Insights into the Human Virome Using CRISPR Spacers from Microbiomes
Abstract
1. Introduction
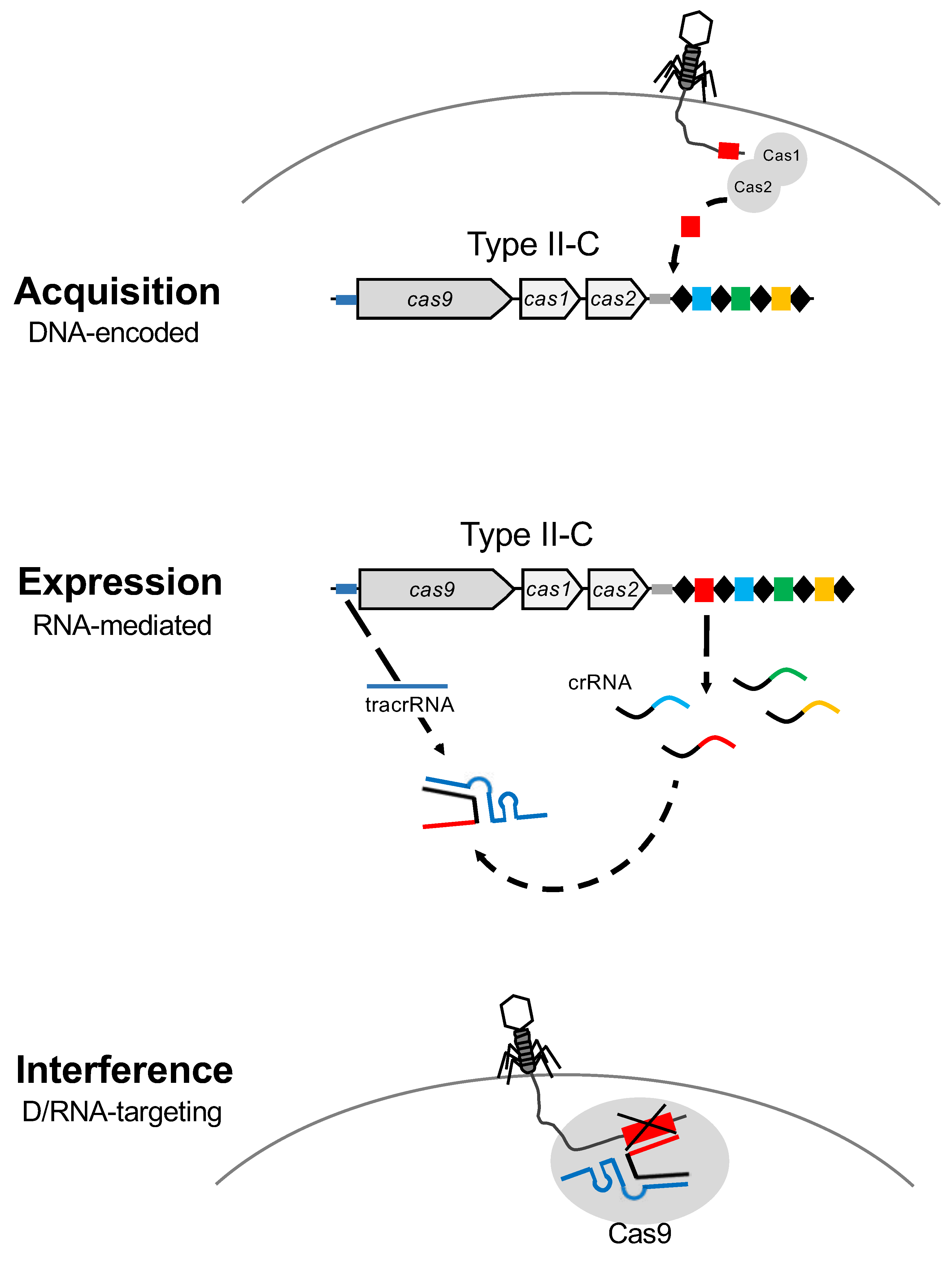
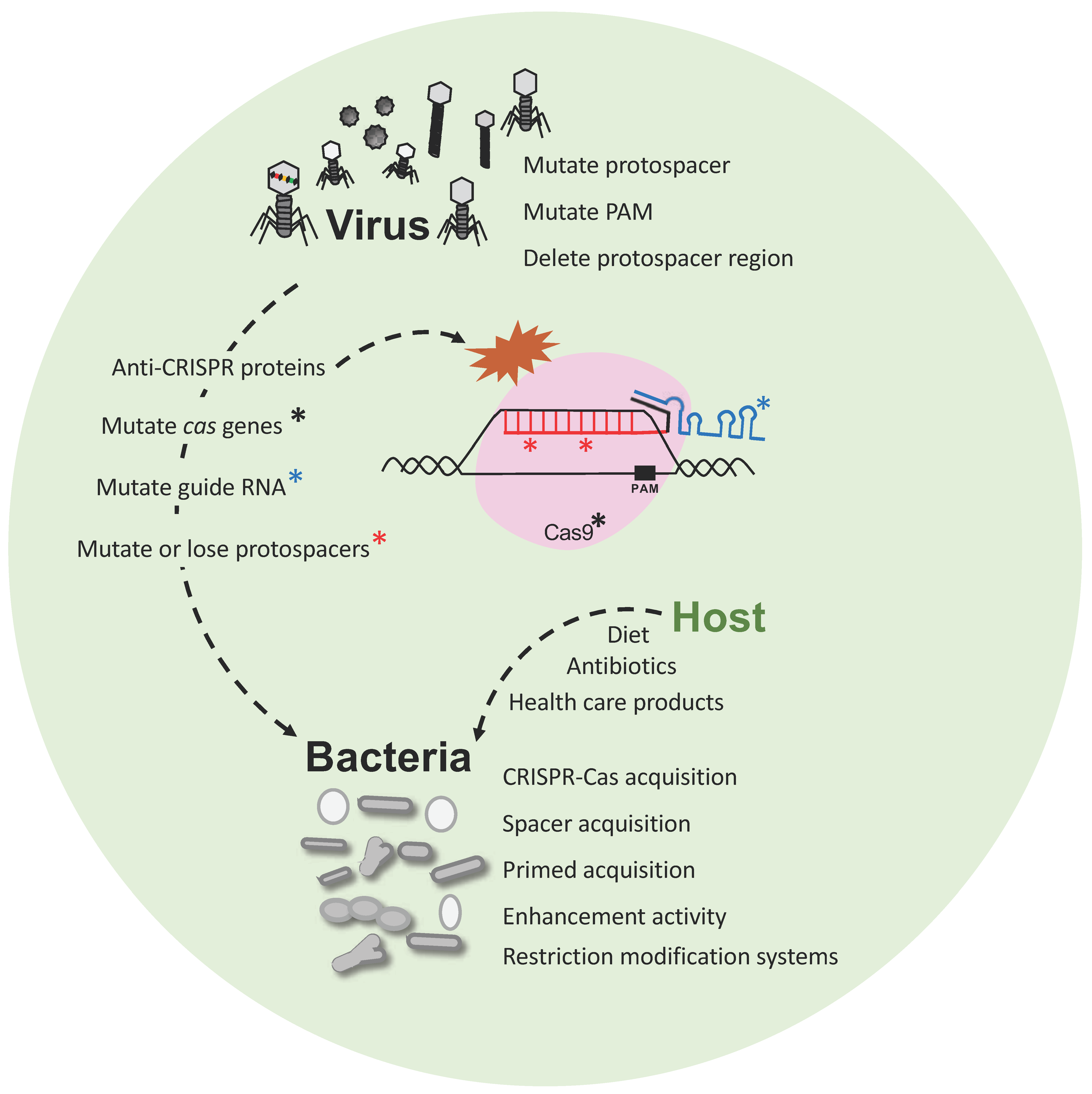
2. CRISPR-Cas as Immune System and Vaccination Record
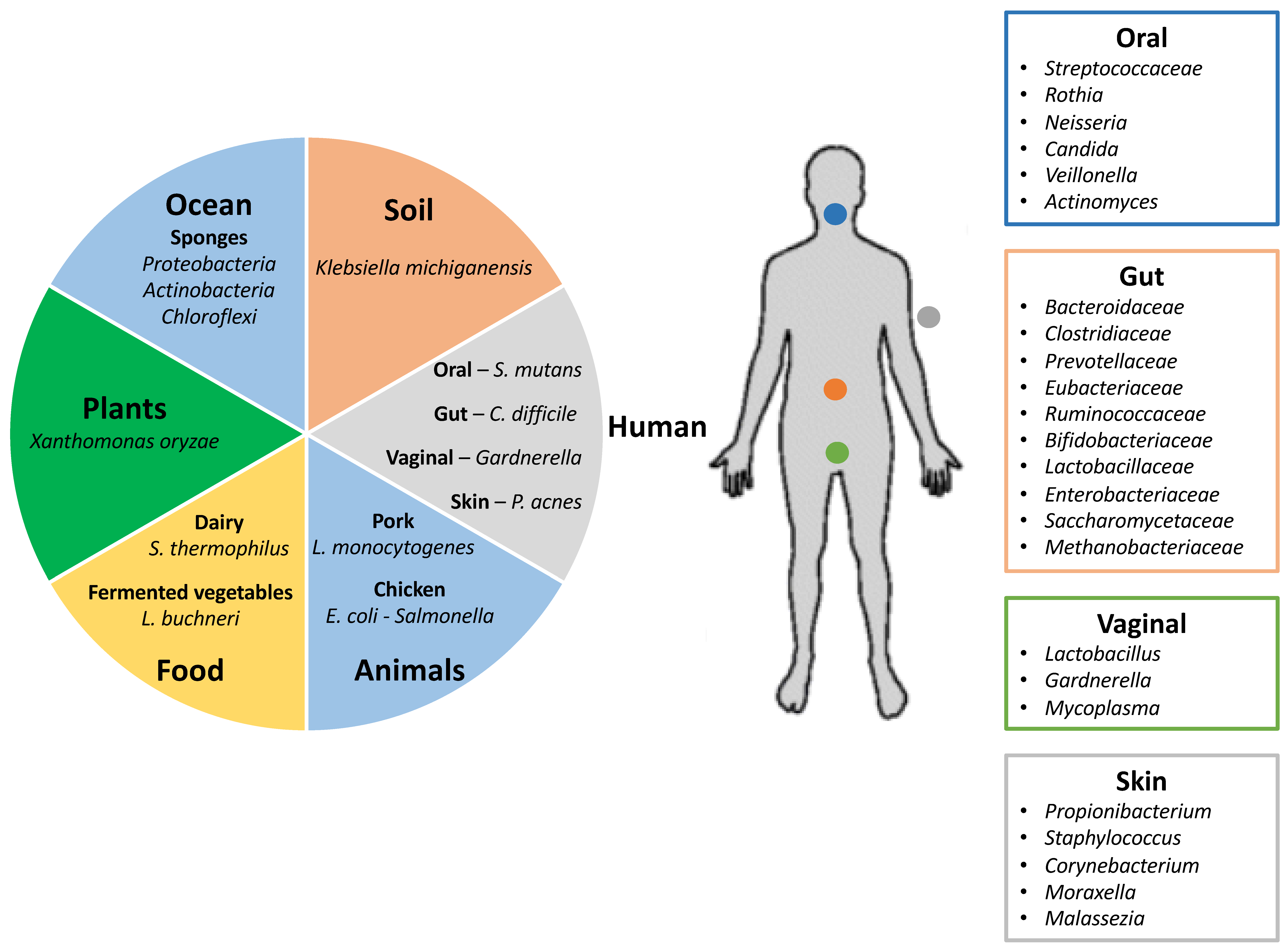
3. Investigating Host-Phage Interactions within Select Body Sites
3.1. Gastrointestinal Tract
3.2. Oral
3.3. Vaginal
3.4. Skin
4. Discussion and Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- The Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Integrative HMP (iHMP) Research Network Consortium. The integrative human microbiome project: Dynamic analysis of microbiome-host omics profiles during periods of human health and disease. Cell Host Microbe 2014, 16, 276–289. [Google Scholar] [CrossRef] [PubMed]
- Kolde, R.; Franzosa, E.A.; Rahnavard, G.; Hall, A.B.; Vlamakis, H.; Stevens, C.; Daly, M.J.; Xavier, R.J.; Huttenhower, C. Host genetic variation and its microbiome interactions within the human microbiome project. Genome Med. 2018, 10, 6. [Google Scholar] [CrossRef] [PubMed]
- Backhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 2015, 17, 690–703. [Google Scholar] [CrossRef] [PubMed]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef] [PubMed]
- Arboleya, S.; Watkins, C.; Stanton, C.; Ross, R.P. Gut bifidobacteria populations in human health and aging. Front. Microbiol. 2016, 7, 1204. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.; Byrd, A.L.; Deming, C.; Conlan, S.; Program, N.C.S.; Kong, H.H.; Segre, J.A. Biogeography and individuality shape function in the human skin metagenome. Nature 2014, 514, 59–64. [Google Scholar] [CrossRef] [PubMed]
- Breitbart, M.; Hewson, I.; Felts, B.; Mahaffy, J.M.; Nulton, J.; Salamon, P.; Rohwer, F. Metagenomic analyses of an uncultured viral community from human feces. J. Bacteriol. 2003, 185, 6220–6223. [Google Scholar] [CrossRef] [PubMed]
- Minot, S.; Bryson, A.; Chehoud, C.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. Rapid evolution of the human gut virome. Proc. Natl. Acad. Sci. USA 2013, 110, 12450–12455. [Google Scholar] [CrossRef] [PubMed]
- Minot, S.; Sinha, R.; Chen, J.; Li, H.; Keilbaugh, S.A.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. The human gut virome: Inter-individual variation and dynamic response to diet. Genome Res. 2011, 21, 1616–1625. [Google Scholar] [CrossRef] [PubMed]
- Reyes, A.; Haynes, M.; Hanson, N.; Angly, F.E.; Heath, A.C.; Rohwer, F.; Gordon, J.I. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 2010, 466, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Lim, E.S.; Zhou, Y.; Zhao, G.; Bauer, I.K.; Droit, L.; Ndao, I.M.; Warner, B.B.; Tarr, P.I.; Wang, D.; Holtz, L.R. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 2015, 21, 1228–1234. [Google Scholar] [CrossRef] [PubMed]
- Findley, K.; Grice, E.A. The skin microbiome: A focus on pathogens and their association with skin disease. PLoS Pathog. 2014, 10, e1004436. [Google Scholar] [CrossRef] [PubMed]
- Barrangou, R.; Fremaux, C.; Deveau, H.; Richards, M.; Boyaval, P.; Moineau, S.; Romero, D.A.; Horvath, P. CRISPR provides acquired resistance against viruses in prokaryotes. Science 2007, 315, 1709–1712. [Google Scholar] [CrossRef] [PubMed]
- Barrangou, R.; Marraffini, L.A. CRISPR-cas systems: Prokaryotes upgrade to adaptive immunity. Mol. Cell 2014, 54, 234–244. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo-Cantabrana, C.; Crawley, A.B.; Sanchez, B.; Barrangou, R. Characterization and exploitation of CRISPR loci in Bifidobacterium longum. Front. Microbiol. 2017, 8, 1851. [Google Scholar] [CrossRef] [PubMed]
- Almeida, F.; Medeiros, M.I.; Rodrigues, D.D.; Allard, M.W.; Falcao, J.P. Molecular characterization of Salmonella typhimurium isolated in brazil by crispr-mvlst. J. Microbiol. Methods 2017, 133, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Andersen, J.M.; Shoup, M.; Robinson, C.; Britton, R.; Olsen, K.E.; Barrangou, R. CRISPR diversity and microevolution in clostridium difficile. Genome Biol. Evol. 2016, 8, 2841–2855. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, N.L.; Petty, N.K.; Ben Zakour, N.L.; Szubert, J.M.; Savill, J.; Beatson, S.A. Genome analysis and CRISPR typing of Salmonella enterica serovar virchow. BMC Genom. 2014, 15, 389. [Google Scholar] [CrossRef] [PubMed]
- Freidlin, P.J.; Nissan, I.; Luria, A.; Goldblatt, D.; Schaffer, L.; Kaidar-Shwartz, H.; Chemtob, D.; Dveyrin, Z.; Head, S.R.; Rorman, E. Structure and variation of crispr and crispr-flanking regions in deleted-direct repeat region Mycobacterium tuberculosis complex strains. BMC Genom. 2017, 18, 168. [Google Scholar] [CrossRef] [PubMed]
- Shariat, N.; Timme, R.E.; Pettengill, J.B.; Barrangou, R.; Dudley, E.G. Characterization and evolution of Salmonella CRISPR-cas systems. Microbiology 2015, 161, 374–386. [Google Scholar] [PubMed]
- Xie, X.; Hu, Y.; Xu, Y.; Yin, K.; Li, Y.; Chen, Y.; Xia, J.; Xu, L.; Liu, Z.; Geng, S.; et al. Genetic analysis of Salmonella enterica serovar gallinarum biovar pullorum based on characterization and evolution of CRISPR sequence. Vet. Microbiol. 2017, 203, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Lloyd-Price, J.; Abu-Ali, G.; Huttenhower, C. The healthy human microbiome. Genome Med. 2016, 8, 51. [Google Scholar] [CrossRef] [PubMed]
- Horn, H.; Slaby, B.; Jahn, M.; Bayer, K.; Moitinho-Silva, L.; Foerster, F.; Abdelmohsen, U.; Hentschel, U. An enrichment of CRISPR and other defense-related features in marine sponge-associated microbial metagenomes. Front. Microbiol. 2016, 7, 1751. [Google Scholar] [CrossRef] [PubMed]
- Chylinski, K.; Le Rhun, A.; Charpentier, E. The tracrRNA and cas9 families of type II CRISPR-Cas immunity systems. RNA Biol. 2013, 10, 726–737. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; Jiang, F.; Taylor, D.W.; Sternberg, S.H.; Kaya, E.; Ma, E.; Anders, C.; Hauer, M.; Zhou, K.; Lin, S.; et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 2014, 343, 1247997. [Google Scholar] [CrossRef] [PubMed]
- Mojica, F.J.; Diez-Villasenor, C.; Garcia-Martinez, J.; Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 2009, 155, 733–740. [Google Scholar] [PubMed]
- Huo, Y.; Nam, K.H.; Ding, F.; Lee, H.; Wu, L.; Xiao, Y.; Farchione, M.D., Jr.; Zhou, S.; Rajashankar, K.; Kurinov, I.; et al. Structures of CRISPR Cas3 offer mechanistic insights into cascade-activated DNA unwinding and degradation. Nat. Struct. Mol. Biol. 2014, 21, 771–777. [Google Scholar] [CrossRef] [PubMed]
- Grissa, I.; Vergnaud, G.; Pourcel, C. Crisprfinder: A web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res. 2007, 35, W52–W57. [Google Scholar] [CrossRef] [PubMed]
- Briner, A.E.; Lugli, G.A.; Milani, C.; Duranti, S.; Turroni, F.; Gueimonde, M.; Margolles, A.; van Sinderen, D.; Ventura, M.; Barrangou, R. Occurrence and diversity of CRISPR-Cas systems in the genus Bifidobacterium. PLoS ONE 2015, 10, e0133661. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Harris, H.M.; McCann, A.; Guo, C.; Argimon, S.; Zhang, W.; Yang, X.; Jeffery, I.B.; Cooney, J.C.; Kagawa, T.F.; et al. Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat. Commun. 2015, 6, 8322. [Google Scholar] [CrossRef] [PubMed]
- Mirzaei, M.K.; Maurice, C.F. Menage a trois in the human gut: Interactions between host, bacteria and phages. Nat. Rev. Microbiol. 2017, 15, 397–408. [Google Scholar] [CrossRef] [PubMed]
- Stern, A.; Mick, E.; Tirosh, I.; Sagy, O.; Sorek, R. CRISPR targeting reveals a reservoir of common phages associated with the human gut microbiome. Genome Res. 2012, 22, 1985–1994. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Muscle: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Gogleva, A.A.; Gelfand, M.S.; Artamonova, II. Comparative analysis of CRISPR cassettes from the human gut metagenomic contigs. BMC Genom. 2014, 15, 202. [Google Scholar] [CrossRef] [PubMed]
- Ventura, M.; Turroni, F.; Lima-Mendez, G.; Foroni, E.; Zomer, A.; Duranti, S.; Giubellini, V.; Bottacini, F.; Horvath, P.; Barrangou, R.; et al. Comparative analyses of prophage-like elements present in bifidobacterial genomes. Appl. Environ. Microbiol. 2009, 75, 6929–6936. [Google Scholar] [CrossRef] [PubMed]
- Mangericao, T.C.; Peng, Z.; Zhang, X. Computational prediction of CRISPR cassettes in gut metagenome samples from chinese type-2 diabetic patients and healthy controls. BMC Syst. Biol. 2016, 10 (Suppl. 1), 5. [Google Scholar] [CrossRef] [PubMed]
- Sanozky-Dawes, R.; Selle, K.; O’Flaherty, S.; Klaenhammer, T.; Barrangou, R. Occurrence and activity of a type II CRISPR-Cas system in Lactobacillus gasseri. Microbiology 2015, 161, 1752–1761. [Google Scholar] [PubMed]
- Duranti, S.; Lugli, G.A.; Mancabelli, L.; Armanini, F.; Turroni, F.; James, K.; Ferretti, P.; Gorfer, V.; Ferrario, C.; Milani, C.; et al. Maternal inheritance of bifidobacterial communities and bifidophages in infants through vertical transmission. Microbiome 2017, 5, 66. [Google Scholar] [CrossRef] [PubMed]
- Janezic, S.; Garneau, J.R.; Monot, M. Comparative genomics of Clostridium difficile. Adv. Exp. Med. Biol. 2018, 1050, 59–75. [Google Scholar] [PubMed]
- Hargreaves, K.R.; Flores, C.O.; Lawley, T.D.; Clokie, M.R. Abundant and diverse clustered regularly interspaced short palindromic repeat spacers in Clostridium difficile strains and prophages target multiple phage types within this pathogen. mBio 2014, 5, e01045-13. [Google Scholar] [CrossRef] [PubMed]
- Rashid, S.J.; Barylski, J.; Hargreaves, K.R.; Millard, A.A.; Vinner, G.K.; Clokie, M.R. Two novel myoviruses from the north of iraq reveal insights into Clostridium difficile phage diversity and biology. Viruses 2016, 8, 310. [Google Scholar] [CrossRef] [PubMed]
- Garneau, J.R.; Sekulovic, O.; Dupuy, B.; Soutourina, O.; Monot, M.; Fortier, L.C. High prevalence and genetic diversity of large phicd211 (phicdif1296t)-like prophages in Clostridioides difficile. Appl. Environ. Microbiol. 2018, 84, e02164-17. [Google Scholar] [CrossRef] [PubMed]
- Shariat, N.; Sandt, C.H.; DiMarzio, M.J.; Barrangou, R.; Dudley, E.G. CRISPR-mvlst subtyping of Salmonella enterica subsp. enterica serovars typhimurium and heidelberg and application in identifying outbreak isolates. BMC Microbiol. 2013, 13, 254. [Google Scholar]
- Sola, C.; Abadia, E.; Le Hello, S.; Weill, F.X. High-throughput CRISPR typing of Mycobacterium tuberculosis complex and salmonella enterica serotype typhimurium. Methods Mol. Biol. 2015, 1311, 91–109. [Google Scholar] [PubMed]
- Verma, D.; Garg, P.K.; Dubey, A.K. Insights into the human oral microbiome. Arch. Microbiol. 2018, 200, 525–540. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Haake, S.K.; Mannon, P.; Lemon, K.P.; Waldron, L.; Gevers, D.; Huttenhower, C.; Izard, J. Composition of the adult digestive tract bacterial microbiome based on seven mouth surfaces, tonsils, throat and stool samples. Genome Biol. 2012, 13, R42. [Google Scholar] [CrossRef] [PubMed]
- Pride, D.T.; Salzman, J.; Haynes, M.; Rohwer, F.; Davis-Long, C.; White, R.A., 3rd; Loomer, P.; Armitage, G.C.; Relman, D.A. Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J. 2012, 6, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Ly, M.; Abeles, S.R.; Boehm, T.K.; Robles-Sikisaka, R.; Naidu, M.; Santiago-Rodriguez, T.; Pride, D.T. Altered oral viral ecology in association with periodontal disease. mBio 2014, 5, e01133-14. [Google Scholar] [CrossRef] [PubMed]
- Pride, D.T.; Sun, C.L.; Salzman, J.; Rao, N.; Loomer, P.; Armitage, G.C.; Banfield, J.F.; Relman, D.A. Analysis of streptococcal CRISPRS from human saliva reveals substantial sequence diversity within and between subjects over time. Genome Res. 2011, 21, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Robles-Sikisaka, R.; Naidu, M.; Ly, M.; Salzman, J.; Abeles, S.R.; Boehm, T.K.; Pride, D.T. Conservation of streptococcal CRISPRS on human skin and saliva. BMC Microbiol. 2014, 14, 146. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Rho, M.; Tang, H.; Doak, T.G.; Ye, Y. CRISPR-Cas systems target a diverse collection of invasive mobile genetic elements in human microbiomes. Genome Biol. 2013, 14, R40. [Google Scholar] [CrossRef] [PubMed]
- Rho, M.; Wu, Y.W.; Tang, H.; Doak, T.G.; Ye, Y. Diverse CRISPRS evolving in human microbiomes. PLoS Genet. 2012, 8, e1002441. [Google Scholar] [CrossRef] [PubMed]
- Naidu, M.; Robles-Sikisaka, R.; Abeles, S.R.; Boehm, T.K.; Pride, D.T. Characterization of bacteriophage communities and crispr profiles from dental plaque. BMC Microbiol. 2014, 14, 175. [Google Scholar] [CrossRef] [PubMed]
- Aagaard, K.; Riehle, K.; Ma, J.; Segata, N.; Mistretta, T.A.; Coarfa, C.; Raza, S.; Rosenbaum, S.; Van den Veyver, I.; Milosavljevic, A.; et al. A metagenomic approach to characterization of the vaginal microbiome signature in pregnancy. PLoS ONE 2012, 7, e36466. [Google Scholar] [CrossRef] [PubMed]
- Beauruelle, C.; Pastuszka, A.; Horvath, P.; Perrotin, F.; Mereghetti, L.; Lanotte, P. CRISPR: A useful genetic feature to follow vaginal carriage of group b streptococcus. Front. Microbiol. 2017, 8, 1981. [Google Scholar] [CrossRef] [PubMed]
- Davies, M.R.; McMillan, D.J.; Van Domselaar, G.H.; Jones, M.K.; Sriprakash, K.S. Phage 3396 from a Streptococcus dysgalactiae subsp. equisimilis pathovar may have its origins in streptococcus pyogenes. J. Bacteriol. 2007, 189, 2646–2652. [Google Scholar] [PubMed]
- Malki, K.; Shapiro, J.W.; Price, T.K.; Hilt, E.E.; Thomas-White, K.; Sircar, T.; Rosenfeld, A.B.; Kuffel, G.; Zilliox, M.J.; Wolfe, A.J.; et al. Genomes of Gardnerella strains reveal an abundance of prophages within the bladder microbiome. PLoS ONE 2016, 11, e0166757. [Google Scholar] [CrossRef] [PubMed]
- Pleckaityte, M.; Zilnyte, M.; Zvirbliene, A. Insights into the CRISPR/Cas system of Gardnerella vaginalis. BMC Microbiol. 2012, 12, 301. [Google Scholar] [CrossRef] [PubMed]
- Martin, R.; Escobedo, S.; Suarez, J.E. Induction, structural characterization, and genome sequence of Lv1, a prophage from a human vaginal Lactobacillus jensenii strain. Int. Microbiol. 2010, 13, 113–121. [Google Scholar] [PubMed]
- Pavlova, S.I.; Kilic, A.O.; Mou, S.M.; Tao, L. Phage infection in vaginal lactobacilli: An in vitro study. Infect. Dis. Obstet. Gynecol. 1997, 5, 36–44. [Google Scholar] [PubMed]
- Kilic, A.O.; Pavlova, S.I.; Alpay, S.; Kilic, S.S.; Tao, L. Comparative study of vaginal Lactobacillus phages isolated from women in the united states and turkey: Prevalence, morphology, host range, and DNA homology. Clin. Diagn. Lab. Immunol. 2001, 8, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Diez-Villasenor, C.; Almendros, C.; Garcia-Martinez, J.; Mojica, F.J. Diversity of CRISPR loci in Escherichia coli. Microbiology 2010, 156, 1351–1361. [Google Scholar] [PubMed]
- Sheludchenko, M.S.; Huygens, F.; Stratton, H.; Hargreaves, M. CRISPR diversity in E. coli isolates from australian animals, humans and environmental waters. PLoS ONE 2015, 10, e0124090. [Google Scholar] [CrossRef] [PubMed]
- Touchon, M.; Charpentier, S.; Clermont, O.; Rocha, E.P.; Denamur, E.; Branger, C. CRISPR distribution within the Escherichia coli species is not suggestive of immunity-associated diversifying selection. J. Bacteriol. 2011, 193, 2460–2467. [Google Scholar] [CrossRef] [PubMed]
- Savitskaya, E.; Lopatina, A.; Medvedeva, S.; Kapustin, M.; Shmakov, S.; Tikhonov, A.; Artamonova, I.I.; Logacheva, M.; Severinov, K. Dynamics of escherichia coli type i-e crispr spacers over 42 000 years. Mol. Ecol. 2017, 26, 2019–2026. [Google Scholar] [CrossRef] [PubMed]
- Dang, T.N.; Zhang, L.; Zollner, S.; Srinivasan, U.; Abbas, K.; Marrs, C.F.; Foxman, B. Uropathogenic Escherichia coli are less likely than paired fecal E. coli to have CRISPR loci. Infect. Genet. Evol. 2013, 19, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Segre, J.A. The skin microbiome. Nat. Rev. Microbiol. 2011, 9, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Brandwein, M.; Steinberg, D.; Meshner, S. Microbial biofilms and the human skin microbiome. Biofilms Microbiomes 2016, 2, 3. [Google Scholar] [CrossRef] [PubMed]
- Schommer, N.N.; Gallo, R.L. Structure and function of the human skin microbiome. Trends Microbiol. 2013, 21, 660–668. [Google Scholar] [CrossRef] [PubMed]
- Chng, K.R.; Tay, A.S.; Li, C.; Ng, A.H.; Wang, J.; Suri, B.K.; Matta, S.A.; McGovern, N.; Janela, B.; Wong, X.F.; et al. Whole metagenome profiling reveals skin microbiome-dependent susceptibility to atopic dermatitis flare. Nat. Microbiol. 2016, 1, 16106. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, M.E.; Schaffer, J.V.; Orlow, S.J.; Gao, Z.; Li, H.; Alekseyenko, A.V.; Blaser, M.J. Cutaneous microbiome effects of fluticasone propionate cream and adjunctive bleach baths in childhood atopic dermatitis. J. Am. Acad. Dermatol. 2016, 75, 481–493. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.A.; Connolly, J.; Hourihane, J.O.; Fallon, P.G.; McLean, W.H.I.; Murray, D.; Jo, J.H.; Segre, J.A.; Kong, H.H.; Irvine, A.D. Skin microbiome before development of atopic dermatitis: Early colonization with commensal staphylococci at 2 months is associated with a lower risk of atopic dermatitis at 1 year. J. Allergy Clin. Immunol. 2017, 139, 166–172. [Google Scholar] [CrossRef] [PubMed]
- Codoner, F.M.; Ramirez-Bosca, A.; Climent, E.; Carrion-Gutierrez, M.; Guerrero, M.; Perez-Orquin, J.M.; Horga de la Parte, J.; Genoves, S.; Ramon, D.; Navarro-Lopez, V.; et al. Gut microbial composition in patients with psoriasis. Sci. Rep. 2018, 8, 3812. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.L.; Fernandez-Penas, P. The microbiome and atopic eczema: More than skin deep. Australas. J. Dermatol. 2017, 58, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Goerke, C.; Wirtz, C.; Fluckiger, U.; Wolz, C. Extensive phage dynamics in Staphylococcus aureus contributes to adaptation to the human host during infection. Mol. Microbiol. 2006, 61, 1673–1685. [Google Scholar] [CrossRef] [PubMed]
- Goerke, C.; Pantucek, R.; Holtfreter, S.; Schulte, B.; Zink, M.; Grumann, D.; Broker, B.M.; Doskar, J.; Wolz, C. Diversity of prophages in dominant Staphylococcus aureus clonal lineages. J. Bacteriol. 2009, 191, 3462–3468. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Liu, J.; Shao, F.; Wang, P.; Duan, G.; Yang, H. Analysis of the features of 45 identified CRISPR loci in 32 Staphylococcus aureus. Biochem. Biophys. Res. Commun. 2015, 464, 894–900. [Google Scholar] [CrossRef] [PubMed]
- Chhibber, S.; Shukla, A.; Kaur, S. Transfersomal phage cocktail is an effective treatment against methicillin-resistant Staphylococcus aureus-mediated skin and soft tissue infections. Antimicrob. Agents Chemother. 2017, 61, e02146-16. [Google Scholar] [CrossRef] [PubMed]
- Bikard, D.; Euler, C.W.; Jiang, W.; Nussenzweig, P.M.; Goldberg, G.W.; Duportet, X.; Fischetti, V.A.; Marraffini, L.A. Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat. Biotechnol. 2014, 32, 1146–1150. [Google Scholar] [CrossRef] [PubMed]
- Guan, J.; Wang, W.; Sun, B. Chromosomal targeting by the type III-A CRISPR-Cas system can reshape genomes in Staphylococcus aureus. mSphere 2017, 2, e00403-17. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Xie, X.; Yin, K.; Tang, Y.; Zhou, X.; Chen, Y.; Xia, J.; Hu, Y.; Ingmer, H.; Li, Y.; et al. Characterization of CRISPR-Cas system in clinical Staphylococcus epidermidis strains revealed its potential association with bacterial infection sites. Microbiol. Res. 2016, 193, 103–110. [Google Scholar] [CrossRef] [PubMed]
- van Belkum, A.; Soriaga, L.B.; LaFave, M.C.; Akella, S.; Veyrieras, J.B.; Barbu, E.M.; Shortridge, D.; Blanc, B.; Hannum, G.; Zambardi, G.; et al. Phylogenetic distribution of CRISPR-Cas systems in antibiotic-resistant Pseudomonas aeruginosa. mBio 2015, 6, e01796-15. [Google Scholar] [CrossRef] [PubMed]
- Zegans, M.E.; Wagner, J.C.; Cady, K.C.; Murphy, D.M.; Hammond, J.H.; O’Toole, G.A. Interaction between bacteriophage dms3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J. Bacteriol. 2009, 191, 210–219. [Google Scholar] [CrossRef] [PubMed]
- Cady, K.C.; Bondy-Denomy, J.; Heussler, G.E.; Davidson, A.R.; O’Toole, G.A. The CRISPR/Cas adaptive immune system of Pseudomonas aeruginosa mediates resistance to naturally occurring and engineered phages. J. Bacteriol. 2012, 194, 5728–5738. [Google Scholar] [CrossRef] [PubMed]
- Heussler, G.E.; Miller, J.L.; Price, C.E.; Collins, A.J.; O’Toole, G.A. Requirements for Pseudomonas aeruginosa type I-F CRISPR-Cas adaptation determined using a biofilm enrichment assay. J. Bacteriol. 2016, 198, 3080–3090. [Google Scholar] [CrossRef] [PubMed]
- Hoyland-Kroghsbo, N.M.; Paczkowski, J.; Mukherjee, S.; Broniewski, J.; Westra, E.; Bondy-Denomy, J.; Bassler, B.L. Quorum sensing controls the Pseudomonas aeruginosa CRISPR-Cas adaptive immune system. Proc. Natl. Acad. Sci. USA 2017, 114, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Fitz-Gibbon, S.; Tomida, S.; Chiu, B.H.; Nguyen, L.; Du, C.; Liu, M.; Elashoff, D.; Erfe, M.C.; Loncaric, A.; Kim, J.; et al. Propionibacterium acnes strain populations in the human skin microbiome associated with acne. J. Investig. Dermatol. 2013, 133, 2152–2160. [Google Scholar] [CrossRef] [PubMed]
- Bruggemann, H.; Lomholt, H.B.; Tettelin, H.; Kilian, M. CRISPR/Cas loci of type II Propionibacterium acnes confer immunity against acquisition of mobile elements present in type I P. acnes. PLoS ONE 2012, 7, e34171. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Yan, R.; Zhong, Q.; Ngo, S.; Bangayan, N.J.; Nguyen, L.; Lui, T.; Liu, M.; Erfe, M.C.; Craft, N.; et al. The diversity and host interactions of Propionibacterium acnes bacteriophages on human skin. ISME J. 2015, 9, 2078–2093. [Google Scholar] [CrossRef] [PubMed]
- Barnard, E.; Shi, B.; Kang, D.; Craft, N.; Li, H. The balance of metagenomic elements shapes the skin microbiome in acne and health. Sci. Rep. 2016, 6, 39491. [Google Scholar] [CrossRef] [PubMed]
- Makarova, K.S.; Zhang, F.; Koonin, E.V. Snapshot: Class 1 CRISPR-Cas systems. Cell 2017, 168, 946. [Google Scholar] [CrossRef] [PubMed]
- Makarova, K.S.; Zhang, F.; Koonin, E.V. Snapshot: Class 2 CRISPR-Cas systems. Cell 2017, 168, 328. [Google Scholar] [CrossRef] [PubMed]
- Pawluk, A.; Davidson, A.R.; Maxwell, K.L. Anti-crispr: Discovery, mechanism and function. Nat. Rev. Microbiol. 2018, 16, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhang, F.; Huang, Z. Structural insights into the inactivation of CRISPR-Cas systems by diverse anti-CRISPR proteins. BMC Biol. 2018, 16, 32. [Google Scholar] [CrossRef] [PubMed]
- Seed, K.D.; Lazinski, D.W.; Calderwood, S.B.; Camilli, A. A bacteriophage encodes its own CRISPR/Cas adaptive response to evade host innate immunity. Nature 2013, 494, 489–491. [Google Scholar] [CrossRef] [PubMed]
- Hannigan, G.D.; Duhaime, M.B.; Koutra, D.; Schloss, P.D. Biogeography and environmental conditions shape bacteriophage-bacteria networks across the human microbiome. PLoS Comput. Biol. 2018, 14, e1006099. [Google Scholar] [CrossRef] [PubMed]
- Forde, A.; Hill, C. Phages of life—The path to pharma. Br. J. Pharmacol. 2018, 175, 412–418. [Google Scholar] [CrossRef] [PubMed]
- Arboleya, S.; Martinez-Camblor, P.; Solis, G.; Suarez, M.; Fernandez, N.; de Los Reyes-Gavilan, C.G.; Gueimonde, M. Intestinal microbiota and weight-gain in preterm neonates. Front. Microbiol. 2017, 8, 183. [Google Scholar] [CrossRef] [PubMed]
- Furfaro, L.L.; Chang, B.J.; Payne, M.S. Applications for bacteriophage therapy during pregnancy and the perinatal period. Front. Microbiol. 2017, 8, 2660. [Google Scholar] [CrossRef] [PubMed]
- Nogacka, A.M.; Salazar, N.; Arboleya, S.; Suarez, M.; Fernandez, N.; Solis, G.; de Los Reyes-Gavilan, C.G.; Gueimonde, M. Early microbiota, antibiotics and health. Cell. Mol. Life Sci. 2018, 75, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Dutilh, B.E.; Cassman, N.; McNair, K.; Sanchez, S.E.; Silva, G.G.; Boling, L.; Barr, J.J.; Speth, D.R.; Seguritan, V.; Aziz, R.K.; et al. A highly abundant bacteriophage discovered in the unknown sequences of human faecal metagenomes. Nat. Commun. 2014, 5, 4498. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Makarova, K.S.; Gussow, A.B.; Krupovic, M.; Segall, A.; Edwards, R.A.; Koonin, E.V. Discovery of an expansive bacteriophage family that includes the most abundant viruses from the human gut. Nat. Microbiol. 2018, 3, 38–46. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hidalgo-Cantabrana, C.; Sanozky-Dawes, R.; Barrangou, R. Insights into the Human Virome Using CRISPR Spacers from Microbiomes. Viruses 2018, 10, 479. https://doi.org/10.3390/v10090479
Hidalgo-Cantabrana C, Sanozky-Dawes R, Barrangou R. Insights into the Human Virome Using CRISPR Spacers from Microbiomes. Viruses. 2018; 10(9):479. https://doi.org/10.3390/v10090479
Chicago/Turabian StyleHidalgo-Cantabrana, Claudio, Rosemary Sanozky-Dawes, and Rodolphe Barrangou. 2018. "Insights into the Human Virome Using CRISPR Spacers from Microbiomes" Viruses 10, no. 9: 479. https://doi.org/10.3390/v10090479
APA StyleHidalgo-Cantabrana, C., Sanozky-Dawes, R., & Barrangou, R. (2018). Insights into the Human Virome Using CRISPR Spacers from Microbiomes. Viruses, 10(9), 479. https://doi.org/10.3390/v10090479





