Identification of Dual Receptor Binding Protein Systems in Lactococcal 936 Group Phages
Abstract
1. Introduction
2. Materials and Methods
2.1. Host Strains and Phages
2.2. Mass Spectrometry Analysis
2.3. Electron Microscopy
2.4. Individual RBP Cloning and Production
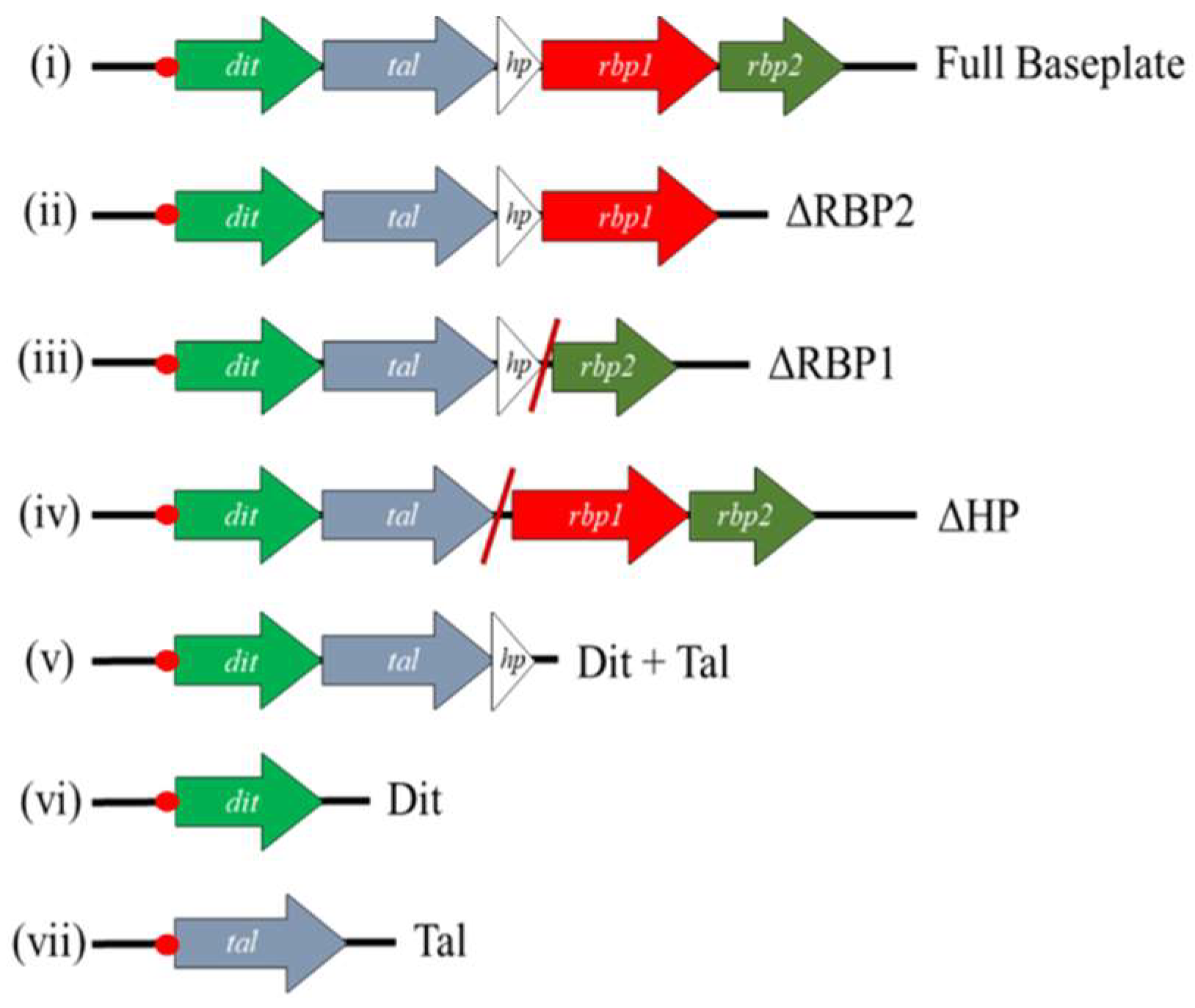
2.5. Block Cloning and Expression of Baseplate Proteins
2.6. Antibody Production, Western Blot Analysis, and Immunogold Labelling
2.7. Adsorption Inhibition Assays
2.8. Fluorescent Binding of RBP1Phi4.2
2.9. SEC MALS Analysis to Determine Baseplate Size
2.10. Negative Stain EM Analysis of Baseplate Complexes
3. Results
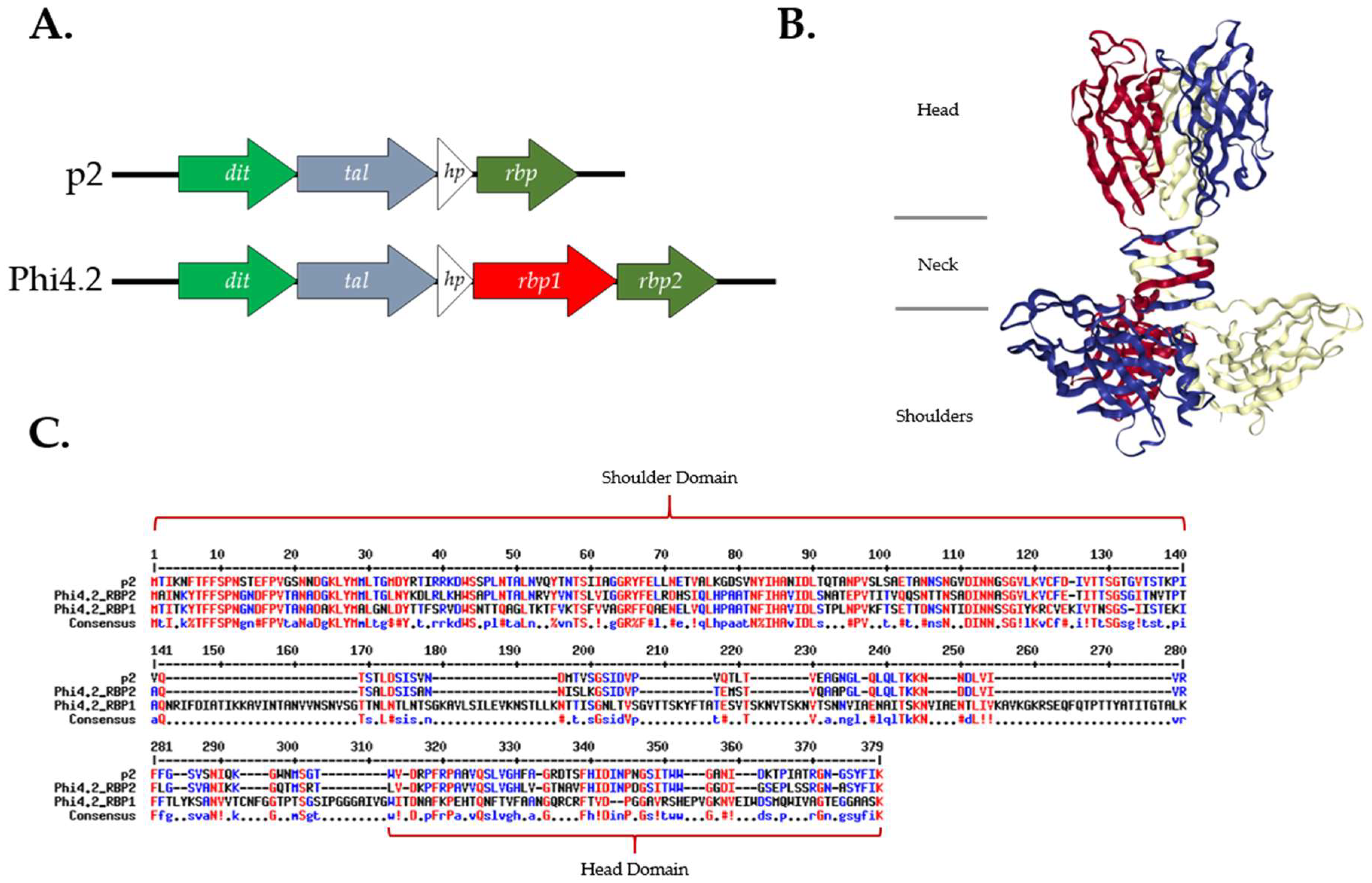
3.1. RBP1Phi4.2 Is Distinct from Current 936 RBP Groups
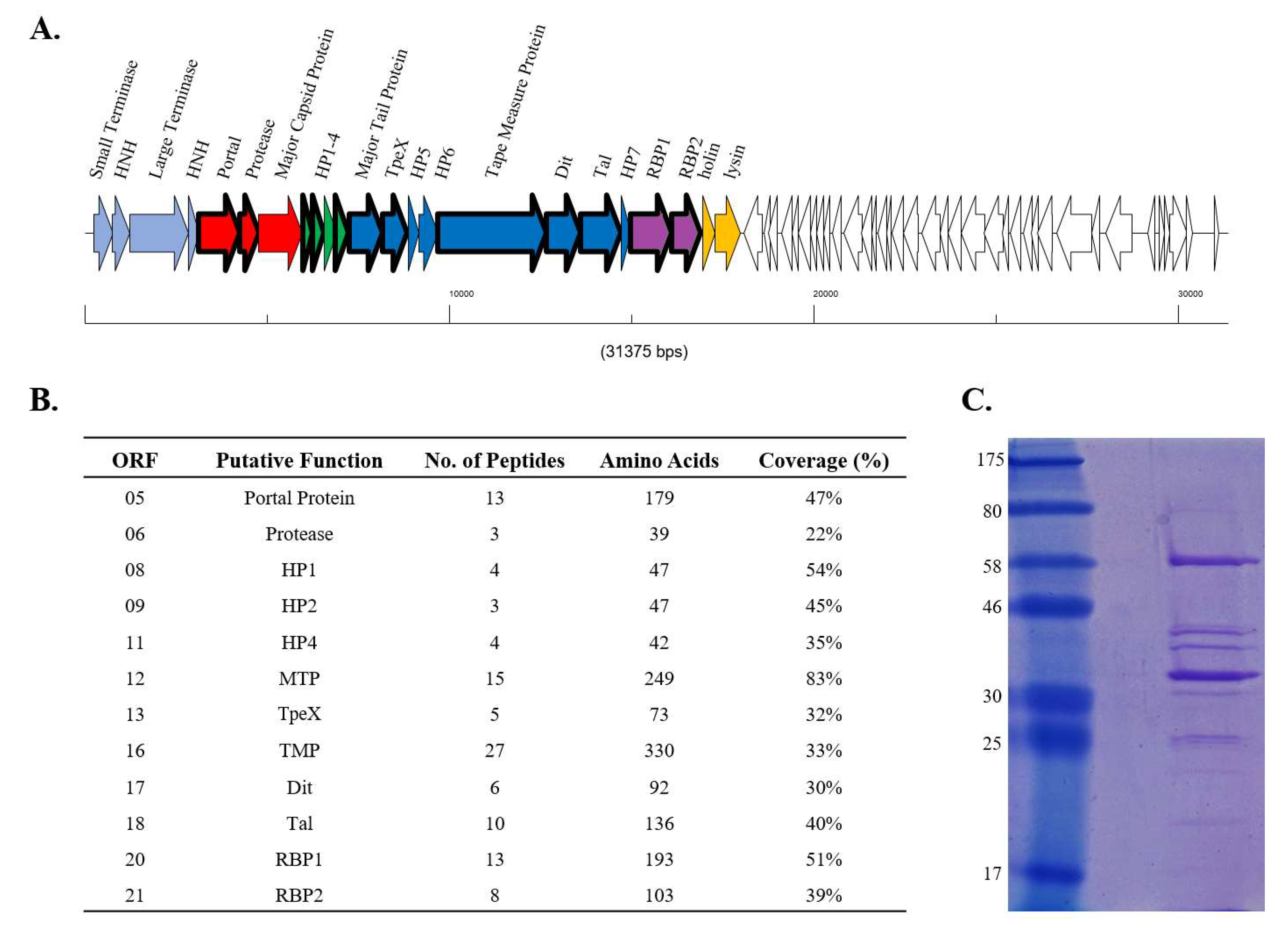
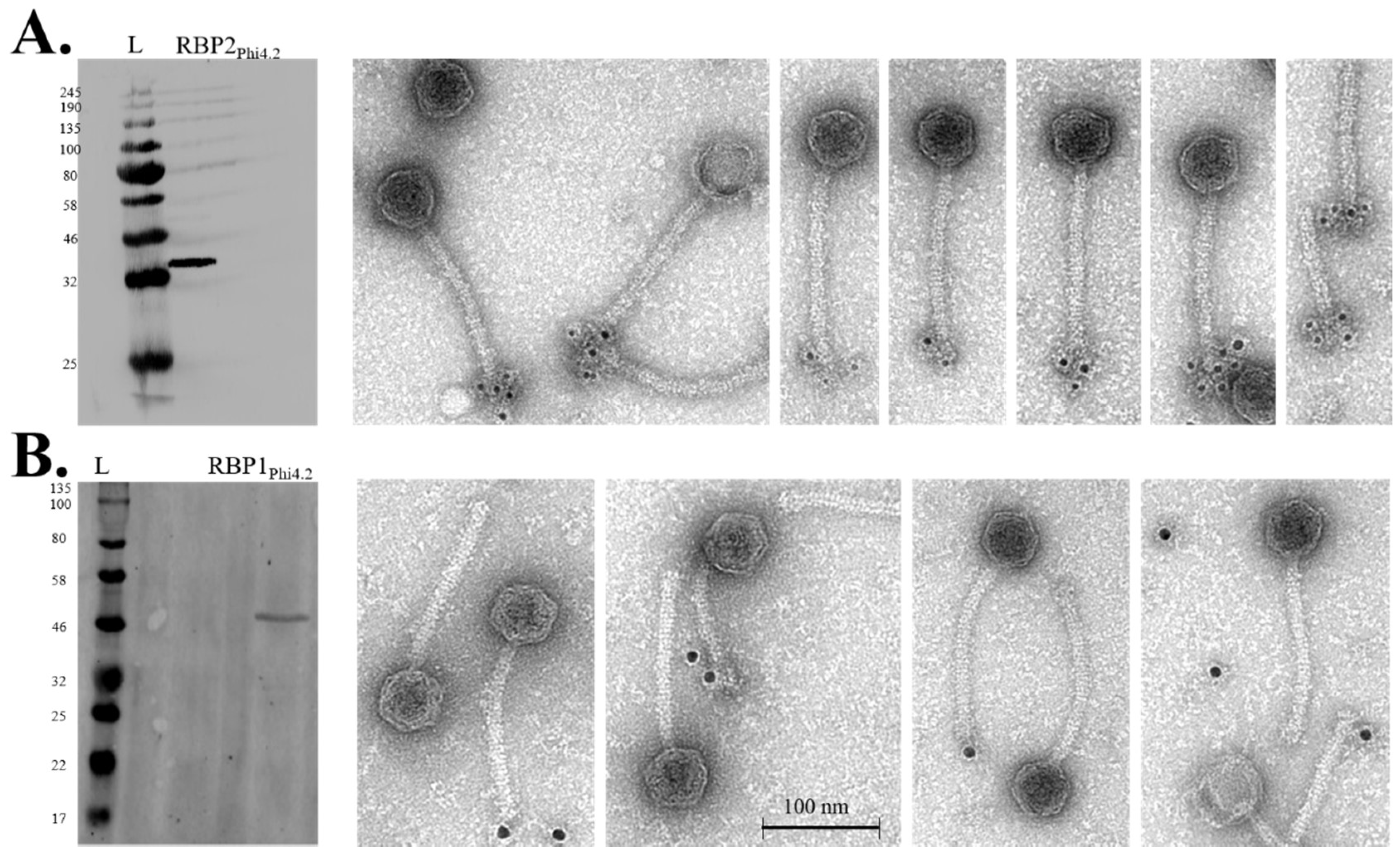
3.2. RBP1Phi4.2 Is a Structural Protein and Part of the Mature Virion
3.3. Phi4.2 Exhibits Distinct Features among 936 Group Phages
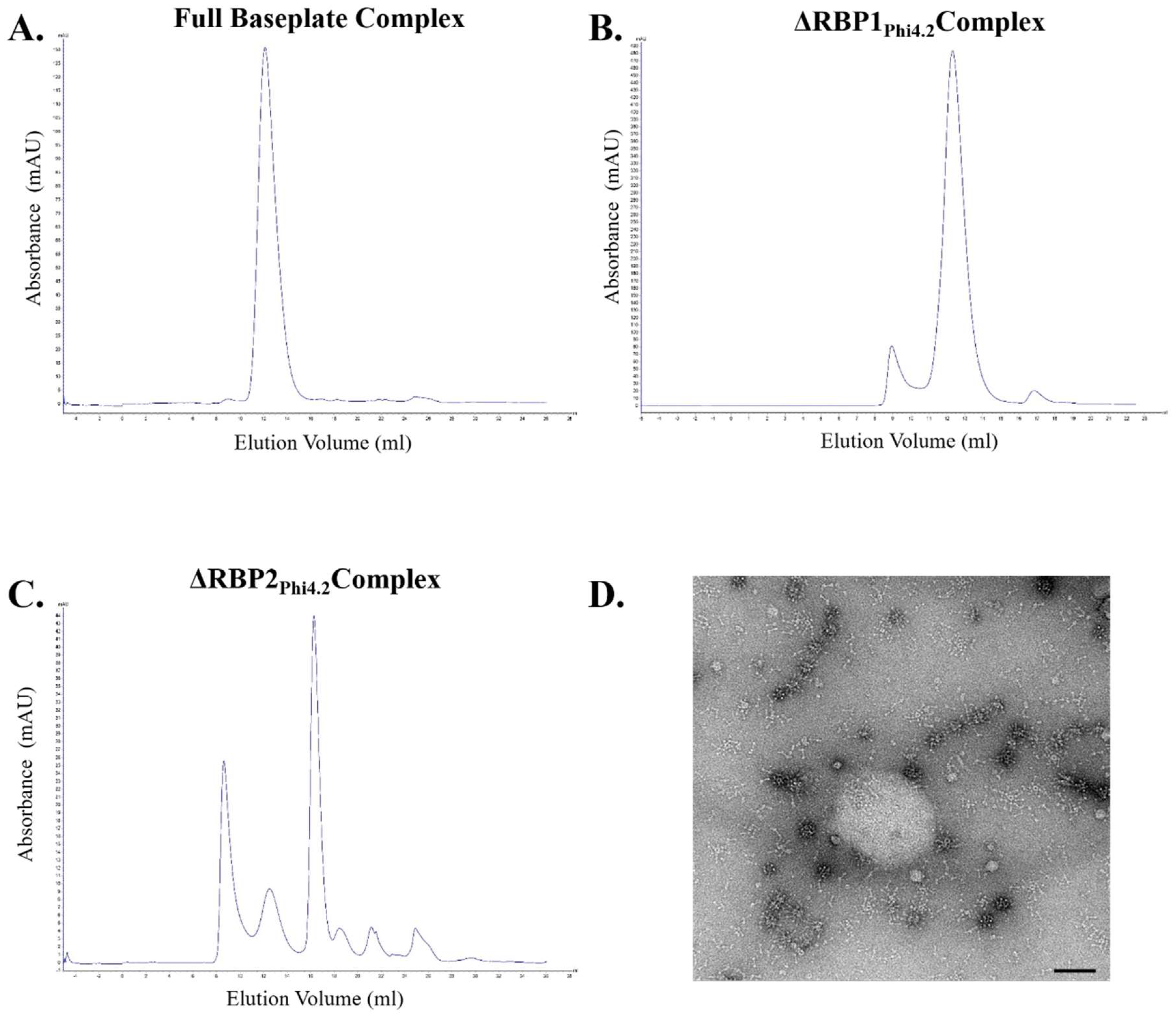
3.4. RBP1Phi4.2 Forms a Complex with Other Baseplate Components
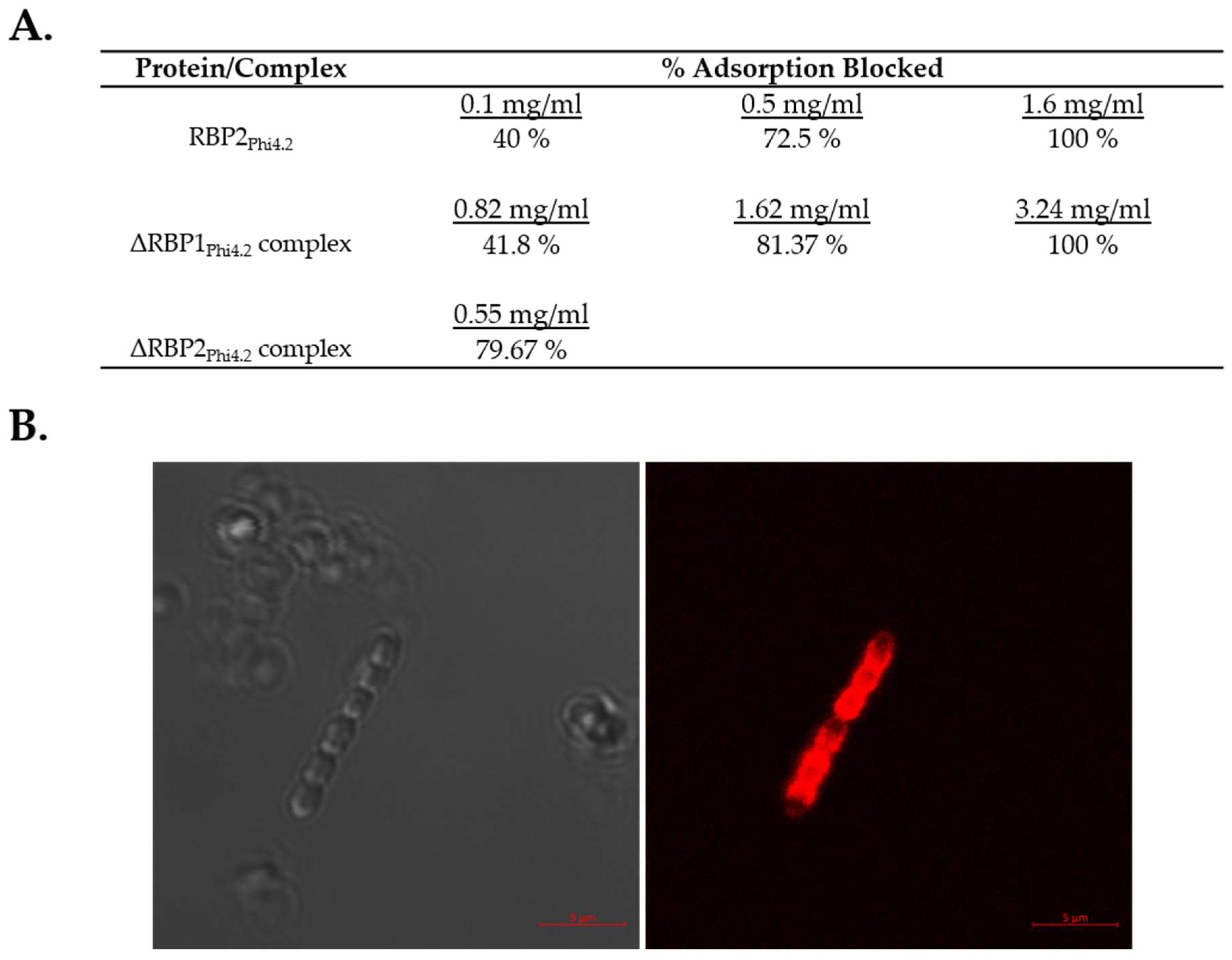
3.5. Both RBP1Phi4.2 and RBP2Phi4.2 Bind Specifically to Lactococcal Host Cells
3.6. Western Blot and Immunogold Labelling
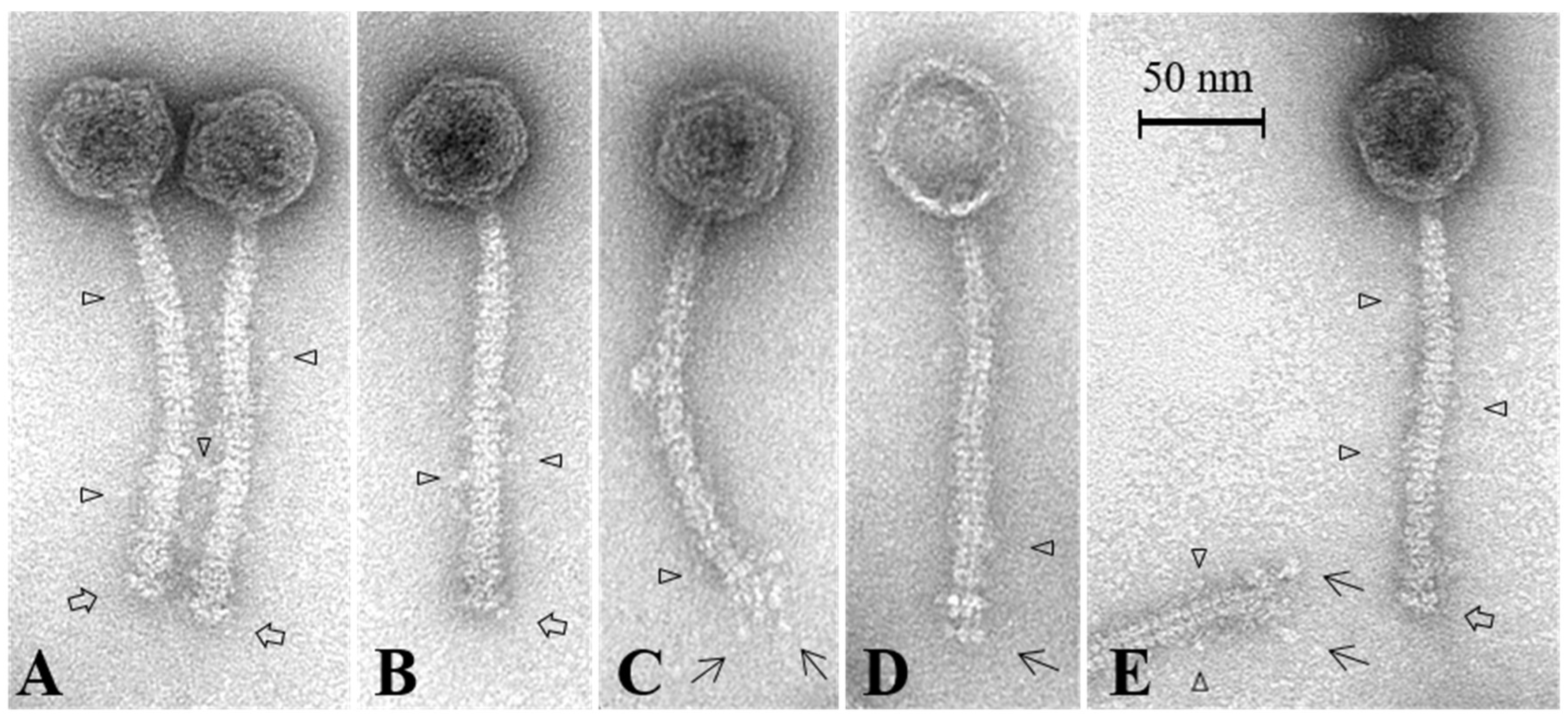
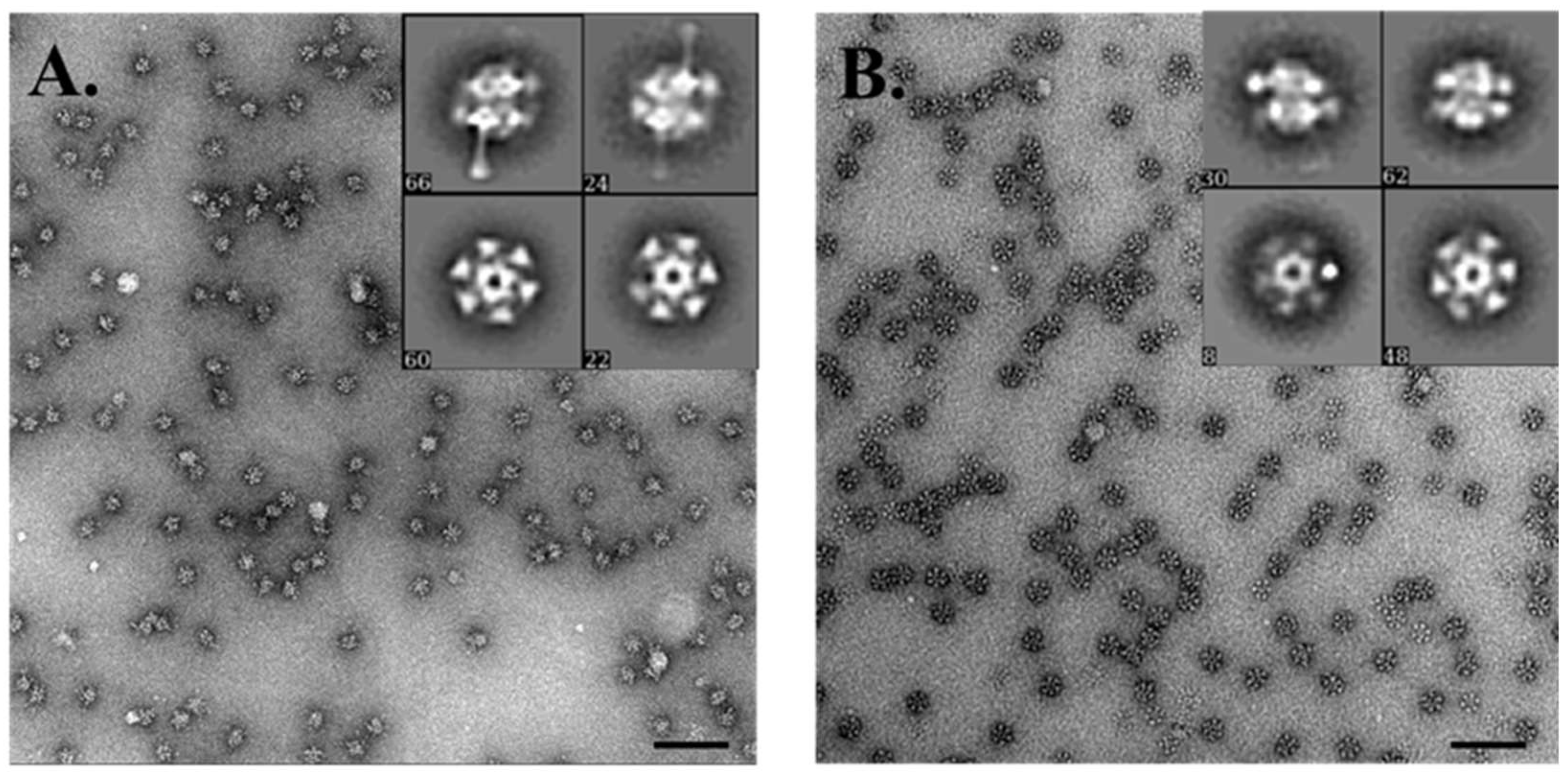
3.7. Electron Microscopy Analysis of the Baseplate of Phi4.2
3.8. SEC/MALS Determination of Baseplate Size
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Garneau, J.E.; Moineau, S. In Bacteriophages of lactic acid bacteria and their impact on milk fermentations. Microb. Cell Fact. 2011, 10, S20. [Google Scholar] [CrossRef] [PubMed]
- Deveau, H.; Labrie, S.J.; Chopin, M.-C.; Moineau, S. Biodiversity and classification of lactococcal phages. Appl. Environ. Microbiol. 2006, 72, 4338–4346. [Google Scholar] [CrossRef] [PubMed]
- Mahony, J.; Murphy, J.; van Sinderen, D. Lactococcal 936-type phages and dairy fermentation problems: From detection to evolution and prevention. Front. Microbiol. 2012, 3, 335. [Google Scholar] [CrossRef] [PubMed]
- Hayes, S.; Vincentelli, R.; Mahony, J.; Nauta, A.; Ramond, L.; Lugli, G.A.; Ventura, M.; van Sinderen, D.; Cambillau, C. Functional carbohydrate binding modules identified in evolved Dits from siphophages infecting various gram-positive bacteria. Mol. Microbiol. 2018, 110, 777–795. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.; Bottacini, F.; Mahony, J.; Kelleher, P.; Neve, H.; Zomer, A.; Nauta, A.; van Sinderen, D. Comparative genomics and functional analysis of the 936 group of lactococcal Siphoviridae phages. Sci. Rep. 2016, 6, 21345. [Google Scholar] [CrossRef] [PubMed]
- Mahony, J.; Van Sinderen, D. Structural aspects of the interaction of dairy phages with their host bacteria. Viruses 2012, 4, 1410–1424. [Google Scholar] [CrossRef] [PubMed]
- Dunne, M.; Hupfeld, M.; Klumpp, J.; Loessner, M. Molecular basis of bacterial host interactions by gram-positive targeting bacteriophages. Viruses 2018, 10, 397. [Google Scholar] [CrossRef] [PubMed]
- Molineux, I.J.; Panja, D. Popping the cork: Mechanisms of phage genome ejection. Nat. Rev. Microbiol. 2013, 11, 194–204. [Google Scholar] [CrossRef] [PubMed]
- Taylor, N.M.; Prokhorov, N.S.; Guerrero-Ferreira, R.C.; Shneider, M.M.; Browning, C.; Goldie, K.N.; Stahlberg, H.; Leiman, P.G. Structure of the T4 baseplate and its function in triggering sheath contraction. Nature 2016, 533, 346–352. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Margolin, W.; Molineux, I.J.; Liu, J. Structural remodeling of bacteriophage T4 and host membranes during infection initiation. Proc. Natl. Acad. Sci. USA 2015, 112, E4919–E4928. [Google Scholar] [CrossRef] [PubMed]
- Veesler, D.; Robin, G.; Lichière, J.; Auzat, I.; Tavares, P.; Bron, P.; Campanacci, V.; Cambillau, C. Crystal structure of bacteriophage SPP1 distal tail protein (gp 19.1): A baseplate hub paradigm in Gram+ infecting phages. J. Biol. Chem. 2010, 285, 36666–36673. [Google Scholar] [CrossRef] [PubMed]
- Vinga, I.; Baptista, C.; Auzat, I.; Petipas, I.; Lurz, R.; Tavares, P.; Santos, M.A.; São-José, C. Role of bacteriophage SPP1 tail spike protein gp21 on host cell receptor binding and trigger of phage DNA ejection. Mol. Microbiol. 2012, 83, 289–303. [Google Scholar] [CrossRef] [PubMed]
- Sciara, G.; Bebeacua, C.; Bron, P.; Tremblay, D.; Ortiz-Lombardia, M.; Lichière, J.; Van Heel, M.; Campanacci, V.; Moineau, S.; Cambillau, C. Structure of lactococcal phage p2 baseplate and its mechanism of activation. Proc. Natl. Acad. Sci. USA 2010, 107, 6852–6857. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, S.; Desmyter, A.; Verrips, C.T.; de Haard, H.J.; Moineau, S.; Cambillau, C. Lactococcal bacteriophage p2 receptor-binding protein structure suggests a common ancestor gene with bacterial and mammalian viruses. Nat. Struct. Mol. Biol. 2006, 13, 85–89. [Google Scholar] [CrossRef] [PubMed]
- Bebeacua, C.; Bron, P.; Lai, L.; Vegge, C.S.; Brondsted, L.; Spinelli, S.; Campanacci, V.; Veesler, D.; van Heel, M.; Cambillau, C. Structure and molecular assignment of lactococcal phage TP901-1 baseplate. J. Biol. Chem. 2010, 285, 39079–39086. [Google Scholar] [CrossRef] [PubMed]
- Veesler, D.; Spinelli, S.; Mahony, J.; Lichière, J.; Blangy, S.; Bricogne, G.; Legrand, P.; Ortiz-Lombardia, M.; Campanacci, V.; van Sinderen, D. Structure of the phage TP901-1 1.8 MDA baseplate suggests an alternative host adhesion mechanism. Proc. Natl. Acad. Sci. USA 2012, 109, 8954–8958. [Google Scholar] [CrossRef] [PubMed]
- Collins, B.; Bebeacua, C.; Mahony, J.; Blangy, S.; Douillard, F.P.; Veesler, D.; Cambillau, C.; van Sinderen, D. Structure and functional analysis of the host-recognition device of lactococcal phage Tuc2009. J. Virol. 2013, 87, 8429–8440. [Google Scholar] [CrossRef] [PubMed]
- Veesler, D.; Cambillau, C. A common evolutionary origin for tailed-bacteriophage functional modules and bacterial machineries. Microbiol. Mol. Biol. Rev. 2011, 75, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Bebeacua, C.; Lai, L.; Vegge, C.S.; Brøndsted, L.; van Heel, M.; Veesler, D.; Cambillau, C. Visualizing a complete siphoviridae by single-particle electron microscopy: The structure of lactococcal phage TP901-1. J. Virol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Campanacci, V.; Veesler, D.; Lichière, J.; Blangy, S.; Sciara, G.; Moineau, S.; Van Sinderen, D.; Bron, P.; Cambillau, C. Solution and electron microscopy characterization of lactococcal phage baseplates expressed in Escherichia coli. J. Struct. Biol. 2010, 172, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Shepherd, D.A.; Veesler, D.; Lichiere, J.; Ashcroft, A.E.; Cambillau, C. Unraveling lactococcal phages baseplate assembly by mass spectrometry. Mol. Cell. Proteom. 2011, 10. [Google Scholar] [CrossRef] [PubMed]
- Lhuillier, S.; Gallopin, M.; Gilquin, B.; Brasilès, S.; Lancelot, N.; Letellier, G.; Gilles, M.; Dethan, G.; Orlova, E.V.; Couprie, J. Structure of bacteriophage SPP1 head-to-tail connection reveals mechanism for viral DNA gating. Proc. Natl. Acad. Sci. USA 2009, 106, 8507–8512. [Google Scholar] [CrossRef] [PubMed]
- Tremblay, D.M.; Tegoni, M.; Spinelli, S.; Campanacci, V.; Blangy, S.; Huyghe, C.; Desmyter, A.; Labrie, S.; Moineau, S.; Cambillau, C. Receptor-binding protein of Lactococcus lactis phages: Identification and characterization of the saccharide receptor-binding site. J. Bacteriol. 2006, 188, 2400–2410. [Google Scholar] [CrossRef] [PubMed]
- Ricagno, S.; Campanacci, V.; Blangy, S.; Spinelli, S.; Tremblay, D.; Moineau, S.; Tegoni, M.; Cambillau, C. Crystal structure of the receptor-binding protein head domain from Lactococcus lactis phage bil170. J. Virol. 2006, 80, 9331–9335. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, S.; Veesler, D.; Bebeacua, C.; Cambillau, C. Structures and host-adhesion mechanisms of lactococcal siphophages. Front. Microbiol. 2014, 5, 3. [Google Scholar] [CrossRef] [PubMed]
- Mahony, J.; Kot, W.; Murphy, J.; Ainsworth, S.; Neve, H.; Hansen, L.H.; Heller, K.J.; Sørensen, S.J.; Hammer, K.; Cambillau, C. Investigation of the relationship between lactococcal host cell wall polysaccharide genotype and 936 phage receptor binding protein phylogeny. Appl. Environ. Microbiol. 2013, 79, 4385–4392. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.; Royer, B.; Mahony, J.; Hoyles, L.; Heller, K.; Neve, H.; Bonestroo, M.; Nauta, A.; van Sinderen, D. Biodiversity of lactococcal bacteriophages isolated from 3 gouda-type cheese-producing plants. J. Dairy Sci. 2013, 96, 4945–4957. [Google Scholar] [CrossRef] [PubMed]
- Marcó, M.B.; Garneau, J.E.; Tremblay, D.; Quiberoni, A.; Moineau, S. Characterization of two virulent phages of Lactobacillus plantarum. Appl. Environ. Microbiol. 2012. [Google Scholar] [CrossRef]
- Casey, E.; Mahony, J.; O’Connell-Motherway, M.; Bottacini, F.; Cornelissen, A.; Neve, H.; Heller, K.J.; Noben, J.-P.; Dal Bello, F.; van Sinderen, D. Molecular characterization of three Lactobacillus delbrueckii subsp. Bulgaricus phages. Appl. Environ. Microbiol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Ceyssens, P.-J.; Mesyanzhinov, V.; Sykilinda, N.; Briers, Y.; Roucourt, B.; Lavigne, R.; Robben, J.; Domashin, A.; Miroshnikov, K.; Volckaert, G. The genome and structural proteome of YUA, a new Pseudomonas aeruginosa phage resembling m6. J. Bacteriol. 2008, 190, 1429–1435. [Google Scholar] [CrossRef] [PubMed]
- Holtappels, M.; Vrancken, K.; Schoofs, H.; Deckers, T.; Remans, T.; Noben, J.-P.; Valcke, R. A comparative proteome analysis reveals flagellin, chemotaxis regulated proteins and amylovoran to be involved in virulence differences between Erwinia amylovora strains. J. Proteom. 2015, 123, 54–69. [Google Scholar] [CrossRef] [PubMed]
- Vegge, C.S.; Vogensen, F.K.; Mc Grath, S.; Neve, H.; van Sinderen, D.; Brøndsted, L. Identification of the lower baseplate protein as the antireceptor of the temperate lactococcal bacteriophages TP901-1 and Tuc2009. J. Bacteriol. 2006, 188, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Kuipers, O.P.; Beerthuyzen, M.M.; de Ruyter, P.G.; Luesink, E.J.; de Vos, W.M. Autoregulation of nisin biosynthesis in Lactococcus lactis by signal transduction. J. Biol. Chem. 1995, 270, 27299–27304. [Google Scholar] [CrossRef] [PubMed]
- Douillard, F.P.; O’Connell-Motherway, M.; Cambillau, C.; van Sinderen, D. Expanding the molecular toolbox for Lactococcus lactis: Construction of an inducible thioredoxin gene fusion expression system. Microb. Cell Fact. 2011, 10, 66. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Dümmler, A.; Lawrence, A.-M.; De Marco, A. Simplified screening for the detection of soluble fusion constructs expressed in E. coli using a modular set of vectors. Microb. Cell Fact. 2005, 4, 34. [Google Scholar] [CrossRef] [PubMed]
- Horton, R.M.; Cai, Z.; Ho, S.N.; Pease, L.R. Gene splicing by overlap extension: Tailor-made genes using the polymerase chain reaction. Biotechniques 1990, 8, 528–535. [Google Scholar] [CrossRef] [PubMed]
- Mc Grath, S.; Neve, H.; Seegers, J.F.; Eijlander, R.; Vegge, C.S.; Brondsted, L.; Heller, K.J.; Fitzgerald, G.F.; Vogensen, F.K.; van Sinderen, D. Anatomy of a lactococcal phage tail. J. Bacteriol. 2006, 188, 3972–3982. [Google Scholar] [CrossRef] [PubMed]
- Garvey, P.; Hill, C.; Fitzgerald, G. The lactococcal plasmid pnp40 encodes a third bacteriophage resistance mechanism, one which affects phage DNA penetration. Appl. Environ. Microbiol. 1996, 62, 676–679. [Google Scholar] [PubMed]
- Lillehaug, D. An improved plaque assay for poor plaque-producing temperate lactococcal bacteriophages. J. Appl. Microbiol. 1997, 83, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Dieterle, M.E.; Spinelli, S.; Sadovskaya, I.; Piuri, M.; Cambillau, C. Evolved distal tail carbohydrate binding modules of Lactobacillus phage J-1: A novel type of anti-receptor widespread among lactic acid bacteria phages. Mol. Microbiol. 2017, 104, 608–620. [Google Scholar] [CrossRef] [PubMed]
- Veesler, D.; Blangy, S.; Siponen, M.; Vincentelli, R.; Cambillau, C.; Sciara, G. Production and biophysical characterization of the cora transporter from Methanosarcina mazei. Anal. Biochem. 2009, 388, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988, 16, 10881–10890. [Google Scholar] [CrossRef] [PubMed]
- Van Sinderen, D.; Karsens, H.; Kok, J.; Terpstra, P.; Ruiters, M.H.; Venema, G.; Nauta, A. Sequence analysis and molecular characterization of the temperate lactococcal bacteriophage r1t. Mol. Microbiol. 1996, 19, 1343–1355. [Google Scholar] [CrossRef] [PubMed]
- Hayes, S.; Murphy, J.; Mahony, J.; Lugli, G.A.; Ventura, M.; Noben, J.-P.; Franz, C.M.; Neve, H.; Nauta, A.; Van Sinderen, D. Biocidal inactivation of Lactococcus lactis bacteriophages: Efficacy and targets of commonly used sanitizers. Front. Microbiol. 2017, 8, 107. [Google Scholar] [CrossRef] [PubMed]
- Goulet, A.; Lai-Kee-Him, J.; Veesler, D.; Auzat, I.; Robin, G.; Shepherd, D.A.; Ashcroft, A.E.; Richard, E.; Lichière, J.; Tavares, P. The opening of the SPP1 bacteriophage tail, a prevalent mechanism in gram-positive-infecting siphophages. J. Biol. Chem. 2011, 286, 25397–25405. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, I.; Osada, K.; Azam, A.H.; Asakawa, H.; Miyanaga, K.; Tanji, Y. The presence of two receptor-binding proteins contributes to the wide host range of staphylococcal Twort-like phages. Appl. Environ. Microbiol. 2016, 82, 5763–5774. [Google Scholar] [CrossRef] [PubMed]
- Bebeacua, C.; Tremblay, D.; Farenc, C.; Chapot-Chartier, M.-P.; Sadovskaya, I.; Van Heel, M.; Veesler, D.; Moineau, S.; Cambillau, C. Structure, adsorption to host, and infection mechanism of virulent lactococcal phage p2. J. Virol. 2013, 87, 12302–12312. [Google Scholar] [CrossRef] [PubMed]
- Dowah, A.S.; Clokie, M.R. Review of the nature, diversity and structure of bacteriophage receptor binding proteins that target gram-positive bacteria. Biophys. Rev. 2018, 10, 535–542. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hayes, S.; Duhoo, Y.; Neve, H.; Murphy, J.; Noben, J.-P.; Franz, C.M.A.P.; Cambillau, C.; Mahony, J.; Nauta, A.; Van Sinderen, D. Identification of Dual Receptor Binding Protein Systems in Lactococcal 936 Group Phages. Viruses 2018, 10, 668. https://doi.org/10.3390/v10120668
Hayes S, Duhoo Y, Neve H, Murphy J, Noben J-P, Franz CMAP, Cambillau C, Mahony J, Nauta A, Van Sinderen D. Identification of Dual Receptor Binding Protein Systems in Lactococcal 936 Group Phages. Viruses. 2018; 10(12):668. https://doi.org/10.3390/v10120668
Chicago/Turabian StyleHayes, Stephen, Yoan Duhoo, Horst Neve, James Murphy, Jean-Paul Noben, Charles M. A. P. Franz, Christian Cambillau, Jennifer Mahony, Arjen Nauta, and Douwe Van Sinderen. 2018. "Identification of Dual Receptor Binding Protein Systems in Lactococcal 936 Group Phages" Viruses 10, no. 12: 668. https://doi.org/10.3390/v10120668
APA StyleHayes, S., Duhoo, Y., Neve, H., Murphy, J., Noben, J.-P., Franz, C. M. A. P., Cambillau, C., Mahony, J., Nauta, A., & Van Sinderen, D. (2018). Identification of Dual Receptor Binding Protein Systems in Lactococcal 936 Group Phages. Viruses, 10(12), 668. https://doi.org/10.3390/v10120668





