A Quorum-Sensing Inhibitor Strain of Vibrio alginolyticus Blocks Qs-Controlled Phenotypes in Chromobacterium violaceum and Pseudomonas aeruginosa
Abstract
1. Introduction
2. Results
2.1. Isolation and Taxonomical Identification of QSI-producing Bacteria from Anemones and Holothurians
2.2. Effect of Cell Pellet Extract from Strain M3-10 on C. Violaceum ATCC 12472 Violacein Production, Pyoverdine Production, Biofilm Formation, and Pseudomonas aeruginosa PAO1 Motility
2.3. Identification of QSI Compounds from V. alginolyticus M3-10
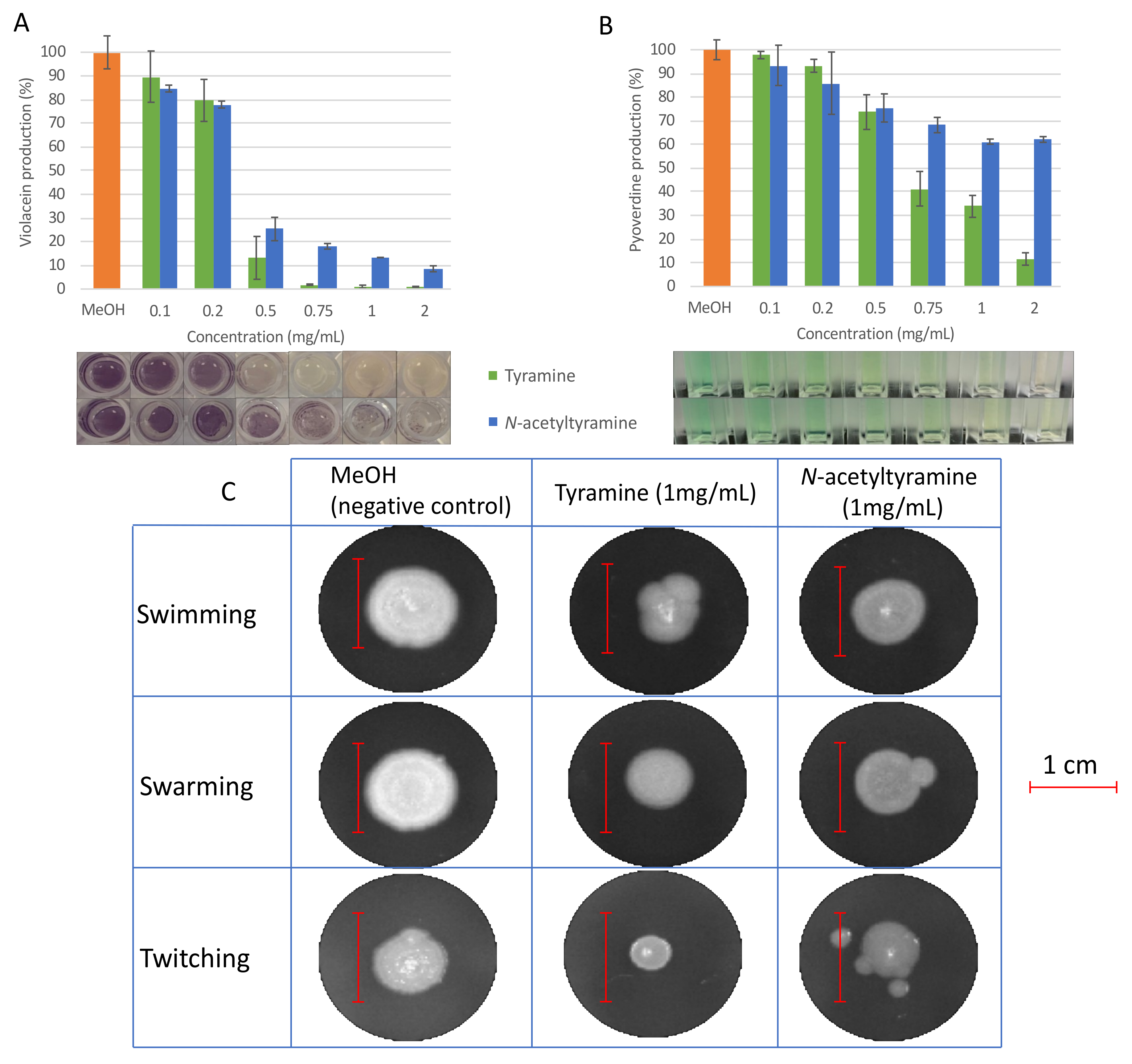
2.4. Evaluation of QSI Activity of Tyramine and N-acetyltyramine Standards
2.5. Whole-Genome Analysis of V. alginolyticus M3-10
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Media, Compounds, and Culture Conditions
4.2. Screening for QS Inhibition Activity
4.3. Preparation of Methanolic Extracts of QSI-producing Bacteria
4.4. Violacein Production Assay
4.5. Pseudomonas Virulence Factor Tests: Pyoverdine, Biofilm, and Motility
4.6. Bioassay-Guided Fractionation and Identification of QSI Compounds by LC–DAD–HRMS and NMR
4.7. QSI inhibition Activity Test of Tyramine and N-acetyltyramine
4.8. Taxonomic Identification and Genome Analysis of Strain M3-10
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Prioritization of Pathogens to Guide Discovery, Research and Development of New Antibiotics for Drug Resistant Bacterial Infections, Including Tuberculosis; World Health Organization: Geneva, Switzerland, 2017. [Google Scholar]
- Garg, N.; Manchanda, G.; Kumar, A. Bacterial quorum sensing: Circuits and applications. Antonie Van Leeuwenhoek 2014, 105, 289–305. [Google Scholar] [CrossRef] [PubMed]
- Jayaraman, A.; Wood, T.K. Bacterial Quorum Sensing: Signals, Circuits, and Implications for Biofilms and Disease. Annu. Rev. Biomed. Eng. 2008, 10, 145–167. [Google Scholar] [CrossRef]
- Whitehead, N.A.; Barnard, A.M.L.; Slater, H.; Simpson, N.J.L.; Salmond, G.P.C. Quorum-sensing in Gram-negative bacteria. FEMS Microbiol. Rev. 2001, 25, 365–404. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Bassler, B.L. Quorum sensing signal-response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576–588. [Google Scholar] [CrossRef]
- Quiñones, B.; Dulla, G.; Lindow, S.E. Exopolysaccharide Production, Motility, and Virulence in Pseudomonas syringae. Society 2005, 18, 682–693. [Google Scholar]
- Whiteley, M.; Diggle, S.P.; Greenberg, E.P.; Wilson, E.O. Bacterial quorum sensing: The progress and promise of an emerging research area. Nature 2017, 551, 313–320. [Google Scholar] [CrossRef]
- Fuqua, C.; Winans, S.C.; Greenberg, E.P. Quorum sensing in bacteria: The LuxR-LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 1994, 176, 269–275. [Google Scholar] [CrossRef]
- Fuqua, C.; Parsek, M.R.; Greenberg, E.P. Regulation of Gene Expression by Cell-to-Cell Communication: Acyl-Homoserine Lactone Quorum Sensing. Annu. Rev. Genet. 2001, 35, 439–468. [Google Scholar] [CrossRef]
- Ng, W.-L.; Bassler, B.L. Bacterial quorum-sensing network architectures. Annu. Rev. Genet. 2009, 43, 197–222. [Google Scholar] [CrossRef]
- Parker, C.T.; Sperandio, V. Cell-to-cell signalling during pathogenesis. Cell. Microbiol. 2009, 11, 363–369. [Google Scholar] [CrossRef]
- Grandclément, C.; Tannières, M.; Moréra, S.; Dessaux, Y.; Faure, D. Quorum quenching: Role in nature and applied developments. FEMS Microbiol. Rev. 2016, 40, 86–116. [Google Scholar] [CrossRef]
- Natrah, F.M.I.; Defoirdt, T.; Sorgeloos, P.; Bossier, P. Disruption of Bacterial Cell-to-Cell Communication by Marine Organisms and its Relevance to Aquaculture. Mar. Biotechnol. 2011, 13, 109–126. [Google Scholar] [CrossRef]
- LaSarre, B.; Federle, M.J. Exploiting Quorum Sensing To Confuse Bacterial Pathogens. Microbiol. Mol. Biol. Rev. 2013, 77, 73–111. [Google Scholar] [CrossRef]
- Defoirdt, T. Quorum-Sensing Systems as Targets for Antivirulence Therapy. Trends Microbiol. 2018, 26, 313–328. [Google Scholar] [CrossRef]
- Ni, N.; Li, M.; Wang, J.; Wang, B. Inhibitors and antagonists of bacterial quorum sensing. Med. Res. Rev. 2009, 29, 65–124. [Google Scholar] [CrossRef]
- Zhao, J.; Li, X.; Hou, X.; Quan, C.; Chen, M. Widespread Existence of Quorum Sensing Inhibitors in Marine Bacteria: Potential Drugs to Combat Pathogens with Novel Strategies. Mar. Drugs 2019, 17, 275. [Google Scholar] [CrossRef]
- Kalia, V.C.; Purohit, H.J. Quenching the quorum sensing system: Potential antibacterial drug targets. Crit. Rev. Microbiol. 2011, 37, 121–140. [Google Scholar] [CrossRef]
- Kalia, V.C.; Patel, S.K.S.; Kang, Y.C.; Lee, J.K. Quorum sensing inhibitors as antipathogens: Biotechnological applications. Biotechnol. Adv. 2019, 37, 68–90. [Google Scholar] [CrossRef]
- Sommer, R.; Joachim, I.; Wagner, S.; Titz, A. New Approaches to Control Infections: Anti-biofilm Strategies against Gram-negative Bacteria. Chim. Int. J. Chem. 2013, 67, 286–290. [Google Scholar] [CrossRef]
- Bhardwaj, A.K.; Vinothkumar, K.; Rajpara, N. Bacterial Quorum Sensing Inhibitors: Attractive Alternatives for Control of Infectious Pathogens Showing Multiple Drug Resistance. Recent Pat. Anti-Infect. Drug Discov. 2013, 8, 68–83. [Google Scholar] [CrossRef]
- Givskov, M.; De Nys, R.; Manefield, M.; Gram, L.; Maximilien, R.; Eberl, L.; Molin, S.; Steinberg, P.D.; Kjelleberg, S. Eukaryotic interference with homoserine lactone-mediated prokaryotic signalling. J. Bacteriol. 1996, 178, 6618–6622. [Google Scholar] [CrossRef]
- Fetzner, S. Quorum quenching enzymes. J. Biotechnol. 2014, 201, 2–14. [Google Scholar] [CrossRef]
- Adonizio, A.; Kong, K.-F.; Mathee, K. Inhibition of quorum sensing-controlled virulence factor production in Pseudomonas aeruginosa by South Florida plant extracts. Antimicrob. Agents Chemother. 2007, 52, 198–203. [Google Scholar] [CrossRef]
- Bodini, S.F.; Manfredini, S.; Epp, M.; Valentini, S.; Santori, F. Quorum sensing inhibition activity of garlic extract and p-coumaric acid. Lett. Appl. Microbiol. 2009, 49, 551–555. [Google Scholar] [CrossRef]
- Choo, J.H.; Rukayadi, Y.; Hwang, J.K. Inhibition of bacterial quorum sensing by vanilla extract. Lett. Appl. Microbiol. 2006, 42, 637–641. [Google Scholar] [CrossRef]
- Tolmacheva, A.A.; Rogozhin, E.A.; Deryabin, D.G. Antibacterial and quorum sensing regulatory activities of some traditional Eastern-European medicinal plants. Acta Pharm. 2014, 64, 173–186. [Google Scholar] [CrossRef]
- Zhu, H.; He, C.-C.; Chu, Q.-H. Inhibition of quorum sensing in Chromobacterium violaceum by pigments extracted from Auricularia auricular. Lett. Appl. Microbiol. 2011, 52, 269–274. [Google Scholar] [CrossRef]
- Zhu, H.; Sun, S.J. Inhibition of bacterial quorum sensing-regulated behaviors by Tremella fuciformis extract. Curr. Microbiol. 2008, 57, 418–422. [Google Scholar] [CrossRef]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef]
- Singh, V.K.; Mishra, A.; Jha, B. Anti-quorum Sensing and Anti-biofilm Activity of Delftia tsuruhatensis Extract by Attenuating the Quorum Sensing-Controlled Virulence Factor Production in Pseudomonas aeruginosa. Front. Cell. Infect. Microbiol. 2017, 7, 337. [Google Scholar] [CrossRef]
- Sun, S.; Dai, X.; Sun, J.; Bu, X.; Weng, C.; Li, H.; Zhu, H. A diketopiperazine factor from Rheinheimera aquimaris QSI02 exhibits anti-quorum sensing activity. Sci. Rep. 2016, 6, 39637. [Google Scholar] [CrossRef]
- Dobretsov, S.; Teplitski, M.; Paul, V. Mini-review: Quorum sensing in the marine environment and its relationship to biofouling. Biofouling 2009, 25, 413–427. [Google Scholar] [CrossRef]
- Teasdale, M.E.; Donovan, K.A.; Forschner-Dancause, S.R.; Rowley, D.C. Gram-Positive Marine Bacteria as a Potential Resource for the Discovery of Quorum Sensing Inhibitors. Mar. Biotechnol. 2011, 13, 722–732. [Google Scholar] [CrossRef]
- Blunt, J.W.; Copp, B.R.; Keyzers, R.A.; Munro, M.H.G.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2015, 32, 116–211. [Google Scholar] [CrossRef]
- Yaniv, K.; Golberg, K.; Kramarsky-Winter, E.; Marks, R.; Pushkarev, A.; Béjà, O.; Kushmaro, A. Functional marine metagenomic screening for anti-quorum sensing and anti-biofilm activity. Biofouling 2017, 33, 1–13. [Google Scholar] [CrossRef]
- Torres, M.; Dessaux, Y.; Llamas, I. Saline Environments as a Source of Potential Quorum Sensing Disruptors to Control Bacterial Infections: A Review. Mar. Drugs 2019, 17, 191. [Google Scholar] [CrossRef]
- Bhatnagar, I.; Kim, S.K. Immense essence of excellence: Marine microbial bioactive compounds. Mar. Drugs 2010, 8, 2673–2701. [Google Scholar] [CrossRef]
- Rosenberg, E.; Koren, O.; Reshef, L.; Efrony, R.; Zilber-Rosenberg, I. The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microbiol. 2007, 5, 355–362. [Google Scholar] [CrossRef]
- Valliappan, K.; Sun, W.; Li, Z. Marine actinobacteria associated with marine organisms and their potentials in producing pharmaceutical natural products. Appl. Microbiol. Biotechnol. 2014, 98, 7365–7377. [Google Scholar] [CrossRef]
- Ma, Z.-P.; Song, Y.; Cai, Z.-H.; Lin, Z.-J.; Lin, G.-H.; Wang, Y.; Zhou, J. Anti-quorum Sensing Activities of Selected Coral Symbiotic Bacterial Extracts From the South China Sea. Front. Cell. Infect. Microbiol. 2018, 8, 144. [Google Scholar] [CrossRef]
- León-Palmero, E.; Joglar, V.; Álvarez, P.A.; Martín-Platero, A.; Llamas, I.; Reche, I. Diversity and antimicrobial potential in sea anemone and holothurian microbiomes. PLoS ONE 2018, 13, e0196178. [Google Scholar] [CrossRef]
- Pérez-Victoria, I.; Martín, J.; Reyes, F. Combined LC/UV/MS and NMR Strategies for the Dereplication of Marine Natural Products. Planta Med. 2016, 82, 857–871. [Google Scholar] [CrossRef]
- Martín, J.; Crespo, G.; González-Menéndez, V.; Pérez-Moreno, G.; Sánchez-Carrasco, P.; Pérez-Victoria, I.; Ruiz-Pérez, L.M.; González-Pacanowska, D.; Vicente, F.; Genilloud, O.; et al. MDN-0104, an Antiplasmodial Betaine Lipid from Heterospora chenopodii. J. Nat. Prod. 2014, 77, 2118–2123. [Google Scholar] [CrossRef]
- Gutiérrez, M.; Capson, T.L.; Guzmán, H.M.; González, J.; Ortega-Barría, E.; Quiñoá, E.; Riguera, R. Antiplasmodial metabolites isolated from the marine octocoral Muricea austera. J. Nat. Prod. 2006, 69, 1379–1383. [Google Scholar] [CrossRef]
- Sun, J.F.; Wu, Y.; Yang, B.; Liu, Y. Chemical Constituents of Marine Sponge Halichondria sp. from South China Sea. Chem. Nat. Compd. 2015, 51, 975–977. [Google Scholar] [CrossRef]
- Rustamova, N.; Bobakulov, K.; Begmatov, N.; Turak, A.; Yili, A.; Aisa, H.A. Secondary metabolites produced by endophytic Pantoea ananatis derived from roots of Baccharoides anthelmintica and their effect on melanin synthesis in murine B16 cells. Nat. Prod. Res. 2019, 1–6. [Google Scholar] [CrossRef]
- Ye, X.; Chai, W.; Lian, X.-Y.; Zhang, Z. Novel propanamide analogue and antiproliferative diketopiperazines from mangrove Streptomyces sp. Q24. Nat. Prod. Res. 2017, 31, 1390–1396. [Google Scholar] [CrossRef]
- Richter, M.; Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA 2009, 106, 19126–19131. [Google Scholar] [CrossRef]
- Dong, Y.H.; Xu, J.L.; Li, X.Z.; Zhang, L.H. AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc. Natl. Acad. Sci. USA 2000, 97, 3526–3531. [Google Scholar] [CrossRef]
- Hraiech, S.; Hiblot, J.; Lafleur, J.; Lepidi, H.; Papazian, L.; Rolain, J.M.; Raoult, D.; Elias, M.; Silby, M.W.; Bzdrenga, J.; et al. Inhaled lactonase reduces Pseudomonas aeruginosa quorum sensing and mortality in rat pneumonia. PLoS ONE 2014, 9, e107125. [Google Scholar] [CrossRef]
- Migiyama, Y.; Kaneko, Y.; Yanagihara, K.; Morohoshi, T.; Morinaga, Y.; Nakamura, S.; Miyazaki, T.; Hasegawa, H.; Izumikawa, K.; Kakeya, H.; et al. Efficacy of AiiM, an N -Acylhomoserine Lactonase, against Pseudomonas aeruginosa in a Mouse Model of Acute Pneumonia. Antimicrob. Agents Chemother. 2013, 57, 3653–3658. [Google Scholar] [CrossRef]
- Chen, R.; Zhou, Z.; Cao, Y.; Bai, Y.; Yao, B. High yield expression of an AHL-lactonase from Bacillus sp. B546 in Pichia pastoris and its application to reduce Aeromonas hydrophila mortality in aquaculture. Microb. Cell Fact. 2010, 9, 39. [Google Scholar] [CrossRef]
- Nhan, D.T.; Cam, D.T.V.; Wille, M.; Defoirdt, T.; Bossier, P.; Sorgeloos, P. Quorum quenching bacteria protect Macrobrachium rosenbergii larvae from Vibrio harveyi infection. J. Appl. Microbiol. 2010, 109, 1007–1016. [Google Scholar] [CrossRef]
- Reina, J.C.; Torres, M.; Llamas, I. Stenotrophomonas maltophilia AHL-degrading strains isolated from marine invertebrate microbiota attenuate the virulence of Pectobacterium carotovorum and Vibrio corallilyticus. Mar. Biotechnol. 2019, 21, 276–290. [Google Scholar] [CrossRef]
- Torres, M.; Romero, M.; Prado, S.; Dubert, J.; Tahrioui, A.; Otero, A.; Llamas, I. N-acylhomoserine lactone-degrading bacteria isolated from hatchery bivalve larval cultures. Microbiol. Res. 2013, 168, 547–554. [Google Scholar] [CrossRef]
- Torres, M.; Rubio-Portillo, E.; Antón, J.; Ramos-Esplá, A.A.; Quesada, E.; Llamas, I. Selection of the N-acylhomoserine lactone-degrading bacterium Alteromonas stellipolaris PQQ-42 and of its potential for biocontrol in aquaculture. Front. Microbiol. 2016, 7, 646. [Google Scholar] [CrossRef]
- Ban, H.; Chai, X.; Lin, Y.; Zhou, Y.; Peng, D.; Zhou, Y.; Zou, Y.; Yu, Z.; Sun, M. Transgenic Amorphophallus konjac expressing synthesized acyl-homoserine lactonase (aiiA) gene exhibit enhanced resistance to soft rot disease. Plant Cell Rep. 2009, 28, 1847–1855. [Google Scholar] [CrossRef]
- Molina, L.; Constantinescu, F.; Michel, L.; Reimmann, C.; Duffy, B.; Défago, G. Degradation of pathogen quorum-sensing molecules by soil bacteria: A preventive and curative biological control mechanism. FEMS Microbiol. Ecol. 2003, 45, 71–81. [Google Scholar] [CrossRef]
- Song, Y.; Cai, Z.H.; Lao, Y.M.; Jin, H.; Ying, K.Z.; Lin, G.H.; Zhou, J. Antibiofilm activity substances derived from coral symbiotic bacterial extract inhibit biofouling by the model strain Pseudomonas aeruginosa PAO1. Microb. Biotechnol. 2018, 11, 1090–1105. [Google Scholar] [CrossRef]
- Tello, E.; Castellanos, L.; Duque, C. Disruption in Quorum-Sensing Systems and Bacterial Biofilm Inhibition by Cembranoid Diterpenes Isolated from the Octocoral Eunicea knighti. J. Nat. Prod. 2012, 75, 1637–1642. [Google Scholar] [CrossRef]
- Saurav, K.; Costantino, V.; Venturi, V.; Steindler, L. Quorum sensing inhibitors from the sea discovered using bacterial N-acyl-homoserine lactone-based biosensors. Mar. Drugs 2017, 15, 53. [Google Scholar] [CrossRef]
- Chen, J.; Wang, B.; Lu, Y.; Guo, Y.; Sun, J.; Wei, B.; Zhang, H.; Wang, H. Quorum Sensing Inhibitors from Marine Microorganisms and Their Synthetic Derivatives. Mar. Drugs 2019, 17, 80. [Google Scholar] [CrossRef]
- Rasch, M.; Buch, C.; Austin, B.; Slierendrecht, W.J.; Ekmann, K.S.; Larsen, J.L.; Johansen, C.; Riedel, K.; Eberl, L.; Givskov, M.; et al. An inhibitor of bacterial quorum sensing reduces mortalities caused by vibriosis in rainbow trout (Oncorhynichus mykiss, Walbaum). Syst. Appl. Microbiol. 2004, 27, 350–359. [Google Scholar] [CrossRef]
- Skindersoe, M.E.; Ettinger-Epstein, P.; Rasmussen, T.B.; Bjarnsholt, T.; de Nys, R.; Givskov, M. Quorum Sensing Antagonism from Marine Organisms. Mar. Biotechnol. 2008, 10, 56–63. [Google Scholar] [CrossRef]
- Saurav, K.; Bar-Shalom, R.; Haber, M.; Burgsdorf, I.; Oliviero, G.; Costantino, V.; Morgenstern, D.; Steindler, L. In search of alternative antibiotic drugs: Quorum-quenching activity in sponges and their bacterial isolates. Front. Microbiol. 2016, 7, 1–18. [Google Scholar] [CrossRef]
- Cunsolo, V.; Cusimano, M.; Vazzana, M.; Russo, D.; Arizza, V.; Vitale, M.; Schillaci, D.; Saletti, R. Immune mediators of sea-cucumber Holothuria tubulosa (Echinodermata) as source of novel antimicrobial and anti-staphylococcal biofilm agents. AMB Express 2013, 3, 35. [Google Scholar]
- Cusimano, M.; Spinello, A.; Barone, G.; Schillaci, D.; Cascioferro, S.; Magistrato, A.; Parrino, B.; Arizza, V.; Vitale, M. A Synthetic Derivative of Antimicrobial Peptide Holothuroidin 2 from Mediterranean Sea Cucumber (Holothuria tubulosa) in the Control of Listeria monocytogenes. Mar. Drugs 2019, 17, 159. [Google Scholar] [CrossRef]
- Golberg, K.; Pavlov, V.; Marks, R.S.; Kushmaro, A. Coral-associated bacteria, quorum sensing disrupters, and the regulation of biofouling. Biofouling 2013, 29, 669–682. [Google Scholar] [CrossRef]
- Tait, K.; Hutchison, Z.; Thompson, F.L.; Munn, C.B. Quorum sensing signal production and inhibition by coral-associated vibrios. Environ. Microbiol. Rep. 2010, 2, 145–150. [Google Scholar] [CrossRef]
- Nathwani, D.; Raman, G.; Sulham, K.; Gavaghan, M.; Menon, V. Clinical and economic consequences of hospital-acquired resistant and multidrug-resistant Pseudomonas aeruginosa infections: A systematic review and meta-analysis. Antimicrob. Resist. Infect. Control 2014, 3, 32. [Google Scholar] [CrossRef]
- Soukarieh, F.; Williams, P.; Stocks, M.J.; Camara, M. Pseudomonas aeruginosa Quorum Sensing Systems as Drug Discovery Targets: Current Position and Future Perspectives. J. Med. Chem. 2018, 23, 10385–10402. [Google Scholar] [CrossRef]
- Wagner, S.; Sommer, R.; Hinsberger, S.; Lu, C.; Hartmann, R.W.; Empting, M.; Titz, A. Novel Strategies for the Treatment of Pseudomonas aeruginosa Infections. J. Med. Chem. 2016, 59, 5929–5969. [Google Scholar] [CrossRef]
- Zhou, J.W.; Luo, H.Z.; Jiang, H.; Jian, T.K.; Chen, Z.Q.; Jia, A.Q. Hordenine: A Novel Quorum Sensing Inhibitor and Antibiofilm Agent against Pseudomonas aeruginosa. J. Agric. Food Chem. 2018, 66, 1620–1628. [Google Scholar] [CrossRef]
- Zhou, J.W.; Ruan, L.Y.; Chen, H.J.; Luo, H.Z.; Jiang, H.; Wang, J.S.; Jia, A.Q. Inhibition of Quorum Sensing and Virulence in Serratia marcescens by Hordenine. J. Agric. Food Chem. 2019, 67, 784–795. [Google Scholar] [CrossRef]
- Teasdale, M.E.; Liu, J.; Wallace, J.; Akhlaghi, F.; Rowley, D.C. Secondary metabolites produced by the marine bacterium halobacillus salinus that Inhibit quorum sensing-controlled phenotypes in gram-negative bacteria. Appl. Environ. Microbiol. 2009, 75, 567–572. [Google Scholar] [CrossRef]
- Martínez-Matamoros, D.; Laiton-Fonseca, M.; Duque, C.; Ramos, F.A.; Castellanos, L. Búsqueda de bacterias marinas como fuente de inhibidores quorum sensing. Vitae 2016, 23, 30–47. [Google Scholar] [CrossRef]
- Prester, L. Biogenic amines in fish, fish products and shellfish: A review. Food Addit. Contam. 2011, 28, 1547–1560. [Google Scholar] [CrossRef]
- Del Rio, B.; Sánchez-Llana, E.; Redruello, B.; Magadan, A.H.; Fernández, M.; Martin, M.C.; Ladero, V.; Alvarez, M.A. Enterococcus faecalis Bacteriophage 156 Is an Effective Biotechnological Tool for Reducing the Presence of Tyramine and Putrescine in an Experimental Cheese Model. Front. Microbiol. 2019, 10, 566. [Google Scholar] [CrossRef]
- Kuley, E.; Özogul, F. Synergistic and antagonistic effect of lactic acid bacteria on tyramine production by food-borne pathogenic bacteria in tyrosine decarboxylase broth. Food Chem. 2011, 127, 1163–1168. [Google Scholar] [CrossRef]
- Marcobal, A.; de las Rivas, B.; Landete, J.M.; Tabera, L.; Muñoz, R. Tyramine and Phenylethylamine Biosynthesis by Food Bacteria. Crit. Rev. Food Sci. Nutr. 2012, 52, 448–467. [Google Scholar] [CrossRef]
- Maifreni, M.; Frigo, F.; Bartolomeoli, I.; Innocente, N.; Biasutti, M.; Marino, M. Identification of the Enterobacteriaceae in Montasio cheese and assessment of their amino acid decarboxylase activity. J. Dairy Res. 2013, 80, 122–127. [Google Scholar] [CrossRef]
- Marino, M.; Maifreni, M.; Moret, S.; Rondinini, G. The capacity of Enterobacteriaceae species to produce biogenic amines in cheese. Lett. Appl. Microbiol. 2000, 31, 169–173. [Google Scholar] [CrossRef]
- Zhao, P.; Li, G.; Shen, Y. New Chemical Constituents from the Endophyte Streptomyces Species LR4612 Cultivated on Maytenus hookeri. Chem. Biodivers. 2006, 3, 337–342. [Google Scholar] [CrossRef]
- Heidari, B.; Mohammadipanah, F. Isolation and identification of two alkaloid structures with radical scavenging activity from Actinokineospora sp. UTMC 968, a new promising source of alkaloid compounds. Mol. Biol. Rep. 2018, 45, 2325–2332. [Google Scholar] [CrossRef]
- Lee, W.; Kim, M.A.; Park, I.W.; Hwang, J.S.; Na, M.K.; Bae, J.S. Novel direct factor Xa inhibitory compounds from Tenebrio molitor with anti-platelet aggregation activity. Food Chem. Toxicol. 2017, 109, 19–27. [Google Scholar] [CrossRef]
- Garcez, W.S.; Martins, D.; Garcez, F.R.; Marques, M.R.; Pereira, A.A.; Oliveira, L.A.; Rondon, J.N.; Peruca, A.D. Effect of spores of saprophytic fungi on phytoalexin accumulation in seeds of frog-eye leaf spot and stem canker-resistant and -susceptible soybean (Glycine max L.) cultivars. J. Agric. Food Chem. 2000, 48, 3662–3665. [Google Scholar] [CrossRef]
- Hernández-Robles, M.F.; Álvarez-Contreras, A.K.; Juárez-García, P.; Natividad-Bonifacio, I.; Curiel-Quesada, E.; Vázquez-Salinas, C.; Quiñones-Ramírez, E.I. Virulence factors and antimicrobial resistance in environmental strains of Vibrio alginolyticus. Int. Microbiol. 2016, 19, 191–198. [Google Scholar]
- Jakšić, S.; Uhitil, S.; Petrak, T.; Bažulić, D.; Gumhalter Karolyi, L. Occurrence of Vibrio spp. in sea fish, shrimps and bivalve molluscs harvested from Adriatic sea. Food Control 2002, 13, 491–493. [Google Scholar] [CrossRef]
- Bassetto, F.; Maschio, N.; Abatangelo, G.; Zavan, B.; Scarpa, C.; Vindigni, V. Collagenase from Vibrio alginolyticus Cultures. Surg. Innov. 2016, 23, 557–562. [Google Scholar] [CrossRef]
- Wang, X.; Huang, Y.; Sheng, Y.; Su, P.; Qiu, Y.; Ke, C.; Feng, D. Antifouling activity towards mussel by small-molecule compounds from a strain of Vibrio alginolyticus bacterium associated with sea anemone Haliplanella sp. J. Microbiol. Biotechnol. 2017, 27, 460–470. [Google Scholar] [CrossRef]
- Padmavathi, A.R.; Abinaya, B.; Pandian, S.K. Phenol, 2,4-bis(1,1-dimethylethyl) of marine bacterial origin inhibits quorum sensing mediated biofilm formation in the uropathogen Serratia marcescens. Biofouling 2014, 30, 1111–1122. [Google Scholar] [CrossRef]
- Morohoshi, T.; Kato, M.; Fukamachi, K.; Kato, N.; Ikeda, T. N-Acylhomoserine lactone regulates violacein production in Chromobacterium violaceum type strain ATCC 12472. FEMS Microbiol. Lett. 2008, 279, 124–130. [Google Scholar] [CrossRef]
- Mcclean, K.H.; Winson, M.K.; Fish, L.; Taylor, A.; Chhabra, S.R.; Camara, M.; Daykin, M.; John, H.; Swift, S.; Bycroft, B.W.; et al. Quorum sensing and Chromobacterium violaceum: Exploitation of violacein production and inhibition for the detection of N-acyl homoserine lactones. Microbiology 1997, 143, 3703–3711. [Google Scholar] [CrossRef]
- Romero, M.; Martin-Cuadrado, A.B.; Roca-Rivada, A.; Cabello, A.M.; Otero, A. Quorum quenching in cultivable bacteria from dense marine coastal microbial communities. FEMS Microbiol. Ecol. 2011, 75, 205–217. [Google Scholar] [CrossRef]
- Ren, D.; Zuo, R.; Wood, T.K. Quorum-sensing antagonist (5Z)-4-bromo-5-(bromomethylene)-3-butyl-2(5H)-furanone influences siderophore biosynthesis in Pseudomonas putida and Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2005, 66, 689–695. [Google Scholar] [CrossRef]
- Turnbull, L.; Whitchurch, C.B. Motility Assay: Twitching Motility. In Pseudomonas Methods and Protocols; Humana Press: New York, NY, USA, 2014; Volume 1149, pp. 73–86. ISBN 978-1-4939-0472-3. [Google Scholar]
- Ha, D.; Kuchma, S.L.; O’Toole, G.A. Plate-Based Assay for Swimming Motility in Pseudomonas aeruginosa. In Pseudomonas Methods and Protocols; Filloux, A., Ramos, J.-L., Eds.; Humana Press: New York, NY, USA, 2014; Volume 1149, pp. 59–65. ISBN 978-1-4939-0472-3. [Google Scholar]
- Ha, D.; Kuchma, S.L.; O’Toole, G.A. Plate-Based Assay for Swarming Motility in Pseudomonas aeruginosa. In Pseudomonas Methods and Protocols; Filloux, A., Ramos, J.-L., Eds.; Humana Press: New York, NY, USA, 2014; Volume 1149, pp. 67–72. ISBN 978-1-4939-0472-3. [Google Scholar]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- O’Toole, G.A. Microtiter Dish Biofilm Formation Assay. J. Vis. Exp. 2011, 47, e2437. [Google Scholar] [CrossRef]
- Martín-Platero, A.M.; Valdivia, E.; Maqueda, M.; Martínez-Bueno, M. Fast, convenient, and economical method for isolating genomic DNA from lactic acid bacteria using a modification of the protein “salting-out” procedure. Anal. Biochem. 2007, 366, 102–104. [Google Scholar] [CrossRef]
- Lane, D. 16S/23S rRna Sequencing. In Nucleic Acid Techniques in Bacterial Systematics; Stackebrandt, E., Goodfellow, M., Eds.; John Wiley and Sons: Hoboken, NY, USA, 1991; pp. 115–175. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Yoon, S.-H.; Ha, S.-M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613–1617. [Google Scholar]
- Marmur, J. A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J. Mol. Biol. 1961, 3, 208–218. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef]
- Rodriguez-R, L.M.; Konstantinidis, K.T. The enveomics collection: A toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Prepr. 2016, 4, e1900v1. [Google Scholar]
- Alikhan, N.-F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Bryant, S.H. CD-Search: Protein domain annotations on the fly. Nucleic Acids Res. 2004, 32, W327–W331. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R.; et al. CDD: A Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef]


| Strain | Identification | Violacein Inhibition Halo (mm) |
|---|---|---|
| M1-9 | Vibrio neocaledonicus | 16 |
| M2-99 | Vibrio antiquarius | 13 |
| M3-10 | Vibrio alginolyticus | 18 |
| M3-21 | V. neocaledonicus | 15 |
| M3-78 | V. alginolyticus | 15 |
| M3-99 | V. alginolyticus | 15 |
| M4-31 | V. alginolyticus | 17 |
| M4-35 | V. alginolyticus | 12 |
| M4-116 | V. alginolyticus | 13 |
| M4-117 | V. neocaledonicus | 12 |
| M4-119 | V. neocaledonicus | 16 |
| M4-126 | V. neocaledonicus | 11 |
| M5-23 | V. neocaledonicus | 13 |
| M5-35 | V. neocaledonicus | 16 |
| M5-47 | V. neocaledonicus | 18 |
| M5-50 | V. neocaledonicus | 13 |
| M6-17 | V. neocaledonicus | 13 |
| M6-26 | V. neocaledonicus | 13 |
| M6-31 | V. alginolyticus | 11 |
| M6-39 | V. zhuhaiensis | 14 |
| M6-50 | V. neocaledonicus | 17 |
| M6-66 | V. alginolyticus | 15 |
| M10-18 | V. neocaledonicus | 14 |
| Extract | Violacein Production (Absorbance at 585 nm) | |
|---|---|---|
| M3-10 | cell pellet | 0.034 |
| supernatant | 0.206 | |
| M4-31 | cell pellet | 0.044 |
| supernatant | 0.282 | |
| M4-119 | cell pellet | 0.06 |
| supernatant | 0.276 | |
| M5-47 | cell pellet | 0.044 |
| supernatant | 0.216 | |
| M6-50 | cell pellet | 0.042 |
| supernatant | 0.280 | |
| MeOH (negative control) | 0.489 |
| Mean in the Absence Of Cell Pellet Extract | Mean in the Presence Of Cell Pellet Extract | |
|---|---|---|
| C. violaceum ATCC 12472 | ||
| Violacein (% production) | 100 [±10.9] | 33.89 * [±17.1] |
| P. aeruginosa PAO1 | ||
| Pyoverdine (% production) | 100 [±16.1] | 74.02 * [±8] |
| Biofilm (% production) | 100 [±12.7] | 71.96 * [±5.9] |
| Swimming (% motility) | 55.6 [±8.1] | 42.8 * [±2.5] |
| Swarming (% motility) | 39.46 [±15.3] | 27.36 [±9.6] |
| Twitching (% motility) | 39.15 [±13.6] | 31.54 [±10.7] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reina, J.C.; Pérez-Victoria, I.; Martín, J.; Llamas, I. A Quorum-Sensing Inhibitor Strain of Vibrio alginolyticus Blocks Qs-Controlled Phenotypes in Chromobacterium violaceum and Pseudomonas aeruginosa. Mar. Drugs 2019, 17, 494. https://doi.org/10.3390/md17090494
Reina JC, Pérez-Victoria I, Martín J, Llamas I. A Quorum-Sensing Inhibitor Strain of Vibrio alginolyticus Blocks Qs-Controlled Phenotypes in Chromobacterium violaceum and Pseudomonas aeruginosa. Marine Drugs. 2019; 17(9):494. https://doi.org/10.3390/md17090494
Chicago/Turabian StyleReina, José Carlos, Ignacio Pérez-Victoria, Jesús Martín, and Inmaculada Llamas. 2019. "A Quorum-Sensing Inhibitor Strain of Vibrio alginolyticus Blocks Qs-Controlled Phenotypes in Chromobacterium violaceum and Pseudomonas aeruginosa" Marine Drugs 17, no. 9: 494. https://doi.org/10.3390/md17090494
APA StyleReina, J. C., Pérez-Victoria, I., Martín, J., & Llamas, I. (2019). A Quorum-Sensing Inhibitor Strain of Vibrio alginolyticus Blocks Qs-Controlled Phenotypes in Chromobacterium violaceum and Pseudomonas aeruginosa. Marine Drugs, 17(9), 494. https://doi.org/10.3390/md17090494