Identification of Prostaglandin Pathway in Dinoflagellates by Transcriptome Data Mining
Abstract
1. Introduction
2. Results
2.1. Dinoflagellate Transcriptomes
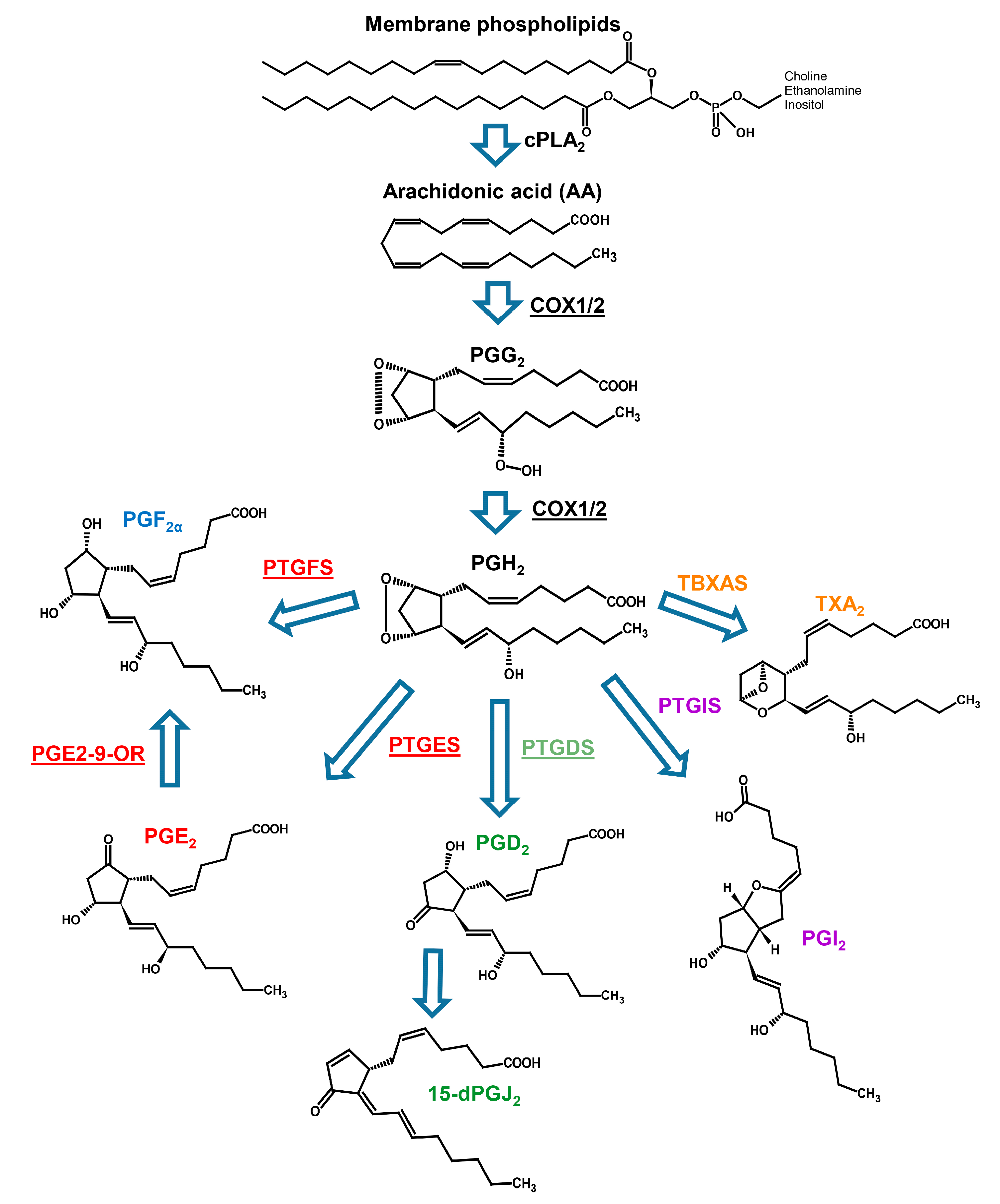
2.2. Prostaglandin Enzyme Identification
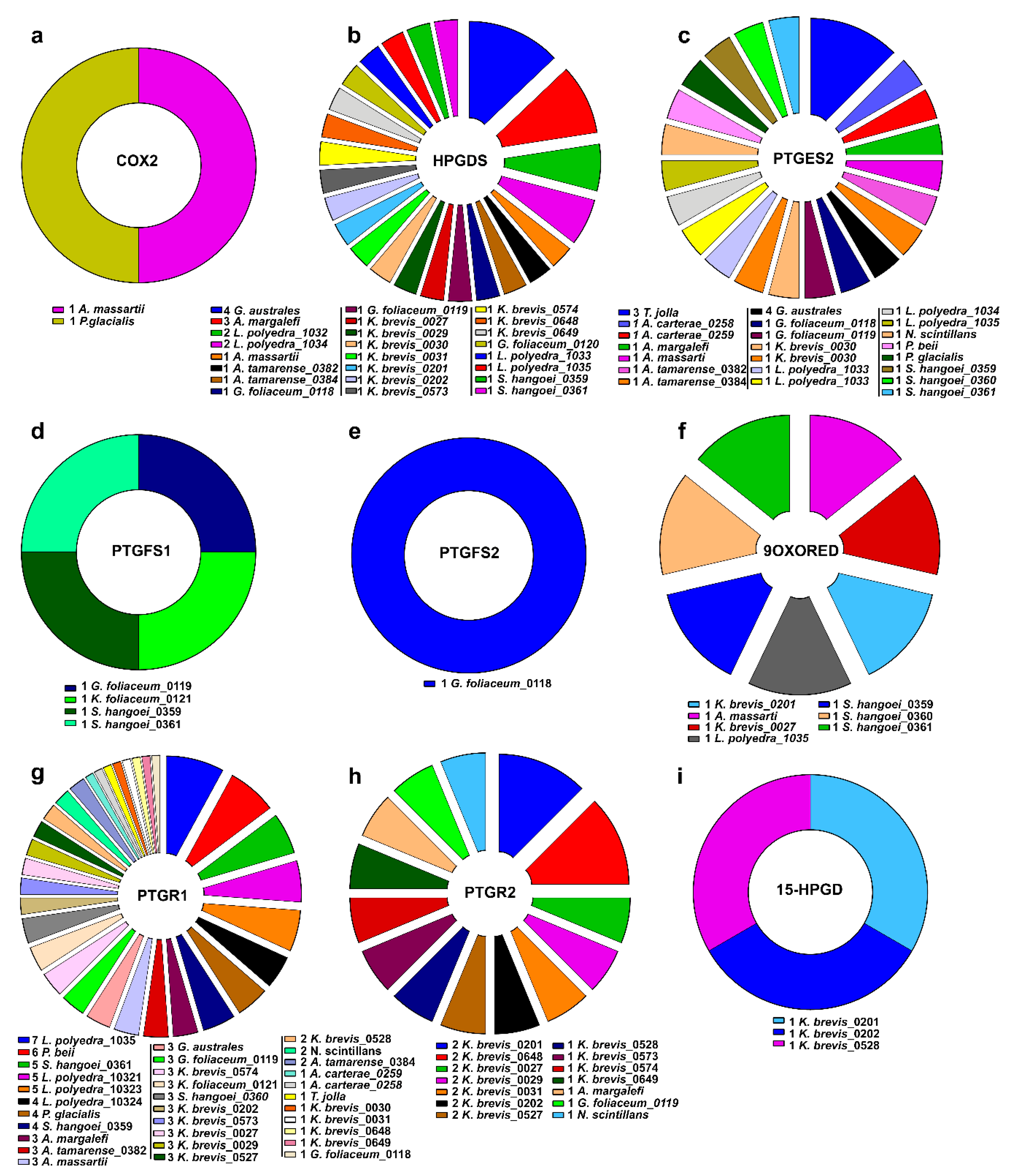
2.3. Clustering of Transcripts Associated to the Pgs-Related Enzymes
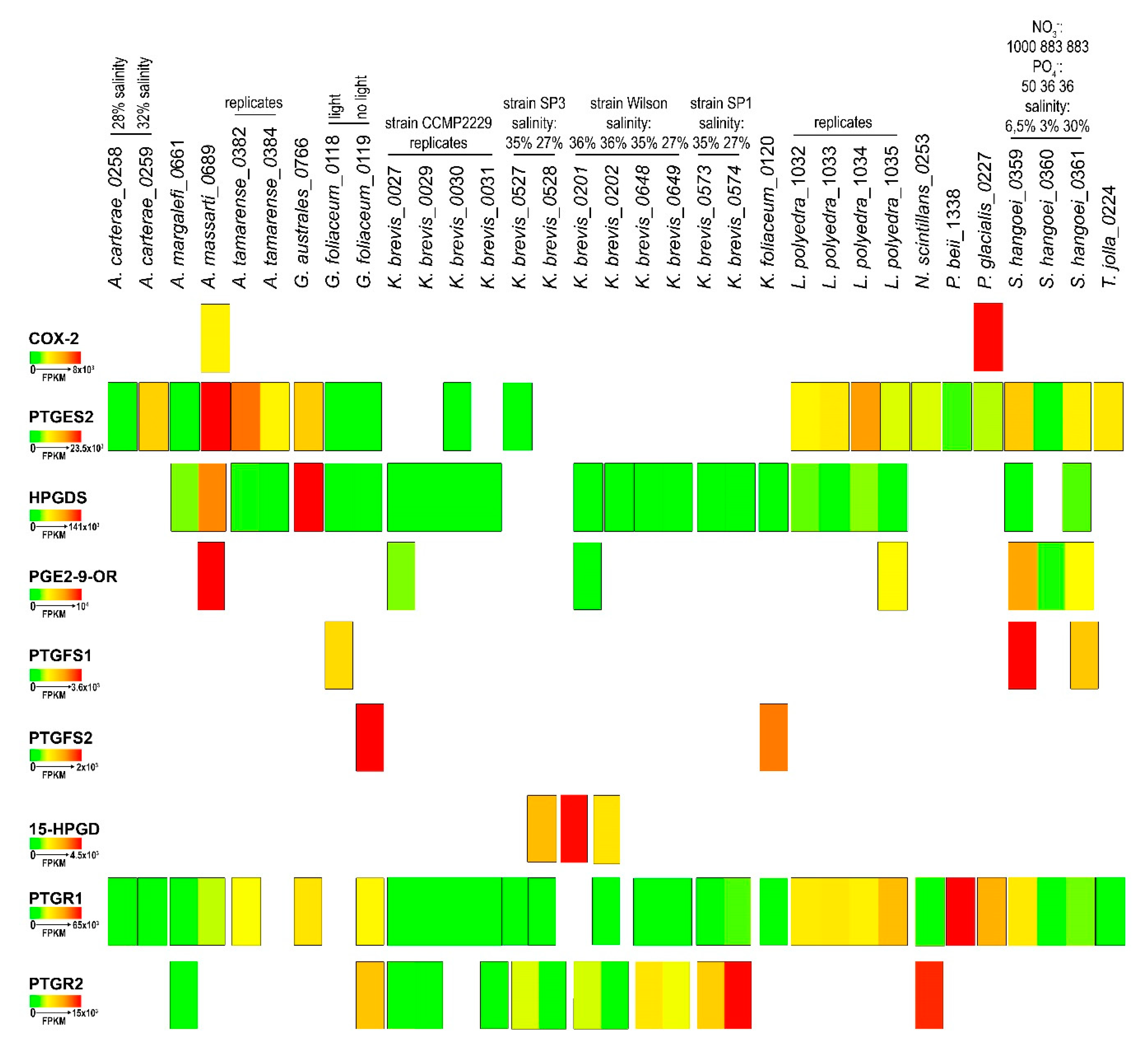
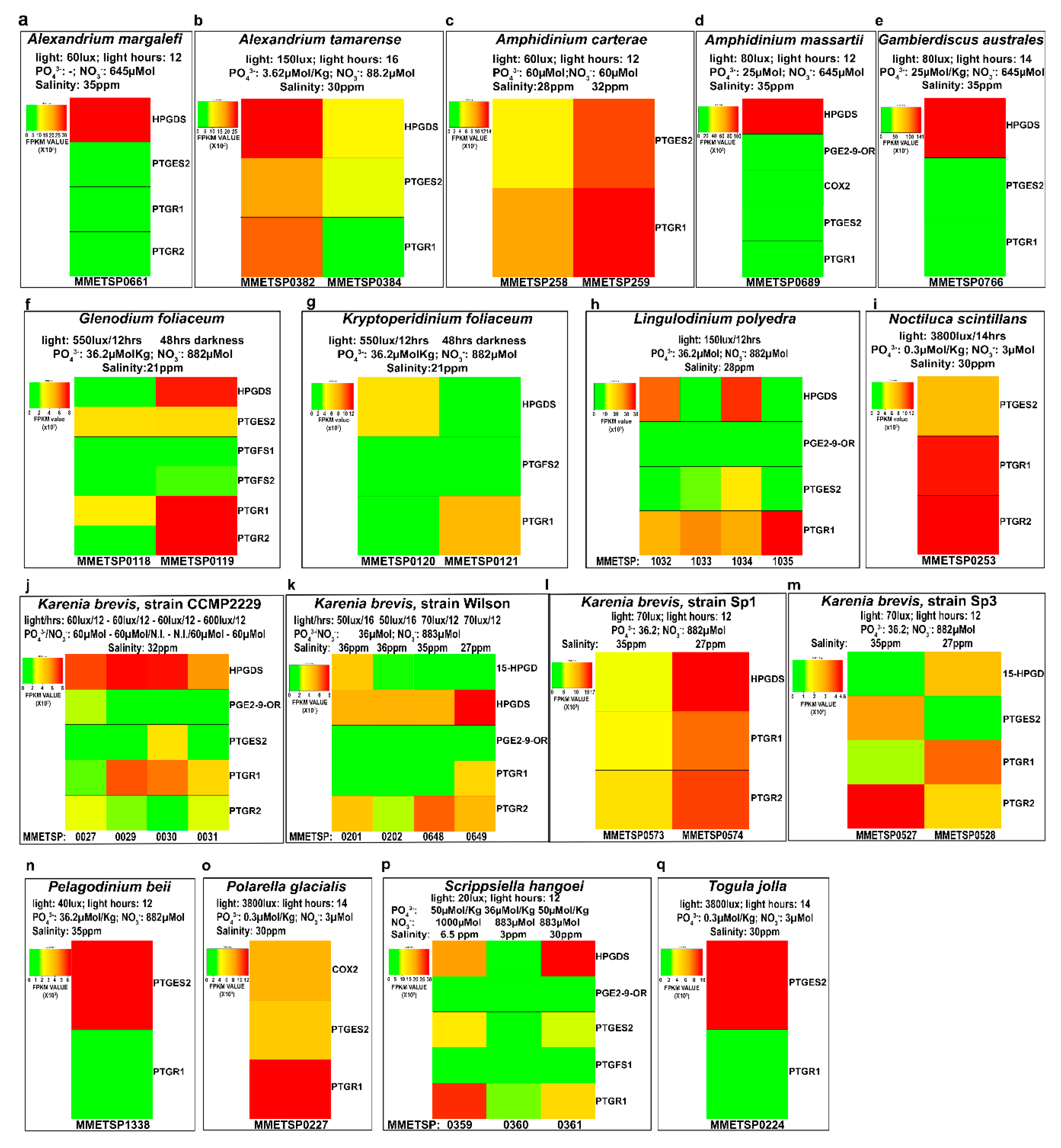
2.4. In-Silico Analysis of Expression Levels
3. Discussion
4. Materials and Methods
4.1. Transcriptomes Collection
4.2. Prostaglandin Enzymes Identification
4.3. Gene Clustering
4.4. Gene Expression Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Wiktorowska-Owczarek, A.; Berezińska, M.; Nowak, J.Z. PUFAs: Structures, Metabolism and Functions. Adv. Clin. Exp. Med. 2015, 24, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Ikawa, M. Algal polyunsaturated fatty acids and effects on plankton ecology and other organisms. UNH Center Freshw. Biol. Res. 2004, 6, 17–44. [Google Scholar]
- Gladyshev, M.I.; Sushchik, N.N.; Makhutova, O.N. Production of EPA and DHA in aquatic ecosystems and their transfer to the land. Prostaglandins Other Lipid Mediat. 2013, 107, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Guschina, I.A.; Harwood, J.L. Lipids and lipid metabolism in eukaryotic algae. Prog. Lipid Res. 2006, 45, 160–186. [Google Scholar] [CrossRef] [PubMed]
- Uttaro, A.D. Biosynthesis of polyunsaturated fatty acids in lower eukaryotes. IUBMB Life 2006, 58, 563–571. [Google Scholar] [CrossRef]
- Andreou, A.; Brodhun, F.; Feussner, I. Biosynthesis of oxylipins in non-mammals. Prog. Lipid Res. 2009, 48, 148–170. [Google Scholar] [CrossRef]
- Peltomaa, E.; Hällfors, H.; Taipale, S.J. Comparison of diatoms and dinoflagellates from different habitats as sources of PUFAs. Mar. Drugs 2019, 17, 233. [Google Scholar] [CrossRef]
- Jónasdóttir, S.H. Fatty acid profiles and production in marine phytoplankton. Mar Drugs 2019, 17, 151. [Google Scholar] [CrossRef]
- Capra, V.; Rovati, G.E.; Mangano, P.; Buccellati, C.; Murphy, R.C.; Sala, A. Transcellular biosynthesis of eicosanoid lipid mediators. BBA—Mol. Cell Biol. Lipids 2015, 1851, 377–382. [Google Scholar] [CrossRef]
- Stonik, V.; Stonik, I. Low-Molecular-Weight metabolites from diatoms: Structures, biological roles and biosynthesis. Mar. Drugs 2015, 13, 3672–3709. [Google Scholar] [CrossRef]
- Clària, J. Cyclooxygenase-2 biology. Curr. Pharm. Des. 2003, 9, 2177–2190. [Google Scholar] [CrossRef] [PubMed]
- Piper, P.J. Introduction to the biosynthesis and metabolism of prostaglandins. Postgrad. Med. J. 1977, 53, 643–646. [Google Scholar] [CrossRef] [PubMed]
- Kawamoto, Y.; Nakamura, Y.; Naito, Y.; Torii, Y.; Kumagai, T.; Osawa, T.; Ohigashi, H.; Satoh, K.; Imagawa, M.; Uchida, K. Cyclopentenone prostaglandins as potential inducers of phase II detoxification enzymes. 15-deoxy-delta(12,14)-prostaglandin j2-induced expression of glutathione S-transferases. J. Biol. Chem. 2000, 275, 11291–11299. [Google Scholar] [CrossRef]
- Wada, M.; DeLong, C.J.; Hong, Y.H.; Rieke, C.J.; Song, I.; Sidhu, R.S.; Yuan, C.; Warnock, M.; Schmaier, A.H.; Yokoyama, C.; et al. Enzymes and receptors of prostaglandin pathways with arachidonic acid-derived versus eicosapentaenoic acid-derived substrates and products. J. Biol. Chem. 2007, 282, 22254–22266. [Google Scholar] [CrossRef]
- Di Costanzo, F.; Di Dato, V.; Ianora, A.; Romano, G. Prostaglandins in marine organisms: A review. Mar. Drugs 2019, 17, 428. [Google Scholar] [CrossRef]
- Di Dato, V.; Orefice, I.; Amato, A.; Fontanarosa, C.; Amoresano, A.; Cutignano, A.; Ianora, A.; Romano, G. Animal-like prostaglandins in marine microalgae. ISME J. 2017, 11, 1722–1726. [Google Scholar] [CrossRef]
- Di Dato, V.; Di Costanzo, F.; Barbarinaldi, R.; Perna, A.; Ianora, A.; Romano, G. Unveiling the presence of biosynthetic pathways for bioactive compounds in the Thalassiosira rotula transcriptome. Sci. Rep. 2019, 9, 9893. [Google Scholar] [CrossRef]
- Yilmaz, E. Biotechnological production of prostaglandin. Biotechnol. Adv. 2001, 19, 387–397. [Google Scholar] [CrossRef]
- Assunção, J.; Guedes, A.C.; Malcata, F.X. Biotechnological and pharmacological applications of biotoxins and other bioactive molecules from dinoflagellates. Mar. Drugs 2017, 15, 393. [Google Scholar] [CrossRef]
- Mendes, A.; Reis, A.; Vasconcelos, R.; Guerra, P.; Lopes da Silva, T. Crypthecodinium cohnii with emphasis on DHA production: A review. J. Appl. Phycol. 2009, 21, 199–214. [Google Scholar] [CrossRef]
- Řezanka, T.; Lukavský, J.; Nedbalová, L.; Sigler, K. Lipidomic profile in three species of dinoflagellates (Amphidinium carterae, Cystodinium sp., and Peridinium aciculiferum) containing very long chain polyunsaturated fatty acids. Phytochemistry 2017, 139, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Beuzenberg, V.; Mountfort, D.; Holland, P.; Shi, F.; MacKenzie, L. Optimization of growth and production of toxins by three dinoflagellates in photobioreactor cultures. J. Appl. Phycol. 2012, 24, 1023–1033. [Google Scholar] [CrossRef]
- Keeling, P.J.; Burki, F.; Wilcox, H.M.; Allam, B.; Allen, E.E.; Amaral-Zettler, L.A.; Armbrust, E.V.; Archibald, J.M.; Bharti, A.K.; Bell, C.J.; et al. The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): Illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 2014, 12, e1001889. [Google Scholar] [CrossRef]
- Tai, H.-H.; Ensor, C.M.; Tong, M.; Zhou, H.; Yan, F. Prostaglandin catabolizing enzymes. Prostaglandins Other Lipid Mediat. 2002, 68–69, 483–493. [Google Scholar] [CrossRef]
- Mesa, J.; Alsina, C.; Oppermann, U.; Parés, X.; Farrés, J.; Porté, S. Human prostaglandin reductase 1 (PGR1): Substrate specificity, inhibitor analysis and site-directed mutagenesis. Chem. Biol. Interact. 2015, 234, 105–113. [Google Scholar] [CrossRef]
- Versteeg, H.H.; van Bergen en Henegouwen, P.M.; van Deventer, S.J.; Peppelenbosch, M.P. Cyclooxygenase-dependent signalling: Molecular events and consequences. FEBS Lett. 1999, 445, 1–5. [Google Scholar] [CrossRef]
- Wang, D.-Z. Neurotoxins from marine dinoflagellates: A brief review. Mar. Drugs 2008, 6, 349–371. [Google Scholar] [CrossRef]
- Waters, A.L.; Hill, R.T.; Place, A.R.; Hamann, M.T. The expanding role of marine microbes in pharmaceutical development. Curr. Opin. Biotechnol. 2010, 21, 780–786. [Google Scholar] [CrossRef]
- Kobayashi, J.; Kubota, T. Bioactive macrolides and polyketides from marine dinoflagellates of the genus Amphidinium. J. Nat. Prod. 2007, 70, 451–460. [Google Scholar] [CrossRef]
- Martínez, K.A.; Lauritano, C.; Druka, D.; Romano, G.; Grohmann, T.; Jaspars, M.; Martín, J.; Díaz, C.; Cautain, B.; de la Cruz, M.; et al. Amphidinol 22, a new cytotoxic and antifungal amphidinol from the dinoflagellate Amphidinium carterae. Mar Drugs 2019, 17, 385. [Google Scholar] [CrossRef]
- Han, R.; Rai, A.; Nakamura, M.; Suzuki, H.; Takahashi, H.; Yamazaki, M.; Saito, K. De novo deep transcriptome analysis of medicinal plants for gene discovery in biosynthesis of plant natural products. In Methods in Enzymology; Elsevier: Amsterdam, The Netherland, 2016; Volume 576, pp. 19–45. ISBN 978-0-12-804539-8. [Google Scholar]
- Wang, D.-Z.; Zhang, S.-F.; Zhang, Y.; Lin, L. Paralytic shellfish toxin biosynthesis in cyanobacteria and dinoflagellates: A molecular overview. J. Proteom. 2016, 135, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Jaeckisch, N.; Yang, I.; Wohlrab, S.; Glöckner, G.; Kroymann, J.; Vogel, H.; Cembella, A.; John, U. Comparative genomic and transcriptomic characterization of the toxigenic marine dinoflagellate Alexandrium ostenfeldii. PLoS ONE 2011, 6, e28012. [Google Scholar] [CrossRef] [PubMed]
- Lauritano, C.; De Luca, D.; Ferrarini, A.; Avanzato, C.; Minio, A.; Esposito, F.; Ianora, A. De novo transcriptome of the cosmopolitan dinoflagellate Amphidinium carterae to identify enzymes with biotechnological potential. Sci. Rep. 2017, 7, 11701. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, S.-F.; Lin, L.; Wang, D.-Z. Whole transcriptomic analysis provides insights into molecular mechanisms for toxin biosynthesis in a toxic dinoflagellate Alexandrium catenella (ACHK-T). Toxins (Basel) 2017, 9, 213. [Google Scholar] [CrossRef]
- Lupette, J.; Jaussaud, A.; Vigor, C.; Oger, C.; Galano, J.-M.; Réversat, G.; Vercauteren, J.; Jouhet, J.; Durand, T.; Maréchal, E. Non-enzymatic synthesis of bioactive isoprostanoids in the diatom Phaeodactylum following oxidative stress. Plant Physiol. 2018, 178, 1344–1357. [Google Scholar] [CrossRef]
| Genus | Species | MMETSP Code | Light | Day Portion in hours | Phosphate µmol/kg | Nitrate µMol | Exp. Salinity (%) | Note | Geo. Area |
|---|---|---|---|---|---|---|---|---|---|
| Akashiwo | sanguinea | 223 | 3800 | 14 | 0.3 | 3 | 30 | New York | |
| Alexandrium | fundyense | 0196C | 200 | 14 | 36.2 | 882 | 28 | Atlantic Ocean | |
| margalefi | 661 | 60 | 12 | 645 | 35 | Tasmania | |||
| minutum | 328 | 100 | 12 | 36.2 | 882 | 25 | Ria da Vigo | ||
| tamarense | 380 | 150 | 16 | 3.62 | 88.2 | 30 | English Channel | ||
| 378 | 150 | 16 | 3.62 | 88.2 | 30 | English Channel | |||
| 384 | 150 | 16 | 3.62 | 88.2 | 30 | English Channel | |||
| 382 | 150 | 16 | 3.62 | 88.2 | 30 | English Channel | |||
| Amoebophrya | - | 795 | 200 | 14 | 36.2 | 882 | 25 | - | |
| Amphidinium | carterae | 258 | 60 | 12 | 60 | 60 | 28 | Massachusetts USA | |
| 259 | 60 | 12 | 60 | N.I. | 32 | Massachusetts USA | |||
| 0398C | 200 | 14 | 36.2 | 882 | 32 | Atlantic Ocean | |||
| massartii | 689 | 80 | 12 | 25 | 645 | 35 | Pacific Ocean | ||
| Gambierdiscus | australes | 766 | 80 | 14 | 25 | 645 | 35 | Pacific Ocean | |
| Glenodinium | foliaceum | 118 | 550 | 12 | 36.2 | 882 | 21 | Antibiotic | Chesapeake Bay |
| 119 | 48hr dark | 36.2 | 882 | 21 | Chesapeake Bay | ||||
| Karenia | brevis | 201 | 50 | 16 | 36 | 883 | 36 | Strain Wilson | Gulf of Mexico |
| 202 | 50 | 16 | 36 | 883 | 36 | Gulf of Mexico | |||
| 648 | 70 | 12 | 36.2 | 882 | 35 | Gulf of Mexico | |||
| 649 | 70 | 12 | 36.2 | 882 | 27 | Gulf of Mexico | |||
| 27 | 60 | 12 | 60 | 60 | 32 | Strain CCMP2229 | |||
| 29 | 60 | 12 | 60 | N.I. | 32 | ||||
| 30 | 60 | 12 | N.I. | 60 | 32 | ||||
| 31 | 600 | 12 | 60 | 60 | 32 | ||||
| 573 | 70 | 12 | 36.2 | 882 | 35 | Strain SP1 | |||
| 574 | 70 | 12 | 36.2 | 882 | 27 | ||||
| 527 | 70 | 12 | 36.2 | 882 | 35 | Strain SP3 | |||
| 528 | 70 | 12 | 36.2 | 882 | 27 | ||||
| Kryptoperidinium | foliaceum | 120 | 550 | 12 | 36.2 | 882 | 21 | Antibiotic | California USA |
| 121 | 48 h dark | 36.2 | 882 | 21 | Antibiotic | California USA | |||
| Lingulodinium | polyedra | 1034 | 150 | 12 | 36.2 | 882 | 28 | Gulf of Mexico | |
| 1033 | 150 | 12 | 36.2 | 882 | 28 | Gulf of Mexico | |||
| 1032 | 150 | 12 | 36.2 | 882 | 28 | Gulf of Mexico | |||
| 1035 | 150 | 12 | 36.2 | 882 | 28 | Gulf of Mexico | |||
| Noctiluca | scintillans | 253 | 3800 | 14 | 0.3 | 3 | 30 | Puget Sound, WA | |
| Pelagodinium | beii | 1338 | 40 | 12 | 36.2 | 882 | 35 | Caribbean Sea | |
| Polarella | glacialis | 227 | 3800 | 14 | 0.3 | 3 | 30 | McMurdo Sound | |
| Pyrocystis | lunula | 229 | 3800 | 14 | 0.3 | 3 | 30 | - | |
| Scrippsiella | hangoei | 359 | 20 | 12 | 50 | 1000 | 6.5 | Selenium 4.55 nMol/L | Baltic Sea |
| 360 | 20 | 12 | 36 | 883 | 3 | Baltic Sea | |||
| 361 | 20 | 12 | 36 | 883 | 30 | Baltic Sea | |||
| Togula | jolla | 224 | 3800 | 14 | 0.3 | 3 | 30 |
| Species | Genus | MMETSP Code | Number of Sequences | Minimum Contig Length | Maximum Contig Length | N50 |
|---|---|---|---|---|---|---|
| Akashiwo | sanguinea | 223 | 380 | 150 | 593 | 197 |
| Alexandrium | fundyense | 0196C | 7872 | 150 | 2586 | 273 |
| margalefi | 661 | 54,023 | 150 | 7976 | 1055 | |
| minutum | 328 | 13,126 | 150 | 2630 | 823 | |
| tamarense | 380 | |||||
| 378 | 44,911 | 150 | 5858 | 701 | ||
| 384 | 106,664 | 150 | 13,283 | 1398 | ||
| 382 | 98,253 | 150 | 9488 | 1506 | ||
| Amoebophrya | - | 795 | 16,699 | 150 | 35,302 | 3178 |
| Amphidinium | carterae | 258 | 44,378 | 150 | 13,731 | 1899 |
| 259 | 45,656 | 150 | 9671 | 1781 | ||
| 0398C | 7775 | 150 | 1411 | 225 | ||
| massartii | 689 | 53,416 | 150 | 11,086 | 1897 | |
| Gambierdiscus | australes | 766 | 53,551 | 150 | 5938 | 1201 |
| Glenodinium | foliaceum | 118 | 80,537 | 150 | 8317 | 1259 |
| 119 | 89,413 | 150 | 51,794 | 950 | ||
| Karenia | brevis | 201 | 89,316 | 150 | 24,712 | 1642 |
| 202 | 77,716 | 150 | 23,631 | 1298 | ||
| 648 | 83,137 | 150 | 28,240 | 1563 | ||
| 649 | 87,365 | 150 | 24,093 | 1566 | ||
| 27 | 87,338 | 150 | 20,088 | 1514 | ||
| 29 | 88,007 | 150 | 17,512 | 1524 | ||
| 30 | 90,601 | 150 | 15,457 | 1493 | ||
| 31 | 79,230 | 150 | 11,278 | 1499 | ||
| 573 | 91,547 | 150 | 24,219 | 1728 | ||
| 574 | 99,942 | 150 | 19,522 | 1792 | ||
| 527 | 81,513 | 150 | 14,879 | 1664 | ||
| 528 | 82,936 | 150 | 18,526 | 1658 | ||
| Kryptoperidinium | foliaceum | 120 | 93,725 | 150 | 13,985 | 1627 |
| 121 | 80,158 | 150 | 5242 | 1190 | ||
| Lingulodinium | polyedra | 1034 | 87,027 | 150 | 9207 | 1260 |
| 1033 | 87,335 | 150 | 15,159 | 1339 | ||
| 1032 | 89,450 | 150 | 11,782 | 1331 | ||
| 1035 | 88,741 | 150 | 22,209 | 1427 | ||
| Noctiluca | scintillans | 253 | 45,249 | 150 | 11,484 | 1594 |
| Pelagodinium | beii | 1338 | 55,559 | 150 | 15,296 | 1545 |
| Polarella | glacialis | 227 | 74,437 | 150 | 23,079 | 1587 |
| Pyrocystis | lunula | 229 | 572 | 150 | 1058 | 198 |
| Scrippsiella | hangoei | 359 | 81,854 | 150 | 16,784 | 1701 |
| 361 | 83,907 | 150 | 33,299 | 1595 | ||
| 360 | 69,908 | 150 | 12,551 | 1281 | ||
| Togula | jolla | 224 | 47,727 | 150 | 8640 | 1570 |
| Enzyme | N° of Species | N° of Transcriptomes |
|---|---|---|
| Cyclooxygenase (COX2) | 2 | 2 |
| Hematopoietic prostaglandin D synthase (HPGDS) | 9 | 18 |
| Prostaglandin E synthase 2 (PTGES2) | 12 | 20 |
| 15-hydroxyprostaglandin dehydrogenase [NAD+] (15-PGDH) | 1 | 2 |
| Prostaglandin-E(2) 9-reductase (PGE2-9-OR) | 4 | 6 |
| Prostaglandin F synthase 1 (PTGFS1) | 3 | 4 |
| Prostaglandin F synthase 2 (PTGFS2) | 1 | 1 |
| Prostaglandin reductase 1 (PTGR1) | 14 | 24 |
| Prostaglandin reductase 2 (PTGR2) | 4 | 7 |
| PTG/HS2 (COX2) | HPGDS | PTGES2 | PGE2-9-OR | 15-PGDH | PTGFS1 | PTGFS2 | PTGR1 | PTGR2 | |
|---|---|---|---|---|---|---|---|---|---|
| A. margalefi | X | X | X | ||||||
| A. tamarense | X | X | X | ||||||
| A. carterae | X | X | |||||||
| A. massartii | X | X | X | X | X | ||||
| G. australes | X | X | X | ||||||
| G. foliaceum | X | X | X | X | X | X | |||
| K. brevis | X | X | X | X | X | X | |||
| K. foliaceum | X | X | X | ||||||
| L polyedra | X | X | X | X | |||||
| N. scintillans | X | X | X | ||||||
| P. beii | X | X | |||||||
| P. glacialis | X | X | X | ||||||
| S. hangoei | X | X | X | X | X | ||||
| T. jolla | X | X |
| Species | MMETSP | 15-HPGD | 9OXORED | HPGDS | PTGES2 | PTGFS1 | PTGFS2 | PTG/HS2 | PTGR1 | PTGR2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SP | CD | SP | CD | SP | CD | SP | CD | SP | CD | SP | CD | SP | CD | SP | CD | SP | CD | ||
| Hit | Hit | Hit | Hit | Hit | Hit | Hit | Hit | Hit | |||||||||||
| A. margalefi | 661 | - | - | 3 | 1 | - | - | - | 4 | 3 | 1 | ||||||||
| A. tamarense | 382 | - | - | 1 | 1 | - | - | - | 4 | 3 | - | ||||||||
| 384 | - | - | 1 | 1 | - | - | - | 2 | - | ||||||||||
| A. carterae | 258 | - | - | - | 2 | 1 | - | - | - | 2 | 1 | - | |||||||
| 259 | - | - | - | 1 | - | - | - | 1 | - | ||||||||||
| A. massartii | 689 | - | 1 | 3 | 1 | 1 | - | - | 2 | 1 | 3 | - | |||||||
| G. australes | 766 | - | - | 4 | 1 | - | - | - | 3 | - | |||||||||
| G. foliaceum | 118 | - | - | 1 | 1 | - | 1 | - | 1 | - | |||||||||
| 119 | - | - | 1 | 2 | 1 | 1 | - | - | 3 | 1 | |||||||||
| K. brevis | 201 | 1 | 1 | 1 | - | - | - | - | - | 3 | 2 | ||||||||
| 202 | 1 | - | 1 | - | - | - | - | 2 | 1 | ||||||||||
| 648 | - | - | 2 | 1 | - | - | - | - | 1 | 2 | |||||||||
| 649 | - | - | 1 | - | - | - | 1 | 1 | |||||||||||
| 573 | - | - | 1 | - | - | - | - | 2 | 1 | ||||||||||
| 574 | - | - | 1 | - | - | - | - | 4 | 3 | 1 | |||||||||
| 27 | - | 1 | 1 | - | - | - | - | 2 | 1 | ||||||||||
| 29 | - | - | 1 | - | - | - | - | 3 | 2 | 2 | 1 | ||||||||
| 30 | - | - | 1 | 1 | - | - | - | 1 | - | ||||||||||
| 31 | - | - | 1 | - | - | - | - | 1 | 1 | ||||||||||
| 527 | - | - | - | 1 | - | - | - | 2 | 2 | 1 | |||||||||
| 528 | 1 | - | - | - | - | - | - | 2 | 1 | ||||||||||
| K. foliaceum | 120 | - | - | 1 | - | - | - | - | - | ||||||||||
| 121 | - | - | - | - | 1 | - | - | 3 | - | ||||||||||
| L. polyedra | 1032 | - | - | 4 | 2 | 3 | 1 | - | - | - | 8 | 5 | - | ||||||
| 1033 | - | - | 1 | 2 | 1 | - | - | - | 6 | 5 | - | ||||||||
| 1034 | - | - | 3 | 2 | 2 | 1 | - | - | - | 5 | 4 | - | |||||||
| 1035 | - | 1 | 1 | 3 | 1 | - | - | - | 10 | 7 | - | ||||||||
| N. scintillans | 253 | - | - | - | 1 | - | - | - | 2 | 1 | |||||||||
| P. beii | 1338 | - | - | - | 1 | - | - | - | 6 | - | |||||||||
| P. glacialis | 227 | - | - | - | 1 | - | - | 4 | 1 | 6 | 4 | - | |||||||
| S. hangoei | 359 | - | 1 | 1 | 1 | 1 | - | - | 4 | - | |||||||||
| 360 | - | 1 | - | 1 | - | - | - | 3 | - | ||||||||||
| 361 | - | 1 | 1 | 1 | 1 | - | - | 5 | - | ||||||||||
| T. jolla | 224 | - | - | - | 3 | - | - | - | 2 | 1 | - | ||||||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Dato, V.; Ianora, A.; Romano, G. Identification of Prostaglandin Pathway in Dinoflagellates by Transcriptome Data Mining. Mar. Drugs 2020, 18, 109. https://doi.org/10.3390/md18020109
Di Dato V, Ianora A, Romano G. Identification of Prostaglandin Pathway in Dinoflagellates by Transcriptome Data Mining. Marine Drugs. 2020; 18(2):109. https://doi.org/10.3390/md18020109
Chicago/Turabian StyleDi Dato, Valeria, Adrianna Ianora, and Giovanna Romano. 2020. "Identification of Prostaglandin Pathway in Dinoflagellates by Transcriptome Data Mining" Marine Drugs 18, no. 2: 109. https://doi.org/10.3390/md18020109
APA StyleDi Dato, V., Ianora, A., & Romano, G. (2020). Identification of Prostaglandin Pathway in Dinoflagellates by Transcriptome Data Mining. Marine Drugs, 18(2), 109. https://doi.org/10.3390/md18020109







