Control of Biofilms with the Fatty Acid Signaling Molecule cis-2-Decenoic Acid
Abstract
:1. Introduction
2. Jamming Bacterial Communication
3. Fatty Acid Signaling Systems
| Compound | New Nomenclature | Structure | Bacterial Species | Function | Reference |
|---|---|---|---|---|---|
| cis-11-Methyl-2-dodecenoic acid (DSF) | 11-Me-C12:Δ2 | 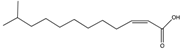 | Xanthomonas campestris, Xanthomonas oryzae, Stenotrophomonas maltophilia, Burkholderia multivorans | Virulence, biofilm formation, floc disaggregation, microcolony formation, tolerance to antibiotics, detoxification, hyphal growth inhibition | [96,97,98,99,104,105] |
| cis-2-Dodecenoic acid (BDSF) | C12:Δ2 |  | Burkholderia cenocepacia, Burkholderia lata Burkholderia stabilis Burkholderia vietnamiensis Burkholderia dolorosa Burkholderia ambifaria Burkholderia anthina Burkholderia pyrrocinia B. multivorans, X. oryzae | Virulence, hyphal growth inhibition | [97,104,106,107] |
| cis-2-Decenoic acid (cis-DA) | C10:Δ2 | 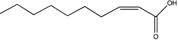 | Pseudomonas aeruginosa | Biofilm formation, biofilm dispersion, persister cell formation, persister cell awakening, tolerance to antimicrobials. | [19,65,108,109,110] |
| cis-2-Tetradecenoic acid | C14:Δ2 | 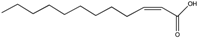 | Xylella fastidiosa | Virulence and aggregation | [100] |
| trans-2-Decenoic acid (SDSF) | C10:Δ2t | 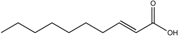 | Streptococcus mutans | Hyphal growth inhibition | [111] |
| cis-11-Methyldodeca-2,5-dienoic acid (CDSF) | 11-Me-C12:Δ2,5 | 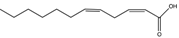 | B. multivorans, B. stabilis B. anthina, B. pyrrocinia, X. oryzae | Hyphal growth inhibition | [97,104] |
| 10-Methyldodecanoic acid | 10-Me-C12 |  | S. maltophilia | Stress tolerance and antibiotic tolerance | [105] |
| 11-Methyldodecanoic acid | 11-Me-C12 | 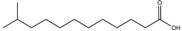 | S. maltophilia | Stress tolerance and antibiotic tolerance | [105] |
| 12-Methyltetradecanoic acid | 12-Me-C14 | 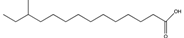 | Xylella fastidiosa | Virulence, biofilm formation, motility | [112,113] |
| 3-Hydroxypalmitic acid | 3OH-PAME |  | Ralstonia solanacearum | Virulence | [114] |
| Farnesoic acid | 3,7,11-Me-C12:Δ2t,6t,1°t | 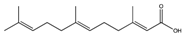 | Candida albicans | Inhibition of germ tube formation | [115] |
4. The Fatty Acid System in P. aeruginosa and Its Role in Biofilm Control
4.1. Dispersion Induced by cis-2-Decenoic Acid as Means of Biofilm Control
4.2. Persister Cell Control by Exposure to cis-DA
4.3. Prevention of Biofilm Formation by Exposure to cis-DA
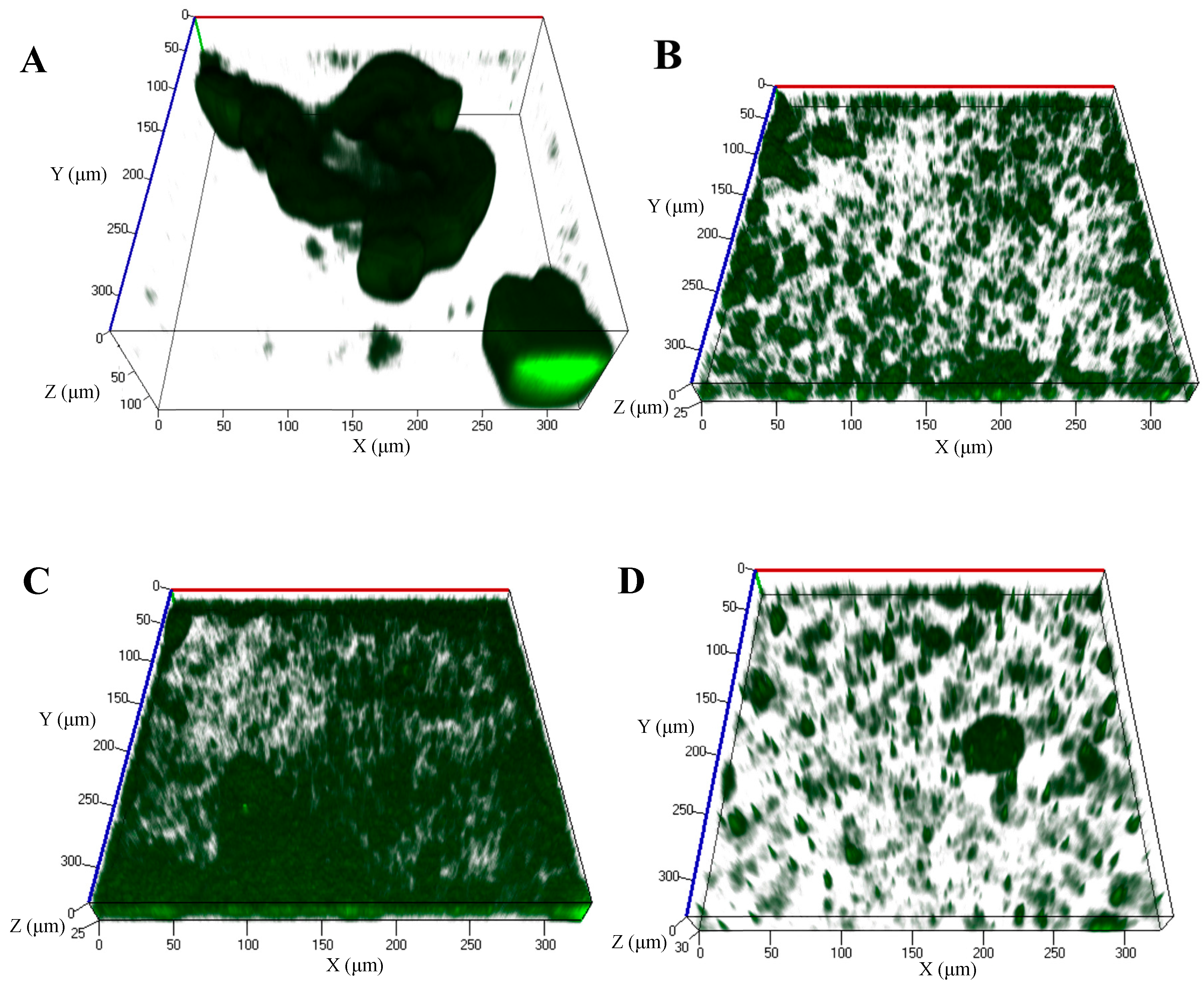
| Biofilm Structure Quantification | Control | cis-DA 1 nM | cis-DA 1 μM | cis-DA 1 mM |
|---|---|---|---|---|
| Average ± SD | Average ± SD | Average ± SD | Average ± SD | |
| Total biomass (μm3/μm2) | 20.66 ± 7.02 | 8.49 ± 2.38 | 12.29 ± 3.5 | 2.42 ± 0.86 |
| Average thickness (μm) | 26.46 ± 14.99 | 12.21 ± 3.73 | 19.37 ± 1.92 | 3.31 ± 1.19 |
| Maximum thickness (μm) | 146.13 ± 27.69 | 61.38 ± 13.22 | 50.80 ± 6.35 | 59.27 ± 15.98 |
| Roughness coefficient (dimensionless, range: zero-infinity) | 1.40 ± 0.38 | 0.90 ± 0.27 | 0.22 ± 0.05 | 1.86 ± 0.07 |
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- De Kievit, T.R.; Gillis, R.; Marx, S.; Brown, C.; Iglewski, B.H.; Kievit, T.R.D.E. Quorum-sensing genes in Pseudomonas aeruginosa biofilms: Their role and expression patterns. Appl. Environ. Microbiol. 2001, 67, 1865–1873. [Google Scholar] [CrossRef] [PubMed]
- Costerton, J.W. Introduction to biofilm. Int. J. Antimicrob. Agents 1999, 11, 217–219. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilms: Microbial life on surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Wimpenny, J.W. An overview of biofilms as functional communities. In Community Structure and Cooperation in Biofilms; Allison Gilbert, P., Lappin-Scott, H., Wilson, M.D., Eds.; Cambridge University Press: Cambridge, UK, 2000; pp. 1–24. [Google Scholar]
- Flemming, H.-C.; Neu, T.R.; Wozniak, D.J. The EPS matrix: The “house of biofilm cells”. J. Bacteriol. 2007, 189, 7945–7947. [Google Scholar] [CrossRef] [PubMed]
- Allesen-Holm, M.; Barken, K.B.; Yang, L.; Klausen, M.; Webb, J.S.; Kjelleberg, S.; Molin, S.; Givskov, M.; Tolker-Nielsen, T. A characterization of DNA release in Pseudomonas aeruginosa cultures and biofilms. Mol. Microbiol. 2006, 59, 1114–1128. [Google Scholar] [CrossRef] [PubMed]
- Sauer, K.; Camper, A.; Ehrlich, G.; Costerton, J.; Davies, D. Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J. Bacteriol. 2002, 184, 1140–1154. [Google Scholar] [CrossRef] [PubMed]
- Stoodley, P.; Sauer, K.; Davies, D.G.; Costerton, J.W. Biofilms as complex differentiated communities. Annu. Rev. Microbiol. 2002, 56, 187–209. [Google Scholar] [CrossRef] [PubMed]
- Allegrucci, M.; Hu, F.Z.; Shen, K.; Hayes, J.; Ehrlich, G.D.; Post, J.C.; Sauer, K. Phenotypic characterization of Streptococcus pneumoniae biofilm development. J. Bacteriol. 2006, 188, 2325–2335. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.; Kaplan, H.B.; Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol. 2000, 54, 49–79. [Google Scholar] [CrossRef] [PubMed]
- Watnick, P.; Kolter, R. Biofilm, city of microbes. J. Bacteriol. 2000, 182, 2675–2679. [Google Scholar] [CrossRef] [PubMed]
- Hinsa, S.M.; Espinosa-Urgel, M.; Ramos, J.L.; O’Toole, G.A. Transition from reversible to irreversible attachment during biofilm formation by Pseudomonas fluorescens WCS365 requires an ABC transporter and a large secreted protein. Mol. Microbiol. 2003, 49, 905–918. [Google Scholar] [CrossRef] [PubMed]
- Caiazza, N.C.; O’Toole, G.A. SadB is required for the transition from reversible to irreversible attachment during biofilm formation by Pseudomonas aeruginosa PA14. J. Bacteriol. 2004, 186, 4476–4485. [Google Scholar] [CrossRef] [PubMed]
- Chiang, P.; Burrows, L.L. Biofilm formation by hyperpiliated mutants of Pseudomonas aeruginosa. J. Bacteriol. 2003, 185, 2374–2378. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, I.W. The biofilm matrix—An immobilized but dynamic microbial environment. Trends Microbiol. 2001, 9, 222–227. [Google Scholar] [CrossRef]
- Gómez-Suárez, C.; Pasma, J.; van der Borden, A.J.; Wingender, J.; Flemming, H.-C.; Busscher, H.J.; van der Mei, H.C. Influence of extracellular polymeric substances on deposition and redeposition of Pseudomonas aeruginosa to surfaces. Microbiology 2002, 148, 1161–1169. [Google Scholar] [CrossRef] [PubMed]
- Davies, D.G.; Parsek, M.R.; Pearson, J.P.; Iglewski, B.H.; Costerton, J.W.; Greenberg, E.P. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 1998, 280, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Harmsen, M.; Yang, L.; Pamp, S.J.; Tolker-Nielsen, T. An update on Pseudomonas aeruginosa biofilm formation, tolerance, and dispersal. FEMS Immunol. Med. Microbiol. 2010, 59, 253–268. [Google Scholar] [CrossRef] [PubMed]
- Davies, D.G.; Marques, C.N.H. A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J. Bacteriol. 2009, 191, 1393–1403. [Google Scholar] [CrossRef] [PubMed]
- Kinniment, S.L.; Wimpenny, J.W. Measurements of the distribution of adenylate concentrations and adenylate energy charge across Pseudomonas aeruginosa biofilms. Appl. Environ. Microbiol. 1992, 58, 1629–1635. [Google Scholar] [PubMed]
- Schierholz, J.M.; Beuth, J.; Konig, D.; Nurnberger, A.; Pulverer, G. Antimicrobial substances and effects on sessile bacteria. Zentralblatt Bakteriol. 1999, 289, 165–177. [Google Scholar] [CrossRef]
- Gilbert, P.; Collier, P.J.; Brown, M.R. Influence of growth rate on susceptibility to antimicrobial agents: Biofilms, cell cycle, dormancy, and stringent response. Antimicrob. Agents Chemother. 1990, 34, 1865–1868. [Google Scholar] [CrossRef] [PubMed]
- Bagge, N.; Hentzer, M.; Andersen, J.B.; Ciofu, O.; Givskov, M.; Høiby, N. Dynamics and spatial distribution of b-lactamase expression in Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 2004, 48, 1168–1174. [Google Scholar] [CrossRef] [PubMed]
- Lenz, A.P.; Williamson, K.S.; Pitts, B.; Stewart, P.S.; Franklin, M.J. Localized gene expression in Pseudomonas aeruginosa biofilms. Appl. Environ. Microbiol. 2008, 74, 4463–4471. [Google Scholar] [CrossRef] [PubMed]
- Vuong, C.; Whitney, A.R.; Deleo, F.R. Polysaccharide intercellular adhesin (PIA) protects Staphylococcus epidermidis against major components of the human innate immune system. Cell. Microbiol. 2004, 6, 269–275. [Google Scholar] [CrossRef] [PubMed]
- Pearson, M.M.; Laurence, C.A.; Guinn, S.E.; Hansen, E.J. Biofilm formation by Moraxella catarrhalis in vitro: Roles of the UspA1 adhesin and the Hag hemagglutinin. Infect. Immun. 2006, 74, 1588–1596. [Google Scholar] [CrossRef] [PubMed]
- Jurcisek, J.A.; Bakaletz, L.O. Biofilms formed by nontypeable Haemophilus influenzae in vivo contain both double-stranded DNA and type IV pilin protein. J. Bacteriol. 2007, 189, 3868–3875. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Fritsch, M.; Hammond, L.; Landreville, R.; Slatculescu, C.; Colavita, A.; Mah, T.-F. Identification of genes involved in Pseudomonas aeruginosa biofilm-specific resistance to antibiotics. PLoS ONE 2013, 8, e61625. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Mah, T.-F. Involvement of a novel efflux system in biofilm-specific resistance to antibiotics. J. Bacteriol. 2008, 190, 4447–4452. [Google Scholar] [CrossRef] [PubMed]
- Diggle, S.P.; Winzer, K.; Chhabra, S.R.; Worrall, K.E.; Camara, M.; Williams, P. The Pseudomonas aeruginosa quinolone signal molecule overcomes the cell density-dependency of the quorum sensing hierarchy, regulates rhl-dependent genes at the onset of stationary phase and can be produced in the absence of LasR. Mol. Microbiol. 2003, 50, 29–43. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Barken, K.B.; Skindersoe, M.E.; Christensen, A.B.; Givskov, M.; Tolker-Nielsen, T. Effects of iron on DNA release and biofilm development by Pseudomonas aeruginosa. Microbiology 2007, 153, 1318–1328. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Nilsson, M.; Gjermansen, M.; Givskov, M.; Tolker-Nielsen, T. Pyoverdine and PQS mediated subpopulation interactions involved in Pseudomonas aeruginosa biofilm formation. Mol. Microbiol. 2009, 74, 1380–1392. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef] [PubMed]
- Rutherford, S.T.; Bassler, B.L. Bacterial quorum sensing: Its role in virulence and possibilities for its control. Cold Spring Harb. Perspect. Med. 2012, 2, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Bassler, B.L. How bacteria talk to each other: Regulation of gene expression by quorum sensing. Curr. Opin. Microbiol. 1999, 2, 582–587. [Google Scholar] [CrossRef]
- Parsek, M.R.; Greenberg, E.P. Sociomicrobiology: The connections between quorum sensing and biofilms. Trends Microbiol. 2005, 13, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Dow, J.M. Communication with a growing family: Diffusible signal factor (DSF) signaling in bacteria. Trends Microbiol. 2011, 19, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Hengge, R. Principles of c-di-GMP signalling in bacteria. Nat. Rev. Microbiol. 2009, 7, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Romling, U.; Amikam, D. Cyclic di-GMP as a second messenger. Curr. Opin. Microbiol. 2006, 9, 218–228. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Fouhy, Y.; Lucey, J.F.; Dow, J.M. Cyclic di-GMP signaling in bacteria: Recent advances and new puzzles. J. Bacteriol. 2006, 188, 8327–8334. [Google Scholar] [CrossRef] [PubMed]
- Simm, R.; Morr, M.; Kader, A.; Nimtz, M.; Romling, U. GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol. Microbiol. 2004, 53, 1123–1134. [Google Scholar] [CrossRef] [PubMed]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Multidrug tolerance of biofilms and persister cells. Curr. Top. Microbiol. Immunol. 2008, 322, 107–131. [Google Scholar] [PubMed]
- Spoering, A.; Lewis, K. Biofilms and planktonic cells of Pseudomonas aeruginosa have similar resistance to killing by antimicrobials. J. Bacteriol. 2001, 183, 6746–6751. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A. Microbiology: A resistance switch. Nature 2002, 416, 695–696. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, P.; Maira-Litran, T.; McBain, A.J.; Rickard, A.H.; Whyte, F.W. The physiology and collective recalcitrance of microbial biofilm communities. Adv. Microb. Physiol. 2002, 46, 202–256. [Google Scholar] [PubMed]
- Prosser, B.L.; Taylor, D.; Dix, B.A.; Cleeland, R. Method of evaluating effects of antibiotics on bacterial biofilm. Antimicrob. Agents Chemother. 1987, 31, 1502–1506. [Google Scholar] [CrossRef] [PubMed]
- Gristina, A.G.; Hobgood, C.D.; Webb, L.X.; Myrvik, Q.N. Adhesive colonization of biomaterials and antibiotic resistance. Biomaterials 1987, 8, 423–426. [Google Scholar] [CrossRef]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Riddle of biofilm resistance. Antimicrob. Agents Chemother. 2001, 45, 999–1007. [Google Scholar] [CrossRef] [PubMed]
- Mah, T.F.; O’Toole, G.A. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. 2001, 9, 34–39. [Google Scholar] [CrossRef]
- Stewart, P.S.; William Costerton, J. Antibiotic resistance of bacteria in biofilms. Lancet 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Stewart, P.S. Mechanisms of antibiotic resistance in bacterial biofilms. Int. J. Med. Microbiol. 2002, 292, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Gupta, K.; Marques, C.N.H.; Petrova, O.E.; Sauer, K. Antimicrobial tolerance of Pseudomonas aeruginosa biofilms is activated during an early developmental stage and requires the two-component hybrid SagS. J. Bacteriol. 2013, 195, 4975–4987. [Google Scholar] [CrossRef] [PubMed]
- Davies, D. Understanding biofilm resistance to antibacterial agents. Nat. Rev. Drug Discov. 2003, 2, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Yassien, M.; Khardori, N.; Ahmedy, A.; Toama, M. Modulation of biofilms of Pseudomonas aeruginosa by quinolones. Antimicrob. Agents Chemother. 1995, 39, 2262–2268. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S. Theoretical aspects of antibiotic diffusion into microbial biofilms. Antimicrob. Agents Chemother. 1996, 40, 2517–2522. [Google Scholar] [PubMed]
- Allison, D.G. The biofilm matrix. Biofouling 2003, 19, 139–150. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Sauer, K. The MerR-like transcriptional regulator BrlR contributes to Pseudomonas aeruginosa biofilm tolerance. J. Bacteriol. 2012, 194, 4823–4836. [Google Scholar] [CrossRef] [PubMed]
- Newberry, K.J.; Brennan, R.G. The structural mechanism for transcription activation by MerR family member multidrug transporter activation, N terminus. J. Biol. Chem. 2004, 279, 20356–20362. [Google Scholar] [CrossRef] [PubMed]
- Heldwein, E.E.; Brennan, R.G. Crystal structure of the transcription activator BmrR bound to DNA and a drug. Nature 2001, 409, 378–382. [Google Scholar] [CrossRef] [PubMed]
- Zheleznova, E.E.; Markham, P.N.; Neyfakh, A.A.; Brennan, R.G. Structural basis of multidrug recognition by BmrR, a transcription activator of a multidrug transporter. Cell 1999, 96, 353–362. [Google Scholar] [CrossRef]
- Drenkard, E.; Ausubel, F.M. Pseudomonas biofilm formation and antibiotic resistance are linked to phenotypic variation. Nature 2002, 416, 740–743. [Google Scholar] [CrossRef] [PubMed]
- Keren, I.; Kaldalu, N.; Spoering, A.; Wang, Y.; Lewis, K. Persister cells and tolerance to antimicrobials. FEMS Microbiol. Lett. 2004, 230, 13–18. [Google Scholar] [CrossRef]
- Marques, C.N.H.; Morozov, A.; Planzos, P.; Zelaya, H.M. The fatty acid signaling molecule cis-2-decenoic acid increases metabolic activity and reverts persister cells to an antimicrobial susceptible state. Appl. Environ. Microbiol. 2014, 80, 6976–6991. [Google Scholar] [CrossRef] [PubMed]
- Keren, I.; Shah, D.; Spoering, A. Specialized persister cells and the mechanism of multidrug tolerance in Escherichia coli. J. Bacteriol. 2004, 186, 8172–8180. [Google Scholar] [CrossRef] [PubMed]
- Sufya, N.; Allison, D.G.; Gilbert, P. Clonal variation in maximum specific growth rate and susceptibility towards antimicrobials. J. Appl. Microbiol. 2003, 95, 1261–1267. [Google Scholar] [CrossRef] [PubMed]
- Hoyle, B.D.; Jass, J.; Costerton, J.W. The biofilm glycocalyx as a resistance factor. J. Antimicrob. Chemother. 1990, 26, 1–5. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, Y.; Yang, L.; Twomey, K.B.; Sass, A.; Tolker-Nielsen, T.; Mahenthiralingam, E.; Dow, J.M.; Ryan, R.P. A sensor kinase recognizing the cell-cell signal BDSF (cis-2-dodecenoic acid) regulates virulence in Burkholderia cenocepacia. Mol. Microbiol. 2010, 77, 1220–1236. [Google Scholar] [CrossRef] [PubMed]
- Rasko, D.A.; Sperandio, V. Novel approaches to bacterial infection therapy by interfering with cell-to-cell signaling. Curr. Protoc. Microbiol. 2009, 12. [Google Scholar] [CrossRef]
- Rasko, D.A.; Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nat. Rev. Drug Discov. 2010, 9, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.K.; Schaefer, A.L.; Parsek, M.R.; Moninger, T.O.; Welsh, M.J.; Greenberg, E.P. Quorum-sensing signals indicate that cystic fibrosis lungs are infected with bacterial biofilms. Nature 2000, 407, 762–764. [Google Scholar] [CrossRef] [PubMed]
- Chugani, S.A.; Whiteley, M.; Lee, K.M.; D’Argenio, D.; Manoil, C.; Greenberg, E.P. QscR, a modulator of quorum-sensing signal synthesis and virulence in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2001, 98, 2752–2757. [Google Scholar] [CrossRef] [PubMed]
- Green, S.K.; Schroth, M.N.; Cho, J.J.; Kominos, S.K.; Vitanza-jack, V.B. Agricultural plants and soil as a reservoir for Pseudomonas aeruginosa. Appl. Microbiol. 1974, 28, 987–991. [Google Scholar] [PubMed]
- Zimakoff, J.; Høiby, N.; Rosendal, K.; Guilbert, J.P. Epidemiology of Pseudomonas aeruginosa infection and the role of contamination of the environment in a cystic fibrosis clinic. J. Hosp. Infect. 1983, 4, 31–40. [Google Scholar] [CrossRef]
- Moreau-Marquis, S.; Stanton, B.A.; O’Toole, G.A. Pseudomonas aeruginosa biofilm formation in the cystic fibrosis airway. Pulm. Pharmacol. Ther. 2008, 21, 595–599. [Google Scholar] [CrossRef] [PubMed]
- Van Delden, C.; Iglewski, B.H. Cell-to-cell signaling and Pseudomonas aeruginosa infections. Emerg. Infect. Dis. 1998, 4, 551–560. [Google Scholar] [CrossRef] [PubMed]
- Williams, P.; Cámara, M. Quorum sensing and environmental adaptation in Pseudomonas aeruginosa: A tale of regulatory networks and multifunctional signal molecules. Curr. Opin. Microbiol. 2009, 12, 182–191. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, P.N.; Koch, G.; Thompson, J.A.; Xavier, K.B.; Cool, R.H.; Quax, W.J. The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol. Mol. Biol. Rev. 2012, 76, 46–65. [Google Scholar] [CrossRef] [PubMed]
- Pearson, J.P.; Pesci, E.C.; Iglewski, B.H. Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J. Bacteriol. 1997, 179, 5756–5767. [Google Scholar] [PubMed]
- Smith, R.S.; Iglewski, B.H. Pseudomonas aeruginosa quorum sensing as a potential antimicrobial target. J. Clin. Investig. 2003, 112, 1460–1465. [Google Scholar] [CrossRef] [PubMed]
- Ishida, T.; Ikeda, T.; Takiguchi, N.; Kuroda, A.; Kato, J.; Ohtake, H. Inhibition of quorum sensing in Pseudomonas aeruginosa by N-acyl cyclopentylamides. Appl. Environ. Microbiol. 2007, 73, 3183–3188. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Song, Z.; Hentzer, M.; Andersen, J.B.; Molin, S.; Givskov, M.; Høiby, N. Synthetic furanones inhibit quorum-sensing and enhance bacterial clearance in Pseudomonas aeruginosa lung infection in mice. J. Antimicrob. Chemother. 2004, 53, 1054–1061. [Google Scholar] [CrossRef] [PubMed]
- Hentzer, M.; Riedel, K.; Rasmussen, T.B.; Heydorn, A.; Andersen, J.B.; Parsek, M.R.; Rice, S.A.; Eberl, L.; Molin, S.; Høiby, N.; et al. Inhibition of quorum sensing in Pseudomonas aeruginosa biofilm bacteria by a halogenated furanone compound. Microbiology 2002, 148, 87–102. [Google Scholar] [CrossRef] [PubMed]
- Bjarnsholt, T.; Jensen, P.Ø.; Rasmussen, T.B.; Christophersen, L.; Calum, H.; Hentzer, M.; Hougen, H.-P.; Rygaard, J.; Moser, C.; Eberl, L.; et al. Garlic blocks quorum sensing and promotes rapid clearing of pulmonary Pseudomonas aeruginosa infections. Microbiology 2005, 151, 3873–3880. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Zuo, R.; Barrios, A.F.G.; Bedzyk, L.A.; Eldridge, G.R.; Pasmore, M.E.; Wood, T.K. Differential gene expression for investigation of Escherichia coli biofilm inhibition by plant extract ursolic acid. Appl. Environ. Microbiol. 2005, 71, 4022–4034. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.; Kim, J.; Park, H.-Y.; Park, H.-J.; Lee, J.H.; Kim, C.K.; Yoon, J. Furanone derivatives as quorum-sensing antagonists of Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2008, 80, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Borlee, B.R.; Goldman, A.D.; Murakami, K.; Samudrala, R.; Wozniak, D.J.; Parsek, M.R. Pseudomonas aeruginosa uses a cyclic-di-GMP-regulated adhesin to reinforce the biofilm extracellular matrix. Mol. Microbiol. 2010, 75, 827–842. [Google Scholar] [CrossRef] [PubMed]
- Sambanthamoorthy, K.; Luo, C.; Pattabiraman, N.; Feng, X.; Koestler, B.; Waters, C.M.; Palys, T.J. Identification of small molecules inhibiting diguanylate cyclases to control bacterial biofilm development. Biofouling 2014, 30, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Lieberman, O.O.J.; Orr, M.M.W.; Wang, Y.; Lee, V.T. High-throughput screening using the differential radial capillary action of ligand assay identifies ebselen as an inhibitor of diguanylate cyclases. ACS Chem. Biol. 2014, 9, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Garo, E.; Eldridge, G.R.; Goering, M.G.; Pulcini, E.D.; Hamilton, M.A.; Costerton, J.W.; James, G.A. Asiatic acid and corosolic acid enhance the susceptibility of Pseudomonas aeruginosa biofilms to tobramycin. Antimicrob. Agents Chemother. 2007, 51, 1813–1817. [Google Scholar] [CrossRef] [PubMed]
- Balaban, N.; Cirioni, O.; Giacometti, A. Treatment of Staphylococcus aureus biofilm infection by the quorum-sensing inhibitor RIP. Antimicrob. Agents Chemother. 2007, 51, 2226–2229. [Google Scholar] [CrossRef] [PubMed]
- LoVetri, K.; Madhyastha, S. Antimicrobial and antibiofilm activity of quorum sensing peptides and Peptide analogues against oral biofilm bacteria. Methods Mol. Biol. 2010, 618, 383–392. [Google Scholar] [PubMed]
- Kjelleberg, S.; Molin, S. Is there a role for quorum sensing signals in bacterial biofilms? Curr. Opin. Microbiol. 2002, 5, 254–258. [Google Scholar] [CrossRef]
- Estrela, A.B.; Abraham, W.-R. Combining Biofilm-Controlling Compounds and Antibiotics as a Promising New Way to Control Biofilm Infections. Pharmaceuticals 2010, 3, 1374–1393. [Google Scholar] [CrossRef]
- Wang, L.H.; He, Y.; Gao, Y.; Wu, J.E.; Dong, Y.H.; He, C.; Wang, S.X.; Weng, L.X.; Xu, J.L.; Tay, L.; et al. A bacterial cell-cell communication signal with cross-kingdom structural analogues. Mol. Microbiol. 2004, 51, 903–912. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wu, J.; Eberl, L.; Zhang, L.H. Structural and functional characterization of diffusible signal factor family quorum-sensing signals produced by members of the Burkholderia cepacia complex. Appl. Environ. Microbiol. 2010, 76, 4675–4683. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.L.; Liu, Y.N.; Barber, C.E.; Dow, J.M.; Wootton, J.C.; Daniels, M.J. Genetic and molecular analysis of a cluster of rpf genes involved in positive regulation of synthesis of extracellular enzymes and polysaccharide in Xanthomonas campestris pathovar campestris. Mol. Gen. Genet. 1991, 226, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Barber, C.E.; Tang, J.L.; Feng, J.X.; Pan, M.Q.; Wilson, T.J.G.; Slater, H.; Dow, J.M.; Williams, P.; Daniels, M.J. A novel regulatory system required for pathogenicity of Xanthomonas campestris is mediated by a small diffusible signal molecule. Mol. Microbiol. 1997, 24, 555–566. [Google Scholar] [CrossRef] [PubMed]
- Beaulieu, E.D.; Ionescu, M.; Chatterjee, S.; Yokota, K.; Trauner, D.; Lindow, S. Characterization of a diffusible signaling factor from Xylella fastidiosa. MBio 2013, 4, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Dow, J.M.; Crossman, L.; Findlay, K.; He, Y.Q.; Feng, J.X.; Tang, J.L. Biofilm dispersal in Xanthomonas campestris is controlled by cell-cell signaling and is required for full virulence to plants. Proc. Natl. Acad. Sci. USA 2003, 100, 10995–11000. [Google Scholar] [CrossRef] [PubMed]
- He, Y.W.; Xu, M.; Lin, K.; Ng, Y.J.A.; Wen, C.M.; Wang, L.H.; Liu, Z.D.; Zhang, H.B.; Dong, Y.H.; Dow, J.M.; et al. Genome scale analysis of diffusible signal factor regulon in Xanthomonas campestris pv. campestris: Identification of novel cell-cell communication-dependent genes and functions. Mol. Microbiol. 2006, 59, 610–622. [Google Scholar] [PubMed]
- Minkwitz, A.; Berg, G. Comparison of antifungal activities and 16S ribosomal DNA sequences of clinical and environmental isolates of Stenotrophomonas maltophilia. J. Clin. Microbiol. 2001, 39, 139–145. [Google Scholar] [CrossRef] [PubMed]
- He, Y.-W.; Wu, J.; Cha, J.-S.; Zhang, L.-H. Rice bacterial blight pathogen Xanthomonas oryzae pv. oryzae produces multiple DSF-family signals in regulation of virulence factor production. BMC Microbiol. 2010, 10, 187. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.-P.; Lee Wong, A.C. Extracellular fatty acids facilitate flagella-independent translocation by Stenotrophomonas maltophilia. Res. Microbiol. 2007, 158, 702–711. [Google Scholar] [CrossRef] [PubMed]
- Boon, C.; Deng, Y.; Wang, L.-H.; He, Y.; Xu, J.-L.; Fan, Y.; Pan, S.Q.; Zhang, L.-H. A novel DSF-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J. 2008, 2, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; McCarthy, Y.; Watt, S.A.; Niehaus, K.; Dow, J.M. Intraspecies signaling involving the diffusible signal factor BDSF (cis-2-dodecenoic acid) influences virulence in Burkholderia cenocepacia. J. Bacteriol. 2009, 191, 5013–5019. [Google Scholar] [CrossRef] [PubMed]
- Sepehr, S.; Rahmani-Badi, A.; Babaie-Naiej, H.; Soudi, M.R. Unsaturated fatty acid, cis-2-decenoic acid, in combination with disinfectants or antibiotics removes pre-established biofilms formed by food-related bacteria. PLoS ONE 2014, 9, e101677. [Google Scholar] [CrossRef] [PubMed]
- Jennings, J.; Courtney, H.; Haggard, W. Cis-2-decenoic acid inhibits S. aureus growth and biofilm in vitro: A pilot study. Clin. Orthop. Relat. Res. 2012, 470, 2663–2670. [Google Scholar] [CrossRef] [PubMed]
- Rahmani-Badi, A.; Sepehr, S.; Mohammadi, P.; Soudi, M.R.; Babaie-Naiej, H. A combination of cis-2-decenoic acid and antibiotics eradicates pre- established catheter-associated biofilms. J. Med. Microbiol. 2014, 63, 1509–1516. [Google Scholar] [CrossRef] [PubMed]
- Vílchez, R.; Lemme, A.; Ballhausen, B.; Thiel, V.; Schulz, S.; Jansen, R.; Wagner-Döbler, I.; Sztajer, H. Streptococcus mutans inhibits Candida albicans hyphal formation by the fatty acid signaling molecule trans-2-decenoic acid (SDSF). Chembiochem 2010, 11, 1552–1162. [Google Scholar] [CrossRef] [PubMed]
- Colnaghi Simionato, A.V.; da Silva, D.S.; Lambais, M.R.; Carrilho, E. Characterization of a putative Xylella fastidiosa diffusible signal factor by HRGC-EI-MS. J. Mass Spectrom. 2007, 42, 490–496. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.; Wistrom, C.; Lindow, S.E. A cell-cell signaling sensor is required for virulence and insect transmission of Xylella fastidiosa. Proc. Natl. Acad. Sci. USA 2008, 105, 2670–2675. [Google Scholar] [CrossRef] [PubMed]
- Flavier, A.B.; Clough, S.J.; Schell, M.A.; Denny, T.P. Identification of 3-hydroxypalmitic acid methyl ester as a novel autoregulator controlling virulence in Ralstonia solanacearum. Mol. Microbiol. 1997, 26, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Oh, K.B.; Miyazawa, H.; Naito, T.; Matsuoka, H. Purification and characterization of an autoregulatory substance capable of regulating the morphological transition in Candida albicans. Proc. Natl. Acad. Sci. USA 2001, 98, 4664–4668. [Google Scholar] [CrossRef] [PubMed]
- Fouhy, Y.; Scanlon, K.; Schouest, K.; Spillane, C.; Crossman, L.; Avison, M.B.; Ryan, R.P.; Dow, J.M. Diffusible signal factor-dependent cell-cell signaling and virulence in the nosocomial pathogen Stenotrophomonas maltophilia. J. Bacteriol. 2007, 189, 4964–4968. [Google Scholar] [CrossRef] [PubMed]
- Inoue, T.; Shingaki, R.; Fukui, K. Inhibition of swarming motility of Pseudomonas aeruginosa by branched-chain fatty acids. FEMS Microbiol. Lett. 2008, 281, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Fouhy, Y.; Garcia, B.F.; Watt, S.A.; Niehaus, K.; Yang, L.; Tolker-Nielsen, T.; Dow, J.M. Interspecies signalling via the Stenotrophomonas maltophilia diffusible signal factor influences biofilm formation and polymyxin tolerance in Pseudomonas aeruginosa. Mol. Microbiol. 2008, 68, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Rahmani-Badi, A.; Sepehr, S.; Fallahi, H.; Heidari-Keshel, S. Dissection of the cis-2-decenoic acid signaling network in Pseudomonas aeruginosa using microarray technique. Front. Microbiol. 2015, 6, 383. [Google Scholar] [CrossRef] [PubMed]
- Amari, D.T.; Marques, C.N.H.; Davies, D.G. The putative enoyl-coenzyme A hydratase DspI is required for production of the Pseudomonas aeruginosa biofilm dispersion autoinducer cis-2-decenoic acid. J. Bacteriol. 2013, 195, 4600–4610. [Google Scholar] [CrossRef] [PubMed]
- Feinbaum, R.L.; Urbach, J.M.; Liberati, N.T.; Djonovic, S.; Adonizio, A.; Carvunis, A.-R.; Ausubel, F.M. Genome-wide identification of Pseudomonas aeruginosa virulence-related genes using a Caenorhabditis elegans infection model. PLoS Pathog. 2012, 8, e1002813. [Google Scholar] [CrossRef] [PubMed]
- Marques, C.N.H.; Amari, D.T.; Varer, M.; Richards, J.S.; Cavender, D.; Davies, D.G. Influence of cis-2-decenoic acid on Propionibacterium acnes biofilm development and antimicrobial susceptibility. In Proceedings of the 5th ASM Conference on Biofilms, Cancun, Mexico, 15–19 November 2009; Volume Abstract A88, p. 67.
- Marques, C.N.H.; Guttenplan, S.B.; Okkotsu, Y.; Davies, D.G. Biofilm dispersion in oral bacterial species. In Proceedings of the 109th General Meeting of the American Society for Microbiology, Philadelphia, PA, USA, 17–21 May 2009; Volume Abstrract Q-28, p. 158.
- Barraud, N.; Schleheck, D.; Klebensberger, J.; Webb, J.S.; Hassett, D.J.; Rice, S.A.; Kjelleberg, S. Nitric oxide signaling in Pseudomonas aeruginosa biofilms mediates phosphodiesterase activity, decreased cyclic di-GMP levels, and enhanced dispersal. J. Bacteriol. 2009, 191, 7333–7342. [Google Scholar] [CrossRef] [PubMed]
- Marques, C.N.H.; Dolacky, S.D.; Guttenplan, S.B.; Payabyab, E.C.; Davies, D.G. Pseudomonas aeruginosa antimicrobial challenge in combination with a biofilm dispersal agent. In Proceedings of the 4th ASM conference on Biofilms, Quebec, QC, Canada, 25–29 March 2007; Volume abstract B-30, p. 166.
- Amari, D.T.; Marques, C.N.H.; Ferreira, A.; Silverman, I.; Davies, D.G. Resuscitation of bacteria by exposure to the intercellular communication molecule, cis-2-decenoic acid. In Proceedings of the 110th General Meeting of the American Society for Microbiology, San Diego, CA, USA, 23–27 May 2010; Volume abstract D-2096, p. 162.
- Fauvart, M.; de Groote, V.N.; Michiels, J. Role of persister cells in chronic infections: Clinical relevance and perspectives on anti-persister therapies. J. Med. Microbiol. 2011, 60, 699–709. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, L.R.; Burns, J.L.; Lory, S.; Lewis, K. Emergence of Pseudomonas aeruginosa strains producing high levels of persister cells in patients with cystic fibrosis. J. Bacteriol. 2010, 192, 6191–6199. [Google Scholar] [CrossRef] [PubMed]
- Balaban, N.Q.; Merrin, J.; Chait, R.; Kowalik, L.; Leibler, S. Bacterial persistence as a phenotypic switch. Science 2004, 305, 1622–1625. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Persister cells and the riddle of biofilm survival. Biochemistry 2005, 70, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Kwan, B.W.; Valenta, J.A.; Benedik, M.J.; Wood, T.K. Arrested protein synthesis increases persister-like cell formation. Antimicrob. Agents Chemother. 2013, 57, 1468–1473. [Google Scholar] [CrossRef] [PubMed]
- Korch, S.B.; Henderson, T.A.; Hill, T.M. Characterization of the hipA7 allele of Escherichia coli and evidence that high persistence is governed by (p)ppGpp synthesis. Mol. Microbiol. 2003, 50, 1199–1213. [Google Scholar] [CrossRef] [PubMed]
- Scherrer, R.; Moyed, H.S. Conditional impairment of cell division and altered lethality in hipA mutants of Escherichia coli K-12. J. Bacteriol. 1988, 170, 3321–3326. [Google Scholar] [PubMed]
- Pascoe, B.; Dams, L.; Wilkinson, T.S.; Harris, L.G.; Bodger, O.; Mack, D.; Davies, A.P. Dormant cells of Staphylococcus aureus are resuscitated by spent culture supernatant. PLoS ONE 2014, 9, e85998. [Google Scholar] [CrossRef] [PubMed]
- Allison, K.R.; Brynildsen, M.P.; Collins, J.J. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature 2011, 473, 216–220. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Bahar, A.A.; Syed, H.; Ren, D. Reverting antibiotic tolerance of Pseudomonas aeruginosa PAO1 persister cells by (Z)-4-bromo-5-(bromomethylene)-3-methylfuran-2(5H)-one. PLoS ONE 2012, 7, e45778. [Google Scholar] [CrossRef] [PubMed]
- Mina, E.; Marques, C.N.H. Effect of cis-2-decenoic acid on virulence of Staphylococcus aureus persister cells. In Proceedings of the 113th General Meeting of the American Society for Microbiology, Denver, CO, USA, 18–21 May 2013; Volume abstract 512, p. 121.
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marques, C.N.H.; Davies, D.G.; Sauer, K. Control of Biofilms with the Fatty Acid Signaling Molecule cis-2-Decenoic Acid. Pharmaceuticals 2015, 8, 816-835. https://doi.org/10.3390/ph8040816
Marques CNH, Davies DG, Sauer K. Control of Biofilms with the Fatty Acid Signaling Molecule cis-2-Decenoic Acid. Pharmaceuticals. 2015; 8(4):816-835. https://doi.org/10.3390/ph8040816
Chicago/Turabian StyleMarques, Cláudia N. H., David G. Davies, and Karin Sauer. 2015. "Control of Biofilms with the Fatty Acid Signaling Molecule cis-2-Decenoic Acid" Pharmaceuticals 8, no. 4: 816-835. https://doi.org/10.3390/ph8040816
APA StyleMarques, C. N. H., Davies, D. G., & Sauer, K. (2015). Control of Biofilms with the Fatty Acid Signaling Molecule cis-2-Decenoic Acid. Pharmaceuticals, 8(4), 816-835. https://doi.org/10.3390/ph8040816






