Novel N4-Like Bacteriophages of Pectobacterium atrosepticum
Abstract
1. Introduction
2. Results
2.1. Isolation of SRE from Potato Crops Symptomatic of Blackleg
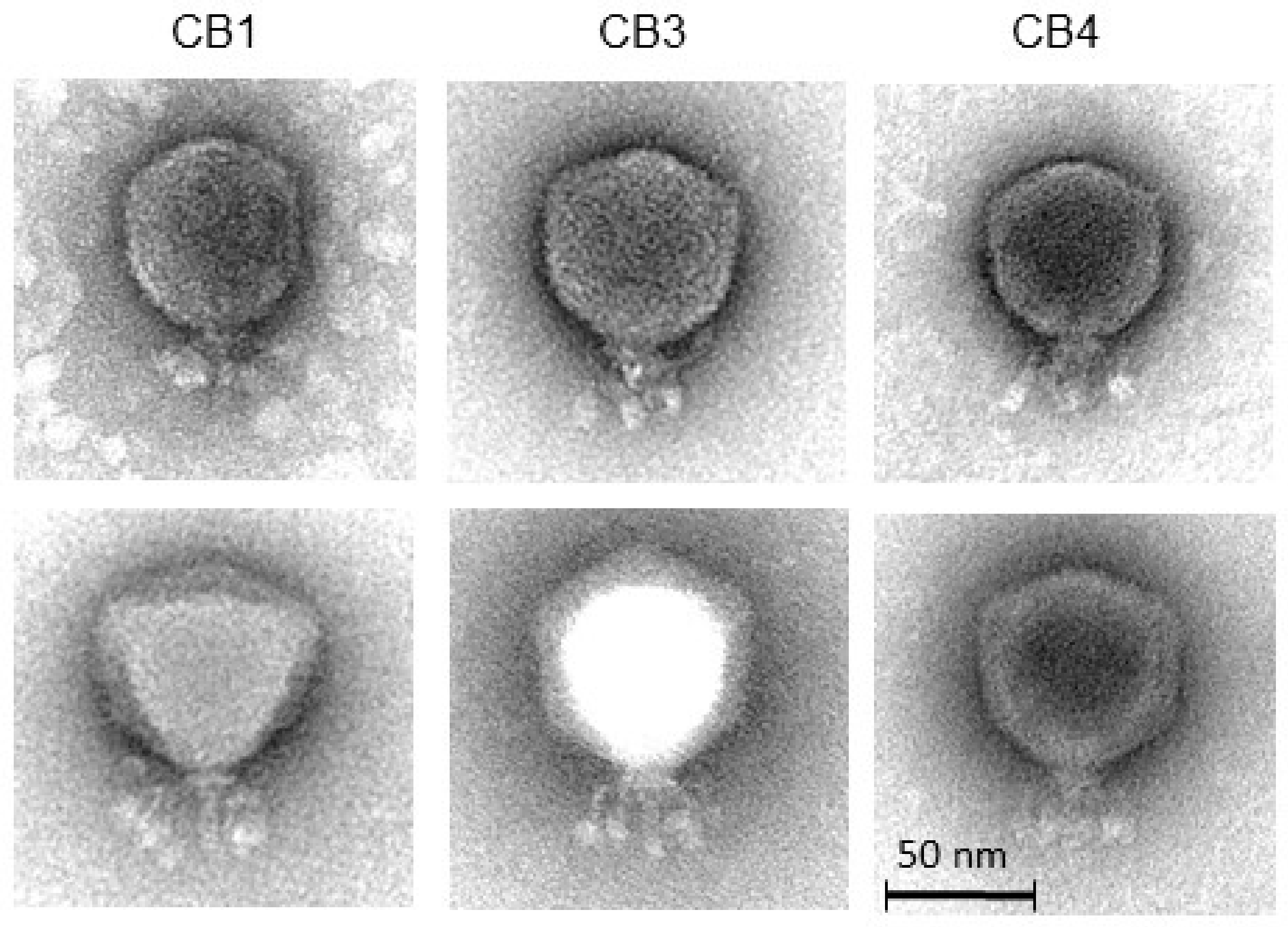
2.2. Isolation of Phages, Host Range and General Characteristics
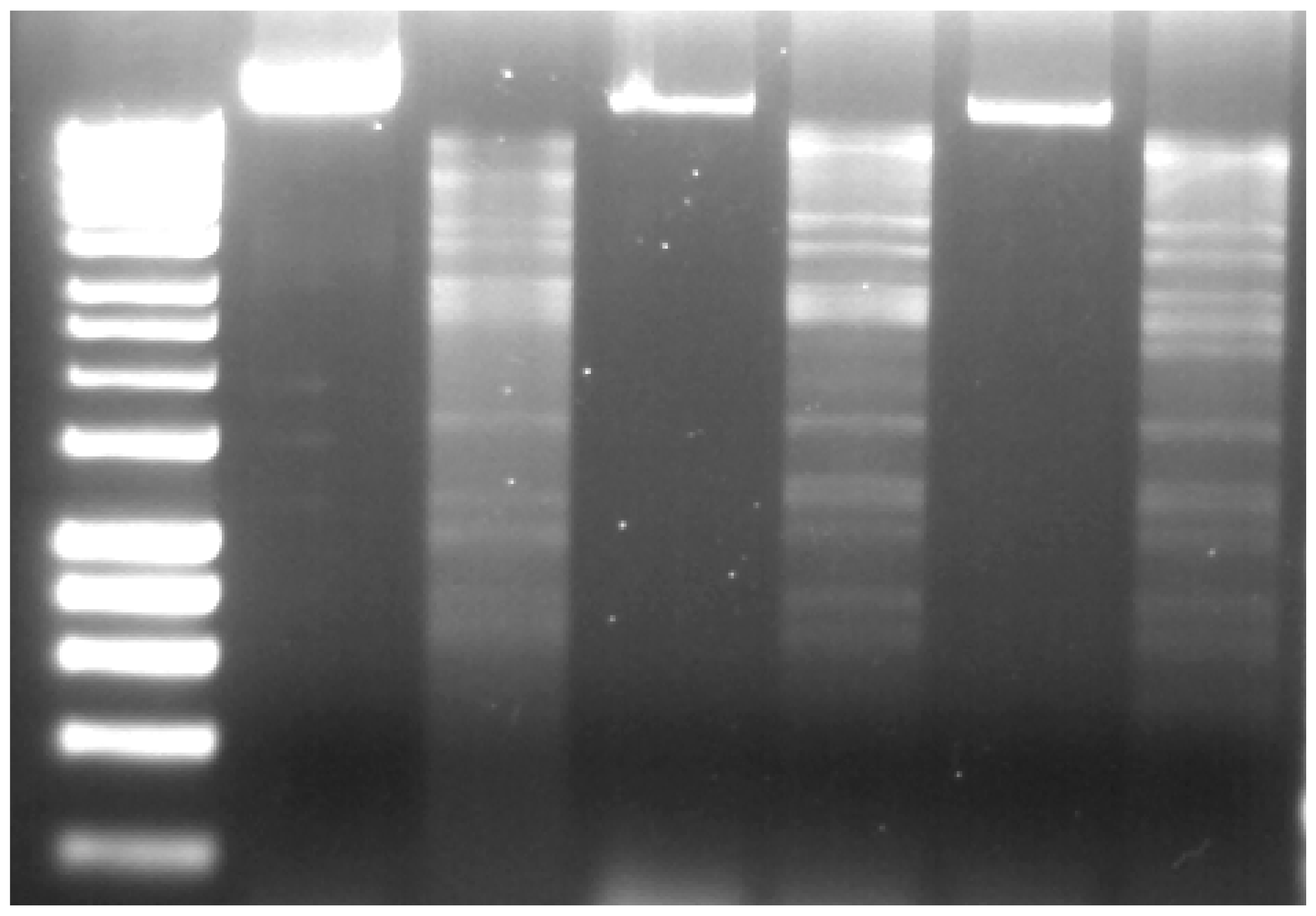
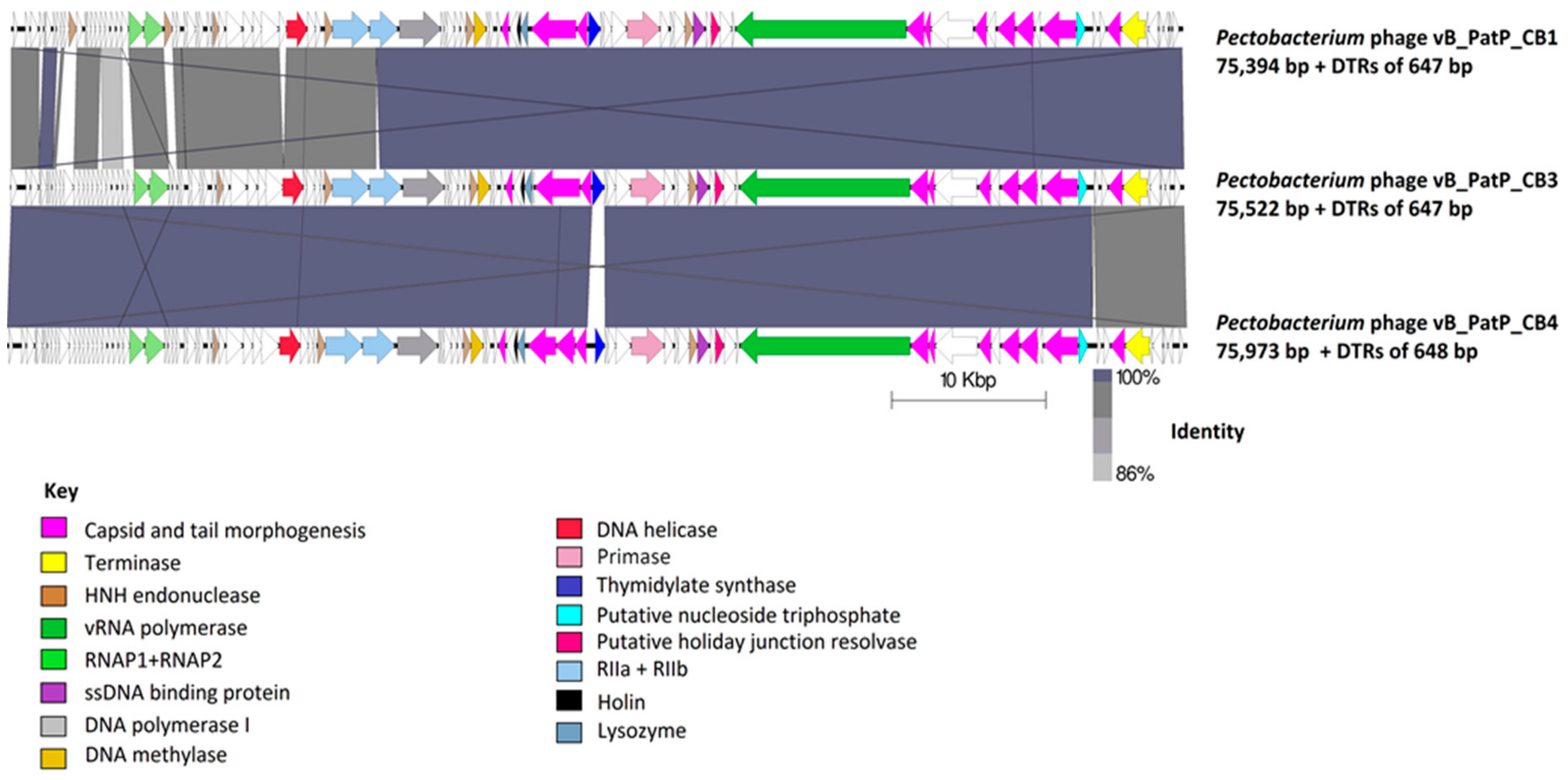
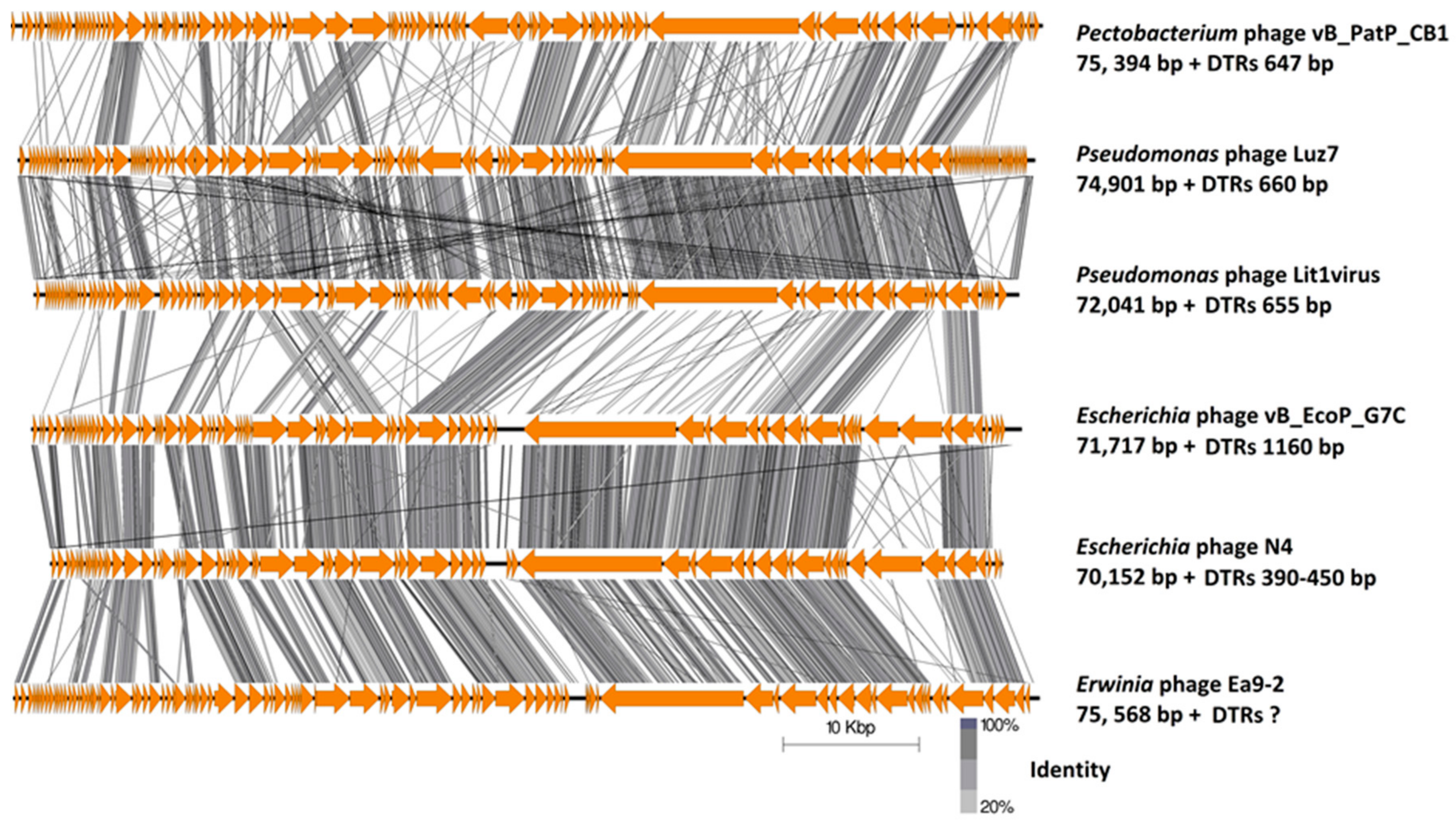
2.3. Genome and Proteome Analysis
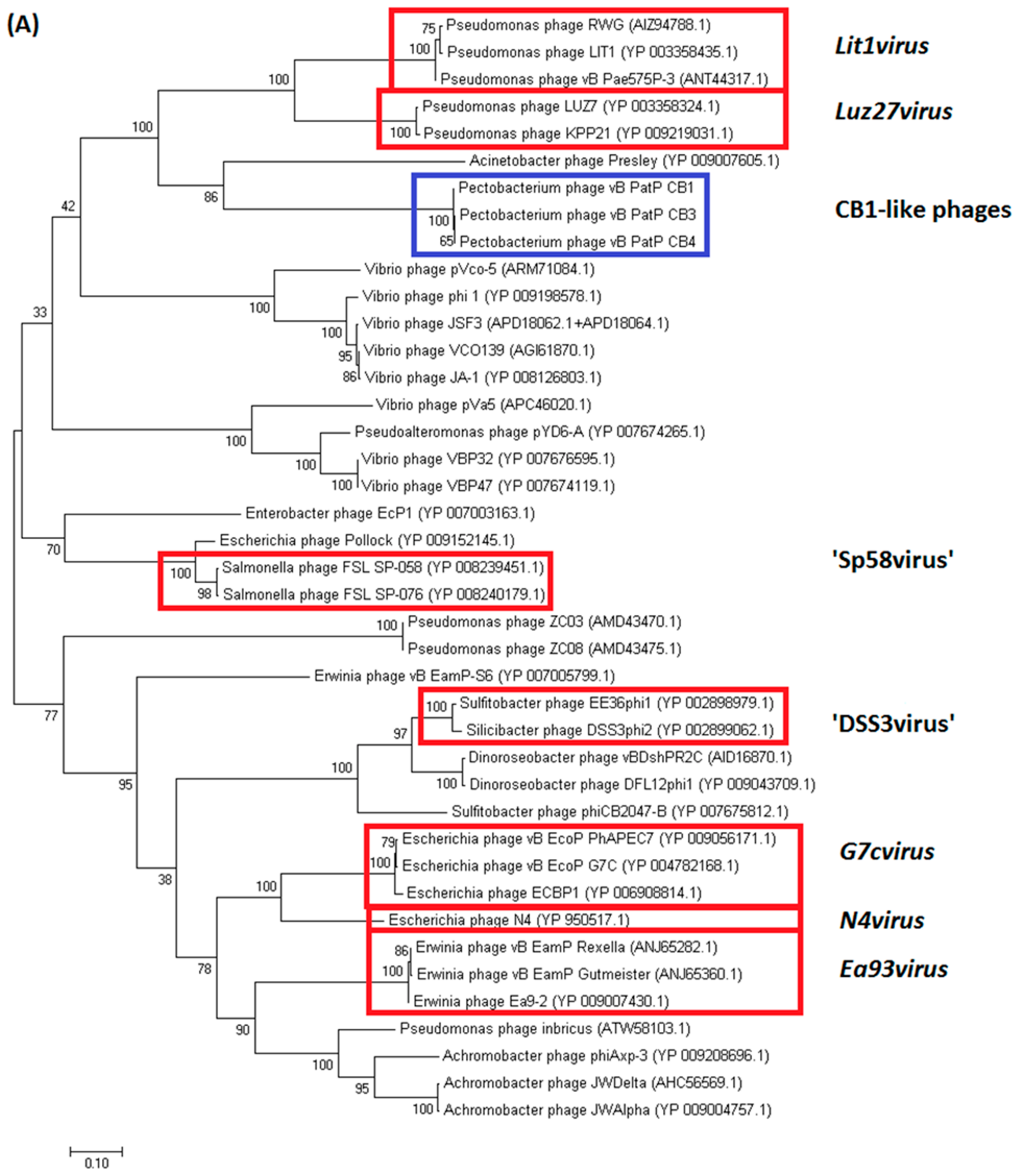
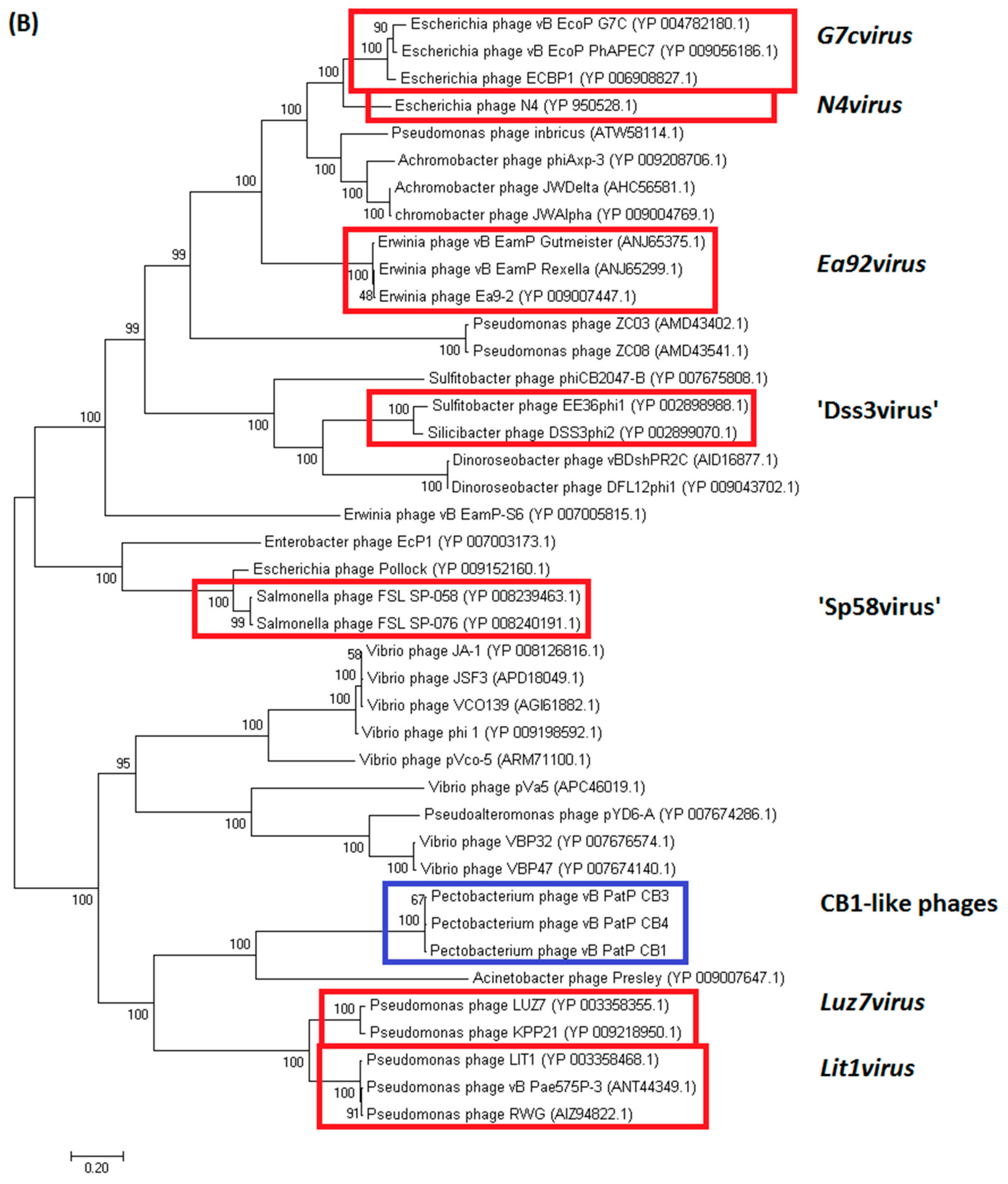
2.3.1. Genomes of Phages CB1, CB3, and CB4 Show an N4virus Organization
2.3.2. Transcription
2.3.3. DNA Replication, Metabolism and Methylation
2.3.4. Cell Lysis
2.3.5. Structural Proteome of Phage CB1
2.3.6. Selfish Genetic Elements
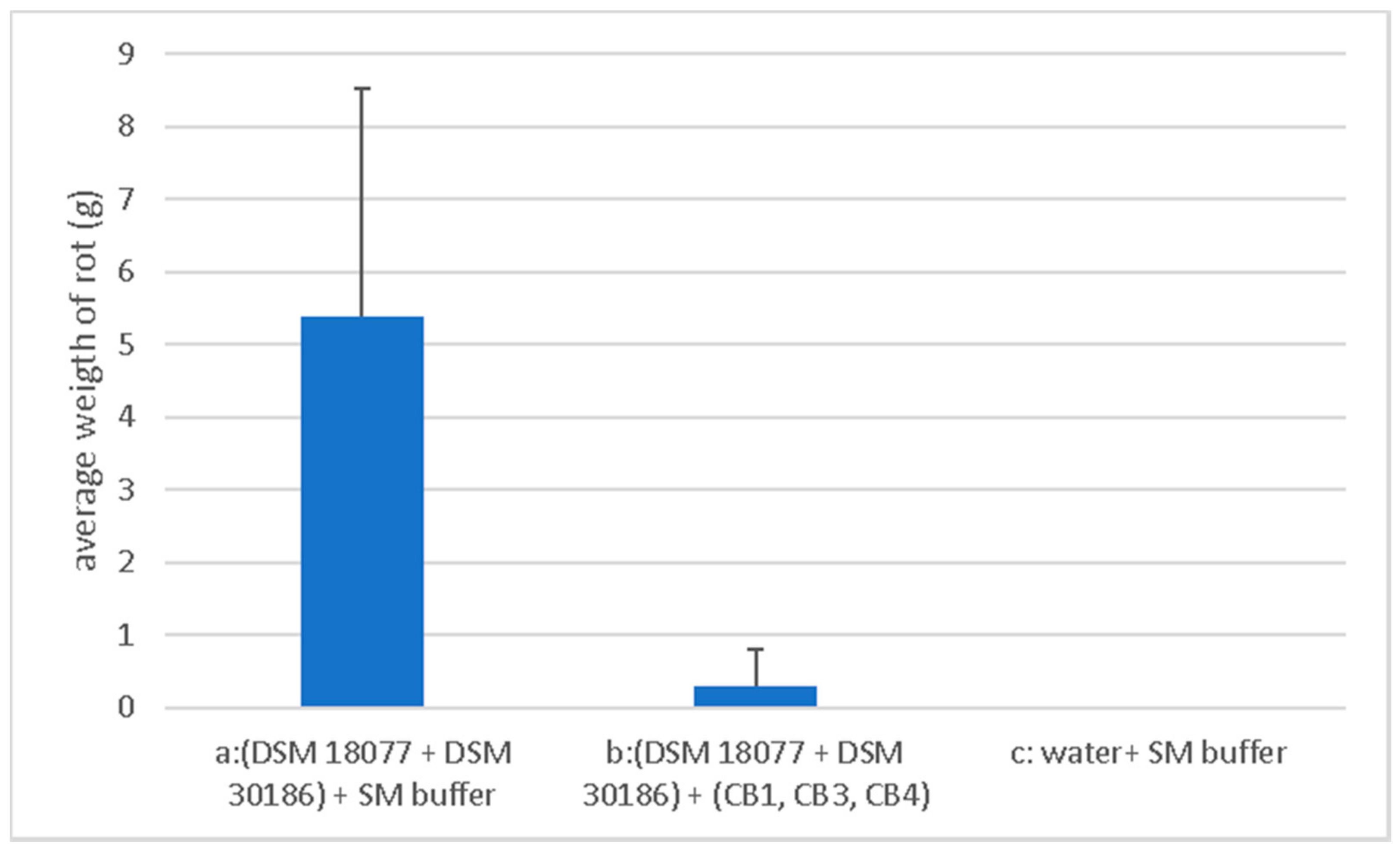
2.4. Phage Biocontrol on Whole Tubers
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Phage and Cultivation Conditions
4.2. Phage Isolation
4.3. Host Range and General Characterization
4.4. CsCl Gradient Purification
4.5. Transmission Electron Microscopy
4.6. DNA Isolation, Restriction and Sequencing
4.7. Bioinformatic Analysis
4.8. Comparative Genomics
4.9. Phage CB1 Virion ESI-MS/MS Proteome Analysis
4.10. Biocontrol Assays to Determine Biocontrol Potential CB1, CB3 and CB4 Mixture
4.11. Statistical Analyses of Data
4.12. Accession Number
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Toth, I.K.; van der Wolf, J.M.; Saddler, G.; Lojkowska, E.; Hélias, V.; Pirhonen, M.; Tsror Lahkim, L.; Elphinstone, J.G. Dickeya species: An emerging problem for potato production in Europe. Plant Pathol. 2011, 60, 385–399. [Google Scholar] [CrossRef]
- Pérombelon, M.C.M. Potato diseases caused by soft rot erwinias: An overview of pathogenesis. Plant Pathol. 2002, 51, 1–12. [Google Scholar] [CrossRef]
- Mansfield, J.; Genin, S.; Magori, S.; Citovsky, V.; Sriariyanum, M.; Ronald, P.; Dow, M.A.X.; Verdier, V.; Beer, S.V.; Machado, M.A.; et al. Top 10 plant pathogenic bacteria in molecular plant pathology. Mol. Plant Pathol. 2012, 13, 614–629. [Google Scholar] [CrossRef] [PubMed]
- Waleron, M.; Waleron, K.; Lojkowska, E. Occurrence of Pectobacterium wasabiae in potato field samples. Eur. J. Plant Pathol. 2013, 137, 149–158. [Google Scholar] [CrossRef][Green Version]
- Van der Wolf, J.M.; de Haan, E.G.; Kastelein, P.; Krijger, M.; de Haas, B.H.; Velvis, H.; Mendes, O.; Kooman-Gersmann, M.; van der Zouwen, P.S. Virulence of Pectobacterium carotovorum subsp. brasiliense on potato compared with that of other Pectobacterium and Dickeya species under climatic conditions prevailing in the Netherlands. Plant Pathol. 2017, 66, 571–583. [Google Scholar] [CrossRef]
- Khayi, S.; Cigna, J.; Chong, T.M.; Quêtu-Laurent, A.; Chan, K.-G.; Hélias, V.; Faure, D. Transfer of the potato plant isolates of Pectobacterium wasabiae to Pectobacterium parmentieri sp. nov. Int. J. Syst. Evol. Microbiol. 2016, 66, 5379–5383. [Google Scholar] [CrossRef] [PubMed]
- Toth, I.K.; Bell, K.S.; Holeva, M.C.; Birch, P.R.J. Soft rot erwiniae: From genes to genomes. Mol. Plant Pathol. 2003, 4, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Czajkowski, R.; Pérombelon, M.C.M.; van Veen, J.A.; van der Wolf, J.M. Control of blackleg and tuber soft rot of potato caused by Pectobacterium and Dickeya species: A review. Plant Pathol. 2011, 60, 999–1013. [Google Scholar] [CrossRef]
- De Boer, S.H. Blackleg of potato. Plant Heal. Instr. 2004. [Google Scholar] [CrossRef]
- Buttimer, C.; McAuliffe, O.; Ross, R.P.P.; Hill, C.; O’Mahony, J.; Coffey, A.; O’Mahony, J.; Coffey, A. Bacteriophages and bacterial plant diseases. Front. Microbiol. 2017, 8, 34. [Google Scholar] [CrossRef] [PubMed]
- Frampton, R.A.; Pitman, A.R.; Fineran, P.C. Advances in bacteriophage-mediated control of plant pathogens. Int. J. Microbiol. 2012, 2012, 326452. [Google Scholar] [CrossRef] [PubMed]
- Czajkowski, R. Bacteriophages of soft rot Enterobacteriaceae—A minireview. FEMS Microbiol. Lett. 2016, 363, fnv230. [Google Scholar] [CrossRef] [PubMed]
- Smolarska, A.; Rabalski, L.; Narajczyk, M.; Czajkowski, R. Isolation and phenotypic and morphological characterization of the first Podoviridae lytic bacteriophages ϕA38 and ϕA41 infecting Pectobacterium parmentieri (former Pectobacterium wasabiae). Eur. J. Plant Pathol. 2018, 150, 413–425. [Google Scholar] [CrossRef]
- Adriaenssens, E.M.; Van Vaerenbergh, J.; Vandenheuvel, D.; Dunon, V.; Ceyssens, P.-J.; De Proft, M.; Kropinski, A.M.; Noben, J.-P.; Maes, M.; Lavigne, R. T4-related bacteriophage LIMEstone isolates for the control of soft rot on potato caused by ‘Dickeya solani’. PLoS ONE 2012, 7, e33227. [Google Scholar] [CrossRef] [PubMed]
- Czajkowski, R.; Ozymko, Z.; de Jager, V.; Siwinska, J.; Smolarska, A.; Ossowicki, A.; Narajczyk, M.; Lojkowska, E. Genomic, proteomic and morphological characterization of two novel broad host lytic bacteriophages ΦPD10.3 and ΦPD23.1 infecting pectinolytic Pectobacterium spp. and Dickeya spp. PLoS ONE 2015, 10, e0119812. [Google Scholar] [CrossRef] [PubMed]
- Ravensdale, M.; Blom, T.J.; Gracia-Garza, J.A.; Svircev, A.M.; Smith, R.J. Bacteriophages and the control of Erwinia carotovora subsp. carotovora. Can. J. Plant Pathol. 2010, 29, 121–130. [Google Scholar] [CrossRef]
- Lim, J.-A.; Jee, S.; Lee, D.H.; Roh, E.; Jung, K.; Oh, C.; Heu, S. Biocontrol of Pectobacterium carotovorum subsp. carotovorum using bacteriophage PP1. J. Microbiol. Biotechnol. 2013, 23, 1147–1153. [Google Scholar] [CrossRef] [PubMed]
- Molina, A.M.; Pesce, A.; Shito, G.C. Un nuovo batteriofago attivo sul ceppo K12 di E. coli. I. Caratteristiche biologiche. Boll. Inst. Sieroter. 1965, 44, 329–337. [Google Scholar]
- Glucksmann, M.A.; Markiewicz, P.; Malone, C.; Rothman-Denes, L.B. Specific sequences and a hairpin structure in the template strand are required for N4 virion RNA polymerase promoter recognition. Cell 1992, 70, 491–500. [Google Scholar] [CrossRef]
- Adriaenssens, E.M.; C Clokie, M.R.; Sullivan, M.B.; Gillis, A.; Jens Kuhn, B.H.; Kropinski, A.M. Taxonomy of prokaryotic viruses: 2016 update from the ICTV bacterial and archaeal viruses subcommittee. Arch. Virol. 2017, 162, 1153–1157. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, J.; Klumpp, J.; Moreno Switt, A.I.; Yagubi, A.; Ackermann, H.-W.; Wiedmann, M.; Svircev, A.; Nash, J.H.E.; Kropinski, A.M. Taxonomic reassessment of N4-like viruses using comparative genomics and proteomics suggests a new subfamily—“Enquartavirinae”. Arch. Virol. 2015, 160, 3053–3062. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, J.; Kropinski, A.M.; Adriaenssens, E.M.; Ackermann, H.-W.; Lavigne, R.; Kuhn, J.H.; Uchiyama, J. To Create a New Genus, Luz7virus, including 2 (Two) New Species within the Family Podoviridae. Available online: https://talk.ictvonline.org/ICTV/proposals/2016.024a-dB.A.v2.Luz7virus.pdf (accessed on 22 January 2018).
- Wittmann, J.; Grose, J.H.; Yagubi, A.I.; Svircev, A.M.; Kropinski, A.M. To Create a New Genus, Ea92virus, Including 2 (Two) New Species within the Family Podoviridae. Available online: https://talk.ictvonline.org/files/ictv_official_taxonomy_updates_since_the_8th_report/m/prokaryote-official/6772 (accessed on 22 January 2018).
- Ackermann, H.W. Frequency of morphological phage descriptions in the year 2000. Brief review. Arch. Virol. 2001, 146, 843–857. [Google Scholar] [CrossRef] [PubMed]
- Kropinski, A.M.; Prangishvili, D.; Lavigne, R. Position paper: The creation of a rational scheme for the nomenclature of viruses of Bacteria and Archaea. Environ. Microbiol. 2009, 11, 2775–2777. [Google Scholar] [CrossRef] [PubMed]
- Fouts, D.E.; Klumpp, J.; Bishop-Lilly, K.A.; Rajavel, M.; Willner, K.M.; Butani, A.; Henry, M.; Biswas, B.; Li, M.; Albert, M.; et al. Whole genome sequencing and comparative genomic analyses of two Vibrio cholerae O139 Bengal-specific Podoviruses to other N4-like phages reveal extensive genetic diversity. Virol. J. 2013, 10, 165. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Fan, H.; An, X.; Fan, H.; Jiang, H.; Chen, Y.; Tong, Y. Scrutinizing virus genome termini by high-throughput sequencing. PLoS ONE 2014, 9, e85806. [Google Scholar] [CrossRef] [PubMed]
- Ohmori, H.; Haynes, L.L.; Rothman-Denes, L.B. Structure of the ends of the coliphage N4 genome. J. Mol. Biol. 1988, 202, 1–10. [Google Scholar] [CrossRef]
- Bell, K.S.; Sebaihia, M.; Pritchard, L.; Holden, M.T.G.; Hyman, L.J.; Holeva, M.C.; Thomson, N.R.; Bentley, S.D.; Churcher, L.J.C.; Mungall, K.; et al. Genome sequence of the enterobacterial phytopathogen Erwinia carotovora subsp. atroseptica and characterization of virulence factors. Proc. Natl. Acad. Sci. USA 2004, 101, 11105–11110. [Google Scholar] [CrossRef] [PubMed]
- Nikolaichik, Y.; Gorshkov, V.; Gogolev, Y.; Valentovich, L.; Evtushenkov, A. Genome sequence of Pectobacterium atrosepticum strain 21A. Genome Announc. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Adriaenssens, E.M.; Rodney Brister, J. How to name and classify your phage: An informal guide. Viruses 2017, 9, 70. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhang, S.; Long, L.; Huang, S. Characterization and complete genome sequences of three N4-Like Roseobacter phages isolated from the south China sea. Curr. Microbiol. 2016, 73, 409–418. [Google Scholar] [CrossRef] [PubMed]
- Ceyssens, P.-J.; Brabban, A.; Rogge, L.; Lewis, M.S.; Pickard, D.; Goulding, D.; Dougan, G.; Noben, J.-P.; Kropinski, A.; Kutter, E.; et al. Molecular and physiological analysis of three Pseudomonas aeruginosa phages belonging to the “N4-like viruses”. Virology 2010, 405, 26–30. [Google Scholar] [CrossRef] [PubMed]
- Kulikov, E.; Kropinski, A.M.; Golomidova, A.; Lingohr, E.; Govorun, V.; Serebryakova, M.; Prokhorov, N.; Letarova, M.; Manykin, A.; Strotskaya, A.; et al. Isolation and characterization of a novel indigenous intestinal N4-related coliphage vB_EcoP_G7C. Virology 2012, 426, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Carter, R.H.; Demidenko, A.A.; Hattingh-Willis, S.; Rothman-Denes, L.B. Phage N4 RNA polymerase II recruitment to DNA by a single-stranded DNA-binding protein. Genes Dev. 2003, 17, 2334–2345. [Google Scholar] [CrossRef] [PubMed]
- Choi, M.; Miller, A.; Cho, N.Y.; Rothman-Denes, L.B. Identification, cloning, and characterization of the bacteriophage N4 gene encoding the single-stranded DNA-binding protein. A protein required for phage replication, recombination, and late transcription. J. Biol. Chem. 1995, 270, 22541–22547. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.; Mahony, J.; Ainsworth, S.; Nauta, A.; van Sinderen, D. Bacteriophage orphan DNA methyltransferases: Insights from their bacterial origin, function, and occurrence. Appl. Environ. Microbiol. 2013, 79, 7547–7555. [Google Scholar] [CrossRef] [PubMed]
- Stojković, E.A.; Rothman-Denes, L.B. Coliphage N4 N-acetylmuramidase defines a new family of murein hydrolases. J. Mol. Biol. 2007, 366, 406–419. [Google Scholar] [CrossRef] [PubMed]
- Savva, C.G.; Dewey, J.S.; Moussa, S.H.; To, K.H.; Holzenburg, A.; Young, R. Stable micron-scale holes are a general feature of canonical holins. Mol. Microbiol. 2014, 91, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Young, R. Phage lysis: Three steps, three choices, one outcome. J. Microbiol. 2014, 52, 243–258. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.H.; McPartland, J.; Kaganman, I.; Bowman, V.D.; Rothman-Denes, L.B.; Rossmann, M.G. Insight into DNA and protein transport in double-stranded DNA viruses: The structure of bacteriophage N4. J. Mol. Biol. 2008, 378, 726–736. [Google Scholar] [CrossRef] [PubMed]
- Hardies, S.C.; Thomas, J.A.; Black, L.; Weintraub, S.T.; Hwang, C.Y.; Cho, B.C. Identification of structural and morphogenesis genes of Pseudoalteromonas phage φRIO-1 and placement within the evolutionary history of Podoviridae. Virology 2016, 489, 116–127. [Google Scholar] [CrossRef] [PubMed]
- Prokhorov, N.S.; Riccio, C.; Zdorovenko, E.L.; Shneider, M.M.; Browning, C.; Knirel, Y.A.; Leiman, P.G.; Letarov, A.V. Function of bacteriophage G7C esterase tailspike in host cell adsorption. Mol. Microbiol. 2017, 105, 385–398. [Google Scholar] [CrossRef] [PubMed]
- Bord Bia the Irish Food Board Potato Industry Welcomes €1 Million Marketing Boost. Available online: https://www.bordbia.ie/corporate/press/2015/pages/Potatoindustry1millionmarketingboost.aspx (accessed on 22 January 2018).
- Shigehisa, R.; Uchiyama, J.; Kato, S.; Takemura-Uchiyama, I.; Yamaguchi, K.; Miyata, R.; Ujihara, T.; Sakaguchi, Y.; Okamoto, N.; Shimakura, H.; et al. Characterization of Pseudomonas aeruginosa phage KPP21 belonging to family Podoviridae genus N4-like viruses isolated in Japan. Microbiol. Immunol. 2016, 60, 64–67. [Google Scholar] [CrossRef] [PubMed]
- Katharios, P.; Kalatzis, P.G.; Kokkari, C.; Sarropoulou, E.; Middelboe, M. Isolation and characterization of a N4-like lytic bacteriophage infecting Vibrio splendidus, a pathogen of fish and bivalves. PLoS ONE 2017, 12, e0190083. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.Z.-M.; Millard, A.D.; Mann, N.H.; Schäfer, H. Comparative genomics defines the core genome of the growing N4-like phage genus and identifies N4-like Roseophage specific genes. Front. Microbiol. 2014, 5, 506. [Google Scholar] [CrossRef] [PubMed]
- García, P.; Martínez, B.; Obeso, J.M.; Rodríguez, A. Bacteriophages and their application in food safety. Lett. Appl. Microbiol. 2008, 47, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Chan, B.K.; Abedon, S.T.; Loc-Carrillo, C. Phage cocktails and the future of phage therapy. Future Microbiol. 2013, 8, 769–783. [Google Scholar] [CrossRef] [PubMed]
- Hélias, V.; Hamon, P.; Huchet, E.; Wolf, J.V.D.; Andrivon, D. Two new effective semiselective crystal violet pectate media for isolation of Pectobacterium and Dickeya. Plant Pathol. 2012, 61, 339–345. [Google Scholar] [CrossRef]
- Perombelon, M.C.M.; Van Der Wolf, J.M. Methods for the Detection and Quantification of Erwinia carotovora subsp. atroseptica (Pectobacterium carotovorum subsp. atrosepticum) on Potatoes: A Laboratory Manual; Scottish Crop Research Institute Occasional Publication: Dundee, UK, 2002. [Google Scholar]
- Darrasse, A.; Priou, S.; Kotoujansky, A.; Bertheau, Y. PCR and restriction fragment length polymorphism of a pel gene as a tool to identify Erwinia carotovora in relation to potato diseases. Appl. Environ. Microbiol. 1994, 60, 1437–1443. [Google Scholar] [PubMed]
- De Boer, S.H.; Ward, L.J. PCR detection of Erwinia carotovora subsp atroseptica associated with potato tissue. Phytopathology 1995, 85, 854–858. [Google Scholar] [CrossRef]
- Kang, H.W.; Kwon, S.W.; Go, S.J. PCR-based specific and sensitive detection of Pectobacterium carotovorum ssp. carotovorum by primers generated from a URP-PCR fingerprinting-derived polymorphic band. Plant Pathol. 2003, 52, 127–133. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Picking bacteriophage Lamda plaques. In Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001; Volume 1, p. 2.32. ISBN 0879695773. [Google Scholar]
- Park, M.; Lee, J.-H.; Shin, H.; Kim, M.; Choi, J.; Kang, D.-H.; Heu, S.; Ryu, S. Characterization and comparative genomic analysis of a novel bacteriophage, SFP10, simultaneously inhibiting both Salmonella enterica and Escherichia coli O157:H7. Appl. Environ. Microbiol. 2012, 78, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Liang, L.; Lin, S.; Jia, S. Isolation and characterization of a virulent bacteriophage AB1 of Acinetobacter baumannii. BMC Microbiol. 2010, 10, 131. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D.W. Purification of bacteriophage lamda particles by isopycnic centrifugation through CsCl gradients. In Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001; Volume 1, p. 2.47. ISBN 0879695773. [Google Scholar]
- Casey, E.; Mahony, J.; O’Connell-Motherway, M.; Bottacini, F.; Cornelissen, A.; Neve, H.; Heller, K.J.; Noben, J.-P.; Dal Bello, F.; van Sinderen, D. Molecular characterization of three Lactobacillus delbrueckii subsp. bulgaricus phages. Appl. Environ. Microbiol. 2014, 80, 5623–5635. [Google Scholar] [CrossRef] [PubMed]
- Pickard, D.J.J. Preparation of bacteriophage lysates and pure DNA. Methods Mol. Biol. 2009, 502, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Delcher, A.L.; Harmon, D.; Kasif, S.; White, O.; Salzberg, S.L. Improved microbial gene identification with GLIMMER. Nucleic Acids Res. 1999, 27, 4636–4641. [Google Scholar] [CrossRef] [PubMed]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2015, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, A.; Chang, H.-Y.; Daugherty, L.; Fraser, M.; Hunter, S.; Lopez, R.; McAnulla, C.; McMenamin, C.; Nuka, G.; Pesseat, S.; et al. The InterPro protein families database: The classification resource after 15 years. Nucleic Acids Res. 2014, 43, D213–D221. [Google Scholar] [CrossRef] [PubMed]
- Söding, J.; Biegert, A.; Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 2005, 33, W244–W248. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [PubMed]
- Juncker, A.S.; Willenbrock, H.; von Heijne, G.; Brunak, S.; Nielsen, H.; Krogh, A. Prediction of lipoprotein signal peptides in Gram-negative bacteria. Protein Sci. 2003, 12, 1652–1662. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Laslett, D.; Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 2004, 32, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Naville, M.; Ghuillot-Gaudeffroy, A.; Marchais, A.; Gautheret, D. ARNold: A web tool for the prediction of Rho-independent transcription terminators. RNA Biol. 2011, 8, 11–13. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef] [PubMed]
- Carver, T.J.; Rutherford, K.M.; Berriman, M.; Rajandream, M.-A.; Barrell, B.G.; Parkhill, J. ACT: The Artemis comparison tool. Bioinformatics 2005, 21, 3422–3423. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Bioinformatics 1992, 8, 275–282. [Google Scholar] [CrossRef]
- Ågren, J.; Sundström, A.; Håfström, T.; Segerman, B. Gegenees: Fragmented alignment of multiple genomes for determining phylogenomic distances and genetic signatures unique for specified target groups. PLoS ONE 2012, 7, e39107. [Google Scholar] [CrossRef] [PubMed]
- Van den Bossche, A.; Ceyssens, P.-J.; De Smet, J.; Hendrix, H.; Bellon, H.; Leimer, N.; Wagemans, J.; Delattre, A.-S.; Cenens, W.; Aertsen, A.; et al. Systematic identification of hypothetical bacteriophage proteins targeting key protein complexes of Pseudomonas aeruginosa. J. Proteome Res. 2014, 13, 4446–4456. [Google Scholar] [CrossRef] [PubMed]

| Bacteria | Bacteriophage Infection | |||
|---|---|---|---|---|
| Species | Strain | CB1 | CB3 | CB4 |
| Pectobacterium atrosepticum | DSMZ 18077 (type strain) | 1.000 * | 2.263 | 0.909 |
| DSMZ 30184 | 2.00 × 10−5 | CS | CS | |
| DSMZ 30185 | 0.371 | 0.126 | 0.136 | |
| DSMZ 30186 | 0.003 | 1.000 * | 1.000 * | |
| CB BL1-1 | CS | CS | CS | |
| CB BL2-1 | 0.571 | 0.737 | 0.336 | |
| CB BL3-1 | 0.529 | 0.605 | 0.218 | |
| CB BL4-1 | 0.571 | 0.789 | 0.245 | |
| CB BL5-1 | CS | 0.158 | CS | |
| CB BL7-1 | CS | CS | CS | |
| CB BL9-1 | 0.027 | 0.007 | 0.164 | |
| CB BL11-1 | CS | 0.279 | 0.127 | |
| CB BL12-2 | 2.86 × 10−6 | 0.079 | 0.255 | |
| CB BL13-1 | 1.71 × 10−5 | 0.063 | 0.436 | |
| CB BL14-1 | CS | CS | CS | |
| CB BL15-1 | - | - | - | |
| CB BL16-1 | 0.005 | CS | CS | |
| CB BL18-1 | CS | 0.037 | CS | |
| CB BL19-1 | 0.005 | CS | CS | |
| Pectobacterium carotovorum subsp. carotovorum | DSMZ 30168 (type strain) | - | - | - |
| DSMZ 30169 | - | - | - | |
| DSMZ 30170 | - | - | - | |
| CB BL19-1-37 | - | - | - | |
| Dickeya chrysanthemi bv. chrysanthemi | LMG 2804 | - | - | - |
| Dickeya dianthicola | PD 482 | - | - | - |
| PD 2174 | - | - | - | |
| GBBC 1538 | - | - | - | |
| Dickeya solani | sp. PRI 2222 | - | - | - |
| LMG 25865 | - | - | - | |
| GBBC 1502 | - | - | - | |
| GBBC 1586 | - | - | - | |
| Phage | Head (nm) | Tail Length (nm) | Collar Width (nm) | Whisker Length * (nm) | Whisker Ball Length (nm) | Whisker Ball Width (nm) |
|---|---|---|---|---|---|---|
| CB1 | 67.8 ± 4.4 (n = 19) | 23.9 ± 1.7 (n = 8) | 18.5 ± 1.3 (n = 8) | 23.4 ± 3.0 (n = 14) | 12.3 ± 1.4 (n = 16) | 6.3 ± 0.7 (n = 17) |
| CB3 | 71.8 ± 1.7 (n = 6) | 25.8 ± 2.3 (n = 8) | 19.6 ± 0.8 (n = 7) | 25.6 ± 1.7 (n = 7) | 12.0 ± 1.5 (n = 9) | 7.4 ± 0.9 (n = 9) |
| CB4 | 70.4 ± 2.5 (n = 12) | 25.5 ± 1.0 (n = 5) | 19.3 ± 0.8 (n = 6) | 23.5 ± 1.6 (n = 3) | 11.8 ± 0.9 (n = 5) | 6.6 ± 0.6 (n = 5) |
| ORF | Predicted Function | Molecular Mass (kDa) | No. of Unique Peptides | Sequence Coverage % |
|---|---|---|---|---|
| CB1_57 | Putative tail protein | 15.59 | 6 | 71 |
| CB1_61 | Putative tail spike protein | 104.98 | 19 | 29 |
| CB1_62 | Putative tail protein | 26.01 | 10 | 55 |
| CB1_77 | Virion associated RNA polymerase (N4 gp50-like) | 399.92 | 37 | 13 |
| CB1_78 | Unknown structural protein | 41.64 | 15 | 52 |
| CB1_79 | Structural protein (N4 gp52-like) | 14.2 | 3 | 19 |
| CB1_81 | Structural protein (N4 gp54-like) | 28.82 | 9 | 53 |
| CB1_83 | Major capsid protein (N4 gp56-like) | 43.33 | 33 | 75 |
| CB1_86 | Portal protein (N4 gp59-like) | 83.31 | 27 | 45 |
| CB1_90 | Structural protein (N4 gp67-like) | 33.72 | 9 | 40 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Buttimer, C.; Hendrix, H.; Lucid, A.; Neve, H.; Noben, J.-P.; Franz, C.; O’Mahony, J.; Lavigne, R.; Coffey, A. Novel N4-Like Bacteriophages of Pectobacterium atrosepticum. Pharmaceuticals 2018, 11, 45. https://doi.org/10.3390/ph11020045
Buttimer C, Hendrix H, Lucid A, Neve H, Noben J-P, Franz C, O’Mahony J, Lavigne R, Coffey A. Novel N4-Like Bacteriophages of Pectobacterium atrosepticum. Pharmaceuticals. 2018; 11(2):45. https://doi.org/10.3390/ph11020045
Chicago/Turabian StyleButtimer, Colin, Hanne Hendrix, Alan Lucid, Horst Neve, Jean-Paul Noben, Charles Franz, Jim O’Mahony, Rob Lavigne, and Aidan Coffey. 2018. "Novel N4-Like Bacteriophages of Pectobacterium atrosepticum" Pharmaceuticals 11, no. 2: 45. https://doi.org/10.3390/ph11020045
APA StyleButtimer, C., Hendrix, H., Lucid, A., Neve, H., Noben, J.-P., Franz, C., O’Mahony, J., Lavigne, R., & Coffey, A. (2018). Novel N4-Like Bacteriophages of Pectobacterium atrosepticum. Pharmaceuticals, 11(2), 45. https://doi.org/10.3390/ph11020045








