Roles of orf60a and orf61 in Development of Bacteriophages λ and Φ24B
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Bacteriophages and Plasmids
2.2. Prophage Induction Experiments
2.3. One-Step Growth Experiment
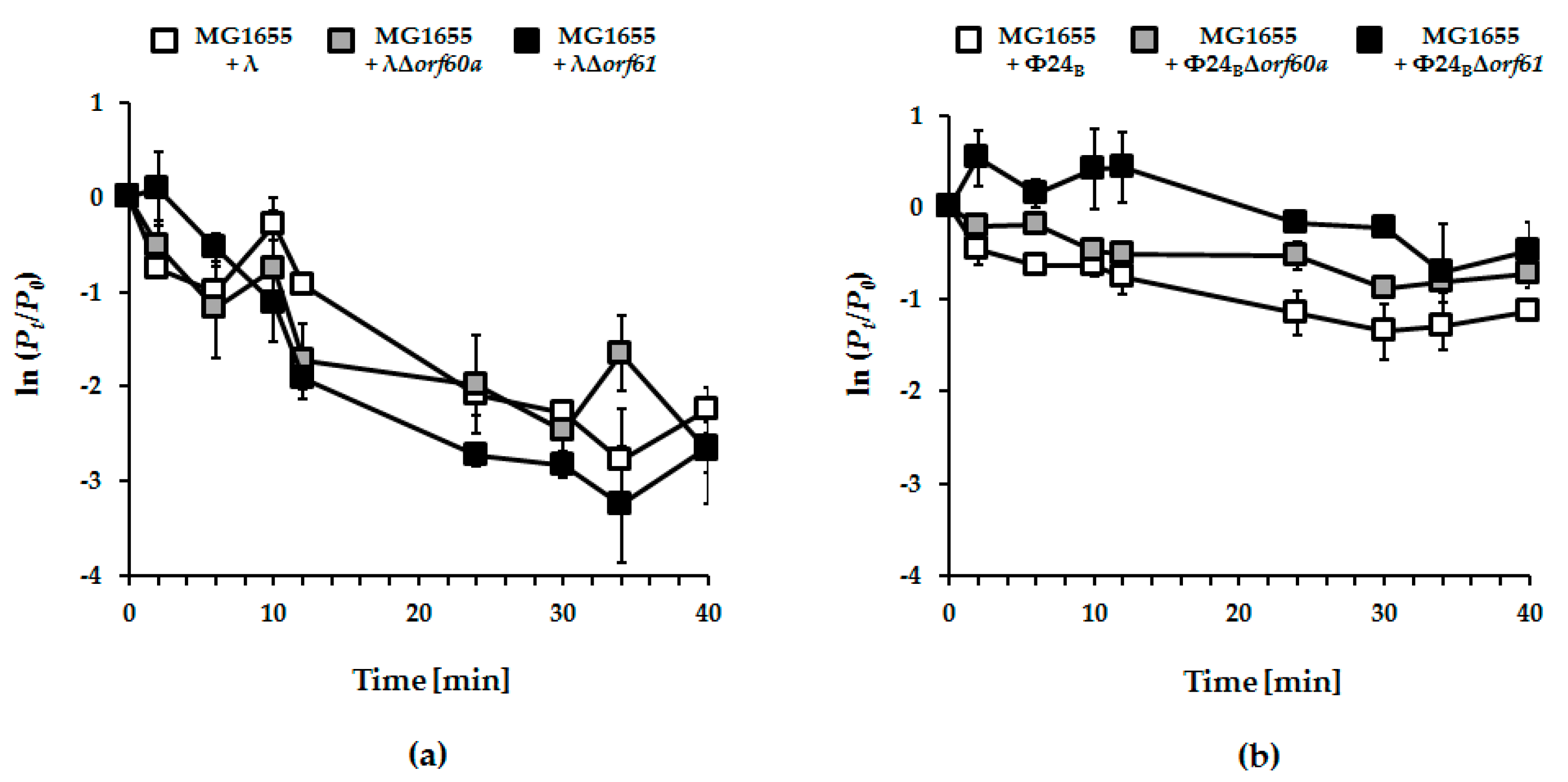
2.4. Measurement of the Efficiency of Phage Adsorption
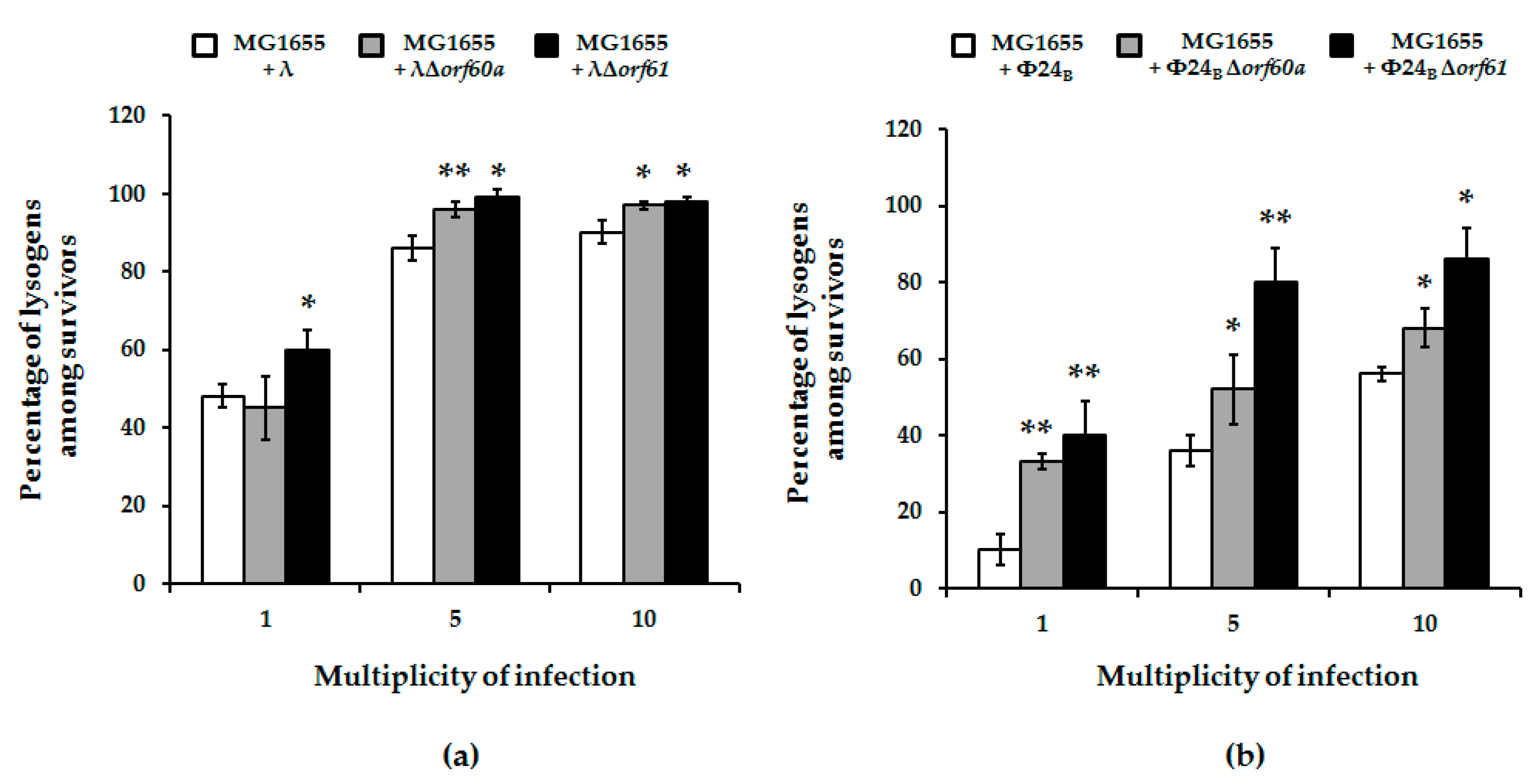
2.5. Efficiency of Lysogenization
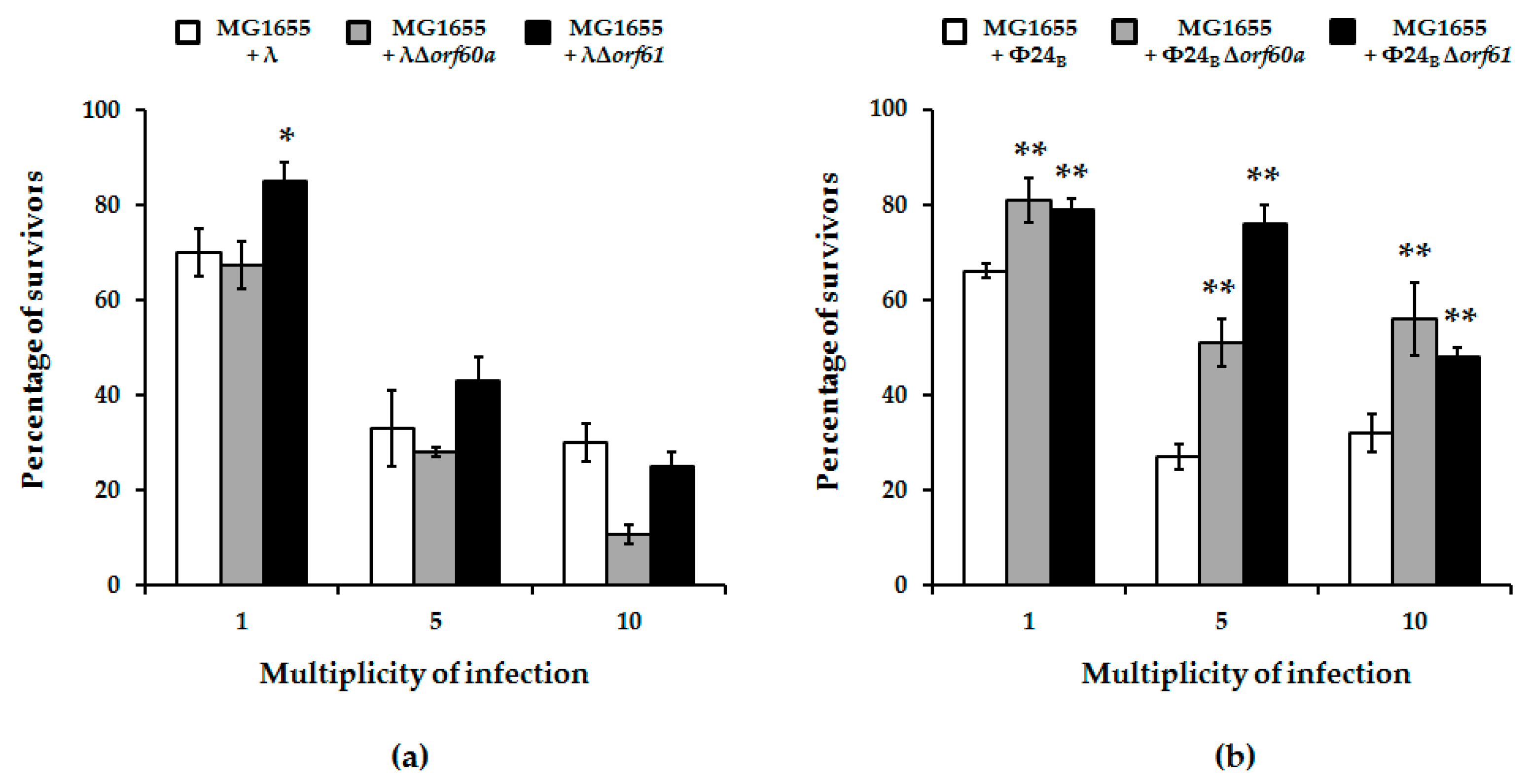
2.6. Survival of Cells after Bacteriophage Infection
2.7. Measurement of Bacterial Viability during Prophage Induction Experiments
3. Results
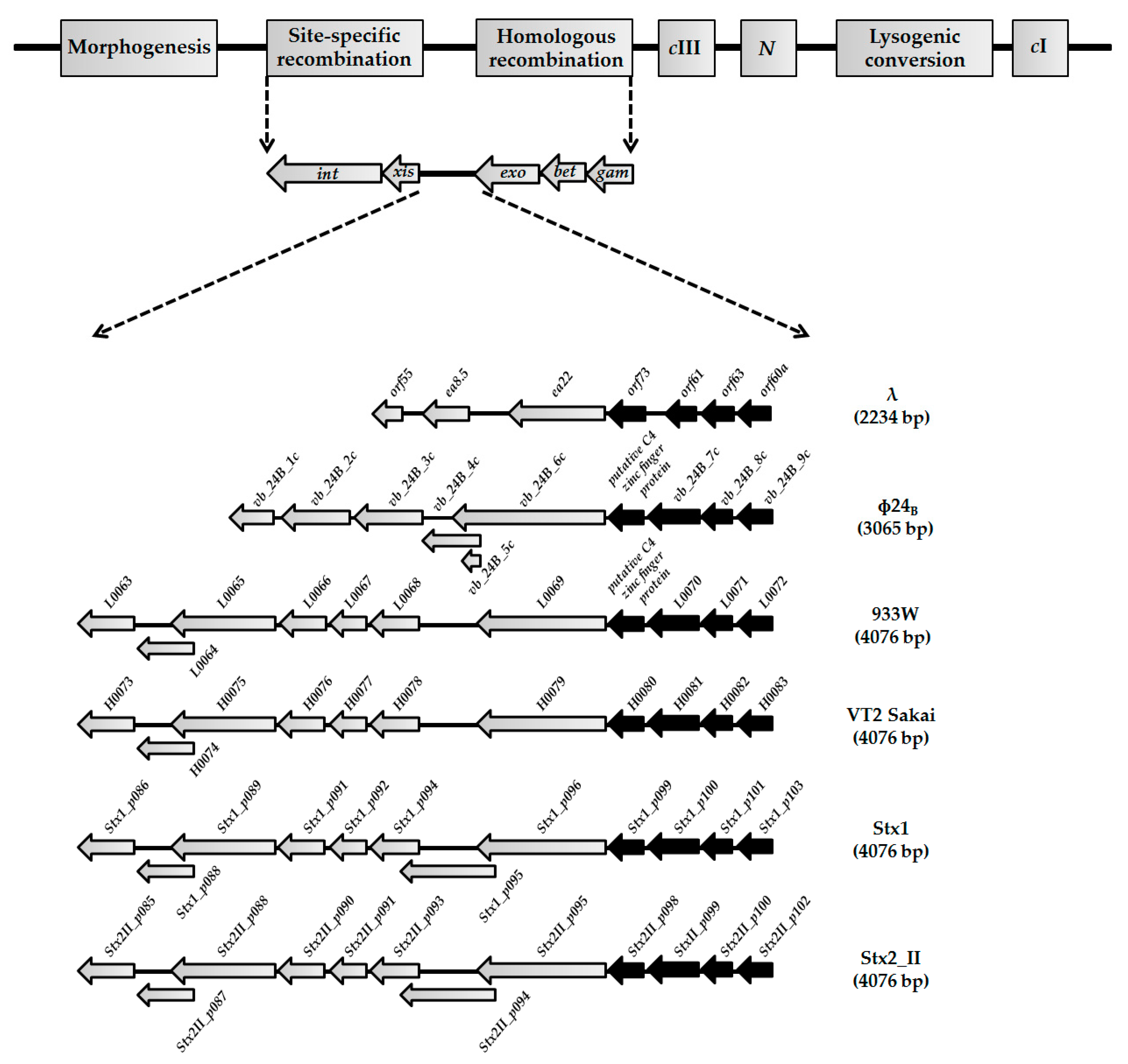
3.1. Sequences of orf60a and orf61, and Their Putative Products, Are Conservative among Lambdoid Phages
3.2. Influence of orf60a and orf61 on Prophage Induction with Various Inductors
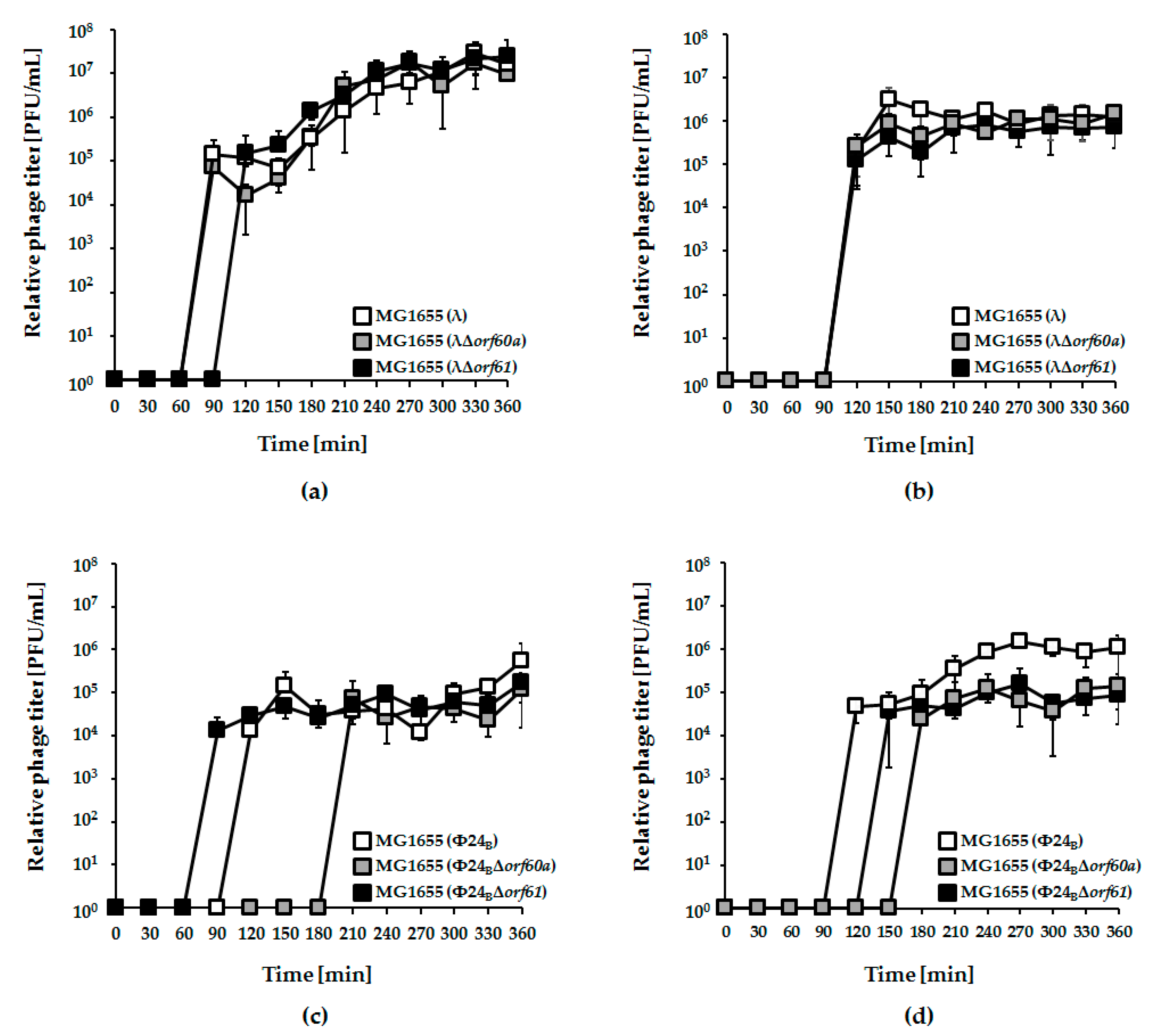
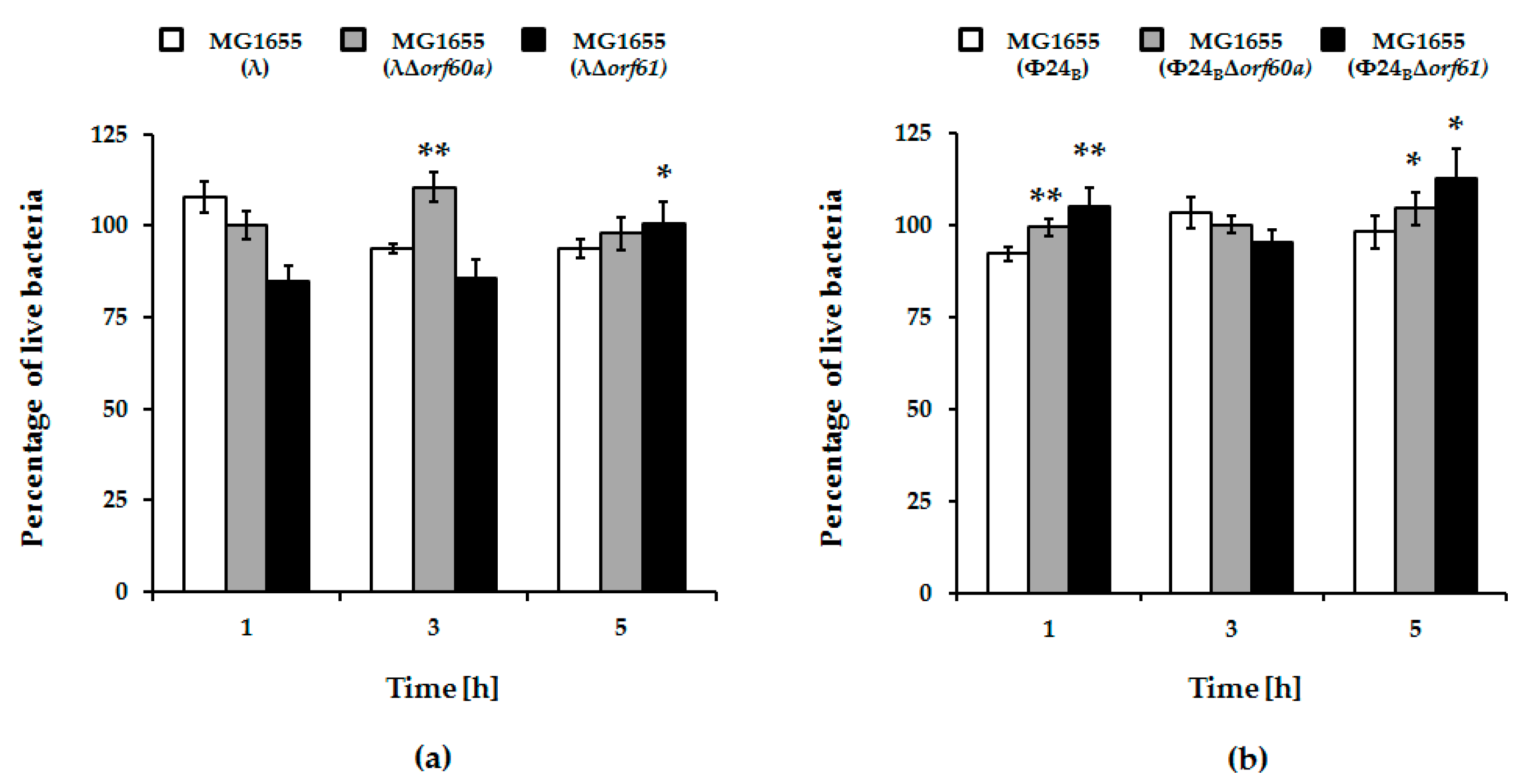
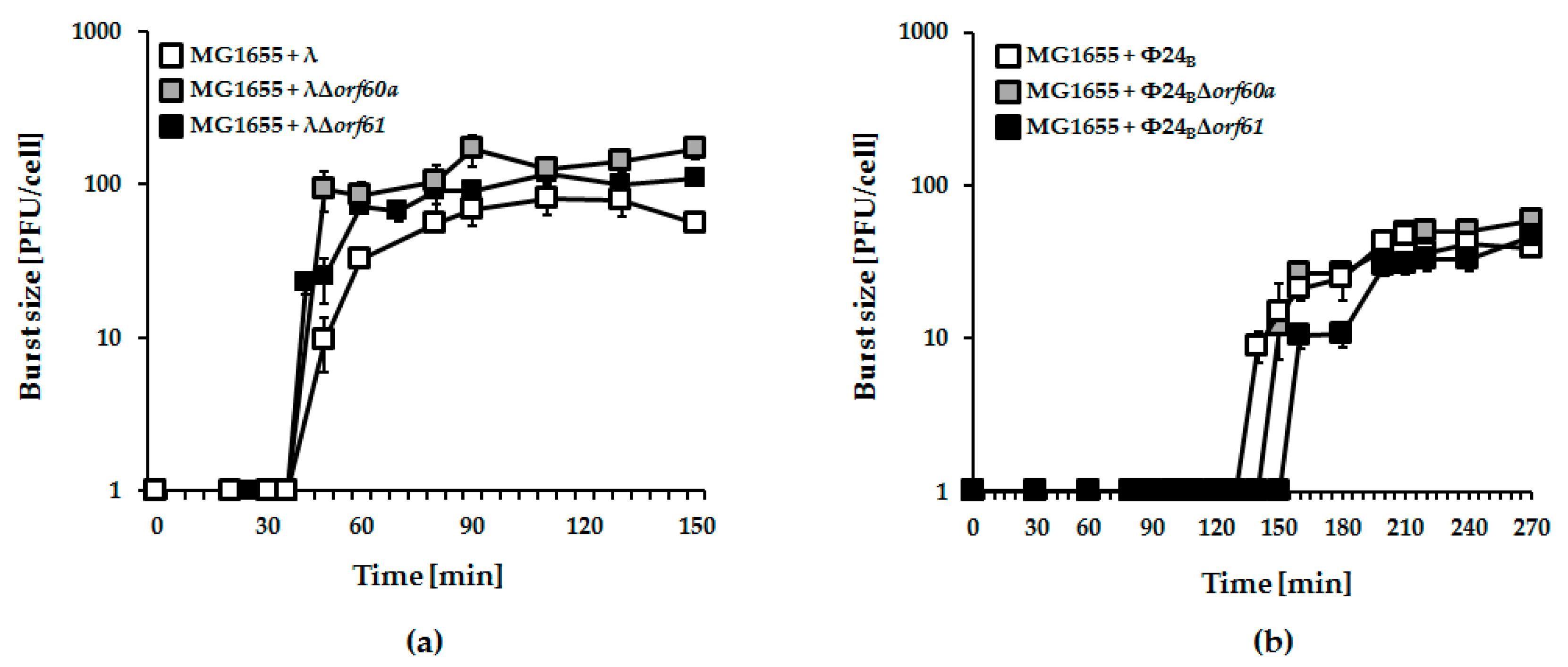
3.3. Effects of orf60a and orf61 Deletions on Phage Infection
3.4. Adsorption of Phage Φ24B is Impaired in the Absence of orf60a or orf61
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Ptashne, M. A Genetic Switch: Phage Lambda Revisited, 3rd ed.; Cold Spring Harbor Laboratory Press: Laurel Hollow, NY, USA, 2004; ISBN 0 87969 716 4. [Google Scholar]
- Wegrzyn, G.; Wegrzyn, A. Genetic switches during bacteriophage lambda development. Prog. Nucleic Acid Res. Mol. Biol. 2005, 79, 1–48. [Google Scholar] [PubMed]
- Wegrzyn, G.; Licznerska, K.; Wegrzyn, A. Phage λ—New insights into regulatory circuits. Adv. Virus Res. 2012, 82, 155–178. [Google Scholar] [PubMed]
- Mauro, S.A.; Koudelka, G.B. Shiga toxin: Expression, distribution, and its role in the environment. Toxins 2011, 3, 608–625. [Google Scholar] [CrossRef] [PubMed]
- Hunt, J.M. Shiga toxin-producing Escherichia coli (STEC). Clin. Lab. Med. 2010, 30, 21–45. [Google Scholar] [CrossRef] [PubMed]
- Gyles, C.L. Shiga toxin-producing Escherichia coli: An overview. J. Anim. Sci. 2007, 85, E45–E62. [Google Scholar] [CrossRef] [PubMed]
- Sergueev, K.; Court, D.; Reaves, L.; Austin, S. E. coli cell-cycle regulation by bacteriophage lambda. J. Mol Biol. 2002, 324, 297–307. [Google Scholar] [CrossRef]
- Bloch, S.; Nejman-Falenczyk, B.; Los, J.M.; Baranska, S.; Lepek, K.; Felczykowsk, A.; Los, M.; Wegrzyn, G.; Wegrzyn, A. Genes from the exo-xis region of λ and Shiga toxin-converting bacteriophages influence lysogenization and prophage induction. Arch. Microbiol. 2013, 195, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Los, J.M.; Los, M.; Wegrzyn, A.; Wegrzyn, G. Role of the bacteriophage lambda exo-xis region in the virus development. Folia Microbiol. 2008, 53, 443–450. [Google Scholar] [CrossRef] [PubMed]
- Bloch, S.; Nejman-Falenczyk, B.; Dydecka, A.; Łoś, J.M.; Felczykowska, A.; Wegrzyn, A.; Wegrzyn, G. Different expression patterns of genes from the exo-xis region of bacteriophage λ and Shiga toxin-converting bacteriophage Φ24B following infection or prophage induction in Escherichia coli. PLoS ONE 2014, 9, e108233. [Google Scholar] [CrossRef] [PubMed]
- Kwan, J.J.; Smirnova, E.; Khazai, S.; Evanics, F.; Maxwell, K.L.; Donaldson, L.W. The solution structures of two prophage homologues of the bacteriophage λ Ea8.5 protein reveal a newly discovered hybrid homeodomain/zinc-finger fold. Biochemistry 2013, 52, 3612–3614. [Google Scholar] [CrossRef] [PubMed]
- Licznerska, K.; Dydecka, A.; Bloch, S.; Topka, G.; Nejman-Falenczyk, B.; Wegrzyn, A.; Wegrzyn, G. The role of the exo-xis region in oxidative stress-mediated induction of Shiga toxin-converting prophages. Oxid. Med. Cell. Longev. 2016, 2016, 8453135. [Google Scholar] [CrossRef] [PubMed]
- Dydecka, A.; Bloch, S.; Rizvi, A.; Perez, S.; Nejman-Falenczyk, B.; Topka, G.; Gasior, T.; Necel, A.; Wegrzyn, G.; Donaldson, L.W.; et al. Bad phages in good bacteria: Role of the mysterious orf63 of λ and Shiga toxin-converting Φ24B bacteriophages. Front. Microbiol. 2017, 8, 1618. [Google Scholar] [CrossRef] [PubMed]
- Nejman-Falenczyk, B.; Bloch, S.; Licznerska, K.; Dydecka, A.; Felczykowska, A.; Topka, G.; Wegrzyn, A.; Wegrzyn, G. A small, microRNA-size, ribonucleic acid regulating gene expression and development of Shiga toxin-converting bacteriophage Φ24B. Sci. Rep. 2015, 5, 10080. [Google Scholar] [CrossRef] [PubMed]
- Jensen, K.F. The Escherichia coli K-12 wild types W3110 and MG1655 have an rph frameshift mutation that leads to pyrimidine starvation due to low pyrE expression levels. J. Bacteriol. 1993, 175, 3401–3407. [Google Scholar] [CrossRef] [PubMed]
- Hendrix, R.W.; Duda, R.L. Bacteriophage lambda PaPa: Not a mother of all lambda phages. Science 1993, 258, 1145–1148. [Google Scholar] [CrossRef]
- Allison, H.E.; Sergeant, M.J.; James, C.E.; Saunders, J.R.; Smith, D.L.; Sharp, R.J.; Marks, T.S.; McCarthy, A.J. Immunity profiles of wild-type and recombinant shiga-like toxin-encoding bacteriophages and characterization of novel double lysogens. Infect. Immun. 2003, 71, 3409–3418. [Google Scholar] [CrossRef] [PubMed]
- Loś, J.M.; Golec, P.; Wegrzyn, G.; Wegrzyn, A.; Loś, M. Simple method for plating Escherichia coli bacteriophages forming very small plaques or no plaques under standard conditions. Appl. Environ. Microbiol. 2008, 74, 5113–5120. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Wang, I.N. Bacteriophage adsorption rate and optimal lysis time. Genetics. 2008, 180, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Wegrzyn, G.; Glass, R.E.; Thomas, M.S. Involvement of the Escherichia coli RNA polymerase α subunit in transcriptional activation by the bacteriophage λ CI and CII proteins. Gene 1992, 122, 1–7. [Google Scholar] [CrossRef]
- Schmidt, H. Shiga-toxin-converting bacteriophages. Res. Microbiol. 2001, 152, 687–695. [Google Scholar] [CrossRef]
- Rajagopala, S.V.; Casjens, S.; Uetz, P. The protein interaction map of bacteriophage lambda. BMC Microbiol. 2011, 11, 213. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Jiang, H.; Gu, Z.; Roberts, J.W. High-resolution view of bacteriophage lambda gene expression by ribosome profiling. Proc. Natl. Acad. Sci. USA 2013, 110, 11928–11933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bloch, S.; Nejman-Falenczyk, B.; Topka, G.; Dydecka, A.; Licznerska, K.; Narajczyk, M.; Necel, A.; Wegrzyn, A.; Wegrzyn, G. UV-Sensitivity of Shiga Toxin-Converting Bacteriophage Virions Φ24B, 933W, P22, P27 and P32. Toxins 2015, 7, 3727–3739. [Google Scholar] [CrossRef] [PubMed]
| Bacterial Strains or Bacteriophages | Relevant Genotype or Description | References |
|---|---|---|
| Escherichia coli Strains | ||
| MG1655 | F– λ– ilvG rfb-50 rph-1 | [15] |
| MG1655 (λ) | MG1655 bearing λ prophage | [8] |
| MG1655 (λΔorf60a) | MG1655 bearing λ prophage with deletion of orf60a | [12] |
| MG1655 (λΔorf61) | MG1655 bearing λ prophage with deletion of orf61 | [12] |
| MG1655 (Φ24B) | MG1655 bearing Φ24B prophage | [8] |
| MG1655 (Φ24BΔorf60a) | MG1655 bearing Φ24B prophage with deletion of vb_24B_9c, the homolog of λorf60a | [12] |
| MG1655 (Φ24BΔorf61) | MG1655 bearing Φ24B prophage with deletion of vb_24B_7c, the homolog of λorf61 | [12] |
| Bacteriophages | ||
| λ | carries a frameshift mutation relative to Ur-lambda | [16] |
| λΔorf60a | λ phage with deletion of orf60a | [12] |
| λΔorf61 | λ phage with deletion of orf61 | [12] |
| Φ24B | Φ24Bstx2::cat | [17] |
| Φ24BΔorf60a | Φ24B phage with deletion of vb_24B_9c, the homolog of Φ24B orf60a | [12] |
| Φ24BΔorf61 | Φ24B phage with deletion of vb_24B_7c, the homolog of Φ24B orf61 | [12] |
| λ | Φ24B | 933W | VT2 Sakai | Stx1 | Stx2_II | |
|---|---|---|---|---|---|---|
| λ | 96 | 96 | 94 | 94 | 94 | |
| Φ24B | 100 | 94 | 94 | 94 | ||
| 933W | 94 | 94 | 94 | |||
| VT2 Sakai | 100 | 100 | ||||
| Stx1 | 100 | |||||
| Stx2_II |
| λ | Φ24B | 933W | VT2 Sakai | Stx1 | Stx2_II | |
|---|---|---|---|---|---|---|
| λ | 95 | 95 | 97 | 97 | 97 | |
| Φ24B | 100 | 94 | 94 | 94 | ||
| 933W | 94 | 94 | 94 | |||
| VT2 Sakai | 100 | 100 | ||||
| Stx1 | 100 | |||||
| Stx2_II |
| λ | Φ24B | 933W | VT2 Sakai | Stx1 | Stx2_II | |
|---|---|---|---|---|---|---|
| λ | 80 | 80 | 80 | 80 | 80 | |
| Φ24B | 100 | 100 | 100 | 100 | ||
| 933W | 100 | 100 | 100 | |||
| VT2 Sakai | 100 | 100 | ||||
| Stx1 | 100 | |||||
| Stx2_II |
| λ | Φ24B | 933W | VT2 Sakai | Stx1 | Stx2_II | |
|---|---|---|---|---|---|---|
| λ | 70 | 70 | 70 | 70 | 70 | |
| Φ24B | 100 | 100 | 100 | 100 | ||
| 933W | 100 | 100 | 100 | |||
| VT2 Sakai | 100 | 100 | ||||
| Stx1 | 100 | |||||
| Stx2_II |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dydecka, A.; Nejman-Faleńczyk, B.; Bloch, S.; Topka, G.; Necel, A.; Donaldson, L.W.; Węgrzyn, G.; Węgrzyn, A. Roles of orf60a and orf61 in Development of Bacteriophages λ and Φ24B. Viruses 2018, 10, 553. https://doi.org/10.3390/v10100553
Dydecka A, Nejman-Faleńczyk B, Bloch S, Topka G, Necel A, Donaldson LW, Węgrzyn G, Węgrzyn A. Roles of orf60a and orf61 in Development of Bacteriophages λ and Φ24B. Viruses. 2018; 10(10):553. https://doi.org/10.3390/v10100553
Chicago/Turabian StyleDydecka, Aleksandra, Bożena Nejman-Faleńczyk, Sylwia Bloch, Gracja Topka, Agnieszka Necel, Logan W. Donaldson, Grzegorz Węgrzyn, and Alicja Węgrzyn. 2018. "Roles of orf60a and orf61 in Development of Bacteriophages λ and Φ24B" Viruses 10, no. 10: 553. https://doi.org/10.3390/v10100553







