1. Introduction
Cotton plays an irreplaceable part in the livelihood of the general population. Xinjiang is China’s largest major producer of long-staple cotton. However, due to the low rainfall and strong light, drip irrigation under the film is often adopted to boost yield, which is prone to mixing with impurities such as plastic film during mechanical picking. In the spinning and weaving processes, the residual film combined with seed cotton can result in a significant number of flaws, which can impact the strength and coloring effect of the yarn and lead to financial losses for the textile sector [
1].
The existing mainstream cotton film removal processes include Mechanical separation, Electrostatic separation, and Optical color separation. Whitelock D. P. et al. investigated major impurity removal equipment in the US cotton industry. A rotating spiked cylinder was utilized to eliminate significant impurities from the seed cotton. These impurities were subsequently gathered in a separate box by means of a grid strip or screen. [
2]. Zhang et al. used computational fluid dynamics (CFD) to model the electrostatic separation of mechanical cotton harvesting and residual plastic film by flying the experimental sample into an electric field at different speeds and applying different electric field forces [
3]. With the increasing prevalence of machine vision, optical color separation has become a popular method for the intelligent classification of agricultural products. In a study conducted by Li et al., a machine vision system was utilized to gather information on the color, shape, and texture of foreign fibers in cotton. The resulting data were adopted to achieve a classification accuracy of 92.34% through multi-class support vector machine (MSVM) [
4].
However, mechanical classification is challenging in the aspect of assuring accuracy and small-size film classification. Electrostatic separation becomes unstable for long-term work because of environmental conditions. Optical color selection relies on color and form characteristics, making it challenging to effectively classify film which is colorless, transparent, or irregularly shaped. Hence, it is imperative to investigate a dependable technique for identifying transparent films in seed cotton.
Hyperspectral imaging combines advanced knowledge from multiple disciplines to achieve a perfect fusion of traditional two-dimensional imaging techniques and spectroscopy. Guo and Ma described the linear relationship between spectra and data by the partial least squares (PLS) method, which could realize the analysis of adulterated rice and the prediction of pork meat fatty acids [
5,
6]. Zhang, Jiang, et al. employed the support vector machine (SVM) in combination with shortwave infrared hyperspectral techniques for cotton foreign matter classification, which significantly improved the detection rate of plastic films in cotton compared to conventional methods [
7,
8,
9].
The above literature has yielded promising results. However, extracting features from hyperspectral images requires manual intervention and has limitations in feature mining. In addition, manual feature extraction of hyperspectral images requires considerable expertise and has subjectivity in feature mining and selection. Therefore, it is highly significative for hyperspectral image features to probe an automatic feature extraction method.
Deep learning is an advanced technology applied to image processing. It has the capability to automatically detect and analyze complex information, which helps to extract deeper features. The use of hyperspectral data greatly enhances the accuracy and efficiency of image recognition [
10,
11]. However, it is important to note that hyperspectral data can be affected by elevated latitudes and severe information redundancy issues. To efficiently extract feature information to support the training of deep learning models, data dimensionality reduction is commonly applied to improve the data processing speed [
12]. Jia et al. employed a method for dimensionality reduction of hyperspectral images by flexible Gabor-based superpixel-level unsupervised linear discriminant analysis (LDA), which reduced a large amount of flexible Gabor (FG) features and increased the peculiarity of image features [
13]. Kang et al. proposed a method based on PCA-EPFs for hyperspectral image (HSI) classification, which used principal component analysis (PCA) to reduce the dimension of the superimposing Edge-preserving features (EPFs). The literature not only represented the EPFs in the mean square sense but also highlighted the divisibility of pixels in EPFs [
14]. To reduce the dimension of hyperspectral remote sensing images, Daniela Lupu et al. established an independent component analysis (ICA) method based on a stochastic higher-order Taylor approximation-based algorithm, which could identify local maxima and facilitate minibatching [
15]. The previous researchers have utilized LDA, PCA, and ICA techniques for reducing the dimensionality of hyperspectral data. The experimental results have demonstrated excellent outcomes, effectively enhancing the efficiency of image processing.
Convolutional neural network (CNN) is the most frequently employed deep learning model that performs excellent classification effect in feature extraction of hyperspectral data; it can be used to solve the problem of plastic film in seed cotton [
16,
17]. LeNet, AlexNet, and VGGNet are frequently employed neural network models in CNN which achieve high classification and recognition accuracy with great fusion with hyperspectral images. Hüseyin Fırat et al. proposed a method to effectively classify hyperspectral remote sensing images (HRSIs) based on PCA dimension reduction and LeNet-5 of the 3D-CNN model. The results showed that a 100% recognition and classification effect was obtained in all experimental data [
18]. Jiang et al. obtained hyperspectral images of different types of pesticide residues and used the fusion of the AlexNet-CNN deep learning network to detect post-harvest pesticide residues in apples. The test results showed that when the number of training epochs was 10, the detection accuracy was 99.09% [
19]. Zhao et al. recorded the waterlogging of cotton after seeding with hyperspectral images. Based on the comparison experiment of GoogLeNet Inception-v3 (GLNI-v3) and VGG-16 conducted by CNN, the classification accuracy of VGG-16 was 97.00% higher than that of GLNI-v3, and the method could provide theoretical support for the evaluation of cotton loss after waterlogging [
20]. The aforementioned literature demonstrates that the CNN-based models (LeNet, AlexNet, VGGNet) mentioned above exhibit strong generalization and adaptive capabilities in processing hyperspectral images, resulting in effective application outcomes.
The combination of hyperspectral imaging and CNN techniques is commonly applied to the classification of remote-sensing images. However, there have been few reports of methods to identify the residual film in seed cotton. The academic paper presents a novel approach for removing film in seed cotton, which combines hyperspectral images and deep learning algorithm. The innovations are listed as follows:
(1) The study establishes an optimal method for dimensionality reduction of hyperspectral data, which can reduce redundant hyperspectral characteristic information and reduce the time and cost of subsequent neural network training.
(2) The study integrates hyperspectral imaging technology and deep learning algorithm to obtain the optimal AlexNet-PCA-12 model which can effectively remove the colorless and transparent film in seed cotton in the practical application.
The remainder of the paper is structured as follows.
Section 2 describes the hyperspectral sorting system, the theory of dimensionality reduction and CNN.
Section 3 illustrates the discussion of the results of dimension reduction and CNN experiments. Conclusions and viewpoints are provided in
Section 4.
3. Results and Discussion
3.1. Design of Intelligent Recognition Algorithm for Film in Seed Cotton
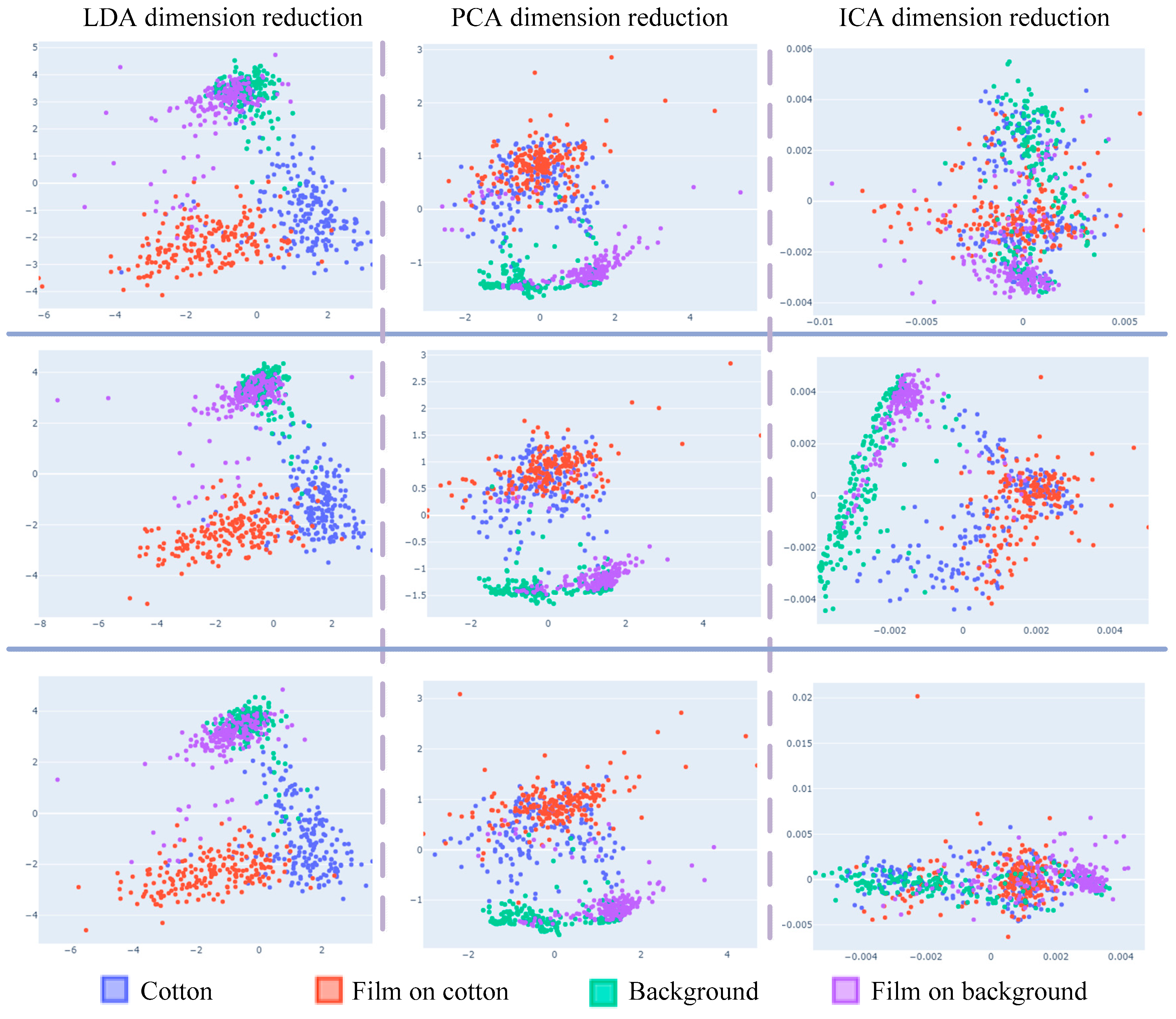
To verify the generalization of the dimensional reduction, scatter plots are presented in
Figure 4. The plots depict the application of different dimensional reduction methods on the same hyperspectral data from the three-dimensional reduction experiments. The scatter plots of dimensionality reduction for different batches of the same data under the same experimental conditions can be concluded as follows:
(1) Considering only the first two samples, LDA data have obvious clustering and separability, but LDA data cannot classify sample “background” accurately.
(2) ICA data classified the four types of samples differently in different batches, so it is not general to data from different batches of dimensionality reduction, and the trained model cannot achieve ideal results on the test.
(3) Considering only the first two samples, PCA dimension reduction has distinct aggregation and separability on “background” and “film on background”, while the data coincidence of the two samples “cotton” and “film on cotton” has no classification.
(4) The result shows that LDA has outstanding classification results with a dimensionality reduction of two for hyperspectral data. However, LDA can only reduce the data to three dimensions. Therefore, when the computer performance is satisfied, PCA obtains higher recognition accuracy when it is used to retain more dimensions.
3.2. CNN Model Training
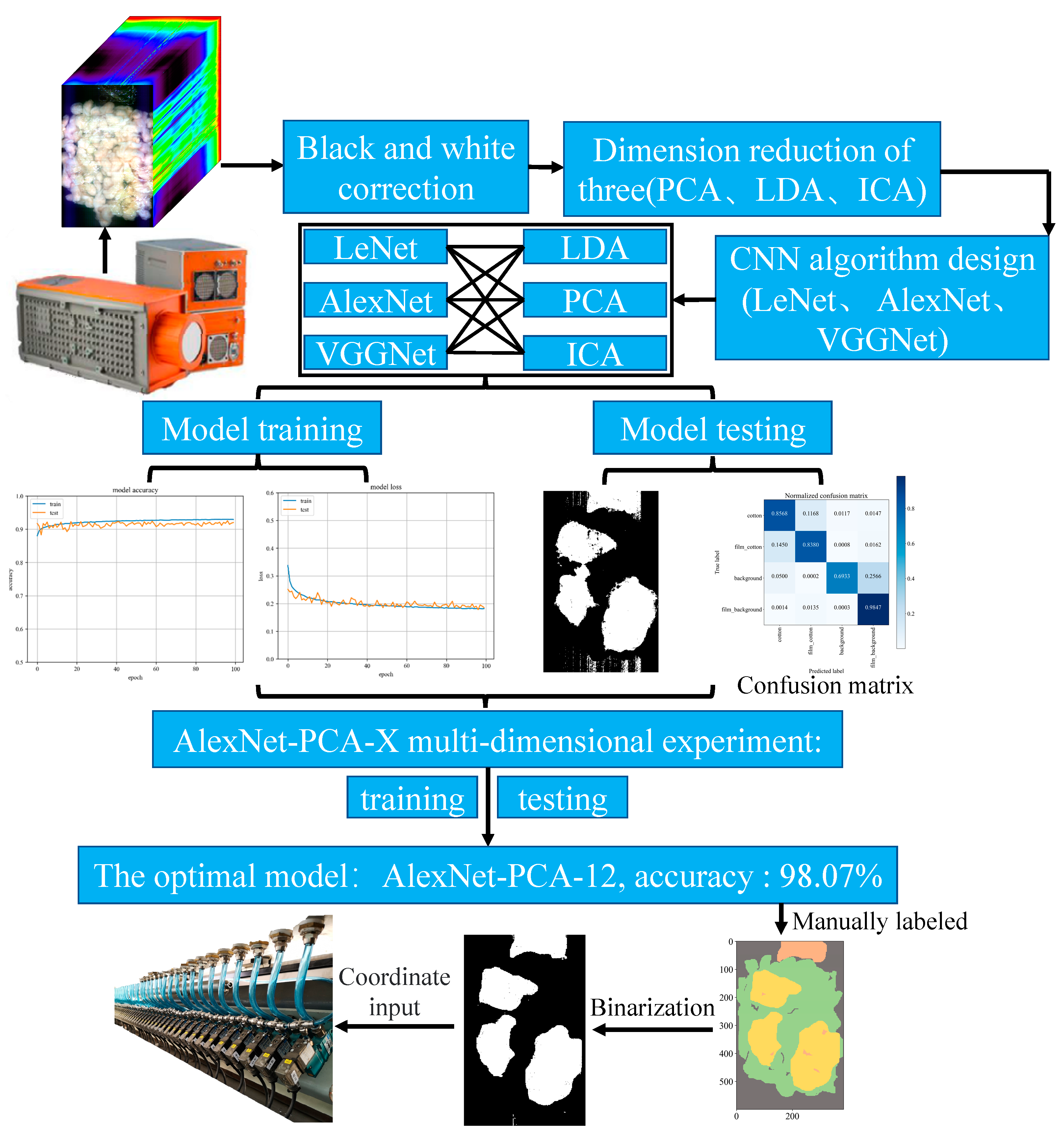
Comparing three different dimension reduction methods (LDA, PCA, and ICA) after the hyperspectral data to a three-dimensional effect, three different structures are adopted CNN (LeNet, AlexNet, and VGGNet) for training and testing accuracy.
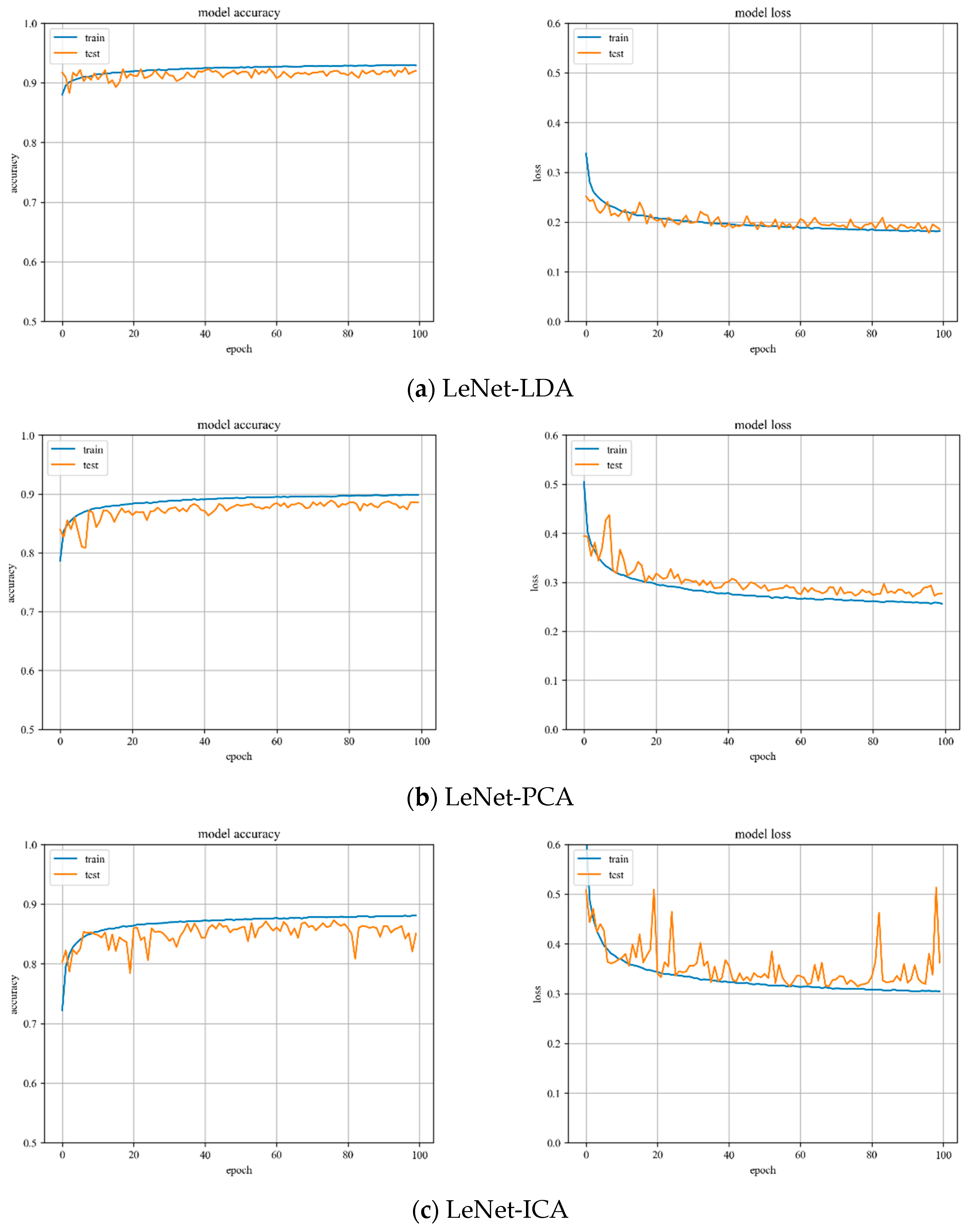
LeNet model training. Variations of training and testing accuracy of LeNet with the number of training epochs, training and testing loss curves are shown in
Figure 5.
In
Figure 5a, LDA recognition accuracy on the test set is about 92%, and the loss value is 0.15~0.2. In
Figure 5b, PCA recognition accuracy on the test set is about 89%, and the loss value is 0.25~0.3. In
Figure 5c, ICA recognition accuracy on the test set is about 85%, and the loss value is 0.3~0.35.
Despite the fact that LDA and PCA are relatively stable to changes throughout the training phase, PCA slightly underperforms the LeNet model with LDA hyperspectral data reduction. However, the LeNet model with ICA hyperspectral data reduction has the worst stability of the three.
AlexNet model training. Variations of training and testing accuracy of AlexNet with the number of training epochs, training and testing loss curves are shown in
Figure 6.
In
Figure 6a, LDA recognition accuracy on the test set is about 93%, and the loss value is 0.15~0.2. In
Figure 6b, PCA recognition accuracy on the test set is about 90%, and the loss value is 0.25~0.3. In
Figure 6c, ICA recognition accuracy on the test set is about 88%, and the loss value is 0.3~0.35.
Despite the fact that LDA and PCA are relatively stable to changes throughout the training phase, PCA slightly underperforms the AlexNet model with LDA hyperspectral data reduction. However, the AlexNet model with ICA hyperspectral data reduction has the worst stability of the three.
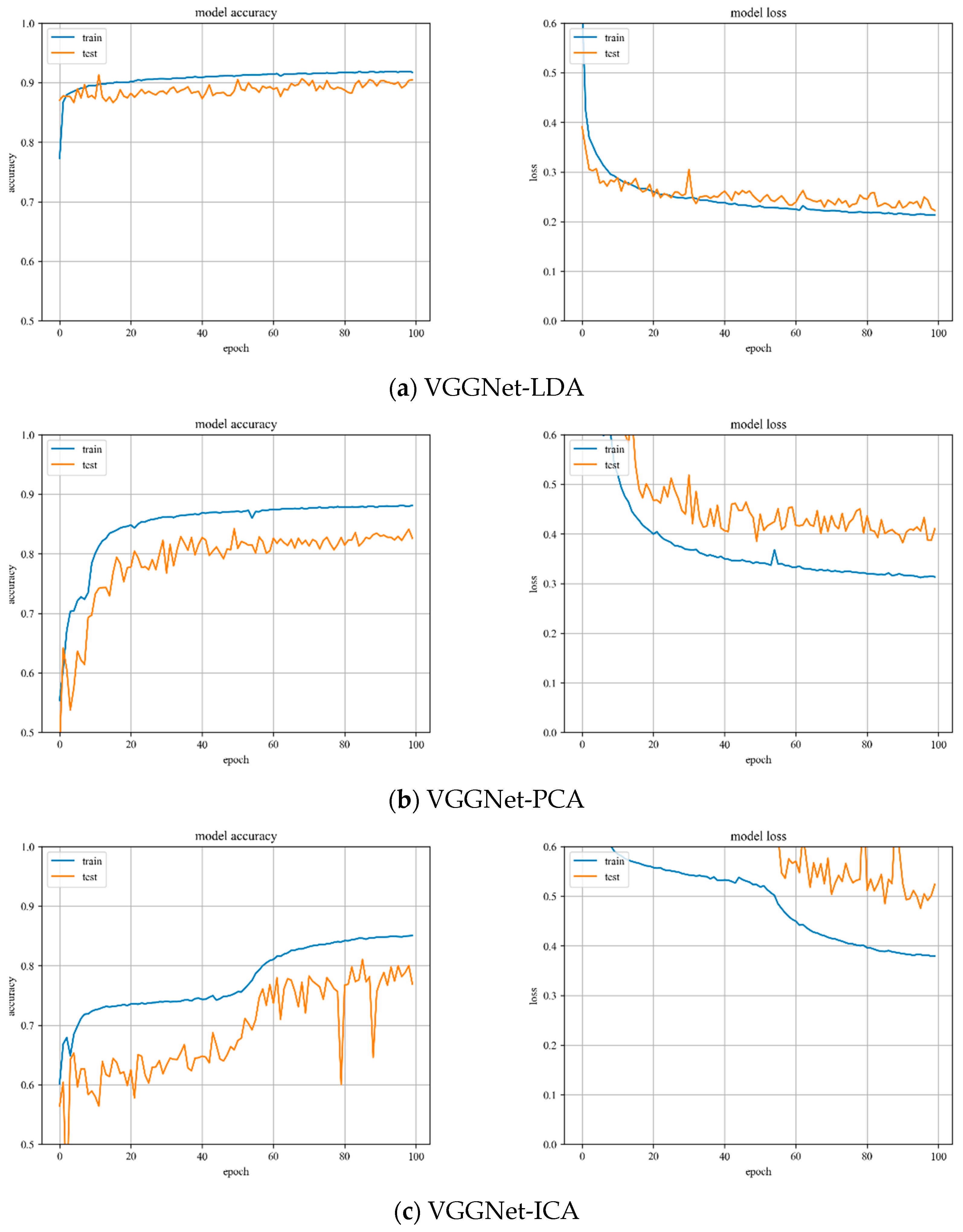
VGGNet model training. Variation of training and testing accuracy of VGGNet with the number of training epochs, training and testing loss curves are shown in
Figure 7.
In
Figure 7a, LDA recognition accuracy on the test set is about 90%, and the loss value is 0.2~0.25. The LDA model is relatively stable to changes throughout the training process and has excellent model stability.
In
Figure 7b, PCA recognition accuracy on the test set is about 84%, and the loss value is about 0.4. In
Figure 7c, ICA recognition accuracy on the test set is about 80%, and the loss value fluctuates widely. Both models have minor stability during the training phase. The ICA results are inferior than the VGGNet model with PCA hyperspectral data reduction.
3.3. CNN Model Testing
The confusion matrix for the test samples for the different algorithmic models are illustrated in
Table 4,
Table 5 and
Table 6. It can be seen that 1 is the cotton, 2 represents the film on cotton, 3 indicates the background, and 4 denotes the film on background, the diagonal expresses the probability of correct classification. The experimental data analysis is as follows:
(1) Using LDA and PCA dimensionality reduction hyperspectral data, it can be determined that the three kinds of CNN models have higher recognition accuracy for test samples. However, there are some errors in the classification of film samples on cotton and film samples on background, which is consistent with the conclusion of the scatter plots above.
(2) Since the hyperspectral data for ICA dimension reduction is not the same batch as the data during training, the extracted dimension information is unstable and the recognition effect is confused. Therefore, it cannot be applied to hyperspectral image recognition, which is consistent with the conclusion of the scatter plots above.
The Overall Accuracy (OA) of the test samples is illustrated in
Table 7, representing the percentage of all samples that are accurately predicted. The results can be summarized as follows:
(1) When the hyperspectral data are reduced to three dimensions, the average OA of LDA is 91.68%, while PCA has an average OA of 87.08%; on the other hand, ICA has a lower average OA of 40.35%. Based on these results, it can be concluded that LDA demonstrates superior performance in terms of dimensionality reduction.
(2) The data in the table show that the CNN-based AlexNet model can achieve excellent recognition effects when the data are dimensionally reduced.
(3) When the dimension reduction of ICA is 3, it exhibits poor performance in terms of average OA compared to the other two dimensionality reduction methods. However, PCA can retain more dimension information to improve the recognition accuracy, which has more potential in practical applications.
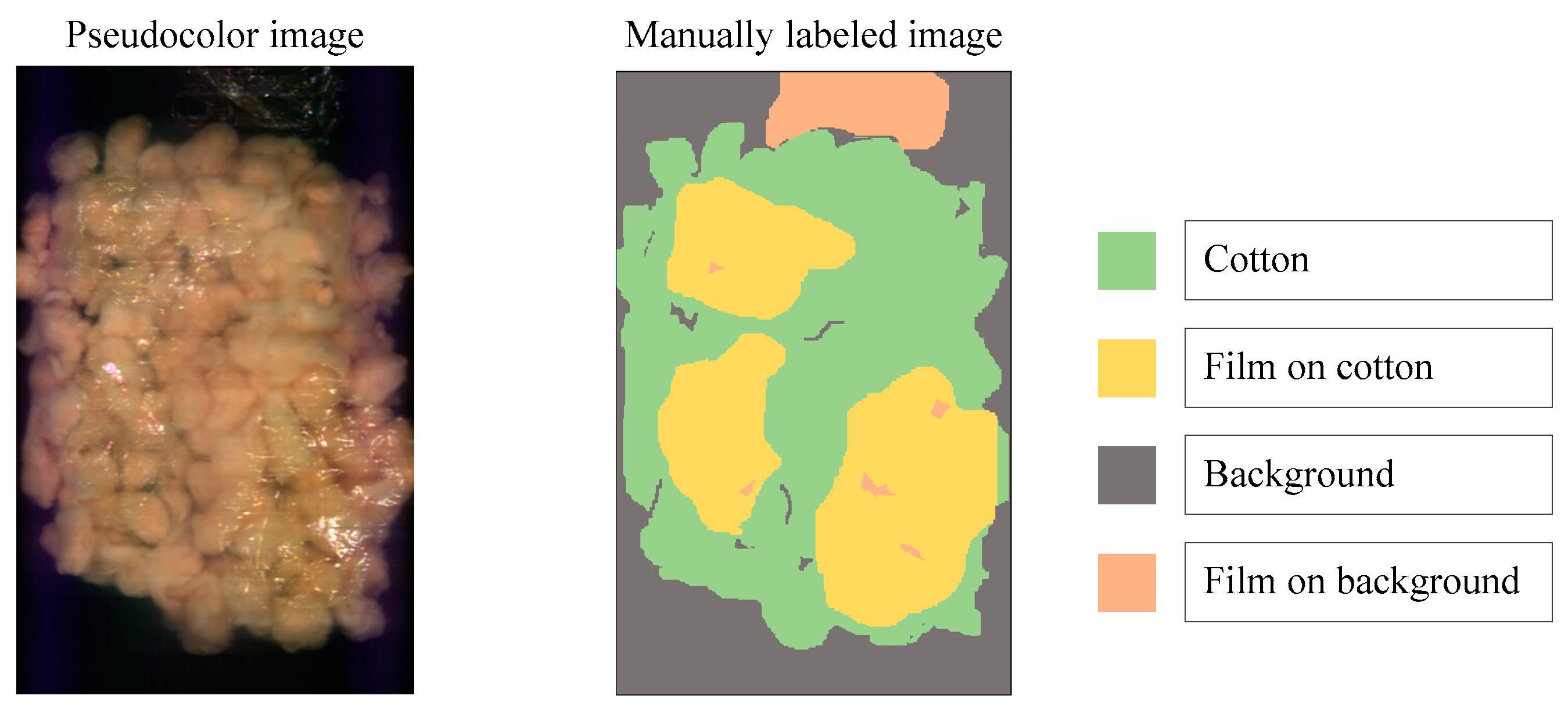
To distinguish the classification effects more intuitively, three bands are selected to display the hyperspectral data as pseudocolor images. Additionally, the spectral toolkit is used in the model tests to plot the predictions in the form of a two-dimensional image. The pseudocolor and manually labeled images are shown in
Figure 8.
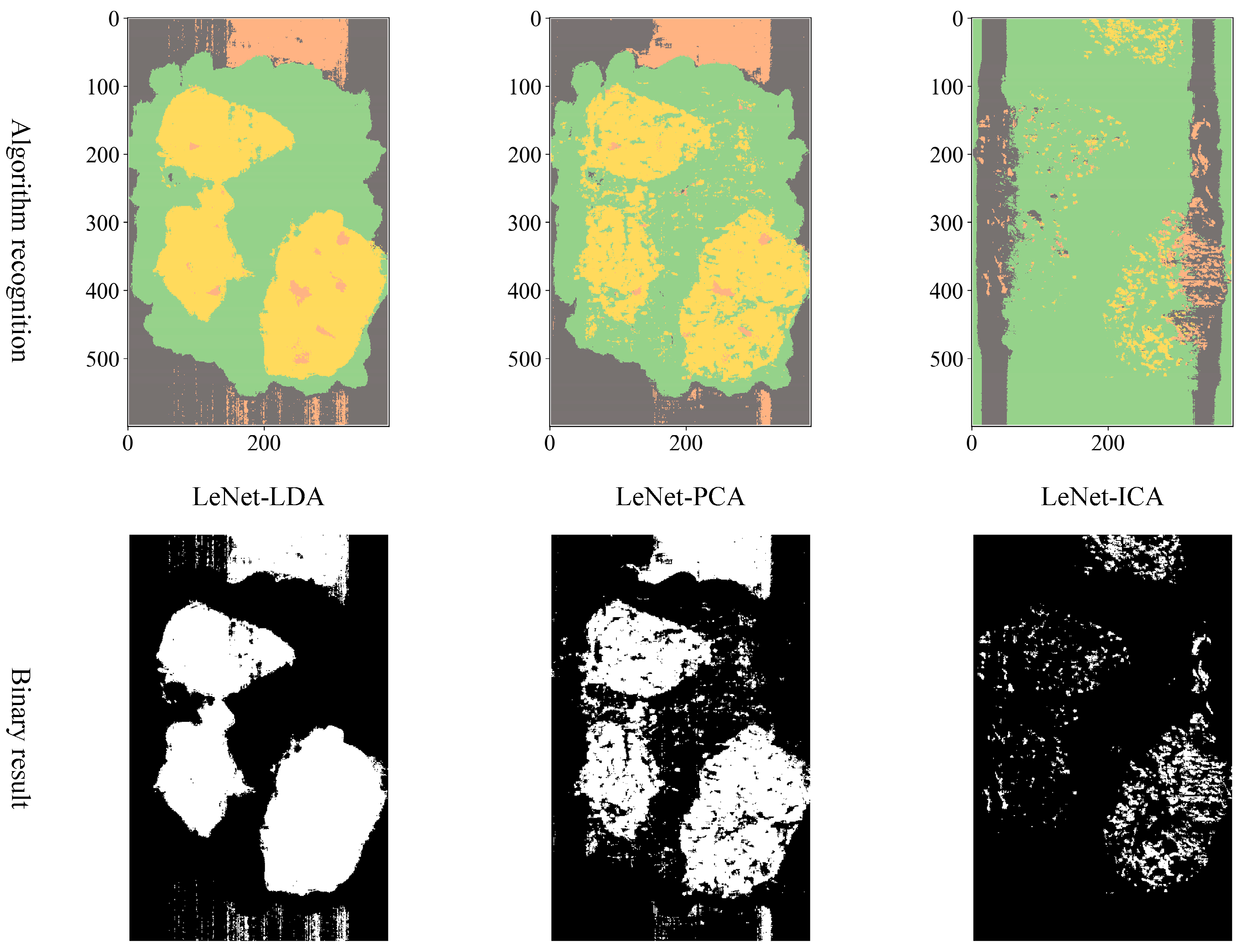
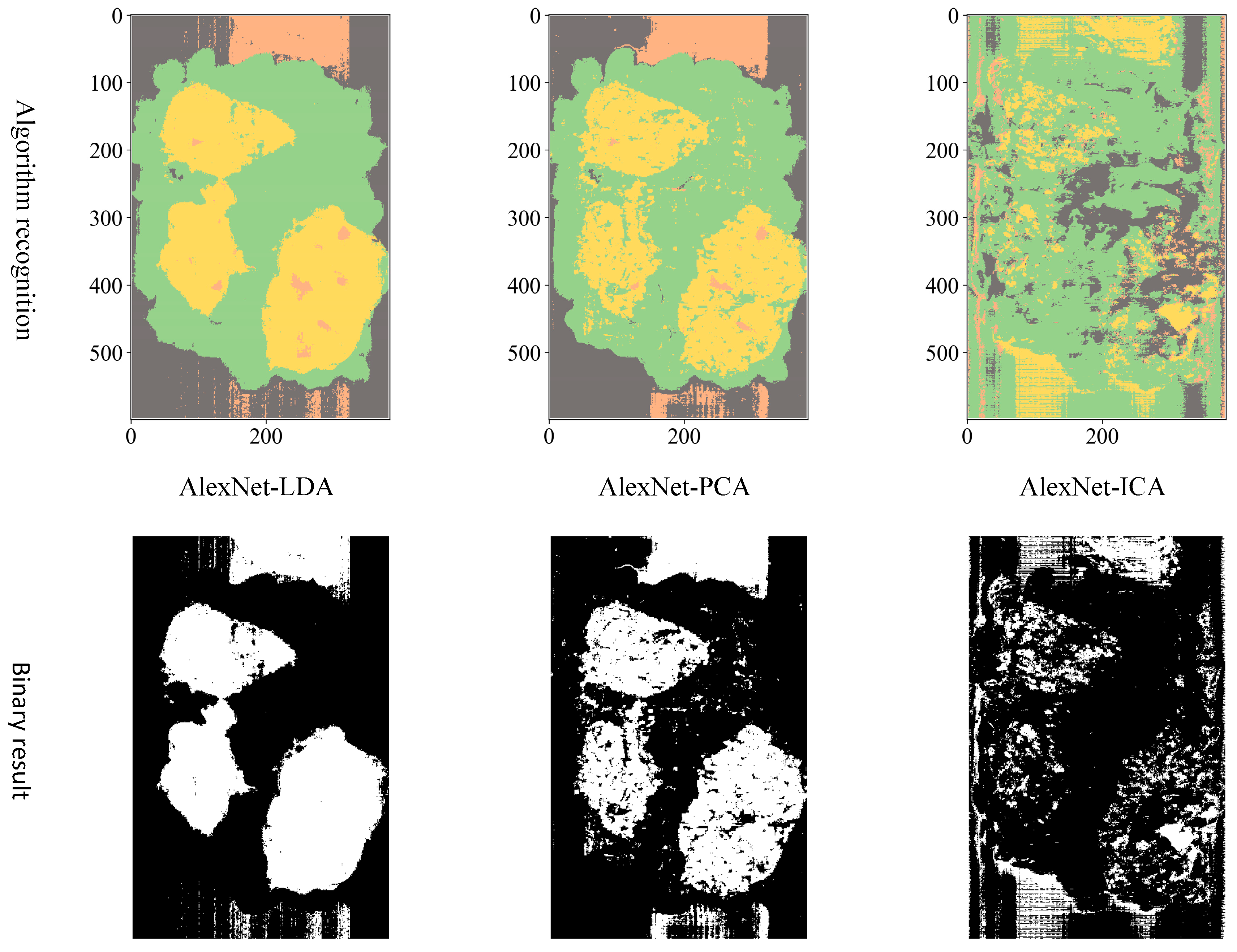
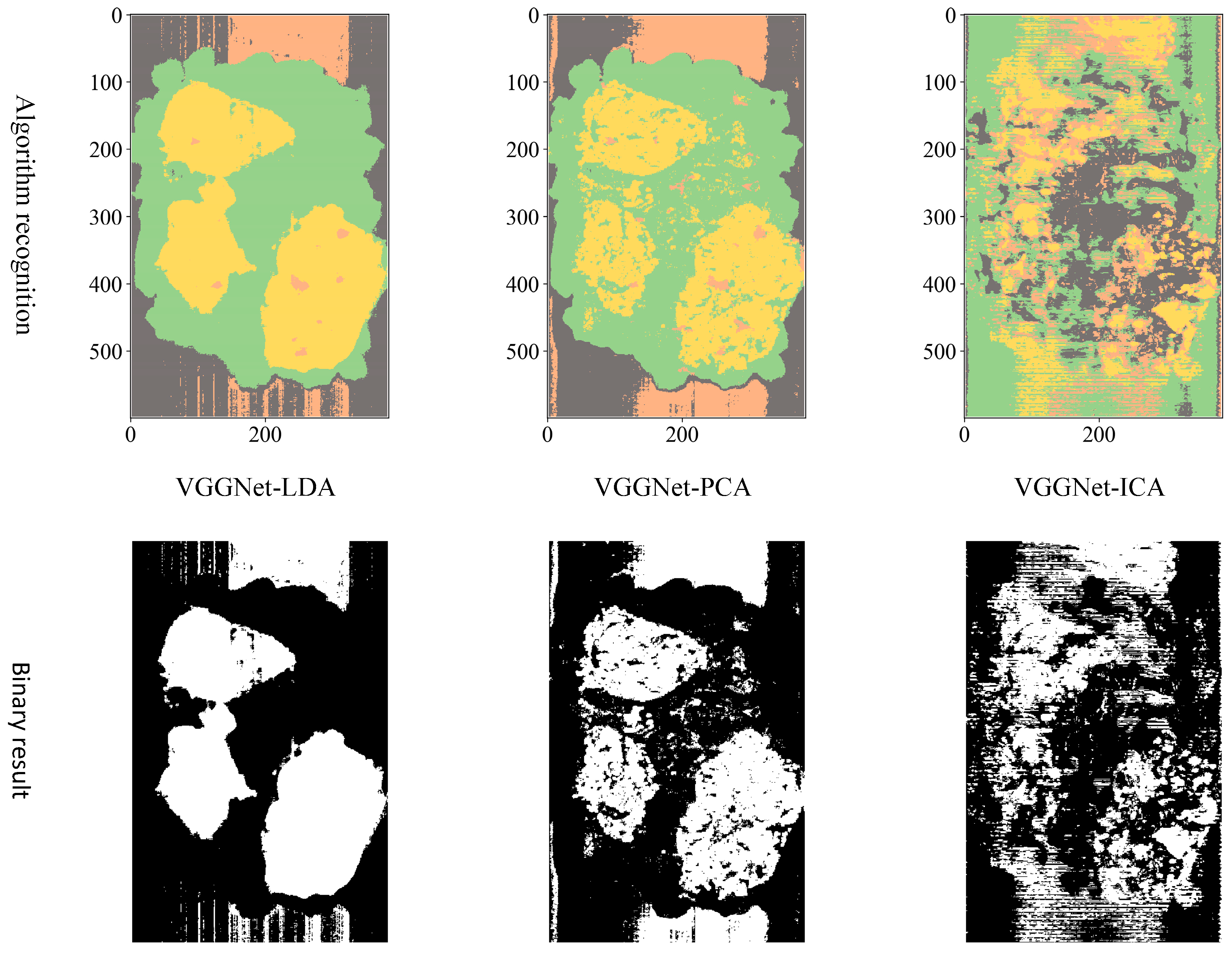
Considering the actual sorting system only needed to locate the spatial coordinate position of the film, the classification results are combined from four categories into two categories: “film on cotton” and “film on background” are classified as film, and “cotton” and “background” are classified as non-film. The binarized images are shown in
Figure 9,
Figure 10 and
Figure 11. It can be seen that the combination of the AlexNet neural network structure and the LDA algorithm indicate the best recognition results, while the VGGNet neural network structure and the ICA algorithm denote the worst recognition results.
Regarding the reduction to three dimensions, the above experiments validate the classification effect of different dimensionality reduction methods on hyperspectral data. The results show that LDA achieves the highest performance in terms of aggregation and separability of features preserved by dimensionality reduction of hyperspectral data. With limited device conditions for hyperspectral images, it is advisable to opt for LDA dimension reduction. However, due to the limitations of the LDA algorithm, the data can only be reduced to three dimensions. Therefore, when the computer performance meets the requirements, PCA achieves higher recognition accuracy when more dimensions are retained. In summary, the AlexNet-PCA multi-dimensional algorithm is experimented with to obtain the highest recognition accuracy for seed cotton mixed with the film.
3.4. AlexNet-PCA Multi-Dimensional Algorithm Experiment
3.4.1. AlexNet-PCA Model Training
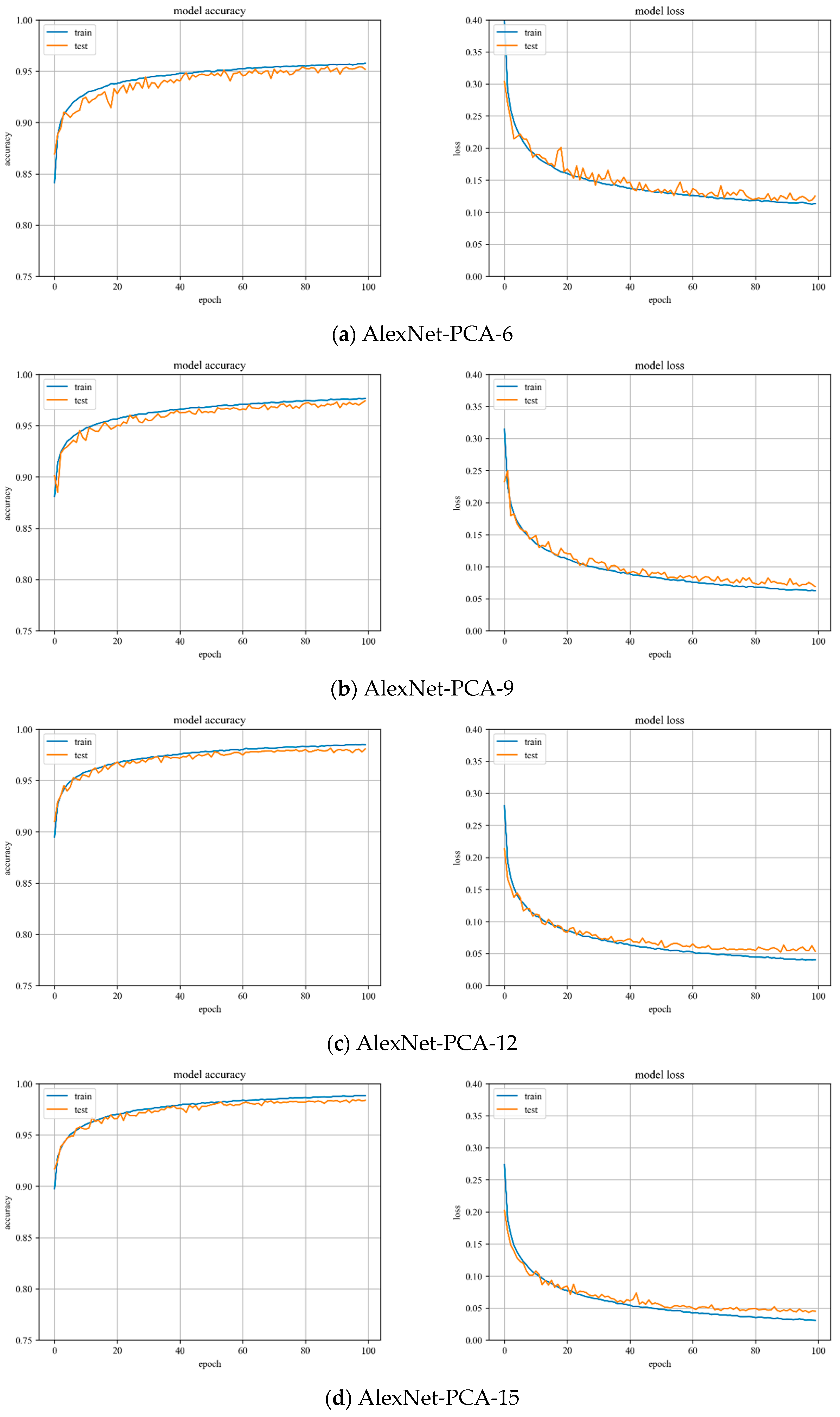
In
Figure 12, the accuracy and loss value curves of the AlexNet model are shown when PCA is used to reduce the dimensionality by 6, 9, 12, and 15. The accuracy curve of the test set starts to converge at the training process of up to 40 iterations and mostly peaks at the training process of up to 60 iterations. The variation is stable throughout the training process and the model has great stability.
3.4.2. AlexNet-PCA Model Testing
As shown in
Table 8, the experimental data analysis can be summarized as follows:
(1) The AlexNet-PCA algorithm for “cotton” and “background” has a minor number of errors in sample recognition classification. It can be attributed to the edge junction containing the reflection spectrum of both the cotton and the background.
(2) Misclassification is observed when using the AlexNet-PCA algorithm to identify the samples of “cotton” and “film on cotton”, “background” and “film on background”. It can be attributed to the weak reflection nature of the film, which leads to an indistinct discrimination of features.
Especially for the PCA dimension selection, a set of linearly increasing dimensions 3, 6, 9 and 12 is chosen for the AlexNet-PCA multi-dimensional algorithm experiment. The linearly increasing dimensions are conducive to the smooth change in the image curve between overall accuracy and dimension; hence, the experimental results are more intuitive.
The Overall Accuracy (OA) of the test sample is shown in
Table 9, representing the percentage of all samples that are accurately predicted. As the number of dimensions retained by PCA increases, the OA of the samples keep increasing.
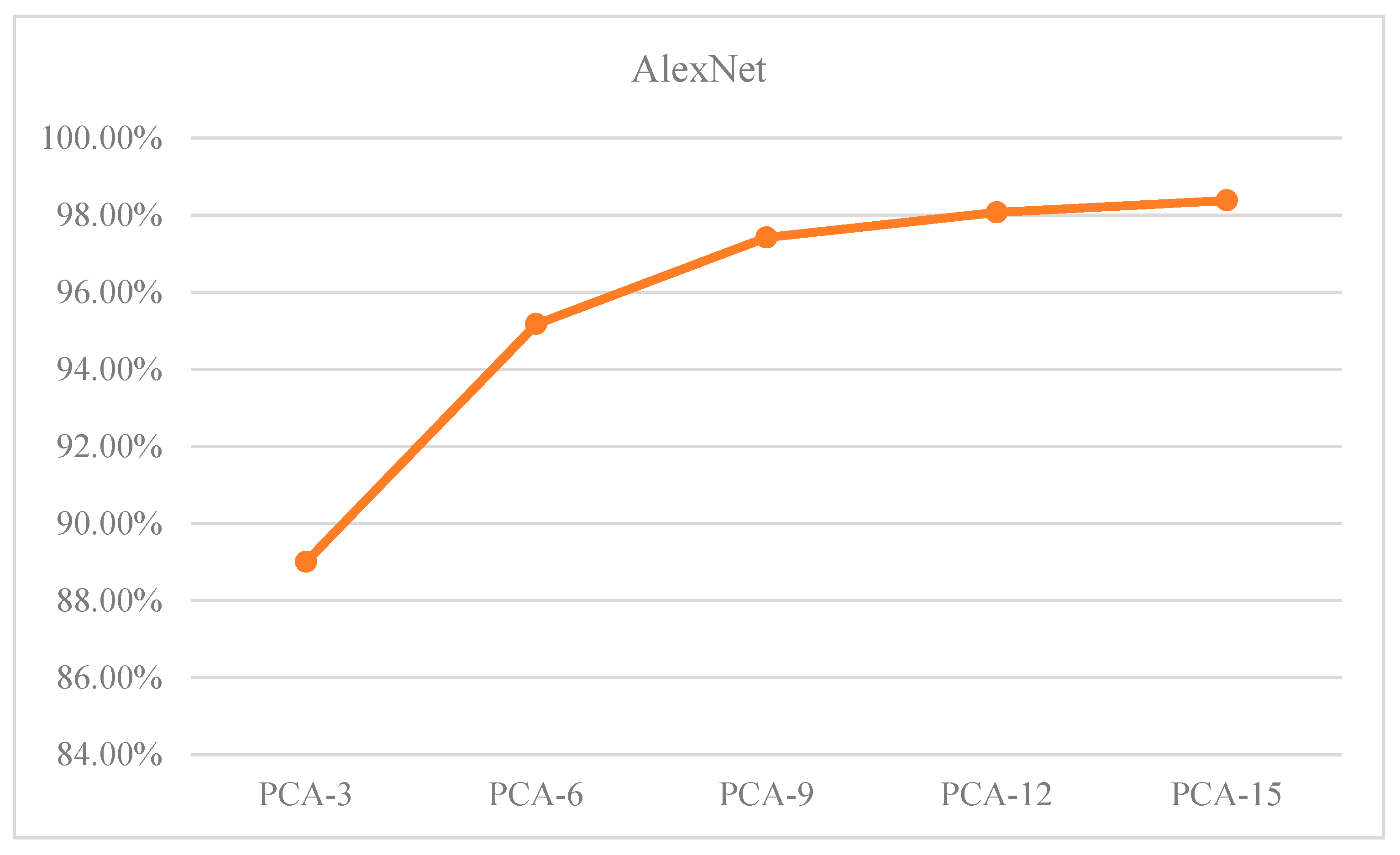
Figure 13 illustrates the OA of the samples as a function of the dimensionality reduction of PCA. The data in
Table 9 and
Figure 13 show teh following:
(1) The increase in PCA dimensionality has an inverse relationship with the increase in accuracy.
(2) When the PCA dimension is set to 12, the proposed algorithm achieves a recognition accuracy of over 98%. Additionally, the overall classification accuracy of the samples begins to converge.
With the increase in PCA dimensionality reduction, the complexity of the neural network model also increases. However, the complexity of the model can lead to overfitting, which in turn can decrease the generalization ability of the model. In the study, we primarily utilize the Dropout method to avoid the overfitting problem. Dropout effectively weakens the connections between neuronal nodes, which reduces the network’s reliance on individual neurons and thereby enhances model generalization ability.
The binarized images are shown in
Figure 14a. As demonstrated in
Figure 14b, the morphological method is utilized to perform an open operation on the binary image, which effectively minimizes the noise caused by light, dust, and artificial marks. As a result, the binary image contains the eliminated artifacts of identified small areas and image edges. The
Figure 14 results show that:
(1) Despite reducing the dimensionality to six utilizing PCA, the post-processing results still exhibit significant imperfections. However, when the dimension reduction is increased to 12, the post-processed image results successfully meet the requirement of providing coordinates. With a dimension reduction of 15, there is no significant difference between the post-processed image results compared to those obtained with a dimension reduction of 12.
(2) Considering the relationship between speed of accuracy improvement, computer performance, image processing results, dimension reduction, and training cost, PCA with a dimension reduction of 12 is the optimal solution for computer performance.
3.4.3. Practical Application Testing of Model AlexNet-PCA-12
As can be seen from the above, the AlexNet-PCA-12 model with the optimal recognition accuracy is obtained experimentally. To verify the feasibility of the research, an application sorting test of the algorithm is conducted in a cotton factory in Aksu, Xinjiang. As depicted in
Figure 15, the computer platform running the algorithm obtains the actual coordinates of the film and inserts them into the industrial control center, which controls the response time of the high-speed spray valve to complete the film removal.
Table 10 shows the data of several sorting experiments: the overall removal rate of film is 97.02%, and the cotton sorting amount can reach 3.0 t/h, which meets the requirements of practical application.
3.5. Summary of Discussions and Results
This chapter focuses on three main tasks: collecting laboratory data, conducting tests on the algorithm, and comparing the visualized recognition results with the experimental results. The recognition effects of LeNet, AlexNet, and VGGNet neural networks combined with LDA, PCA, and ICA dimension reductions are compared and analyzed. Finally, the feasibility of the proposed optimal model is verified for practical applications.
4. Conclusions
Based on hyperspectral images and the deep learning intelligent recognition algorithm, a novel intelligent recognition method for seed cotton mixed with colorless and transparent film is proposed in this paper. The main research topics include the construction of hyperspectral classification systems, dimensionality reduction for hyperspectral data processing, construction of algorithmic recognition models, and the practical application sorting tests.
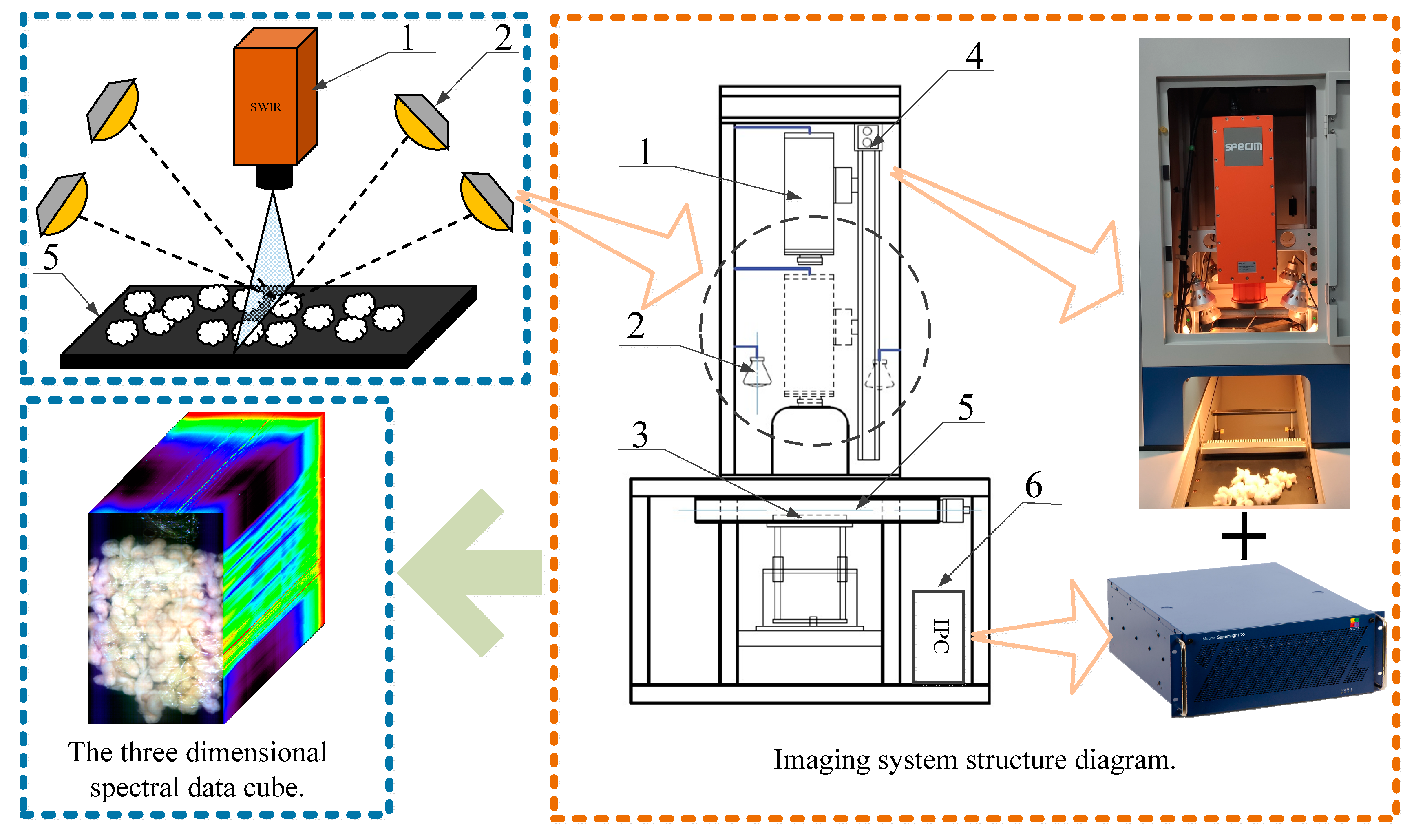
(1) The basic principles of hyperspectral imaging are studied and a hyperspectral classification system is designed for the intelligent classification of seed cotton mixed with the film. The system can obtain 288 hyperspectral data bands with a resolution of 384 pixels × 600 pixels and a spectral range of 1000~2500 nm, which provides an excellent data basis for the recognition of film in seed cotton.
(2) LDA, PCA, and ICA are utilized to reduce the dimension of hyperspectral data to settle the problems of high latitude, large amounts of data, and redundant information of hyperspectral data. Experimental results suggest that LDA and PCA generalized better than ICA. LDA is the best method when the dimensionality reduction is the same as the that of the three. PCA dimensionality reduction is more advantageous when computer performance is satisfied.
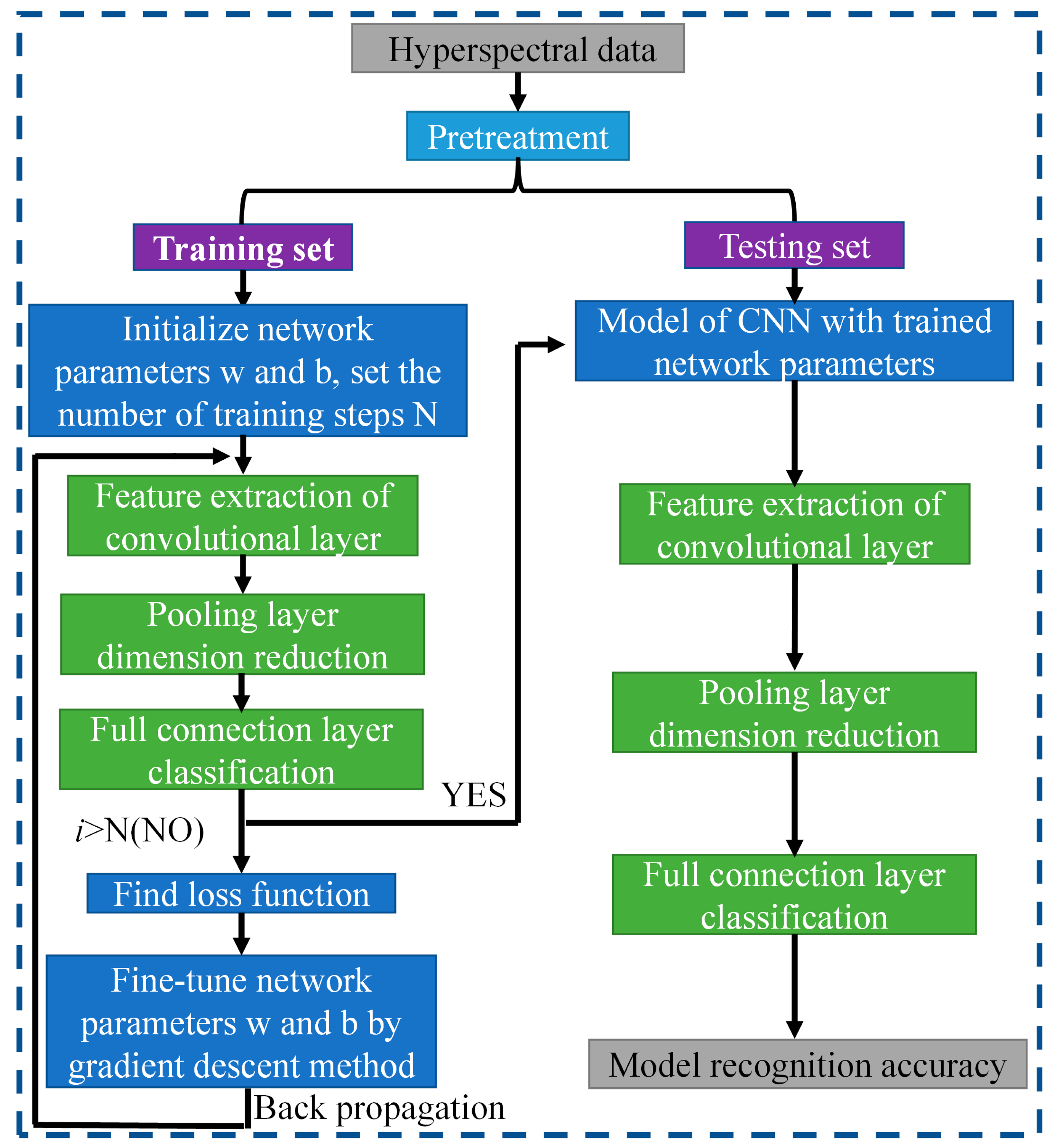
(3) An algorithm is successfully completed for hyperspectral image recognition of film in seed cotton. Based on the convolutional neural network architectures of LeNet, AlexNet, and VGGNet, the network model is constructed for hyperspectral image recognition applications in the seed cotton film domain.
(4) The combination test of the hyperspectral data dimension reduction algorithm (LDA, PCA, ICA) and the CNN model (LeNet, AlexNet, VGGNet) is completed. The experimental results illustrate that when the computer performance is satisfied, AlexNet-PCA-12 can achieve the best cost-to-performance ratio for both recognition and dimensionality reduction, and the recognition accuracy of the algorithm can reach 98.07%; the overall removal rate of film is 97.02% with the data of several sorting experiments in Aksu, Xinjiang.
On the whole, considering the influence of environmental factors such as light, humidity and dust in the practical application sorting tests, data under different environmental variables should be collected to further improve the generalization of the model. However, the research has potential applications in various fields, including but not limited to tea stalks removal, fruit and vegetable flaw separation, and pesticide residue detection in agricultural products. Further research can explore the use of photoelectric separation technology to enhance agricultural development.