Urbanisation Shapes the Diversity, Composition, and Functional Profile of Endophytic Bacteriome in Common Urban Tree Species
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area and Sampling Plots
2.2. Leaf Sampling and DNA Extraction
2.3. Amplicon Sequencing and Bioinformatics
2.4. Alpha and Beta Diversity Analyses
2.5. Differential Abundance and Biomarker Analysis
2.6. Bioinformatic Processing
2.7. Functional Profiling
2.8. Statistical Evaluation
3. Results
3.1. Taxonomic Diversity
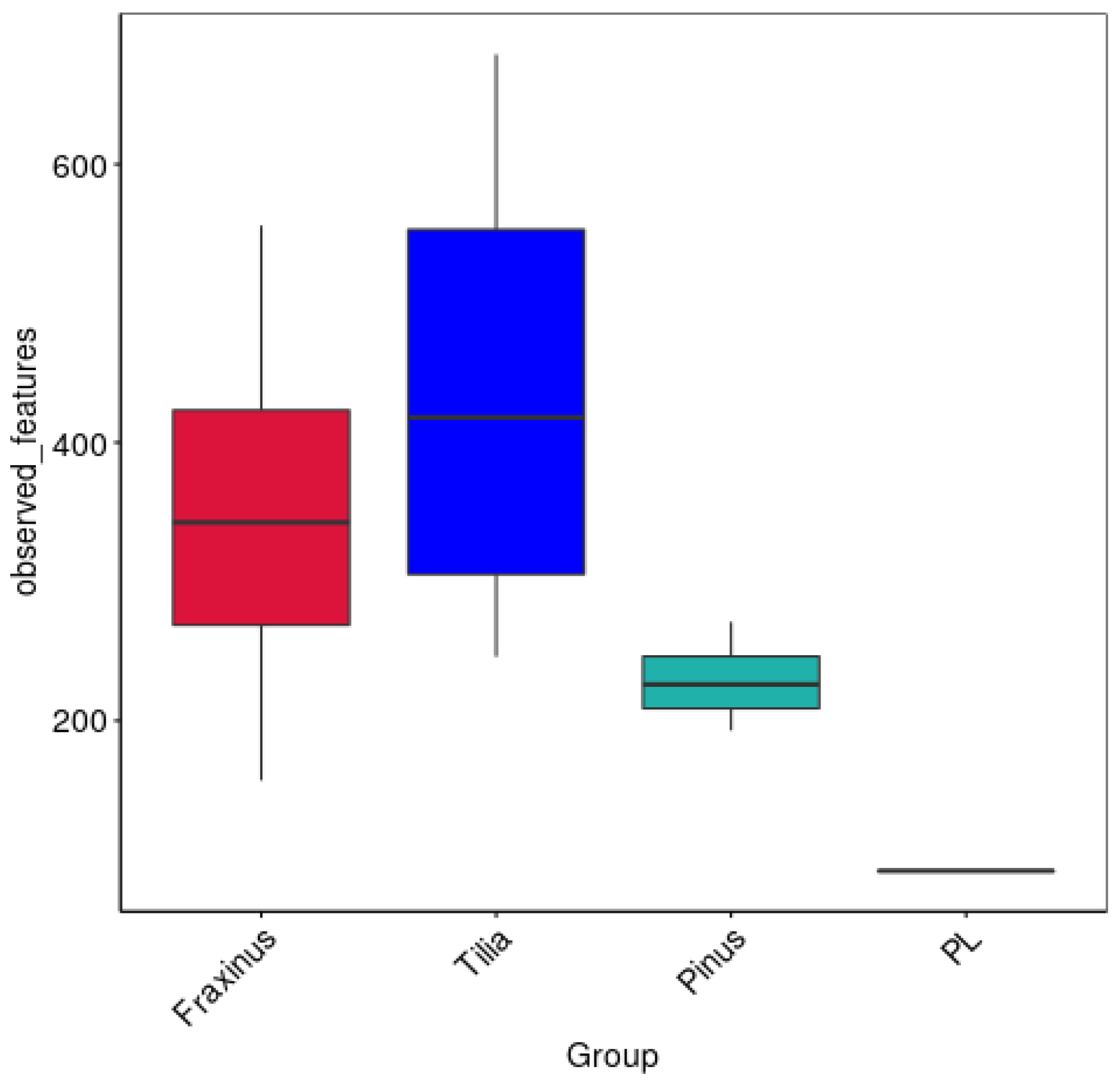
3.1.1. Differences in Endophytic Richness Among Tree Species
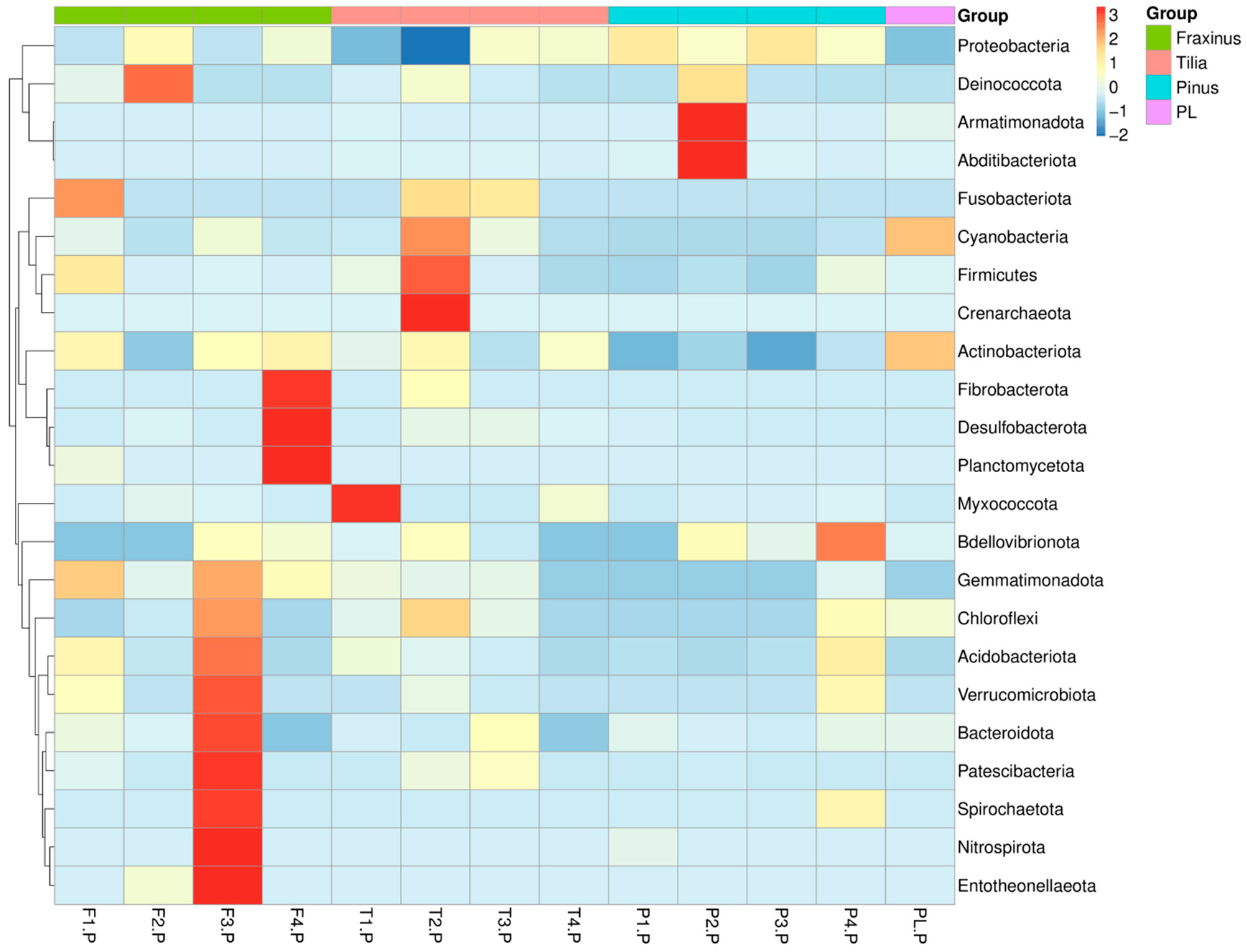
3.1.2. Phylum Level—Endophytic Bacterial Community Composition
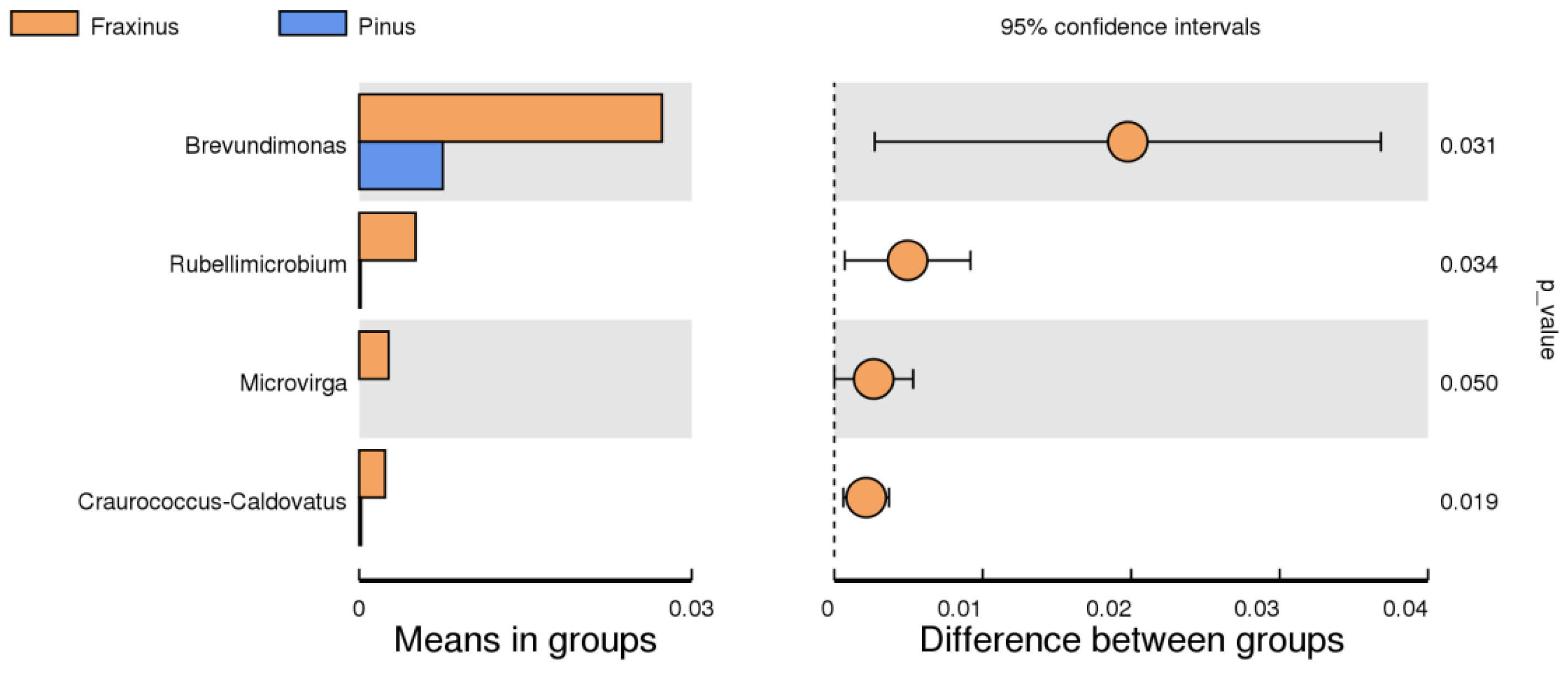
3.1.3. Genus Level—Endophytic Community Composition
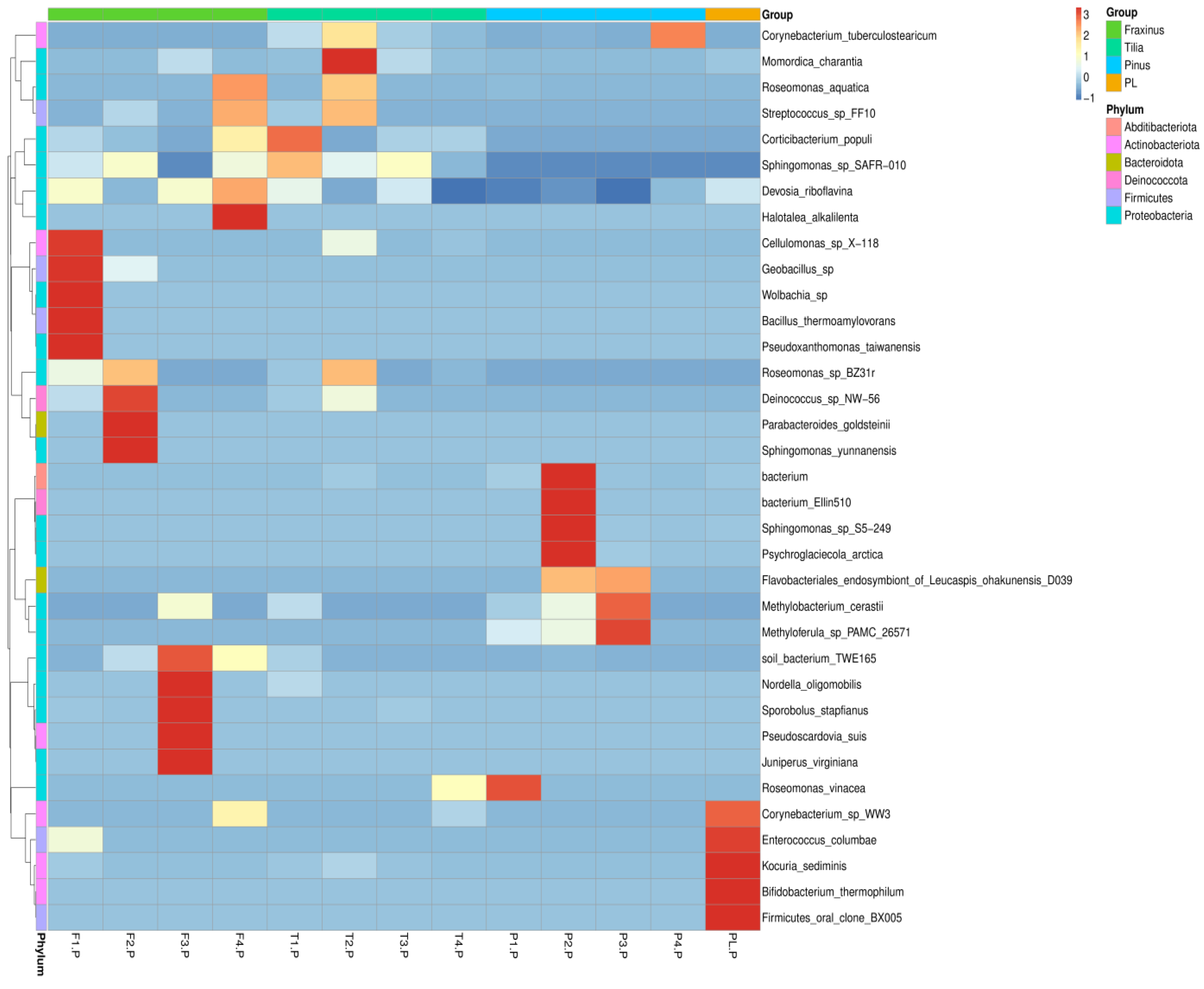
3.1.4. Species-Level Endophytic Community Composition
3.1.5. Host-Driven Structuring of the Endophytic Microbiome
3.2. Alpha and Beta Diversity of Endophytic Microbiome
3.2.1. Alpha Diversity Indices
3.2.2. Comparison of Shannon Diversity Index Among Tree Species Groups and Pooled Sample
3.2.3. Beta Diversity
3.2.4. NMDS Analysis of Endophytic Bacterial Microbiome Associated with the Studied Urban Tree Species and a Pooled Sample PL
3.3. Functional Prediction
4. Discussion
4.1. Endophytic Bacteriome Diversity Is Shaped by Host Species and Urban Site Conditions
4.2. Distinct Taxonomic Signatures Among Tree Species
4.3. Bacterial Community Variation Driven by Host Identity
4.4. Beta Diversity and NMDS Confirm Tree-Specific Community Structures
4.5. Urban Pollution Modulates Endophytic Bacteriome Composition
4.6. Functional Implications of Predicted Microbial Metabolism
4.7. Ecological and Practical Implications
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Grimm, N.B.; Faeth, S.H.; Golubiewski, N.E.; Redman, C.L.; Wu, J.; Bai, X.; Briggs, J.M. Global change and the ecology of cities. Science 2008, 319, 756–760. [Google Scholar] [CrossRef]
- McKinney, M.L. Effects of urbanization on species richness: A review of plants and animals. Urban Ecosyst. 2008, 11, 161–176. [Google Scholar] [CrossRef]
- Nowak, D.J.; Hirabayashi, S.; Bodine, A.; Greenfield, E. Tree and forest effects on air quality and human health in the United States. Environ. Pollut. 2014, 193, 119–129. [Google Scholar] [CrossRef]
- Sjöman, H.; Nielsen, A.B. Selecting trees for urban paved sites in Scandinavia—A review of information on stress tolerance and its relation to the requirements of tree planners. Urban For. Urban Green. 2010, 9, 281–293. [Google Scholar] [CrossRef]
- Calfapietra, C.; Fares, S.; Manes, F.; Morani, A.; Sgrigna, G.; Loreto, F. Role of Biogenic Volatile Organic Compounds (BVOCs) emitted by urban trees on ozone concentration in cities: A review. Environ. Pollut. 2013, 183, 71–80. [Google Scholar] [CrossRef] [PubMed]
- Compant, S.; Samad, A.; Faist, H.; Sessitsch, A. A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J. Adv. Res. 2019, 19, 29–37. [Google Scholar] [CrossRef]
- Berg, G.; Rybakova, D.; Fischer, D.; Cernava, T.; Vergès, M.-C.; Charles, T.; Chen, X.; Cocolin, L.; Eversole, K.; Corral, G.H.; et al. Microbiome definition re-visited: Old concepts and new challenges. Microbiome 2020, 8, 103. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Brettell, L.E.; Singh, B.K. Linking the phyllosphere microbiome to plant health. Trends Plant Sci. 2020, 25, 841–844. [Google Scholar] [CrossRef]
- Faticov, M.; Amorim, J.H.; Abdelfattah, A.; van Dijk, L.J.A.; Carvalho, A.C.; Laforest-Lapointe, I.; Tack, A.J.M. Local climate, air quality and leaf litter cover shape foliar fungal communities on an urban tree. AMBIO 2024, 53, 1673. [Google Scholar] [CrossRef]
- Espenshade, J.; Thijs, S.; Gawronski, S.W.; Bové, H.; Weyens, N.; Vangronsveld, J. Influence of Urbanization on Epiphytic Bacterial Communities of the Platanus × hispanica Tree Leaves in a Biennial Study. Front. Microbiol. 2019, 10, 675. [Google Scholar] [CrossRef]
- Gandolfi, I.; Canedoli, C.; Rosatelli, A.; Covino, S.; Cappelletti, D.; Sebastiani, B.; Tatangelo, V.; Corengia, D.; Pittino, F.; Padoa-Schioppa, E.; et al. Microbiomes of urban trees: Unveiling contributions to atmospheric pollution mitigation. Front. Microbiol. 2024, 15, 1470376. [Google Scholar] [CrossRef]
- Laforest-Lapointe, I.; Paquette, A.; Messier, C.; Kembel, S.W. Leaf bacterial diversity mediates plant diversity and ecosystem function relationships. Nature 2017, 546, 145–147. [Google Scholar] [CrossRef]
- Reinhold-Hurek, B.; Bünger, W.; Burbano, C.S.; Sabale, M.; Hurek, T. Roots shaping their microbiome: Global hotspots for microbial activity. Annu. Rev. Phytopathol. 2015, 53, 403–424. [Google Scholar] [CrossRef] [PubMed]
- Altaye, G.; Agete, A.; Talebi, E. Seasonal dynamics of endophytic fungal diversity Grevillea robusta (Silver oak) trees. Biotechnol. J. Environ. Microorg. (BJEM) 2023, 2, 223–232. [Google Scholar] [CrossRef]
- Cregger, M.A.; Veach, A.M.; Yang, Z.K.; Crouch, M.J.; Vilgalys, R.; Tuskan, G.A.; Schadt, C.W. The Populus holobiont: Dissecting the effects of plant niches and genotype on the microbiome. Microbiome 2018, 6, 31. [Google Scholar] [CrossRef]
- Xu, L.; Naylor, D.; Dong, Z.; Simmons, T.; Pierroz, G.; Hixson, K.K.; Kim, Y.M.; Zink, E.M.; Engbrecht, K.M.; Wang, Y.; et al. Drought delays development of the sorghum root microbiome and enriches for monoderm bacteria. Proc. Natl. Acad. Sci. USA 2018, 115, E4284–E4293. [Google Scholar] [CrossRef]
- Petrova, S. The added value of urban trees (Tilia tomentosa Moench, Fraxinus excelsior L. and Pinus nigra J.F. Arnold) in terms of air pollutants removal. Forests 2024, 15, 1034. [Google Scholar] [CrossRef]
- Petrova, S.; Velcheva, I.; Nikolov, B.; Vasileva, T.; Bivolarski, V. Antioxidant Responses and Adaptation Mechanisms of Tilia tomentosa Moench, Fraxinus excelsior L. and Pinus nigra J. F. Arnold towards Urban Air Pollution. Forests 2022, 13, 1689. [Google Scholar] [CrossRef]
- Petrova, S.; Petkova, M. Plant Traits of Tilia tomentosa Moench, Fraxinus excelsior L., and Pinus nigra J.F. Arnold as a Proxy of Urbanization. Forests 2023, 14, 800. [Google Scholar] [CrossRef]
- Hallmann, J.; Quadt-Hallmann, A.; Mahaffee, W.F.; Kloepper, J.W. Bacterial endophytes in agricultural crops. Can. J. Microbiol. 1997, 43, 895–914. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Simpson, G.; Blanchet, F.; Kindt, R.; Legendre, P.; Minchin, P.; O’Hara, R.; Solymos, P.; Stevens, M.; Szoecs, E.; et al. Vegan: Community Ecology Package. R Package Version 2.6-4, 2022. Available online: https://CRAN.R-project.org/package=vegan (accessed on 15 December 2025).
- Paulson, J.N.; Stine, O.C.; Bravo, H.C.; Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 2013, 10, 1200–1202. [Google Scholar] [CrossRef]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
- Louca, S.; Parfrey, L.W.; Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science 2016, 353, 1272–1277. [Google Scholar] [CrossRef] [PubMed]
- Dahal, R.H.; Kim, J. Brevundimonas humi sp. nov., an alphaproteobacterium isolated from forest soil. Int. J. Syst. Evol. Microbiol. 2018, 68, 709–714. [Google Scholar] [CrossRef]
- Madhaiyan, M.; Poonguzhali, S.; Kwon, S.-W.; Sa, T.M. Bacillus methylotrophicus sp. nov., a methanol-utilizing, plant-growth-promoting bacterium isolated from rice rhizosphere soil. Int. J. Syst. Evol. Microbiol. 2010, 60, 2490–2495. [Google Scholar] [CrossRef]
- Baldrian, P. Forest microbiome: Diversity, complexity and dynamics. FEMS Microbiol. Rev. 2017, 41, 109–130. [Google Scholar] [CrossRef]
- Sazonova, O.I.; Ivanova, A.A.; Vetrova, A.A.; Zvonarev, A.N.; Streletskii, R.A.; Vasenev, V.I.; Myazin, V.A.; Makhinya, K.I.; Kozlova, E.V.; Korneykova, M.V. Impact of Anthropogenic Factors on the Diversity of Microbial Communities of PM10 Air and PM100 of Tilia L. Phylloplane in an Urban Ecosystem. Biology 2024, 13, 969. [Google Scholar] [CrossRef]
- Mendes, R.; Kruijt, M.; de Bruijn, I.; Dekkers, E.; van der Voort, M.; Schneider, J.H.; Piceno, Y.M.; DeSantis, T.Z.; Andersen, G.L.; Bakker, P.A.; et al. Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 2011, 332, 1097–1100. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; van Themaat, E.V.L.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef]
- Flores-Duarte, N.J.; Pérez-Pérez, J.; Navarro-Torre, S.; Mateos-Naranjo, E.; Redondo-Gómez, S.; Pajuelo, E.; Rodríguez-Llorente, I.D. Improved Medicago sativa Nodulation under Stress Assisted by Variovorax sp. Endophytes. Plants 2022, 11, 1091. [Google Scholar] [CrossRef]
- McPherson, M.R.; Wang, P.; Marsh, E.L.; Mitchell, R.B.; Schachtman, D.P. Isolation and analysis of microbial communities in soil, rhizosphere, and roots in perennial grass experiments. J. Vis. Exp. 2018, 137, 57932. [Google Scholar] [CrossRef]
- Xu, N.; Zhao, Q.; Zhang, Z.; Zhang, Q.; Wang, Y.; Qin, G.; Ke, M.; Qiu, D.; Peijnenburg, W.J.G.M.; Lu, T.; et al. Phyllosphere Microorganisms: Sources, Drivers, and Their Interactions with Plant Hosts. J. Agric. Food Chem. 2022, 70, 4860–4870. [Google Scholar] [CrossRef]
- Santoyo, G.; Moreno-Hagelsieb, G.; del Carmen Orozco-Mosqueda, M.; Glick, B.R. Plant growth-promoting bacterial endophytes. Microbiol. Res. 2016, 183, 92–99. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zhang, X.; Fu, S.; Zhang, W. Environmental Filtering Drives Local Soil Fungal Beta Diversity More Than Dispersal Limitation in Six Forest Types along a Latitudinal Gradient in Eastern China. Forests 2019, 10, 863. [Google Scholar] [CrossRef]
- Shen, S.Y.; Fulthorpe, R. Seasonal variation of bacterial endophytes in urban trees. Front. Microbiol. 2015, 6, 427. [Google Scholar] [CrossRef] [PubMed]
- Al Ashhab, A.; Meshner, S.; Alexander-Shani, R.; Dimerets, H.; Brandwein, M.; Bar-Lavan, Y.; Winters, G. Temporal and Spatial Changes in Phyllosphere Microbiome of Acacia Trees Growing in Arid Environments. Front. Microbiol. 2021, 12, 656269. [Google Scholar] [CrossRef] [PubMed]
- Arunrat, N.; Sansupa, C.; Sereenonchai, S.; Hatano, R. Stability of soil bacteria in undisturbed soil and continuous maize cultivation in Northern Thailand. Front. Microbiol. 2023, 14, 1285445. [Google Scholar] [CrossRef] [PubMed]
- Sangiorgio, D.; Cáliz, J.; Mattana, S.; Barceló, A.; De Cinti, B.; Elustondo, D.; Hellsten, S.; Magnani, F.; Matteucci, G.; Merilä, P.; et al. Host species and temperature drive beech and Scots pine phyllosphere microbiota across European forests. Commun. Earth Environ. 2024, 5, 747. [Google Scholar] [CrossRef]
- Vandenkoornhuyse, P.; Quaiser, A.; Duhamel, M.; Le Van, A.; Dufresne, A. The importance of the microbiome of the plant holobiont. New Phytol. 2015, 206, 1196–1206. [Google Scholar] [CrossRef]
- Han, F.; Ye, X.; Chen, Q.; Long, H.; Rao, Y. The oxidative degradation of diclofenac using the activation of peroxymonosulfate by BiFeO3 microspheres—Kinetics, role of visible light and decay pathways. Sep. Purif. Technol. 2020, 232, 115967. [Google Scholar] [CrossRef]
- Carro, L.; Golinska, P.; Nouioui, I.; Bull, A.T.; Igual, J.M.; Andrews, B.A.; Klenk, H.P.; Goodfellow, M. Micromonospora acroterricola sp. nov., a novel actinobacterium isolated from a high-altitude Atacama Desert soil. Int. J. Syst. Evol. Microbiol. 2019, 69, 3426–3436. [Google Scholar] [CrossRef]
- Jones, A.L.; Taylor, J.E.; Ennos, R.A. Foliar Fungal Endophyte Communities of Scottish Plantation Pines. J. Fungi 2025, 11, 148. [Google Scholar] [CrossRef]
- Liu-Xu, L.; Ma, L.; Farvardin, A.; García-Agustín, P.; Llorens, E. Exploring the impact of plant genotype and fungicide treatment on endophytic communities in tomato stems. Front. Microbiol. 2024, 15, 1453669. [Google Scholar] [CrossRef]
- Laforest-Lapointe, I.; Messier, C.; Kembel, S.W. Host species identity, site and time drive temperate tree phyllosphere bacterial community structure. Microbiome 2016, 4, 27. [Google Scholar] [CrossRef]
- Lugtenberg, B.; Kamilova, F. Plant-growth-promoting rhizobacteria. Annu. Rev. Microbiol. 2009, 63, 541–556. [Google Scholar] [CrossRef]
- Kim, S.J.; Kwon, K.K.; Kim, B.K.; Kim, J.F.; Lee, H.S. Genome sequence of Novosphingobium pentaromativorans US6-1T, a high-molecular-weight polycyclic aromatic hydrocarbon-degrading marine bacterium. J. Bacteriol. 2009, 191, 6224–6225. [Google Scholar] [CrossRef]
- Jia, X.; Hong, L.; Wang, Y.; Zhang, Q.; Wang, Y.; Jia, M.; Luo, Y.; Wang, T.; Ye, J.; Wang, H. Effect of microbial diversity and their functions on soil nutrient cycling in the rhizosphere zone of Dahongpao mother tree and cutting Dahongpao. Front. Plant Sci. 2025, 16, 1574020. [Google Scholar] [CrossRef]
- Gan, H.M.; Hudson, A.O.; Rahman, A.Y.A.; Chan, K.G. Comparative genomics reveals environmental adaptation of Novosphingobium. Microorganisms 2022, 10, 500. [Google Scholar] [CrossRef]
- Sohn, J.H.; Kwon, K.K.; Kang, J.H.; Jung, H.B.; Kim, S.J. Novosphingobium pentaromativorans sp. nov., a high-molecular-weight polycyclic aromatic hydrocarbon-degrading bacterium isolated from petroleum-contaminated sediment. Int. J. Syst. Evol. Microbiol. 2004, 54, 1483–1487. [Google Scholar] [CrossRef]
- Aylward, F.O.; McDonald, B.R.; Adams, S.M.; Valenzuela, A.; Schmidt, R.A.; Goodwin, L.A.; Woyke, T.; Currie, C.R. Comparison of 26 sphingomonad genomes reveals diverse environmental adaptations and biodegradative potential. Front. Microbiol. 2013, 4, 373. [Google Scholar] [CrossRef]
- Lyu, Y.; Zheng, W.; Zheng, T.; Tian, Y. Biodegradation of polycyclic aromatic hydrocarbons by Novosphingobium pentaromativorans US6-1. PLoS ONE 2014, 9, e101438. [Google Scholar] [CrossRef] [PubMed]
- Petrova, S.; Velcheva, I.; Nikolov, B. Nature-Based Solutions to Reduce Air Pollution: A Case Study from Plovdiv, Bulgaria, Using Trees, Herbs, Mosses and Lichens. Forests 2024, 15, 928. [Google Scholar] [CrossRef]







| Urbanisation Intensity | Built-Up Area (%) | Traffic (Cars Per Hour) | Experimental Plot |
|---|---|---|---|
| Low | 0–10 | 0–60 | Plot 4 |
| Medium | 10–40 | 61–300 | Plot 2 |
| High | 40–70 | 301–900 | Plot 1 |
| Very high | >70 | >900 | Plot 3 |
| Sample | Raw PE | Combined | Qualified | No Chime | Base (nt) | Avglen (nt) | GC | Q20 | Q30 |
|---|---|---|---|---|---|---|---|---|---|
| F1.P | 64,588 | 64,274 | 63,333 | 58,417 | 21,977,137 | 376.21 | 53.14% | 98.68% | 95.10% |
| F2.P | 66,148 | 65,793 | 64,779 | 62,919 | 23,691,812 | 376.54 | 53.51% | 98.89% | 95.83% |
| F3.P | 79,697 | 78,923 | 77,523 | 51,791 | 19,546,406 | 377.41 | 54.11% | 98.86% | 95.92% |
| F4.P | 65,816 | 65,429 | 64,400 | 61,858 | 23,170,827 | 374.58 | 54.37% | 98.90% | 95.96% |
| T1.P | 58,405 | 58,071 | 56,991 | 52,010 | 19,552,157 | 375.93 | 53.88% | 98.86% | 95.87% |
| T2.P | 91,504 | 90,878 | 89,848 | 69,674 | 26,208,769 | 376.16 | 54.51% | 98.99% | 96.32% |
| T3.P | 75,574 | 75,183 | 73,835 | 45,838 | 17,215,999 | 375.58 | 53.21% | 98.84% | 95.89% |
| T4.P | 47,877 | 47,656 | 47,027 | 35,583 | 13,395,823 | 376.47 | 55.58% | 98.92% | 96.10% |
| P1.P | 65,335 | 64,893 | 63,534 | 51,120 | 19,277,943 | 377.11 | 53.75% | 98.80% | 95.74% |
| P2.P | 68,135 | 66,196 | 65,404 | 58,723 | 22,142,632 | 377.07 | 54.38% | 98.99% | 96.27% |
| P3.P | 64,483 | 64,121 | 63,028 | 54,908 | 20,727,695 | 377.50 | 53.72% | 98.87% | 95.87% |
| P4.P | 65,047 | 63,751 | 62,646 | 53,696 | 20,246,834 | 377.06 | 53.79% | 98.89% | 95.97% |
| PL.P | 64,312 | 63,993 | 63,327 | 61,426 | 23,028,129 | 374.89 | 54.61% | 98.90% | 96.03% |
| (A) Tilia tomentosa vs. Fraxinus excelsior | |||
| Genus | Enriched in | Reported Significance | Known Ecological Role |
| Staphylococcus | Tilia tomentosa | 0.041 | stress tolerance/environmental persistence |
| Additibacterium | Tilia tomentosa | p < 0.05 | environmental resilience |
| Friedmanniella | Tilia tomentosa | p < 0.05 | Actinobacteria plant association |
| Pseudolabrys | Tilia tomentosa | p < 0.05 | nitrogen cycling |
| Arenimonas | Tilia tomentosa | p < 0.05 | oxidative stress tolerance |
| Lawsonella | Tilia tomentosa | p < 0.05 | environmental persistence |
| Fenollaria | Tilia tomentosa | p < 0.05 | anaerobic metabolism |
| Variovorax | Tilia tomentosa | p < 0.05 | plant growth promotion, hormone regulation |
| Thermicanus | Fraxinus excelsior | p < 0.05 | environmental adaptation |
| (B) Fraxinus excelsior vs. Pinus nigra | |||
| Genus | Enriched in Host | Reported Significance | Putative Ecological Function |
| Brevundimonas | Fraxinus excelsior | 0.031 | plant-growth promotion, stress tolerance |
| Microvirga | Fraxinus excelsior | <0.05 | nitrogen fixation, rhizosphere association |
| Rubellimicrobium | Fraxinus excelsior | <0.05 | environmental resilience, organic compound metabolism |
| Craurococcus | Fraxinus excelsior | 0.019–0.050 | stress adaptation (reported soil taxa) |
| Arthrobacter | Fraxinus excelsior | <0.05 | pollutant tolerance, nutrient cycling |
| Paenibacillus | Fraxinus excelsior | <0.05 | PGPR, pathogen suppression |
| Tumebacillus | Fraxinus excelsior | <0.05 | environmental persistence |
| Enterococcus | Fraxinus excelsior | <0.05 | plant-associated metabolism |
| Novosphingobium | Pinus nigra | <0.05 | hydrocarbon degradation, xenobiotic metabolism |
| Bacteroides | Pinus nigra | <0.05 | organic matter turnover |
| Finegoldia | Pinus nigra | <0.05 | stress-adapted niche taxa |
| (C) Tilia tomentosa vs. Pinus nigra | |||
| Genus | Enriched in Host | Statistical Significance | Putative Ecological Function |
| Arthrobacter | Tilia tomentosa | p < 0.05 | Nutrient cycling; pollutant resistance |
| Microvirga | Tilia tomentosa | p < 0.05 | Nitrogen fixation; plant association |
| Adhaeribacter | Tilia tomentosa | p < 0.05 | Organic matter degradation |
| Enterococcus | Tilia tomentosa | p < 0.05 | Plant-associated metabolism |
| Blautia | Tilia tomentosa | p < 0.05 | Carbon metabolism |
| Novosphingobium | Pinus nigra | p < 0.05 | Xenobiotic and aromatic hydrocarbon degradation |
| Roseburia | Pinus nigra | p < 0.05 | Metabolic specialisation |
| Lineage 1174-901-12 | Pinus nigra | p < 0.05 | Environmental stress tolerance |
| Sample | Chao1 | Dominance | Goods_Coverage | Observed_Features | Pielou_e | Shannon | Simpson |
|---|---|---|---|---|---|---|---|
| F1.P | 560.000 | 0.097 | 1.000 | 556 | 0.632 | 5.765 | 0.903 |
| F2.P | 308.000 | 0.298 | 1.000 | 306 | 0.423 | 3.489 | 0.702 |
| F3.P | 380.667 | 0.089 | 1.000 | 379 | 0.677 | 5.798 | 0.911 |
| F4.P | 158.500 | 0.267 | 1.000 | 157 | 0.523 | 3.813 | 0.733 |
| T1.P | 511.60 | 0.195 | 1.000 | 511 | 0.502 | 4.517 | 0.805 |
| T2.P | 686.630 | 0.129 | 0.999 | 679 | 0.534 | 5.025 | 0.871 |
| T3.P | 325.231 | 0.102 | 1.000 | 325 | 0.571 | 4.761 | 0.898 |
| T4.P | 250.500 | 0.396 | 1.000 | 246 | 0.381 | 3.030 | 0.604 |
| P1.P | 215.364 | 0.130 | 1.000 | 214 | 0.570 | 4.414 | 0.870 |
| P2.P | 239.667 | 0.039 | 1.000 | 238 | 0.735 | 5.799 | 0.961 |
| P3.P | 193.300 | 0.082 | 1.000 | 193 | 0.620 | 4.710 | 0.918 |
| P4.P | 271.000 | 0.177 | 1.000 | 271 | 0.508 | 4.108 | 0.823 |
| PL.P | 92.00 | 0.201 | 1.000 | 92 | 0.519 | 3.383 | 0.799 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Petkova, M.; Shilev, S.; Nikolov, B.; Petrova, S. Urbanisation Shapes the Diversity, Composition, and Functional Profile of Endophytic Bacteriome in Common Urban Tree Species. Forests 2026, 17, 313. https://doi.org/10.3390/f17030313
Petkova M, Shilev S, Nikolov B, Petrova S. Urbanisation Shapes the Diversity, Composition, and Functional Profile of Endophytic Bacteriome in Common Urban Tree Species. Forests. 2026; 17(3):313. https://doi.org/10.3390/f17030313
Chicago/Turabian StylePetkova, Mariana, Stefan Shilev, Bogdan Nikolov, and Slaveya Petrova. 2026. "Urbanisation Shapes the Diversity, Composition, and Functional Profile of Endophytic Bacteriome in Common Urban Tree Species" Forests 17, no. 3: 313. https://doi.org/10.3390/f17030313
APA StylePetkova, M., Shilev, S., Nikolov, B., & Petrova, S. (2026). Urbanisation Shapes the Diversity, Composition, and Functional Profile of Endophytic Bacteriome in Common Urban Tree Species. Forests, 17(3), 313. https://doi.org/10.3390/f17030313








