Microalgal Enzymes with Biotechnological Applications
Abstract
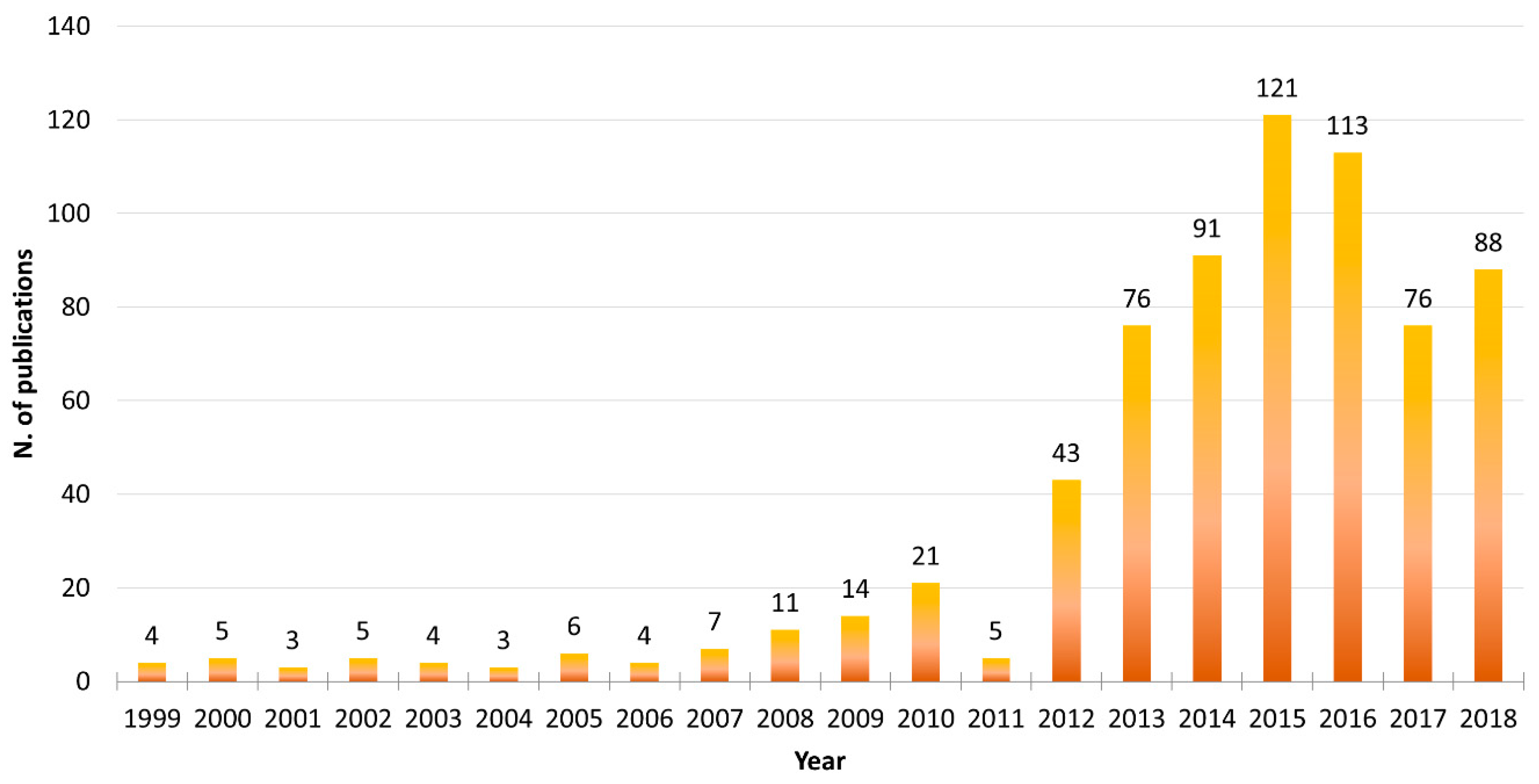
1. Introduction
2. Enzymes from Microalgae
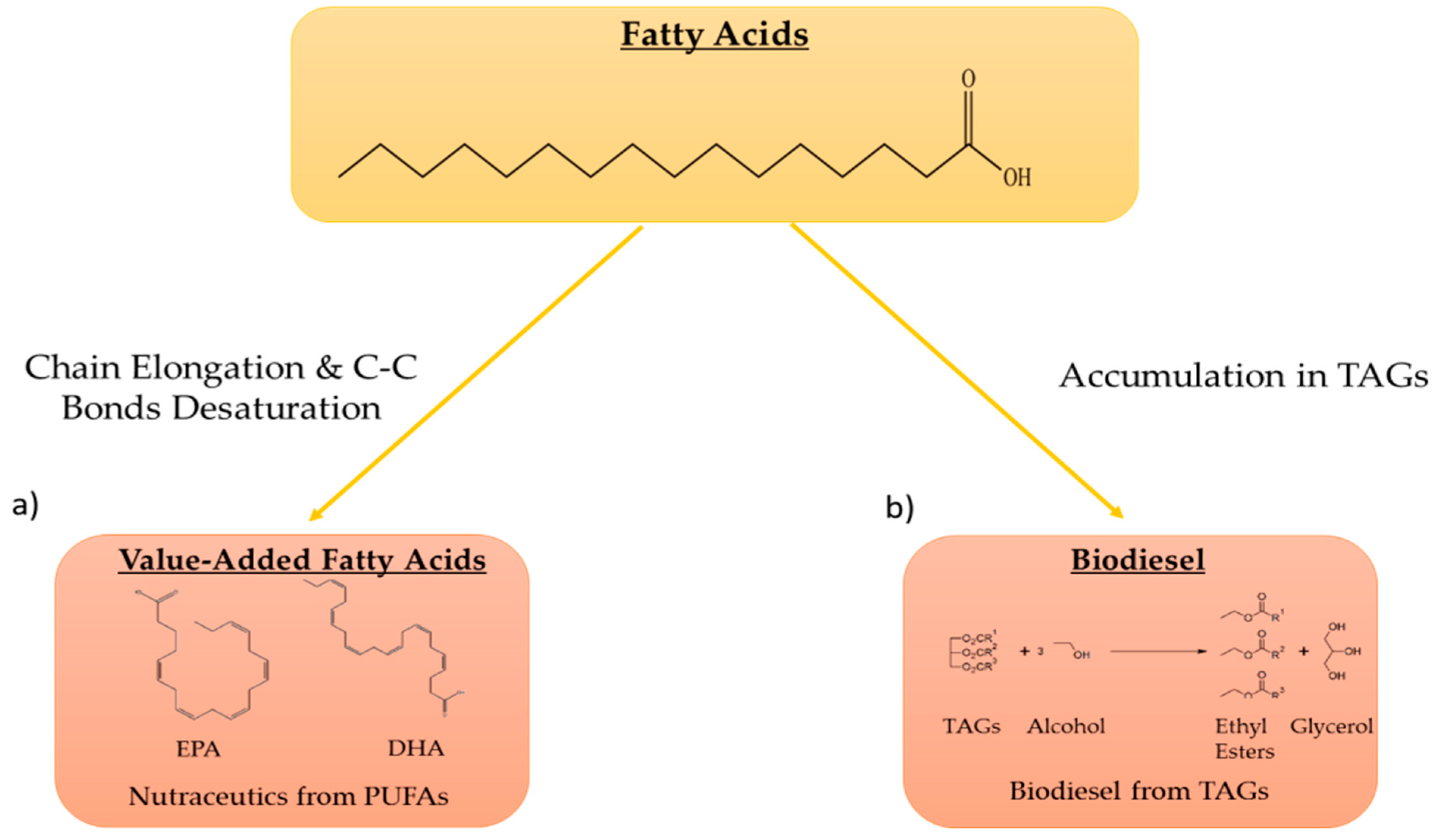
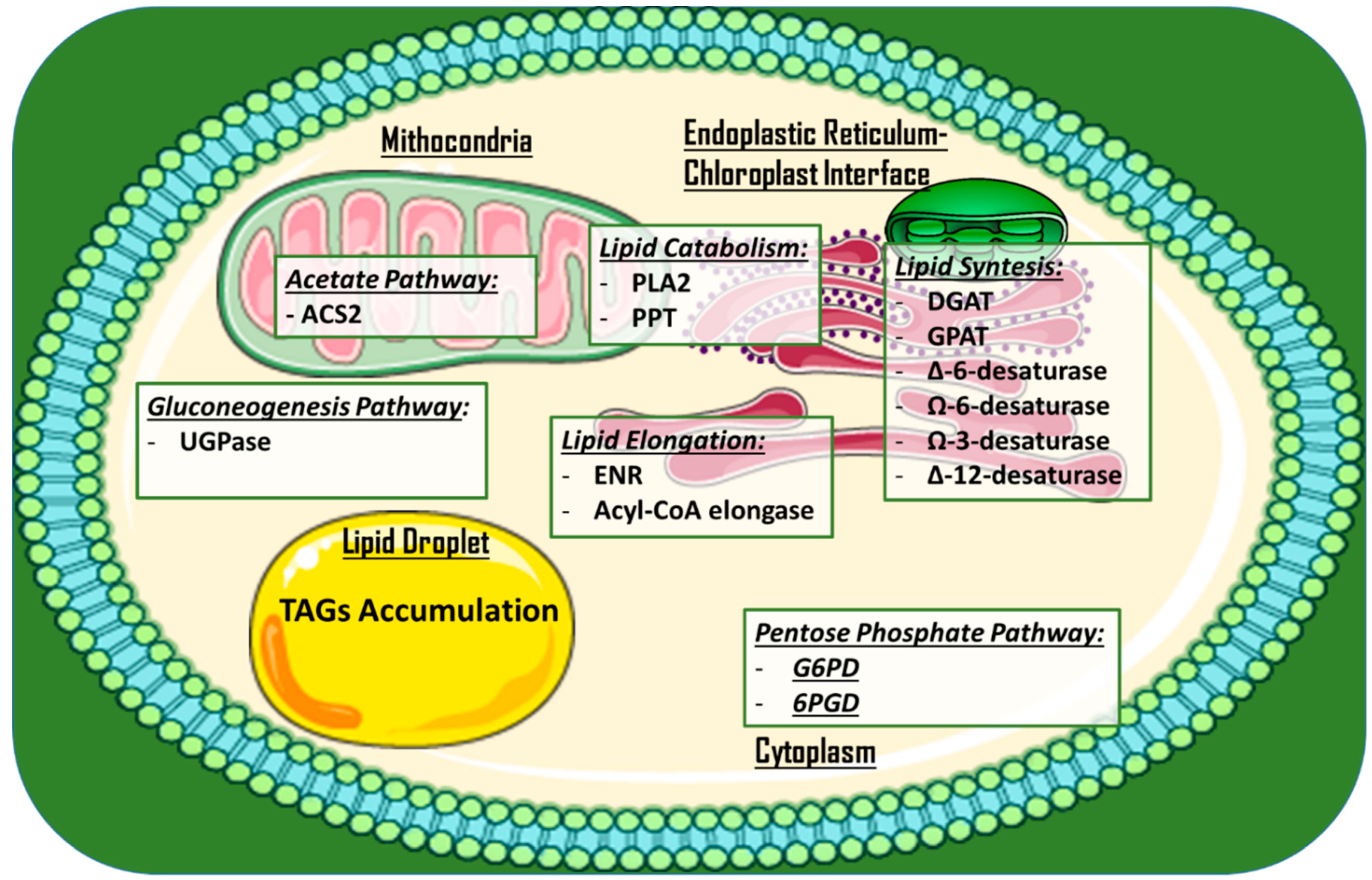
2.1. Enzymes for High-Value Added Lipids and Biodiesel Production
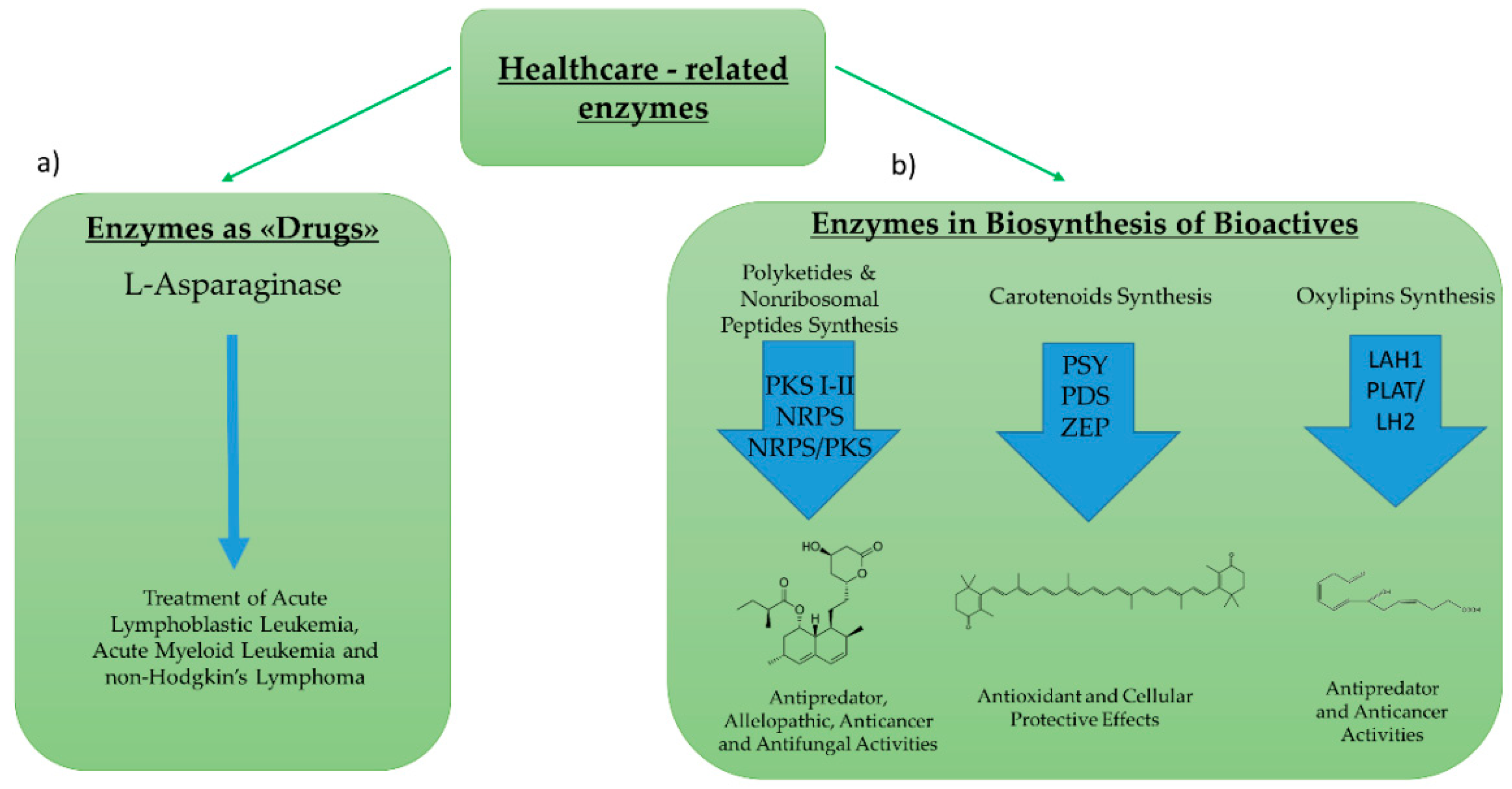
2.2. Enzymes for Healthcare Application
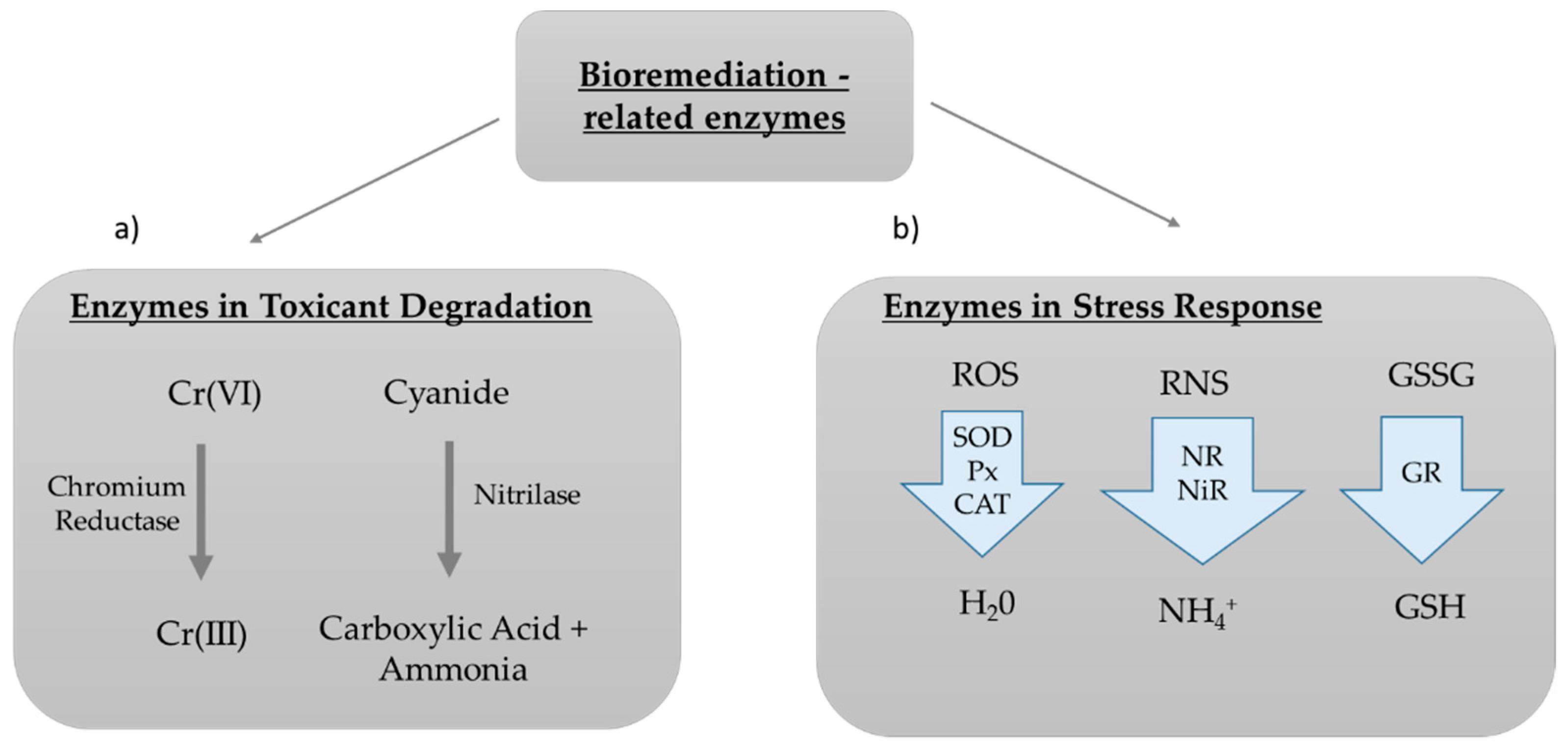
2.3. Enzymes for Bioremediation
3. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schneider, S.H.; Root, T.L.; Mastrandrea, M.D. Encyclopedia of climate and weather; Oxford University Press: Oxford, UK, 2011. [Google Scholar]
- Bagchi, B. Water in Biological and Chemical Processes; Cambridge University Press: Cambridge, UK, 2013. [Google Scholar]
- Faber, K. Biotransformations in Organic Chemistry; Springer: Heidelberg, Germany, 2011. [Google Scholar]
- Cao, Y.; Tan, H. Effects of cellulase on the modification of cellulose. Carbohydr. Res. 2002, 337, 1291–1296. [Google Scholar] [CrossRef]
- Nigam, P.S. Microbial enzymes with special characteristics for biotechnological applications. Biomolecules 2013, 3, 597–611. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, P. Enzymes in food processing: a condensed overview on strategies for better biocatalysts. Enzyme Res. 2010, 2010, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Vellard, M. The enzyme as drug: application of enzymes as pharmaceuticals. Curr. Opin. Biotechnol. 2003, 14, 444–450. [Google Scholar] [CrossRef]
- Andersen, R.A. Algal culturing techniques; Academic Press: Cambridge, MA, USA, 2005. [Google Scholar]
- Khan, M.I.; Shin, J.H.; Kim, J.D. The promising future of microalgae: current status, challenges, and optimization of a sustainable and renewable industry for biofuels, feed, and other products. Microb. Cell Fact. 2018, 17, 36. [Google Scholar] [CrossRef] [PubMed]
- Martínez Andrade, K.; Lauritano, C.; Romano, G.; Ianora, A. Marine Microalgae with Anti-Cancer Properties. Mar. Drugs 2018, 16, 165. [Google Scholar] [CrossRef]
- Bhalamurugan, G.L.; Valerie, O.; Mark, L. Valuable bioproducts obtained from microalgal biomass and their commercial applications: A review. Environ. Eng. Res. 2018, 23, 229–241. [Google Scholar] [CrossRef]
- Doron, L.; Segal, N.; Shapira, M. Transgene Expression in Microalgae-From Tools to Applications. Front. Plant Sci. 2016, 7, 505. [Google Scholar] [CrossRef]
- Brasil, B.d.S.A.F.; de Siqueira, F.G.; Salum, T.F.C.; Zanette, C.M.; Spier, M.R. Microalgae and cyanobacteria as enzyme biofactories. Algal Res. 2017, 25, 76–89. [Google Scholar] [CrossRef]
- Lauritano, C.; Ianora, A. Grand Challenges in Marine Biotechnology: Overview of Recent EU-Funded Projects. In Grand Challenges in Marine Biotechnology; Rampellotto, P.H., Trincone, A., Eds.; Springer: Heidelberg, Germany, 2018; pp. 425–449. [Google Scholar]
- Mogharabi, M.; Faramarzi, M.A. Are Algae the Future Source of Enzymes? Trends Pept. Protein Sci. 2016, 1, 1–6. [Google Scholar]
- Chisti, Y. Biodiesel from microalgae. Biotechnol. Adv. 2007, 25, 294–306. [Google Scholar] [CrossRef]
- Sanghvi, A.M.; Martin Lo, Y. Present and Potential Industrial Applications of Macro- and Microalgae. Recent Patents Food, Nutr. Agric. 2010, 2, 187–194. [Google Scholar]
- Bellou, S.; Baeshen, M.N.; Elazzazy, A.M.; Aggeli, D.; Sayegh, F.; Aggelis, G. Microalgal lipids biochemistry and biotechnological perspectives. Biotechnol. Adv. 2014, 32, 1476–1493. [Google Scholar] [CrossRef]
- Moriyama, T.; Toyoshima, M.; Saito, M.; Wada, H.; Sato, N. Revisiting the Algal “Chloroplast Lipid Droplet”: The Absence of an Entity That Is Unlikely to Exist. Plant Physiol. 2018, 176, 1519–1530. [Google Scholar] [CrossRef]
- Fukuda, H.; Kondo, A.; Noda, H. Biodiesel fuel production by transesterification of oils. J. Biosci. Bioeng. 2001, 92, 405–416. [Google Scholar] [CrossRef]
- Wells, M.L.; Potin, P.; Craigie, J.S.; Raven, J.A.; Merchant, S.S.; Helliwell, K.E.; Smith, A.G.; Camire, M.E.; Brawley, S.H. Algae as nutritional and functional food sources: revisiting our understanding. J. Appl. Phycol. 2017, 29, 949–982. [Google Scholar] [CrossRef]
- Caporgno, M.P.; Mathys, A. Trends in microalgae incorporation into innovative food products with potential health benefits. Front. Nutr. 2018, 5, 1–10. [Google Scholar] [CrossRef]
- Merchant, S.S.; Kropat, J.; Liu, B.; Shaw, J.; Warakanont, J. TAG, You’re it! Chlamydomonas as a reference organism for understanding algal triacylglycerol accumulation. Curr. Opin. Biotechnol. 2012, 23, 352–363. [Google Scholar] [CrossRef]
- Xu, Y.; Caldo, K.M.P.; Pal-Nath, D.; Ozga, J.; Lemieux, M.J.; Weselake, R.J.; Chen, G. Properties and Biotechnological Applications of Acyl-CoA:diacylglycerol Acyltransferase and Phospholipid:diacylglycerol Acyltransferase from Terrestrial Plants and Microalgae. Lipids 2018, 53, 663–688. [Google Scholar] [CrossRef]
- Lung, S.-C.; Weselake, R.J. Diacylglycerol acyltransferase: a key mediator of plant triacylglycerol synthesis. Lipids 2006, 41, 1073–1088. [Google Scholar] [CrossRef]
- Shockey, J.M.; Gidda, S.K.; Chapital, D.C.; Kuan, J.-C.; Dhanoa, P.K.; Bland, J.M.; Rothstein, S.J.; Mullen, R.T.; Dyer, J.M. Tung tree DGAT1 and DGAT2 have nonredundant functions in triacylglycerol biosynthesis and are localized to different subdomains of the endoplasmic reticulum. Plant Cell 2006, 18, 2294–2313. [Google Scholar] [CrossRef]
- Guo, X.; Fan, C.; Chen, Y.; Wang, J.; Yin, W.; Wang, R.R.C.; Hu, Z. Identification and characterization of an efficient acyl-CoA: Diacylglycerol acyltransferase 1 (DGAT1) gene from the microalga Chlorella ellipsoidea. BMC Plant Biol. 2017, 17, 1–16. [Google Scholar] [CrossRef]
- Wei, H.; Shi, Y.; Ma, X.; Pan, Y.; Hu, H.; Li, Y.; Luo, M.; Gerken, H.; Liu, J. A type-I diacylglycerol acyltransferase modulates triacylglycerol biosynthesis and fatty acid composition in the oleaginous microalga, Nannochloropsis oceanica. Biotechnol. Biofuels 2017, 10, 1–18. [Google Scholar] [CrossRef]
- Wagner, M.; Hoppe, K.; Czabany, T.; Heilmann, M.; Daum, G.; Feussner, I.; Fulda, M. Identification and characterization of an acyl-CoA:diacylglycerol acyltransferase 2 (DGAT2) gene from the microalga Ostreococcus tauri. Plant Physiol. Biochem. 2010, 48, 407–416. [Google Scholar] [CrossRef]
- Niu, Y.-F.; Zhang, M.-H.; Li, D.-W.; Yang, W.-D.; Liu, J.-S.; Bai, W.-B.; Li, H.-Y. Improvement of Neutral Lipid and Polyunsaturated Fatty Acid Biosynthesis by Overexpressing a Type 2 Diacylglycerol Acyltransferase in Marine Diatom Phaeodactylum tricornutum. Mar. Drugs 2013, 11, 4558–4569. [Google Scholar] [CrossRef]
- Manandhar-Shrestha, K.; Hildebrand, M. Characterization and manipulation of a DGAT2 from the diatom Thalassiosira pseudonana: Improved TAG accumulation without detriment to growth, and implications for chloroplast TAG accumulation. Algal Res. 2015, 12, 239–248. [Google Scholar] [CrossRef]
- Klaitong, P.; Fa-aroonsawat, S.; Chungjatupornchai, W. Accelerated triacylglycerol production and altered fatty acid composition in oleaginous microalga Neochloris oleoabundans by overexpression of diacylglycerol acyltransferase 2. Microb. Cell Fact. 2017, 16, 1–10. [Google Scholar] [CrossRef]
- Li, D.-W.; Cen, S.-Y.; Liu, Y.-H.; Balamurugan, S.; Zheng, X.-Y.; Alimujiang, A.; Yang, W.-D.; Liu, J.-S.; Li, H.-Y. A type 2 diacylglycerol acyltransferase accelerates the triacylglycerol biosynthesis in heterokont oleaginous microalga Nannochloropsis oceanica. J. Biotechnol. 2016, 229, 65–71. [Google Scholar] [CrossRef]
- Xin, Y.; Lu, Y.; Lee, Y.-Y.; Wei, L.; Jia, J.; Wang, Q.; Wang, D.; Bai, F.; Hu, H.; Hu, Q.; et al. Producing Designer Oils in Industrial Microalgae by Rational Modulation of Co-evolving Type-2 Diacylglycerol Acyltransferases. Mol. Plant 2017, 10, 1523–1539. [Google Scholar] [CrossRef]
- Deng, X.-D.; Gu, B.; Li, Y.-J.; Hu, X.-W.; Guo, J.-C.; Fei, X.-W. The roles of acyl-CoA: diacylglycerol acyltransferase 2 genes in the biosynthesis of triacylglycerols by the green algae Chlamydomonas reinhardtii. Mol. Plant 2012, 5, 945–947. [Google Scholar] [CrossRef]
- La Russa, M.; Bogen, C.; Uhmeyer, A.; Doebbe, A.; Filippone, E.; Kruse, O.; Mussgnug, J.H. Functional analysis of three type-2 DGAT homologue genes for triacylglycerol production in the green microalga Chlamydomonas reinhardtii. J. Biotechnol. 2012, 162, 13–20. [Google Scholar] [CrossRef]
- Cui, Y.; Zhao, J.; Wang, Y.; Qin, S.; Lu, Y. Characterization and engineering of a dual-function diacylglycerol acyltransferase in the oleaginous marine diatom Phaeodactylum tricornutum. Biotechnol. Biofuels 2018, 11, 32. [Google Scholar] [CrossRef]
- Xue, J.; Balamurugan, S.; Li, D.-W.; Liu, Y.-H.; Zeng, H.; Wang, L.; Yang, W.-D.; Liu, J.-S.; Li, H.-Y. Glucose-6-phosphate dehydrogenase as a target for highly efficient fatty acid biosynthesis in microalgae by enhancing NADPH supply. Metab. Eng. 2017, 41, 212–221. [Google Scholar] [CrossRef]
- Zhu, B.-H.; Tu, C.-C.; Shi, H.-P.; Yang, G.-P.; Pan, K.-H. Overexpression of endogenous delta-6 fatty acid desaturase gene enhances eicosapentaenoic acid accumulation in Phaeodactylum tricornutum. Process Biochem. 2017, 57, 43–49. [Google Scholar] [CrossRef]
- Fukuda, S.; Hirasawa, E.; Takemura, T.; Takahashi, S.; Chokshi, K.; Pancha, I.; Tanaka, K.; Imamura, S. Accelerated triacylglycerol production without growth inhibition by overexpression of a glycerol-3-phosphate acyltransferase in the unicellular red alga Cyanidioschyzon merolae. Sci. Rep. 2018, 8, 1–12. [Google Scholar] [CrossRef]
- Rengel, R.; Smith, R.T.; Haslam, R.P.; Sayanova, O.; Vila, M.; León, R. Overexpression of acetyl-CoA synthetase (ACS) enhances the biosynthesis of neutral lipids and starch in the green microalga Chlamydomonas reinhardtii. Algal Res. 2018, 31, 183–193. [Google Scholar] [CrossRef]
- Osada, K.; Maeda, Y.; Yoshino, T.; Nojima, D.; Bowler, C.; Tanaka, T. Enhanced NADPH production in the pentose phosphate pathway accelerates lipid accumulation in the oleaginous diatom Fistulifera solaris. Algal Res. 2017, 23, 126–134. [Google Scholar] [CrossRef]
- Shin, Y.S.; Jeong, J.; Nguyen, T.H.T.; Kim, J.Y.H.; Jin, E.; Sim, S.J. Targeted knockout of phospholipase A2 to increase lipid productivity in Chlamydomonas reinhardtii for biodiesel production. Bioresour. Technol. 2019, 271, 368–374. [Google Scholar] [CrossRef]
- Los, D.A.; Murata, N. Structure and expression of fatty acid desaturases. Biochim. Biophys. Acta Lipids Lipid Metab. 1998, 1394, 3–15. [Google Scholar] [CrossRef]
- De Jaeger, L.; Springer, J.; Wolbert, E.J.H.; Martens, D.E.; Eggink, G.; Wijffels, R.H. Gene silencing of stearoyl-ACP desaturase enhances the stearic acid content in Chlamydomonas reinhardtii. Bioresour. Technol. 2017, 245, 1616–1626. [Google Scholar] [CrossRef]
- Daboussi, F.; Leduc, S.; Maréchal, A.; Dubois, G.; Guyot, V.; Perez-Michaut, C.; Amato, A.; Falciatore, A.; Juillerat, A.; Beurdeley, M.; et al. Genome engineering empowers the diatom Phaeodactylum tricornutum for biotechnology. Nat. Commun. 2014, 5, 3831. [Google Scholar] [CrossRef]
- Sorigué, D.; Légeret, B.; Cuiné, S.; Blangy, S.; Moulin, S.; Billon, E.; Richaud, P.; Brugière, S.; Couté, Y.; Nurizzo, D.; et al. An algal photoenzyme converts fatty acids to hydrocarbons. Science 2017, 357, 903–907. [Google Scholar] [CrossRef]
- Misra, N.; Panda, P.K.; Parida, B.K.; Mishra, B.K. dEMBF: A comprehensive database of enzymes of microalgal biofuel feedstock. PLoS One 2016, 11, 146–158. [Google Scholar] [CrossRef]
- Batool, T.; Makky, E.A.; Jalal, M.; Yusoff, M.M. A comprehensive review on L-asparaginase and its applications. Appl. Biochem. Biotechnol. 2016, 178, 900–923. [Google Scholar] [CrossRef]
- Ali, U.; Naveed, M.; Ullah, A.; Ali, K.; Shah, S.A.; Fahad, S.; Mumtaz, A.S. L-asparaginase as a critical component to combat Acute Lymphoblastic Leukemia (ALL): A novel approach to target ALL. Eur. J. Pharmacol. 2016, 771, 199–210. [Google Scholar] [CrossRef]
- Roberts, J.; Prager, M.D.; Bachynsky, N. The antitumor activity of Escherichia coli L-asparaginase. Cancer Res. 1966, 26, 2213–2217. [Google Scholar]
- Peterson, R.E.; Ciegler, A. L-asparaginase production by various bacteria. Appl. Microbiol. 1969, 17, 929–930. [Google Scholar]
- Thenmozhi, C.; Sankar, R.; Karuppiah, V.; Sampathkumar, P. L-asparaginase production by mangrove derived Bacillus cereus MAB5: Optimization by response surface methodology. Asian Pac. J. Trop. Med. 2011, 4, 486–491. [Google Scholar] [CrossRef]
- Ahmad, N.; Pandit, N.; Maheshwari, S. L-asparaginase gene-a therapeutic approach towards drugs for cancer cell. Int. J. Biosci. 2012, 2, 1–11. [Google Scholar]
- Vidya, J.; Sajitha, S.; Ushasree, V.; Sindhu, R.; Binod, P.; Madhavan, A.; Pandey, A. Genetic and metabolic engineering approaches for the production and delivery of L-asparaginases: An overview. Bioresour. Technol. 2017, 245, 1775–1781. [Google Scholar] [CrossRef]
- Paul, J.H. Isolation and characterization of a Chlamydomonas L-asparaginase. Biochem. J. 1982, 203, 109–115. [Google Scholar] [CrossRef]
- Ebrahiminezhad, A.; Rasoul-Amini, S.; Ghoshoon, M.B.; Ghasemi, Y. Chlorella vulgaris, a novel microalgal source for L-asparaginase production. Biocatal. Agric. Biotechnol. 2014, 3, 214–217. [Google Scholar] [CrossRef]
- Sasso, S.; Pohnert, G.; Lohr, M.; Mittag, M.; Hertweck, C. Microalgae in the postgenomic era: a blooming reservoir for new natural products. FEMS Microbiol. Rev. 2012, 36, 761–785. [Google Scholar] [CrossRef]
- Berry, J. Marine and freshwater microalgae as a potential source of novel herbicides. In Herbicides and Environment, Kortekamp A.; InTechOpen: London, UK, 2011. [Google Scholar]
- Kobayashi, J. Amphidinolides and its related macrolides from marine dinoflagellates. J. Antibiot. (Tokyo) 2008, 61, 271–284. [Google Scholar] [CrossRef]
- Kellmann, R.; Stüken, A.; Orr, R.J.S.; Svendsen, H.M.; Jakobsen, K.S. Biosynthesis and molecular genetics of polyketides in marine dinoflagellates. Mar. Drugs 2010, 8, 1011–1048. [Google Scholar] [CrossRef]
- Kohli, G.S.; John, U.; Van Dolah, F.M.; Murray, S.A. Evolutionary distinctiveness of fatty acid and polyketide synthesis in eukaryotes. ISME J. 2016, 10, 1877–1890. [Google Scholar] [CrossRef]
- Jenke-Kodama, H.; Sandmann, A.; Müller, R.; Dittmann, E. Evolutionary implications of bacterial polyketide synthases. Mol. Biol. Evol. 2005, 22, 2027–2039. [Google Scholar] [CrossRef]
- Lauritano, C.; De Luca, D.; Ferrarini, A.; Avanzato, C.; Minio, A.; Esposito, F.; Ianora, A. De novo transcriptome of the cosmopolitan dinoflagellate Amphidinium carterae to identify enzymes with biotechnological potential. Sci. Rep. 2017, 7, 11701. [Google Scholar] [CrossRef]
- Meyer, J.M.; Rödelsperger, C.; Eichholz, K.; Tillmann, U.; Cembella, A.; McGaughran, A.; John, U. Transcriptomic characterisation and genomic glimps into the toxigenic dinoflagellate Azadinium spinosum, with emphasis on polykeitde synthase genes. BMC Genomics 2015, 16, 27. [Google Scholar] [CrossRef]
- Kohli, G.S.; Campbell, K.; John, U.; Smith, K.F.; Fraga, S.; Rhodes, L.L.; Murray, S.A. Role of modular polyketide synthases in the production of polyether ladder compounds in ciguatoxin-producing Gambierdiscus polynesiensis and G. excentricus (Dinophyceae). J. Eukaryot. Microbiol. 2017, 64, 691–706. [Google Scholar] [CrossRef]
- Monroe, E.A.; Van Dolah, F.M. The toxic dinoflagellate Karenia brevis encodes novel Type I-like polyketide synthases containing discrete catalytic domains. Protist 2008, 159, 471–482. [Google Scholar] [CrossRef]
- Lauritano, C.; De Luca, D.; Amoroso, M.; Benfatto, S.; Maestri, S.; Racioppi, C.; Esposito, F.; Lanora, A. New molecular insights on the response of the green alga Tetraselmis suecica to nitrogen starvation. Sci. Rep. 2019, 9, 3336. [Google Scholar] [CrossRef]
- López-Legentil, S.; Song, B.; DeTure, M.; Baden, D.G. Characterization and localization of a hybrid non-ribosomal peptide synthetase and polyketide synthase gene from the toxic dinoflagellate Karenia brevis. Mar. Biotechnol. 2010, 12, 32–41. [Google Scholar] [CrossRef]
- Fiedor, J.; Burda, K. Potential role of carotenoids as antioxidants in human health and disease. Nutrients 2014, 6, 466–488. [Google Scholar] [CrossRef]
- Musser, A.J.; Maiuri, M.; Brida, D.; Cerullo, G.; Friend, R.H.; Clark, J. The nature of singlet exciton fission in carotenoid aggregates. J. Am. Chem. Soc. 2015, 137, 5130–5139. [Google Scholar] [CrossRef]
- Gong, M.; Bassi, A. Carotenoids from microalgae: A review of recent developments. Biotechnol. Adv. 2016, 34, 1396–1412. [Google Scholar] [CrossRef]
- Rammuni, M.N.; Ariyadasa, T.U.; Nimarshana, P.H.V.; Attalage, R.A. Comparative assessment on the extraction of carotenoids from microalgal sources: Astaxanthin from H. pluvialis and β-carotene from D. salina. Food Chem. 2019, 277, 128–134. [Google Scholar] [CrossRef]
- Lamers, P.P.; Janssen, M.; De Vos, R.C.H.; Bino, R.J.; Wijffels, R.H. Exploring and exploiting carotenoid accumulation in Dunaliella salina for cell-factory applications. Trends Biotechnol. 2008, 26, 631–638. [Google Scholar] [CrossRef]
- Prieto, A.; Pedro Cañavate, J.; García-González, M. Assessment of carotenoid production by Dunaliella salina in different culture systems and operation regimes. J. Biotechnol. 2011, 151, 180–185. [Google Scholar] [CrossRef]
- Besson, A.; Formosa-Dague, C.; Guiraud, P. Flocculation-flotation harvesting mechanism of Dunaliella salina: From nanoscale interpretation to industrial optimization. Water Res. 2019, 155, 352–361. [Google Scholar] [CrossRef]
- Tan, C.; Qin, S.; Zhang, Q.; Jiang, P.; Zhao, F. Establishment of a micro-particle bombardment transformation system for Dunaliella salina. J. Microbiol. 2005, 43, 361–365. [Google Scholar]
- Simon, D.P.; Anila, N.; Gayathri, K.; Sarada, R. Heterologous expression of β-carotene hydroxylase in Dunaliella salina by Agrobacterium-mediated genetic transformation. Algal Res. 2016, 18, 257–265. [Google Scholar] [CrossRef]
- Saini, D.K.; Chakdar, H.; Pabbi, S.; Shukla, P. Enhancing production of microalgal biopigments through metabolic and genetic engineering. Crit. Rev. Food Sci. Nutr. 2019, 1–15. [Google Scholar] [CrossRef]
- Huang, J.C.; Wang, Y.; Sandmann, G.; Chen, F. Isolation and characterization of a carotenoid oxygenase gene from Chlorella zofingiensis (Chlorophyta). Appl. Microbiol. Biotechnol. 2006, 71, 473–479. [Google Scholar] [CrossRef]
- Cordero, B.F.; Couso, I.; Leon, R.; Rodriguez, H.; Vargas, M.A. Isolation and characterization of a lycopene ε-cyclase gene of Chlorella (Chromochloris) zofingiensis. Regulation of the carotenogenic pathway by nitrogen and light. Mar. Drugs 2012, 10, 2069–2088. [Google Scholar] [CrossRef]
- Couso, I.; Vila, M.; Vigara, J.; Cordero, B.F.; Vargas, M.Á.; Rodríguez, H.; León, R. Synthesis of carotenoids and regulation of the carotenoid biosynthesis pathway in response to high light stress in the unicellular microalga Chlamydomonas reinhardtii. Eur. J. Phycol. 2012, 47, 223–232. [Google Scholar] [CrossRef]
- Cunningham, F.X.; Pogson, B.; Sun, Z.; McDonald, K.A.; DellaPenna, D.; Gantt, E.; Gantt, E. Functional analysis of the beta and epsilon lycopene cyclase enzymes of Arabidopsis reveals a mechanism for control of cyclic carotenoid formation. Plant Cell 1996, 8, 1613–1626. [Google Scholar] [CrossRef]
- Cordero, B.F.; Couso, I.; León, R.; Rodríguez, H.; Vargas, M.Á. Enhancement of carotenoids biosynthesis in Chlamydomonas reinhardtii by nuclear transformation using a phytoene synthase gene isolated from Chlorella zofingiensis. Appl. Microbiol. Biotechnol. 2011, 91, 341–351. [Google Scholar] [CrossRef]
- Fraser, P.D.; Misawa, N.; Linden, H.; Yamano, S.; Kobayashi, K.; Sandmann, G. Expression in Escherichia coli, purification, and reactivation of the recombinant Erwinia uredovora phytoene desaturase. J. Biol. Chem. 1992, 267, 19891–19895. [Google Scholar]
- Steinbrenner, J.; Sandmann, G. Transformation of the green alga Haematococcus pluvialis with a phytoene desaturase for accelerated astaxanthin biosynthesis. Appl. Environ. Microbiol. 2006, 72, 7477–7484. [Google Scholar] [CrossRef]
- Galarza, J.I.; Gimpel, J.A.; Rojas, V.; Arredondo-Vega, B.O.; Henríquez, V. Over-accumulation of astaxanthin in Haematococcus pluvialis through chloroplast genetic engineering. Algal Res. 2018, 31, 291–297. [Google Scholar] [CrossRef]
- Srinivasan, R.; Babu, S.; Gothandam, K.M. Accumulation of phytoene, a colorless carotenoid by inhibition of phytoene desaturase (PDS) gene in Dunaliella salina V-101. Bioresour. Technol. 2017, 242, 311–318. [Google Scholar] [CrossRef]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013, 339, 819–823. [Google Scholar] [CrossRef]
- Baek, K.; Yu, J.; Jeong, J.; Sim, S.J.; Bae, S.; Jin, E. Photoautotrophic production of macular pigment in a Chlamydomonas reinhardtii strain generated by using DNA-free CRISPR-Cas9 RNP-mediated mutagenesis. Biotechnol. Bioeng. 2018, 115, 719–728. [Google Scholar] [CrossRef]
- Frommolt, R.; Goss, R.; Wilhelm, C. The de-epoxidase and epoxidase reactions of Mantoniella squamata (Prasinophyceae) exhibit different substrate-specific reaction kinetics compared to spinach. Planta 2001, 213, 446–456. [Google Scholar] [CrossRef]
- Gimpel, J.A.; Henríquez, V.; Mayfield, S.P. In Metabolic Engineering of Eukaryotic Microalgae: Potential and Challenges Come with Great Diversity. Front. Microbiol. 2015, 6, 1376. [Google Scholar] [CrossRef]
- De los Reyes, C.; Ávila-Román, J.; Ortega, M.J.; de la Jara, A.; García-Mauriño, S.; Motilva, V.; Zubía, E. Oxylipins from the microalgae Chlamydomonas debaryana and Nannochloropsis gaditana and their activity as TNF-α inhibitors. Phytochemistry 2014, 102, 152–161. [Google Scholar] [CrossRef]
- Lauritano, C.; Romano, G.; Roncalli, V.; Amoresano, A.; Fontanarosa, C.; Bastianini, M.; Braga, F.; Carotenuto, Y.; Ianora, A. New oxylipins produced at the end of a diatom bloom and their effects on copepod reproductive success and gene expression levels. Harmful Algae 2016, 55, 221–229. [Google Scholar] [CrossRef]
- Ávila-Román, J.; Talero, E.; de los Reyes, C.; García-Mauriño, S.; Motilva, V. Microalgae-derived oxylipins decrease inflammatory mediators by regulating the subcellular location of NFκB and PPAR-γ. Pharmacol. Res. 2018, 128, 220–230. [Google Scholar] [CrossRef]
- Pohnert, G. Phospholipase A2 activity triggers the wound-activated chemical defense in the diatom Thalassiosira rotula. Plant Physiol. 2002, 129, 103–111. [Google Scholar] [CrossRef]
- Matos, A.R.; Pham-Thi, A.T. Lipid deacylating enzymes in plants: Old activities, new genes. Plant Physiol. Biochem. 2009, 47, 491–503. [Google Scholar] [CrossRef]
- Cutignano, A.; Lamari, N.; D’ippolito, G.; Manzo, E.; Cimino, G.; Fontana, A. Lipoxygenase products in marine diatoms: A concise analytical method to explore the functional potential of oxylipins. J. Phycol. 2011, 47, 233–243. [Google Scholar] [CrossRef]
- Adelfi, M.G.; Vitale, R.M.; d’Ippolito, G.; Nuzzo, G.; Gallo, C.; Amodeo, P.; Manzo, E.; Pagano, D.; Landi, S.; Picariello, G.; et al. Patatin-like lipolytic acyl hydrolases and galactolipid metabolism in marine diatoms of the genus Pseudo-nitzschia. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2019, 1864, 181–190. [Google Scholar] [CrossRef]
- Li, J.; Han, D.; Wang, D.; Ning, K.; Jia, J.; Wei, L.; Jing, X.; Huang, S.; Chen, J.; Li, Y.; et al. Choreography of Transcriptomes and Lipidomes of Nannochloropsis Reveals the Mechanisms of Oil Synthesis in Microalgae. Plant Cell 2014, 26, 1645–1665. [Google Scholar] [CrossRef]
- De la Noüe, J.; Laliberté, G.; Proulx, D. Algae and waste water. J. Appl. Phycol. 1992, 4, 247–254. [Google Scholar] [CrossRef]
- Mathimani, T.; Pugazhendhi, A. Utilization of algae for biofuel, bio-products and bio-remediation. Biocatal. Agric. Biotechnol. 2019, 17, 326–330. [Google Scholar] [CrossRef]
- Kuo, C.-M.; Chen, T.-Y.; Lin, T.-H.; Kao, C.-Y.; Lai, J.-T.; Chang, J.-S.; Lin, C.-S. Cultivation of Chlorella sp. GD using piggery wastewater for biomass and lipid production. Bioresour. Technol. 2015, 194, 326–333. [Google Scholar] [CrossRef]
- Kim, H.-C.; Choi, W.J.; Chae, A.N.; Park, J.; Kim, H.J.; Song, K.G. Evaluating integrated strategies for robust treatment of high saline piggery wastewater. Water Res. 2016, 89, 222–231. [Google Scholar] [CrossRef]
- Hemalatha, M.; Sravan, J.S.; Yeruva, D.K.; Venkata Mohan, S. Integrated ecotechnology approach towards treatment of complex wastewater with simultaneous bioenergy production. Bioresour. Technol. 2017, 242, 60–67. [Google Scholar] [CrossRef]
- Rugnini, L.; Costa, G.; Congestri, R.; Antonaroli, S.; Sanità di Toppi, L.; Bruno, L. Phosphorus and metal removal combined with lipid production by the green microalga Desmodesmus sp.: An integrated approach. Plant Physiol. Biochem. 2018, 125, 45–51. [Google Scholar] [CrossRef]
- Sharma, B.; Dangi, A.K.; Shukla, P. Contemporary enzyme based technologies for bioremediation: a review. J. Environ. Manage. 2018, 210, 10–22. [Google Scholar] [CrossRef]
- Thatoi, H.; Das, S.; Mishra, J.; Rath, B.P.; Das, N. Bacterial chromate reductase, a potential enzyme for bioremediation of hexavalent chromium: a review. J. Environ. Manage. 2014, 146, 383–399. [Google Scholar] [CrossRef]
- Sivaperumal, P.; Kamala, K.; Rajaram, R. Bioremediation of industrial waste through enzyme producing marine microorganisms. In Advances in Food and Nutrition Research; Academic Press: Cambridge, MA, USA, 2017; Volume 80, pp. 165–179. ISBN 9780128095874. [Google Scholar]
- Joutey, N.T.; Sayel, H.; Bahafid, W.; El Ghachtouli, N. Mechanisms of hexavalent chromium resistance and removal by microorganisms. Rev. Environ. Contam. Toxicol. 2015, 233, 45–69. [Google Scholar]
- Yen, H.-W.; Chen, P.-W.; Hsu, C.-Y.; Lee, L. The use of autotrophic Chlorella vulgaris in chromium (VI) reduction under different reduction conditions. J. Taiwan Inst. Chem. Eng. 2017, 74, 1–6. [Google Scholar] [CrossRef]
- Han, X.; Wong, Y.S.; Wong, M.H.; Tam, N.F.Y. Biosorption and bioreduction of Cr(VI) by a microalgal isolate, Chlorella miniata. J. Hazard. Mater. 2007, 146, 65–72. [Google Scholar] [CrossRef]
- Jácome-Pilco, C.R.; Cristiani-Urbina, E.; Flores-Cotera, L.B.; Velasco-García, R.; Ponce-Noyola, T.; Cañizares-Villanueva, R.O. Continuous Cr(VI) removal by Scenedesmus incrassatulus in an airlift photobioreactor. Bioresour. Technol. 2009, 100, 2388–2391. [Google Scholar] [CrossRef]
- Pradhan, D.; Sukla, L.B.; Mishra, B.B.; Devi, N. Biosorption for removal of hexavalent chromium using microalgae Scenedesmus sp. J. Clean. Prod. 2019, 209, 617–629. [Google Scholar] [CrossRef]
- Raczynska, J.E.; Vorgias, C.E.; Antranikian, G.; Rypniewski, W. Crystallographic analysis of a thermoactive nitrilase. J. Struct. Biol. 2011, 173, 294–302. [Google Scholar] [CrossRef]
- Park, J.M.; Trevor Sewell, B.; Benedik, M.J. Cyanide bioremediation: the potential of engineered nitrilases. Appl. Microbiol. Biotechnol. 2017, 101, 3029–3042. [Google Scholar] [CrossRef]
- Cirulis, J.T.; Scott, J.A.; Ross, G.M. Management of oxidative stress by microalgae. Can. J. Physiol. Pharmacol. 2013, 91, 15–21. [Google Scholar] [CrossRef]
- Lauritano, C.; Orefice, I.; Procaccini, G.; Romano, G.; Ianora, A. Key genes as stress indicators in the ubiquitous diatom Skeletonema marinoi. BMC Genomics 2015, 16, 411. [Google Scholar] [CrossRef]
- Couto, N.; Wood, J.; Barber, J. The role of glutathione reductase and related enzymes on cellular redox homoeostasis network. Free Radic. Biol. Med. 2016, 95, 27–42. [Google Scholar] [CrossRef]
- Rogato, A.; Amato, A.; Iudicone, D.; Chiurazzi, M.; Ferrante, M.I.; d’Alcalà, M.R. The diatom molecular toolkit to handle nitrogen uptake. Mar. Genomics 2015, 24, 95–108. [Google Scholar] [CrossRef]
- Suzuki, S.; Yamaguchi, H.; Nakajima, N.; Kawachi, M. Raphidocelis subcapitata (=Pseudokirchneriella subcapitata) provides an insight into genome evolution and environmental adaptations in the Sphaeropleales. Sci. Rep. 2018, 8, 8058. [Google Scholar] [CrossRef]
- Sauser, K.R.; Liu, J.K.; Wong, T.-Y. Identification of a copper-sensitive ascorbate peroxidase in the unicellular green alga Selenastrum capricornutum. Biometals 1997, 10, 163–168. [Google Scholar] [CrossRef]
- Morelli, E.; Scarano, G. Copper-induced changes of non-protein thiols and antioxidant enzymes in the marine microalga Phaeodactylum tricornutum. Plant Sci. 2004, 167, 289–296. [Google Scholar] [CrossRef]
- Álvarez, R.; del Hoyo, A.; García-Breijo, F.; Reig-Armiñana, J.; del Campo, E.M.; Guéra, A.; Barreno, E.; Casano, L.M. Different strategies to achieve Pb-tolerance by the two Trebouxia algae coexisting in the lichen Ramalina farinacea. J. Plant Physiol. 2012, 169, 1797–1806. [Google Scholar] [CrossRef]
- Kennedy, J.; Margassery, L.M.; Morrissey, J.P.; O’Gara, F.; Dobson, A.D.W. Metagenomic strategies for the discovery of novel enzymes with biotechnological application from marine ecosystems. Mar. Enzym. Biocatal. 2013, 109–130. [Google Scholar]
- Trincone, A. Enzymatic processes in marine biotechnology. Mar. Drugs 2017, 15, 93. [Google Scholar] [CrossRef]
- Raut, N.; Al-Balushi, T.; Panwar, S.; Vaidya, R.S.; Shinde, G.B. Microalgal Biofuel. In Biofuels-Status and Perspective; InTech: London, UK, 2015. [Google Scholar]
- Ruiz, J.; Olivieri, G.; de Vree, J.; Bosma, R.; Willems, P.; Reith, J.H.; Eppink, M.H.M.; Kleinegris, D.M.M.; Wijffels, R.H.; Barbosa, M.J. Towards industrial products from microalgae. Energy Environ. Sci. 2016, 9, 3036–3043. [Google Scholar] [CrossRef]
- Gifuni, I.; Pollio, A.; Safi, C.; Marzocchella, A.; Olivieri, G. Current Bottlenecks and Challenges of the Microalgal Biorefinery. Trends Biotechnol. 2018, 37, 242–252. [Google Scholar] [CrossRef]
- Lauritano, C.; Ferrante, M.I.; Rogato, A. Marine Natural Products from Microalgae: An -Omics Overview. Mar. Drugs 2019, 17, 269. [Google Scholar] [CrossRef]
- Mardis, E.R. A decade’s perspective on DNA sequencing technology. Nature 2011, 470, 198–203. [Google Scholar] [CrossRef]
| Ref. | Enzymes | Microalgae | Strain Source | Microalgal Class | Main Results |
|---|---|---|---|---|---|
| [39] | ∆6-Desaturase | Phaeodactylum tricornutum | M | BA | Neutral lipid production enhanced and increase of EPA content |
| [41] | acetyl-CoA synthetase | Chlamydomonas reinhardtii | F | CH | Increase in neutral lipid production |
| [27] | acyl-CoA diacylglycerol acyltransferase 1 | Chlorella ellipsoidea | F | TR | Sequence identification and function of TAG accumultation characterized |
| [28] | acyl-CoA diacylglycerol acyltransferase 1A | Nannochloropsis oceanica | M | EU | Increase in TAGs production both in nitrogen-replete and -deplete conditions |
| [36] | acyl-CoA diacylglycerol acyltransferase 2 | Chlamydomonas reinhardtii | F | CH | No TAGs overproduction |
| [35] | acyl-CoA diacylglycerol acyltransferase 2 | Chlamydomonas reinhardtii | F | CH | Five DGAT2 homologous genes identification and the overexpression of CrDGAT2-1 and CrDGAT2-5 resulting in a significant increase in lipid production |
| [33] | acyl-CoA diacylglycerol acyltransferase 2 | Nannochloropsis oceanica | M | EU | Increase in neutral lipid production |
| [32] | acyl-CoA diacylglycerol acyltransferase 2 | Neochloris oleoabundans | F | CH | Change of lipid profile |
| [29] | acyl-CoA diacylglycerol acyltransferase 2 | Ostreococcus tauri | M | MA | Gene identification and enzyme characterization in heterologous systems |
| [30] | acyl-CoA diacylglycerol acyltransferase 2 | Phaeodactylum tricornutum | M | BA | Increase in neutral lipid production with enrichment EPA-PUFAs content |
| [31] | acyl-CoA diacylglycerol acyltransferase 2 | Thalassiosira pseudonana | M | CO | Increase in TAGs production with focus on the intracellular enzyme localization |
| [34] | acyl-CoA diacylglycerol acyltransferase 2A, 2C, 2D | Nannochloropsis oceanica | M | EU | Differential DGAT2 isoforms expression in different engineered strains with individual specialized lipid profiles |
| [47] | fatty acid photodecarboxylase | Chlorella variabilis | F | TR | Enzyme identification and alkane synthase activity tested |
| [38] | glucose-6-phosphate dehydrogenase | Phaeodactylum tricornutum | M | BA | Modest increase in neutral lipid production with a lipid composition switch from polyunsaturated to monounsaturated |
| [42] | glucose-6-phosphate dehydrogenase; phosphogluconate dehydrogenase | Fistulifera solaris | M | BA | Slight increase in TAGs production |
| [40] | glycerol-3-phosphate acyltransferase 1, 2 | Cyanidioschyzon merolae | F | CY | Significant increase in TAGs production |
| [43] | phospholipase A2 | Chlamydomonas reinhardtii | F | CH | Increase in TAGs production |
| [45] | stearoyl-ACP desaturase | Chlamydomonas reinhardtii | F | CH | Production of TAGs enriched in stearic acid |
| [46] | UDP-glucose pyrophosphorylase, glycerol-3-phosphate dehydrogenase, enoyl-ACP reductase, long chain acyl-CoA elongase, putative palmitoyl-protein thioesterase, Ω-3 fatty acid desaturase and ∆-12-fatty acid desaturase | Phaeodactylum tricornutum | M | BA | Significant increase in TAGs production (45-fold increase for UDP-glucose pyrophosphorylase mutant) |
| [37] | wax esther synthase/acyl-CoA diacylglycerol acyltransferase | Phaeodactylum tricornutum | M | BA | Increase in neutral lipids and wax esters production |
| Patent Code (Year) | Enzymes | Microalgae | Strain Source | Microalgal Class | Notes |
| CN107299090A (2017) | wax esther synthase/acyl-CoA diacylglycerol acyltransferase | Phaeodactylum tricornutum | M | BA | Neutral lipids and wax esters production enhanced |
| CN101289659A (2010) | ∆6-Desaturase | Nannochloropsis spp. | M | EU | The enzyme sequence was identified and the enzyme characterized in bacterial systems |
| KR101829048B1 (2018) | Ω6-Desaturase | Arctic Chlamydomonas sp. ArF0006 | F | CH | The enzyme sequence was identified and the enzyme characterized in bacterial systems |
| Reference | Enzymes | Microalgae | Strain Source | Microalgal Class | Main Results |
|---|---|---|---|---|---|
| [78] | β-carotene hydroxylase | Dunaliella salina | M | CH | Increase in violaxanthin and zeaxanthin production |
| [81] | β-carotene oxygenase | Chlorella zofingiensis | S | TR | Increase in canthaxanthin, zeaxanthin and astaxanthin production under combined nitrogen starvation and high light stress |
| [56] | l-asparaginase | Chlamidomonas spp. | F | CH | Enzyme purified and tested |
| [57] | l-asparaginase | Chlorella vulgaris | F, S | TR | Screening of 40 microalgal isolates searching for new l-asparaginase sources |
| [82] | lycopene-β-cyclase | Chlamidomonas reinhardtii | F | CH | Increased gene expression under high light stress |
| [99] | lypolitic acid hydrolase 1 | Pseudo-nitzschia multistrata, Pseudo-nitzschia arenysensis | M | BA | Enzyme finding, characterization and retrieval of homologous sequences in other diatoms |
| [69] | non-ribosomal peptide synthase | Karenia brevis | M | DY | Gene cluster identification and chloroplastic localization identification |
| [68] | polycystin-1, Lipoxygenase, Alpha-Toxin/lipoxygenase homology 2 | Tetraselmis suecica | M | CR | Three putative enzyme sequences identification and in silico domain assessment and structure prediction |
| [88] | phytoene desaturase | Dunaliella salina | M | CH | Increase in phytoene production |
| [84] | phytoene synthase | Chlamidomonas reinhardtii | F | CH | Increase in violaxanthin (2.0 fold) and lutein (2.2-fold) production |
| [86] | phytoene desaturase | Haematococcus pluvialis | F | CH | Increase in astaxanthin production |
| [87] | phytoene desaturase | Haematococcus pluvialis | F | CH | Increase in astaxanthin production |
| [64] | polyketide synthase | Amphidinium carterae | M | DY | Identification of a transcript coding for type I PKS β-ketosynthase domain |
| [65] | polyketide synthase | Azadinium spinosum | M | DY | Identification of type I PKS domains using a combination of genomic and transcriptomic anayses |
| [66] | polyketide synthase | Gamberdiscus polynesiensis, Gamberdiscus excentricus | M | CH | Identification of transcripts coding for type I and type II PKS domains |
| [67] | polyketide synthase | Karenia brevis | M | DY | Identification of eight transcripts, six of which coding for type I PKS catalytic domains |
| [90] | zeaxanthin epoxidase | Chlamydomonas reinhardtii | F | CH | Increase in zeaxanthin production of 47-fold |
| Reference | Enzymes | Microalgae | Strain Source | Microalgal Class | Main Results |
|---|---|---|---|---|---|
| [111] | Putative Cr Reductase | Chlorella vulgaris | F | TR | Enzymatic Cr conversion (from Cr(VI) to Cr(III)) detected |
| [68] | Nitrilase | Tetraselmis suecica | M | CR | Putative enzyme sequence identification |
| [122] | Putative ascorbate peroxidase | Selenastrum capricornutum | F | CH | High sensitivity to Cu concentration activity |
| [123] | superoxide-dismutase, catalase, glutathione reductase | Phaeodactylum tricornutum | M | BA | Higher detected enzymatic activity after Cu accumulation |
| [124] | superoxide-dismutase, catalase, glutathione reductase, ascorbate peroxidase | Trebouxia 1 (TR1), Trebouxia 9 (TR9) | L | TR | Constitutive higher enzymatic activity detected in TR1, while exposed to Pb brings TR1 and TR9 enzymatic activities to comparable levels |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vingiani, G.M.; De Luca, P.; Ianora, A.; Dobson, A.D.W.; Lauritano, C. Microalgal Enzymes with Biotechnological Applications. Mar. Drugs 2019, 17, 459. https://doi.org/10.3390/md17080459
Vingiani GM, De Luca P, Ianora A, Dobson ADW, Lauritano C. Microalgal Enzymes with Biotechnological Applications. Marine Drugs. 2019; 17(8):459. https://doi.org/10.3390/md17080459
Chicago/Turabian StyleVingiani, Giorgio Maria, Pasquale De Luca, Adrianna Ianora, Alan D.W. Dobson, and Chiara Lauritano. 2019. "Microalgal Enzymes with Biotechnological Applications" Marine Drugs 17, no. 8: 459. https://doi.org/10.3390/md17080459
APA StyleVingiani, G. M., De Luca, P., Ianora, A., Dobson, A. D. W., & Lauritano, C. (2019). Microalgal Enzymes with Biotechnological Applications. Marine Drugs, 17(8), 459. https://doi.org/10.3390/md17080459








