Orexin Receptor Multimerization versus Functional Interactions: Neuropharmacological Implications for Opioid and Cannabinoid Signalling and Pharmacogenetics
Abstract
1. Introduction
The Molecular Biology of the Orexins
2. Orexin Receptor Variants
3. Orexin Signalling
3.1. Coupling to G Proteins and Other Effectors
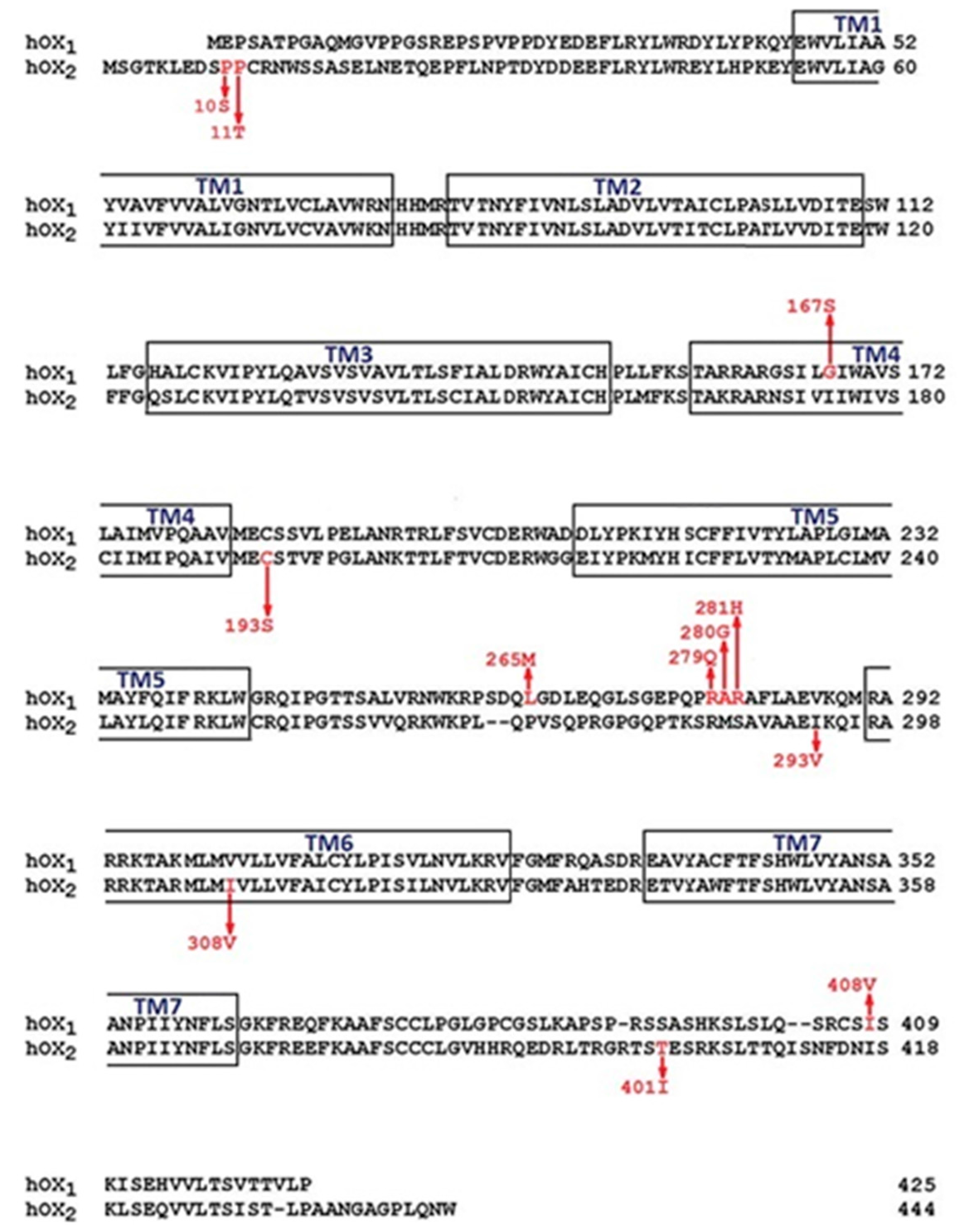
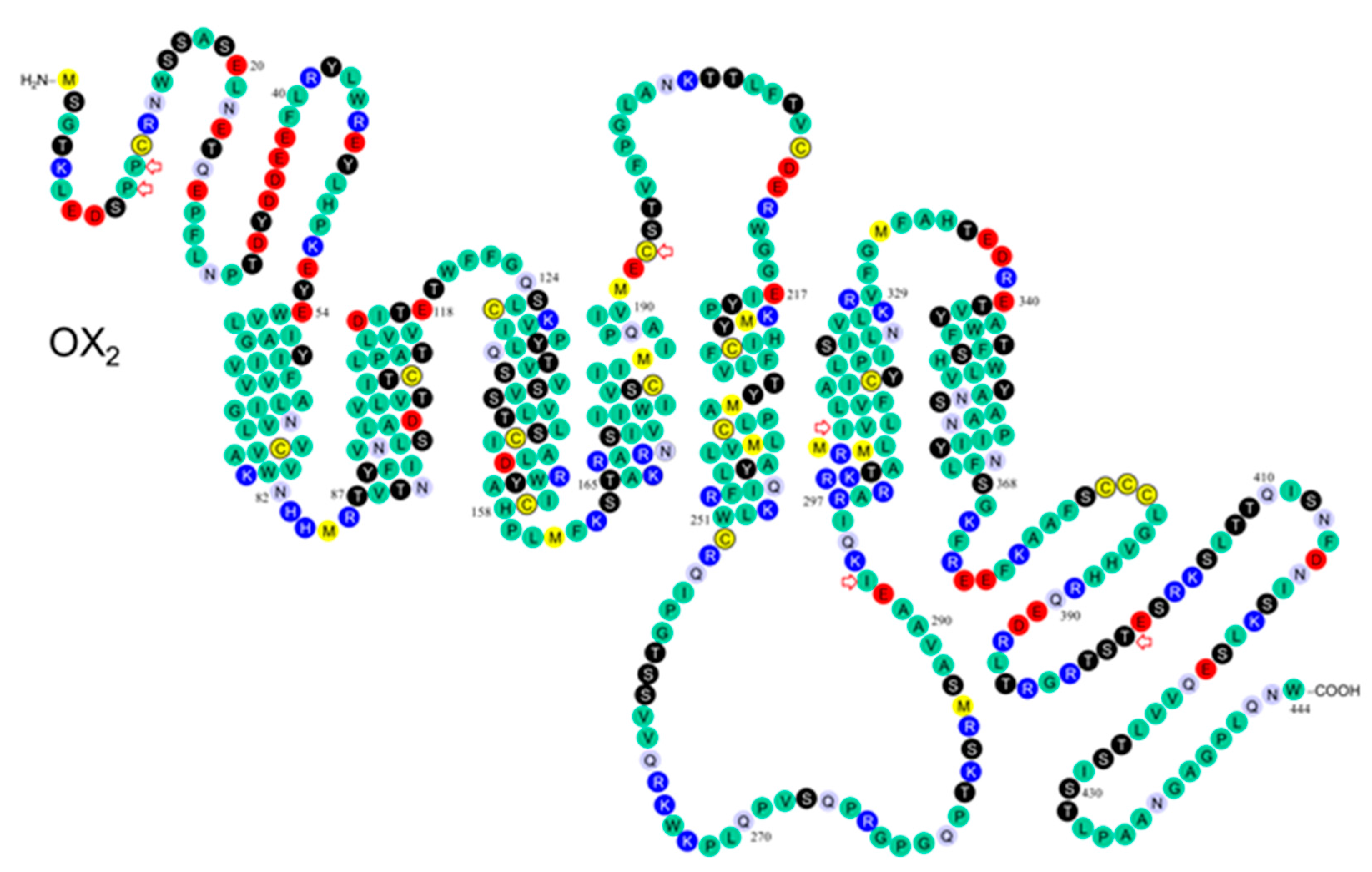
3.2. Post-Translational Modifications
3.3. Binding Pocket
4. Modelling Orexin Receptor Hetero- and Homodimerization
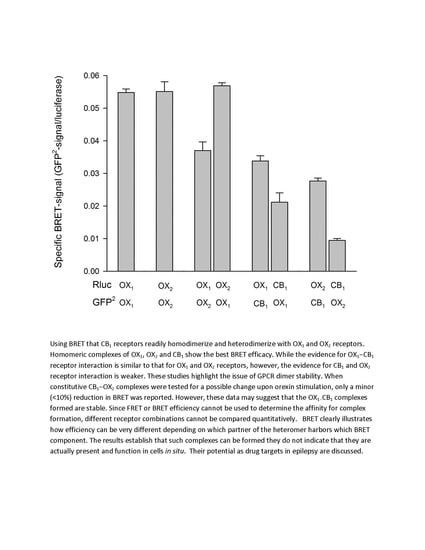
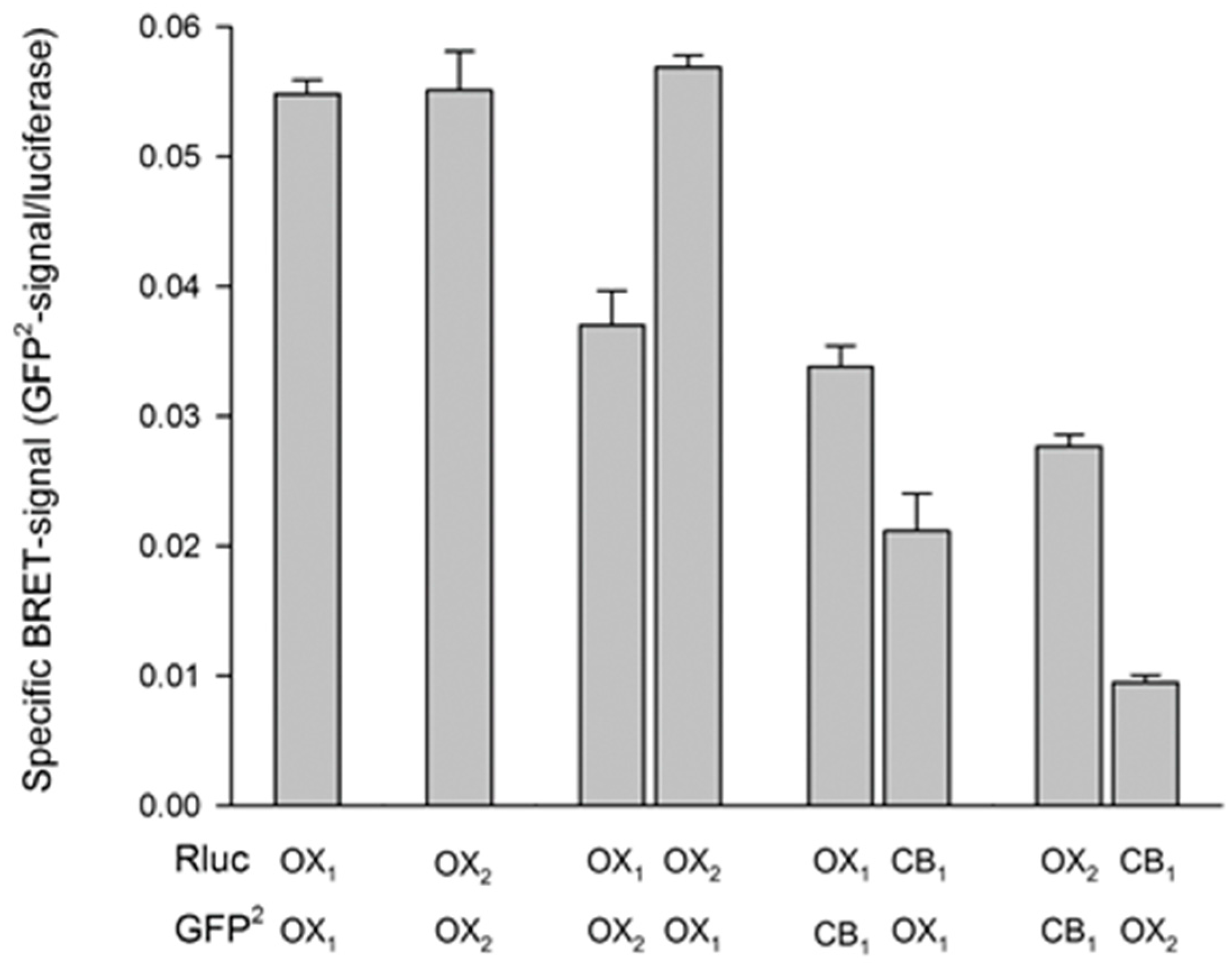
4.1. Evidence for Orexin Receptor Dimerization
4.2. Evidence for OX1-CRF1 Receptor Dimerization
4.3. OX1 Orexin Receptor and κ Opioid Receptor (κOR) Heteromerization
4.4. Interaction of OX1 Receptors and CB1 Cannabinoid Receptors
5. Relevance of CBl/OX1 Expression in the Hippocampus to Disease
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gencik, M.; Dahmen, N.; Wieczorek, S.; Kasten, M.; Bierbrauer, J.; Anghelescu, I.; Szegedi, A.; Menezes Saecker, A.M.; Epplen, J.T. A prepro-orexin gene polymorphism is associated with narcolepsy. Neurology 2001, 56, 115–117. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Xhaard, H.; Sakurai, T.; Rainero, I.; Kukkonen, J.P. OX1 and OX2 orexin/hypocretin receptor pharmacogenetics. Front. Neurosci. 2014, 8, 57. [Google Scholar] [CrossRef] [PubMed]
- Winrow, C.J.; Renger, J.J. Discovery and development of orexin receptor antagonists as therapeutics for insomnia. Br. J. Pharmacol. 2014, 171, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Cole, D.E.; Capra, V.; Siminovitch, K.A.; Rovati, G.E.; Burnham, W.M.; Rana, B.K. Pharmacogenetics of the G protein-coupled receptors. Methods Mol. Biol. 2014, 1175, 189–242. [Google Scholar] [PubMed]
- Thompson, M.D.; Comings, D.E.; Abu-Ghazalah, R.; Jereseh, Y.; Lin, L.; Wade, J.; Sakurai, T.; Tokita, S.; Yoshida, T.; Tanaka, H.; et al. Variants of the orexin2/hcrt2 receptor gene identified in patients with excessive daytime sleepiness and patients with Tourette’s syndrome comorbidity. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2004, 129, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Kukkonen, J.P. Orexin/hypocretin signalling. Curr. Top Behav. Neurosci. 2017, 33, 17–50. [Google Scholar] [PubMed]
- Kukkonen, J.P.; Leonard, C.S. Orexin/hypocretin receptor signalling cascades. Br. J. Pharmacol. 2014, 171, 314–331. [Google Scholar] [CrossRef] [PubMed]
- Leonard, C.S.; Kukkonen, J.P. Orexin/hypocretin receptor signalling: A functional perspective. Br. J. Pharmacol. 2014, 171, 294–313. [Google Scholar] [CrossRef] [PubMed]
- Olafsdottir, B.R.; Rye, D.B.; Scammell, T.E.; Matheson, J.K.; Stefansson, K.; Gulcher, J.R. Polymorphisms in hypocretin/orexin pathway genes and narcolepsy. Neurology 2001, 57, 1896–1899. [Google Scholar] [CrossRef] [PubMed]
- Peyron, C.; Faraco, J.; Rogers, W.; Ripley, B.; Overeem, S.; Charnay, Y.; Nevsimalova, S.; Aldrich, M.; Reynolds, D.; Albin, R.; et al. A mutation in a case of early onset narcolepsy and a generalized absence of hypocretin peptides in human narcoleptic brains. Nat. Med. 2000, 6, 991–997. [Google Scholar] [PubMed]
- Rainero, I.; Rubino, E.; Gallone, S.; Fenoglio, P.; Picci, L.R.; Giobbe, L.; Ostacoli, L.; Pinessi, L. Evidence for an association between migraine and the hypocretin receptor 1 gene. J. Headache Pain 2011, 12, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Rainero, I.; Ostacoli, L.; Rubino, E.; Gallone, S.; Picci, L.R.; Fenoglio, P.; Negro, E.; Rosso, C.; De Martino, P.; De Marchi, M.; et al. Association between major mood disorders and the hypocretin receptor 1 gene. J. Affect. Disord. 2011, 130, 487–491. [Google Scholar] [CrossRef] [PubMed]
- Meerabux, J.; Iwayama, Y.; Sakurai, T.; Ohba, H.; Toyota, T.; Yamada, K.; Nagata, R.; Irukayama-Tomobe, Y.; Shimizu, H.; Yoshitsugu, K.; et al. Association of an orexin 1 receptor 408Val variant with polydipsia-hyponatremia in schizophrenic subjects. Biol. Psychiatry 2005, 58, 401–407. [Google Scholar] [CrossRef] [PubMed]
- Fukunaka, Y.; Shinkai, T.; Hwang, R.; Hori, H.; Utsunomiya, K.; Sakata, S.; Naoe, Y.; Shimizu, K.; Matsumoto, C.; Ohmori, O.; et al. The orexin 1 receptor (HCRTR1) gene as a susceptibility gene contributing to polydipsia-hyponatremia in schizophrenia. Neuromol. Med. 2007, 9, 292–297. [Google Scholar] [CrossRef] [PubMed]
- Annerbrink, K.; Westberg, L.; Olsson, M.; Andersch, S.; Sjodin, I.; Holm, G.; Allgulander, C.; Eriksson, E. Panic disorder is associated with the Val308Iso polymorphism in the hypocretin receptor gene. Psychiatr. Genet. 2011, 21, 85–89. [Google Scholar] [CrossRef] [PubMed]
- Schurks, M.; Limmroth, V.; Geissler, I.; Tessmann, G.; Savidou, I.; Engelbergs, J.; Kurth, T.; Diener, H.C.; Rosskopf, D. Association between migraine and the G1246A polymorphism in the hypocretin receptor 2 gene. Headache 2007, 47, 1195–1199. [Google Scholar] [CrossRef] [PubMed]
- Baumber, L.; Sjostrand, C.; Leone, M.; Harty, H.; Bussone, G.; Hillert, J.; Trembath, R.C.; Russell, M.B. A genome-wide scan and HCRTR2 candidate gene analysis in a European cluster headache cohort. Neurology 2006, 66, 1888–1893. [Google Scholar] [CrossRef] [PubMed]
- Schurks, M.; Kurth, T.; Geissler, I.; Tessmann, G.; Diener, H.C.; Rosskopf, D. Cluster headache is associated with the G1246A polymorphism in the hypocretin receptor 2 gene. Neurology 2006, 66, 1917–1919. [Google Scholar] [CrossRef] [PubMed]
- Rainero, I.; Gallone, S.; Rubino, E.; Ponzo, P.; Valfre, W.; Binello, E.; Fenoglio, P.; Gentile, S.; Anoaica, M.; Gasparini, M.; et al. Haplotype analysis confirms the association between the HCRTR2 gene and cluster headache. Headache 2008, 48, 1108–1114. [Google Scholar] [CrossRef] [PubMed]
- Schurks, M.; Kurth, T.; Geissler, I.; Tessmann, G.; Diener, H.C.; Rosskopf, D. The G1246A polymorphism in the hypocretin receptor 2 gene is not associated with treatment response in cluster headache. Cephalalgia 2007, 27, 363–367. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, T.; Amemiya, A.; Ishii, M.; Matsuzaki, I.; Chemelli, R.M.; Tanaka, H.; Williams, S.C.; Richardson, J.A.; Kozlowski, G.P.; et al. Orexins and orexin receptors: A family of hypothalamic neuropeptides and G protein-coupled receptors that regulate feeding behavior. Cell 1998, 92, 573–585. [Google Scholar] [CrossRef]
- de Lecea, L.; Kilduff, T.S.; Peyron, C.; Gao, X.; Foye, P.E.; Danielson, P.E.; Fukuhara, C.; Battenberg, E.L.; Gautvik, V.T.; Bartlett, F.S. The hypocretins: Hypothalamus-specific peptides with neuroexcitatory activity. Proc. Natl. Acad. Sci. USA 1998, 95, 322–327. [Google Scholar] [CrossRef] [PubMed]
- Kukkonen, J.P. Physiology of the orexinergic/hypocretinergic system: A revisit in 2012. Am. J. Physiol. Cell Physiol. 2013, 304, 2–32. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Faraco, J.; Li, R.; Kadotani, H.; Rogers, W.; Lin, X.; Qiu, X.; de Jong, P.J.; Nishino, S.; Mignot, E. The sleep disorder canine narcolepsy is caused by a mutation in the hypocretin (orexin) receptor 2 gene. Cell 1999, 98, 365–376. [Google Scholar] [CrossRef]
- Hungs, M.; Fan, J.; Lin, L.; Lin, X.; Maki, R.A.; Mignot, E. Identification and functional analysis of mutations in the hypocretin (orexin) genes of narcoleptic canines. Genome Res. 2001, 11, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Chemelli, R.M.; Willie, J.T.; Sinton, C.M.; Elmquist, J.K.; Scammell, T.; Lee, C.; Richardson, J.A.; Williams, S.C.; Xiong, Y.; Kisanuki, Y.; et al. Narcolepsy in orexin knockout mice: Molecular genetics of sleep regulation. Cell 1999, 98, 437–451. [Google Scholar] [CrossRef]
- Willie, J.T.; Chemelli, R.M.; Sinton, C.M.; Tokita, S.; Williams, S.C.; Kisanuki, Y.Y.; Marcus, J.N.; Lee, C.; Elmquist, J.K.; Kohlmeier, K.A.; et al. Distinct narcolepsy syndromes in Orexin receptor-2 and Orexin null mice: Molecular genetic dissection of non-REM and REM sleep regulatory processes. Neuron 2003, 38, 715–730. [Google Scholar] [CrossRef]
- Nishino, S.; Ripley, B.; Overeem, S.; Lammers, G.J.; Mignot, E. Hypocretin (orexin) deficiency in human narcolepsy. Lancet 2000, 355, 39–40. [Google Scholar] [CrossRef]
- Ida, T.; Nakahara, K.; Katayama, T.; Murakami, N.; Nakazato, M. Effect of lateral cerebroventricular injection of the appetite-stimulating neuropeptide, orexin and neuropeptide Y, on the various behavioral activities of rats. Brain Res. 1999, 821, 526–529. [Google Scholar] [CrossRef]
- Takahashi, N.; Okumura, T.; Yamada, H.; Kohgo, Y. Stimulation of gastric acid secretion by centrally administered orexin-A in conscious rats. Biochem. Biophys. Res. Commun. 1999, 254, 623–627. [Google Scholar] [CrossRef] [PubMed]
- Karteris, E.; Machado, R.J.; Chen, J.; Zervou, S.; Hillhouse, E.W.; Randeva, H.S. Food deprivation differentially modulates orexin receptor expression and signalling in rat hypothalamus and adrenal cortex. Am. J. Physiol. Endocrinol. Metab. 2005, 288, E1089–1100. [Google Scholar] [CrossRef] [PubMed]
- Malendowicz, L.K.; Tortorella, C.; Nussdorfer, G.G. Orexins stimulate corticosterone secretion of rat adrenocortical cells, through the activation of the adenylate cyclase-dependent signalling cascade. J. Steroid Biochem. Mol. Biol. 1999, 70, 185–188. [Google Scholar] [CrossRef]
- Lubkin, M.; Stricker-Krongrad, A. Independent feeding and metabolic actions of orexins in mice. Biochem. Biophys. Res. Commun. 1998, 253, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, Y.; Ueta, Y.; Serino, R.; Nomura, M.; Shibuya, I.; Yamashita, H. Effects of food restriction on the hypothalamic prepro-orexin gene expression in genetically obese mice. Brain Res. Bull. 2000, 51, 515–521. [Google Scholar] [CrossRef]
- Malherbe, P.; Roche, O.; Marcuz, A.; Kratzeisen, C.; Wettstein, J.G.; Bissantz, C. Mapping the binding pocket of dual antagonist almorexant to human orexin 1 and orexin 2 receptors: Comparison with the selective OX1 antagonist SB-674042 and the selective OX2 antagonist N-ethyl-2-[(6-methoxy-pyridin-3-yl)-(toluene-2-sulfonyl)-amino]-N-pyridin-3-ylmethyl-acetamide (EMPA). Mol. Pharmacol. 2010, 78, 81–93. [Google Scholar] [PubMed]
- Putula, J.; Kukkonen, J.P. Mapping of the binding sites for the OX1 orexin receptor antagonist, SB-334867, using orexin/hypocretin receptor chimaeras. Neurosci. Lett. 2012, 506, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Tran, D.T.; Bonaventure, P.; Hack, M.; Mirzadegan, T.; Dvorak, C.; Letavic, M.; Carruthers, N.; Lovenberg, T.; Sutton, S.W. Chimeric, mutant orexin receptors show key interactions between orexin receptors, peptides and antagonists. Eur. J. Pharmacol. 2011, 667, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Heifetz, A.; Morris, G.B.; Biggin, P.C.; Barker, O.; Fryatt, T.; Bentley, J.; Hallett, D.; Manikowski, D.; Pal, S.; Reifegerste, R.; et al. Study of human Orexin-1 and -2 G-protein-coupled receptors with novel and published antagonists by modeling, molecular dynamics simulations, and site-directed mutagenesis. Biochemistry 2012, 51, 3178–3197. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Babaoglu, K.; Brautigam, C.A.; Clark, L.; Shao, Z.; Scheuermann, T.H.; Harrell, C.M.; Gotter, A.L.; Roecker, A.J.; Winrow, C.J.; et al. Structure and ligand-binding mechanism of the human OX1 and OX2 orexin receptors. Nat. Struct. Mol. Biol. 2016, 23, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Mobarec, J.C.; Kolb, P.; Rosenbaum, D.M. Crystal structure of the human OX2 orexin receptor bound to the insomnia drug suvorexant. Nature 2015, 519, 247–250. [Google Scholar] [CrossRef] [PubMed]
- Karhu, L.; Turku, A.; Xhaard, H. Modeling of the OX1R-orexin-A complex suggests two alternative binding modes. BMC Struct. Biol. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Rainero, I.; Gallone, S.; Valfre, W.; Ferrero, M.; Angilella, G.; Rivoiro, C.; Rubino, E.; De Martino, P.; Savi, L.; Ferrone, M.; et al. A polymorphism of the hypocretin receptor 2 gene is associated with cluster headache. Neurology 2004, 63, 1286–1288. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Bowen, R.A.; Wong, B.Y.; Antal, J.; Liu, Z.; Yu, H.; Siminovitch, K.; Kreiger, N.; Rohan, T.E.; Cole, D.E. Whole genome amplification of buccal cell DNA: Genotyping concordance before and after multiple displacement amplification. Clin. Chem. Lab. Med. 2005, 43, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Mardis, E.R. New strategies and emerging technologies for massively parallel sequencing: Applications in medical research. Genome Med. 2009, 1, 40. [Google Scholar] [CrossRef] [PubMed]
- Krawitz, P.M.; Schweiger, M.R.; Rodelsperger, C.; Marcelis, C.; Kolsch, U.; Meisel, C.; Stephani, F.; Kinoshita, T.; Murakami, Y.; Bauer, S.; et al. Identity-by-descent filtering of exome sequence data identifies PIGV mutations in hyperphosphatasia mental retardation syndrome. Nat. Genet. 2010, 42, 827–829. [Google Scholar] [CrossRef] [PubMed]
- Barton, G.J. ALSCRIPT: A tool to format multiple sequence alignments. Protein Eng. 1993, 6, 37–40. [Google Scholar] [CrossRef] [PubMed]
- Milligan, G. G protein-coupled receptor hetero-dimerization: Contribution to pharmacology and function. Br. J. Pharmacol. 2009, 158, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Hendy, G.N.; Percy, M.E.; Bichet, D.G.; Cole, D.E. G protein-coupled receptor mutations and human genetic disease. Methods Mol. Biol. 2014, 1175, 153–187. [Google Scholar] [PubMed]
- Thompson, M.D.; Cole, D.E.; Jose, P.A.; Chidiac, P. G protein-coupled receptor accessory proteins and signalling: Pharmacogenomic insights. Methods Mol Biol. 2014, 1175, 121–152. [Google Scholar] [PubMed]
- Milasta, S.; Evans, N.A.; Ormiston, L.; Wilson, S.; Lefkowitz, R.J.; Milligan, G. The sustainability of interactions between the orexin-1 receptor and beta-arrestin-2 is defined by a single C-terminal cluster of hydroxy amino acids and modulates the kinetics of ERK MAPK regulation. Biochem. J. 2005, 387, 573–584. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, W.C.; Seeber, R.M.; Eidne, K.A.; Pfleger, K.D. Molecular determinants of orexin receptor-arrestin-ubiquitin complex formation. Br. J. Pharmacol. 2014, 171, 364–374. [Google Scholar] [CrossRef] [PubMed]
- Duguay, D.; Belanger-Nelson, E.; Mongrain, V.; Beben, A.; Khatchadourian, A.; Cermakian, N. Dynein light chain Tctex-Type 1 modulates orexin signalling through its interaction with orexin 1 receptor. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- EI Firar, A.; Voisin, T.; Rouyer-Fessard, C.; Ostuni, M.A.; Couvineau, A.; Laburthe, M. Discovery of a functional immunoreceptor tyrosine-based switch motif in a 7-transmembrane-spanning receptor: Role in the orexin receptor OX1R-driven apoptosis. FASEB J. 2009, 23, 4069–4080. [Google Scholar] [CrossRef] [PubMed]
- Dalrymple, M.B.; Jaeger, W.C.; Eidne, K.A.; Pfleger, K.D. Temporal profiling of orexin receptor-arrestin-ubiquitin complexes reveals differences between receptor subtypes. J. Biol. Chem. 2011, 286, 16726–16733. [Google Scholar] [CrossRef] [PubMed]
- Holmqvist, T.; Johansson, L.; Ostman, M.; Ammoun, S.; Akerman, K.E.; Kukkonen, J.P. OX1 orexin receptors couple to adenylyl cyclase regulation via multiple mechanisms. J. Biol. Chem. 2005, 280, 6570–6579. [Google Scholar] [CrossRef] [PubMed]
- Magga, J.; Bart, G.; Oker-Blom, C.; Kukkonen, J.P.; Akerman, K.E.; Nasman, J. Agonist potency differentiates G protein activation and Ca2+ signalling by the orexin receptor type 1. Biochem. Pharmacol. 2006, 71, 827–836. [Google Scholar] [CrossRef] [PubMed]
- Kukkonen, J.P. OX2 orexin/hypocretin receptor signal transduction in recombinant Chinese hamster ovary cells. Cell. Signal. 2016, 28, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Randeva, H.S.; Karteris, E.; Grammatopoulos, D.; Hillhouse, E.W. Expression of orexin-A and functional orexin type 2 receptors in the human adult adrenals: Implications for adrenal function and energy homeostasis. J. Clin. Endocrinol. Metab. 2001, 86, 4808–4813. [Google Scholar] [CrossRef] [PubMed]
- Wess, J. Molecular basis of receptor/G-protein-coupling selectivity. Pharmacol. Ther. 1998, 80, 231–264. [Google Scholar] [CrossRef]
- Jantti, M.H.; Putula, J.; Turunen, P.M.; Nasman, J.; Reijonen, S.; Lindqvist, C.; Kukkonen, J.P. Autocrine endocannabinoid signalling through CB1 receptors potentiates OX1 orexin receptor signalling. Mol. Pharmacol. 2013, 83, 621–632. [Google Scholar] [CrossRef] [PubMed]
- Milligan, G.; Canals, M.; Pediani, J.D.; Ellis, J.; Lopez-Gimenez, J.F. The role of GPCR dimerisation/oligomerisation in receptor signalling. In GPCRs: From Deorphanization to Lead Structure Identification; Springer: Berlin/Heidelberg, Germany, 2006; Available online: https://link.springer.com/chapter/10.1007/2789_2006_007 (accessed on 28 September 2017).
- Thompson, M.D.; Burnham, W.M.; Cole, D.E. The G protein-coupled receptors: Pharmacogenetics and disease. Crit. Rev. Clin. Lab. Sci. 2005, 42, 311–392. [Google Scholar] [CrossRef] [PubMed]
- Feldman, H.J.; Hogue, C.W. A fast method to sample real protein conformational space. Proteins 2000, 39, 112–131. [Google Scholar] [CrossRef]
- Galtier, N.; Gouy, M.; Gautier, C. SEAVIEW and PHYLO_WIN: Two graphic tools for sequence alignment and molecular phylogeny. Comput. Appl. Biosci. 1996, 12, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Bulenger, S.; Marullo, S.; Bouvier, M. Emerging role of homo- and heterodimerization in G-protein-coupled receptor biosynthesis and maturation. Trends Pharmacol. Sci. 2005, 26, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.R.; Ward, R.J.; Pediani, J.D.; Milligan, G. The orexin OX1 receptor exists predominantly as a homodimer in the basal state: Potential regulation of receptor organization by both agonist and antagonist ligands. Biochem. J. 2011, 439, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Ward, R.J.; Pediani, J.D.; Milligan, G. Heteromultimerization of cannabinoid CB(1) receptor and orexin OX(1) receptor generates a unique complex in which both protomers are regulated by orexin A. J. Biol. Chem. 2011, 286, 37414–37428. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Pan, Y.; Zhang, R.; Bai, B.; Chen, J.; Randeva, H.S. Heterodimerization of mouse orexin type 2 receptor variants and the effects on signal transduction. Biochim. Biophys. Acta 2014, 1843, 652–663. [Google Scholar] [CrossRef] [PubMed]
- Jantti, M.H.; Mandrika, I.; Kukkonen, J.P. Human orexin/hypocretin receptors form constitutive homo- and heteromeric complexes with each other and with human CB1 cannabinoid receptors. Biochem. Biophys. Res. Commun. 2014, 445, 486–490. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Storm van’s Gravesande, K.; Galczenski, H.; Burnham, W.M.; Siminovitch, K.A.; Zamel, N.; Slutsky, A.; Drazen, J.M.; George, S.R.; Evans, J.F.; et al. A cysteinyl leukotriene 2 receptor variant is associated with atopy in the population of Tristan da Cunha. Pharmacogenetics 2003, 13, 641–649. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Capra, V.; Takasaki, J.; Maresca, G.; Rovati, G.E.; Slutsky, A.S.; Lilly, C.; Zamel, N.; McIntyre Burnham, W.; Cole, D.E.; et al. A functional G300S variant of the cysteinyl leukotriene 1 receptor is associated with atopy in a Tristan da Cunha isolate. Pharmacogenet. Genom. 2007, 17, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Takasaki, J.; Capra, V.; Rovati, G.E.; Siminovitch, K.A.; Burnham, W.M.; Hudson, T.J.; Bosse, Y.; Cole, D.E. G-protein-coupled receptors and asthma endophenotypes: The cysteinyl leukotriene system in perspective. Mol. Diagn. Ther. 2006, 10, 353–366. [Google Scholar] [CrossRef] [PubMed]
- Capra, V.; Thompson, M.D.; Sala, A.; Cole, D.E.; Folco, G.; Rovati, G.E. Cysteinyl-leukotrienes and their receptors in asthma and other inflammatory diseases: Critical update and emerging trends. Med. Res. Rev. 2007, 27, 469–527. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.D.; Capra, V.; Clunes, M.T.; Rovati, G.E.; Stankova, J.; Maj, M.C.; Duffy, D.L. Cysteinyl leukotrienes pathway genes, atopic asthma and drug response: From population isolates to large genome-wide association studies. Front Pharmacol. 2016, 7, 299. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Blache, D.; Vercoe, P.E.; Adam, C.L.; Blackberry, M.A.; Findlay, P.A.; Eidne, K.A.; Martin, G.B. Expression of orexin receptors in the brain and peripheral tissues of the male sheep. Regul. Pept. 2005, 124, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chen, J.; Bai, B.; Du, H.; Liu, Y.; Liu, H. Heterodimerization of human apelin and kappa opioid receptors: Roles in signal transduction. Cell. Signal. 2012, 24, 991–1001. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhang, R.; Chen, X.; Wang, C.; Cai, X.; Liu, H.; Jiang, Y.; Liu, C.; Bai, B. Heterodimerization of human orexin receptor 1 and kappa opioid receptor promotes protein kinase A/cAMP-response element binding protein signalling via a Galphas-mediated mechanism. Cell. Signal. 2015, 27, 1426–1438. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Acuna-Goycolea, C.; Li, Y.; Cheng, H.M.; Obrietan, K.; van den Pol, A.N. Cannabinoids excite hypothalamic melanin-concentrating hormone but inhibit hypocretin/orexin neurons: Implications for cannabinoid actions on food intake and cognitive arousal. J. Neurosci. 2007, 27, 4870–4881. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Pearsall, E.A.; Hurst, D.P.; Zhang, Y.; Chu, J.; Zhou, Y.; Reggio, P.H.; Loh, H.H.; Law, P.Y. Palmitoylation and membrane cholesterol stabilize mu-opioid receptor homodimerization and G protein coupling. BMC Cell Biol. 2012, 13. [Google Scholar] [CrossRef] [PubMed]
- Shang, Y.; Filizola, M. Opioid receptors: Structural and mechanistic insights into pharmacology and signalling. Eur. J. Pharmacol. 2015, 763, 206–213. [Google Scholar] [CrossRef] [PubMed]
- Haj-Dahmane, S.; Shen, R.Y. The wake-promoting peptide orexin-B inhibits glutamatergic transmission to dorsal raphe nucleus serotonin neurons through retrograde endocannabinoid signalling. J. Neurosci. 2005, 25, 896–905. [Google Scholar] [CrossRef] [PubMed]
- Turunen, P.M.; Jantti, M.H.; Kukkonen, J.P. OX1 orexin/hypocretin receptor signalling through arachidonic acid and endocannabinoid release. Mol. Pharmacol. 2012, 82, 156–167. [Google Scholar] [CrossRef] [PubMed]
- Ho, Y.C.; Lee, H.J.; Tung, L.W.; Liao, Y.Y.; Fu, S.Y.; Teng, S.F.; Liao, H.T.; Mackie, K.; Chiou, L.C. Activation of orexin 1 receptors in the periaqueductal gray of male rats leads to antinociception via retrograde endocannabinoid (2-arachidonoylglycerol)-induced disinhibition. J. Neurosci. 2011, 31, 14600–14610. [Google Scholar] [CrossRef] [PubMed]
- Wager-Miller, J.; Westenbroek, R.; Mackie, K. Dimerization of G protein-coupled receptors: CB1 cannabinoid receptors as an example. Chem. Phys. Lipids 2002, 121, 83–89. [Google Scholar] [CrossRef]
- Kearn, C.S.; Blake-Palmer, K.; Daniel, E.; Mackie, K.; Glass, M. Concurrent stimulation of cannabinoid CB1 and dopamine D2 receptors enhances heterodimer formation: A mechanism for receptor cross-talk? Mol. Pharmacol. 2005, 67, 1697–1704. [Google Scholar] [CrossRef] [PubMed]
- Hern, J.A.; Baig, A.H.; Mashanov, G.I.; Birdsall, B.; Corrie, J.E.; Lazareno, S.; Molloy, J.E.; Birdsall, N.J. Formation and dissociation of M1 muscarinic receptor dimers seen by total internal reflection fluorescence imaging of single molecules. Proc. Natl. Acad. Sci. USA 2010, 107, 2693–2698. [Google Scholar] [CrossRef] [PubMed]
- Pin, J.P.; Kniazeff, J.; Liu, J.; Binet, V.; Goudet, C.; Rondard, P.; Prezeau, L. Allosteric functioning of dimeric class C G-protein-coupled receptors. FEBS J. 2005, 272, 2947–2955. [Google Scholar] [CrossRef] [PubMed]
- Maurel, D.; Comps-Agrar, L.; Brock, C.; Rives, M.L.; Bourrier, E.; Ayoub, M.A.; Bazin, H.; Tinel, N.; Durroux, T.; Prezeau, L.; et al. Cell-surface protein-protein interaction analysis with time-resolved FRET and snap-tag technologies: Application to GPCR oligomerization. Nat. Methods 2008, 5, 561–567. [Google Scholar] [CrossRef] [PubMed]
- Kilpatrick, L.E.; Hill, S.J. The use of fluorescence correlation spectroscopy to characterize the molecular mobility of fluorescently labelled G protein-coupled receptors. Biochem. Soc. Trans. 2016, 44, 624–629. [Google Scholar] [CrossRef] [PubMed]
- Hague, C.; Uberti, M.A.; Chen, Z.; Hall, R.A.; Minneman, K.P. Cell surface expression of α1D-adrenergic receptors is controlled by heterodimerization with α1B-adrenergic receptors. J. Biol. Chem. 2004, 279, 15541–15549. [Google Scholar] [CrossRef] [PubMed]
- Prinster, S.C.; Holmqvist, T.G.; Hall, R.A. α2C-adrenergic receptors exhibit enhanced surface expression and signalling upon association with β2-adrenergic receptors. J. Pharmacol. Exp. Ther. 2006, 318, 974–981. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Randeva, H.S. Genomic organization of mouse orexin receptors: Characterization of two novel tissue-specific splice variants. Mol. Endocrinol. 2004, 18, 2790–2804. [Google Scholar] [CrossRef] [PubMed]
- Cvejic, S.; Devi, L.A. Dimerization of the delta opioid receptor: Implication for a role in receptor internalization. J. Biol. Chem. 1997, 272, 26959–26964. [Google Scholar] [CrossRef] [PubMed]
- Grant, M.; Patel, R.C.; Kumar, U. The role of subtype-specific ligand binding and the C-tail domain in dimer formation of human somatostatin receptors. J. Biol. Chem. 2004, 279, 38636–38643. [Google Scholar] [CrossRef] [PubMed]
- Navarro, G.; Quiroz, C.; Moreno-Delgado, D.; Sierakowiak, A.; McDowell, K.; Moreno, E.; Rea, W.; Cai, N.S.; Aguinaga, D.; Howell, L.A.; et al. Orexin-corticotropin-releasing factor receptor heteromers in the ventral tegmental area as targets for cocaine. J. Neurosci. 2015, 35, 6639–6653. [Google Scholar] [CrossRef] [PubMed]
- Scott, M.M.; Marcus, J.N.; Pettersen, A.; Birnbaum, S.G.; Mochizuki, T.; Scammell, T.E.; Nestler, E.J.; Elmquist, J.K.; Lutter, M. Hcrtr1 and 2 signalling differentially regulates depression-like behaviors. Behav. Brain Res. 2011, 222, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Salomon, R.M.; Ripley, B.; Kennedy, J.S.; Johnson, B.; Schmidt, D.; Zeitzer, J.M.; Nishino, S.; Mignot, E. Diurnal variation of cerebrospinal fluid hypocretin-1 (Orexin-A) levels in control and depressed subjects. Biol. Psychiatry 2003, 54, 96–104. [Google Scholar] [CrossRef]
- Nollet, M.; Gaillard, P.; Tanti, A.; Girault, V.; Belzung, C.; Leman, S. Neurogenesis-independent antidepressant-like effects on behavior and stress axis response of a dual orexin receptor antagonist in a rodent model of depression. Neuropsychopharmacology 2012, 37, 2210–2221. [Google Scholar] [CrossRef] [PubMed]
- Rogala, B.; Li, Y.; Li, S.; Chen, X.; Kirouac, G.J. Effects of a post-shock injection of the kappa opioid receptor antagonist norbinaltorphimine (norBNI) on fear and anxiety in rats. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Schwarzer, C. 30 years of dynorphins—new insights on their functions in neuropsychiatric diseases. Pharmacol. Ther. 2009, 123, 353–370. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.D.; McDonald, P.H. The orexin 1 receptor modulates kappa opioid receptor function via a JNK-dependent mechanism. Cell. Signal. 2015, 27, 1449–1456. [Google Scholar] [CrossRef] [PubMed]
- Baimel, C.; Lau, B.K.; Qiao, M.; Borgland, S.L. Projection-target-defined effects of orexin and dynorphin on VTA dopamine neurons. Cell Res. 2017, 18, 1346–1355. [Google Scholar] [CrossRef] [PubMed]
- Bruchas, M.R.; Chavkin, C. Kinase cascades and ligand-directed signalling at the kappa opioid receptor. Psychopharmacology 2010, 210, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Nocjar, C.; Zhang, J.; Feng, P.; Panksepp, J. The social defeat animal model of depression shows diminished levels of orexin in mesocortical regions of the dopamine system, and of dynorphin and orexin in the hypothalamus. Neuroscience 2012, 218, 138–153. [Google Scholar] [CrossRef] [PubMed]
- Rios, C.; Gomes, I.; Devi, L.A. Mu opioid and CB1 cannabinoid receptor interactions: Reciprocal inhibition of receptor signalling and neuritogenesis. Br. J. Pharmacol. 2006, 148, 387–395. [Google Scholar] [CrossRef] [PubMed]
- Carriba, P.; Ortiz, O.; Patkar, K.; Justinova, Z.; Stroik, J.; Themann, A.; Muller, C.; Woods, A.S.; Hope, B.T.; Ciruela, F.; et al. Striatal adenosine A2A and cannabinoid CB1 receptors form functional heteromeric complexes that mediate the motor effects of cannabinoids. Neuropsychopharmacology 2007, 32, 2249–2259. [Google Scholar] [CrossRef] [PubMed]
- Rozenfeld, R.; Gupta, A.; Gagnidze, K.; Lim, M.P.; Gomes, I.; Lee-Ramos, D.; Nieto, N.; Devi, L.A. AT1R-CB1R heteromerization reveals a new mechanism for the pathogenic properties of angiotensin II. EMBO J. 2011, 30, 2350–2363. [Google Scholar] [CrossRef] [PubMed]
- Kano, M.; Ohno-Shosaku, T.; Hashimotodani, Y.; Uchigashima, M.; Watanabe, M. Endocannabinoid-mediated control of synaptic transmission. Physiol. Rev. 2009, 89, 309–380. [Google Scholar] [CrossRef] [PubMed]
- Ellis, J.; Pediani, J.D.; Canals, M.; Milasta, S.; Milligan, G. Orexin-1 receptor-cannabinoid CB1 receptor heterodimerization results in both ligand-dependent and -independent coordinated alterations of receptor localization and function. J. Biol. Chem. 2006, 281, 38812–38824. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Gilliam, A.; Maitra, R.; Damaj, M.I.; Tajuba, J.M.; Seltzman, H.H.; Thomas, B.F. Synthesis and biological evaluation of bivalent ligands for the cannabinoid 1 receptor. J. Med. Chem. 2010, 53, 7048–7060. [Google Scholar] [CrossRef] [PubMed]
- Le Naour, M.; Akgun, E.; Yekkirala, A.; Lunzer, M.M.; Powers, M.D.; Kalyuzhny, A.E.; Portoghese, P.S. Bivalent ligands that target mu opioid (MOP) and cannabinoid1 (CB1) receptors are potent analgesics devoid of tolerance. J. Med. Chem. 2013, 56, 5505–5513. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Wang, X.Q.; Chen, Y.N.; Yang, N.; Lang, S.Y.; Zuo, P.P.; Zhang, J.T.; Li, R.S. Changes and overlapping distribution in the expression of CB1/OX1-GPCRs in rat hippocampus by kainic acid-induced status epilepticus. Brain Res. 2015, 1597, 14–27. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, L.S.; DeLorenzo, R.J. Acetaminophen inhibits status epilepticus in cultured hippocampal neurons. Neuroreport 2011, 22, 15–18. [Google Scholar] [CrossRef] [PubMed]
- Pugin, D.; Foreman, B.; De Marchis, G.M.; Fernandez, A.; Schmidt, J.M.; Czeisler, B.M.; Mayer, S.A.; Agarwal, S.; Lesch, C.; Lantigua, H.; et al. Is pentobarbital safe and efficacious in the treatment of super-refractory status epilepticus: A cohort study. Crit. Care 2014, 18, R103. [Google Scholar] [CrossRef] [PubMed]
- Kogan, N.M.; Mechoulam, R. Cannabinoids in health and disease. Dialogues Clin. Neurosci. 2007, 9, 413–430. [Google Scholar] [PubMed]
- Maejima, T.; Ohno-Shosaku, T.; Kano, M. Endogenous cannabinoid as a retrograde messenger from depolarized postsynaptic neurons to presynaptic terminals. Neurosci. Res. 2001, 40, 205–210. [Google Scholar] [CrossRef]
- Ludanyi, A.; Eross, L.; Czirjak, S.; Vajda, J.; Halasz, P.; Watanabe, M.; Palkovits, M.; Magloczky, Z.; Freund, T.F.; Katona, I. Downregulation of the CB1 cannabinoid receptor and related molecular elements of the endocannabinoid system in epileptic human hippocampus. J. Neurosci. 2008, 28, 2976–2990. [Google Scholar] [CrossRef] [PubMed]
- Falenski, K.W.; Carter, D.S.; Harrison, A.J.; Martin, B.R.; Blair, R.E.; DeLorenzo, R.J. Temporal characterization of changes in hippocampal cannabinoid CB1 receptor expression following pilocarpine-induced status epilepticus. Brain Res. 2009, 1262, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Bojnik, E.; Turunc, E.; Armagan, G.; Kanit, L.; Benyhe, S.; Yalcin, A.; Borsodi, A. Changes in the cannabinoid (CB1) receptor expression level and G-protein activation in kainic acid induced seizures. Epilepsy Res. 2012, 99, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Khaspekov, L.G.; Brenz Verca, M.S.; Frumkina, L.E.; Hermann, H.; Marsicano, G.; Lutz, B. Involvement of brain-derived neurotrophic factor in cannabinoid receptor-dependent protection against excitotoxicity. Eur. J. Neurosci. 2004, 19, 1691–1698. [Google Scholar] [CrossRef] [PubMed]
- Cristino, L.; Busetto, G.; Imperatore, R.; Ferrandino, I.; Palomba, L.; Silvestri, C.; Petrosino, S.; Orlando, P.; Bentivoglio, M.; Mackie, K.; et al. Obesity-driven synaptic remodeling affects endocannabinoid control of orexinergic neurons. Proc. Natl. Acad. Sci. USA 2013, 110, E2229–2238. [Google Scholar] [CrossRef] [PubMed]
- Doreulee, N.; Alania, M.; Vashalomidze, G.; Skhirtladze, E.; Kapanadze, T. Orexinergic system and pathophysiology of epilepsy. Georgian Med. News 2010, 188, 74–79. [Google Scholar]

| OX1 aa | Location | SNP | Numbering/ Peyron | Numbering/ 0lafsdottir | Frequency | Findings | Ref. |
|---|---|---|---|---|---|---|---|
| 167 Gly/Ser | TM4 | rs44603792 | - | 652 G/A | 0002 | - | [9] |
| 265 Leu/Met | IC3 | rs41501244 | 793 C/A | - | 0.001 | - | [10] |
| 279 Arg/Gln | IC3 | rs7516785 | - | 989 G/A | 0.008 | - | [9] |
| 280 Gly/Ala 281 Arg/His | IC3 IC3 | NP_001516 rs41439244 | 842 GIA | 995 G/A | 0.008 | - | [9,10] |
| 408 Ile/Val | C-term | rs227 I933 | 1222 GIA | 1375 G/A | 0.56 | - | [9,10] |
| - | - | - | - | - | - | 1.4 X migraine | [11] |
| - | - | - | - | - | - | 1.6 X mood disorders | [12] |
| - | - | - | - | - | - | Polydipsia- hyponatremia in schizophrenia | [13,14] |
| - | - | - | - | - | - | Panic not linked | [15] |
| OX2 aa | Location | SNP | Numbering/ Peyron | Numbering/ Olafsdottir | Frequency | Findings | Ref. |
|---|---|---|---|---|---|---|---|
| 10 Pro/Ser | N-term | rs41271310 | 28 C/T | 352 C/T 0.003 | 0.003 | Not linked | [5,9,10] |
| - | - | - | - | - | - | with narcolepsy | - |
| 11 Pro/Thr | N-term | rs4127312 | 31 C/A | 355 C/A | 0.005 | Not linked | [5,9,10] |
| - | - | - | - | - | - | with narcolepsy | - |
| 193 Cys/Ser | TM4 | rs41381449 | 577 T/A | - | 0.002 | - | [10] |
| 293 Ile/Val | IC3 | rs74720047 | 1201 G/A | 0.002 | - | [9] | |
| 308 Ile/Val | TM6 | rs26533 | 922 G/A | 1246 G/A | 0.84 | - | [9,10] |
| - | - | - | - | - | - | Ile assoc. with panic | [15] |
| - | - | - | - | - | - | Migraine not associated | [16] |
| - | - | - | - | - | - | Some association | [17,18,19] |
| - | - | - | - | - | - | with CH | - |
| - | - | - | - | - | - | No effect on triptan response | [20] |
| 401Thr/lle | C-term | rs41321149 | 1202 C/T | 0.00012 | - | - | [10] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thompson, M.D.; Sakurai, T.; Rainero, I.; Maj, M.C.; Kukkonen, J.P. Orexin Receptor Multimerization versus Functional Interactions: Neuropharmacological Implications for Opioid and Cannabinoid Signalling and Pharmacogenetics. Pharmaceuticals 2017, 10, 79. https://doi.org/10.3390/ph10040079
Thompson MD, Sakurai T, Rainero I, Maj MC, Kukkonen JP. Orexin Receptor Multimerization versus Functional Interactions: Neuropharmacological Implications for Opioid and Cannabinoid Signalling and Pharmacogenetics. Pharmaceuticals. 2017; 10(4):79. https://doi.org/10.3390/ph10040079
Chicago/Turabian StyleThompson, Miles D., Takeshi Sakurai, Innocenzo Rainero, Mary C. Maj, and Jyrki P. Kukkonen. 2017. "Orexin Receptor Multimerization versus Functional Interactions: Neuropharmacological Implications for Opioid and Cannabinoid Signalling and Pharmacogenetics" Pharmaceuticals 10, no. 4: 79. https://doi.org/10.3390/ph10040079
APA StyleThompson, M. D., Sakurai, T., Rainero, I., Maj, M. C., & Kukkonen, J. P. (2017). Orexin Receptor Multimerization versus Functional Interactions: Neuropharmacological Implications for Opioid and Cannabinoid Signalling and Pharmacogenetics. Pharmaceuticals, 10(4), 79. https://doi.org/10.3390/ph10040079