Bromoperoxidase Producing Bacillus spp. Isolated from the Hypobranchial Glands of A Muricid Mollusc Are Capable of Tyrian Purple Precursor Biogenesis
Abstract
1. Introduction
2. Results
2.1. Bacterial Isolation
2.2. Molecular Identification of Cultivated Bacteria
2.3. Putative Bromoperoxidase Gene Screening by PCR
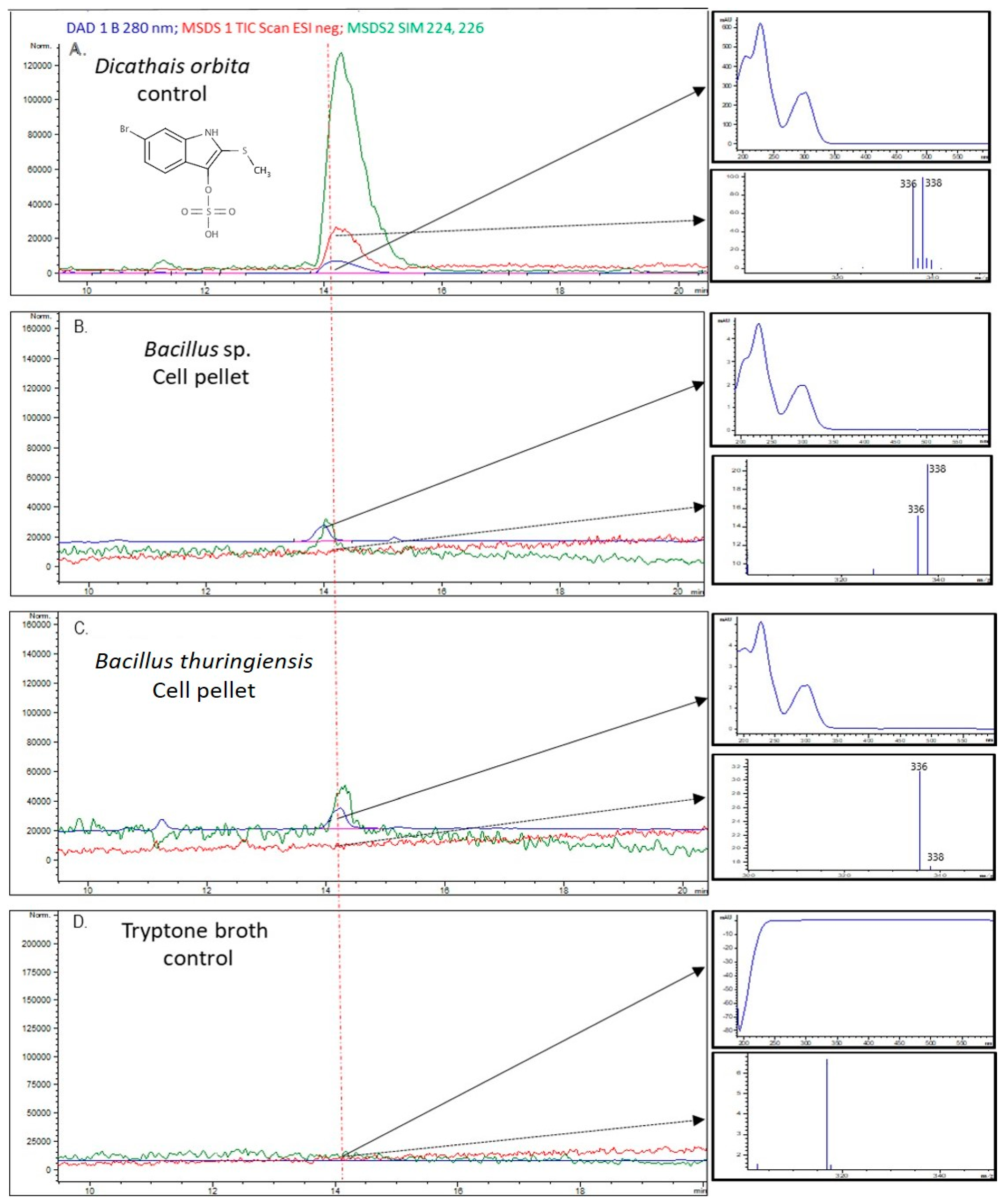
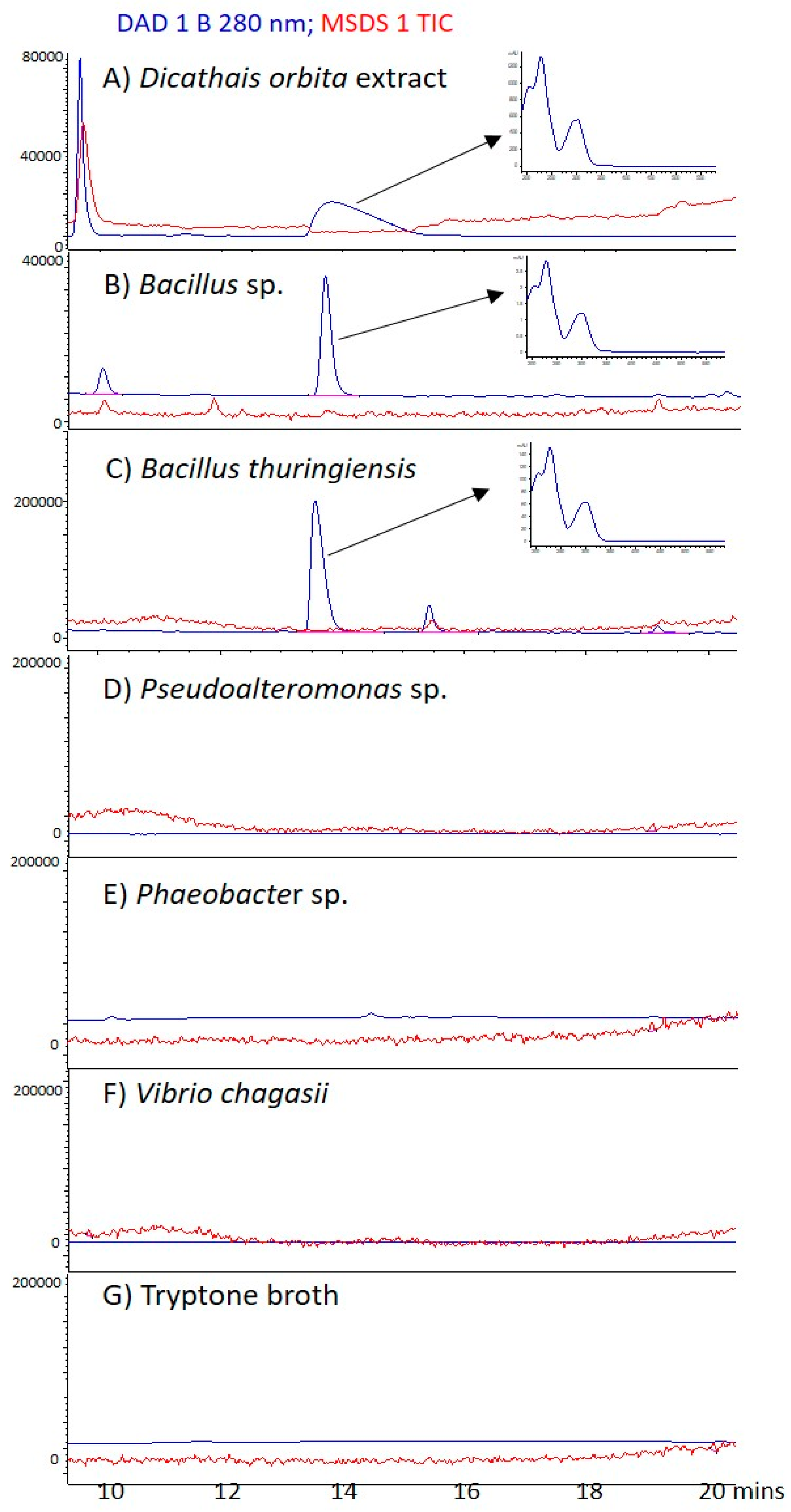
2.4. Bacterial Extract Analysis for Brominated Compounds by Liquid Chromatography–Mass Spectrometry
3. Discussion
4. Materials and Methods
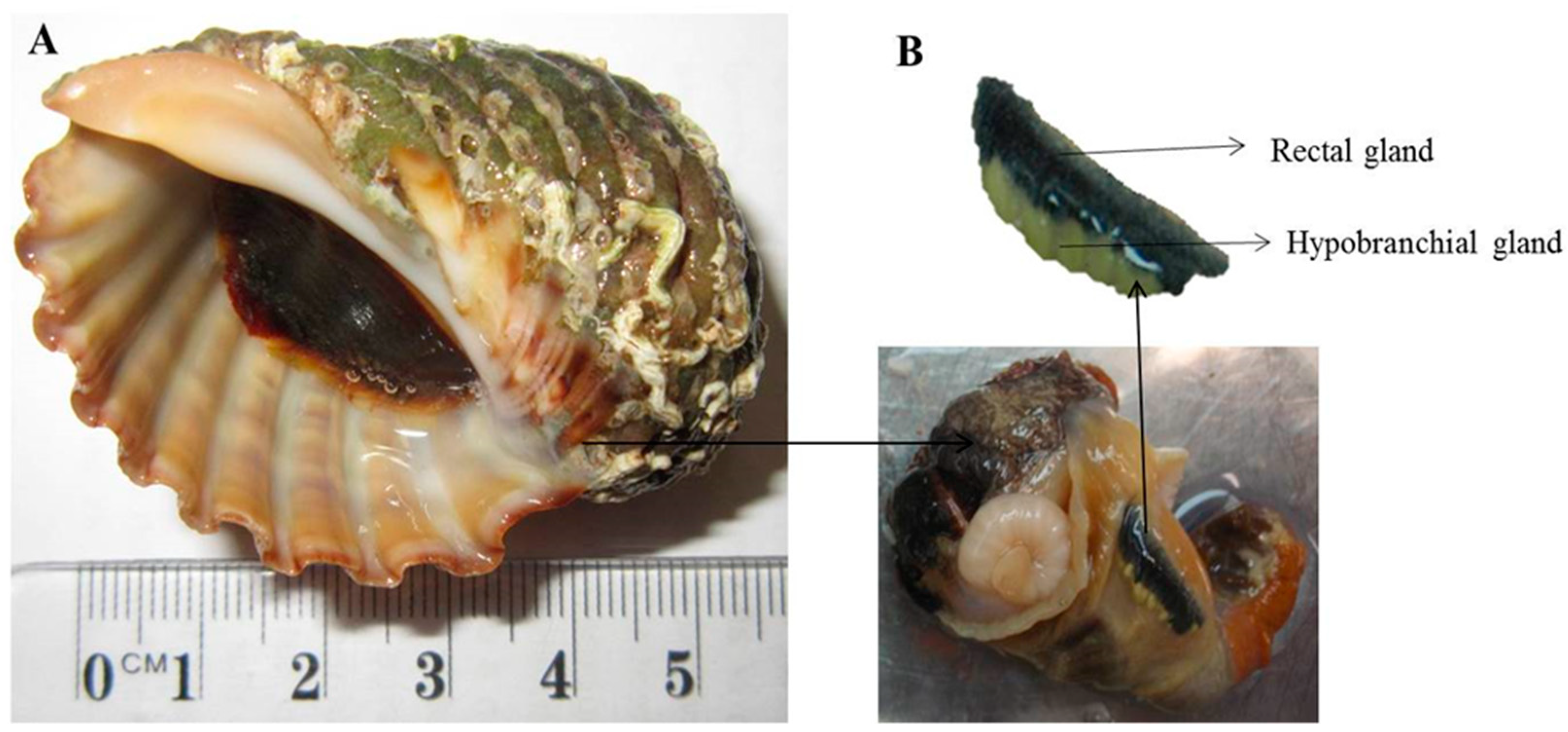
4.1. Sample Collection, Preparation and Culturing
4.2. 16S rRNA Sequencing of Bacterial Isolates
4.3. Bromoperoxidase Gene Screening
4.4. Liquid Chromatography–Mass Spectrometry (LC–MS) Analysis of Bacterial Extracts
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pawlik, J.R. Marine invertebrate chemical defenses. Chem. Rev. 1993, 93, 1911–1922. [Google Scholar] [CrossRef]
- Engel, S.; Jensen, P.R.; Fenical, W. Chemical ecology of marine microbial defense. J. Chem. Ecol. 2002, 28, 1971–1985. [Google Scholar] [CrossRef]
- Boltovskoy, D.; Cataldo, D.H. Population dynamics of Limnoperna fortunei, an invasive fouling mollusc, in the lower Parana river (Argentina). Biofouling 1999, 14, 255–263. [Google Scholar] [CrossRef]
- Haefner, B. Drugs from the deep: Marine natural products as drug candidates. Drug Discov. Today 2003, 8, 536–544. [Google Scholar] [CrossRef]
- Molinski, T.F.; Dalisay, D.S.; Lievens, S.L.; Saludes, J.P. Drug development from marine natural products. Nat. Rev. Drug. Discov. 2009, 8, 69–85. [Google Scholar] [CrossRef]
- Donia, M.; Hamann, M.T. Marine natural products and their potential applications as anti-infective agents. Lancet. Infect. Dis. 2003, 3, 338–348. [Google Scholar] [CrossRef]
- Nagle, D.G.; Zhou, Y.D. Marine natural products as inhibitors of hypoxic signaling in tumors. Phytochem. Rev. 2009, 8, 415–429. [Google Scholar] [CrossRef]
- Malaker, A.; Ahmad, S.A.I. Therapeutic potency of anticancer peptides derived from marine organism. Int. J. Eng. Appl. Sci. 2013, 2, 82–94. [Google Scholar]
- Prota, G. Nitrogenous pigments in marine invertebrates. Nat. Prod. Rep. 2012, 3, 141–174. [Google Scholar]
- Westley, C.; Benkendorff, K. Sex-specific tyrian purple genesis: Precursor and pigment distribution in the reproductive system of the marine mollusc, Dicathais orbita. J. Chem. Ecol. 2008, 34, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Benkendorff, K.; Westley, C.B.; Gallardo, C.S. Observations on the production of purple pigments in the egg capsules, hypobranchial and reproductive glands from seven species of Muricidae (Gastropoda: Mollusca). Invertebr. Reprod. Dev. 2004, 46, 93–102. [Google Scholar] [CrossRef]
- Pereira, D.M.; Valentao, P.; Andrade, P.B. Marine natural pigments: Chemistry, distribution and analysis. Dyes Pigment 2014, 111, 124–134. [Google Scholar] [CrossRef]
- Benkendorff, K.; Rudd, D.; Nongmaithem, B.D.; Liu, L.; Young, F.; Edwards, V.; Avila, C.; Abbott, C.A. Are the traditional medical uses of Muricidae molluscs substantiated by their pharmacological properties and bioactive compounds? Mar. Drugs 2015, 13, 5237–5275. [Google Scholar] [CrossRef] [PubMed]
- Nongmaithem, B.D.; Mouatt, P.; Smith, J.; Rudd, D.; Russell, M.; Sullivan, C.; Benkendorff, K. Volatile and bioactive compounds in opercula from Muricidae molluscs supports their use in ceremonial incense and traditional medicines. Sci. Rep. 2017, 7, 17404. [Google Scholar] [CrossRef] [PubMed]
- Kalaitzaki, A.; Vafiadou, A.; Frony, A.; Reese, D.; Drivaliari, A.; Liritzis, I. Po-pu-re: Workshops, use and archaeometric analysis in pre-roman central eastern mediterranean. MAA 2017, 17. [Google Scholar] [CrossRef]
- Sterman, B.; Taubes-Sterman, J. The Rarest Blue; Lyons Press: Guilford, CO, USA, 2012. [Google Scholar]
- Cooksey, C. Tyrian purple: The first four thousand years. Sci. Prog 2013, 96, 171–186. [Google Scholar] [CrossRef]
- Dutta, S.; Roychoudhary, S.; Sarangi, B.K. Effect of different physico-chemical parameters for natural indigo production during fermentation of indigofera plant biomass. Biotech. 2017, 7, 322. [Google Scholar] [CrossRef]
- Comlekcioglu, N.; Efe, L.; Karaman, S. Extraction of indigo from some Isatis species and dyeing standardization using low-technology methods. Braz. Arch. Biol. Technol. 2015, 58, 96–102. [Google Scholar] [CrossRef]
- Lim, H.K.; Chung, E.J.; Kim, J.C.; Choi, G.J.; Jang, K.S.; Chung, Y.R.; Cho, K.Y.; Lee, S.W. Characterization of a forest soil metagenome clone that confers indirubin and indigo production on Escherichia coli. Appl. Environ. Microbiol. 2005, 71, 7768–7777. [Google Scholar] [CrossRef]
- Qu, Y.; Pi, W.; Ma, F.; Zhou, J.; Zhang, X. Influence and optimization of growth substrates on indigo formation by a novel isolate Acinetobacter sp. Pp-2. Bioresour. Technol. 2010, 101, 4527–4532. [Google Scholar] [CrossRef] [PubMed]
- Qu, Y.; Zhang, X.; Ma, Q.; Ma, F.; Zhang, Q.; Li, X.; Zhou, H.; Zhou, J. Indigo biosynthesis by Comamonas sp. Mq. Biotechnol. Lett. 2012, 34, 353–357. [Google Scholar] [CrossRef]
- Benkendorff, K. Natural product research in the Australian marine invertebrate Dicathais orbita. Mar. Drugs 2013, 11, 1370–1398. [Google Scholar] [CrossRef]
- KremnerPigments. Tyrian Purple, Genuine. Available online: https://shop.kremerpigments.com/en/pigments/kremer-made-and-historic-pigments/4915/tyrian-purple-genuine (accessed on 2 November 2018).
- Ahmad, T.; Rudd, D.; Smith, J.; Kotiw, M.; Mouatt, P.; Seymour, L.; Liu, L.; Benkendorff, K. Anti-inflammatory activity and structure-activity relationships of brominated indoles from a marine mollusc. Mar. Drugs 2017, 15, 133. [Google Scholar] [CrossRef]
- Ahmad, T.B.; Rudd, D.; Benkendorff, K.; Mahdi, L.K.; Pratt, K.-A.; Dooley, L.; Wei, C.; Kotiw, M. Brominated indoles from a marine mollusc inhibit inflammation in a murine model of acute lung injury. PLoS ONE 2017, 12, e0186904. [Google Scholar] [CrossRef]
- Esmaeelian, B.; Abbott, C.A.; Le Leu, R.K.; Benkendorff, K. 6-bromoisatin found in muricid mollusc extracts inhibits colon cancer cell proliferation and induces apoptosis, preventing early stage tumor formation in a colorectal cancer rodent model. Mar. Drugs 2014, 12, 17–35. [Google Scholar] [CrossRef]
- Gaboriaud-Kolar, N.; Myrianthopoulos, V.; Vougogiannopoulou, K.; Gerolymatos, P.; Horne, D.A.; Jove, R.; Mikros, E.; Nam, S.; Skaltsounis, A.-L. Natural-based indirubins display potent cytotoxicity toward wild-type and t315i-resistant leukemia cell lines. J. Nat. Prod 2016, 79, 2464–2471. [Google Scholar] [CrossRef]
- Gaboriaud-Kolar, N.; Vougogiannopoulou, K.; Skaltsounis, A.-L. Indirubin derivatives: A patent review (2010–present). Expert Opin. Ther. Pat. 2015, 25, 583–593. [Google Scholar] [CrossRef]
- Esmaeelian, B.; Benkendorff, K.; Le Leu, R.K.; Abbott, C.A. Simultaneous assessment of the efficacy and toxicity of marine mollusc–derived brominated indoles in an in vivo model for early stage colon cancer. Integr. Cancer Ther. 2017, 1–15. [Google Scholar] [CrossRef]
- Esmaeelian, B.; Benkendorff, K.; Johnston, M.R.; Abbott, C.A. Purified brominated indole derivatives from Dicathais orbita induce apoptosis and cell cycle arrest in colorectal cancer cell lines. Mar.Drugs 2013, 11, 3802–3822. [Google Scholar] [CrossRef]
- Cooksey, C.J. Tyrian purple: 6,6′-dibromoindigo and related compounds. Molecules (Basel, Switzerland) 2001, 6, 736–769. [Google Scholar] [CrossRef]
- Cooksey, C.J.; Sinclair, R.S. Colour variations in Tyrian purple dyeing. Dyes in History and Archaeology 2005, 20, 127–135. [Google Scholar]
- Baten, A.; Ngangbam, A.; Waters, D.; Benkendorff, K. Transcriptome of the Australian mollusc Dicathais orbita provides insights into the biosynthesis of indoles and choline esters. Mar. Drugs 2016, 14, 135. [Google Scholar] [CrossRef]
- Jannun, R.; Coe, E.L. Bromoperoxidase from the marine snail, Murex-trunculus. Comp. Biochem. Physiol. B 1987, 88, 917–922. [Google Scholar] [CrossRef]
- Westley, C.B.; Lewis, M.C.; Benkendorff, K. Histomorphology of the hypobranchial gland in Dicathais orbita (Gmelin, 1791) (Neogastropoda: Muricidae). J. Molluscan Stud. 2010, 76, 186–195. [Google Scholar] [CrossRef]
- Westley, C.; Benkendorff, K. The distribution of precursors and biosynthetic enzymes required for Tyrian purple genesis in the hypobranchial gland, gonoduct, an egg masses of Dicathais orbita (Gmelin, 1791) (Neogastropoda: Muricidae). Nautilus 2009, 123, 148–153. [Google Scholar]
- Berrue, F.; Withers, S.T.; Haltli, B.; Withers, J.; Kerr, R.G. Chemical screening method for the rapid identification of microbial sources of marine invertebrate-associated metabolites. Mar. Drugs 2011, 9, 369–381. [Google Scholar] [CrossRef]
- Lane, A.L.; Moore, B.S. A sea of biosynthesis: Marine natural products meet the molecular age. Nat. Prod. Rep. 2011, 28, 411–428. [Google Scholar] [CrossRef]
- Ngangbam, A.K.; Baten, A.; Waters, D.L.E.; Whalan, S.; Benkendorff, K. Characterization of bacterial communities associated with the Tyrian purple producing gland in a marine gastropod. PLoS ONE 2015, 10(10), e0140725. [Google Scholar] [CrossRef]
- Ngangbam, A.K.; Waters, D.L.E.; Whalan, S.; Baten, A.; Benkendorff, K. Indole producing bacteria from the biosynthetic organs of muricid mollusc could contribute to Tyrian purple production. J. Shellfish Res. 2015, 34, 443–454. [Google Scholar] [CrossRef]
- Lingens, F. Purification and molecular and catalytic properties of bromoperoxidase from Streptomyces phaeochromogenes. J. Gen. Microbiol. 1985, 131, 1911–1916. [Google Scholar]
- Van Pee, K.H.; Sury, G.; Lingens, F. Purification and properties of a nonheme bromoperoxidase from Streptomyces aureofaciens. Biol. Chem. Hoppe. Seyler 1987, 368, 1225–1232. [Google Scholar] [CrossRef]
- Zeiner, R.; van Pee, K.H.; Lingens, F. Purification and partial characterization of multiple bromoperoxidases from Streptomyces griseus. J. Gen. Microbiol. 1988, 134, 3141–3149. [Google Scholar] [CrossRef]
- Knoch, M.; Van Pee, K.H.; Vining, L.C.; Lingens, F. Purification, properties and immunological detection of a bromoperoxidase-catalase from Streptomyces venezuelae and from a chloramphenicol-nonproducing mutant. J. Gen. Microbiol. 1989, 135, 2493–2502. [Google Scholar] [CrossRef]
- Wiesner, W.; van Pee, K.-H.; Lingens, F. Purification and properties of bromoperoxidase from Pseudomonas pyrrocinia. Biol. Chem. Hoppe. Seyler 1985, 366, 1085–1092. [Google Scholar] [CrossRef]
- Beleneva, I.; Kukhlevskii, A. Characterization of Vibrio gigantis and Vibrio pomeroyi isolated from invertebrates of Peter the Great Bay, Sea of Japan. Microbiology 2010, 79, 402–407. [Google Scholar] [CrossRef]
- Alcaide, E.; Amaro, C.; Todoli, R.; Oltra, R. Isolation and characterization of Vibrio parahaemolyticus causing infection in Iberian toothcarp Aphanius iberus. Dis. Aquat. Organ. 1999, 35, 77–80. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Patten, C.L.; Glick, B.R. Role of Pseudomonas putida indoleacetic acid in development of the host plant root system. Appl. Environ. Microbiol. 2002, 68, 3795–3801. [Google Scholar] [CrossRef] [PubMed]
- Benkendorff, K. Aquaculture and the production of pharmaceuticals and nutraceuticals. In New Technologies in Aquaculture: Improving Production Efficiency, Quality and Environmental Management; Burnell, G., Allan, G., Eds.; Woodhead Publishing: Cambridge, UK, 2009; pp. 866–891. [Google Scholar]
- Sipkema, D.; Osinga, R.; Schatton, W.; Mendola, D.; Tramper, J.; Wijffels, R.H. Large-scale production of pharmaceuticals by marine sponges: Sea, cell, or synthesis? Biotechnol. Bioeng. 2005, 90, 201–222. [Google Scholar] [CrossRef]
- Romano, G.; Costantini, M.; Sansone, C.; Lauritano, C.; Ruocco, N.; Ianora, A. Marine microorganisms as a promising and sustainable source of bioactive molecules. Mar. Environ. Res 2017, 128, 58–69. [Google Scholar] [CrossRef]
- Perez, L.B.; Fenical, W. Chapter 8 - Accessing marine microbial diversity for drug discovery. In Microbial Resources; Kurtboke, I., Ed.; Academic Press, 2017; pp. 169–187. [Google Scholar]
- Joint, I.; Muhling, M.; Querellou, J. Culturing marine bacteria - an essential prerequisite for biodiscovery. Microb. Biotechnol. 2010, 3, 564–575. [Google Scholar] [CrossRef]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev. 1995, 59, 143–169. [Google Scholar] [PubMed]
- Overmann, J.; Abt, B.; Sikorski, J. Present and future of culturing bacteria. Annu. Rev. Microbiol 2017, 71, 711–730. [Google Scholar] [CrossRef] [PubMed]
- Gulder, T.A.; Moore, B.S. Chasing the treasures of the sea - bacterial marine natural products. Curr. Opin. Microbiol. 2009, 12, 252–260. [Google Scholar] [CrossRef]
- Adnani, N.; Chevrette, M.G.; Adibhatla, S.N.; Zhang, F.; Yu, Q.; Braun, D.R.; Nelson, J.; Simpkins, S.W.; McDonald, B.R.; Myers, C.L. Coculture of marine invertebrate-associated bacteria and interdisciplinary technologies enable biosynthesis and discovery of a new antibiotic, keyicin. ACS Chem. Biol 2017, 12, 3093–3102. [Google Scholar] [CrossRef]
- Brinkmann, C.; Marker, A.; Kurtboke, D. An overview on marine sponge-symbiotic bacteria as unexhausted sources for natural product discovery. Diversity 2017, 9, 40. [Google Scholar] [CrossRef]
- Rizzo, C.; Lo Giudice, A. Marine invertebrates: Underexplored sources of bacteria producing biologically active molecules. Diversity 2018, 10, 52. [Google Scholar] [CrossRef]
- Vartoukian, S.R.; Palmer, R.M.; Wade, W.G. Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol Lett. 2010, 309, 1–7. [Google Scholar] [CrossRef]
- Pham, V.H.; Kim, J. Cultivation of unculturable soil bacteria. Trends Biotechnol. 2012, 30, 475–484. [Google Scholar] [CrossRef] [PubMed]
- Davis, K.E.; Joseph, S.J.; Janssen, P.H. Effects of growth medium, inoculum size, and incubation time on culturability and isolation of soil bacteria. Appl. Environ. Microbiol. 2005, 71, 826–834. [Google Scholar] [CrossRef]
- Montalvo, N.F.; Davis, J.; Vicente, J.; Pittiglio, R.; Ravel, J.; Hill, R.T. Integration of culture-based and molecular analysis of a complex sponge-associated bacterial community. PLoS ONE 2014, 9, e90517. [Google Scholar] [CrossRef] [PubMed]
- Xing, M.; Hou, Z.; Qu, Y.; Liu, B. Enhancing the culturability of bacteria from the gastrointestinal tract of farmed adult turbot Scophthalmus maximus. Chin. J. Oceanol. Limnol. 2014, 32, 316–325. [Google Scholar] [CrossRef]
- Li, Z.Y.; Liu, Y. Marine sponge Craniella austrialiensis-associated bacterial diversity revelation based on 16s rdna library and biologically active actinomycetes screening, phylogenetic analysis. Lett. Appl. Microbiol. 2006, 43, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Valles-Regino, R.; Mouatt, P.; Rudd, D.; Yee, L.H.; Benkendorff, K. Extraction and quantification of bioactive Tyrian purple precursors: A comparative and validation study from the hypobranchial gland of a muricid Dicathais orbita. Molecules 2016, 21, 1672. [Google Scholar] [CrossRef] [PubMed]
- Imming, P.; Imhof, I.; Zentgraf, M. An improved synthetic procedure for 6, 6′-dibromoindigo (Tyrian purple). Synth. Commun. 2001, 31, 3721–3727. [Google Scholar] [CrossRef]
- Schatz, P.F. Indigo and Tyrian purple-in nature and in the lab. J. Chem. Educ. 2001, 78, 1442. [Google Scholar] [CrossRef]
- Wolk, J.L.; Frimer, A.A. A simple, safe and efficient synthesis of Tyrian purple (6, 6′-dibromoindigo). Molecules 2010, 5561–5580. [Google Scholar] [CrossRef]
- Flores-Garza, R.; Gonzalez, A.V.; Flores-Rodriguez, P.; Cruz-Ramirez, N.L. Density, sex ratio, size, weight, and recruitment of Plicopurpura pansa (Gastropoda: Muricidae) in Costa Rica, Guerrero, Mexico. OJMS 2012, 2, 157. [Google Scholar] [CrossRef]
- Read, T.D.; Peterson, S.N.; Tourasse, N.; Baillie, L.W.; Paulsen, I.T.; Nelson, K.E.; Tettelin, H.; Fouts, D.E.; Eisen, J.A.; Gill, S.R. The genome sequence of Bacillus anthracis Ames and comparison to closely related bacteria. Nature 2003, 423, 81–86. [Google Scholar] [CrossRef]
- Itoh, N.; Morinaga, N.; Kouzai, T. Purification and characterization of a novel metal-containing nonheme bromoperoxidase from Pseudomonas putida. Biochim. Biophys. Acta. 1994, 1207, 208–216. [Google Scholar] [CrossRef]
- Van Pee, K.H.; Lingens, F. Purification of bromoperoxidase from Pseudomonas aureofaciens. J. Bacteriol. 1985, 161, 1171–1175. [Google Scholar]
- Butler, A.; Carter-Franklin, J.N. The role of vanadium bromoperoxidase in the biosynthesis of halogenated marine natural products. Nat. Prod. Rep. 2004, 21, 180–188. [Google Scholar] [CrossRef]
- Martinez, J.S.; Carroll, G.L.; Tschirret-Guth, R.A.; Altenhoff, G.; Little, R.D.; Butler, A. On the regiospecificity of vanadium bromoperoxidase. J. Am. Chem. Soc. 2001, 123, 3289–3294. [Google Scholar] [CrossRef]
- Jiang, Z.; Boyd, K.G.; Mearns-Spragg, A.; Adams, D.R.; Wright, P.C.; Burgess, J.G. Two diketopiperazines and one halogenated phenol from cultures of the marine bacterium, Pseudoalteromonas luteoviolacea. Nat. Prod. Lett. 2000, 14, 435–440. [Google Scholar] [CrossRef]
- Feher, D.; Barlow, R.; McAtee, J.; Hemscheidt, T.K. Highly brominated antimicrobial metabolites from a marine Pseudoalteromonas sp. J. Nat. Prod. 2010, 73, 1963–1966. [Google Scholar] [CrossRef]
- Leyton, Y.; Riquelme, C. Marine Bacillus spp. Associated with the egg capsule of Concholepas concholepas (common name “loco”) have an inhibitory activity toward the pathogen Vibrio parahaemolyticus. Microb. Ecol. 2010, 60, 599–605. [Google Scholar] [CrossRef]
- Pandey, A.; Naik, M.M.; Dubey, S.K. Organic metabolites produced by Vibrio parahaemolyticus strain an3 isolated from goan mullet inhibit bacterial fish pathogens. Afr. J. Biotechnol. 2010, 9, 7134–7140. [Google Scholar]
- Thompson, F.; Thompson, C.; Hoste, B.; Vandemeulebroecke, K.; Gullian, M.; Swings, J. Vibrio fortis sp. nov. and Vibrio hepatarius sp. nov., isolated from aquatic animals and the marine environment. Int. J. Syst. Evol. Microbiol. 2003, 53, 1495–1501. [Google Scholar] [CrossRef]
- Pujalte, M.-J.; Garay, E. Proposal of Vibrio mediterranei sp. nov.: A new marine member of the genus Vibrio. Int. J. Syst. Bacteriol. 1986, 36, 278–281. [Google Scholar] [CrossRef]
- Jaksic, S.; Uhitil, S.; Petrak, T.; Bazulic, D.; Gumhalter Karolyi, L. Occurrence of Vibrio spp. In sea fish, shrimps and bivalve molluscs harvested from Adriatic Sea. Food Control 2002, 13, 491–493. [Google Scholar] [CrossRef]
- Gomez-Gil, B.; Tron-Mayen, L.; Roque, A.; Turnbull, J.F.; Inglis, V.; Guerra-Flores, A.L. Species of Vibrio isolated from hepatopancreas, haemolymph and digestive tract of a population of healthy juvenile Penaeus vannamei. Aquaculture 1998, 163, 1–9. [Google Scholar] [CrossRef]
- Phatarpekar, P.V.; Kenkre, V.D.; Sreepada, R.A.; Desai, U.M.; Achuthankutty, C.T. Bacterial flora associated with larval rearing of the giant freshwater prawn, Macrobrachium rosenbergii. Aquaculture 2002, 203, 279–291. [Google Scholar] [CrossRef]
- Parija, S.C. Culture media. In Textbook of Microbiology & Immunology; Elsevier Health Sciences, 2014. [Google Scholar]
- Marinho, P.R.; Moreira, A.P.; Pellegrino, F.L.; Muricy, G.; Bastos Mdo, C.; Santos, K.R.; Giambiagi-deMarval, M.; Laport, M.S. Marine Pseudomonas putida: A potential source of antimicrobial substances against antibiotic-resistant bacteria. Mem. Inst. Oswaldo Cruz 2009, 104, 678–682. [Google Scholar] [CrossRef] [PubMed]
- Lilles, M.M. Halogenation Enzymes in Bacteria Associated with the Red-banded Acorn Worm, Ptychodera jamaicensis. Master’s Thesis, San Jose State University, San Jose, CA, USA, Fall. 2011. [Google Scholar]
| Closest Match and Accession Number | Agar Media 1 | GenBank Accession Number | Length (Base Pair) | Identity (%) | Gram Stain | Hypobranchial Gland Preparation 2 | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| MA * | TCBS | CA | MAH | BA | ||||||
| Vibrio sp. (KM369853.1) | + | + | - | - | + | KR338844 | 1149 | 100 | Gram- | HGVS |
| Vibrio chagasii (NR117891.1) | + | + | + | + | + | KR338845 | 1083 | 99 | Gram- | HGDS |
| Vibrio alginolyticus (KF886646.1) | + | + | + | + | + | KR338846 | 1115 | 99 | Gram- | HGDS |
| Vibrio sp. (KP126921.1) | + | + | - | - | + | KR338847 | 510 | 100 | Gram- | HGVS |
| Aliivibrio sp. (FR744854.1) | + | + | - | - | + | KR338848 | 1087 | 99 | Gram- | HGVS |
| Vibrio azureus (JF412237.1) | + | + | - | - | + | KR338849 | 1119 | 99 | Gram- | HGVS |
| Vibrio pomeroyi (KM014017.1) | + | + | - | - | + | KR338850 | 1090 | 99 | Gram- | HGVS |
| Vibrio sp. (KM369851.1) | + | + | + | + | - | KR338851 | 1004 | 100 | Gram- | HGDS |
| Phaeobacter sp. (GQ906799.1) | + | + | - | + | - | KR338852 | 1044 | 100 | Gram- | HGDS |
| Vibrio sp. (GQ406789.1) | + | + | + | + | - | KR338853 | 1046 | 99 | Gram- | HGDS |
| Vibrio sp. NB0059, (KP770076.1) | + | + | + | + | + | KR338854 | 1096 | 100 | Gram- | HGDS |
| Shewanella sp. (JF825445.1) | + | + | + | + | + | KR338855 | 1058 | 98 | Gram- | HGDS |
| Vibrio mediterranei, (HF541948.1) | + | + | - | + | + | KR338856 | 1116 | 99 | Gram- | HGDS |
| Vibrio sp. (KF188532.1) | + | - | - | + | - | KR338857 | 1096 | 100 | Gram- | HGH |
| Vibrio sp. (KF188531.1) | + | - | - | + | - | KR338858 | 1066 | 99 | Gram- | HGH |
| Vibrio sp. (HG942391.1) | + | - | - | + | - | KR338859 | 626 | 99 | Gram- | HGH |
| Vibrio sp. (KP126921.1) | + | + | - | + | + | KR338860 | 951 | 97 | Gram- | HGVS |
| Vibrio sp. (KM369860.1) | + | + | - | + | + | KR338861 | 796 | 99 | Gram- | HGVS |
| Vibrio sp. (KM369853.1) | + | + | - | + | + | KR338862 | 1091 | 100 | Gram- | HGVS |
| Vibrio sp. (GQ406789.1) | + | + | - | + | + | KR338863 | 1079 | 100 | Gram- | HGVS |
| Halobacillus sp. (FM992846.1) | + | + | - | + | + | KR338864 | 1109 | 100 | Gram + | HGVS |
| Vibrio sp. (FJ457587.1) | + | + | - | + | + | KR338865 | 1004 | 100 | Gram- | HGVS |
| Vibrio sp. (KM369853.1) | + | + | - | + | + | KR338866 | 1038 | 99 | Gram- | HGVS |
| Vibrio splendidus (AB038030.1) | + | + | - | + | + | KR338867 | 1083 | 100 | Gram- | HGVS |
| Vibrio harveyi (KR003734.1) | + | + | - | + | + | KR338868 | 973 | 100 | Gram- | HGVS |
| Bacillus sp. (KJ756140.1) | + | - | - | + | - | KR338869 | 1034 | 100 | Gram + | HGH |
| Bacillus thuringiensis (KC355253.1) | + | - | - | + | - | KR855712 | 1180 | 99 | Gram + | HGH |
| Vibrio sp. (HF937138.1) | + | - | - | + | - | KR338870 | 866 | 99 | Gram- | HGH |
| Vibrio sp. (EU340847.1) | + | + | + | + | + | KR338871 | 1110 | 99 | Gram- | HGDS |
| Pseudoalteromonas sp. (KP301110.1) | + | + | + | + | + | KR338872 | 973 | 99 | Gram- | HGDS |
| Vibrio sp. (FJ457361.1) | + | + | - | - | + | KR338873 | 1056 | 99 | Gram- | HGVS |
| Vibrio jasicida (AB562594.1) | + | + | - | - | + | KR338874 | 907 | 99 | Gram- | HGVS |
| Bacterial Isolates | GenBank Accession Number | Length (Base Pair) | Identity (%) | Closest Match, Accession Number, Position and Protein ID |
|---|---|---|---|---|
| Bacillus sp. | KR338869 | 628 | 97 | Bacillus thuringiensis MC28- bromoperoxidase (CP003687.1); 2239980-2240816 and AFU13721.1 |
| Bacillus thuringiensis | KR855712 | 634 | 97 | Bacillus thuringiensis MC28- bromoperoxidase (CP003687.1); 2239980-2240816 and AFU13721.1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ngangbam, A.K.; Mouatt, P.; Smith, J.; Waters, D.L.E.; Benkendorff, K. Bromoperoxidase Producing Bacillus spp. Isolated from the Hypobranchial Glands of A Muricid Mollusc Are Capable of Tyrian Purple Precursor Biogenesis. Mar. Drugs 2019, 17, 264. https://doi.org/10.3390/md17050264
Ngangbam AK, Mouatt P, Smith J, Waters DLE, Benkendorff K. Bromoperoxidase Producing Bacillus spp. Isolated from the Hypobranchial Glands of A Muricid Mollusc Are Capable of Tyrian Purple Precursor Biogenesis. Marine Drugs. 2019; 17(5):264. https://doi.org/10.3390/md17050264
Chicago/Turabian StyleNgangbam, Ajit Kumar, Peter Mouatt, Joshua Smith, Daniel L. E. Waters, and Kirsten Benkendorff. 2019. "Bromoperoxidase Producing Bacillus spp. Isolated from the Hypobranchial Glands of A Muricid Mollusc Are Capable of Tyrian Purple Precursor Biogenesis" Marine Drugs 17, no. 5: 264. https://doi.org/10.3390/md17050264
APA StyleNgangbam, A. K., Mouatt, P., Smith, J., Waters, D. L. E., & Benkendorff, K. (2019). Bromoperoxidase Producing Bacillus spp. Isolated from the Hypobranchial Glands of A Muricid Mollusc Are Capable of Tyrian Purple Precursor Biogenesis. Marine Drugs, 17(5), 264. https://doi.org/10.3390/md17050264





