Dynamical Behavior of Disordered Regions in Disease-Related Proteins Revealed by Quasielastic Neutron Scattering
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Methods
3. Results
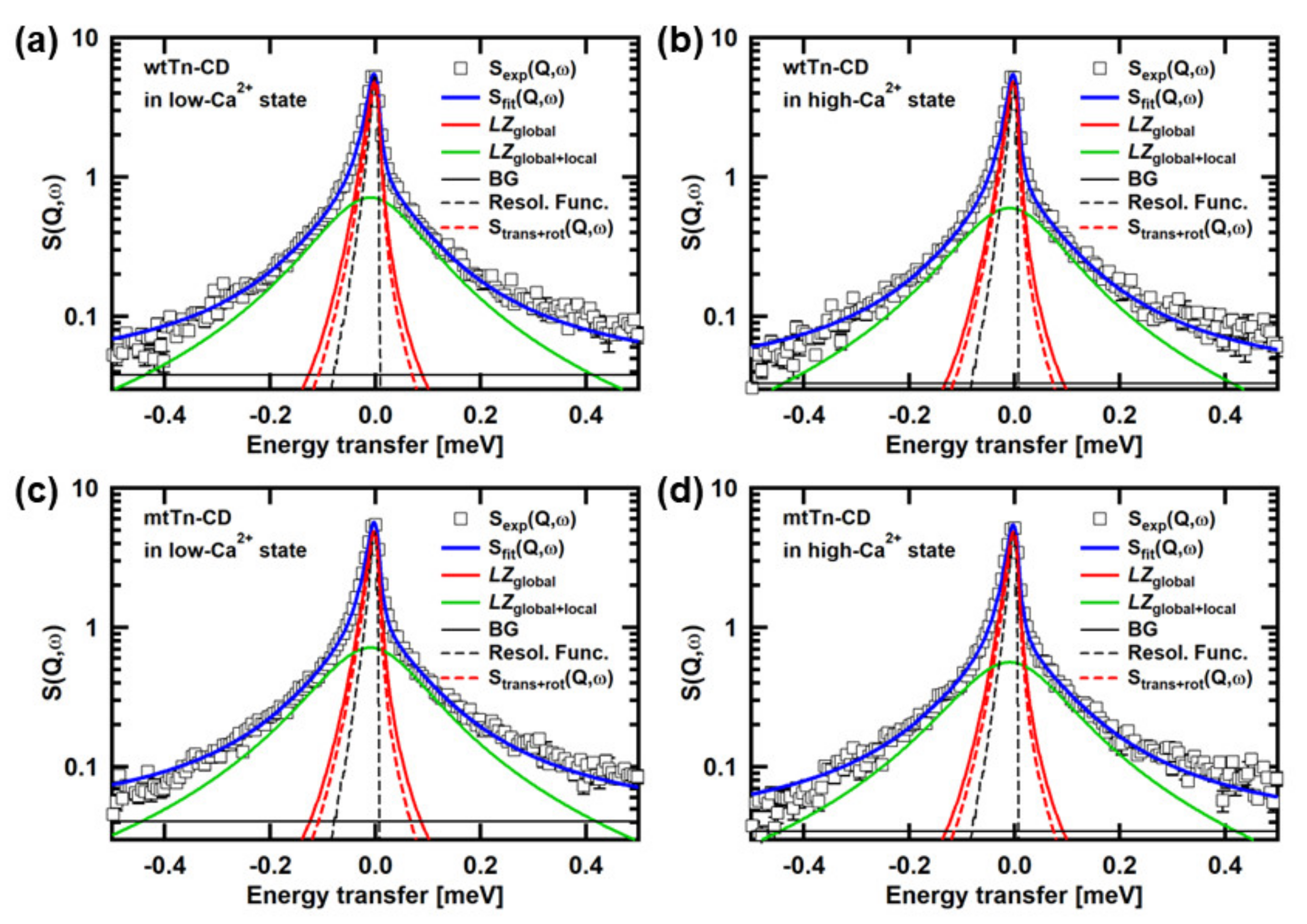
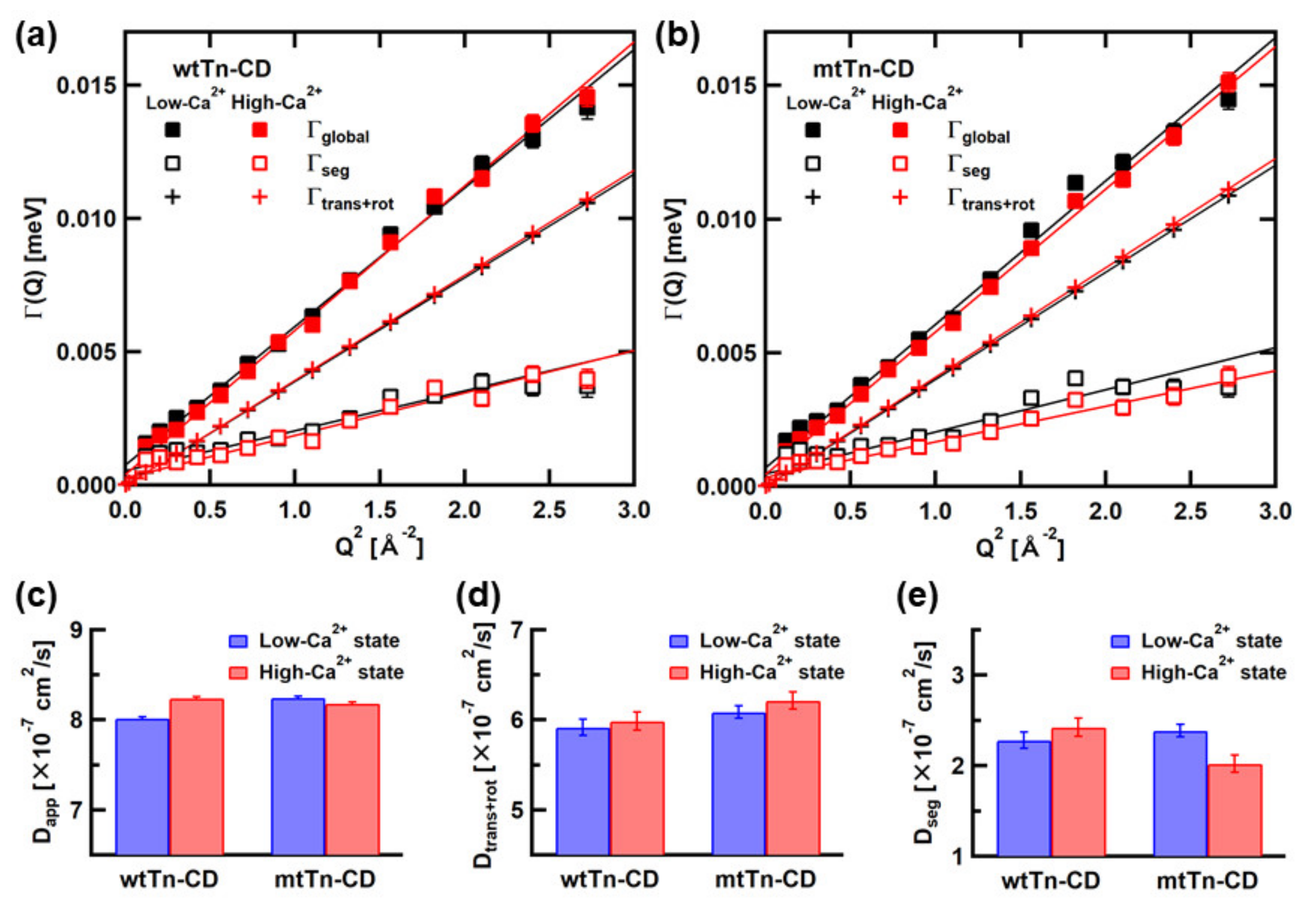
3.1. Analysis of the QENS Spectra of the Tn-CD
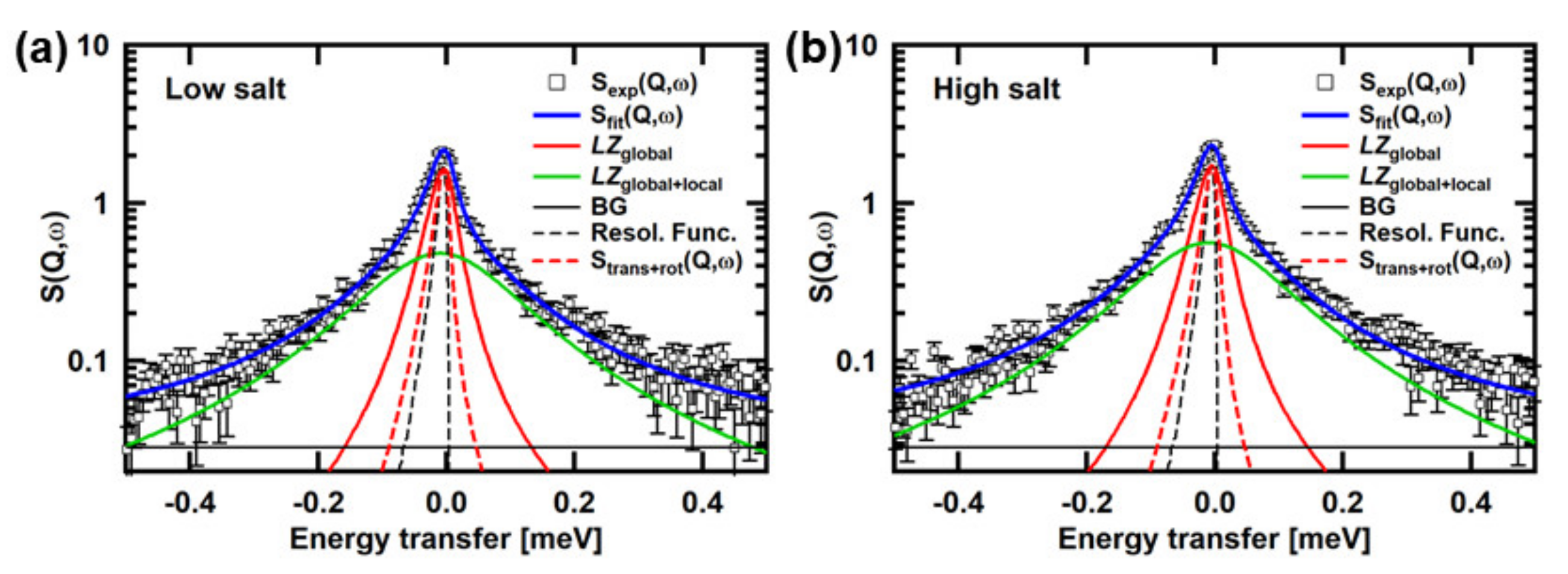
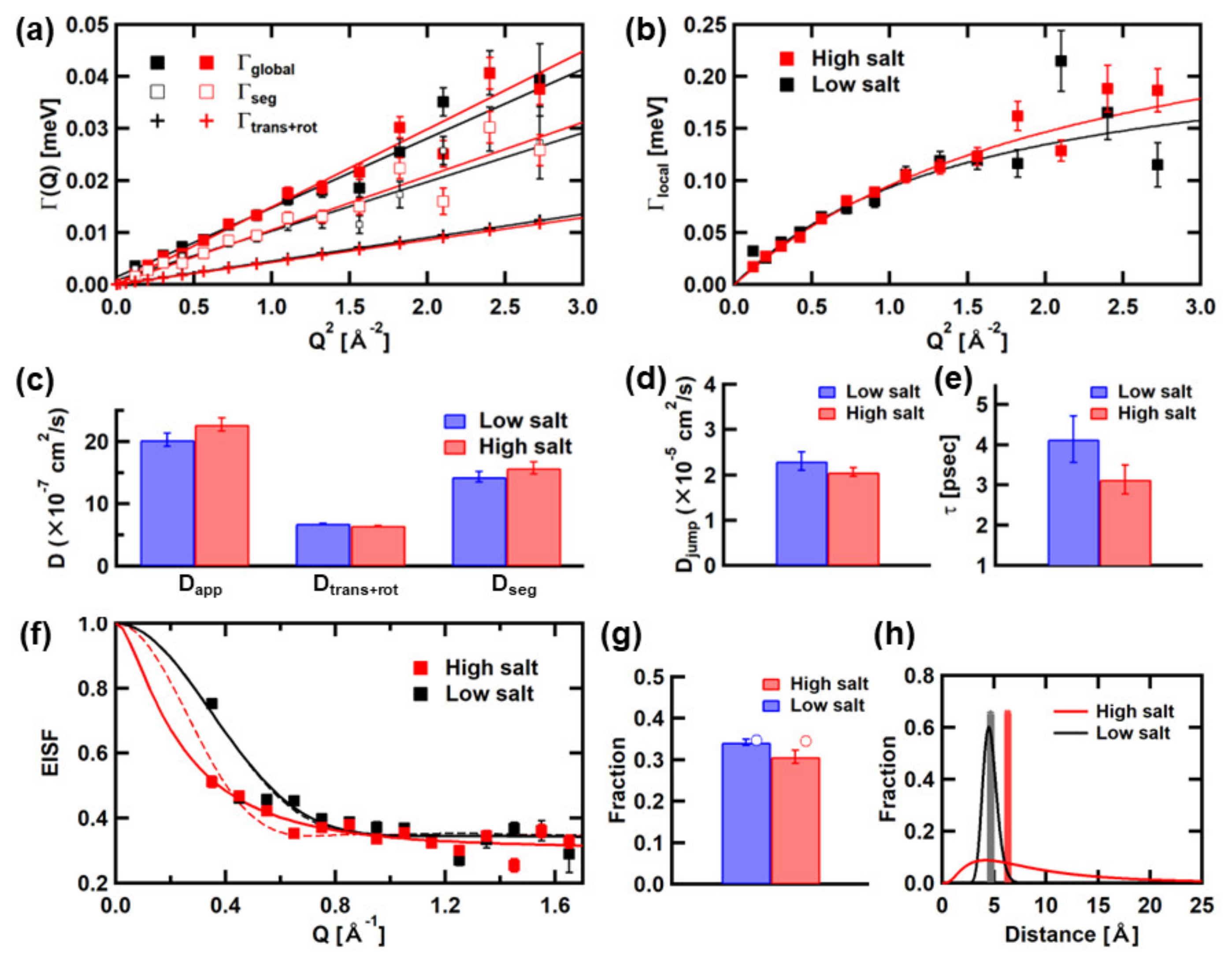
3.2. Analysis of the QENS Spectra of αSyn
4. Discussion
5. Conclusions
Funding
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kendrew, D.C.; Bodo, G.; Dintzis, H.M.; Parrish, R.G.; Wyckoff, H.; Phillips, D.C. A three-dimensional model of the myoglobin molecule obtained by X-ray analysis. Nature 1958, 181, 662–666. [Google Scholar] [CrossRef] [PubMed]
- Perutz, M.F.; Rossmann, M.G.; Cullis, A.F.; Muirhead, H.; Will, G.; North, A.C.T. Structure of haemoglobin. A three-dimensional Fourier synthesis at 5.5Å resolution obtained by X-ray analysis. Nature 1960, 185, 416–422. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically unstructured proteins: Re-assessing the proteins structure-function paradigm. J. Mol. Biol. 1999, 293, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Gillespie, J.R.; Fink, A.L. Why are “natively unfolded” proteins unstructured under physiologic conditions? Proteins 2000, 41, 415–427. [Google Scholar] [CrossRef]
- Dunker, A.K.; Lawson, J.D.; Brown, C.J.; Williams, R.M.; Romeo, P.; Oh, J.S.; Oldfield, C.J.; Campen, A.M.; Ratliff, C.M.; Hipps, K.W.; et al. Intrinsically disorderd protein. J. Mol. Graph. Model. 2001, 19, 26–59. [Google Scholar] [CrossRef]
- Tompa, P. Intrinsically unstructured proteins. Trends Biochem. Sci. 2002, 27, 509–516. [Google Scholar] [CrossRef]
- van der Lee, R.; Buljan, M.; Lang, B.; Weatheritt, R.J.; Daughdrill, G.W.; Dunker, A.K.; Fuxreiter, M.; Gough, J.; Gsponer, J.; Jones, D.T.; et al. Classification of intrinsically disordered regions and proteins. Chem. Rev. 2014, 114, 6589–6631. [Google Scholar] [CrossRef]
- Knowles, T.P.J.; Vendruscolo, M.; Dobson, C.M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 2014, 15, 384–396. [Google Scholar] [CrossRef]
- Uversky, V.N.; Oldfield, C.J.; Dunker, A.K. Intrinsically disordered proteins in human diseases: Introducing the D2 concept. Annu. Rev. Biophys. 2008, 37, 215–245. [Google Scholar] [CrossRef]
- Smith, J.C. Protein dynamics: Comparison of simulations with inelastic neutron scattering experiments. Q. Rev. Biophys. 1991, 24, 227–291. [Google Scholar] [CrossRef]
- Grimaldo, M.; Roosen-Runge, F.; Zhang, F.; Schreiber, F.; Seydel, T. Dynamics of proteins in solution. Q. Rev. Biophys. 2019, 52, e7, 1–63. [Google Scholar] [CrossRef]
- Fujiwara, S.; Chatake, T.; Matsuo, T.; Kono, F.; Tominaga, T.; Shibata, K.; Sato-Tomita, A.; Shibayama, N. Ligation-dependent picosecondd dynamics in human hemoglobin as revealed by quasielastic neutron scattering. J. Phys. Chem. B 2017, 121, 8069–8077. [Google Scholar] [CrossRef]
- Matsuo, T.; Tominaga, T.; Kono, F.; Shibata, K.; Fujiwara, S. Modulation of the picosecond dynamics of troponin by the cardiomyopathy-causing mutation K247R of tropoin T observed by quasielastic neutron scattering. BBA–Proteins Proteom. 2017, 1865, 1781–1789. [Google Scholar] [CrossRef]
- Fujiwara, S.; Kono, F.; Matsuo, T.; Sugimoto, Y.; Matsumoto, T.; Narita, A.; Shibata, K. Dynamic properties of human α-synuclein related to propensity to amyloid fibril formation. J. Mol. Biol. 2019, 431, 3229–3245. [Google Scholar] [CrossRef]
- Fujiwara, S.; Matsuo, T.; Sugimoto, Y.; Shibata, K. Segmental motions of proteins under non-native states evaluated using quasielastic neutron scatteing. J. Phys. Chem. Lett. 2019, 10, 7505–7509. [Google Scholar] [CrossRef]
- Ebashi, S.; Endo, M.; Otsuki, I. Control of muscle contraction. Q. Rev. Biophys. 1969, 2, 351–384. [Google Scholar] [CrossRef]
- Li, M.X.; Wang, X.; Sykes, B.D. Structural based insights into the role of troponin in cardiac muscle pathophysiology. J. Muscle Res. Cell Motil. 2004, 25, 559–579. [Google Scholar] [CrossRef]
- Harada, K.; Morimoto, S. Inherited cardiomyopathies as a troponin disease. Jpn. J. Physiol. 2004, 54, 307–318. [Google Scholar] [CrossRef]
- Maroteaus, L.; Campanelli, J.T.; Scheller, R.H. Synuclein: A neutron-specific protein localized to the nucleus and presynaptic nerve terminal. J. Neurosci. 1988, 8, 2804–2815. [Google Scholar] [CrossRef]
- Bendor, J.T.; Logan, T.P.; Edwards, R.H. The function of α-synuclein. Neuron 2013, 79, 1044–1066. [Google Scholar] [CrossRef]
- Breydo, L.; Wu, J.W.; Uversky, V.N. α-Synuclein misfolding and Parkinson’s disease. Biochim. Biophys. Acta 2012, 1822, 261–285. [Google Scholar] [CrossRef]
- García-Castro, M.; Reguero, J.R.; Batalla, A.; Díaz-Molina, B.; González, P.; Alvarez, V.; Cortina, A.; Cubero, G.I.; Coto, E. Hypertrophic cardiomyopathy: Low frequency of mutations in the β-myosin heavy chain (MYH7) and cardiac Troponin T (TNNT2) genes among Spanish patients. Clin. Chem. 2003, 49, 1279–1285. [Google Scholar] [CrossRef]
- Møller, D.V.; Andersen, P.S.; Hedley, P.; Ersbøl, M.K.; Bundgaard, H.; Moolman-Smook, J.; Christiansen, M.; Køber, L. The role of sarcomere gene mutations in patients with idiopathic dilated cardiomyopathy. Eur. J. Hum. Genet. 2009, 17, 1241–1249. [Google Scholar] [CrossRef]
- Matsumoto, F.; Maeda, K.; Chatake, T.; Maéda, Y.; Fujiwara, S. Functional aberration of myofibrils by cardiomyopathy mutations in the coiled-coil region of the troponin-core domain. Biochem. Biophys. Res. Commun. 2009, 382, 109–205. [Google Scholar] [CrossRef]
- Takeda, S.; Yamashita, A.; Maeda, K.; Maéda, Y. Structure of the core domain of human cardiac troponin in the Ca2+-saturated form. Nature 2003, 424, 35–41. [Google Scholar] [CrossRef]
- Shibata, K.; Takahashi, N.; Kawakita, Y.; Matsuura, M.; Yamada, T.; Tominaga, T.; Kambara, W.; Kobayashi, M.; Inamura, Y.; Nakatani, T.; et al. The Performance of TOF Near Backscattering Spectrometer DNA in MLF, J-PARC. JPS Conf. Proc. 2015, 8, 036022. [Google Scholar] [CrossRef]
- Pérez, J.; Zanotti, J.M.; Durand, D. Evolution of the internal dynamics of two globular proteins from dry powder to solution. Biophys. J. 1999, 77, 454–469. [Google Scholar] [CrossRef]
- Roosen-Runge, F.; Henning, M.; Zhang, F.; Jacobs, R.M.J.; Sztucki, M.; Schrober, H.; Seydel, T.; Shreiber, F. Protein self-diffusion in crowded solutions. Proc. Natl. Acad. Sci. USA 2011, 108, 11815–11820. [Google Scholar] [CrossRef]
- Matsuo, T.; Takeda, S.; Oda, T.; Fujiwara, S. Structures of the troponin core domain containing the cardiomyopathy-causing mutants studied by small-angle X-ray scattering. Biophys. Physicobiol. 2015, 12, 145–158. [Google Scholar] [CrossRef]
- Franke, D.; Svergun, D.I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J. Appl. Cryst. 2009, 42, 342–346. [Google Scholar] [CrossRef]
- Franke, D.; Petoukhov, M.V.; Konarev, P.V.; Panjkovich, A.; Tuukkanen, A.; Mertens, H.D.T.; Kikhney, A.G.; Hajizadeh, N.R.; Franklin, J.M.; Jeffries, C.M.; et al. ATSAS 2.8: A comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Cryst. 2017, 50, 1212–1225. [Google Scholar] [CrossRef] [PubMed]
- Ortega, A.; Amoros, D.; de la Torre, J.G. Prediction of hydrodynamic and other solution properties of rigid proteins from atomic and residue-level models. Biophys. J. 2011, 101, 892–898. [Google Scholar] [CrossRef] [PubMed]
- de la Torre, J.G.; del Rio, G.; Ortega, A. Improved calculation of rotational diffusion and intrinsic viscosity of bead models for macromolecules and nanoparticles. J. Phys. Chem. B 2007, 111, 955–961. [Google Scholar] [CrossRef] [PubMed]
- Cho, C.H.; Urquidi, J.; Singh, S.; Robinson, G.W. Thermal offset viscosities of liquid H2O, D2O, and T2O. J. Phys. Chem. B 1999, 103, 1991–1994. [Google Scholar] [CrossRef]
- Ostroff, A.G.; Snowden, J.B.S.; Woessner, D.E. Viscosities of protonated and deuterated water solutions of alkali metal chlorides. J. Phys. Chem. 1969, 73, 2784–2785. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera-A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Gibrat, G.; Assairi, F.L.; Blouquit, Y.; Craescu, C.T.; Bellissent-Funel, M.C. Biophysical study of thermal denaturation of apo-calmodulin: Dynamics of native and unfolded states. Biophys. J. 2008, 95, 5247–5256. [Google Scholar] [CrossRef][Green Version]
- Volino, F.; Dianoux, A.J. Neutron Incoherent-Scattering Law for Diffusion in a Potential of Spherical Symmetry: General Formalism and Application to Diffusion inside a Sphere. Mol. Phys. 1980, 41, 271–279. [Google Scholar] [CrossRef]
- Ando, T.; Skolnick, J. Crowding and hydrodynamic interactions likely dominate in vivo macromolecular motion. Proc. Natl. Acad. Sci. USA 2010, 107, 18457–18462. [Google Scholar] [CrossRef]
- Bucciarelli, S.; Myung, J.S.; Farago, B.; Das, S.; Vliegenthart, G.A.; Holderer, O.; Winkler, R.G.; Schurtenberger, P.; Gompper, G.; Stradner, A. Dramatic influence of patchy attractions on short-time protein diffusion under crowded conditions. Sci. Adv. 2016, 2, e1601432. [Google Scholar] [CrossRef]
- Lionberger, R.A.; Russel, W.B. High frequency modulus of hard sphere colloids. J. Rheol. 1994, 38, 1885–1908. [Google Scholar] [CrossRef]
- Lee, J.C.; Gray, H.B.; Winkler, J.R. Tertiary contact information in α-synuclein probed by electron transfer. J. Am. Chem. Soc. 2005, 127, 16388–16389. [Google Scholar] [CrossRef]
- Grupi, A.; Haas, E. Segmental conformational disorder and dynamics in the intrinsically disordered protein α-synuclein and its chain length dependence. J. Mol. Biol. 2011, 405, 1267–1283. [Google Scholar] [CrossRef]
- Woodard, J.; Srivastava, K.R.; Rahamim, G.; Grupi, A.; Hogan, S.; Witalka, D.J.; Nawrocki, G.; Haas, E.; Feig, M.; Lapidus, L.J. Intramolecular diffusion in α-synuclein: It depends on how you measure it. Biophys. J. 2018, 115, 1190–1199. [Google Scholar] [CrossRef]
- Wilkins, D.K.; Grimshaw, S.B.; Receveur, V.; Dobson, C.M.; Jones, J.A.; Smith, L.J. Hydrodynamic radii of native and denatured proteins measured by pulse field gradient NMR techniques. Biochemistry 1999, 38, 15431–16424. [Google Scholar] [CrossRef]
- Banchio, A.J.; Nägele, G. Shorttime transport properties in dense suspensions: From neutral to charge-stabilized colloidal spheres. J. Chem. Phys. 2008, 128, 104903. [Google Scholar] [CrossRef]
- Ameseder, F.; Radulescu, A.; Khaneft, M.; Lohstroh, W.; Stadler, A.M. Homogeneous and heterogeneous dynamics in native and denatured bovine serum albumin. Phys. Chem. Chem. Phys. 2018, 20, 5128–5139. [Google Scholar] [CrossRef]
- Hassani, A.N.; Haris, L.; Appel, M.; Seydel, T.; Stadler, A.M.; Kneller, G.R. Multiscale relaxation dynamics and diffusion of myelin basic protein in solution studied by quasielastic neutron scattering. J. Chem. Phys. 2022, 156, 025102. [Google Scholar] [CrossRef]



| wtTn-CD Low Ca2+ | wtTn-CD High Ca2+ | mtTn-CD Low Ca2+ | mtTn-CD High Ca2+ | αSyn Low Salt | αSyn High Salt | |
|---|---|---|---|---|---|---|
| DT (×10−7 cm2/s) | 4.64 ± 0.01 | 4.68 ± 0.02 | 4.79 ± 0.02 | 4.87 ± 0.02 | 5.59 ± 0.03 | 4.70 ± 0.04 |
| DR (×106 1/s) | 1.62 ± 0.13 | 1.64 ± 0.17 | 1.76 ± 0.16 | 1.86 ± 0.18 | 2.54 ± 0.03 | 1.63 ± 0.06 |
| Dapp (×10−7 cm2/s) | Dtrans+rot (×10−7 cm2/s) | Dseg (×10−7 cm2/s) | |
|---|---|---|---|
| wtTn-CD, Low-Ca2+ state | 8.02 ± 0.15 | 5.92 ± 0.02 | 2.28 ± 0.16 |
| wtTn-CD, High-Ca2+ state | 8.24 ± 0.14 | 5.99 ± 0.02 | 2.43 ± 0.15 |
| mtTn-CD, Low-Ca2+ state | 8.24 ± 0.14 | 6.09 ± 0.02 | 2.39 ± 0.16 |
| mtTn-CD, High-Ca2+ state | 8.18 ± 0.14 | 6.22 ± 0.02 | 2.02 ± 0.15 |
| Dapp (×10−7 cm2/s) | Dtrans+rot (×10−7 cm2/s) | Dseg (×10−7 cm2/s) | |
|---|---|---|---|
| Low salt | 20.3 ± 1.1 | 6.88 ± 0.01 | 14.3 ± 0.9 |
| High salt | 21.7 ± 0.9 | 6.49 ± 0.02 | 15.1 ± 0.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fujiwara, S. Dynamical Behavior of Disordered Regions in Disease-Related Proteins Revealed by Quasielastic Neutron Scattering. Medicina 2022, 58, 795. https://doi.org/10.3390/medicina58060795
Fujiwara S. Dynamical Behavior of Disordered Regions in Disease-Related Proteins Revealed by Quasielastic Neutron Scattering. Medicina. 2022; 58(6):795. https://doi.org/10.3390/medicina58060795
Chicago/Turabian StyleFujiwara, Satoru. 2022. "Dynamical Behavior of Disordered Regions in Disease-Related Proteins Revealed by Quasielastic Neutron Scattering" Medicina 58, no. 6: 795. https://doi.org/10.3390/medicina58060795
APA StyleFujiwara, S. (2022). Dynamical Behavior of Disordered Regions in Disease-Related Proteins Revealed by Quasielastic Neutron Scattering. Medicina, 58(6), 795. https://doi.org/10.3390/medicina58060795






