Modification of Bacteriophages to Increase Their Association with Lung Epithelium Cells In Vitro
Abstract
1. Introduction
2. Results
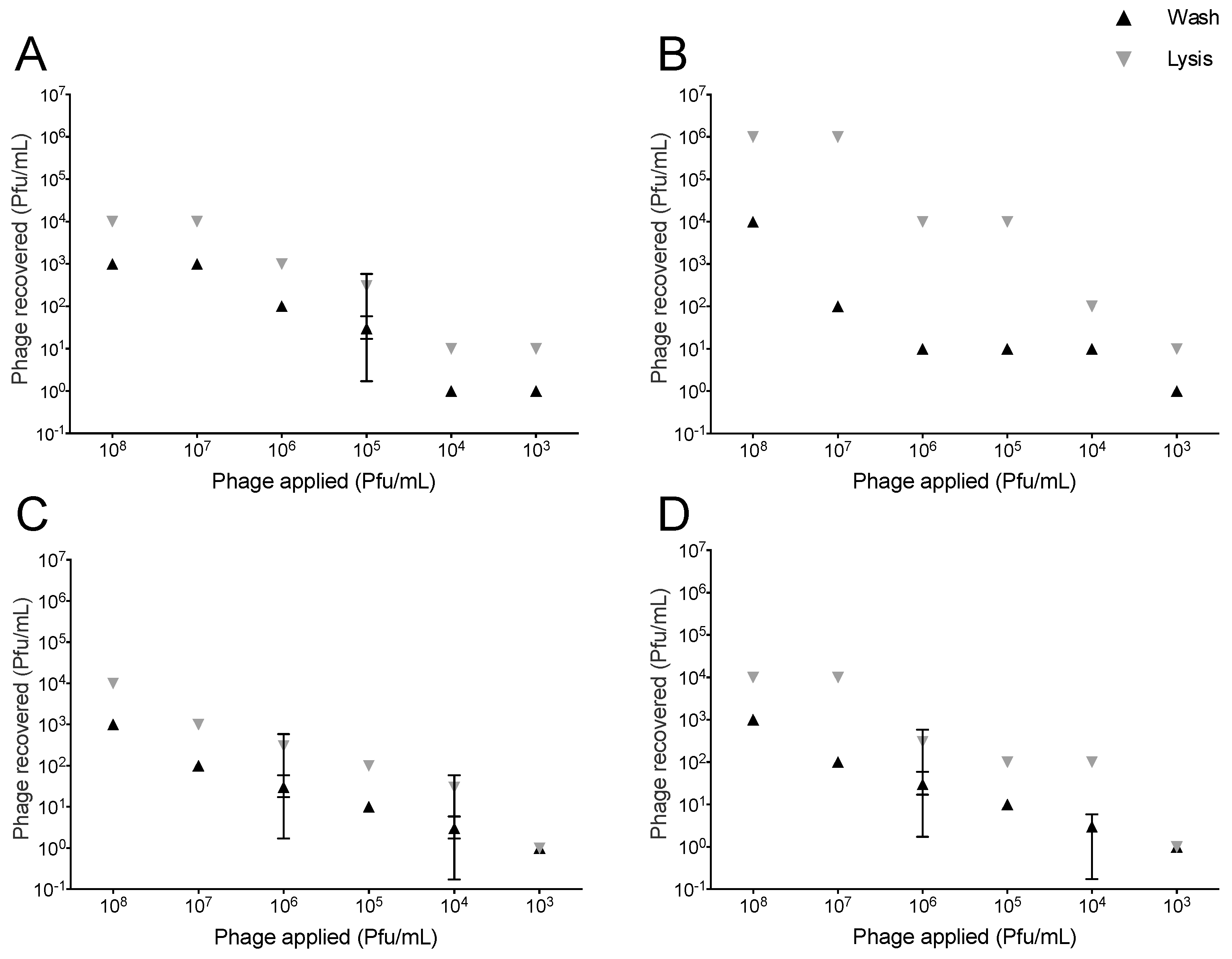
2.1. Association of T4-Like and T7 with Lung Epithelial Cells
2.2. Modification of Phage Capsids with “Homing” Peptides
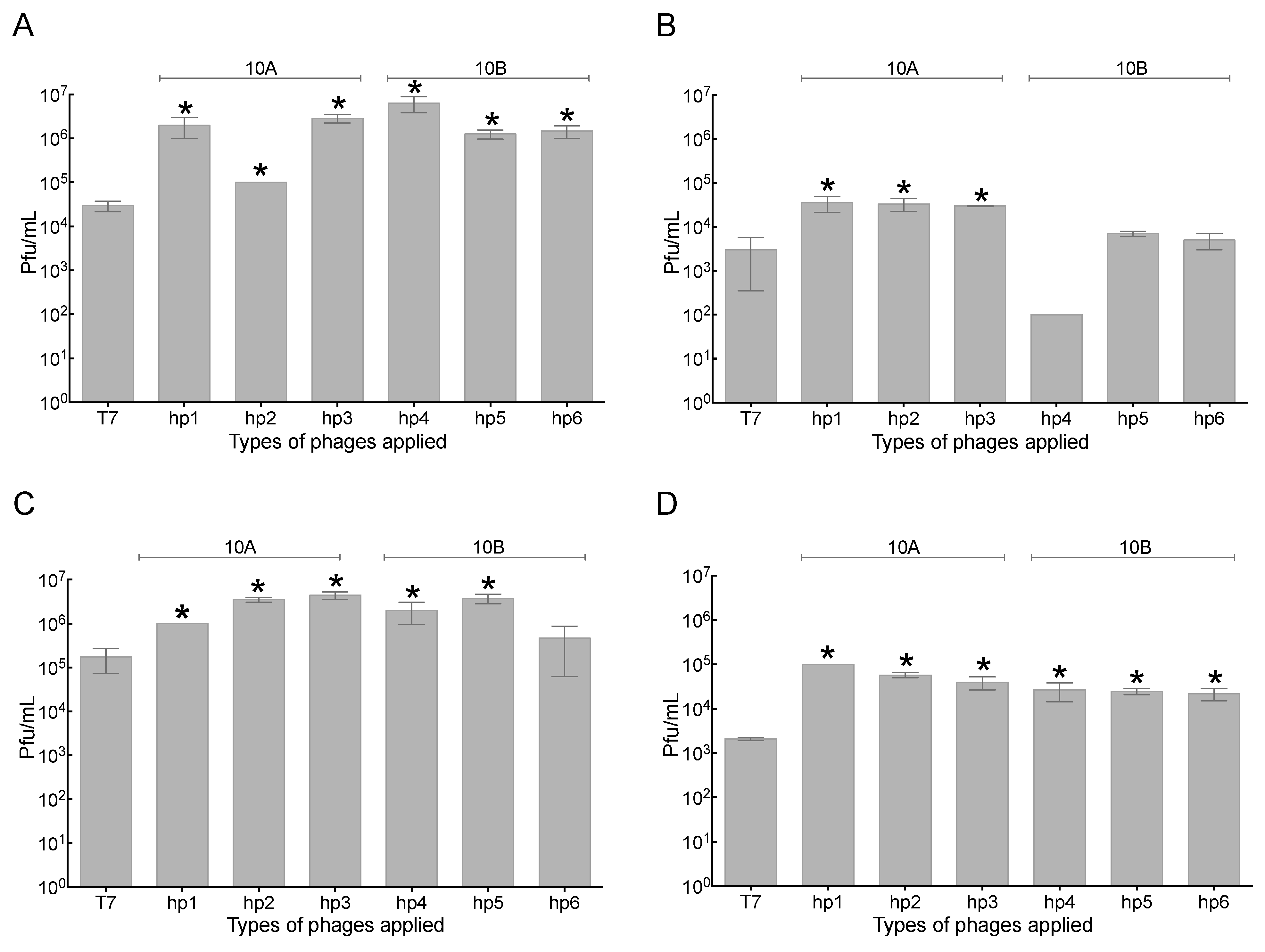
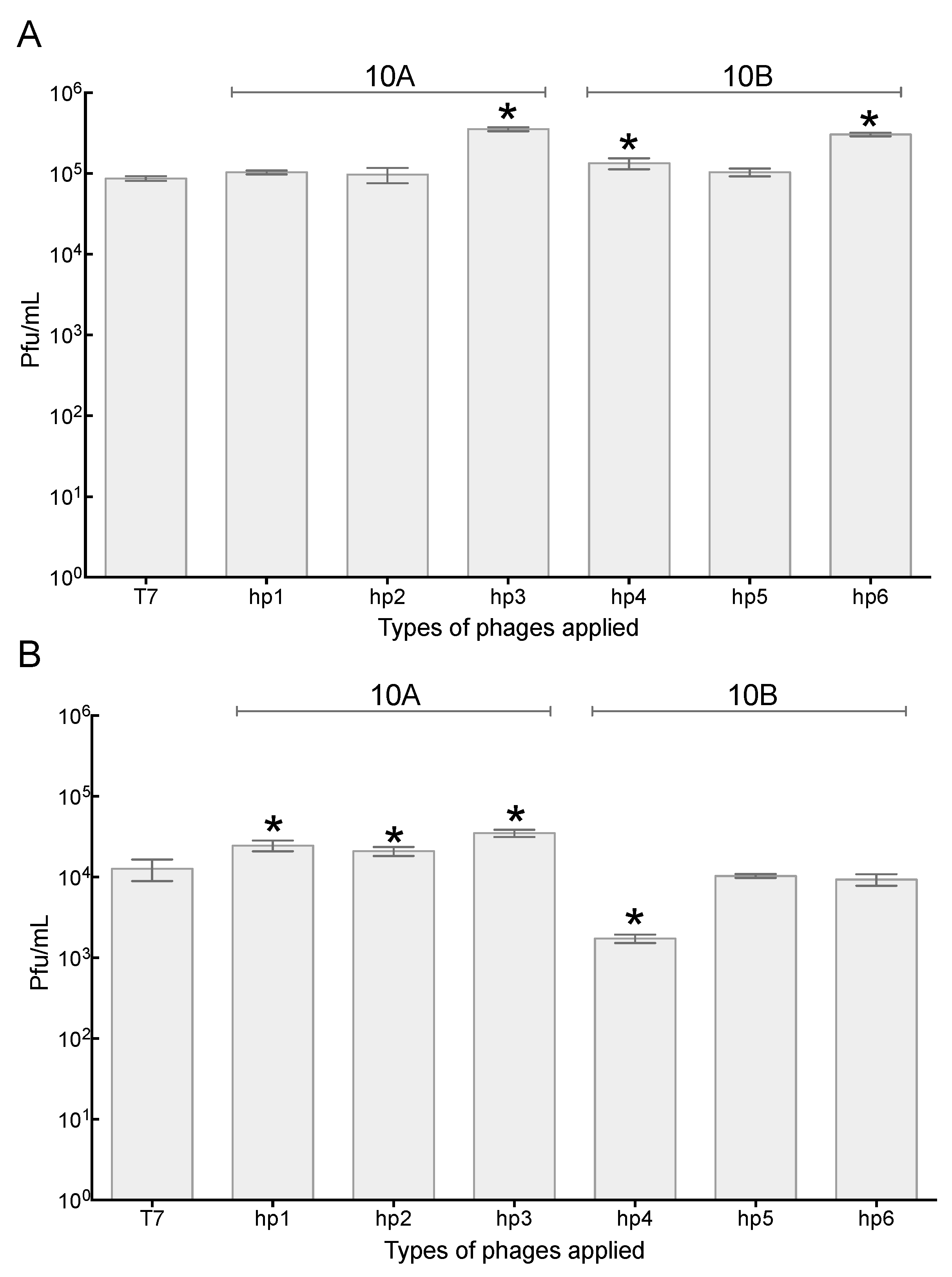
2.3. Increased Association of Phages with A549 Human Lung Epithelial Cells
2.4. Association of Phage with 3T3 Mouse Embryonic Fibroblast Cells
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Phage Stocks, Tissue Culture Cell Lines, and Growth Conditions
4.2. Design of Homologous Recombination Vectors for Homing Peptide Insertion
4.3. Selection of T7 Mutants
4.4. Phage Association Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- The United States Centers for Disease Control and Prevention. GET SMART: Know When Antibiotics Work. AJN 2008, 6. [Google Scholar] [CrossRef]
- Shallcross, L.J.; Davies, S.C. The World Health Assembly resolution on antimicrobial resistance. J. Antimicrob. Chemother. 2014, 69, 2883–2885. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Antimicrobial Use in Aquaculture and Antimicrobial Resistance. Aeromonas Resist. How It Eff. Hum. 2006, 1–107. [Google Scholar]
- Levy, S.B.; Bergman, M.M. The Antibiotic Paradox: How the Misuse of Antibiotics Destroys Their Curative Powers, 2nd Edition. Clin. Infect. Dis. 2003, 36, 238. [Google Scholar] [CrossRef]
- Nathan, C. Antibiotics at the crossroads. Nature 2004, 431, 899–902. [Google Scholar] [CrossRef]
- Yewale, V.N. Antimicrobial resistance—A ticking bomb! Indian Pediatr. 2014, 51, 171–172. [Google Scholar] [CrossRef]
- Schlesinger, L.A.; Kotter, J.P. Choosing Strategies for Change—HBR.org. Harv. Bus. Rev. 2008, 1–8. [Google Scholar]
- Moellering, R.C. NDM-1—A Cause for Worldwide Concern. N. Engl. J. Med. 2010, 363, 2377–2379. [Google Scholar] [CrossRef]
- Alanis, A.J. Resistance to antibiotics: Are we in the post-antibiotic era? Arch. Med. Res. 2005, 36, 697–705. [Google Scholar] [CrossRef]
- Czaplewski, L.; Bax, R.; Clokie, M.; Dawson, M.; Fairhead, H.; Fischetti, V.A.; Foster, S.; Gilmore, B.F.; Hancock, R.E.W.; Harper, D.; et al. Alternatives to antibiotics-a pipeline portfolio review. Lancet Infect. Dis. 2016, 16, 239–251. [Google Scholar] [CrossRef]
- NIAID’s Antibacterial Resistance Program: Current Status and Future Directions. NIAID 2014, 1, 13.
- Abedon, S.T.; García, P.; Mullany, P.; Aminov, R. Editorial: Phage therapy: Past, present and future. Front. Microbiol. 2017, 8, 981. [Google Scholar] [CrossRef] [PubMed]
- Roach, D.R.; Leung, C.Y.; Henry, M.; Morello, E.; Singh, D.; Di Santo, J.P.; Weitz, J.S.; Debarbieux, L. Synergy between the Host Immune System and Bacteriophage Is Essential for Successful Phage Therapy against an Acute Respiratory Pathogen. Cell Host Microbe 2017, 22, 38–47.e4. [Google Scholar] [CrossRef] [PubMed]
- Gelman, D.; Eisenkraft, A.; Chanishvili, N.; Nachman, D.; Coppenhagem Glazer, S.; Hazan, R. The history and promising future of phage therapy in the military service. J. Trauma Acute Care Surg. 2018, 85, S18–S26. [Google Scholar] [CrossRef]
- Van der Merwe, R.G.; Warren, R.M.; Sampson, S.L.; Gey van Pittius, N.C. Phage-based detection of bacterial pathogens. Analyst 2014, 139, 2617–2626. [Google Scholar] [CrossRef]
- Moye, Z.D.; Woolston, J.; Sulakvelidze, A. Bacteriophage applications for food production and processing. Viruses 2018, 10, 205. [Google Scholar] [CrossRef]
- Coffey, B.; Mills, S.; Coffey, A.; McAuliffe, O.; Paul Ross, R. Phage and their lysins as biocontrol agents for food safety applications. Annu. Rev. Food Sci. Technol. 2010, 1, 449–468. [Google Scholar] [CrossRef]
- Lone, A.; Anany, H.; Hakeem, M.; Aguis, L.; Avdjian, A.C.; Bouget, M.; Atashi, A.; Brovko, L.; Rochefort, D.; Griffiths, M.W. Development of prototypes of bioactive packaging materials based on immobilized bacteriophages for control of growth of bacterial pathogens in foods. Int. J. Food Microbiol. 2016, 217, 49–58. [Google Scholar] [CrossRef]
- Wang, C.; Sauvageau, D.; Elias, A. Immobilization of Active Bacteriophages on Polyhydroxyalkanoate Surfaces. ACS Appl. Mater. Interfaces 2016, 8, 1128–1138. [Google Scholar] [CrossRef]
- Zhang, S.; Chen, D.C.; Chen, L.M. Facing a new challenge: The adverse effects of antibiotics on gut microbiota and host immunity. Chin. Med. J. 2019, 132, 1135–1138. [Google Scholar] [CrossRef]
- Willing, B.P.; Russell, S.L.; Finlay, B.B. Shifting the balance: Antibiotic effects on host-microbiota mutualism. Nat. Rev. Microbiol. 2011, 9, 233–243. [Google Scholar] [CrossRef]
- Becattini, S.; Taur, Y.; Pamer, E.G. Antibiotic-Induced Changes in the Intestinal Microbiota and Disease. Trends Mol. Med. 2016, 22, 458–478. [Google Scholar] [CrossRef]
- Robak, O.H.; Heimesaat, M.M.; Kruglov, A.A.; Prepens, S.; Ninnemann, J.; Gutbier, B.; Reppe, K.; Hochrein, H.; Suter, M.; Kirschning, C.J.; et al. Antibiotic treatment-induced secondary IgA deficiency enhances susceptibility to Pseudomonas aeruginosa pneumonia. J. Clin. Investig. 2018, 128, 3535–3545. [Google Scholar] [CrossRef]
- Chibani-Chennoufi, S.; Sidoti, J.; Bruttin, A.; Kutter, E.; Sarker, S.; Brüssow, H. In vitro and in vivo bacteriolytic activities of Escherichia coli phages: Implications for phage therapy. Antimicrob. Agents Chemother. 2004, 48, 2558–2569. [Google Scholar] [CrossRef]
- Sarker, S.A.; McCallin, S.; Barretto, C.; Berger, B.; Pittet, A.C.; Sultana, S.; Krause, L.; Huq, S.; Bibiloni, R.; Bruttin, A.; et al. Oral T4-like phage cocktail application to healthy adult volunteers from Bangladesh. Virology 2012, 434, 222–232. [Google Scholar] [CrossRef] [PubMed]
- Granowitz, E.V.; Brown, R.B. Antibiotic Adverse Reactions and Drug Interactions. Crit. Care Clin. 2008, 24, 421–442. [Google Scholar] [CrossRef] [PubMed]
- Morales-Alvarez, M.C. Nephrotoxicity of Antimicrobials and Antibiotics. Adv. Chronic Kidney Dis. 2020, 27, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Grill, M.F.; Maganti, R.K. Neurotoxic effects associated with antibiotic use: Management considerations. Br. J. Clin. Pharmacol. 2011, 72, 381–393. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.K.; Yuan, J.; Li, M.; Sutton, S.S.; Rao, G.A.; Jacob, S.; Bennett, C.L. Cardiac risks associated with antibiotics: Azithromycin and levofloxacin. Expert Opin. Drug Saf. 2015, 14, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Clokie, M.R.J.; Kropinski, A.M. Bacteriophages: Methods and protocols Volume 1: Isolation, Characterization, and Interactions; Humana Press: Totowa, NJ, USA, 2009. [Google Scholar]
- Carlton, R.M. Phage therapy: Past history and future prospects. Arch. Immunol. Ther. Exp. 1999, 47, 267–274. [Google Scholar]
- Abedon, S.T. Bacteriophage secondary infection. Virol. Sin. 2015, 30, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Capparelli, R.; Nocerino, N.; Iannaccone, M.; Ercolini, D.; Parlato, M.; Chiara, M.; Iannelli, D. Bacteriophage therapy of Salmonella enterica: A fresh appraisal of bacteriophage therapy. J. Infect. Dis. 2010, 201, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Skurnik, M.; Strauch, E. Phage therapy: Facts and fiction. Int. J. Med. Microbiol. 2006, 296, 5–14. [Google Scholar] [CrossRef]
- Abedon, S.T.; Thomas-Abedon, C. Phage therapy pharmacology. Curr. Pharm. Biotechnol. 2010, 11, 28–47. [Google Scholar] [CrossRef] [PubMed]
- Górski, A.; Dąbrowska, K.; Hodyra-Stefaniak, K.; Borysowski, J.; Międzybrodzki, R.; Weber-Dąbrowska, B. Phages targeting infected tissues: Novel approach to phage therapy. Future Microbiol. 2015, 10, 199–204. [Google Scholar] [CrossRef]
- Keller, R.; Engley, F.B. Fate of Bacteriophage Particles Introduced into Mice by Various Routes. Proc. Soc. Exp. Biol. Med. 1958, 98, 577–580. [Google Scholar] [CrossRef] [PubMed]
- Bogovazova, G.G.; Voroshilova, N.N.; Bondarenko, V.M. The efficacy of Klebsiella pneumoniae bacteriophage in the therapy of experimental Klebsiella infection. Zh. Mikrobiol. Epidemiol. Immunobiol. 1991, 4, 5–8. [Google Scholar]
- Abedon, S.T. Bacteriophages as Drugs: The Pharmacology of Phage Therapy. In Phage Therapy: Current Research and Applications; Międzybrodzki, J.B.R., Górski, A., Eds.; Caister Academic Press: Norfolk, UK, 2014; pp. 69–100. [Google Scholar]
- Serwer, P.; Wright, E.T.; Lee, J.C. High murine blood persistence of phage T3 and suggested strategy for phage therapy. BMC Res. Notes 2019, 12, 1–5. [Google Scholar] [CrossRef]
- Hodyra-Stefaniak, K.; Lahutta, K.; Majewska, J.; Kaźmierczak, Z.; Lecion, D.; Harhala, M.; Kęska, W.; Owczarek, B.; Jończyk-Matysiak, E.; Kłopot, A.; et al. Bacteriophages engineered to display foreign peptides may become short-circulating phages. Microb. Biotechnol. 2019, 12, 730–741. [Google Scholar] [CrossRef]
- Gregory, A.C.; Zablocki, O.; Howell, A.; Bolduc, B.; Sullivan, M.B. The human gut virome database. bioRxiv 2019, 655910. [Google Scholar]
- Hoyles, L.; McCartney, A.L.; Neve, H.; Gibson, G.R.; Sanderson, J.D.; Heller, K.J.; van Sinderen, D. Characterization of virus-like particles associated with the human faecal and caecal microbiota. Res. Microbiol. 2014, 165, 803–812. [Google Scholar] [CrossRef]
- Shkoporov, A.N.; Ryan, F.J.; Draper, L.A.; Forde, A.; Stockdale, S.R.; Daly, K.M.; McDonnell, S.A.; Nolan, J.A.; Sutton, T.D.S.; Dalmasso, M.; et al. Reproducible protocols for metagenomic analysis of human faecal phageomes. Microbiome 2018, 6, 68. [Google Scholar] [CrossRef]
- Dabrowska, K.; Switała-Jelen, K.; Opolski, A.; Weber-Dabrowska, B.; Gorski, A. A review: Bacteriophage penetration in vertebrates. J. Appl. Microbiol. 2005, 98, 7–13. [Google Scholar] [CrossRef]
- Górski, A.; Wazna, E.; Dąbrowska, B.W.; Dąbrowska, K.; Świtała-Jeleń, K.; Miȩdzybrodzki, R. Bacteriophage translocation. FEMS Immunol. Med. Microbiol. 2006, 46, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Weber-Dabrowska, B.; Dabrowski, M.; Slopek, S. Studies on bacteriophage penetration in patients subjected to phage therapy. Arch. Immunol. Ther. Exp. 1987, 35, 563–568. [Google Scholar]
- Sutton, T.D.S.; Hill, C. Gut Bacteriophage: Current Understanding and Challenges. Front. Endocrinol. 2019, 10, 784. [Google Scholar] [CrossRef]
- Navarro, F.; Muniesa, M. Phages in the human body. Front. Microbiol. 2017, 8, 566. [Google Scholar] [CrossRef]
- König, J.; Wells, J.; Cani, P.D.; García-Ródenas, C.L.; MacDonald, T.; Mercenier, A.; Whyte, J.; Troost, F.; Brummer, R.J. Human intestinal barrier function in health and disease. Clin. Transl. Gastroenterol. 2016, 7, e196. [Google Scholar] [CrossRef] [PubMed]
- Huh, H.; Wong, S.; St. Jean, J.; Slavcev, R. Bacteriophage interactions with mammalian tissue: Therapeutic applications. Adv. Drug Deliv. Rev. 2019, 145, 4–17. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, S.; Baker, K.; Padman, B.S.; Patwa, R.; Dunstan, R.A.; Weston, T.A.; Schlosser, K.; Bailey, B.; Lithgow, T.; Lazarou, M.; et al. Bacteriophage transcytosis provides a mechanism to cross epithelial cell layers. MBio 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Aird, W.C. Endothelial cell heterogeneity. Cold Spring Harb. Perspect. Med. 2012, 2, a006429. [Google Scholar] [CrossRef]
- Garlanda, C.; Dejana, E. Heterogeneity of endothelial cells: Specific markers. Arterioscler. Thromb. Vasc. Biol. 1997, 17, 1193–1202. [Google Scholar] [CrossRef] [PubMed]
- Aird, W.C. Phenotypic heterogeneity of the endothelium: II. Representative vascular beds. Circ. Res. 2007, 100, 174–190. [Google Scholar] [CrossRef] [PubMed]
- Rajotte, D.; Arap, W.; Hagedorn, M.; Koivunen, E.; Pasqualini, R.; Ruoslahti, E. Molecular Heterogeneity of the Vascular Endothelium Revealed by In Vivo Phage Display. J. Clin. Investig 1998, 102, 430–437. [Google Scholar] [CrossRef] [PubMed]
- Costantini, T.W.; Putnam, J.G.; Sawada, R.; Baird, A.; Loomis, W.H.; Eliceiri, B.P.; Bansal, V.; Coimbra, R. Targeting the gut barrier: Identification of a homing peptide sequence for delivery into the injured intestinal epithelial cell. Surgery 2009, 146, 206–212. [Google Scholar] [CrossRef]
- Curnis, F.; Arrigoni, G.; Sacchi, A.; Fischetti, L.; Corti, A.; Arap, W.; Pasqualini, R. Differential binding of drugs containing the NGR motif to CD13 isoforms in tumor vessels, epithelia, and myeloid cells. Cancer Res. 2002, 62, 867–874. [Google Scholar]
- Morris, C.J.; Smith, M.W.; Griffiths, P.C.; McKeown, N.B.; Gumbleton, M. Enhanced pulmonary absorption of a macromolecule through coupling to a sequence-specific phage display-derived peptide. J. Control. Release 2011, 151, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Kim, Y.J.; Jon, S. A high-affinity peptide for nicotinic acetylcholine receptor-α1 and its potential use in pulmonary drug delivery. J. Control. Release 2014, 192, 141–147. [Google Scholar] [CrossRef]
- Wu, M.; Pasula, R.; Smith, P.A.; Martin, W.J. Mapping alveolar binding sites in vivo using phage peptide libraries. Gene Ther. 2003, 10, 1429–1436. [Google Scholar] [CrossRef][Green Version]
- Laakkonen, P.; Vuorinen, K. Homing peptides as targeted delivery vehicles. Integr. Biol. 2010, 2, 326–337. [Google Scholar] [CrossRef]
- Pasqualini, R.; Ruoslahti, E. Organ targeting in vivo using phage display peptide libraries. Nature 1996, 380, 364–366. [Google Scholar] [CrossRef] [PubMed]
- Arap, W.; Haedicke, W.; Bernasconi, M.; Kain, R.; Rajotte, D.; Krajewski, S.; Ellerby, H.M.; Bredesen, D.E.; Pasqualini, R.; Ruoslahti, E. Targeting the prostate for destruction through a vascular address. Proc. Natl. Acad. Sci. USA 2002, 99, 1527–1531. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Hoffman, J.A.; Ruoslahti, E. Molecular profiling of heart endothelial cells. Circulation 2005, 112, 1601–1611. [Google Scholar] [CrossRef] [PubMed]
- Pasqualini, R.; Koivunen, E.; Ruoslahti, E. αv Integrins as Receptors for Tumor Targeting by Circulating Ligands. Nat. Biotechnol. 1997, 15, 542–546. [Google Scholar] [CrossRef] [PubMed]
- Porkka, K.; Laakkonen, P.; Hoffman, J.A.; Bernasconi, M.; Ruoslahti, E. A fragment of the HMGN2 protein homes to the nuclei of tumor cells and tumor endothelial cells in vivo. Proc. Natl. Acad. Sci. USA 2002, 99, 7444–7449. [Google Scholar] [CrossRef] [PubMed]
- Laakkonen, P.; Akerman, M.E.; Biliran, H.; Yang, M.; Ferrer, F.; Karpanen, T.; Hoffman, R.M.; Ruoslahti, E. Antitumor activity of a homing peptide that targets tumor lymphatics and tumor cells. Proc. Natl. Acad. Sci. USA 2004, 101, 9381–9386. [Google Scholar] [CrossRef]
- Zhang, L.; Giraudo, E.; Hoffman, J.A.; Hanahan, D.; Ruoslahti, E. Lymphatic zip codes in premalignant lesions and tumors. Cancer Res. 2006, 66, 5696–5706. [Google Scholar] [CrossRef]
- Rajotte, D.; Ruoslahti, E. Membrane dipeptidase is the receptor for a lung-targeting peptide identified by in vivo phage display. J. Biol. Chem. 1999, 274, 11593–11598. [Google Scholar] [CrossRef]
- Samoylova, T.I.; Smith, B.F. Elucidation of muscle-binding peptides by phage display screening. Muscle Nerve 1999, 22, 460–466. [Google Scholar] [CrossRef]
- Kolonin, M.G.; Saha, P.K.; Chan, L.; Pasqualini, R.; Arap, W. Reversal of obesity by targeted ablation of adipose tissue. Nat. Med. 2004, 10, 625–632. [Google Scholar] [CrossRef]
- Ruoslahti, E. Peptides as targeting elements and tissue penetration devices for nanoparticles. Adv. Mater. 2012, 24, 3747–3756. [Google Scholar] [CrossRef]
- Ellerby, H.M.; Arap, W.; Ellerby, L.M.; Kain, R.; Andrusiak, R.; Del Rio, G.; Krajewski, S.; Lombardo, C.R.; Rao, R.; Ruoslahti, E.; et al. Anti-cancer activity of targeted pro-apoptotic peptides. Nat. Med. 1999, 5, 1032–1038. [Google Scholar] [CrossRef]
- Corti, A.; Curnis, F.; Rossoni, G.; Marcucci, F.; Gregorc, V. Peptide-mediated targeting of cytokines to tumor vasculature: The NGR-hTNF example. BioDrugs 2013, 27, 591–603. [Google Scholar] [CrossRef]
- Grigonyte, A.M.; Harrison, C.; MacDonald, P.R.; Montero-Blay, A.; Tridgett, M.; Duncan, J.; Sagona, A.P.; Constantinidou, C.; Jaramillo, A.; Millard, A. Comparison of CRISPR and marker-based methods for the engineering of phage T7. Viruses 2020, 12, 193. [Google Scholar] [CrossRef]
- Liu, Y.; Huang, H.; Wang, H.; Zhang, Y. A novel approach for T7 bacteriophage genome integration of exogenous DNA. J. Biol. Eng. 2020, 14, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Jensen, J.D.; Parks, A.R.; Adhya, S.; Rattray, A.J.; Court, D.L. λ Recombineering Used to Engineer the Genome of Phage T7. Antibiotics 2020, 9, 805. [Google Scholar] [CrossRef] [PubMed]
- Chan, L.Y.; Kosuri, S.; Endy, D. Refactoring bacteriophage T7. Mol. Syst. Biol. 2005, 1, 2005.0018. [Google Scholar] [CrossRef] [PubMed]
- Studier, F.W. The genetics and physiology of bacteriophage T7. Virology 1969, 39, 562–574. [Google Scholar] [CrossRef]
- Brown, D.M.; Ruoslahti, E. Metadherin, a cell surface protein in breast tumors that mediates lung metastasis. Cancer Cell 2004, 5, 365–374. [Google Scholar] [CrossRef]
- Habib, G.M.; Barriost, R.; Shi, Z.Z.; Lieberman, M.W. Four distinct membrane-bound dipeptidase RNAs are differentially expressed and show discordant regulation with γ-glutamyl transpeptidase. J. Biol. Chem. 1996, 271, 16273–16280. [Google Scholar] [CrossRef]
- Inamura, T.; Pardridge, W.M.; Kumagai, Y.; Black, K.L. Differential tissue expression of immunoreactive dehydropeptidase I, a peptidyl leukotriene metabolizing enzyme. Prostaglandins Leukot. Essent. Fat. Acids 1994, 50, 85–92. [Google Scholar] [CrossRef]
- Payne, R.J.H.; Jansen, V.A.A. Understanding bacteriophage therapy as a density-dependent kinetic process. J. Theor. Biol. 2001, 208, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Kolonin, M.G.; Sun, J.; Do, K.; Vidal, C.I.; Ji, Y.; Baggerly, K.A.; Pasqualini, R.; Amp, W.; Kolonin, M.G.; Sun, J.; et al. Synchronous selection of homing peptides for multiple tissues by in vivo phage display. FASEB J. 2006, 20, 979–981. [Google Scholar] [CrossRef] [PubMed]
- Giordano, R.J.; Lahdenranta, J.; Zhen, L.; Chukwueke, U.; Petrache, I.; Langley, R.R.; Fidler, I.J.; Pasqualini, R.; Tuder, R.M.; Arap, W. Targeted induction of lung endothelial cell apoptosis causes emphysema-like changes in the mouse. J. Biol. Chem. 2008, 283, 29447–29460. [Google Scholar] [CrossRef]
- Deng, X.; Wang, L.; You, X.; Dai, P.; Zeng, Y. Advances in the T7 phage display system (Review). Mol. Med. Rep. 2018, 17, 714–720. [Google Scholar] [CrossRef]
- Slootweg, E.J.; Keller, H.J.H.G.; Hink, M.A.; Borst, J.W.; Bakker, J.; Schots, A. Fluorescent T7 display phages obtained by translational frameshift. Nucleic Acids Res. 2006, 34, e137. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.; Harjai, K.; Chhibber, S. Bacteriophage-aided intracellular killing of engulfed methicillin-resistant Staphylococcus aureus (MRSA) by murine macrophages. Appl. Microbiol. Biotechnol. 2014, 98, 4653–4661. [Google Scholar] [CrossRef]
- Zhang, L.; Sun, L.; Wei, R.; Gao, Q.; He, T.; Xu, C.; Liu, X.; Wang, R. Intracellular Staphylococcus aureus control by virulent bacteriophages within MAC-T bovine mammary epithelial cells. Antimicrob. Agents Chemother. 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Lehti, T.A.; Pajunen, M.I.; Skog, M.S.; Finne, J. Internalization of a polysialic acid-binding Escherichia coli bacteriophage into eukaryotic neuroblastoma cells. Nat. Commun. 2017, 8, 1–12. [Google Scholar] [CrossRef]
- Dabrowska, K.; Opolski, A.; Wietrzyk, J.; Switala-Jelen, K.; Boratynski, J.; Nasulewicz, A.; Lipinska, L.; Chybicka, A.; Kujawa, M.; Zabel, M.; et al. Antitumor activity of bacteriophages in murine experimental cancer models caused possibly by inhibition of β3 integrin signaling pathway. Acta Virol. 2004, 48, 241–248. [Google Scholar]
- Akerman, M.E.; Chan, W.C.W.; Laakkonen, P.; Bhatia, S.N.; Ruoslahti, E. Nanocrystal targeting in vivo. Proc. Natl. Acad. Sci. USA 2002, 99, 12617–12621. [Google Scholar] [CrossRef]
- Karmali, P.P.; Kotamraju, V.R.; Kastantin, M.; Black, M.; Missirlis, D.; Tirrell, M.; Ruoslahti, E. Targeting of albumin-embedded paclitaxel nanoparticles to tumors. Nanomed. Nanotechnol. Biol. Med. 2009, 5, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Uchida, M.; Kosuge, H.; Terashima, M.; Willits, D.A.; Liepold, L.O.; Young, M.J.; McConnell, M.V.; Douglas, T. Protein cage nanoparticles bearing the LyP-1 peptide for enhanced imaging of macrophage-rich vascular lesions. ACS Nano 2011, 5, 2493–2502. [Google Scholar] [CrossRef] [PubMed]
- Soda, Y.; Marumoto, T.; Friedmann-Morvinski, D.; Soda, M.; Liu, F.; Michiue, H.; Pastorino, S.; Yang, M.; Hoffman, R.M.; Kesari, S.; et al. Transdifferentiation of glioblastoma cells into vascular endothelial cells. Proc. Natl. Acad. Sci. USA 2011, 108, 4274–4280. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.Y.; Yang, W.H.; Dong, X.K.; Cong, L.M.; Li, N.; Li, Y.; Wen, Z.B.; Yin, Z.; Lan, Z.J.; Li, W.P.; et al. Inhalation Study of Mycobacteriophage D29 Aerosol for Mice by Endotracheal Route and Nose-Only Exposure. J. Aerosol Med. Pulm. Drug Deliv. 2016, 29, 393–405. [Google Scholar] [CrossRef] [PubMed]
- Przystal, J.M.; Waramit, S.; Pranjol, M.Z.I.; Yan, W.; Chu, G.; Chongchai, A.; Samarth, G.; Olaciregui, N.G.; Tabatabai, G.; Carcaboso, A.M.; et al. Efficacy of systemic temozolomide-activated phage-targeted gene therapy in human glioblastoma. EMBO Mol. Med. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Kropinski, A.M.; Mazzocco, A.; Waddell, T.E.; Lingohr, E.; Johnson, R.P. Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol. Biol. 2009, 501, 69–76. [Google Scholar]
| T7 Mutant | Structural Protein Subunit | Peptide |
|---|---|---|
| T7-hp1 | A | GFE-1 |
| T7-hp2 | A | GFE-2 |
| T7-hp3 | A | MTDH |
| T7-hp4 | B | GFE-1 |
| T7-hp5 | B | GFE-2 |
| T7-hp6 | B | MTDH |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grigonyte, A.M.; Hapeshi, A.; Constantinidou, C.; Millard, A. Modification of Bacteriophages to Increase Their Association with Lung Epithelium Cells In Vitro. Pharmaceuticals 2021, 14, 308. https://doi.org/10.3390/ph14040308
Grigonyte AM, Hapeshi A, Constantinidou C, Millard A. Modification of Bacteriophages to Increase Their Association with Lung Epithelium Cells In Vitro. Pharmaceuticals. 2021; 14(4):308. https://doi.org/10.3390/ph14040308
Chicago/Turabian StyleGrigonyte, Aurelija M., Alexia Hapeshi, Chrystala Constantinidou, and Andrew Millard. 2021. "Modification of Bacteriophages to Increase Their Association with Lung Epithelium Cells In Vitro" Pharmaceuticals 14, no. 4: 308. https://doi.org/10.3390/ph14040308
APA StyleGrigonyte, A. M., Hapeshi, A., Constantinidou, C., & Millard, A. (2021). Modification of Bacteriophages to Increase Their Association with Lung Epithelium Cells In Vitro. Pharmaceuticals, 14(4), 308. https://doi.org/10.3390/ph14040308






