Capsid Structure of dsRNA Fungal Viruses
Abstract
1. Introduction
2. Structure of dsRNA Virus Capsids
2.1. Totiviruses
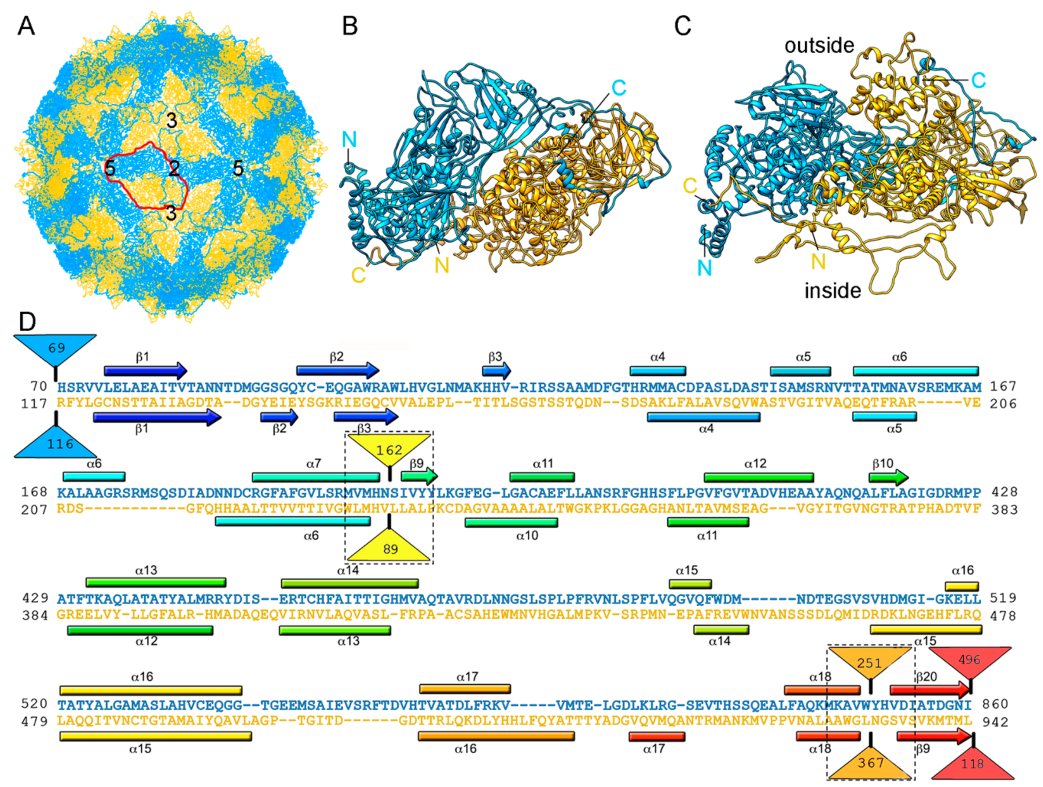
2.2. Chrysoviruses
2.3. Partitiviruses
2.4. Quadriviruses
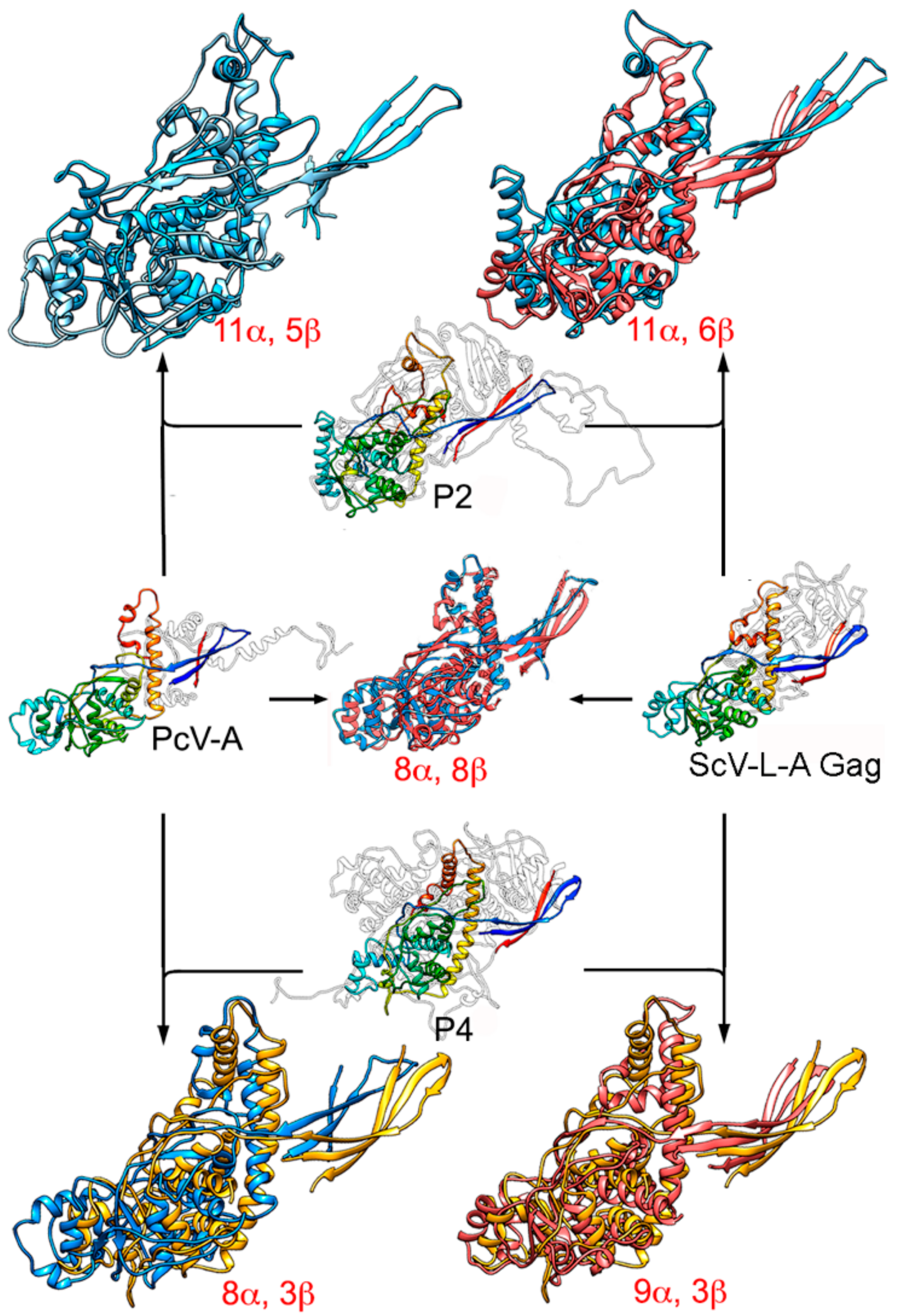
3. Evolutionary Relationships Based on Structural Comparisons
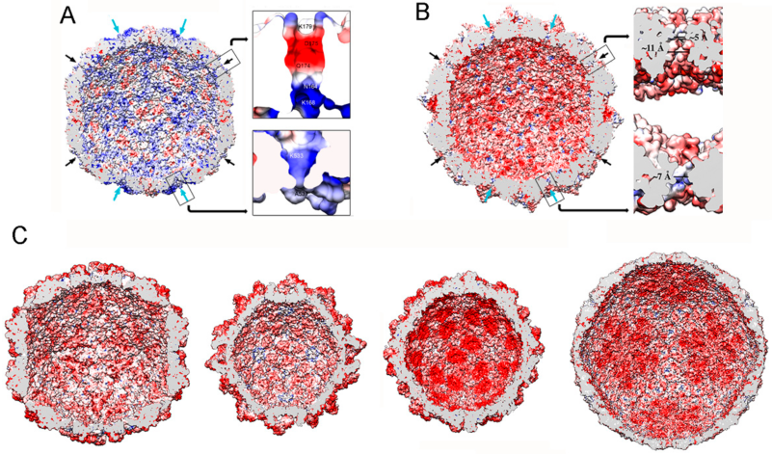
4. RdRp and dsRNA Organization within Mycovirus Capsids
5. Concluding Remarks and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Patton, J.T. Segmented double-stranded RNA viruses. Structure and Molecular Biology; Caister Academic Press: Norfolk, UK, 2008. [Google Scholar]
- Hill, C.L.; Booth, T.F.; Prasad, B.V.; Grimes, J.M.; Mertens, P.P.; Sutton, G.C.; Stuart, D.I. The structure of a cypovirus and the functional organization of dsRNA viruses. Nat. Struct. Biol. 1999, 6, 565–568. [Google Scholar] [PubMed]
- Castón, J.R.; Trus, B.L.; Booy, F.P.; Wickner, R.B.; Wall, J.S.; Steven, A.C. Structure of L-A virus: A specialized compartment for the transcription and replication of double-stranded RNA. J. Cell Biol. 1997, 138, 975–985. [Google Scholar] [CrossRef] [PubMed]
- Grimes, J.M.; Burroughs, J.N.; Gouet, P.; Diprose, J.M.; Malby, R.; Zientara, S.; Mertens, P.P.; Stuart, D.I. The atomic structure of the bluetongue virus core. Nature 1998, 395, 470–478. [Google Scholar] [CrossRef] [PubMed]
- Prasad, B.V.; Rothnagel, R.; Zeng, C.Q.; Jakana, J.; Lawton, J.A.; Chiu, W.; Estes, M.K. Visualization of ordered genomic RNA and localization of transcriptional complexes in rotavirus. Nature 1996, 382, 471–473. [Google Scholar] [CrossRef] [PubMed]
- Shaw, A.L.; Samal, S.K.; Subramanian, K.; Prasad, B.V. The structure of aquareovirus shows how the different geometries of the two layers of the capsid are reconciled to provide symmetrical interactions and stabilization. Structure 1996, 4, 957–967. [Google Scholar] [CrossRef]
- Lu, G.; Zhou, Z.H.; Baker, M.L.; Jakana, J.; Cai, D.; Wei, X.; Chen, S.; Gu, X.; Chiu, W. Structure of double-shelled rice dwarf virus. J. Virol. 1998, 72, 8541–8549. [Google Scholar] [PubMed]
- Caspar, D.L.D.; Klug, A. Physical principles in the construction of regular viruses. Cold Spring Harbor. Symp. Quant. Biol. 1962, 27, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Harrison, S.C. Principles of virus structure. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Griffin, D.E., Lamb, R.A., Martin, M.A., Roizman, B., Strauss, S.E., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; Volume 1, pp. 59–98. [Google Scholar]
- Mertens, P. The dsRNA viruses. Virus Res. 2004, 101, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Arnold, M.M.; Sen, A.; Greenberg, H.B.; Patton, J.T. The battle between rotavirus and its host for control of the interferon signaling pathway. PLoS Pathog. 2013, 9, e1003064. [Google Scholar] [CrossRef] [PubMed]
- Nuss, D.L. Mycoviruses, RNA silencing, and viral RNA recombination. Adv. Virus Res. 2011, 80, 25–48. [Google Scholar] [PubMed]
- Cheng, R.H.; Castón, J.R.; Wang, G.J.; Gu, F.; Smith, T.J.; Baker, T.S.; Bozarth, R.F.; Trus, B.L.; Cheng, N.; Wickner, R.B.; et al. Fungal virus capsids, cytoplasmic compartments for the replication of double-stranded RNA, formed as icosahedral shells of asymmetric gag dimers. J. Mol. Biol. 1994, 244, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Lawton, J.A.; Estes, M.K.; Prasad, B.V. Mechanism of genome transcription in segmented dsRNA viruses. Adv. Virus Res. 2000, 55, 185–229. [Google Scholar] [PubMed]
- Patton, J.T.; Spencer, E. Genome replication and packaging of segmented double-stranded RNA viruses. Virology 2000, 277, 217–225. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Baker, M.L.; Jiang, W.; Estes, M.K.; Prasad, B.V. Rotavirus architecture at subnanometer resolution. J. Virol. 2009, 83, 1754–1766. [Google Scholar] [CrossRef] [PubMed]
- Trask, S.D.; McDonald, S.M.; Patton, J.T. Structural insights into the coupling of virion assembly and rotavirus replication. Nat. Rev. Microbiol. 2012, 10, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Castón, J.R.; Luque, D.; Trus, B.L.; Rivas, G.; Alfonso, C.; González, J.M.; Carrascosa, J.L.; Annamalai, P.; Ghabrial, S.A. Three-dimensional structure and stoichiometry of helmintosporium victoriae190s totivirus. Virology 2006, 347, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Walker, S.B.; Chipman, P.R.; Nibert, M.L.; Baker, T.S. Reovirus polymerase lambda 3 localized by cryo-electron microscopy of virions at a resolution of 7.6 A. Nat. Struct. Biol. 2003, 10, 1011–1018. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Jin, L.; Zhou, Z.H. 3.88 A structure of cytoplasmic polyhedrosis virus by cryo-electron microscopy. Nature 2008, 453, 415–419. [Google Scholar] [CrossRef] [PubMed]
- Lawton, J.A.; Estes, M.K.; Prasad, B.V. Three-dimensional visualization of mRNA release from actively transcribing rotavirus particles. Nat. Struct. Biol. 1997, 4, 118–121. [Google Scholar] [CrossRef] [PubMed]
- McClain, B.; Settembre, E.; Temple, B.R.; Bellamy, A.R.; Harrison, S.C. X-ray crystal structure of the rotavirus inner capsid particle at 3.8 A resolution. J. Mol. Biol. 2010, 397, 587–599. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Zhu, J.; Hui, W.H.; Zhang, X.; Honig, B.; Fang, Q.; Zhou, Z.H. Backbone model of an aquareovirus virion by cryo-electron microscopy and bioinformatics. J. Mol. Biol. 2010, 397, 852–863. [Google Scholar] [CrossRef] [PubMed]
- Duquerroy, S.; Da Costa, B.; Henry, C.; Vigouroux, A.; Libersou, S.; Lepault, J.; Navaza, J.; Delmas, B.; Rey, F.A. The picobirnavirus crystal structure provides functional insights into virion assembly and cell entry. EMBO J. 2009, 28, 1655–1665. [Google Scholar] [CrossRef] [PubMed]
- Jaalinoja, H.T.; Huiskonen, J.T.; Butcher, S.J. Electron cryomicroscopy comparison of the architectures of the enveloped bacteriophages phi6 and phi8. Structure 2007, 15, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Huiskonen, J.T.; de Haas, F.; Bubeck, D.; Bamford, D.H.; Fuller, S.D.; Butcher, S.J. Structure of the bacteriophage phi6 nucleocapsid suggests a mechanism for sequential RNA packaging. Structure 2006, 14, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Naitow, H.; Tang, J.; Canady, M.; Wickner, R.B.; Johnson, J.E. L-A virus at 3.4 A resolution reveals particle architecture and mRNA decapping mechanism. Nat. Struct. Biol. 2002, 9, 725–728. [Google Scholar] [CrossRef] [PubMed]
- Janssen, M.E.; Takagi, Y.; Parent, K.N.; Cardone, G.; Nibert, M.L.; Baker, T.S. Three-dimensional structure of a protozoal double-stranded RNA virus that infects the enteric pathogen giardia lamblia. J. Virol. 2015, 89, 1182–1194. [Google Scholar] [CrossRef] [PubMed]
- Parent, K.N.; Takagi, Y.; Cardone, G.; Olson, N.H.; Ericsson, M.; Yang, M.; Lee, Y.; Asara, J.M.; Fichorova, R.N.; Baker, T.S.; et al. Structure of a protozoan virus from the human genitourinary parasite trichomonas vaginalis. MBio 2013, 4, e00056–013. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Dong, L.; Lin, L.; Ochoa, W.F.; Sinkovits, R.S.; Havens, W.M.; Nibert, M.L.; Baker, T.S.; Ghabrial, S.A.; Tao, Y.J. Atomic structure reveals the unique capsid organization of a dsRNA virus. Proc. Natl. Acad. Sci. USA 2009, 106, 4225–4230. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Ochoa, W.F.; Li, H.; Havens, W.M.; Nibert, M.L.; Ghabrial, S.A.; Baker, T.S. Structure of fusarium poae virus 1 shows conserved and variable elements of partitivirus capsids and evolutionary relationships to picobirnavirus. J. Struct. Biol. 2010, 172, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, N.; Salaipeth, L.; Kanematsu, S.; Iwasaki, K.; Suzuki, N. Megabirnavirus structure reveals a putative 120-subunit capsid formed by asymmetrical dimers with distinctive large protrusions. J. Gen. Virol. 2015, 96, 2435–2441. [Google Scholar] [CrossRef] [PubMed]
- Luque, D.; González, J.M.; Garriga, D.; Ghabrial, S.A.; Havens, W.M.; Trus, B.; Verdaguer, N.; Carrascosa, J.L.; Castón, J.R. The T = 1 capsid protein of penicillium chrysogenum virus is formed by a repeated helix-rich core indicative of gene duplication. J. Virol. 2010, 84, 7256–7266. [Google Scholar] [CrossRef] [PubMed]
- Castón, J.R.; Luque, D.; Gómez-Blanco, J.; Ghabrial, S.A. Chrysovirus structure: Repeated helical core as evidence of gene duplication. Adv. Virus Res. 2013, 86, 87–108. [Google Scholar] [PubMed]
- Gómez-Blanco, J.; Luque, D.; Gonzalez, J.M.; Carrascosa, J.L.; Alfonso, C.; Trus, B.; Havens, W.M.; Ghabrial, S.A.; Castón, J.R. Cryphonectria nitschkei virus 1 structure shows that the capsid protein of chrysoviruses is a duplicated helix-rich fold conserved in fungal double-stranded RNA viruses. J. Virol. 2012, 86, 8314–8318. [Google Scholar] [CrossRef] [PubMed]
- Luque, D.; Mata, C.P.; Gonzalez-Camacho, F.; González, J.M.; Gómez-Blanco, J.; Alfonso, C.; Rivas, G.; Havens, W.M.; Kanematsu, S.; Suzuki, N.; et al. Heterodimers as the structural unit of the T = 1 capsid of the fungal double-stranded RNA rosellinia necatrix quadrivirus 1. J. Virol. 2016, 90, 11220–11230. [Google Scholar] [CrossRef] [PubMed]
- Mata, C.P.; Luque, D.; Gómez-Blanco, J.; Rodríguez, J.M.; González, J.M.; Suzuki, N.; Ghabrial, S.A.; Carrascosa, J.L.; Trus, B.L.; Castón, J.R. Acquisition of functions on the outer capsid surface during evolution of double-stranded RNA fungal viruses. PLoS Pathog. 2017, 13, e1006755. [Google Scholar] [CrossRef] [PubMed]
- Coulibaly, F.; Chevalier, C.; Gutsche, I.; Pous, J.; Navaza, J.; Bressanelli, S.; Delmas, B.; Rey, F.A. The birnavirus crystal structure reveals structural relationships among icosahedral viruses. Cell 2005, 120, 761–772. [Google Scholar] [CrossRef] [PubMed]
- Castón, J.R.; Martínez-Torrecuadrada, J.L.; Maraver, A.; Lombardo, E.; Rodríguez, J.F.; Casal, J.I.; Carrascosa, J.L. C terminus of infectious bursal disease virus major capsid protein VP2 is involved in definition of the t number for capsid assembly. J. Virol. 2001, 75, 10815–10828. [Google Scholar] [CrossRef] [PubMed]
- Luque, D.; Rivas, G.; Alfonso, C.; Carrascosa, J.L.; Rodríguez, J.F.; Castón, J.R. Infectious bursal disease virus is an icosahedral polyploid dsrna virus. Proc. Natl. Acad. Sci. USA 2009, 106, 2148–2152. [Google Scholar] [CrossRef] [PubMed]
- Luque, D.; Saugar, I.; Rejas, M.T.; Carrascosa, J.L.; Rodríguez, J.F.; Castón, J.R. Infectious bursal disease virus: Ribonucleoprotein complexes of a double-stranded RNA virus. J. Mol. Biol. 2009, 386, 891–901. [Google Scholar] [CrossRef] [PubMed]
- Reinisch, K.M.; Nibert, M.L.; Harrison, S.C. Structure of the reovirus core at 3.6 Å resolution. Nature 2000, 404, 960–967. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, A.; Miyazaki, N.; Taka, J.; Naitow, H.; Ogawa, A.; Fujimoto, Z.; Mizuno, H.; Higashi, T.; Watanabe, Y.; Omura, T.; et al. The atomic structure of rice dwarf virus reveals the self-assembly mechanism of component proteins. Structure 2003, 11, 1227–1238. [Google Scholar] [CrossRef] [PubMed]
- Settembre, E.C.; Chen, J.Z.; Dormitzer, P.R.; Grigorieff, N.; Harrison, S.C. Atomic model of an infectious rotavirus particle. Embo J. 2011, 30, 408–416. [Google Scholar] [CrossRef] [PubMed]
- Nemecek, D.; Boura, E.; Wu, W.; Cheng, N.; Plevka, P.; Qiao, J.; Mindich, L.; Heymann, J.B.; Hurley, J.H.; Steven, A.C. Subunit folds and maturation pathway of a dsrna virus capsid. Structure 2013, 21, 1374–1383. [Google Scholar] [CrossRef] [PubMed]
- El Omari, K.; Sutton, G.; Ravantti, J.J.; Zhang, H.; Walter, T.S.; Grimes, J.M.; Bamford, D.H.; Stuart, D.I.; Mancini, E.J. Plate tectonics of virus shell assembly and reorganization in phage phi8, a distant relative of mammalian reoviruses. Structure 2013, 21, 1384–1395. [Google Scholar] [CrossRef] [PubMed]
- Luque, D.; Gómez-Blanco, J.; Garriga, D.; Brilot, A.F.; González, J.M.; Havens, W.M.; Carrascosa, J.L.; Trus, B.L.; Verdaguer, N.; Ghabrial, S.A.; et al. Cryo-EM near-atomic structure of a dsRNA fungal virus shows ancient structural motifs preserved in the dsRNA viral lineage. Proc. Natl. Acad. Sci. USA 2014, 111, 7641–7646. [Google Scholar] [CrossRef] [PubMed]
- Reinisch, K.M. The dsRNA viridae and their catalytic capsids. Nat. Struct. Biol. 2002, 9, 714–716. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A.; Castón, J.R.; Jiang, D.; Nibert, M.L.; Suzuki, N. 50-plus years of fungal viruses. Virology 2015, 479–480, 356–368. [Google Scholar] [CrossRef] [PubMed]
- Wickner, R.B. Viruses of yeast, fungi, and parasitic microorganisms. In Fields. Virology, 4th ed.; Knipe, D.M., Howley, P.M., Griffin, D.E., Martin, M.A., Lamb, R.A., Roizman, B., Strauss, S.E., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2001; Volume 1, pp. 629–658. [Google Scholar]
- Wickner, R.B.; Fujimura, T.; Esteban, R. Viruses and prions of saccharomyces cerevisiae. Adv. Virus Res. 2013, 86, 1–36. [Google Scholar] [PubMed]
- Dinman, J.D.; Icho, T.; Wickner, R.B. A-1 ribosomal frameshift in a double-stranded RNA virus of yeast forms a gag-pol fusion protein. Proc. Natl. Acad. Sci. USA 1991, 88, 174–178. [Google Scholar] [CrossRef] [PubMed]
- Ribas, J.C.; Wickner, R.B. The gag domain of the gag-pol fusion protein directs incorporation into the l-a double-stranded rna viral particles in saccharomyces cerevisiae. J. Biol. Chem. 1998, 273, 9306–9311. [Google Scholar] [CrossRef] [PubMed]
- Dinman, J.D.; Wickner, R.B. Ribosomal frameshifting efficiency and gag/gag-pol ratio are critical for yeast M1 double-stranded RNA virus propagation. J. Virol. 1992, 66, 3669–3676. [Google Scholar] [PubMed]
- Tang, J.; Naitow, H.; Gardner, N.A.; Kolesar, A.; Tang, L.; Wickner, R.B.; Johnson, J.E. The structural basis of recognition and removal of cellular mRNA 7-methyl G ‘caps’ by a viral capsid protein: A unique viral response to host defense. J. Mol. Recognit. 2005, 18, 158–168. [Google Scholar] [CrossRef] [PubMed]
- Fujimura, T.; Esteban, R. Cap-snatching mechanism in yeast L-A double-stranded RNA virus. Proc. Natl. Acad. Sci. USA 2011, 108, 17667–17671. [Google Scholar] [CrossRef] [PubMed]
- Dunn, S.E.; Li, H.; Cardone, G.; Nibert, M.L.; Ghabrial, S.A.; Baker, T.S. Three-dimensional structure of victorivirus HvV190s suggests coat proteins in most totiviruses share a conserved core. PLoS Pathog. 2013, 9, e1003225. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Ghabrial, S.A. Organization and expression of the double-stranded RNA genome of helminthosporium victoriae 190s virus, a totivirus infecting a plant pathogenic filamentous fungus. Proc. Natl. Acad. Sci. USA 1996, 93, 12541–12546. [Google Scholar] [CrossRef] [PubMed]
- Soldevila, A.I.; Ghabrial, S.A. Expression of the totivirus helminthosporium victoriae 190s virus RNA-dependent rna polymerase from its downstream open reading frame in dicistronic constructs. J. Virol. 2000, 74, 997–1003. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Ochoa, W.F.; Sinkovits, R.S.; Poulos, B.T.; Ghabrial, S.A.; Lightner, D.V.; Baker, T.S.; Nibert, M.L. Infectious myonecrosis virus has a totivirus-like, 120-subunit capsid, but with fiber complexes at the fivefold axes. Proc. Natl. Acad. Sci. USA 2008, 105, 17526–17531. [Google Scholar] [CrossRef] [PubMed]
- Nibert, M.L.; Takagi, Y. Fibers come and go: Differences in cell-entry components among related dsRNA viruses. Curr. Opin. Virol. 2013, 3, 20–26. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A.; Castón, J.R.; Coutts, R.H.A.; Hillman, B.I.; Jiang, D.; Kim, D.H.; Moriyama, H.; Ictv Report, C. ICTV virus taxonomy profile: Chrysoviridae. J. Gen. Virol. 2018, 99, 19–20. [Google Scholar] [CrossRef] [PubMed]
- Castón, J.R.; Ghabrial, S.A.; Jiang, D.; Rivas, G.; Alfonso, C.; Roca, R.; Luque, D.; Carrascosa, J.L. Three-dimensional structure of penicillium chrysogenum virus: A double-stranded RNA virus with a genuine T = 1 capsid. J. Mol. Biol. 2003, 331, 417–431. [Google Scholar] [CrossRef]
- Jiang, D.; Ghabrial, S.A. Molecular characterization of penicillium chrysogenum virus: Reconsideration of the taxonomy of the genus chrysovirus. J. Gen. Virol. 2004, 85, 2111–2121. [Google Scholar] [CrossRef] [PubMed]
- Nibert, M.L.; Ghabrial, S.A.; Maiss, E.; Lesker, T.; Vainio, E.J.; Jiang, D.; Suzuki, N. Taxonomic reorganization of family partitiviridae and other recent progress in partitivirus research. Virus Res. 2014, 188, 128–141. [Google Scholar] [CrossRef] [PubMed]
- Vainio, E.J.; Chiba, S.; Ghabrial, S.A.; Maiss, E.; Roossinck, M.; Sabanadzovic, S.; Suzuki, N.; Xie, J.; Nibert, M.; Ictv Report Consortium. ICTV virus taxonomy profile: Partitiviridae. J. Gen. Virol. 2018, 99, 17–18. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Pan, J.; Havens, W.M.; Ochoa, W.F.; Guu, T.S.; Ghabrial, S.A.; Nibert, M.L.; Tao, Y.J.; Baker, T.S. Backbone trace of partitivirus capsid protein from electron cryomicroscopy and homology modeling. Biophys. J. 2010, 99, 685–694. [Google Scholar] [CrossRef] [PubMed]
- Ochoa, W.F.; Havens, W.M.; Sinkovits, R.S.; Nibert, M.L.; Ghabrial, S.A.; Baker, T.S. Partitivirus structure reveals a 120-subunit, helix-rich capsid with distinctive surface arches formed by quasisymmetric coat-protein dimers. Structure 2008, 16, 776–786. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Cheng, J.; Tang, J.; Fu, Y.; Jiang, D.; Baker, T.S.; Ghabrial, S.A.; Xie, J. A novel partitivirus that confers hypovirulence on plant pathogenic fungi. J. Virol. 2014, 88, 10120–10133. [Google Scholar] [CrossRef] [PubMed]
- Krol, M.A.; Olson, N.H.; Tate, J.; Johnson, J.E.; Baker, T.S.; Ahlquist, P. RNA-controlled polymorphism in the in vivo assembly of 180-subunit and 120-subunit virions from a single capsid protein. Proc. Natl. Acad. Sci. USA 1999, 96, 13650–13655. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Johnson, J.M.; Dryden, K.A.; Young, M.J.; Zlotnick, A.; Johnson, J.E. The role of subunit hinges and molecular “switches” in the control of viral capsid polymorphism. J. Struct. Biol. 2006, 154, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Kainov, D.E.; Butcher, S.J.; Bamford, D.H.; Tuma, R. Conserved intermediates on the assembly pathway of double-stranded RNA bacteriophages. J. Mol. Biol. 2003, 328, 791–804. [Google Scholar] [CrossRef]
- Lin, Y.H.; Chiba, S.; Tani, A.; Kondo, H.; Sasaki, A.; Kanematsu, S.; Suzuki, N. A novel quadripartite dsRNA virus isolated from a phytopathogenic filamentous fungus, rosellinia necatrix. Virology 2012, 426, 42–50. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.H.; Hisano, S.; Yaegashi, H.; Kanematsu, S.; Suzuki, N. A second quadrivirus strain from the phytopathogenic filamentous fungus rosellinia necatrix. Arch. Virol. 2013, 158, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Kondo, H.; Kanematsu, S.; Suzuki, N. Viruses of the white root rot fungus, rosellinia necatrix. Adv. Virus Res. 2013, 86, 177–214. [Google Scholar] [PubMed]
- Arjona-Lopez, J.M.; Telengech, P.; Jamal, A.; Hisano, S.; Kondo, H.; Yelin, M.D.; Arjona-Girona, I.; Kanematsu, S.; López-Herrera, C.J.; Suzuki, N. Novel, diverse RNA viruses from mediterranean isolates of the phytopathogenic fungus, rosellinia necatrix: Insights into evolutionary biology of fungal viruses. Environ. Microbiol. 2018, 20, 1464–1483. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.L.; Jiang, W.; Rixon, F.J.; Chiu, W. Common ancestry of herpesviruses and tailed DNA bacteriophages. J. Virol. 2005, 79, 14967–14970. [Google Scholar] [CrossRef] [PubMed]
- Bamford, D.H.; Grimes, J.M.; Stuart, D.I. What does structure tell us about virus evolution? Curr. Opin. Struct. Biol. 2005, 15, 655–663. [Google Scholar] [CrossRef] [PubMed]
- Rossmann, M.; Johnson, J. Icosahedral rna virus structure. Annu. Rev. Biochem. 1989, 58, 533–573. [Google Scholar] [CrossRef] [PubMed]
- Abrescia, N.G.; Bamford, D.H.; Grimes, J.M.; Stuart, D.I. Structure unifies the viral universe. Annu. Rev. Biochem. 2012, 81, 795–822. [Google Scholar] [CrossRef] [PubMed]
- Benson, S.D.; Bamford, J.K.; Bamford, D.H.; Burnett, R.M. Does common architecture reveal a viral lineage spanning all three domains of life? Mol. Cell 2004, 16, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Krupovic, M.; Bamford, D.H. Virus evolution: How far does the double beta-barrel viral lineage extend? Nat. Rev. Microbiol. 2008, 6, 941–948. [Google Scholar] [CrossRef] [PubMed]
- Rissanen, I.; Grimes, J.M.; Pawlowski, A.; Mantynen, S.; Harlos, K.; Bamford, J.K.; Stuart, D.I. Bacteriophage P23-77 capsid protein structures reveal the archetype of an ancient branch from a major virus lineage. Structure 2013, 21, 718–726. [Google Scholar] [CrossRef] [PubMed]
- Pietila, M.K.; Laurinmaki, P.; Russell, D.A.; Ko, C.C.; Jacobs-Sera, D.; Hendrix, R.W.; Bamford, D.H.; Butcher, S.J. Structure of the archaeal head-tailed virus HSTV-1 completes the HK97 fold story. Proc. Natl. Acad. Sci. USA 2013, 110, 10604–10609. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.M.; White, J.L.; Grutter, M.G.; Burnett, R.M. Three-dimensional structure of the adenovirus major coat protein hexon. Science 1986, 232, 1148–1151. [Google Scholar] [CrossRef] [PubMed]
- Nandhagopal, N.; Simpson, A.A.; Gurnon, J.R.; Yan, X.; Baker, T.S.; Graves, M.V.; Van Etten, J.L.; Rossmann, M.G. The structure and evolution of the major capsid protein of a large, lipid-containing DNA virus. Proc. Natl. Acad. Sci. USA 2002, 99, 14758–14763. [Google Scholar] [CrossRef] [PubMed]
- Abrescia, N.G.; Cockburn, J.J.; Grimes, J.M.; Sutton, G.C.; Diprose, J.M.; Butcher, S.J.; Fuller, S.D.; San Martin, C.; Burnett, R.M.; Stuart, D.I.; et al. Insights into assembly from structural analysis of bacteriophage PRD1. Nature 2004, 432, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Lomonossoff, G.P.; Johnson, J.E. The synthesis and structure of comovirus capsids. Prog. Biophys. Mol. Biol. 1991, 55, 107–137. [Google Scholar] [CrossRef]
- Ravantti, J.; Bamford, D.; Stuart, D.I. Automatic comparison and classification of protein structures. J. Struct. Biol. 2013, 183, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Suhanovsky, M.M.; Teschke, C.M. Nature’s favorite building block: Deciphering folding and capsid assembly of proteins with the HK97-fold. Virology 2015, 479–480, 487–497. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Jih, J.; Jiang, J.; Zhou, Z.H. Atomic structure of the human cytomegalovirus capsid with its securing tegument layer of pp150. Science 2017, 356. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Cheng, L. Cryo-em shows the polymerase structures and a nonspooled genome within a dsrna virus. Science 2015, 349, 1347–1350. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Ding, K.; Yu, X.; Chang, W.; Sun, J.; Zhou, Z.H. In situ structures of the segmented genome and RNA polymerase complex inside a dsRNA virus. Nature 2015, 527, 531–534. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhou, N.; Chen, W.; Zhu, B.; Wang, X.; Xu, B.; Wang, J.; Liu, H.; Cheng, L. Near-atomic resolution structure determination of a cypovirus capsid and polymerase complex using cryo-EM at 200kV. J. Mol. Biol. 2017, 429, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Estrozi, L.F.; Settembre, E.C.; Goret, G.; McClain, B.; Zhang, X.; Chen, J.Z.; Grigorieff, N.; Harrison, S.C. Location of the dsRNA-dependent polymerase, VP1, in rotavirus particles. J. Mol. Biol. 2013, 425, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Havens, W.M.; Nibert, M.L.; Ghabrial, S.A. An RNA cassette from helminthosporium victoriae virus 190s necessary and sufficient for stop/restart translation. Virology 2015, 474, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Sato, Y.; Castón, J.R.; Suzuki, N. The biological attributes, genome architecture and packaging of diverse multi-component fungal viruses. Curr. Opin. Virol. 2018, 33, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Dryden, K.; Wang, G.; Yeager, M.; Nibert, M.; Coombs, K.; Furlong, D.; Fields, B.; Baker, T. Early steps in reovirus infection are associated with dramatic changes in supramolecular structure and protein conformation: Analysis of virions and subviral particles by cryoelectron microscopy and image reconstruction. J. Cell Biol. 1993, 122, 1023–1041. [Google Scholar] [CrossRef] [PubMed]
- Gouet, P.; Diprose, J.M.; Grimes, J.M.; Malby, R.; Burroughs, J.N.; Zientara, S.; Stuart, D.I.; Mertens, P.P. The highly ordered double-stranded RNA genome of bluetongue virus revealed by crystallography. Cell 1999, 97, 481–490. [Google Scholar] [CrossRef]
- Pesavento, J.B.; Lawton, J.A.; Estes, M.E.; Venkataram Prasad, B.V. The reversible condensation and expansion of the rotavirus genome. Proc. Natl. Acad. Sci. USA 2001, 98, 1381–1386. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Hisano, S.; Tani, A.; Kondo, H.; Kanematsu, S.; Suzuki, N. A capsidless ssrna virus hosted by an unrelated dsRNA virus. Nat. Microbiol. 2016, 1, 15001. [Google Scholar] [CrossRef] [PubMed]
- Hisano, S.; Zhang, R.; Faruk, M.I.; Kondo, H.; Suzuki, N. A neo-virus lifestyle exhibited by a (+)ssRNA virus hosted in an unrelated dsrna virus: Taxonomic and evolutionary considerations. Virus Res. 2018, 244, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Kozlakidis, Z.; Herrero, N.; Coutts, R.H. The complete nucleotide sequence of a totivirus from aspergillus foetidus. Arch. Virol. 2013, 158, 263–266. [Google Scholar] [CrossRef] [PubMed]
- Nerva, L.; Ciuffo, M.; Vallino, M.; Margaria, P.; Varese, G.C.; Gnavi, G.; Turina, M. Multiple approaches for the detection and characterization of viral and plasmid symbionts from a collection of marine fungi. Virus Res. 2016, 219, 22–38. [Google Scholar] [CrossRef] [PubMed]
- Osaki, H.; Sasaki, A.; Nomiyama, K.; Tomioka, K. Multiple virus infection in a single strain of fusarium poae shown by deep sequencing. Virus Genes 2016, 52, 835–847. [Google Scholar] [CrossRef] [PubMed]
- Kanhayuwa, L.; Kotta-Loizou, I.; Ozkan, S.; Gunning, A.P.; Coutts, R.H. A novel mycovirus from aspergillus fumigatus contains four unique dsrnas as its genome and is infectious as dsRNA. Proc. Natl. Acad. Sci. USA 2015, 112, 9100–9105. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Dong, K.; Zhou, L.; Wang, G.; Hong, N.; Jiang, D.; Xu, W. A dsRNA virus with filamentous viral particles. Nat. Commun. 2017, 8, 168. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Li, Z.; Lai, M.; Shu, S.; Du, Y.; Zhou, Z.H.; Sun, R. In situ structures of the genome and genome-delivery apparatus in a single-stranded RNA virus. Nature 2017, 541, 112–116. [Google Scholar] [CrossRef] [PubMed]
- Huiskonen, J.T. Image processing for cryogenic transmission electron microscopy of symmetry-mismatched complexes. Biosci. Rep. 2018. [Google Scholar] [CrossRef] [PubMed]

| Virus Family | dsRNA Features | Capsid Features | ||||
|---|---|---|---|---|---|---|
| Nº Segments | Size (kbp) | MW c (MDa) | CP (residue) | Φ d/ir e (nm) | dsRNA Density (bp/100 nm3) f | |
| HSV a | 1 | ~152 | 103.7 | 1374 | ~130/43 | 46 |
| Reoviridae | ||||||
| Orthoreovirus | 10 | ~23.5 | 16 | 1275 | ~60/24.5 | 38 |
| Rotavirus | 11 | ~18.5 | 12.6 | 880 | ~52/23.5 | 34 |
| Orbivirus, BTV | 10 | ~19.2 | 13.1 | 901 | ~52/22 | 43 |
| Aquareovirus, GCRV | 11 | ~23.6 | 16 | 1027 | ~60/23 | 46 |
| Phytoreovirus, RDV | 12 | ~25.7 | 17.5 | 1019 | ~57/26 | 35 |
| Cypovirus, CPV | 10 | ~31.4 | 21.4 | 1333 | ~58/24 | 54 |
| Mycoreovirus, MyRV1 | 11 | 23.4 | 16 | |||
| Picobirnaviridae | 2 | ~4.2 | 2.9 | 590 | ~35/14 | |
| Cystoviridae, phage ϕ6 | 3 | ~13.4 | 9.1 | 769 | ~50/20 | 40 |
| Totiviridae, ScV-L-A | 1 | ~4.6 | 3.1 | 680 | ~43/17 | 22 |
| Partitiviridae, PsV-S | 1 (2) b | ~1.7 (3.3) | 1.2 (2.2) | 420 | ~35/12 | 23 |
| Chrysoviridae, PcV | 1 (4) b | ~3.2 (12.6) | 2.2 (8.6) | 109 | ~40/16 | 19 |
| Megabirnaviridae, RnMBV1 | 1 (2) b | ~8.1 (16.2) | 5.5 (11) | 135 | ~52/19 | 28 |
| Quadriviridae, RnQV1 | 1–2 (4) | ~4.3 (17.1) | 2.9 (11.7) | 1356 + 1061 | ~47/16 | 25 (50) g |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luque, D.; Mata, C.P.; Suzuki, N.; Ghabrial, S.A.; Castón, J.R. Capsid Structure of dsRNA Fungal Viruses. Viruses 2018, 10, 481. https://doi.org/10.3390/v10090481
Luque D, Mata CP, Suzuki N, Ghabrial SA, Castón JR. Capsid Structure of dsRNA Fungal Viruses. Viruses. 2018; 10(9):481. https://doi.org/10.3390/v10090481
Chicago/Turabian StyleLuque, Daniel, Carlos P. Mata, Nobuhiro Suzuki, Said A. Ghabrial, and José R. Castón. 2018. "Capsid Structure of dsRNA Fungal Viruses" Viruses 10, no. 9: 481. https://doi.org/10.3390/v10090481
APA StyleLuque, D., Mata, C. P., Suzuki, N., Ghabrial, S. A., & Castón, J. R. (2018). Capsid Structure of dsRNA Fungal Viruses. Viruses, 10(9), 481. https://doi.org/10.3390/v10090481