Triterpenoids Biosynthesis Regulation for Leaf Coloring of Wheel Wingnut (Cyclocaryapaliurus)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Determination of Triterpenoids
2.3. RNA Extraction, Quantification and Sequencing
2.4. Transcriptome Data Analysis
3. Results
3.1. Triterpenoid Individuals Analysis
3.2. RNA-seq and Data Analysis
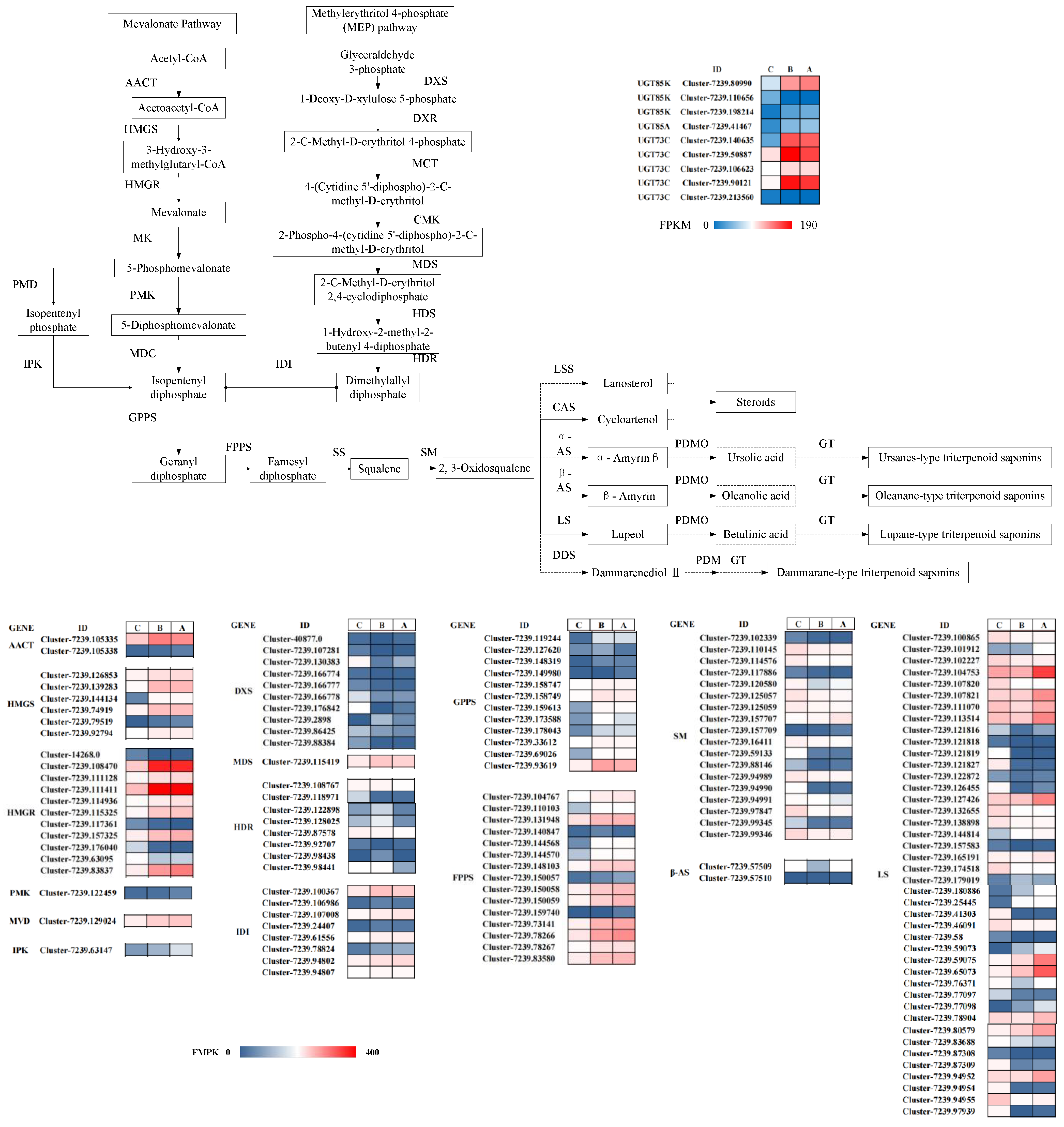
3.3. Transcriptomic and Metabolomic Association Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Li, X.; Fu, X.; Shang, X.; Yang, W.; Fang, S. Natural population structure and genetic differentiation for heterodicogamous plant: Cyclocarya paliurus (Batal.) Iljinskaja (Juglandaceae). Tree Genet. Genomes 2017, 13, 80. [Google Scholar] [CrossRef]
- Sun, C.; Zhou, Y.; Fang, S.; Shang, X. Ecological Gradient Analysis and Environmental Interpretation of Cyclocarya paliurus Communities. Forests 2021, 12, 146. [Google Scholar] [CrossRef]
- Sun, C.; Zhong, W.; Fu, X.; Shang, X.; Fang, S. A study on growth and aboveground biomass production of juvenile Cyclocarya paliurus plantations. J. Nanjing For. Univ. (Nat. Sci. Ed.) 2021, 10, 387. Available online: https://kns.cnki.net/kcms/detail/32.1161.S.20210825.0918.002.html/ (accessed on 25 August 2021).
- Fang, S.Z.; Fu, X.X. Progress and Prospects on Silviculture and Utilization of Cyclocarya paliurus Resources. J. Nanjing For. Univ. (Nat. Sci. Ed.) 2007, 31, 95–100. [Google Scholar]
- Chen, X.; Chen, B.; Shang, X.; Fang, S. Rna in situ hybridization and expression of related genes regulating the accumulation of triterpenoids in Cyclocarya paliurus. Tree Physiol. 2021, 41, 2189–2197. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Shang, X.; Ding, H.; Cao, Y.; Fang, S. Natural Variations in Flavonoids and Triterpenoids of Cyclocarya paliurus Leaves. J. For. Res. 2020, 31, 805–814. [Google Scholar] [CrossRef]
- Tian, L.; Xu, C.; Shang, X.; Fu, X. Evaluation and selection on superior individuals for medicinal use of Cyclocarya paliurus. J. Nanjing For. Univ. (Nat. Sci. Ed.) 2021, 45, 21–28. [Google Scholar] [CrossRef]
- Han, M.; Yang, C.; Zhou, J.; Zhu, J.; Meng, J.; Shen, T.; Li, H. Analysis of flavonoids and anthocyanin biosynthesis-related genes expression reveals the mechanism of petal color fading of Malus hupehensis (Rosaceae). Braz. J. Bot. 2020, 43, 81–89. [Google Scholar] [CrossRef]
- Xu, Y.; Chen, Z.; Jia, L.; Weng, X. Advances in understanding of the biosynthetic pathway and regulatory mechanism of triterpenoid saponins in plants. Sci. Sin. Vitae 2021, 51, 525–555. [Google Scholar] [CrossRef]
- Sheng, X.; Chen, H.; Wang, J.; Zheng, Y.; Li, J. Joint transcriptomic and metabolic analysis of flavonoids in Cyclocarya paliurus leaves. ACS Omega 2021, 6, 9028–9038. [Google Scholar] [CrossRef] [PubMed]
- Liao, W.; Mei, Z.; Miao, L.; Liu, P.; Gao, R. Comparative transcriptome analysis of root, stem, and leaf tissues of Entada phaseoloides reveals potential genes involved in triterpenoid saponin biosynthesis. BMC Genom. 2019, 21, 639. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.Y.; Zhang, S.X.; Peng, D.Y.; Wang, C.K.; Zhao, D.R.; Ma, K.L.; Wu, J.W.; Huang, L.Q. Transcriptome Analysis of Clinopodium chinense (Benth.) O. Kuntze and Identification of Genes Involved in Triterpenoid Saponin Biosynthesis. Int. J. Mol. Sci. 2019, 20, 2643. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuwahara, Y.; Nakajima, D.; Shinpo, S.; Nakamura, M.; Kawano, N.; Kawahara, N.; Yamazaki, M.; Saito, K.; Suzuki, H.; Hirakawa, H. Identification of potential genes involved in triterpenoid saponins biosynthesis in Gleditsia sinensis by transcriptome and metabolome analyses. J. Nat. Med. 2019, 73, 369–380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Y.; Gao, Y.; Chen, Z. Metabolomics analysis of the soapberry (Sapindus mukorossi Gaertn.) pericarp during fruit development and ripening based on UHPLC-HRMS. Sci. Rep. 2021, 11, 11657. [Google Scholar] [CrossRef]
- Yuan, H.J.; Zeng, X.Q.; Shi, J.; Xu, Q.J.; Wang, Y.L.; Jabu, D.Z.; Sang, Z.; Nyima, T. Time-course comparative metabolite profiling under osmotic stress in tolerant and sensitive Tibetian hulless barley. BioMed Res. Int. 2018, 9415409, 1–12. [Google Scholar]
- Zhang, Q.; Wang, L.; Liu, Z.; Zhao, Z.; Zhao, J.; Wang, Z.; Zhou, G.; Liu, P.; Liu, M. Transcriptome and metabolome profiling unveil the mechanisms of Ziziphus jujuba Mill. peel coloration. Food Chem. 2020, 312, 125903. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full length transcriptome assembly from RNA Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [Green Version]
- Yu, R.; Liang, X.; Wei, Z.; Wang, Y.; Luo, X.; Wang, R.; Zhu, X.; Xie, Y.; Karanja, B.; Liu, L. De novo taproot transcriptome sequencing and analysis of major genes involved in sucrose metabolism in Radish (Raphanus sativus L.). Front. Plant Sci. 2016, 7, 585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI BLAST: A New Generation of Protein Database Search Programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [Green Version]
- Qiao, Z.Q.; Liu, S.S.; Zeng, H.J.; Li, Y.X.; Wang, X.Y.; Chen, Y.; Wang, X.M.; Cai, N. Exploring the molecular mechanism underlying the stable purple-red leaf phenotype in lagerstroemia indica cv. ebony embers. Int. J. Mol. Sci. 2019, 20, 5636. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Colin, N.D. RSEM: Accurate transcript quantification from RNA Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [Green Version]
- Wu, Z.; Gao, T.; Zhong, R.; Lin, Z.; Jiang, C.; Ouyang, S.; Zhao, M.; Che, C.; Zhang, J.; Yin, Z. Antihyperlipidaemic effect of triterpenic acid-enriched fraction from Cyclocarya paliurus leaves in hyperlipidaemic rats. Pharm. Biol. 2017, 55(1), 712–721. [Google Scholar] [CrossRef] [Green Version]
- Fang, Z.J.; Shen, S.N.; Wang, J.M.; Wu, Y.J.; Zhou, C.X.; Mo, J.X.; Lin, L.G.; Gan, L.S. Triterpenoids from Cyclocarya paliurus that Enhance Glucose Uptake in 3T3-L1 Adipocyt. Molecules 2019, 24, 187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, K.N.; Jiang, C.H.; Tian, Y.S.; Xiao, N.; Wu, Z.F.; Ma, Y.L.; Lin, Z.; Fang, S.Z.; Shang, X.L.; Liu, K.; et al. Two triterpeniods from Cyclocarya paliurus (Batal) Iljinsk (Juglandaceae) promote glucose uptake in 3T3-L1 adipocytes: The relationship to AMPK activation. Phytomedicine 2015, 22, 837–846. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.M.; Chen, P.; Shang, X.L.; Yang, W.X.; Fang, S.Z. Genotype-Environment Interactions for Tree Growth and Leaf Phytochemical Content of Cyclocarya paliurus (Batal.) Iljinskaja. Forests 2021, 12, 735. [Google Scholar] [CrossRef]
- Tang, M.; Zhao, L.C.; Zhi, Y.H.; Zhang, Q.H. Chemical constituents and pharmacological activities of Cyclocarya paliurus (Batal.)Iljinskaja:research advances. J. Int. Pharm. Res. 2017, 44, 851–859. [Google Scholar]
- Wang, Y.R.; Cui, B.S.; Han, S.W.; Li, S. New dammarane triterpenoid saponins from the leaves of Cyclocarya paliurus. J. Asian Nat. Prod. Res. 2018, 20, 1019–1027. [Google Scholar] [CrossRef]
- Xuan, T.Y.; Tan, J.; Sun, H.H.; Yang, C.; Lv, W.Y.; Zhang, J.H.; Zhang, K.Q.; Nie, Z.Q.; Ye, Z.J.; He, X.A.; et al. Cyclocarioside O-Q, three novel seco-dammarane triterpenoid glycosides from the leaves of Cyclocarya paliurus. Nat. Prod. Res. 2021, 35, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Yue, X.; Shang, X.; Fang, S. Nitrogen Forms Alter Triterpenoid Accumulation and Related Gene Expression in Cyclocarya paliurus (Batalin) Iljinsk. Seedlings For. 2020, 11, 631. [Google Scholar] [CrossRef]
- Augustin, J.M.; Drok, S.; Shinoda, T.; Sanmiya, K.; Nielsen, J.K.; Khakimov, B.; Olsen, C.E.; Hansen, E.H.; Kuzina, V.; Ekstrom, C.T.; et al. UDP glycosyltransferases from the UGT73C subfamily in Barbarea vulgaris catalyze sapogenin 3-O-glucosylation in saponin-mediated insect resistance. Plant Physiol 2012, 160, 1881–1895. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rahimi, S.; Kim, J.; Mijakovic, I.; Jung, K.H.; Choi, G.; Kim, S.C.; Kim, Y.J. Triterpenoid-biosynthetic UDP-glycosyltransferases from plants. Biotech. Adv. 2019, 37, 107394. [Google Scholar] [CrossRef] [PubMed]
- Thimmappa, R.; Geisler, K.; Louveau, T.; O’Maille, P.; Osbourn, A. Triterpene biosynthesis in plants. Annu. Rev. Plant Biol. 2014, 65, 225–257. [Google Scholar] [CrossRef] [PubMed]


| Compounds | A (Ion Abundance) | B (Ion Abundance) | C (Ion Abundance) | CvsB | BvsA | CvsA |
|---|---|---|---|---|---|---|
| Madasiatic acid | 3.64 × 107 | 1.86 × 107 | 8.59 × 106 | 1.1 | 1.0 | 2.1 |
| 3-O-(2-O-acetyl-glucosyl)oleanolic acid | 1.60 × 107 | 1.27 × 107 | 8.06 × 106 | 1.0 | ||
| 2α-hydroxyursolic acid | 4.95 × 106 | 2.53 × 106 | 1.63 × 106 | 1.0 | 1.6 | |
| 11-keto-ursolic acid | 1.46 × 106 | 1.84 × 106 | 1.58 × 106 | |||
| α-amyrenone | 1.21 × 106 | 1.52 × 106 | 1.55 × 106 | |||
| Pomolic acid | 1.34 × 107 | 4.91 × 106 | 1.38 × 106 | 1.8 | 1.5 | 3.3 |
| Betulin | 1.05 × 106 | 1.06 × 106 | 1.17 × 106 | |||
| Cyclocarioside III | 4.77 × 105 | 9.55 × 105 | 8.04 × 105 | −1.0 | ||
| 3,24-dihydroxy-17,21-semiacetal-12(13) oleanolic fruit | 3.35 × 106 | 8.01 × 105 | 5.10 × 105 | 2.1 | 2.7 | |
| 3β,19α-dihydroxyolean-12-en-28-oic acid | 3.73 × 105 | 3.83 × 105 | 4.64 × 105 | 1.8 | ||
| Alphitolic acid | 4.72 × 106 | 1.20 × 106 | 3.37 × 105 | 1.9 | 2.0 | 3.8 |
| Maslinic acid | 4.70 × 106 | 1.25 × 106 | 3.36 × 105 | 1.8 | 1.9 | 3.8 |
| 2-hydroxyoleanolic acid | 4.50 × 106 | 1.10 × 106 | 3.25 × 105 | −1.0 | 2.0 | 3.8 |
| Betulinic acid | 3.79 × 104 | 7.14 × 104 | 1.47 × 105 | −2.0 | ||
| Lupenone | 3.51 × 104 | 4.78 × 104 | 1.27 × 105 | −1.9 | ||
| p-coumaroyleuscaphic acid | 7.96 × 104 | 1.24 × 105 | 1.24 × 105 | |||
| Cyclocarioside I | 2.21 × 104 | 6.08 × 104 | 7.51 × 104 | 1.5 | −1.5 | −1.8 |
| Cyclocaric acid A | 3.67 × 104 | 6.32 × 104 | 7.12 × 104 | −1.0 | ||
| Madecassic acid | 9.26 × 104 | 1.06 × 105 | 6.74 × 104 | −1.4 | ||
| Camaldulenic acid | 2.01 × 105 | 1.11 × 105 | 4.05 × 104 | 1.7 | 2.3 | |
| Taraxerol | 4.55 × 104 | 4.32 × 104 | 2.87 × 104 | |||
| Ursolic acid | 2.18 × 105 | 1.07 × 105 | 2.74 × 104 | 1.0 | 3.0 | |
| Rosamultic acid | 1.86 × 104 | 9.82 × 103 | 2.57 × 104 | |||
| 24,30-dihydroxy-12(13)-enolupinol | 1.93 × 105 | 8.47 × 104 | 2.56 × 104 | 2.6 | 1.2 | 2.9 |
| 3β-[(arabinosyl)oxy]-28-norurs-12,17-dien | 1.88 × 104 | 2.13 × 104 | 2.45 × 104 | |||
| 3β-hydroxy-28-norurs-17,19,21-trien | 2.09 × 104 | 2.61 × 104 | 1.91 × 104 | |||
| oleanolic acid-3-O-glucosyl(1→2)glucoside | 1.16 × 104 | 1.46 × 104 | 1.90 × 104 | |||
| 3,11-dioxo-19α-hydroxyurs-12-en-28-oic acid | 2.81 × 104 | 1.70 × 104 | 1.80 × 104 | |||
| oleanolic acid-3-O-glucuronide | 1.29 × 104 | 1.80 × 104 | 1.14 × 104 | |||
| Terminolic acid | 3.15 × 105 | 5.13 × 104 | 8.71 × 103 | 2.6 | 5.2 | |
| Arjunic acid | 7.44 × 104 | 9.31 × 103 | 6.67 × 103 | 3.0 | 3.5 | |
| Cyclocarioside II | 4.01 × 103 | 6.27 × 103 | 5.99 × 103 | |||
| Tormentic acid | 2.90 × 104 | 9.49 × 103 | 5.50 × 103 | 1.6 | 2.4 | |
| Rutundic acid | 3.29 × 104 | 8.37 × 103 | 4.60 × 103 | 2.0 | 2.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, C.; Fang, S.; Shang, X. Triterpenoids Biosynthesis Regulation for Leaf Coloring of Wheel Wingnut (Cyclocaryapaliurus). Forests 2021, 12, 1733. https://doi.org/10.3390/f12121733
Sun C, Fang S, Shang X. Triterpenoids Biosynthesis Regulation for Leaf Coloring of Wheel Wingnut (Cyclocaryapaliurus). Forests. 2021; 12(12):1733. https://doi.org/10.3390/f12121733
Chicago/Turabian StyleSun, Caowen, Shengzuo Fang, and Xulan Shang. 2021. "Triterpenoids Biosynthesis Regulation for Leaf Coloring of Wheel Wingnut (Cyclocaryapaliurus)" Forests 12, no. 12: 1733. https://doi.org/10.3390/f12121733
APA StyleSun, C., Fang, S., & Shang, X. (2021). Triterpenoids Biosynthesis Regulation for Leaf Coloring of Wheel Wingnut (Cyclocaryapaliurus). Forests, 12(12), 1733. https://doi.org/10.3390/f12121733







