef] [PubMed]
- Katzelnick, L.C.; Coloma, J.; Harris, E. Dengue: Knowledge gaps, unmet needs, and research priorities. Lancet Infect. Dis. 2017, 17, e88–e100. [Google Scholar] [CrossRef]
- Dejnirattisai, W.; Supasa, P.; Wongwiwat, W.; Rouvinski, A.; Barba-Spaeth, G.; Duangchinda, T.; Sakuntabhai, A.; Cao-Lormeau, V.M.; Malasit, P.; Rey, F.A.; et al. Dengue virus sero-cross-reactivity drives antibody-dependent enhancement of infection with zika virus. Nat. Immunol. 2016, 17, 1102–1108. [Google Scholar] [CrossRef] [PubMed]
- Stettler, K.; Beltramello, M.; Espinosa, D.A.; Graham, V.; Cassotta, A.; Bianchi, S.; Vanzetta, F.; Minola, A.; Jaconi, S.; Mele, F.; et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science 2016, 353, 823–826. [Google Scholar] [CrossRef] [PubMed]
- Paul, L.M.; Carlin, E.R.; Jenkins, M.M.; Tan, A.L.; Barcellona, C.M.; Nicholson, C.O.; Michael, S.F.; Isern, S. Dengue virus antibodies enhance Zika virus infection. Clin. Transl. Immunol. 2016, 5, e117. [Google Scholar] [CrossRef] [PubMed]
- Priyamvada, L.; Quicke, K.M.; Hudson, W.H.; Onlamoon, N.; Sewatanon, J.; Edupuganti, S.; Pattanapanyasat, K.; Chokephaibulkit, K.; Mulligan, M.J.; Wilson, P.C.; et al. Human antibody responses after dengue virus infection are highly cross-reactive to Zika virus. Proc. Natl. Acad. Sci. USA 2016, 113, 7852–7857. [Google Scholar] [CrossRef] [PubMed]
- Bardina, S.V.; Bunduc, P.; Tripathi, S.; Duehr, J.; Frere, J.J.; Brown, J.A.; Nachbagauer, R.; Foster, G.A.; Krysztof, D.; Tortorella, D.; et al. Enhancement of Zika virus pathogenesis by preexisting antiflavivirus immunity. Science 2017, 356, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Grifoni, A.; Pham, J.; Sidney, J.; O’Rourke, P.H.; Paul, S.; Peters, B.; Martini, S.R.; de Silva, A.D.; Ricciardi, M.J.; Magnani, D.M.; et al. Prior Dengue virus exposure shapes T cell immunity to Zika virus in humans. J. Virol. 2017, 91, e01469-17. [Google Scholar] [CrossRef] [PubMed]
- Wen, J.; Tang, W.W.; Sheets, N.; Ellison, J.; Sette, A.; Kim, K.; Shresta, S. Identification of Zika virus epitopes reveals immunodominant and protective roles for dengue virus cross-reactive CD8+ T cells. Nat. Microbiol. 2017, 2, 17036. [Google Scholar] [CrossRef] [PubMed]
- Herrera, B.B.; Tsai, W.Y.; Chang, C.A.; Hamel, D.J.; Wang, W.K.; Lu, Y.; Mboup, S.; Kanki, P.J. Sustained specific and cross-reactive T cell responses to Zika and Dengue virus NS3 in West Africa. J. Virol. 2018, 92, e01992-17. [Google Scholar] [CrossRef] [PubMed]
- Weiskopf, D.; Angelo, M.A.; de Azeredo, E.L.; Sidney, J.; Greenbaum, J.A.; Fernando, A.N.; Broadwater, A.; Kolla, R.V.; De Silva, A.D.; de Silva, A.M.; et al. Comprehensive analysis of dengue virus-specific responses supports an HLA-linked protective role for CD8+ T cells. Proc. Natl. Acad. Sci. USA 2013, 110, e2046–e2053. [Google Scholar] [CrossRef] [PubMed]
- Weiskopf, D.; Bangs, D.J.; Sidney, J.; Kolla, R.V.; De Silva, A.D.; de Silva, A.M.; Crotty, S.; Peters, B.; Sette, A. Dengue virus infection elicits highly polarized CX3CR1+ cytotoxic CD4+ T cells associated with protective immunity. Proc. Natl. Acad. Sci. USA 2015, 112, e4256–4263. [Google Scholar] [CrossRef] [PubMed]
- Rivino, L. T cell immunity to dengue virus and implications for vaccine design. Expert Rev. Vaccines 2016, 15, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Alam, A.; Ali, S.; Ahamad, S.; Malik, M.Z.; Ishrat, R. From ZikV genome to vaccine: In silico approach for the epitope-based peptide vaccine against Zika virus envelope glycoprotein. Immunology 2016, 149, 386–399. [Google Scholar] [CrossRef] [PubMed]
- Dar, H.; Zaheer, T.; Rehman, M.T.; Ali, A.; Javed, A.; Khan, G.A.; Babar, M.M.; Waheed, Y. Prediction of promiscuous T-cell epitopes in the Zika virus polyprotein: An in silico approach. Asian Pac. J. Trop. Med. 2016, 9, 844–850. [Google Scholar] [CrossRef] [PubMed]
- Usman Mirza, M.; Rafique, S.; Ali, A.; Munir, M.; Ikram, N.; Manan, A.; Salo-Ahen, O.M.; Idrees, M. Towards peptide vaccines against Zika virus: Immunoinformatics combined with molecular dynamics simulations to predict antigenic epitopes of Zika viral proteins. Sci. Rep. 2016, 6, 37313. [Google Scholar] [CrossRef] [PubMed]
- Elong Ngono, A.; Vizcarra, E.A.; Tang, W.W.; Sheets, N.; Joo, Y.; Kim, K.; Gorman, M.J.; Diamond, M.S.; Shresta, S. Mapping and role of the CD8+ T cell response during primary Zika virus infection in mice. Cell Host Microbe 2017, 21, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Li, S.; Zhang, Y.; Han, X.; Jia, B.; Liu, H.; Liu, D.; Tan, S.; Wang, Q.; Bi, Y.; et al. CD8+ T cell immune response in immunocompetent mice during Zika virus infection. J. Virol. 2017, 91, e00900-17. [Google Scholar] [CrossRef] [PubMed]
- Pardy, R.D.; Rajah, M.M.; Condotta, S.A.; Taylor, N.G.; Sagan, S.M.; Richer, M.J. Analysis of the T cell response to Zika Virus and identification of a novel CD8+ T cell epitope in immunocompetent mice. PLoS Pathog. 2017, 13, e1006184. [Google Scholar] [CrossRef] [PubMed]
- Winkler, C.W.; Myers, L.M.; Woods, T.A.; Messer, R.J.; Carmody, A.B.; McNally, K.L.; Scott, D.P.; Hasenkrug, K.J.; Best, S.M.; Peterson, K.E. Adaptive immune responses to Zika virus are important for controlling virus infection and preventing infection in brain and testes. J. Immunol. 2017, 198, 3526–3535. [Google Scholar] [CrossRef] [PubMed]
- Aubry, M.; Teissier, A.; Huart, M.; Merceron, S.; Vanhomwegen, J.; Roche, C.; Vial, A.L.; Teururai, S.; Sicard, S.; Paulous, S.; et al. Zika virus seroprevalence, French Polynesia, 2014–2015. Emerg. Infect. Dis. 2017, 23, 669–672. [Google Scholar] [CrossRef] [PubMed]
- De Alwis, R.; Williams, K.L.; Schmid, M.A.; Lai, C.Y.; Patel, B.; Smith, S.A.; Crowe, J.E.; Wang, W.K.; Harris, E.; de Silva, A.M. Dengue viruses are enhanced by distinct populations of serotype cross-reactive antibodies in human immune sera. PLoS Pathog. 2014, 10, e1004386. [Google Scholar] [CrossRef] [PubMed]
- De Alwis, R.; de Silva, A.M. Measuring antibody neutralization of dengue virus (DENV) using a flow cytometry-based technique. Methods Mol. Biol. 2014, 1138, 27–39. [Google Scholar] [PubMed]
- Calvo, E.P.; Sanchez-Quete, F.; Duran, S.; Sandoval, I.; Castellanos, J.E. Easy and inexpensive molecular detection of dengue, chikungunya and zika viruses in febrile patients. Acta Trop. 2016, 163, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Andreatta, M.; Karosiene, E.; Rasmussen, M.; Stryhn, A.; Buus, S.; Nielsen, M. Accurate pan–specific prediction of peptide-MHC class II binding affinity with improved binding core identification. Immunogenetics 2015, 67, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, M.; Andreatta, M. NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Med. 2016, 8, 33. [Google Scholar] [CrossRef] [PubMed]
- Simmons, C.P.; Dong, T.; Chau, N.V.; Dung, N.T.; Chau, T.N.; Thao le, T.T.; Dung, N.T.; Hien, T.T.; Rowland-Jones, S.; Farrar, J. Early T-cell responses to dengue virus epitopes in Vietnamese adults with secondary dengue virus infections. J. Virol. 2005, 79, 5665–5675. [Google Scholar] [CrossRef] [PubMed]
- Duangchinda, T.; Dejnirattisai, W.; Vasanawathana, S.; Limpitikul, W.; Tangthawornchaikul, N.; Malasit, P.; Mongkolsapaya, J.; Screaton, G. Immunodominant T-cell responses to dengue virus NS3 are associated with DHF. Proc. Natl. Acad. Sci. USA 2010, 107, 16922–16927. [Google Scholar] [CrossRef] [PubMed]
- Rivino, L.; Kumaran, E.A.; Jovanovic, V.; Nadua, K.; Teo, E.W.; Pang, S.W.; Teo, G.H.; Gan, V.C.; Lye, D.C.; Leo, Y.S.; et al. Differential targeting of viral components by CD4+ versus CD8+ T lymphocytes in dengue virus infection. J. Virol. 2013, 87, 2693–2706. [Google Scholar] [CrossRef] [PubMed]
- Weiskopf, D.; Angelo, M.A.; Bangs, D.J.; Sidney, J.; Paul, S.; Peters, B.; de Silva, A.D.; Lindow, J.C.; Diehl, S.A.; Whitehead, S.; et al. The Human CD8+ T Cell Responses Induced by a Live Attenuated Tetravalent Dengue Vaccine Are Directed against Highly Conserved Epitopes. J. Virol. 2015, 89, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Dikhit, M.R.; Ansari, M.Y.; Vijaymahantesh; Kalyani; Mansuri, R.; Sahoo, B.R.; Dehury, B.; Amit, A.; Topno, R.K.; Sahoo, G.C.; et al. Computational prediction and analysis of potential antigenic CTL epitopes in Zika virus: A first step towards vaccine development. Infect. Genet. Evol. 2016, 45, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Imrie, A.; Meeks, J.; Gurary, A.; Sukhbataar, M.; Kitsutani, P.; Effler, P.; Zhao, Z. Differential functional avidity of dengue virus-specific T-cell clones for variant peptides representing heterologous and previously encountered serotypes. J. Virol. 2007, 81, 10081–10091. [Google Scholar] [CrossRef] [PubMed]
- Tai, W.; He, L.; Wang, Y.; Sun, S.; Zhao, G.; Luo, C.; Li, P.; Zhao, H.; Fremont, D.H.; Li, F.; et al. Critical neutralizing fragment of Zika virus EDIII elicits cross-neutralization and protection against divergent Zika viruses. Emerg. Microbes Infect. 2018, 7, 7. [Google Scholar] [CrossRef] [PubMed]
- Collins, M.H.; McGowan, E.; Jadi, R.; Young, E.; Lopez, C.A.; Baric, R.S.; Lazear, H.M.; de Silva, A.M. Lack of durable cross-neutralizing antibodies against Zika virus from Dengue virus infection. Emerg. Infect. Dis. 2017, 23, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Katzelnick, L.C.; Gresh, L.; Halloran, M.E.; Mercado, J.C.; Kuan, G.; Gordon, A.; Balmaseda, A.; Harris, E. Antibody-dependent enhancement of severe dengue disease in humans. Science. 2017, 358, 929–932. [Google Scholar] [CrossRef] [PubMed]
- Priyamvada, L.; Hudson, W.; Ahmed, R.; Wrammert, J. Humoral cross-reactivity between Zika and dengue viruses: Implications for protection and pathology. Emerg. Microbes Infect. 2017, 6, e33. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.C.; Martin, E.; Shimada, M.; Godfrey, S.B.; Fricke, J.; Locastro, S.; Lai, N.Y.; Liebesny, P.; Carlson, J.M.; Brumme, C.J.; et al. Aminopeptidase substrate preference affects HIV epitope presentation and predicts immune escape patterns in HIV-infected individuals. J. Immunol. 2012, 188, 5924–5934. [Google Scholar] [CrossRef] [PubMed]
- Ricciardi, M.J.; Magnani, D.M.; Grifoni, A.; Kwon, Y.C.; Gutman, M.J.; Grubaugh, N.D.; Gangavarapu, K.; Sharkey, M.; Silveira, C.G.T.; et al. Ontogeny of the B- and T-cell response in a primary Zika virus infection of a dengue-naive individual during the 2016 outbreak in Miami, FL. PLoS Negl. Trop. Dis. 2017, 11, e0006000. [Google Scholar] [CrossRef] [PubMed]
- Swanstrom, J.A.; Plante, J.A.; Plante, K.S.; Young, E.F.; McGowan, E.; Gallichotte, E.N.; Widman, D.G.; Heise, M.T.; de Silva, A.M.; Baric, R.S. Dengue virus envelope dimer epitope monoclonal antibodies isolated from dengue patients are protective against Zika virus. MBio 2016, 7, e01123-16. [Google Scholar] [CrossRef] [PubMed]
- Morens, D.M.; Halstead, S.B. Measurement of antibody-dependent infection enhancement of four dengue virus serotypes by monoclonal and polyclonal antibodies. J. Gen. Virol. 1990, 71, 2909–29014. [Google Scholar] [CrossRef] [PubMed]
- Guy, B.; Chanthavanich, P.; Gimenez, S.; Sirivichayakul, C.; Sabchareon, A.; Begue, S.; Yoksan, S.; Luxemburger, C.; Lang, J. Evaluation by flow cytometry of antibody-dependent enhancement (ADE) of dengue infection by sera from Thai children immunized with a live-attenuated tetravalent dengue vaccine. Vaccine 2004, 22, 3563–3574. [Google Scholar] [CrossRef] [PubMed]
- Barba-Spaeth, G.; Dejnirattisai, W.; Rouvinski, A.; Vaney, M.C.; Medits, I.; Sharma, A.; Simon-Loriere, E.; Sakuntabhai, A.; Cao-Lormeau, V.M.; Haouz, A.; et al. Structural basis of potent Zika-dengue virus antibody cross-neutralization. Nature 2016, 536, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Robbiani, D.F.; Bozzacco, L.; Keeffe, J.R.; Khouri, R.; Olsen, P.C.; Gazumyan, A.; Schaefer-Babajew, D.; Avila-Rios, S.; Nogueira, L.; Patel, R.; et al. Recurrent potent human neutralizing antibodies to Zika virus in Brazil and Mexico. Cell 2017, 169, 597–609. [Google Scholar] [CrossRef] [PubMed]
- Rogers, T.F.; Goodwin, E.C.; Briney, B.; Sok, D.; Beutler, N.; Strubel, A.; Nedellec, R.; Le, K.; Brown, M.E.; Burton, D.R.; et al. Zika virus activates de novo and cross-reactive memory B cell responses in dengue-experienced donors. Sci. Immunol. 2017, 2, 6809. [Google Scholar] [CrossRef] [PubMed]
- McCracken, M.K.; Gromowski, G.D.; Friberg, H.L.; Lin, X.; Abbink, P.; De La Barrera, R.; Eckles, K.H.; Garver, L.S.; Boyd, M.; Jetton, D.; et al. Impact of prior flavivirus immunity on Zika virus infection in rhesus macaques. PLoS Pathog. 2017, 13, e1006487. [Google Scholar] [CrossRef] [PubMed]
- Pantoja, P.; Perez-Guzman, E.X.; Rodriguez, I.V.; White, L.J.; Gonzalez, O.; Serrano, C.; Giavedoni, L.; Hodara, V.; Cruz, L.; Arana, T.; et al. Zika virus pathogenesis in rhesus macaques is unaffected by pre-existing immunity to dengue virus. Nat. Commun. 2017, 8, 15674. [Google Scholar] [CrossRef] [PubMed]
- Dan, J.M.; Lindestam Arlehamn, C.S.; Weiskopf, D.; da Silva Antunes, R.; Havenar-Daughton, C.; Reiss, S.M.; Brigger, M.; Bothwell, M.; Sette, A.; Crotty, S. A Cytokine-Independent Approach To Identify Antigen-Specific Human Germinal Center T Follicular Helper Cells and Rare Antigen-Specific CD4+ T Cells in Blood. J. Immunol. 2016, 197, 983–993. [Google Scholar] [CrossRef] [PubMed]

| Group | Nb of Samples | ZIKV Status 1,2 | DENV Status 1,2 | Age 3 (Year) | Sex (% Male) |
|---|---|---|---|---|---|
| A | 14 | Negative | Positive | 17 (84–1) | 36 |
| B | 9 | Positive | Negative | 30 (184–1) | 56 |
| C | 5 | Positive | Positive | 27 (213–5) | 20 |
| D | 4 | Acute | Positive | 33 (205–4) | 25 |
| E | 2 | Acute | Acute | 27 (252–8) | 50 |
| Peptide 1 | Sequence | SFC/Million PBMC 2 | HLA | Score 3 |
|---|---|---|---|---|
| C869–4 | KDLAAMLRI | 65 | B*55:01 | 1.4 |
| B*40:02 | 2.0 | |||
| C899–7 | AAMLRIINA | 75 | A*02:01 | 4.5 |
| NS1637–7 | MENIMWRSVEGELNA | 310 | DRB1*04:05 | 50 |
| NS1687–6 | WRSVEGELN | 206 | B*40:02 | 48 |
| B*18:01 | 48 | |||
| NS11671–75 | VWLKVREDY | 75 | A*29:02 | 1.3 |
| NS12802–88 | CPGTKVHVE | 170 | B*35:01 | 8.5 |
| B*35:31 | 6.5 | |||
| NS32192–33 | TVILAPTRVVAAEME | 165 | DRB1*08:02 | 1.5 |
| NS33113–25 | AAIFMTATPPGTRDA | 470 | DRB1*04:01 | 4.0 |
| NS33133–21 | IFMTATPPG | 30 | A*24:02 | 5.0 |
| NS33143–22 | FMTATPPGT | 85 | A*02:17 | 5.5 |
| NS4B1121–20 | AIILLVAHY | 88 | A*29:02 | 0.6 |
| A*11:01 | 3.5 | |||
| NS4B1151–23 | LLVAHYMYL | 68 | A*02:05 | 0.3 |
| NS4B1161–24 | LVAHYMYLI | 35 | A*69:01 | 0.15 |
| A*02:05 | 0.2 | |||
| NS52983–06 | NHPYRTWAY 4 | 358 | B*35:01 | 3.0 |
| NS52993–07 | HPYRTWAYH 4 | 308 | B*35:01 | 0.4 |
| NS53023–10 | RTWAYHGSY | 205 | A*01:01 | 0.5 |
| NS53483–56 | TPYGQQRVF 4 | 1681 | B*35:31 | 0.7 |
| B*35:01 | 0.3 | |||
| NS54254–33 | EAVNDPRFW | 465 | B*44:03 | 5.0 |
| B*15:17 | 7.0 | |||
| B*35:01 | 5.0 | |||
| NS54614–75 | KKQGEFGKAKGSRAI | 405 | DRB1*07:01 | 32 |
| NS54734–87 | RAIWYMWLGARFLEF | 505 | DRB1*07:01 | 16 |
| NS56096–17 | YALNTFTNL | 42 | B*35:43 | 0.4 |
| B*35:31 | 0.25 |
| Peptide 1 | Sequence | ZIKV | DENV/ZIKV | Identity (%) | |||||
|---|---|---|---|---|---|---|---|---|---|
| Donors | SFC/million PBMC 2 | Donors | SFC/million PBMC 2 | DENV Serotypes | |||||
| 1 | 2 | 3 | 4 | ||||||
| C49-63 | AILAFLRFTAIKPSL | 60 | 60 | 28,63 | 365 | 60 | 53 | 60 | 40 |
| E67-81 | DMASDSRCPTQGEAY | 33 | 465 | 67 | 53 | 67 | 53 | ||
| E87-101 | DTQYVCKRTLVDRGW | 56 | 505 | 67 | 54 | 73 | 67 | ||
| NS119-33 | VFVYNDVEAWRDRYK | 21,46,60 | 195 | 28,56 | 380 | 47 | 33 | 47 | 40 |
| NS155-69 | CGISSVSRMENIMWR | 35,46 | 125 | 56 | 275 | 67 | 66 | 60 | 60 |
| NS191-105 | GSVKNPMWRGPQRLP | 21,35,46,60 | 275 | 28 | 165 | 13 | 33 | 20 | 33 |
| NS1107-121 | PVNELPHGWKAWGKS | 28,53 | 430 | 40 | 47 | 47 | 50 | ||
| NS1147-161 | HRAWNSFLVEDHGFG | 46 | 40 | 33,53 | 445 | 67 | 73 | 67 | 76 |
| NS1163-177 | FHTSVWLKVREDYSL | 46 | 35 | 20,28,55 | 450 | 47 | 46 | 53 | 47 |
| NS1195-209 | HSDLGYWIESEKNDT | 28,33 | 615 | 80 | 73 | 66 | 73 | ||
| NS2B117-131 | AAGAWYVYVKTGKRS | 55 | 445 | 33 | 33 | 27 | 27 | ||
| NS3131-145 | PAGTSGSPILDKCGR | 21,42 | 405 | 26,55,63 | 495 | 53 | 61 | 53 | 54 |
| NS3143-157 | CGRVIGLYGNGVVIK | 21 | 350 | 20,55,63,66 | 550 | 60 | 67 | 72 | 80 |
| NS3311-325 | AAIFMTATPPGTRDA | 28,33 | 470 | 80 | 80 | 93 | 80 | ||
| NS513-27 | KARLNQMSALEFYSY | 55,69 | 405 | 53 | 47 | 53 | 40 | ||
| NS5293-307 | WFFDENHPYRTWAYH | 55,69 | 1620 | 67 | 67 | 60 | 67 | ||
| NS5297-311 | ENHPYRTWAYHGSYE | 55,69 | 1330 | 80 | 80 | 73 | 80 | ||
| NS5325-339 | VVRLLSKPWDVVTGV | 28,55,66 | 495 | 73 | 80 | 73 | 67 | ||
| NS5345-359 | TDTTPYGQQRVFKEK | 33,55,69 | 4195 | 93 | 93 | 93 | 93 | ||
| NS5373-387 | QVMSMVSSWLWKELG | 60 | 130 | 55,66,69 | 340 | 40 | 53 | 47 | 47 |
| NS5461-475 | KKQGEFGKAKGSRAI | 28,53 | 405 | 93 | 93 | 93 | 87 | ||
| NS5465-479 | EFGKAKGSRAIWYMW | 28,53,55,56 | 1085 | 100 | 100 | 100 | 93 | ||
| NS5473-487 | RAIWYMWLGARFLEF | 28,55 | 505 | 100 | 100 | 93 | 100 | ||
| NS5481-495 | GARFLEFEALGFLNE | 28,53,56,63 | 1870 | 93 | 100 | 93 | 100 | ||
| NS5546-560 | RFDLENEALITNQME | 28,53,66 | 515 | 60 | 47 | 53 | 60 | ||
| NS5573-586 | TYQNKVVKVLRPAEK | 28,53,56 | 615 | 73 | 67 | 73 | 80 | ||
| NS5849-863 | CGSLIGHRPRTTWAE | 60 | 90 | 33,55 | 340 | 67 | 67 | 67 | 67 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Delgado, F.G.; Torres, K.I.; Castellanos, J.E.; Romero-Sánchez, C.; Simon-Lorière, E.; Sakuntabhai, A.; Roth, C. Improved Immune Responses Against Zika Virus After Sequential Dengue and Zika Virus Infection in Humans. Viruses 2018, 10, 480. https://doi.org/10.3390/v10090480
Delgado FG, Torres KI, Castellanos JE, Romero-Sánchez C, Simon-Lorière E, Sakuntabhai A, Roth C. Improved Immune Responses Against Zika Virus After Sequential Dengue and Zika Virus Infection in Humans. Viruses. 2018; 10(9):480. https://doi.org/10.3390/v10090480
Chicago/Turabian StyleDelgado, Félix G., Karina I. Torres, Jaime E. Castellanos, Consuelo Romero-Sánchez, Etienne Simon-Lorière, Anavaj Sakuntabhai, and Claude Roth. 2018. "Improved Immune Responses Against Zika Virus After Sequential Dengue and Zika Virus Infection in Humans" Viruses 10, no. 9: 480. https://doi.org/10.3390/v10090480
APA StyleDelgado, F. G., Torres, K. I., Castellanos, J. E., Romero-Sánchez, C., Simon-Lorière, E., Sakuntabhai, A., & Roth, C. (2018). Improved Immune Responses Against Zika Virus After Sequential Dengue and Zika Virus Infection in Humans. Viruses, 10(9), 480. https://doi.org/10.3390/v10090480





