Integrated Analysis of Fatty Acid Metabolism and Transcriptome Involved in Olive Fruit Development to Improve Oil Composition
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Transcriptome Processing and Bioinformatics Analysis
2.3. Targeted Fatty Acid Metabolite Analysis
2.4. Statistical Analyses
3. Results and Discussion
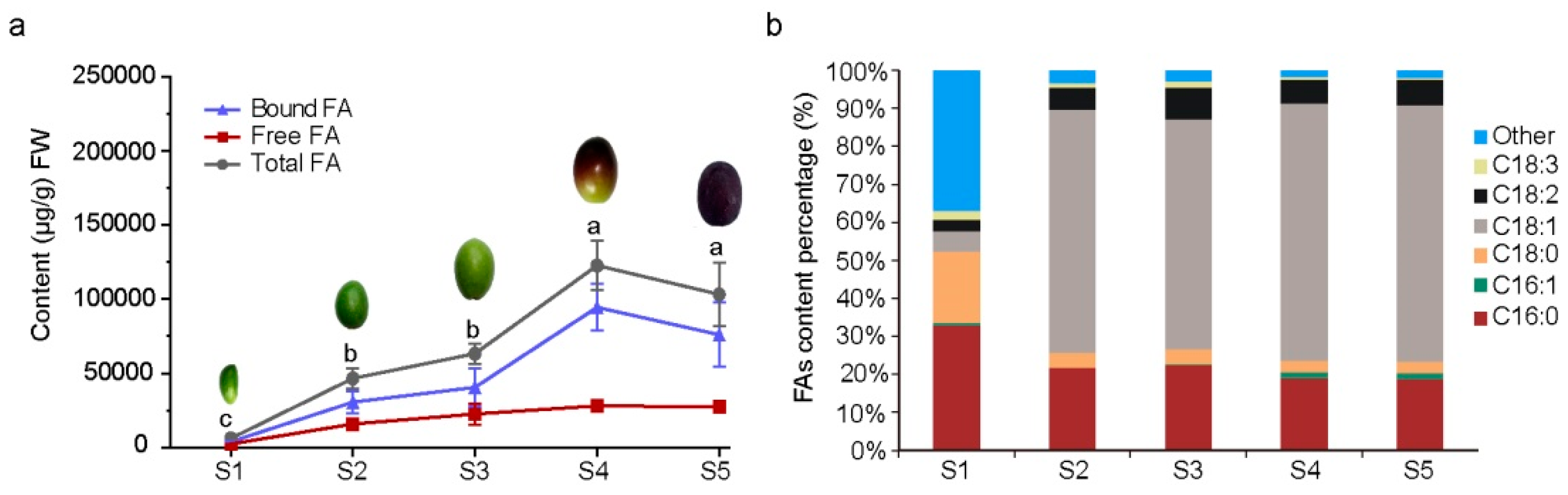
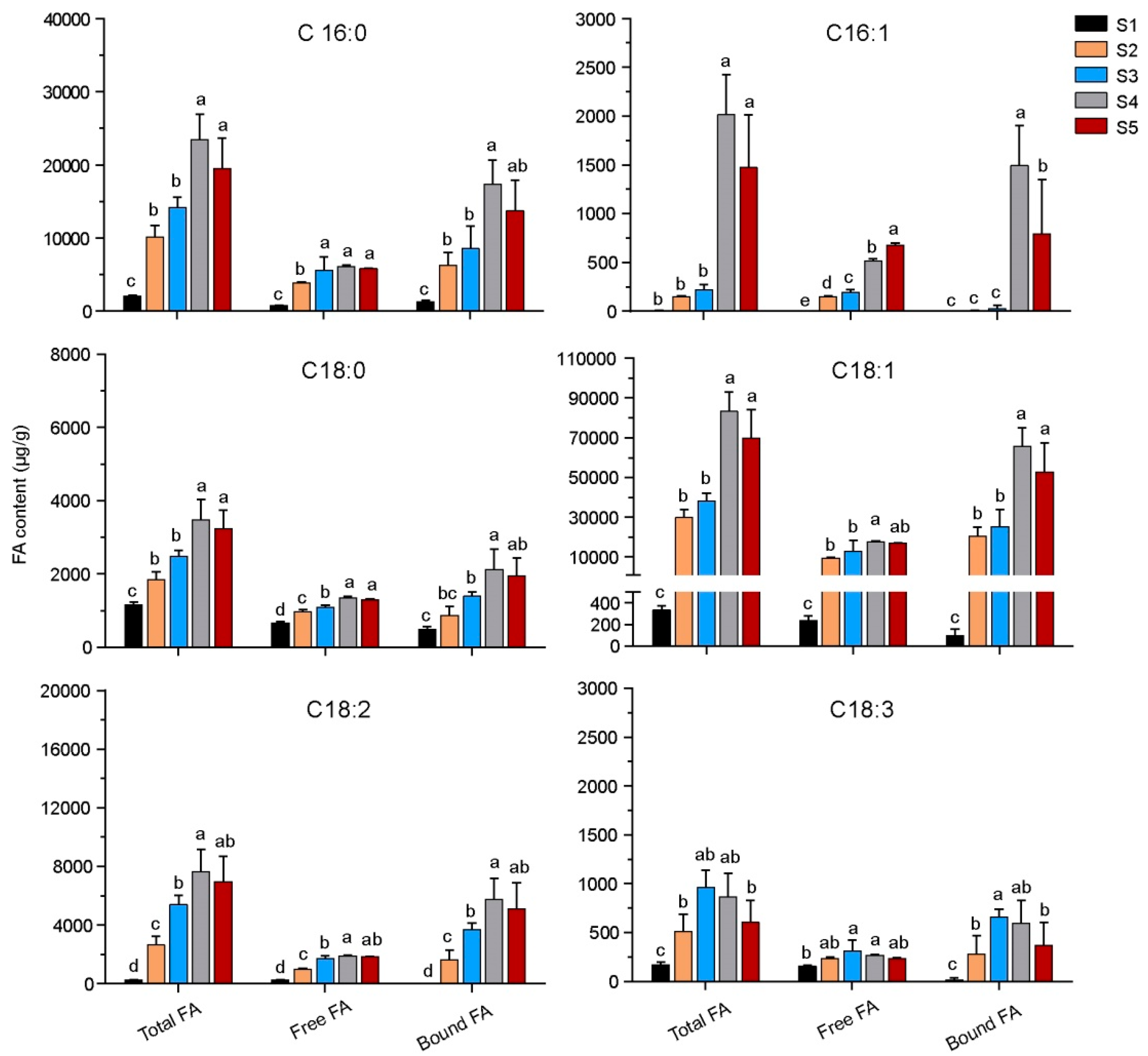
3.1. Fatty Acids Content of Olive Pulp during Maturation
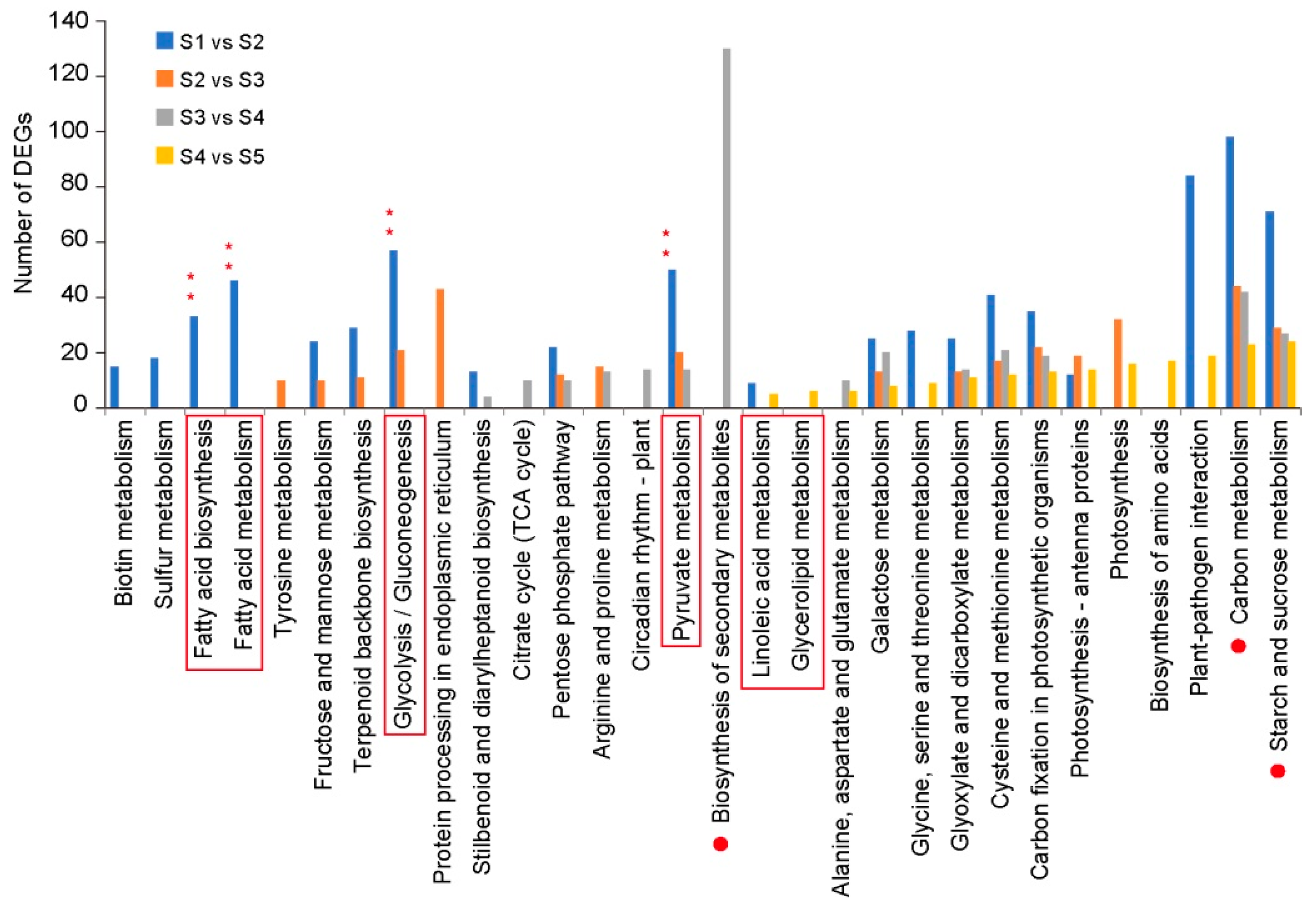
3.2. Transcriptome Data Analysis
3.3. Identification and Characterization of Genes Involved in Glycolysis at Five Developmental Stages
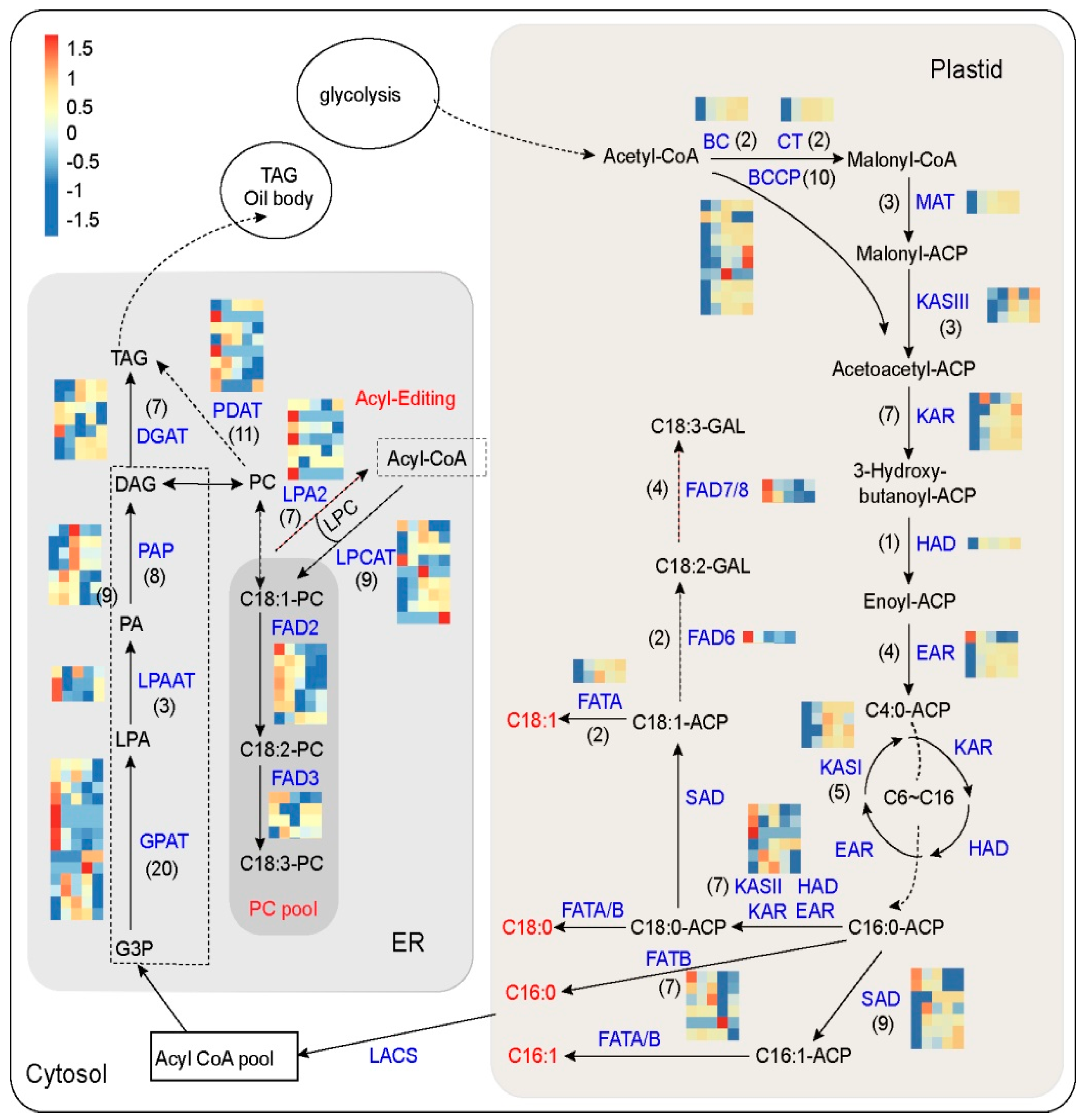
3.4. Identification of Genes Involved in FA and TAG Metabolism at Five Developmental Stages
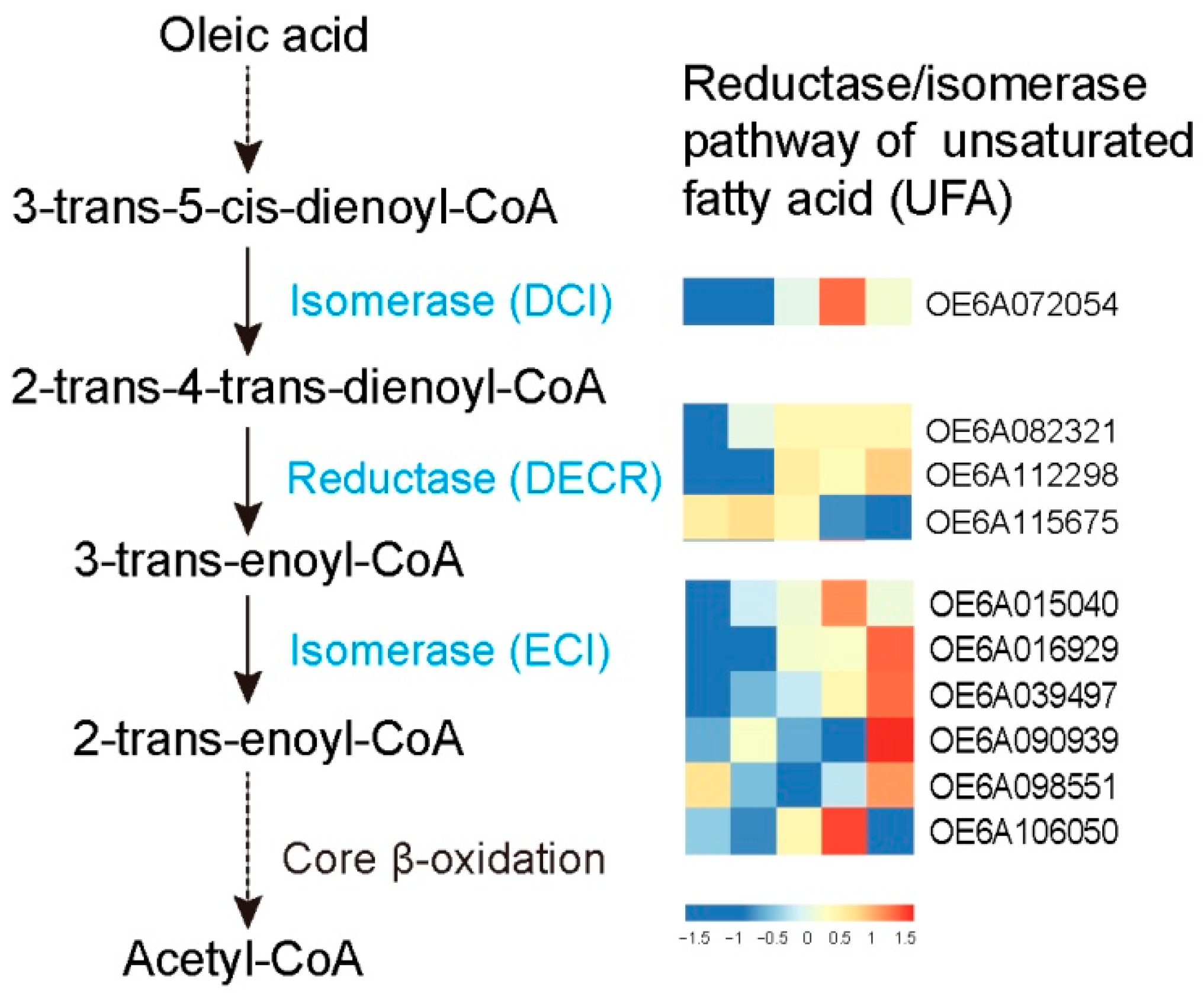
3.5. FA Degradation at Different Stages
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rosa, R.D.L.; Belaj, A.; Mérida, A.M.; Trelles, O.; López, C.B. Development of EST-derived SSR Markers with Long-core Repeat in Olive and Their Use for Paternity Testing. J. Am. Soc. Hortic. Sci. 2013, 138, 290–296. [Google Scholar] [CrossRef]
- Diez, C.M.; Trujillo, I.; Martinez-Urdiroz, N.; Barranco, D.; Rallo, L.; Marfil, P.; Gaut, B.S. Olive domestication and diversification in the Mediterranean Basin. New Phytol. 2015, 206, 436–447. [Google Scholar] [CrossRef] [PubMed]
- Gavriilidou, V.; Boskou, D. Chemical interesterification of olive-tristearin blends for margarines. Int. J. Food Sci. Technol. 2007, 26, 451–456. [Google Scholar] [CrossRef]
- Parvini, F.; Sicardo, M.D.; Hosseini-Mazinani, M.; Martinez-Rivas, J.M.; Hernandez, M.L. Transcriptional Analysis of Stearoyl-Acyl Carrier Protein Desaturase Genes from Olive (Olea europaea) in Relation to the Oleic Acid Content of the Virgin Olive Oil. J. Agric. Food Chem. 2016, 64, 7770–7781. [Google Scholar] [CrossRef] [PubMed]
- Ben Brahim, S.; Kelebek, H.; Ammar, S.; Abichou, M.; Bouaziz, M. LC-MS phenolic profiling combined with multivariate analysis as an approach for the characterization of extra virgin olive oils of four rare Tunisian cultivars during ripening. Food Chem. 2017, 229, 9–19. [Google Scholar] [CrossRef]
- Cruz, F.; Julca, I.; Gomez-Garrido, J.; Loska, D.; Marcet-Houben, M.; Cano, E.; Galan, B.; Frias, L.; Ribeca, P.; Derdak, S.; et al. Genome sequence of the olive tree, Olea europaea. GigaScience 2016, 5, 29. [Google Scholar] [CrossRef]
- Dabbou, S.; Chaieb, I.; Rjiba, I.; Issaoui, M.; Echbili, A.; Nakbi, A.; Gazzah, N.; Hammami, M. Multivariate Data Analysis of Fatty Acid Content in the Classification of Olive Oils Developed Through Controlled Crossbreeding. J. Am. Oil Chem. Soc. 2011, 89, 667–674. [Google Scholar] [CrossRef]
- Flores, G.; Blanch, G.P.; del Castillo, M.L.R. Effect of postharvest methyl jasmonate treatment on fatty acid composition and phenolic acid content in olive fruits during storage. J. Sci. Food Agric. 2017, 97, 2767–2772. [Google Scholar] [CrossRef]
- Cheng, C.; Wang, D.; Xia, H.; Wang, F.; Yang, X.; Pan, D.; Wang, S.; Yang, L.; Lu, H.; Shu, G.; et al. A comparative study of the effects of palm olein, cocoa butter and extra virgin olive oil on lipid profile, including low-density lipoprotein subfractions in young healthy Chinese people. Int. J. Food Sci. Nutr. 2019, 70, 355–366. [Google Scholar] [CrossRef]
- Bates, P.D.; Stymne, S.; Ohlrogge, J. Biochemical pathways in seed oil synthesis. Curr. Opin. Plant. Biol. 2013, 16, 358–364. [Google Scholar] [CrossRef] [Green Version]
- Lung, S.-C.; Weselake, R.J. Diacylglycerol acyltransferase: A key mediator of plant triacylglycerol synthesis. Lipids 2006, 41, 1073–1088. [Google Scholar] [CrossRef] [PubMed]
- Contreras, C.; Mariotti, R.; Mousavi, S.; Baldoni, L.; Torres, M. Characterization and validation of olive FAD and SAD gene families: Expression analysis in different tissues and during fruit development. Mol. Biol. Rep. 2020, 47, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Wei, X.; Liu, W.; Min, X.; Jin, X.; Ndayambaza, B.; Wang, Y. Genome-wide identification and expression analysis of the fatty acid desaturase genes in Medicago truncatula. Biochem. Biophys. Res. Commun. 2018, 499, 361–367. [Google Scholar] [CrossRef]
- Chia, T.Y.; Pike, M.J.; Rawsthorne, S. Storage oil breakdown during embryo development of Brassica napus (L.). J. Exp. Bot. 2005, 56, 1285–1296. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, M.L.; Sicardo, M.D.; Martinez-Rivas, J.M. Differential Contribution of Endoplasmic Reticulum and Chloroplast omega-3 Fatty Acid Desaturase Genes to the Linolenic Acid Content of Olive (Olea europaea) Fruit. Plant. Cell Physiol. 2016, 57, 138–151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bodoira, R.; Torres, M.; Pierantozzi, P.; Taticchi, A.; Servili, M.; Maestri, D. Oil biogenesis and antioxidant compounds from "Arauco" olive (Olea europaea L.) cultivar during fruit development and ripening. Eur. J. Lipid Sci. Technol. 2015, 117, 377–388. [Google Scholar] [CrossRef]
- Graham, I.A. Seed storage oil mobilization. Annu. Rev. Plant. Biol. 2008, 59, 115–142. [Google Scholar] [CrossRef]
- Guodong, R.; Jianguo, Z.; Xiaoxia, L.; Ying, L. Identification of putative genes for polyphenol biosynthesis in olive fruits and leaves using full-length transcriptome sequencing. Food Chem. 2019, 300, 125246. [Google Scholar] [CrossRef] [PubMed]
- Clark, T.A. Full-Length cDNA Sequencing on the PacBio Sequel Platform. Available online: http://www.pacb.com/applications/rnasequencing/2017 (accessed on 10 December 2015).
- Hackl, T.; Hedrich, R.; Schultz, J.; Forster, F. Proovread: Large-scale high-accuracy PacBio correction through iterative short read consensus. Bioinformatics 2014, 30, 392. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Varet, H.; Brillet-Gueguen, L.; Coppee, J.Y.; Dillies, M.A. SARTools: A DESeq2- and EdgeR-Based R Pipeline for Comprehensive Differential Analysis of RNA-Seq Data. PLoS ONE 2016, 11, e0157022. [Google Scholar] [CrossRef] [Green Version]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, L.-X.; Zou, L.; Tan, M.-L.; Deng, Y.-Y.; Yan, J.; Yan, Z.-Y.; Zhao, G. Free amino acids, fatty acids and phenolic compounds in Tartary buckwheat of different hull colour. Czech. J. Food Sci. 2017, 35, 214–222. [Google Scholar] [CrossRef] [Green Version]
- Zhou, L.; Wang, H.; Chen, X.; Li, Y.; Hussain, N.; Cui, L.; Wu, D.; Jiang, L. Identification of candidate genes involved in fatty acids degradation at the late maturity stage in Brassica napus based on transcriptomic analysis. Plant. Growth Regul. 2017, 83, 385–396. [Google Scholar] [CrossRef]
- Wang, X.; Liang, H.; Guo, D.; Guo, L.; Duan, X.; Jia, Q.; Hou, X. Integrated analysis of transcriptomic and proteomic data from tree peony (P. ostii) seeds reveals key developmental stages and candidate genes related to oil biosynthesis and fatty acid metabolism. Hortic. Res. 2019, 6, 111. [Google Scholar] [CrossRef] [Green Version]
- Weselake, R.J.; Taylor, D.C.; Rahman, M.H.; Shah, S.; Laroche, A.; McVetty, P.B.E.; Harwood, J.L. Increasing the flow of carbon into seed oil. Biotechnol. Adv. 2009, 27, 866–878. [Google Scholar] [CrossRef]
- Thelen, J.J.; Ohlrogge, J.B. Both antisense and sense expression of biotin carboxyl carrier protein isoform 2 inactivates the plastid acetyl-coenzyme A carboxylase in Arabidopsis thaliana. Plant. J. 2002, 32, 419–431. [Google Scholar] [CrossRef]
- Huang, J.; Zhang, T.; Zhang, Q.; Chen, M.; Wang, Z.; Zheng, B.; Xia, G.; Yang, X.; Huang, C.; Huang, Y. The mechanism of high contents of oil and oleic acid revealed by transcriptomic and lipidomic analysis during embryogenesis in Carya cathayensis Sarg. BMC Genom. 2016, 17, 113. [Google Scholar] [CrossRef] [Green Version]
- Wu, G.Z.; Xue, H.W. Arabidopsis beta-ketoacyl-[acyl carrier protein] synthase i is crucial for fatty acid synthesis and plays a role in chloroplast division and embryo development. Plant. Cell 2010, 22, 075564. [Google Scholar] [CrossRef] [Green Version]
- Gupta, M.; Dekelver, R.C.; Palta, A.; Clifford, C.; Petolino, J.F. Oil Modification via Transcriptional Activation of Canola KASII Using an Engineered Zinc Finger Transcription Factor. Vitr. Cell. Dev. Biol. Anim. 2012, 48, 25. [Google Scholar]
- Herńndez, M.L.; Padilla, M.A.N.; Mancha, M.; Martínez-Rivas, J.M. Expression Analysis IdentifiesFAD2-2as the Olive Oleate Desaturase Gene Mainly Responsible for the Linoleic Acid Content in Virgin Olive Oil. J. Agric. Food Chem. 2009, 57, 6199–6206. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Perez, A.J.; Venegas-Caleron, M.; Vaistij, F.E.; Salas, J.J.; Larson, T.R.; Garces, R.; Graham, I.A.; Martinez-Force, E. Reduced expression of FatA thioesterases in Arabidopsis affects the oil content and fatty acid composition of the seeds. Planta 2012, 235, 629–639. [Google Scholar] [CrossRef] [PubMed]
- Salas, J.J.; Ohlrogge, J.B. Characterization of substrate specificity of plant FatA and FatB acyl-ACP thioesterases. Arch. Biochem. Biophys. 2002, 403, 25–34. [Google Scholar] [CrossRef]
- Bonaventure, G.; Salas, J.; Pollard, J.; Michael, R.; Ohlrogge, J. Disruption of the FATB Gene in Arabidopsis Demonstrates an Essential Role of Saturated Fatty Acids in Plant Growth. Plant. Cell 2003, 15, 1020–1033. [Google Scholar] [CrossRef] [Green Version]
- Janssen, J.H.; Spoelder, J.; Koehorst, J.J.; Schaap, P.J.; Barbosa, M.J. Time-dependent transcriptome profile of genes involved in triacylglycerol (TAG) and polyunsaturated fatty acid synthesis in Nannochloropsis gaditana during nitrogen starvation. J. Appl. Phycol. 2020, 32, 1153–1164. [Google Scholar] [CrossRef] [Green Version]
- Dahlqvist, A.; Stahl, U.; Lenman, M.; Banas, A.; Lee, M.; Sandager, L.; Ronne, H.; Stymne, S. Phospholipid:diacylglycerol acyltransferase: An enzyme that catalyzes the acyl-CoA-independent formation of triacylglycerol in yeast and plants. Proc. Natl. Acad. Sci. USA 2000, 97, 6487–6492. [Google Scholar] [CrossRef] [Green Version]
- Lu, S.; Song, T.; Kosma, D.K.; Parsons, E.P.; Rowland, O.; Jenks, M.A. Arabidopsis CER8 encodes LONG-CHAIN ACYL-COA SYNTHETASE 1 (LACS1) that has overlapping functions with LACS2 in plant wax and cutin synthesis. Plant. J. 2009, 59, 553–564. [Google Scholar] [CrossRef]
- Zhao, L.; Katavic, V.; Li, F.; Haughn, G.W.; Kunst, L. Insertional mutant analysis reveals that long-chain acyl-CoA synthetase 1 (LACS1), but not LACS8, functionally overlaps with LACS9 in Arabidopsis seed oil biosynthesis. Plant. J. 2010, 64, 1048–1058. [Google Scholar] [CrossRef]
- Fulda, M.; Schnurr, J.; Abbadi, A.; Heinz, E.; Browse, J. Peroxisomal Acyl-CoA Synthetase Activity Is Essential for Seedling Development in Arabidopsis thaliana. Plant. Cell 2004, 16, 394–405. [Google Scholar] [CrossRef] [Green Version]



| Summary of Sequence | |
|---|---|
| Subreads base (G) | 10.55 |
| Average subreads length (bp) | 1791 |
| Consensus reads | 187,517 |
| Total error-corrected consensus reads | 186,150 |
| Circular consensus sequence (CCSs) | 492,350 |
| Full-length non chimera (FLNC) reads | 399,263 |
| Total mapped of GMAP on cultivar Farga | 175,700 (94.39%) |
| Sample | Total Reads | Total Mapped | Unmapped | Multiple Mapped | Uniquely Mapped |
|---|---|---|---|---|---|
| S1 | 260,230,962 | 237,461,682 (91.25%) | 22,769,280 (8.75%) | 29,826,494 (11.46%) | 19,259,528 (7.40%) |
| S2 | 331,998,532 | 303,326,286 (91.36%) | 28,672,246 (8.64%) | 34,811,070 (10.49%) | 18,038,372 (5.43%) |
| S3 | 306,350,130 | 280,941,378 (91.71%) | 25,408,752 (8.29%) | 36,341,656 (11.86%) | 18,510,870 (6.04%) |
| S4 | 333,794,400 | 312,544,680 (93.63%) | 21,249,720 (6.37%) | 34,543,262 (10.35%) | 17,451,618 (5.23%) |
| S5 | 264,183,768 | 241,789,036 (91.52%) | 22,394,732 (8.48%) | 31,835,116 (12.05%) | 18,850,240 (7.14%) |
| Candidate Genes | S1 vs. S2 | S2 vs. S3 | S3 vs. S4 | S4 vs. S5 | Gene Name |
|---|---|---|---|---|---|
| OE6A002753 | 2.0107 | LPAAT | |||
| OE6A020169 | 4.19 | −3.2021 | LPAAT | ||
| OE6A007392 | −1.5115 | PDAT1 | |||
| OE6A093626 | −1.9341 | DGAT | |||
| OE6A108115 | DGAT | ||||
| OE6A119597 | 1.8299 | −1.9289 | DGAT | ||
| OE6A030092 | −2.1724 | DGAT | |||
| OE6A102251 | 1.6139 | LPCAT | |||
| OE6A111035 | −1.2815 | −1.1866 | LPCAT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Guo, L.; Zhang, J.; Xue, L.; Luo, Y.; Rao, G. Integrated Analysis of Fatty Acid Metabolism and Transcriptome Involved in Olive Fruit Development to Improve Oil Composition. Forests 2021, 12, 1773. https://doi.org/10.3390/f12121773
Liu X, Guo L, Zhang J, Xue L, Luo Y, Rao G. Integrated Analysis of Fatty Acid Metabolism and Transcriptome Involved in Olive Fruit Development to Improve Oil Composition. Forests. 2021; 12(12):1773. https://doi.org/10.3390/f12121773
Chicago/Turabian StyleLiu, Xiaoxia, Liqin Guo, Jianguo Zhang, Li Xue, Ying Luo, and Guodong Rao. 2021. "Integrated Analysis of Fatty Acid Metabolism and Transcriptome Involved in Olive Fruit Development to Improve Oil Composition" Forests 12, no. 12: 1773. https://doi.org/10.3390/f12121773
APA StyleLiu, X., Guo, L., Zhang, J., Xue, L., Luo, Y., & Rao, G. (2021). Integrated Analysis of Fatty Acid Metabolism and Transcriptome Involved in Olive Fruit Development to Improve Oil Composition. Forests, 12(12), 1773. https://doi.org/10.3390/f12121773







