Sensing Senses: Optical Biosensors to Study Gustation
Abstract
1. Introduction
2. Taste—Function and Mechanism of Action
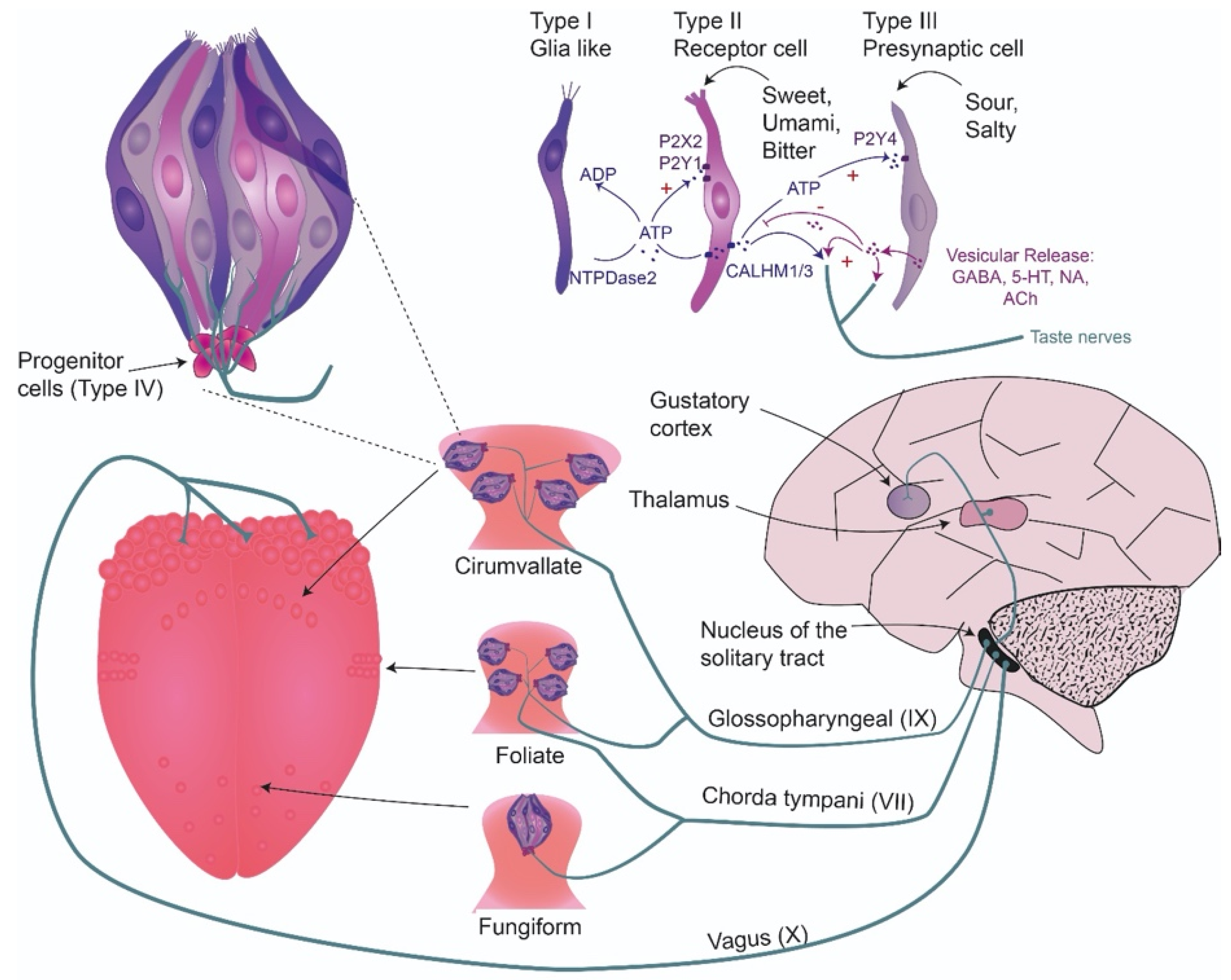
2.1. Taste System Anatomy and Transduction in the Periphery
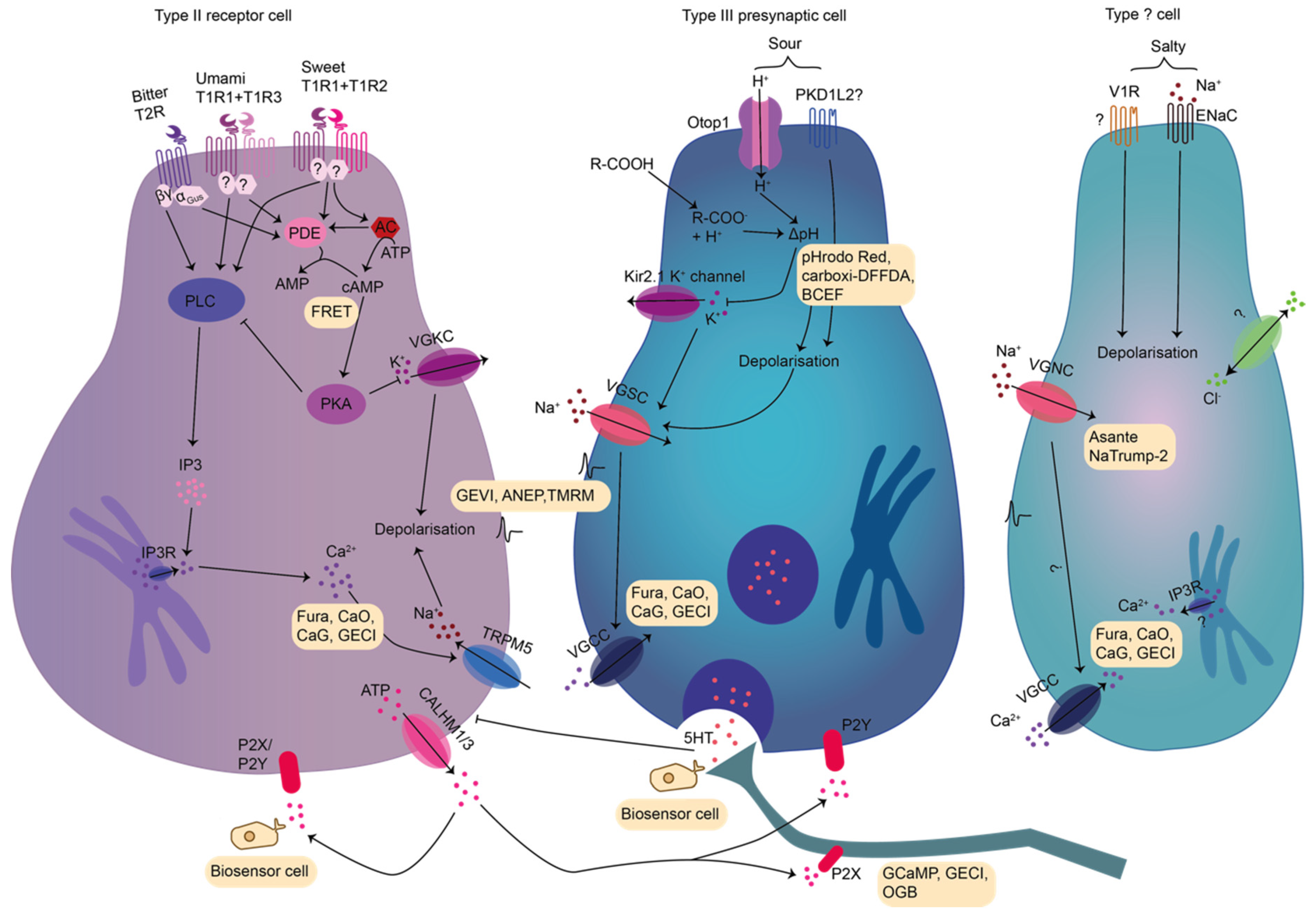
2.1.1. Taste Transduction of Saltiness
2.1.2. Taste Transduction of Sour
2.1.3. Taste Transduction of Bitter
2.1.4. Taste Transduction of Sweet
2.1.5. Taste Transduction of Umami and Additional Taste Qualities
2.1.6. Communication of Taste Cells
2.2. In Search of an Appropriate System to Study Taste Signaling with Biosensors
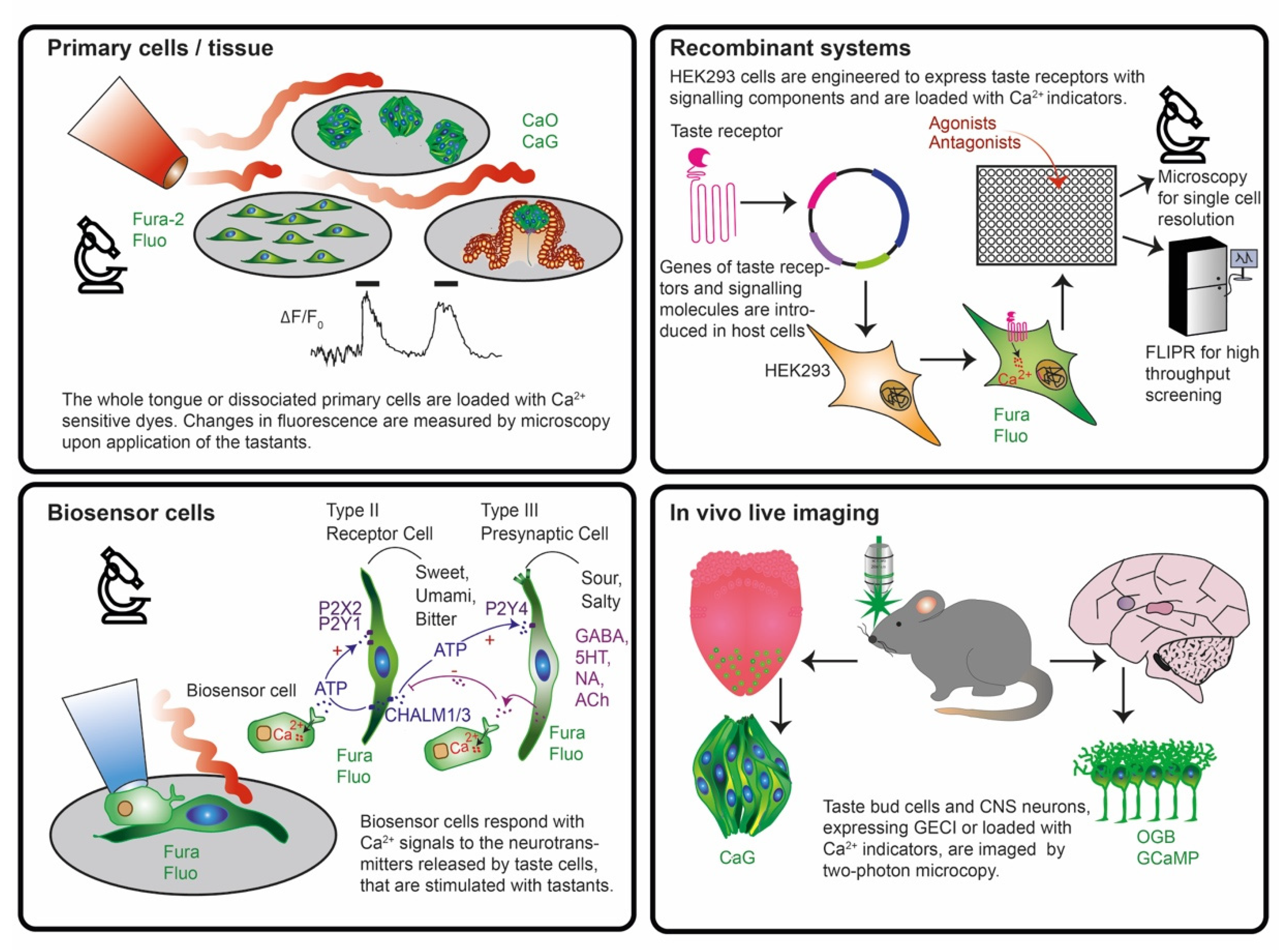
- Isolated animal primary taste cells, taste buds, tongue epithelia and slices were used in combination with fluorescent dyes in ex vivo live imaging experiments for unravelling the intracellular signal transduction pathways and intercellular communication.
- Recombinant systems expressing taste receptors and downstream signaling molecules in non-taste cell lines were employed upon loading with chemical dyes in plate reader experiments to study receptor structure, binding sites, selectivity and sensitivity in high throughput.
- Biosensor cells expressing specific neurotransmitter/hormone receptors were used upon loading with fluorescent dyes and juxtaposed to taste cells/tissue to monitor with live imaging experiments the release of neurotransmitters such as ATP, serotonin, noradrenaline, GABA and acetylcholine.
- Expression of genetically encoded biosensors in neurons of mice to monitor brain activity patterns in response to flavor application in vivo and to label specific cell types in a reporter gene mode.
3. Use of Molecular Optical Biosensors for Taste Research
3.1. Application of Molecular Optical Biosensors for Peripheral Processing of Taste
3.1.1. Chemical Ca2+ Sensors
3.1.2. Genetically Encoded Ca2+ Indicators (GECIs)
3.1.3. Molecular pH Biosensors
3.2. Biosensors to Study Taste Representation in the Central Nervous System
4. Additional Optical Biosensors in the Taste Field
4.1. Voltage-Sensitive Dyes
4.2. cAMP Sensors
5. Recombinant Systems to Study Taste Receptor Function
6. Reporter Genes to Mark Specific Taste Cell Populations in Mice
7. Biosensor Cells to Determine Neurotransmitter Release from Taste Bud Cells
8. Conclusions and Future Perspectives
8.1. New Approaches to Study Taste Physiology
8.2. Development of New In Vitro Taste Systems
9. Literature Research
Author Contributions
Funding
Conflicts of Interest
References
- Ali, J.; Najeeb, J.; Asim Ali, M.; Farhan Aslam, M.; Raza, A. Biosensors: Their Fundamentals, Designs, Types and Most Recent Impactful Applications: A Review. J. Biosens. Bioelectron. 2017, 8. [Google Scholar] [CrossRef]
- Liu, Q.; Wu, C.; Cai, H.; Hu, N.; Zhou, J.; Wang, P. Cell-based biosensors and their application in biomedicine. Chem. Rev. 2014, 114, 6423–6461. [Google Scholar] [CrossRef]
- Podrażka, M.; Bączyńska, E.; Kundys, M.; Jeleń, P.S.; Witkowska Nery, E. Electronic Tongue-A Tool for All Tastes? Biosensors (Basel) 2017, 8, 3. [Google Scholar] [CrossRef]
- Ha, D.; Sun, Q.; Su, K.; Wan, H.; Li, H.; Xu, N.; Sun, F.; Zhuang, L.; Hu, N.; Wang, P. Recent achievements in electronic tongue and bioelectronic tongue as taste sensors. Sens. Actuators B Chem. 2015, 207, 1136–1146. [Google Scholar] [CrossRef]
- Wu, C.; Lillehoj, P.B.; Wang, P. Bioanalytical and chemical sensors using living taste, olfactory, and neural cells and tissues: A short review. Analyst 2015, 140, 7048–7061. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Du, Y.-W.; Huang, L.; Ben-Shoshan Galeczki, Y.; Dagan-Wiener, A.; Naim, M.; Niv, M.Y.; Wang, P. Biomimetic Sensors for the Senses: Towards Better Understanding of Taste and Odor Sensation. Sensors (Basel) 2017, 17, 2881. [Google Scholar] [CrossRef] [PubMed]
- Wasilewski, T.; Kamysz, W.; Gębicki, J. Bioelectronic tongue: Current status and perspectives. Biosens. Bioelectron. 2020, 150, 111923. [Google Scholar] [CrossRef]
- Bioinspired Smell and Taste Sensors, 1st ed.; Wang, P., Liu, Q., Wu, C., Hsia, K.J., Eds.; Springer: Dordrecht, The Netherlands; Beijing, China, 2015; ISBN 9789401773324. [Google Scholar]
- Woertz, K.; Tissen, C.; Kleinebudde, P.; Breitkreutz, J. Taste sensing systems (electronic tongues) for pharmaceutical applications. Int. J. Pharm. 2011, 417, 256–271. [Google Scholar] [CrossRef]
- Gutiérrez, M.; Llobera, A.; Vila-Planas, J.; Capdevila, F.; Demming, S.; Büttgenbach, S.; Mínguez, S.; Jiménez-Jorquera, C. Hybrid electronic tongue based on optical and electrochemical microsensors for quality control of wine. Analyst 2010, 135, 1718–1725. [Google Scholar] [CrossRef]
- Laffitte, A.; Neiers, F.; Briand, L. Functional roles of the sweet taste receptor in oral and extraoral tissues. Curr. Opin. Clin. Nutr. Metab. Care 2014, 17, 379–385. [Google Scholar] [CrossRef]
- Lu, P.; Zhang, C.-H.; Lifshitz, L.M.; ZhuGe, R. Extraoral bitter taste receptors in health and disease. J. Gen. Physiol. 2017, 149, 181–197. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Ishimaru, Y. Oral and extra-oral taste perception. Semin. Cell Dev. Biol. 2013, 24, 240–246. [Google Scholar] [CrossRef]
- Beauchamp, G.K. Why do we like sweet taste: A bitter tale? Physiol. Behav. 2016, 164, 432–437. [Google Scholar] [CrossRef]
- Chaudhari, N.; Roper, S.D. The cell biology of taste. J. Cell Biol. 2010, 190, 285–296. [Google Scholar] [CrossRef]
- Mennella, J.A.; Spector, A.C.; Reed, D.R.; Coldwell, S.E. The Bad Taste of Medicines: Overview of Basic Research on Bitter Taste. Clin. Ther. 2013, 35, 1225–1246. [Google Scholar] [CrossRef] [PubMed]
- Ley, J.P. Masking Bitter Taste by Molecules. Chemosens. Percept. 2008, 1, 58–77. [Google Scholar] [CrossRef]
- Clark, J.E. Taste and flavour: Their importance in food choice and acceptance. Proc. Nutr. Soc. 1998, 57, 639–643. [Google Scholar] [CrossRef] [PubMed]
- Riedel, K.; Sombroek, D.; Fiedler, B.; Siems, K.; Krohn, M. Human cell-based taste perception—A bittersweet job for industry. Nat. Prod. Rep. 2017, 34, 484–495. [Google Scholar] [CrossRef]
- Roper, S.D.; Chaudhari, N. Taste buds: Cells, signals and synapses. Nat. Rev. Neurosci. 2017, 18, 485–497. [Google Scholar] [CrossRef]
- Lindemann, B. Receptor seeks ligand: On the way to cloning the molecular receptors for sweet and bitter taste. Nat. Med. 1999, 5, 381–382. [Google Scholar] [CrossRef]
- Montmayeur, J.-P.; Matsunami, H. Receptors for bitter and sweet taste. Curr. Opin. Neurobiol. 2002, 12, 366–371. [Google Scholar] [CrossRef]
- Witt, M.; Reutter, K.; Miller, I.J., Jr. Morphology of the Peripheral Taste System. In Handbook of Olfaction and Gustation; CRC Press: Boca Raton, FL, USA, 2003; pp. 1142–1188. [Google Scholar]
- Kikut-Ligaj, D.; Trzcielińska-Lorych, J. How taste works: Cells, receptors and gustatory perception. Cell. Mol. Biol. Lett. 2015, 20, 699–716. [Google Scholar] [CrossRef] [PubMed]
- Scott, K. Taste Recognition: Food for Thought. Neuron 2005, 48, 455–464. [Google Scholar] [CrossRef]
- Roper, S.D. Taste buds as peripheral chemosensory processors. Semin. Cell Dev. Biol. 2013, 24, 71–79. [Google Scholar] [CrossRef]
- Bannwarth, M.; Correa, I.R.; Sztretye, M.; Pouvreau, S.; Fellay, C.; Aebischer, A.; Royer, L.; Rois, E.; Johnsson, K. Indo-1 derivatives for local calcium sensing. ACS Chem. Biol. 2009, 4, 179–190. [Google Scholar] [CrossRef] [PubMed]
- Vandenbeuch, A.; Anderson, C.B.; Parnes, J.; Enjyoji, K.; Robson, S.C.; Finger, T.E.; Kinnamon, S.C. Role of the ectonucleotidase NTPDase2 in taste bud function. Proc. Natl. Acad. Sci. USA 2013, 110, 14789–14794. [Google Scholar] [CrossRef]
- Yee, C.L.; Yang, R.; Böttger, B.; Finger, T.E.; Kinnamon, J.C. “Type III” cells of rat taste buds: Immunohistochemical and ultrastructural studies of neuron-specific enolase, protein gene product 9.5, and serotonin. J. Comp. Neurol. 2001, 440, 97–108. [Google Scholar] [CrossRef]
- Miura, H.; Kusakabe, Y.; Harada, S. Cell lineage and differentiation in taste buds. Arch. Histol. Cytol. 2006, 69, 209–225. [Google Scholar] [CrossRef]
- Finger, T.E.; Danilova, V.; Barrows, J.; Bartel, D.L.; Vigers, A.J.; Stone, L.; Hellekant, G.; Kinnamon, S.C. ATP signaling is crucial for communication from taste buds to gustatory nerves. Science 2005, 310, 1495–1499. [Google Scholar] [CrossRef]
- Romanov, R.A.; Rogachevskaja, O.A.; Bystrova, M.F.; Jiang, P.; Margolskee, R.F.; Kolesnikov, S.S. Afferent neurotransmission mediated by hemichannels in mammalian taste cells. EMBO J. 2007, 26, 657–667. [Google Scholar] [CrossRef]
- Dando, R.; Roper, S.D. Cell-to-cell communication in intact taste buds through ATP signalling from pannexin 1 gap junction hemichannels. J. Physiol. (Lond) 2009, 587, 5899–5906. [Google Scholar] [CrossRef]
- Taruno, A.; Vingtdeux, V.; Ohmoto, M.; Ma, Z.; Dvoryanchikov, G.; Li, A.; Adrien, L.; Zhao, H.; Leung, S.; Abernethy, M.; et al. CALHM1 ion channel mediates purinergic neurotransmission of sweet, bitter and umami tastes. Nature 2013, 495, 223–226. [Google Scholar] [CrossRef] [PubMed]
- Romanov, R.A.; Lasher, R.S.; High, B.; Savidge, L.E.; Lawson, A.; Rogachevskaja, O.A.; Zhao, H.; Rogachevsky, V.V.; Bystrova, M.F.; Churbanov, G.D.; et al. Chemical synapses without synaptic vesicles: Purinergic neurotransmission via a CALHM1 channel-mitochondrial signaling complex. Sci. Signal. 2018, 11. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-J.; Maruyama, Y.; Lu, K.-S.; Pereira, E.; Plonsky, I.; Baur, J.E.; Wu, D.; Roper, S.D. Mouse taste buds use serotonin as a neurotransmitter. J. Neurosci. 2005, 25, 843–847. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-J.; Maruyama, Y.; Dvoryanchikov, G.; Pereira, E.; Chaudhari, N.; Roper, S.D. The role of pannexin 1 hemichannels in ATP release and cell-cell communication in mouse taste buds. Proc. Natl. Acad. Sci. USA 2007, 104, 6436–6441. [Google Scholar] [CrossRef]
- Larson, E.D.; Vandenbeuch, A.; Voigt, A.; Meyerhof, W.; Kinnamon, S.C.; Finger, T.E. The Role of 5-HT3 Receptors in Signaling from Taste Buds to Nerves. J. Neurosci. 2015, 35, 15984–15995. [Google Scholar] [CrossRef]
- Huang, Y.A.; Maruyama, Y.; Roper, S.D. Norepinephrine is coreleased with serotonin in mouse taste buds. J. Neurosci. 2008, 28, 13088–13093. [Google Scholar] [CrossRef]
- Huang, Y.A.; Pereira, E.; Roper, S.D. Acid stimulation (sour taste) elicits GABA and serotonin release from mouse taste cells. PLoS ONE 2011, 6, e25471. [Google Scholar] [CrossRef]
- Dvoryanchikov, G.; Huang, Y.A.; Barro-Soria, R.; Chaudhari, N.; Roper, S.D. GABA, its receptors, and GABAergic inhibition in mouse taste buds. J. Neurosci. 2011, 31, 5782–5791. [Google Scholar] [CrossRef]
- Dando, R.; Roper, S.D. Acetylcholine is released from taste cells, enhancing taste signalling. J. Physiol. (Lond) 2012, 590, 3009–3017. [Google Scholar] [CrossRef]
- Gilbertson, T.A.; Damak, S.; Margolskee, R.F. The molecular physiology of taste transduction. Curr. Opin. Neurobiol. 2000, 10, 519–527. [Google Scholar] [CrossRef]
- Roper, S.D. Parallel processing in mammalian taste buds? Physiol. Behav. 2009, 97, 604–608. [Google Scholar] [CrossRef]
- Roper, S.D. Signal transduction and information processing in mammalian taste buds. Pflugers Arch. 2007, 454, 759–776. [Google Scholar] [CrossRef] [PubMed]
- Oka, Y.; Butnaru, M.; Buchholtz, L.v.; Ryba, N.J.P.; Zuker, C.S. High salt recruits aversive taste pathways. Nature 2013, 494, 472–475. [Google Scholar] [CrossRef] [PubMed]
- Chandrashekar, J.; Mueller, K.L.; Hoon, M.A.; Adler, E.; Feng, L.; Guo, W.; Zuker, C.S.; Ryba, N.J.P. T2Rs Function as Bitter Taste Receptors. Cell 2000, 100, 703–711. [Google Scholar] [CrossRef]
- Chandrashekar, J.; Kuhn, C.; Oka, Y.; Yarmolinsky, D.A.; Hummler, E.; Ryba, N.J.P.; Zuker, C.S. The cells and peripheral representation of sodium taste in mice. Nature 2010, 464, 297–301. [Google Scholar] [CrossRef]
- DeFazio, R.A.; Dvoryanchikov, G.; Maruyama, Y.; Kim, J.W.; Pereira, E.; Roper, S.D.; Chaudhari, N. Separate populations of receptor cells and presynaptic cells in mouse taste buds. J. Neurosci. 2006, 26, 3971–3980. [Google Scholar] [CrossRef]
- Hacker, K.; Laskowski, A.; Feng, L.; Restrepo, D.; Medler, K. Evidence for two populations of bitter responsive taste cells in mice. J. Neurophysiol. 2008, 99, 1503–1514. [Google Scholar] [CrossRef]
- Ossebaard, C.A.; Smith, D.V. Effect of amiloride on the taste of NaCl, Na-gluconate and KCl in humans: Implications for Na+ receptor mechanisms. Chem. Senses 1995, 20, 37–46. [Google Scholar] [CrossRef]
- Feldman, G.M.; Mogyorosi, A.; Heck, G.L.; Desimone, J.A.; Santos, C.R.; Clary, R.A.; Lyall, V. Salt-evoked lingual surface potential in humans. J. Neurophysiol. 2003, 90, 2060–2064. [Google Scholar] [CrossRef]
- Lyall, V.; Heck, G.L.; Vinnikova, A.K.; Ghosh, S.; Phan, T.-H.T.; Alam, R.I.; Russell, O.F.; Malik, S.A.; Bigbee, J.W.; Desimone, J.A. The mammalian amiloride-insensitive non-specific salt taste receptor is a vanilloid receptor-1 variant. J. Physiol. (Lond) 2004, 558, 147–159. [Google Scholar] [CrossRef]
- Lyall, V.; Heck, G.L.; Vinnikova, A.K.; Ghosh, S.; Phan, T.-H.T.; Desimone, J.A. A novel vanilloid receptor-1 (VR-1) variant mammalian salt taste receptor. Chem. Senses 2005, 30 (Suppl. 1), i42–i43. [Google Scholar] [CrossRef]
- Roebber, J.K.; Roper, S.D.; Chaudhari, N. The Role of the Anion in Salt (NaCl) Detection by Mouse Taste Buds. J. Neurosci. 2019, 39, 6224–6232. [Google Scholar] [CrossRef]
- Huang, Y.A.; Maruyama, Y.; Stimac, R.; Roper, S.D. Presynaptic (Type III) cells in mouse taste buds sense sour (acid) taste. J. Physiol. (Lond) 2008, 586, 2903–2912. [Google Scholar] [CrossRef] [PubMed]
- Kataoka, S.; Yang, R.; Ishimaru, Y.; Matsunami, H.; Sévigny, J.; Kinnamon, J.C.; Finger, T.E. The candidate sour taste receptor, PKD2L1, is expressed by type III taste cells in the mouse. Chem. Senses 2008, 33, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Ramos Da Conceicao Neta, E.R.; Johanningsmeier, S.D.; McFeeters, R.F. The Chemistry and Physiology of Sour Taste—A Review. J. Food Sci. 2007, 72, R33–R38. [Google Scholar] [CrossRef] [PubMed]
- Richter, T.A.; Caicedo, A.; Roper, S.D. Sour taste stimuli evoke Ca2+ and pH responses in mouse taste cells. J. Physiol. (Lond) 2003, 547, 475–483. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, T.; Okada, Y.; Sato, T. Ionic basis of receptor potential of frog taste cells induced by acid stimuli. J. Physiol. (Lond) 1988, 405, 699–711. [Google Scholar] [CrossRef]
- Kinnamon, S.C.; Roper, S.D. Membrane properties of isolated mudpuppy taste cells. J. Gen. Physiol. 1988, 91, 351–371. [Google Scholar] [CrossRef]
- Gilbertson, T.A.; Roper, S.D.; Kinnamon, S.C. Proton currents through amiloride-sensitive Na+ channels in isolated hamster taste cells: Enhancement by vasopressin and cAMP. Neuron 1993, 10, 931–942. [Google Scholar] [CrossRef]
- Miyamoto, T.; Fujiyama, R.; Okada, Y.; Sato, T. Sour transduction involves activation of NPPB-sensitive conductance in mouse taste cells. J. Neurophysiol. 1998, 80, 1852–1859. [Google Scholar] [CrossRef] [PubMed]
- Ugawa, S.; Minami, Y.; Guo, W.; Saishin, Y.; Takatsuji, K.; Yamamoto, T.; Tohyama, M.; Shimada, S. Receptor that leaves a sour taste in the mouth. Nature 1998, 395, 555–556. [Google Scholar] [CrossRef] [PubMed]
- Stevens, D.R.; Seifert, R.; Bufe, B.; Müller, F.; Kremmer, E.; Gauss, R.; Meyerhof, W.; Kaupp, U.B.; Lindemann, B. Hyperpolarization-activated channels HCN1 and HCN4 mediate responses to sour stimuli. Nature 2001, 413, 631–635. [Google Scholar] [CrossRef] [PubMed]
- Huang, A.L.; Chen, X.; Hoon, M.A.; Chandrashekar, J.; Guo, W.; Tränkner, D.; Ryba, N.J.P.; Zuker, C.S. The cells and logic for mammalian sour taste detection. Nature 2006, 442, 934–938. [Google Scholar] [CrossRef] [PubMed]
- Lyall, V.; Alam, R.I.; Phan, D.Q.; Ereso, G.L.; Phan, T.H.; Malik, S.A.; Montrose, M.H.; Chu, S.; Heck, G.L.; Feldman, G.M.; et al. Decrease in rat taste receptor cell intracellular pH is the proximate stimulus in sour taste transduction. Am. J. Physiol. Cell Physiol. 2001, 281, C1005–C1013. [Google Scholar] [CrossRef]
- Nelson, T.M.; Lopezjimenez, N.D.; Tessarollo, L.; Inoue, M.; Bachmanov, A.A.; Sullivan, S.L. Taste function in mice with a targeted mutation of the pkd1l3 gene. Chem. Senses 2010, 35, 565–577. [Google Scholar] [CrossRef]
- Teng, B.; Wilson, C.E.; Tu, Y.-H.; Joshi, N.R.; Kinnamon, S.C.; Liman, E.R. Cellular and Neural Responses to Sour Stimuli Require the Proton Channel Otop1. Curr. Biol. 2019, 29, 3647–3656.e5. [Google Scholar] [CrossRef]
- Go, Y.; Satta, Y.; Takenaka, O.; Takahata, N. Lineage-specific loss of function of bitter taste receptor genes in humans and nonhuman primates. Genetics 2005, 170, 313–326. [Google Scholar] [CrossRef]
- Adler, E.; Hoon, M.A.; Mueller, K.L.; Chandrashekar, J.; Ryba, N.J.P.; Zuker, C.S. A Novel Family of Mammalian Taste Receptors. Cell 2000, 100, 693–702. [Google Scholar] [CrossRef]
- Kuhn, C.; Bufe, B.; Winnig, M.; Hofmann, T.; Frank, O.; Behrens, M.; Lewtschenko, T.; Slack, J.P.; Ward, C.D.; Meyerhof, W. Bitter taste receptors for saccharin and acesulfame K. J. Neurosci. 2004, 24, 10260–10265. [Google Scholar] [CrossRef]
- Meyerhof, W.; Batram, C.; Kuhn, C.; Brockhoff, A.; Chudoba, E.; Bufe, B.; Appendino, G.; Behrens, M. The molecular receptive ranges of human TAS2R bitter taste receptors. Chem. Senses 2010, 35, 157–170. [Google Scholar] [CrossRef] [PubMed]
- Caicedo, A.; Roper, S.D. Taste receptor cells that discriminate between bitter stimuli. Science 2001, 291, 1557–1560. [Google Scholar] [CrossRef] [PubMed]
- Matsunami, H.; Montmayeur, J.P.; Buck, L.B. A family of candidate taste receptors in human and mouse. Nature 2000, 404, 601–604. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.T.; Gannon, K.S.; Margolskee, R.F. Transduction of bitter and sweet taste by gustducin. Nature 1996, 381, 796–800. [Google Scholar] [CrossRef]
- Choi, M.; Lee, W.M.; Yun, S.H. Intravital Microscopic Interrogation of Peripheral Taste Sensation. Sci. Rep. 2015, 5, 8661. [Google Scholar] [CrossRef]
- Ming, D.; Ninomiya, Y.; Margolskee, R.F. Blocking taste receptor activation of gustducin inhibits gustatory responses to bitter compounds. Proc. Natl. Acad. Sci. USA 1999, 96, 9903–9908. [Google Scholar] [CrossRef]
- Huang, L.; Shanker, Y.G.; Dubauskaite, J.; Zheng, J.Z.; Yan, W.; Rosenzweig, S.; Spielman, A.I.; Max, M.; Margolskee, R.F. Gγ13 colocalizes with gustducin in taste receptor cells and mediates IP 3 responses to bitter denatonium. Nat. Neurosci. 1999, 2, 1055–1062. [Google Scholar] [CrossRef]
- Liu, D.; Liman, E.R. Intracellular Ca2+ and the phospholipid PIP2 regulate the taste transduction ion channel TRPM5. Proc. Natl. Acad. Sci. USA 2003, 100, 15160–15165. [Google Scholar] [CrossRef] [PubMed]
- Chandrashekar, J.; Hoon, M.A.; Ryba, N.J.P.; Zuker, C.S. The receptors and cells for mammalian taste. Nature 2006, 444, 288–294. [Google Scholar] [CrossRef] [PubMed]
- Avau, B.; Depoortere, I. The bitter truth about bitter taste receptors: Beyond sensing bitter in the oral cavity. Acta Physiol. (Oxf) 2016, 216, 407–420. [Google Scholar] [CrossRef] [PubMed]
- Nelson, G.; Hoon, M.A.; Chandrashekar, J.; Zhang, Y.; Ryba, N.J.P.; Zuker, C.S. Mammalian Sweet Taste Receptors. Cell 2001, 106, 381–390. [Google Scholar] [CrossRef]
- Kitagawa, M.; Kusakabe, Y.; Miura, H.; Ninomiya, Y.; Hino, A. Molecular Genetic Identification of a Candidate Receptor Gene for Sweet Taste. Biochem. Biophys. Res. Commun. 2001, 283, 236–242. [Google Scholar] [CrossRef] [PubMed]
- Max, M.; Shanker, Y.G.; Huang, L.; Rong, M.; Liu, Z.; Campagne, F.; Weinstein, H.; Damak, S.; Margolskee, R.F. Tas1r3, encoding a new candidate taste receptor, is allelic to the sweet responsiveness locus Sac. Nat. Genet. 2001, 28, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Damak, S.; Rong, M.; Yasumatsu, K.; Kokrashvili, Z.; Varadarajan, V.; Zou, S.; Jiang, P.; Ninomiya, Y.; Margolskee, R.F. Detection of sweet and umami taste in the absence of taste receptor T1r3. Science 2003, 301, 850–853. [Google Scholar] [CrossRef] [PubMed]
- Yee, K.K.; Sukumaran, S.K.; Kotha, R.; Gilbertson, T.A.; Margolskee, R.F. Glucose transporters and ATP-gated K+ (KATP) metabolic sensors are present in type 1 taste receptor 3 (T1r3)-expressing taste cells. Proc. Natl. Acad. Sci. USA 2011, 108, 5431–5436. [Google Scholar] [CrossRef] [PubMed]
- Avenet, P.; Lindemann, B. Patch-clamp study of isolated taste receptor cells of the frog. J. Membr. Biol. 1987, 97, 223–240. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, K.; Koyama, N. High activity of adenyl cyclase in olfactory and gustatory organs. Biochem. Biophys. Res. Commun. 1972, 48, 30–34. [Google Scholar] [CrossRef]
- Naim, M.; Ronen, T.; Striem, B.J.; Levinson, M.; Zehavi, U. Adenylate cyclase responses to sucrose stimulation in membranes of pig circumvallate taste papillae. Comp. Biochem. Physiol. Part B Comp. Biochem. 1991, 100, 455–458. [Google Scholar] [CrossRef]
- Striem, B.J.; Pace, U.; Zehavi, U.; Naim, M.; Lancet, D. Sweet tastants stimulate adenylate cyclase coupled to GTP-binding protein in rat tongue membranes. Biochem. J. 1989, 260, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Striem, B.J.; Naim, M.; Lindemann, B. Generation of Cyclic AMP in Taste Buds of the Rat Circumvallate Papilla in Response to Sucrose. Cell Physiol. Biochem. 1991, 1, 46–54. [Google Scholar] [CrossRef]
- Li, X.; Staszewski, L.; Xu, H.; Durick, K.; Zoller, M.; Adler, E. Human receptors for sweet and umami taste. Proc. Natl. Acad. Sci. USA 2002, 99, 4692–4696. [Google Scholar] [CrossRef] [PubMed]
- Nelson, G.; Chandrashekar, J.; Hoon, M.A.; Feng, L.; Zhao, G.; Ryba, N.J.P.; Zuker, C.S. An amino-acid taste receptor. Nature 2002, 416, 199–202. [Google Scholar] [CrossRef] [PubMed]
- Kinnamon, S.C. Umami taste transduction mechanisms. Am. J. Clin. Nutr. 2009, 90, 753S–755S. [Google Scholar] [CrossRef] [PubMed]
- Chandrashekar, J.; Yarmolinsky, D.; Buchholtz, L.v.; Oka, Y.; Sly, W.; Ryba, N.J.P.; Zuker, C.S. The taste of carbonation. Science 2009, 326, 443–445. [Google Scholar] [CrossRef] [PubMed]
- Dramane, G.; Abdoul-Azize, S.; Hichami, A.; Vögtle, T.; Akpona, S.; Chouabe, C.; Sadou, H.; Nieswandt, B.; Besnard, P.; Khan, N.A. STIM1 regulates calcium signaling in taste bud cells and preference for fat in mice. J. Clin. Investig. 2012, 122, 2267–2282. [Google Scholar] [CrossRef] [PubMed]
- Ozdener, M.H.; Subramaniam, S.; Sundaresan, S.; Sery, O.; Hashimoto, T.; Asakawa, Y.; Besnard, P.; Abumrad, N.A.; Khan, N.A. CD36- and GPR120-mediated Ca(2)(+) signaling in human taste bud cells mediates differential responses to fatty acids and is altered in obese mice. Gastroenterology 2014, 146, 995–1005. [Google Scholar] [CrossRef]
- Ohsu, T.; Amino, Y.; Nagasaki, H.; Yamanaka, T.; Takeshita, S.; Hatanaka, T.; Maruyama, Y.; Miyamura, N.; Eto, Y. Involvement of the calcium-sensing receptor in human taste perception. J. Biol. Chem. 2010, 285, 1016–1022. [Google Scholar] [CrossRef]
- Maruyama, Y.; Yasuda, R.; Kuroda, M.; Eto, Y. Kokumi Substances, Enhancers of Basic Tastes, Induce Responses in Calcium-Sensing Receptor Expressing Taste Cells. PLoS ONE 2012, 7, e34489. [Google Scholar] [CrossRef]
- Ishida, Y.; Ugawa, S.; Ueda, T.; Murakami, S.; Shimada, S. Vanilloid receptor subtype-1 (VR1) is specifically localized to taste papillae. Mol. Brain Res. 2002, 107, 17–22. [Google Scholar] [CrossRef]
- Smutzer, G.; Devassy, R.K. Integrating TRPV1 Receptor Function with Capsaicin Psychophysics. Adv. Pharmacol. Sci. 2016, 2016, 1512457. [Google Scholar] [CrossRef]
- Hacker, K.; Medler, K.F. Mitochondrial calcium buffering contributes to the maintenance of Basal calcium levels in mouse taste cells. J. Neurophysiol. 2008, 100, 2177–2191. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Huang, Y.A.; Dando, R.; Roper, S.D. Autocrine and paracrine roles for ATP and serotonin in mouse taste buds. J. Neurosci. 2009, 29, 13909–13918. [Google Scholar] [CrossRef] [PubMed]
- Eguchi, K.; Ohtubo, Y.; Yoshii, K. Functional expression of M3, a muscarinic acetylcholine receptor subtype, in taste bud cells of mouse fungiform papillae. Chem. Senses 2008, 33, 47–55. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Simon, S.A.; Baggett, H.C. Identification of muscarinic acetylcholine receptors in isolated canine lingual epithelia via voltage clamp measurements. Arch. Oral Biol. 1992, 37, 685–690. [Google Scholar] [CrossRef]
- Ogura, T.; Lin, W. Acetylcholine and acetylcholine receptors in taste receptor cells. Chem. Senses 2005, 30 (Suppl. 1), i41. [Google Scholar] [CrossRef]
- Huang, A.Y.; Wu, S.Y. Substance P as a putative efferent transmitter mediates GABAergic inhibition in mouse taste buds. Br. J. Pharmacol. 2018, 175, 1039–1053. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, D.; Xu, M.; Liu, F.; Millar, S.E.; Barlow, L.A. β-Catenin Signaling Biases Multipotent Lingual Epithelial Progenitors to Differentiate and Acquire Specific Taste Cell Fates. PLoS Genet. 2015, 11, e1005208. [Google Scholar] [CrossRef] [PubMed]
- Lawton, D.M.; Furness, D.N.; Lindemann, B.; Hackney, C.M. Localization of the glutamate-aspartate transporter, GLAST, in rat taste buds. Eur. J. Neurosci. 2000, 12, 3163–3171. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.T.; Ruiz-Avila, L.; Margolskee, R.F. Directing Gene Expression to Gustducin-Positive Taste Receptor Cells. J. Neurosci. 1999, 19, 5802–5809. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhao, Z.; Margolskee, R.; Liman, E. The transduction channel TRPM5 is gated by intracellular calcium in taste cells. J. Neurosci. 2007, 27, 5777–5786. [Google Scholar] [CrossRef] [PubMed]
- Clapp, T.R.; Medler, K.F.; Damak, S.; Margolskee, R.F.; Kinnamon, S.C. Mouse taste cells with G protein-coupled taste receptors lack voltage-gated calcium channels and SNAP-25. BMC Biol. 2006, 4, 7. [Google Scholar] [CrossRef] [PubMed]
- Tomchik, S.M.; Berg, S.; Kim, J.W.; Chaudhari, N.; Roper, S.D. Breadth of tuning and taste coding in mammalian taste buds. J. Neurosci. 2007, 27, 10840–10848. [Google Scholar] [CrossRef] [PubMed]
- Beidler, L.M.; Fishman, I.Y.; Hardiman, C.W. Species Differences in Taste Responses. Am. J. Physiol. 1955, 181, 235–239. [Google Scholar] [CrossRef] [PubMed]
- Go, Y. Lineage-Specific Expansions and Contractions of the Bitter Taste Receptor Gene Repertoire in Vertebrates. Mol. Biol. Evol. 2006, 23, 964–972. [Google Scholar] [CrossRef]
- Hochheimer, A.; Krohn, M.; Rudert, K.; Riedel, K.; Becker, S.; Thirion, C.; Zinke, H. Endogenous gustatory responses and gene expression profile of stably proliferating human taste cells isolated from fungiform papillae. Chem. Senses 2014, 39, 359–377. [Google Scholar] [CrossRef]
- Behrens, M.; Meyerhof, W. Bitter taste receptors and human bitter taste perception. Cell. Mol. Life Sci. 2006, 63, 1501–1509. [Google Scholar] [CrossRef]
- Li, X.; Li, W.; Wang, H.; Bayley, D.L.; Cao, J.; Reed, D.R.; Bachmanov, A.A.; Huang, L.; Legrand-Defretin, V.; Beauchamp, G.K.; et al. Cats Lack a Sweet Taste Receptor. J. Nutr. 2006, 136, 1932S–1934S. [Google Scholar] [CrossRef]
- Jiang, P.; Cui, M.; Zhao, B.; Liu, Z.; Snyder, L.A.; Benard, L.M.J.; Osman, R.; Margolskee, R.F.; Max, M. Lactisole interacts with the transmembrane domains of human T1R3 to inhibit sweet taste. J. Biol. Chem. 2005, 280, 15238–15246. [Google Scholar] [CrossRef]
- Inoue, M.; McCaughey, S.A.; Bachmanov, A.A.; Beauchamp, G.K. Whole nerve chorda tympani responses to sweeteners in C57BL/6ByJ and 129P3/J mice. Chem. Senses 2001, 26, 915–923. [Google Scholar] [CrossRef][Green Version]
- Sclafani, A. The sixth taste? Appetite 2004, 43, 1–3. [Google Scholar] [CrossRef]
- Sclafani, A.; Zukerman, S.; Glendinning, J.I.; Margolskee, R.F. Fat and carbohydrate preferences in mice: The contribution of alpha-gustducin and Trpm5 taste-signaling proteins. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2007, 293, R1504–R1513. [Google Scholar] [CrossRef] [PubMed]
- Barlow, L.A. Progress and renewal in gustation: New insights into taste bud development. Development 2015, 142, 3620–3629. [Google Scholar] [CrossRef]
- Ozdener, M.H.; Rawson, N.E. Primary culture of mammalian taste epithelium. Methods Mol. Biol. 2013, 945, 95–107. [Google Scholar] [CrossRef]
- Aihara, E.; Mahe, M.M.; Schumacher, M.A.; Matthis, A.L.; Feng, R.; Ren, W.; Noah, T.K.; Matsu-ura, T.; Moore, S.R.; Hong, C.I.; et al. Characterization of stem/progenitor cell cycle using murine circumvallate papilla taste bud organoid. Sci. Rep. 2015, 5, 17185. [Google Scholar] [CrossRef] [PubMed]
- Ren, W.; Lewandowski, B.C.; Watson, J.; Aihara, E.; Iwatsuki, K.; Bachmanov, A.A.; Margolskee, R.F.; Jiang, P. Single Lgr5- or Lgr6-expressing taste stem/progenitor cells generate taste bud cells ex vivo. Proc. Natl. Acad. Sci. USA 2014, 111, 16401–16406. [Google Scholar] [CrossRef]
- Ren, W.; Aihara, E.; Lei, W.; Gheewala, N.; Uchiyama, H.; Margolskee, R.F.; Iwatsuki, K.; Jiang, P. Transcriptome analyses of taste organoids reveal multiple pathways involved in taste cell generation. Sci. Rep. 2017, 7, 4004. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Du, L.; Zou, L.; Zhao, L.; Huang, L.; Wang, P. Recent advances in taste cell- and receptor-based biosensors. Sens. Actuators B Chem. 2014, 201, 75–85. [Google Scholar] [CrossRef]
- Gee, K.R.; Archer, E.A.; Lapham, L.A.; Leonard, M.E.; Zhou, Z.-L.; Bingham, J.; Diwu, Z. New ratiometric fluorescent calcium indicators with moderately attenuated binding affinities. Bioorganic Med. Chem. Lett. 2000, 10, 1515–1518. [Google Scholar] [CrossRef]
- Grynkiewicz, G.; Poenie, M.; Tsien, R.Y. A new generation of Ca2+ indicators with greatly improved fluorescence properties. J. Biol. Chem. 1985, 260, 3440–3450. [Google Scholar]
- Tsien, R.Y.; Rink, T.J.; Poenie, M. Measurement of cytosolic free Ca2+ in individual small cells using fluorescence microscopy with dual excitation wavelengths. Cell Calcium 1985, 6, 145–157. [Google Scholar] [CrossRef]
- Akabas, M.H.; Dodd, J.; Al-Awqati, Q. A bitter substance induces a rise in intracellular calcium in a subpopulation of rat taste cells. Science 1988, 242, 1047–1050. [Google Scholar] [CrossRef]
- Ogura, T.; Mackay-Sim, A.; Kinnamon, S.C. Bitter Taste Transduction of Denatonium in the Mudpuppy Necturus maculosus. J. Neurosci. 1997, 17, 3580–3587. [Google Scholar] [CrossRef]
- Bootman, M.D.; Rietdorf, K.; Collins, T.; Walker, S.; Sanderson, M. Ca2+-sensitive fluorescent dyes and intracellular Ca2+ imaging. Cold Spring Harb. Protoc. 2013, 2013, 83–99. [Google Scholar] [CrossRef] [PubMed]
- Di Virgilio, F.; Steinberg, T.H.; Silverstein, S.C. Inhibition of Fura-2 sequestration and secretion with organic anion transport blockers. Cell Calcium 1990, 11, 57–62. [Google Scholar] [CrossRef]
- Oakes, S.G.; Martin, W.J.; Lisek, C.A.; Powis, G. Incomplete hydrolysis of the calcium indicator precursor fura-2 pentaacetoxymethyl ester (fura-2 AM) by cells. Anal. Biochem. 1988, 169, 159–166. [Google Scholar] [CrossRef]
- Jakob, I.; Hauser, I.A.; Thévenod, F.; Lindemann, B. MDR1 in taste buds of rat vallate papilla: Functional, immunohistochemical, and biochemical evidence. Am. J. Physiol. 1998, 274, C182–C191. [Google Scholar] [CrossRef]
- Bernhardt, S.J.; Naim, M.; Zehavi, U.; Lindemann, B. Changes in IP3 and cytosolic Ca2+ in response to sugars and non-sugar sweeteners in transduction of sweet taste in the rat. J. Physiol. (Lond) 1996, 490, 325–336. [Google Scholar] [CrossRef] [PubMed]
- Baryshnikov, S.G.; Rogachevskaja, O.A.; Kolesnikov, S.S. Calcium signaling mediated by P2Y receptors in mouse taste cells. J. Neurophysiol. 2003, 90, 3283–3294. [Google Scholar] [CrossRef] [PubMed]
- Clapp, T.R.; Trubey, K.R.; Vandenbeuch, A.; Stone, L.M.; Margolskee, R.F.; Chaudhari, N.; Kinnamon, S.C. Tonic activity of Gα-gustducin regulates taste cell responsivity. FEBS Lett. 2008, 582, 3783–3787. [Google Scholar] [CrossRef]
- Sinclair, M.S.; Perea-Martinez, I.; Dvoryanchikov, G.; Yoshida, M.; Nishimori, K.; Roper, S.D.; Chaudhari, N. Oxytocin Signaling in Mouse Taste Buds. PLoS ONE 2010, 5, e11980. [Google Scholar] [CrossRef]
- Huang, Y.A.; Grant, J.; Roper, S. Glutamate may be an efferent transmitter that elicits inhibition in mouse taste buds. PLoS ONE 2012, 7, e30662. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.V.; Bobkov, Y.V.; Kolesnikov, S.S. Adenosine triphosphate mobilizes cytosolic calcium and modulates ionic currents in mouse taste receptor cells. Neurosci. Lett. 2000, 290, 165–168. [Google Scholar] [CrossRef]
- Szebenyi, S.A.; Laskowski, A.I.; Medler, K.F. Sodium/calcium exchangers selectively regulate calcium signaling in mouse taste receptor cells. J. Neurophysiol. 2010, 104, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Medler, K.F.; Margolskee, R.F.; Kinnamon, S.C. Electrophysiological Characterization of Voltage-Gated Currents in Defined Taste Cell Types of Mice. J. Neurosci. 2003, 23, 2608–2617. [Google Scholar] [CrossRef]
- Herness, S.; Zhao, F.-l.; Kaya, N.; Lu, S.-g.; Shen, T.; Sun, X.-D. Adrenergic signalling between rat taste receptor cells. J. Physiol. (Lond) 2002, 543, 601–614. [Google Scholar] [CrossRef]
- Ogura, T.; Kinnamon, S.C. IP(3)-Independent release of Ca(2+) from intracellular stores: A novel mechanism for transduction of bitter stimuli. J. Neurophysiol. 1999, 82, 2657–2666. [Google Scholar] [CrossRef] [PubMed]
- Roberts, C.D.; Dvoryanchikov, G.; Roper, S.D.; Chaudhari, N. Interaction between the second messengers cAMP and Ca2+ in mouse presynaptic taste cells. J. Physiol. (Lond) 2009, 587, 1657–1668. [Google Scholar] [CrossRef]
- Kawaguchi, H.; Yamanaka, A.; Uchida, K.; Shibasaki, K.; Sokabe, T.; Maruyama, Y.; Yanagawa, Y.; Murakami, S.; Tominaga, M. Activation of Polycystic Kidney Disease-2-like 1 (PKD2L1)-PKD1L3 Complex by Acid in Mouse Taste Cells. J. Biol. Chem. 2010, 285, 17277–17281. [Google Scholar] [CrossRef]
- Banik, D.D.; Martin, L.E.; Freichel, M.; Torregrossa, A.-M.; Medler, K.F. TRPM4 and TRPM5 are both required for normal signalling in taste receptor cells. PNAS 2018, 115, E772–E781. [Google Scholar] [CrossRef]
- Hayashi, Y.; Zviman, M.M.; Brand, J.G.; Teeter, J.H.; Restrepo, D. Measurement of membrane potential and [Ca2+]i in cell ensembles: Application to the study of glutamate taste in mice. Biophys. J. 1996, 71, 1057–1070. [Google Scholar] [CrossRef][Green Version]
- Wong, S.T.; Henley, J.R.; Kanning, K.C.; Huang, K.-h.; Bothwell, M.; Poo, M.-m. A p75 NTR and Nogo receptor complex mediates repulsive signaling by myelin-associated glycoprotein. Nat. Neurosci. 2002, 5, 1302–1308. [Google Scholar] [CrossRef] [PubMed]
- Ishimaru, Y.; Inada, H.; Kubota, M.; Zhuang, H.; Tominaga, M.; Matsunami, H. Transient receptor potential family members PKD1L3 and PKD2L1 form a candidate sour taste receptor. Proc. Natl. Acad. Sci. USA 2006, 103, 12569–12574. [Google Scholar] [CrossRef] [PubMed]
- Paredes, R.M.; Etzler, J.C.; Watts, L.T.; Lechleiter, J.D. Chemical Calcium Indicators. Methods 2008, 46, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Cherkashin, A.P.; Kolesnikova, A.S.; Tarasov, M.V.; Romanov, R.A.; Rogachevskaja, O.A.; Bystrova, M.F.; Kolesnikov, S.S. Expression of calcium-activated chloride channels Ano1 and Ano2 in mouse taste cells. Pflugers Arch-Eur. J. Physiol. 2016, 468, 305–319. [Google Scholar] [CrossRef] [PubMed]
- Caicedo, A.; Jafri, M.S.; Roper, S.D. In Situ Ca 2+ Imaging Reveals Neurotransmitter Receptors for Glutamate in Taste Receptor Cells. J. Neurosci. 2000, 20, 7978–7985. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Maruyama, Y.; Pereira, E.; Margolskee, R.F.; Chaudhari, N.; Roper, S.D. Umami responses in mouse taste cells indicate more than one receptor. J. Neurosci. 2006, 26, 2227–2234. [Google Scholar] [CrossRef]
- Caicedo, A.; Pereira, E.; Margolskee, R.F.; Roper, S.D. Role of the G-Protein Subunit α-Gustducin in Taste Cell Responses to Bitter Stimuli. J. Neurosci. 2003, 23, 9947–9952. [Google Scholar] [CrossRef]
- Richter, T.A.; Dvoryanchikov, G.A.; Chaudhari, N.; Roper, S.D. Acid-sensitive two-pore domain potassium (K2P) channels in mouse taste buds. J. Neurophysiol. 2004, 92, 1928–1936. [Google Scholar] [CrossRef]
- Caicedo, A.; Kim, K.-N.; Roper, S.D. Individual mouse taste cells respond to multiple chemical stimuli. J. Physiol. (Lond) 2002, 544, 501–509. [Google Scholar] [CrossRef]
- Oheim, M.; Van ‘t Hoff, M.; Feltz, A.; Zamaleeva, A.; Mallet, J.-M.; Collot, M. New red-fluorescent calcium indicators for optogenetics, photoactivation and multi-color imaging. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2014, 1843, 2284–2306. [Google Scholar] [CrossRef]
- Boens, N.; Qin, W.; Basarić, N.; Orte, A.; Talavera, E.M.; Alvarez-Pez, J.M. Photophysics of the fluorescent pH indicator BCECF. J. Phys. Chem. A 2006, 110, 9334–9343. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Stewart, R.E.; Lyall, V.; Feldman, G.M.; Heck, G.L.; DeSimone, J.A. Acid-induced responses in hamster chorda tympani and intracellular pH tracking by taste receptor cells. Am. J. Physiol. 1998, 275, C227–C238. [Google Scholar] [CrossRef] [PubMed]
- Trubey, K.R.; Culpepper, S.; Maruyama, Y.; Kinnamon, S.C.; Chaudhari, N. Tastants evoke cAMP signal in taste buds that is independent of calcium signaling. Am. J. Physiol. Cell Physiol. 2006, 291, C237–C244. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Simon, S.A. Acidic stimuli activates two distinct pathways in taste receptor cells from rat fungiform papillae. Brain Res. 2001, 923, 58–70. [Google Scholar] [CrossRef]
- Hayato, R.; Ohtubo, Y.; Yoshii, K. Functional expression of ionotropic purinergic receptors on mouse taste bud cells. J. Physiol. (Lond) 2007, 584, 473–488. [Google Scholar] [CrossRef]
- Rebello, M.R.; Maliphol, A.B.; Medler, K.F. Ryanodine Receptors Selectively Interact with L Type Calcium Channels in Mouse Taste Cells. PLoS ONE 2013, 8, e68174. [Google Scholar] [CrossRef][Green Version]
- Rebello, M.R.; Maliphol A., B.; Medler, K.F. Ryanodine Receptors Selectively Interact with L Type Calcium Channels in Mouse Taste Cells. PLoS ONE 2013, 8, e68174. [Google Scholar] [CrossRef][Green Version]
- Laskowski, A.I.; Medler, K.F. Sodium-calcium exchangers contribute to the regulation of cytosolic calcium levels in mouse taste cells. J. Physiol. (Lond) 2009, 587, 4077–4089. [Google Scholar] [CrossRef]
- Ogura, T.; Margolskee, R.F.; Kinnamon, S.C. Taste receptor cell responses to the bitter stimulus denatonium involve Ca2+ influx via store-operated channels. J. Neurophysiol. 2002, 87, 3152–3155. [Google Scholar] [CrossRef]
- Kudo, K.-i.; Kawabata, F.; Nomura, T.; Aridome, A.; Nishimura, S.; Tabata, S. Isolation of chicken taste buds for real-time Ca2+ imaging. Anim. Sci. J. 2014, 85, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Whitaker, M. Genetically encoded probes for measurement of intracellular calcium. Methods Cell Biol. 2010, 99, 153–182. [Google Scholar] [CrossRef] [PubMed]
- Mank, M.; Griesbeck, O. Genetically encoded calcium indicators. Chem. Rev. 2008, 108, 1550–1564. [Google Scholar] [CrossRef] [PubMed]
- Zhong, C.; Schleifenbaum, J. Genetically Encoded Calcium Indicators: A New Tool in Renal Hypertension Research. Front. Med. (Lausanne) 2019, 6, 128. [Google Scholar] [CrossRef] [PubMed]
- Pérez Koldenkova, V.; Nagai, T. Genetically encoded Ca2+ indicators: Properties and evaluation. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2013, 1833, 1787–1797. [Google Scholar] [CrossRef]
- SHIMOMURA, O.; JOHNSON, F.H.; SAIGA, Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J. Cell. Comp. Physiol. 1962, 59, 223–239. [Google Scholar] [CrossRef]
- Chalfie, M.; Tu, Y.; Euskirchen, G.; Ward, W.W.; Prasher, D.C. Green fluorescent protein as a marker for gene expression. Science 1994, 263, 802–805. [Google Scholar] [CrossRef]
- Von Molitor, E.; Nürnberg, E.; Ertongur-Fauth, T.; Scholz, P.; Riedel, K.; Hafner, M.; Rudolf, R.; Cesetti, T. Analysis of Calcium Signaling in Live Human Tongue Cell 3D-Cultures upon Tastant Perfusion. Cell Calcium 2020, 102164. [Google Scholar] [CrossRef]
- Kim, Y.S.; Chu, Y.; Han, L.; Li, M.; Li, Z.; LaVinka, P.C.; Sun, S.; Tang, Z.; Park, K.; Caterina, M.J.; et al. Central Terminal Sensitization of TRPV1 by Descending Serotonergic Facilitation Modulates Chronic Pain. Neuron 2014, 81, 873–887. [Google Scholar] [CrossRef]
- Kim, Y.S.; Anderson, M.; Park, K.; Zheng, Q.; Agarwal, A.; Gong, C.; Saijilafu; Young, L.; He, S.; LaVinka, P.C.; et al. Coupled Activation of Primary Sensory Neurons Contributes to Chronic Pain. Neuron 2016, 91, 1085–1096. [Google Scholar] [CrossRef]
- Rink, T.J.; Tsien, R.Y.; Pozzan, T. Cytoplasmic pH and free Mg2+ in lymphocytes. J. Cell Biol. 1982, 95, 189–196. [Google Scholar] [CrossRef]
- Lyall, V.; Alam, R.I.; Phan, T.-H.T.; Russell, O.F.; Malik, S.A.; Heck, G.L.; Desimone, J.A. Modulation of Rat Chorda Tympani NaCl Responses and Intracellular Na+ Activity in Polarized Taste Receptor Cells by pH. J. Gen. Physiol. 2002, 120, 793–815. [Google Scholar] [CrossRef] [PubMed]
- Bachmanov, A.A.; Tordoff, M.G.; Beauchamp, G.K. Ethanol Consumption and Taste Preferences in C57BL/6ByJ and 129/J Mice. Alcohol. Clin. Exp. Res. 1996, 20, 201–206. [Google Scholar] [CrossRef] [PubMed]
- Tu, Y.-H.; Cooper, A.J.; Teng, B.; Chang, R.B.; Artiga, D.J.; Turner, H.N.; Mulhall, E.M.; Ye, W.; Smith, A.D.; Liman, E.R. An evolutionarily conserved gene family encodes proton-selective ion channels. Science 2018, 359, 1047–1050. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, R.B.; Norgren, R. Central projections of gustatory nerves in the rat. J. Comp. Neurol. 1984, 222, 560–577. [Google Scholar] [CrossRef]
- Ohla, K.; Yoshida, R.; Roper, S.D.; Di Lorenzo, P.M.; Victor, J.D.; Boughter, J.D.; Fletcher, M.; Katz, D.B.; Chaudhari, N. Recognizing Taste: Coding Patterns Along the Neural Axis in Mammals. Chem. Senses 2019, 44, 237–247. [Google Scholar] [CrossRef]
- Gutierrez, R.; Simon, S.A. Chemosensory processing in the taste-reward pathway. Flavour Fragr. J. 2011, 26, 231–238. [Google Scholar] [CrossRef][Green Version]
- Carleton, A.; Accolla, R.; Simon, S.A. Coding in the mammalian gustatory system. Trends Neurosci. 2010, 33, 326–334. [Google Scholar] [CrossRef]
- Lemon, C.H.; Katz, D.B. The neural processing of taste. BMC Neurosci. 2007, 8 (Suppl. 3), S5. [Google Scholar] [CrossRef]
- Simon, S.A.; Araujo, I.E.d.; Gutierrez, R.; Nicolelis, M.A.L. The neural mechanisms of gustation: A distributed processing code. Nat. Rev. Neurosci. 2006, 7, 890–901. [Google Scholar] [CrossRef]
- Fletcher, M.L.; Ogg, M.C.; Lu, L.; Ogg, R.J.; Boughter, J.D. Overlapping Representation of Primary Tastes in a Defined Region of the Gustatory Cortex. J. Neurosci. 2017, 37, 7595–7605. [Google Scholar] [CrossRef]
- Chen, X.; Gabitto, M.; Peng, Y.; Ryba, N.J.P.; Zuker, C.S. A gustotopic map of taste qualities in the mammalian brain. Science 2011, 333, 1262–1266. [Google Scholar] [CrossRef] [PubMed]
- Prinster, A.; Cantone, E.; Verlezza, V.; Magliulo, M.; Sarnelli, G.; Iengo, M.; Cuomo, R.; Di Salle, F.; Esposito, F. Cortical representation of different taste modalities on the gustatory cortex: A pilot study. PLoS ONE 2017, 12, e0190164. [Google Scholar] [CrossRef] [PubMed]
- Chikazoe, J.; Lee, D.H.; Kriegeskorte, N.; Anderson, A.K. Distinct representations of basic taste qualities in human gustatory cortex. Nat. Commun. 2019, 10, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Yoshimura, H.; Sugai, T.; Fukuda, M.; Segami, N.; Onoda, N. Cortical spatial aspects of optical intrinsic signals in response to sucrose and NaCl stimuli. Neuroreport 2004, 15, 17–20. [Google Scholar] [CrossRef]
- Accolla, R.; Bathellier, B.; Petersen, C.C.H.; Carleton, A. Differential spatial representation of taste modalities in the rat gustatory cortex. J. Neurosci. 2007, 27, 1396–1404. [Google Scholar] [CrossRef]
- Ali, F.; Kwan, A.C. Interpreting in vivo calcium signals from neuronal cell bodies, axons, and dendrites: A review. Neurophotonics 2020, 7, 11402. [Google Scholar] [CrossRef]
- Lentz, T.B.; Gray, S.J.; Samulski, R.J. Viral vectors for gene delivery to the central nervous system. Neurobiol. Dis. 2012, 48, 179–188. [Google Scholar] [CrossRef]
- Hammond, S.L.; Leek, A.N.; Richman, E.H.; Tjalkens, R.B. Cellular selectivity of AAV serotypes for gene delivery in neurons and astrocytes by neonatal intracerebroventricular injection. PLoS ONE 2017, 12, e0188830. [Google Scholar] [CrossRef]
- Barretto, R.P.J.; Gillis-Smith, S.; Chandrashekar, J.; Yarmolinsky, D.A.; Schnitzer, M.J.; Ryba, N.J.P.; Zuker, C.S. The neural representation of taste quality at the periphery. Nature 2015, 517, 373–376. [Google Scholar] [CrossRef]
- Chen, T.-W.; Wardill, T.J.; Sun, Y.; Pulver, S.R.; Renninger, S.L.; Baohan, A.; Schreiter, E.R.; Kerr, R.A.; Orger, M.B.; Jayaraman, V.; et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 2013, 499, 295–300. [Google Scholar] [CrossRef]
- Sahu, P.; Mazumder, N. Advances in adaptive optics–based two-photon fluorescence microscopy for brain imaging. Lasers Med. Sci. 2020, 35, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Lecoq, J.; Orlova, N.; Grewe, B.F. Wide. Fast. Deep: Recent Advances in Multiphoton Microscopy of In Vivo Neuronal Activity. J. Neurosci. 2019, 39, 9042–9052. [Google Scholar] [CrossRef] [PubMed]
- Wu, A.; Dvoryanchikov, G.; Pereira, E.; Chaudhari, N.; Roper, S.D. Breadth of tuning in taste afferent neurons varies with stimulus strength. Nat. Commun. 2015, 6, 8171. [Google Scholar] [CrossRef]
- Sugita, M.; Yamamoto, K.; Hirono, C.; Shiba, Y. Information processing in brainstem bitter taste-relaying neurons defined by genetic tracing. Neuroscience 2013, 250, 166–180. [Google Scholar] [CrossRef] [PubMed]
- Lavi, K.; Jacobson, G.A.; Rosenblum, K.; Lüthi, A. Encoding of Conditioned Taste Aversion in Cortico-Amygdala Circuits. Cell Rep. 2018, 24, 278–283. [Google Scholar] [CrossRef] [PubMed]
- Fu, O.; Iwai, Y.; Kondoh, K.; Misaka, T.; Minokoshi, Y.; Nakajima, K.-I. SatB2-Expressing Neurons in the Parabrachial Nucleus Encode Sweet Taste. Cell Rep. 2019, 27, 1650–1656.e4. [Google Scholar] [CrossRef]
- Susaki, E.A.; Tainaka, K.; Perrin, D.; Yukinaga, H.; Kuno, A.; Ueda, H.R. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat. Protoc. 2015, 10, 1709–1727. [Google Scholar] [CrossRef]
- Lemon, C.H. Perceptual and Neural Responses to Sweet Taste in Humans and Rodents. Chem. Percept. 2015, 8, 46–52. [Google Scholar] [CrossRef]
- Rolls, E.T. Brain mechanisms underlying flavour and appetite. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2006, 361, 1123–1136. [Google Scholar] [CrossRef]
- Kobayashi, M. Functional Organization of the Human Gustatory Cortex. J. Oral Biosci. 2006, 48, 244–260. [Google Scholar] [CrossRef]
- Vendrell-Llopis, N.; Yaksi, E. Evolutionary conserved brainstem circuits encode category, concentration and mixtures of taste. Sci. Rep. 2015, 5, 17825. [Google Scholar] [CrossRef]
- Vandenbeuch, A.; Kinnamon, S.C. Why do taste cells generate action potentials? J. Biol. 2009, 8, 42. [Google Scholar] [CrossRef] [PubMed]
- Roper, S. Regenerative impulses in taste cells. Science 1983, 220, 1311–1312. [Google Scholar] [CrossRef]
- Kinnamon, S.C.; Roper, S.D. Passive and active membrane properties of mudpuppy taste receptor cells. J. Physiol. (Lond) 1987, 383, 601–614. [Google Scholar] [CrossRef] [PubMed]
- Cummings, T.A.; Daniels, C.; Kinnamon, S.C. Sweet taste transduction in hamster: Sweeteners and cyclic nucleotides depolarize taste cells by reducing a K+ current. J. Neurophysiol. 1996, 75, 1256–1263. [Google Scholar] [CrossRef] [PubMed]
- Avenet, P.; Lindemann, B. Noninvasive recording of receptor cell action potentials and sustained currents from single taste buds maintained in the tongue: The response to mucosal NaCl and amiloride. J. Membrain Biol. 1991, 124, 33–41. [Google Scholar] [CrossRef]
- Furue, H.; Yoshii, K. In situ tight-seal recordings of taste substance-elicited action currents and voltage-gated Ba currents from single taste bud cells in the peeled epithelium of mouse tongue. Brain Res. 1997, 776, 133–139. [Google Scholar] [CrossRef]
- Chen, Y.; Sun, X.D.; Herness, S. Characteristics of action potentials and their underlying outward currents in rat taste receptor cells. J. Neurophysiol. 1996, 75, 820–831. [Google Scholar] [CrossRef]
- Herness, M.S.; Sun, X.-D. Voltage-dependent sodium currents recorded from dissociated rat taste cells. J. Membarin Biol. 1995, 146, 73–84. [Google Scholar] [CrossRef]
- Noguchi, T.; Ikeda, Y.; Miyajima, M.; Yoshii, K. Voltage-gated channels involved in taste responses and characterizing taste bud cells in mouse soft palates. Brain Res. 2003, 982, 241–259. [Google Scholar] [CrossRef]
- Varkevisser, B.; Peterson, D.; Ogura, T.; Kinnamon, S.C. Neural networks distinguish between taste qualities based on receptor cell population responses. Chem Senses 2001, 26, 499–505. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yoshida, R.; Shigemura, N.; Sanematsu, K.; Yasumatsu, K.; Ishizuka, S.; Ninomiya, Y. Taste responsiveness of fungiform taste cells with action potentials. J. Neurophysiol. 2006, 96, 3088–3095. [Google Scholar] [CrossRef]
- DAVILA, H.V.; SALZBERG, B.M.; COHEN, L.B.; WAGGONER, A.S. A Large Change in Axon Fluorescence that Provides a Promising Method for Measuring Membrane Potential. Nat. New Biol. 1973, 241, 159–160. [Google Scholar] [CrossRef]
- Xu, Y.; Zou, P.; Cohen, A.E. Voltage imaging with genetically encoded indicators. Curr. Opin. Chem. Biol. 2017, 39, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Antic, S.D.; Empson, R.M.; Knöpfel, T. Voltage imaging to understand connections and functions of neuronal circuits. J. Neurophysiol. 2016, 116, 135–152. [Google Scholar] [CrossRef] [PubMed]
- Preuss, S.; Stein, W. Comparison of Two Voltage-Sensitive Dyes and Their Suitability for Long-Term Imaging of Neuronal Activity. PLoS ONE 2013, 8. [Google Scholar] [CrossRef]
- Fluhler, E.; Burnham, V.G.; Loew, L.M. Spectra, membrane binding, and potentiometric responses of new charge shift probes. Biochemistry 1985, 24, 5749–5755. [Google Scholar] [CrossRef]
- Loew, L.M.; Simpson, L.L. Charge-shift probes of membrane potential: A probable electrochromic mechanism for p-aminostyrylpyridinium probes on a hemispherical lipid bilayer. Biophys. J. 1981, 34, 353–365. [Google Scholar] [CrossRef]
- Li, J.H.-Y.; Lindemann, B. Multi-photon microscopy of cell types in the viable taste disk of the frog. Cell Tissue Res. 2003, 313, 11–27. [Google Scholar] [CrossRef]
- Loew, L.M. Potentiometric dyes: Imaging electrical activity of cell membranes. Pure Appl. Chem. 1996, 68, 1405–1409. [Google Scholar] [CrossRef]
- Bachtel, A.D.; Gray, R.A.; Stohlman, J.M.; Bourgeois, E.B.; Pollard, A.E.; Rogers, J.M. A novel approach to dual excitation ratiometric optical mapping of cardiac action potentials with di-4-ANEPPS using pulsed LED excitation. IEEE Trans. Biomed. Eng. 2011, 58, 2120–2126. [Google Scholar] [CrossRef] [PubMed]
- Vitha, M.F.; Clarke, R.J. Comparison of excitation and emission ratiometric fluorescence methods for quantifying the membrane dipole potential. Biochim. Biophys. Acta (BBA) Biomembr. 2007, 1768, 107–114. [Google Scholar] [CrossRef]
- Canepari, M.; Vogt, K.; Zecevic, D. Combining Voltage and Calcium Imaging from Neuronal Dendrites. Cell Mol. Neurobiol. 2008, 28, 1079–1093. [Google Scholar] [CrossRef] [PubMed]
- Ehrenberg, B.; Montana, V.; Wei, M.D.; Wuskell, J.P.; Loew, L.M. Membrane potential can be determined in individual cells from the nernstian distribution of cationic dyes. Biophys. J. 1988, 53, 785–794. [Google Scholar] [CrossRef]
- Scaduto, R.C.; Grotyohann, L.W. Measurement of Mitochondrial Membrane Potential Using Fluorescent Rhodamine Derivatives. Biophys. J. 1999, 76, 469–477. [Google Scholar] [CrossRef]
- Ohtubo, Y.; Suemitsu, T.; Shiobara, S.; Matsumoto, T.; Kumazawa, T.; Yoshii, K.Y. Optical recordings of taste responses from fungiform papillae of mouse in situ. J. Physiol. (Lond) 2001, 530, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.H.; St-Pierre, F.; Sun, X.; Ding, X.; Lin, M.Z.; Clandinin, T.R. Subcellular Imaging of Voltage and Calcium Signals Reveals Neural Processing In Vivo. Cell 2016, 166, 245–257. [Google Scholar] [CrossRef]
- Bando, Y.; Grimm, C.; Cornejo, V.H.; Yuste, R. Genetic voltage indicators. BMC Biol. 2019, 17, 1–12. [Google Scholar] [CrossRef]
- Abdelfattah, A.S.; Kawashima, T.; Singh, A.; Novak, O.; Liu, H.; Shuai, Y.; Huang, Y.-C.; Campagnola, L.; Seeman, S.C.; Yu, J.; et al. Bright and photostable chemigenetic indicators for extended in vivo voltage imaging. Science 2019, 365, 699–704. [Google Scholar] [CrossRef] [PubMed]
- Abdelfattah, A.S.; Farhi, S.L.; Zhao, Y.; Brinks, D.; Zou, P.; Ruangkittisakul, A.; Platisa, J.; Pieribone, V.A.; Ballanyi, K.; Cohen, A.E.; et al. A Bright and Fast Red Fluorescent Protein Voltage Indicator That Reports Neuronal Activity in Organotypic Brain Slices. J. Neurosci. 2016, 36, 2458–2472. [Google Scholar] [CrossRef]
- Quicke, P.; Song, C.; McKimm, E.J.; Milosevic, M.M.; Howe, C.L.; Neil, M.; Schultz, S.R.; Antic, S.D.; Foust, A.J.; Knöpfel, T. Single-Neuron Level One-Photon Voltage Imaging With Sparsely Targeted Genetically Encoded Voltage Indicators. Front. Cell. Neurosci. 2019, 13, 39. [Google Scholar] [CrossRef] [PubMed]
- Beck, C.; Gong, Y. A high-speed, bright, red fluorescent voltage sensor to detect neural activity. Sci. Rep. 2019, 9, 15878. [Google Scholar] [CrossRef] [PubMed]
- Sugai, T.; Yamamoto, R.; Yoshimura, H.; Kato, N. Multimodal Cross-Talk of Olfactory and Gustatory Information in the Endopiriform Nucleus in Rats. Chem Senses 2012, 37, 681–688. [Google Scholar] [CrossRef] [PubMed]
- Margolskee, R.F. The biochemistry and molecular biology of taste transduction. Curr. Opin. Neurobiol. 1993, 3, 526–531. [Google Scholar] [CrossRef]
- Eric Walters, D. How are bitter and sweet tastes related? Trends Food Sci. Technol. 1996, 7, 399–403. [Google Scholar] [CrossRef]
- Kolesnikov, S.S.; Margolskee, R.F. A cyclic-nucleotide-suppressible conductance activated by transducin in taste cells. Nature 1995, 376, 85–88. [Google Scholar] [CrossRef]
- Misaka, T.; Kusakabe, Y.; Emori, Y.; Gonoi, T.; Arai, S.; Abe, K. Taste buds have a cyclic nucleotide-activated channel, CNGgust. J. Biol. Chem. 1997, 272, 22623–22629. [Google Scholar] [CrossRef]
- Adams, S.R.; Harootunian, A.T.; Buechler, Y.J.; Taylor, S.S.; Tsien, R.Y. Fluorescence ratio imaging of cyclic AMP in single cells. Nature 1991, 349, 694–697. [Google Scholar] [CrossRef]
- Patel, N.; Gold, M.G. The genetically encoded tool set for investigating cAMP: More than the sum of its parts. Front. Pharmacol. 2015, 6, 164. [Google Scholar] [CrossRef]
- Zaccolo, M.; Giorgi, F.D.; Cho, C.Y.; Feng, L.; Knapp, T.; Negulescu, P.A.; Taylor, S.S.; Tsien, R.Y.; Pozzan, T. A genetically encoded, fluorescent indicator for cyclic AMP in living cells. Nat. Cell Biol. 2000, 2, 25–29. [Google Scholar] [CrossRef]
- Zaccolo, M. Use of chimeric fluorescent proteins and fluorescence resonance energy transfer to monitor cellular responses. Circ. Res. 2004, 94, 866–873. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.Y.; Falcone, J.L.; Curci, S.; Hofer, A.M. Interrogating cyclic AMP signaling using optical approaches. Cell Calcium 2017, 64, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Zaccolo, M.; Musheshe, N.; Lobo, M.J.; Schmidt, M. Targeting FRET-based reporters for cAMP and PKA activity using AKAP79. Sensors 2018, 18, E2164. [Google Scholar]
- Rudolf, R.; Magalhães, P.J.; Pozzan, T. Direct in vivo monitoring of sarcoplasmic reticulum Ca2+ and cytosolic cAMP dynamics in mouse skeletal muscle. J Cell Biol 2006, 173, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Buck, M.A.; Fraser, C.M. Muscarinic acetylcholine receptor subtypes which selectively couple to phospholipase C: Pharmacological and biochemical properties. Biochem. Biophys. Res. Commun. 1990, 173, 666–672. [Google Scholar] [CrossRef]
- Bhuckory, S.; Kays, J.C.; Dennis, A.M. In Vivo Biosensing Using Resonance Energy Transfer. Biosensors (Basel) 2019, 9, 76. [Google Scholar] [CrossRef]
- Ueda, T.; Ugawa, S.; Yamamura, H.; Imaizumi, Y.; Shimada, S. Functional Interaction between T2R Taste Receptors and G-Protein α Subunits Expressed in Taste Receptor Cells. J. Neurosci. 2003, 23, 7376–7380. [Google Scholar] [CrossRef]
- Ishii, S.; Misaka, T.; Kishi, M.; Kaga, T.; Ishimaru, Y.; Abe, K. Acetic acid activates PKD1L3–PKD2L1 channel—A candidate sour taste receptor. Biochem. Biophys. Res. Commun. 2009, 385, 346–350. [Google Scholar] [CrossRef]
- Riera, C.E.; Vogel, H.; Simon, S.A.; Le Coutre, J. Artificial sweeteners and salts producing a metallic taste sensation activate TRPV1 receptors. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2007, 293, R626–R634. [Google Scholar] [CrossRef]
- Pérez, C.A.; Huang, L.; Rong, M.; Kozak, J.A.; Preuss, A.K.; Zhang, H.; Max, M.; Margolskee, R.F. A transient receptor potential channel expressed in taste receptor cells. Nat. Neurosci. 2002, 5, 1169–1176. [Google Scholar] [CrossRef]
- Medina, J.; Nakagawa, Y.; Nagasawa, M.; Fernandez, A.; Sakaguchi, K.; Kitaguchi, T.; Kojima, I. Positive Allosteric Modulation of the Calcium-sensing Receptor by Physiological Concentrations of Glucose. J. Biol. Chem. 2016, 291, 23126–23135. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, C.; Ishimaru, Y.; Katano, Y.; Misaka, T.; Yamasoba, T.; Asakura, T.; Abe, K. The single pore residue Asp523 in PKD2L1 determines Ca2+ permeation of the PKD1L3/PKD2L1 complex. Biochem. Biophys. Res. Commun. 2011, 404, 946–951. [Google Scholar] [CrossRef] [PubMed]
- Brockhoff, A.; Behrens, M.; Niv, M.Y.; Meyerhof, W. Structural requirements of bitter taste receptor activation. Proc. Natl. Acad. Sci. USA 2010, 107, 11110–11115. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Cui, M.; Zhao, B.; Snyder, L.A.; Benard, L.M.J.; Osman, R.; Max, M.; Margolskee, R.F. Identification of the cyclamate interaction site within the transmembrane domain of the human sweet taste receptor subunit T1R3. J. Biol. Chem. 2005, 280, 34296–34305. [Google Scholar] [CrossRef] [PubMed]
- Gee, K.R.; Brown, K.A.; Chen, W.-N.U.; Bishop-Stewart, J.; Gray, D.; Johnson, I. Chemical and physiological characterization of fluo-4 Ca2+-indicator dyes. Cell Calcium 2000, 27, 97–106. [Google Scholar] [CrossRef]
- Offermanns, S.; Simon, M.I. G alpha 15 and G alpha 16 couple a wide variety of receptors to phospholipase C. J. Biol. Chem. 1995, 270, 15175–15180. [Google Scholar] [CrossRef]
- Krautwurst, D.; Yau, K.-W.; Reed, R.R. Identification of Ligands for Olfactory Receptors by Functional Expression of a Receptor Library. Cell 1998, 95, 917–926. [Google Scholar] [CrossRef]
- Brockhoff, A.; Behrens, M.; Roudnitzky, N.; Appendino, G.; Avonto, C.; Meyerhof, W. Receptor agonism and antagonism of dietary bitter compounds. J. Neurosci. 2011, 31, 14775–14782. [Google Scholar] [CrossRef]
- Behrens, M.; Blank, K.; Meyerhof, W. Blends of Non-caloric Sweeteners Saccharin and Cyclamate Show Reduced Off-Taste due to TAS2R Bitter Receptor Inhibition. Cell Chem. Biol. 2017, 24, 1199–1204.e2. [Google Scholar] [CrossRef]
- McLaughlin, S.K.; McKinnon, P.J.; Margolskee, R.F. Gustducin is a taste-cell-specific G protein closely related to the transducins. Nature 1992, 357, 563–569. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, S.K.; McKinnon, P.J.; Robichon, A.; Spickofsky, N.; Margolskee, R.F. Gustducin and transducin: A tale of two G proteins. Ciba Found. Symp. 1993, 179, 186–200. [Google Scholar] [CrossRef] [PubMed]
- Kusakabe, Y.; Yasuoka, A.; Asano-Miyoshi, M.; Iwabuchi, K.; Matsumoto, I.; Arai, S.; Emori, Y.; Abe, K. Comprehensive study on G protein alpha-subunits in taste bud cells, with special reference to the occurrence of Galphai2 as a major Galpha species. Chem. Senses 2000, 25, 525–531. [Google Scholar] [CrossRef] [PubMed]
- Tizzano, M.; Dvoryanchikov, G.; Barrows, J.K.; Kim, S.; Chaudhari, N.; Finger, T.E. Expression of Galpha14 in sweet-transducing taste cells of the posterior tongue. BMC Neurosci. 2008, 9, 110. [Google Scholar] [CrossRef] [PubMed]
- Soranzo, N.; Bufe, B.; Sabeti, P.C.; Wilson, J.F.; Weale, M.E.; Marguerie, R.; Meyerhof, W.; Goldstein, D.B. Positive Selection on a High-Sensitivity Allele of the Human Bitter-Taste Receptor TAS2R16. Curr. Biol. 2005, 15, 1257–1265. [Google Scholar] [CrossRef]
- Bolsover, S.; Ibrahim, O.; O’luanaigh, N.; Williams, H.; Cockcroft, S. Use of fluorescent Ca2+ dyes with green fluorescent protein and its variants: Problems and solutions. Biochem. J. 2001, 356, 345–352. [Google Scholar] [CrossRef]
- Schreder, B.; Lukacs, Z.; Schmitt, M.; Schreier, P.; Humpf, H.-U. 1-Naphthoic acid: A new type of asymmetric chromophore for exciton-coupled circular dichroism (ECCD). Tetrahedron Asymmetry 1996, 7, 1543–1546. [Google Scholar] [CrossRef]
- Slack, J.P.; Brockhoff, A.; Batram, C.; Menzel, S.; Sonnabend, C.; Born, S.; Galindo, M.M.; Kohl, S.; Thalmann, S.; Ostopovici-Halip, L.; et al. Modulation of bitter taste perception by a small molecule hTAS2R antagonist. Curr. Biol. 2010, 20, 1104–1109. [Google Scholar] [CrossRef]
- Servant, G.; Brust, P.; Ray, S.; Hung, N. Cell-Based Fluorescent Assays for Identifying Alpha and Delta ENaC Modulators. Google Patents US20090181404A1, 16 July 2009. [Google Scholar]
- Callamaras, N.; Chang, H. High throughput cell-based assay for monitoring sodium channel activity and discovery of salty taste modulating compounds. Google Patents AU 2002308481A8, 1 February 2007. [Google Scholar]
- Landin, A.M.; Kim, J.W.; Chaudhari, N. Liposome-mediated transfection of mature taste cells. J. Neurobiol. 2005, 65, 12–21. [Google Scholar] [CrossRef]
- Kishi, M.; Emori, Y.; Tsukamoto, Y.; Abe, K. Primary culture of rat taste bud cells that retain molecular markers for taste buds and permit functional expression of foreign genes. Neuroscience 2001, 106, 217–225. [Google Scholar] [CrossRef]
- Cavnar, S.P.; Salomonsson, E.; Luker, K.E.; Luker, G.D.; Takayama, S. Transfer, imaging, and analysis plate for facile handling of 384 hanging drop 3D tissue spheroids. J. Lab. Autom. 2014, 19, 208–214. [Google Scholar] [CrossRef]
- Costa, E.C.; Gaspar, V.M.; Coutinho, P.; Correia, I.J. Optimization of liquid overlay technique to formulate heterogenic 3D co-cultures models. Biotechnol. Bioeng. 2014, 111, 1672–1685. [Google Scholar] [CrossRef] [PubMed]
- Gualda, E.J.; Simão, D.; Pinto, C.; Alves, P.M.; Brito, C. Imaging of human differentiated 3D neural aggregates using light sheet fluorescence microscopy. Front. Cell. Neurosci. 2014, 8, 221. [Google Scholar] [CrossRef] [PubMed]
- Foster, S.R.; Roura, E.; Thomas, W.G. Extrasensory perception: Odorant and taste receptors beyond the nose and mouth. Pharmacol. Ther. 2014, 142, 41–61. [Google Scholar] [CrossRef]
- Brockhoff, A.; Behrens, M.; Massarotti, A.; Appendino, G.; Meyerhof, W. Broad tuning of the human bitter taste receptor hTAS2R46 to various sesquiterpene lactones, clerodane and labdane diterpenoids, strychnine, and denatonium. J. Agric. Food Chem. 2007, 55, 6236–6243. [Google Scholar] [CrossRef]
- Behrens, M.; Brockhoff, A.; Kuhn, C.; Bufe, B.; Winnig, M.; Meyerhof, W. The human taste receptor hTAS2R14 responds to a variety of different bitter compounds. Biochem. Biophys. Res. Commun. 2004, 319, 479–485. [Google Scholar] [CrossRef]
- Behrens, M.; Brockhoff, A.; Batram, C.; Kuhn, C.; Appendino, G.; Meyerhof, W. The human bitter taste receptor hTAS2R50 is activated by the two natural bitter terpenoids andrographolide and amarogentin. J. Agric. Food Chem. 2009, 57, 9860–9866. [Google Scholar] [CrossRef]
- Greene, T.A.; Alarcon, S.; Thomas, A.; Berdougo, E.; Doranz, B.J.; Breslin, P.A.S.; Rucker, J.B. Probenecid Inhibits the Human Bitter Taste Receptor TAS2R16 and Suppresses Bitter Perception of Salicin. PLoS ONE 2011, 6. [Google Scholar] [CrossRef]
- Bufe, B.; Hofmann, T.; Krautwurst, D.; Raguse, J.-D.; Meyerhof, W. The human TAS2R16 receptor mediates bitter taste in response to beta-glucopyranosides. Nat. Genet. 2002, 32, 397–401. [Google Scholar] [CrossRef]
- Xu, H.; Staszewski, L.; Tang, H.; Adler, E.; Zoller, M.; Li, X. Different functional roles of T1R subunits in the heteromeric taste receptors. Proc. Natl. Acad. Sci. USA 2004, 101, 14258–14263. [Google Scholar] [CrossRef]
- Galindo-Cuspinera, V.; Winnig, M.; Bufe, B.; Meyerhof, W.; Breslin, P.A.S. A TAS1R receptor-based explanation of sweet ‘water-taste’. Nature 2006, 441, 354–357. [Google Scholar] [CrossRef]
- Haruyama, N.; Cho, A.; Kulkarni, A.B. Overview: Engineering transgenic constructs and mice. Curr. Protoc. Cell Biol. 2009, 19, 19.10.1-19.10.9. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Chen, L.-X.; Peng, X.-H.; Wang, C.; Qin, B.-Y.; Tan, D.; Han, C.-X.; Yang, H.; Ren, X.-N.; Liu, F.; et al. Overview of the reporter genes and reporter mouse models. Animal Model. Exp. Med. 2018, 1, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.W.; Roberts, C.; Maruyama, Y.; Berg, S.; Roper, S.; Chaudhari, N. Faithful expression of GFP from the PLCbeta2 promoter in a functional class of taste receptor cells. Chem. Senses 2006, 31, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyaya, B.; Di Cristo, G.; Higashiyama, H.; Knott, G.W.; Kuhlman, S.J.; Welker, E.; Huang, Z.J. Experience and Activity-Dependent Maturation of Perisomatic GABAergic Innervation in Primary Visual Cortex during a Postnatal Critical Period. J. Neurosci. 2004, 24, 9598–9611. [Google Scholar] [CrossRef] [PubMed]
- Tamura, T.-a.; Hashimoto, M.; Aruga, J.; Konishi, Y.; Nakagawa, M.; Ohbayashi, T.; Shimada, M.; Mikoshiba, K. Promoter structure and gene expression of the mouse inositol 1,4,5-trisphosphate receptor type 3 gene. Gene 2001, 275, 169–176. [Google Scholar] [CrossRef]
- Damak, S.; Mosinger, B.; Margolskee, R.F. Transsynaptic transport of wheat germ agglutinin expressed in a subset of type II taste cells of transgenic mice. BMC Neurosci. 2008, 9, 96. [Google Scholar] [CrossRef]
- Vandenbeuch, A.; Clapp, T.R.; Kinnamon, S.C. Amiloride-sensitive channels in type I fungiform taste cells in mouse. BMC Neurosci. 2008, 9, 1. [Google Scholar] [CrossRef]
- Feng, P.; Zhao, H.; Chai, J.; Huang, L.; Wang, H. Expression and secretion of TNF-α in mouse taste buds: A novel function of a specific subset of type II taste cells. PLoS ONE 2012, 7, e43140. [Google Scholar] [CrossRef]
- Parker, M.R.; Feng, D.; Chamuris, B.; Margolskee, R.F. Expression and nuclear translocation of glucocorticoid receptors in type 2 taste receptor cells. Neurosci. Lett. 2014, 571, 72–77. [Google Scholar] [CrossRef]
- Takai, S.; Yasumatsu, K.; Inoue, M.; Iwata, S.; Yoshida, R.; Shigemura, N.; Yanagawa, Y.; Drucker, D.J.; Margolskee, R.F.; Ninomiya, Y. Glucagon-like peptide-1 is specifically involved in sweet taste transmission. FASEB J. 2015, 29, 2268–2280. [Google Scholar] [CrossRef]
- Yoshida, R.; Noguchi, K.; Shigemura, N.; Jyotaki, M.; Takahashi, I.; Margolskee, R.F.; Ninomiya, Y. Leptin suppresses mouse taste cell responses to sweet compounds. Diabetes 2015, 64, 3751–3762. [Google Scholar] [CrossRef] [PubMed]
- Mueller, K.L.; Hoon, M.A.; Erlenbach, I.; Chandrashekar, J.; Zuker, C.S.; Ryba, N.J.P. The receptors and coding logic for bitter taste. Nature 2005, 434, 225–229. [Google Scholar] [CrossRef] [PubMed]
- Zapata-Hommer, O.; Griesbeck, O. Efficiently folding and circularly permuted variants of the Sapphire mutant of GFP. BMC Biotechnol. 2003, 3, 5. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, R.; Miyauchi, A.; Yasuo, T.; Jyotaki, M.; Murata, Y.; Yasumatsu, K.; Shigemura, N.; Yanagawa, Y.; Obata, K.; Ueno, H.; et al. Discrimination of taste qualities among mouse fungiform taste bud cells. J. Physiol. (Lond) 2009, 587, 4425–4439. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, R.; Takai, S.; Sanematsu, K.; Margolskee, R.F.; Shigemura, N.; Ninomiya, Y. Bitter Taste Responses of Gustducin-positive Taste Cells in Mouse Fungiform and Circumvallate Papillae. Neuroscience 2018, 369, 29–39. [Google Scholar] [CrossRef]
- Dvoryanchikov, G.; Hernandez, D.; Roebber, J.K.; Hill, D.L.; Roper, S.D.; Chaudhari, N. Transcriptomes and neurotransmitter profiles of classes of gustatory and somatosensory neurons in the geniculate ganglion. Nat. Commun. 2017, 8, 1–16. [Google Scholar] [CrossRef]
- Vandenbeuch, A.; Tizzano, M.; Anderson, C.B.; Stone, L.M.; Goldberg, D.; Kinnamon, S.C. Evidence for a role of glutamate as an efferent transmitter in taste buds. BMC Neurosci. 2010, 11, 77. [Google Scholar] [CrossRef]
- Chang, R.B.; Waters, H.; Liman, E.R. A proton current drives action potentials in genetically identified sour taste cells. Proc. Natl. Acad. Sci. USA 2010, 107, 22320–22325. [Google Scholar] [CrossRef]
- Horio, N.; Yoshida, R.; Yasumatsu, K.; Yanagawa, Y.; Ishimaru, Y.; Matsunami, H.; Ninomiya, Y. Sour taste responses in mice lacking PKD channels. PLoS ONE 2011, 6, e20007. [Google Scholar] [CrossRef]
- La Sala, M.S.; Hurtado, M.D.; Brown, A.R.; Bohórquez, D.V.; Liddle, R.A.; Herzog, H.; Zolotukhin, S.; Dotson, C.D. Modulation of taste responsiveness by the satiation hormone peptide YY. FASEB J. 2013, 27, 5022–5033. [Google Scholar] [CrossRef]
- Ping, W.; Xu, G.; Qin, L.; Xu, Y..; Li, Y.; Li, R. Cell-based biosensors and its application in biomedicine. Sens. Actuators B Chem. 2005, 108, 576–584. [Google Scholar] [CrossRef]
- Ziegler, C. Cell-based biosensors. Fresenius’ J. Anal. Chem. 2000, 366, 552–559. [Google Scholar] [CrossRef] [PubMed]
- Gui, Q.; Lawson, T.; Shan, S.; Yan, L.; Liu, Y. The Application of Whole Cell-Based Biosensors for Use in Environmental Analysis and in Medical Diagnostics. Sensors (Basel) 2017, 17, 1623. [Google Scholar] [CrossRef] [PubMed]
- Gupta, N.; Renugopalakrishnan, V.; Liepmann, D.; Paulmurugan, R.; Malhotra, B.D. Cell-based biosensors: Recent trends, challenges and future perspectives. Biosens. Bioelectron. 2019, 141, 111435. [Google Scholar] [CrossRef] [PubMed]
- Berg, K.A.; Clarke, W.P.; Sailstad, C.; Saltzman, A.; Maayani, S. Signal transduction differences between 5-hydroxytryptamine type 2A and type 2C receptor systems. Mol. Pharmacol. 1994, 46, 477–484. [Google Scholar] [PubMed]
- Akiyoshi, J.; Isogawa, K.; Yamada, K.; Nagayama, H.; Fujii, I. Effects of antidepressants on intracellular Ca2+ mobilization in CHO cells transfected with the human 5-HT2C receptors. Biol. Psychiatry 1996, 39, 1000–1008. [Google Scholar] [CrossRef]
- Kawashima, E.; Estoppey, D.; Virginio, C.; Fahmi, D.; Rees, S.; Surprenant, A.; North, R.A. A novel and efficient method for the stable expression of heteromeric ion channels in mammalian cells. Recept. Channels 1998, 5, 53–60. [Google Scholar]
- Jiao, X.; Gonzalez-Cabrera, P.J.; Xiao, L.; Bradley, M.E.; Abel, P.W.; Jeffries, W.B. Tonic inhibitory role for cAMP in alpha(1a)-adrenergic receptor coupling to extracellular signal-regulated kinases ½. J. Pharmacol. Exp. Ther. 2002, 303, 247–256. [Google Scholar] [CrossRef]
- Romanov, R.A.; Rogachevskaja, O.A.; Khokhlov, A.A.; Kolesnikov, S.S. Voltage dependence of ATP secretion in mammalian taste cells. J. Gen. Physiol. 2008, 132, 731–744. [Google Scholar] [CrossRef]
- Huang, A.Y.; Wu, S.Y. Calcitonin Gene-Related Peptide Reduces Taste-Evoked ATP Secretion from Mouse Taste Buds. J. Neurosci. 2015, 35, 12714–12724. [Google Scholar] [CrossRef]
- Dando, R.; Dvoryanchikov, G.; Pereira, E.; Chaudhari, N.; Roper, S.D. Adenosine enhances sweet taste through A2B receptors in the taste bud. J. Neurosci. 2012, 32, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Leenders, A.G.M.; Sheng, Z.-H. Modulation of neurotransmitter release by the second messenger-activated protein kinases: Implications for presynaptic plasticity. Pharmacol. Ther. 2005, 105, 69–84. [Google Scholar] [CrossRef]
- Liu, Z.; Liu, J.; Zhang, S.; Xing, X.-H.; Su, Z. Microbial fuel cell based biosensor for in situ monitoring of anaerobic digestion process. Bioresour. Technol. 2011, 102, 10221–10229. [Google Scholar] [CrossRef] [PubMed]
- HUANG, Y.-J.; MARUYAMA, Y.; LU, K.-S.; PEREIRA, E.; PLONSKY, I.; BAUR, J.E.; WU, D.; Roper, S.D. USING BIOSENSORS TO DETECT THE RELEASE OF SEROTONIN FROM TASTE BUDS DURING TASTE STIMULATION. Arch. Ital. Biol. 2005, 143, 87–96. [Google Scholar] [PubMed]
- Huang, Y.A.; Roper, S.D. Intracellular Ca(2+) and TRPM5-mediated membrane depolarization produce ATP secretion from taste receptor cells. J. Physiol. (Lond) 2010, 588, 2343–2350. [Google Scholar] [CrossRef] [PubMed]
- Glendinning, J.I.; Bloom, L.D.; Onishi, M.; Zheng, K.H.; Damak, S.; Margolskee, R.F.; Spector, A.C. Contribution of α-Gustducin to Taste-guided Licking Responses of Mice. Chem Senses 2005, 30, 299–316. [Google Scholar] [CrossRef]
- Ruiz-Avila, L.; McLaughlin, S.K.; Wildman, D.; McKinnon, P.J.; Robichon, A.; Spickofsky, N.; Margolskee, R.F. Coupling of bitter receptor to phosphodiesterase through transducin in taste receptor cells. Nature 1995, 376, 80–85. [Google Scholar] [CrossRef]
- Glendinning, J.I.; Frim, Y.G.; Hochman, A.; Lubitz, G.S.; Basile, A.J.; Sclafani, A. Glucose elicits cephalic-phase insulin release in mice by activating KATP channels in taste cells. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2017, 312, R597–R610. [Google Scholar] [CrossRef]
- Dotson, C.D.; Roper, S.D.; Spector, A.C. PLCbeta2-independent behavioral avoidance of prototypical bitter-tasting ligands. Chem Senses 2005, 30, 593–600. [Google Scholar] [CrossRef]
- Deuschle, K.; Okumoto, S.; Fehr, M.; Looger, L.L.; Kozhukh, L.; Frommer, W.B. Construction and optimization of a family of genetically encoded metabolite sensors by semirational protein engineering. Protein Sci. 2005, 14, 2304–2314. [Google Scholar] [CrossRef]
- Filadi, R.; Pozzan, T. Generation and functions of second messengers microdomains. Cell Calcium 2015, 58, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Mehta, S.; Zhang, J. Dynamic visualization of calcium-dependent signaling in cellular microdomains. Cell Calcium 2015, 58, 333–341. [Google Scholar] [CrossRef]
- Lee, J.S.; Cho, A.-N.; Jin, Y.; Kim, J.; Kim, S.; Cho, S.-W. Bio-artificial tongue with tongue extracellular matrix and primary taste cells. Biomaterials 2018, 151, 24–37. [Google Scholar] [CrossRef]
- Chessel, A.; Carazo Salas, R.E. From observing to predicting single-cell structure and function with high-throughput/high-content microscopy. Essays Biochem. 2019, 63, 197–208. [Google Scholar] [CrossRef] [PubMed]
- Booij, T.H.; Price, L.S.; Danen, E.H.J. 3D Cell-Based Assays for Drug Screens: Challenges in Imaging, Image Analysis, and High-Content Analysis. SLAS Discov. 2019, 24, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Nürnberg, E.; Vitacolonna, M.; Klicks, J.; Von Molitor, E.; Cesetti, T.; Keller, F.; Bruch, R.; Ertongur-Fauth, T.; Riedel, K.; Scholzt, P.; et al. Routine Optical Clearing of 3D-Cell Cultures: Simplicity Forward. Front. Mol. Biosci. 2020, 7, 20. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Lu, G.; Chiang, E.Y.; Chernov-Rogan, T.; Grogan, J.L.; Chen, J. High-throughput electrophysiological assays for voltage gated ion channels using SyncroPatch 768PE. PLoS ONE 2017, 12, e0180154. [Google Scholar] [CrossRef]
| Cell/Tissue | Species | Imaging | Sensor | Detect | Stimuli | Microscopy Technique | Source |
|---|---|---|---|---|---|---|---|
| Dissociated taste cells | Hamster | Ex vivo | BCECF-AM | pH | Sour | Conventional | [164] |
| Single fungiform papilla | Rat | [66] | |||||
| Slices | Mouse | BCECF-D/AM + CaO + lucifer yellow | Ca2+, pH | Confocal | [59] | ||
| Epithelium | Mouse | CaGD | Ca2+ | Salt | [46,48] | ||
| Dissociated cells, slices | Mouse | Sour, kokumi bitter, umami, K+ | [56,100,114,159] | ||||
| Slices | Rat | Sweet umami bitter, salt, sour | [74,157,161,165] | ||||
| Isolated taste buds | Rat | CaG-AM | Sour | Conventional | [166] | ||
| Isolated taste buds, slices | Mouse PLCβ2-GFP | Fura-2 CaGD/CaOD | Adenosine | [42] | |||
| Dissociated cells, isolated taste buds, slices | Mouse | GABA | Confocal | [41] | |||
| Dissociated cells | Mouse GAD-GFP | Fura-2 | Sour | Not stated | [150] | ||
| Dissociated cells, isolated taste buds | Mouse | ATP | Conventional | [140,144,167] | |||
| Dissociated cells, cell aggregates | Rat | Bitter | [50,103,133,141] | ||||
| Dissociated cells | Mouse | Bitter, umami, sweet, ryanodine | Confocal | [37,168,169,170] | |||
| Dissociated cells, isolated taste buds | Mouse PLCβ2-GFP | Glutamate | [143] | ||||
| Dissociated cells | Mouse GAD67-GFP | IBMX, Forskolin | Conventional | [149] | |||
| Dissociated cells | Mouse T1R3-GFP/TRPM5-GFP | K+ | [113] | ||||
| Dissociated cells | Mouse | Bitter, adrenergic agonist, K+ | [147] | ||||
| Dissociated cells, isolated taste buds | Mouse | Oxytocine | Not stated | [142] | |||
| Dissociated cells, isolated taste buds | Rat | Sweet, Forskolin | Conventional | [139] | |||
| Dissociated cells | Mouse/Mudpuppy | Bitter | [171] | ||||
| Dissociated cells | Mouse/Human | Fatty acid | [98] | ||||
| FACS isolated CD36 pos. cells | Mouse | Primary culture | Fura-2-AM | Fatty acid, | Confocal | [97] | |
| Primary culture of taste cells | Human | Sweet, bitter | Plate reader | [125] | |||
| Dissociated cells | Mouse | Ex vivo | Fura-2 AsanteNaTrump-2 | Ca2+, Na+ | Bitter, sweet, umami | Confocal | [151] |
| Slices | Mouse | GCaMP3 in type II and III cells | Ca2+ | Salt + AF-568,647 or fluorescein | [55] | ||
| Tongue | Mouse | In vivo | CaGD | Sweet, salt, sour, bitter | Two-photon | [77] | |
| 3D culture (organoids) | Mouse | 2D cell culture | Fura-2 | Sweet, salt, sour, bitter | Conventional | [127] | |
| Isolated taste bud | Chicken | Ex vivo | Fluo-4-AM | Bitter, umami | Confocal | [172] | |
| Dissociated cells, isolated taste buds | Mouse | Fluo-4M Np-EGTA-AM | Ca2++ uncaging | ATP | Conventional | [156] |
| Region | Species | Transgenic Model | Tracing | Sensor | Detect | Stimuli | Microscopy Technique | Source |
|---|---|---|---|---|---|---|---|---|
| Genic.gangl. | Mouse | Tr. mouse 5HT3A-GFP | Fura-2/Fluoro-Gold | Ca2+ | 5HT | Confocal | [38] | |
| Tr. mouse Pirt-GCamMP3 | GCaMP3 | Sweet, bitter, umami, salt, sour | [205] | |||||
| Tr. mouse Thy1-GCaMP3 | AVV-GCaMP3 (retrograde/NTS) | Two-photon | [201] | |||||
| NTS | Tr. mouse T2R5, tWGA-DsRed | Zif268 | Sweet, bitter | Conventional | [206] | |||
| PBN | Mouse SatB2-Cre Vglut2-ihres-Cre | AAV1-Cre-GCaMP6s | AAV8-Cre-synaptophysin-mCherry AAV5-DIO-ChR2-EYFPin PBN | GCaMP6s | Ca2+ c-fos | Bitter | Miniaturized | [208] |
| Brain stem | Zebra fish | Tr. fish Elav3-GCaMP5 | GCaMP5 | Ca2+ | Sweet, bitter, umami, sour | Two-photon | [213] | |
| GC | Mouse | AVV1-GCaMP6s | AVV1-mCherry/microruby dextran (anterograde-thalamus) | GCaMP6s | Sweet, bitter, salt, sour | [192] | ||
| AAV2/1-GCaMP6s | CAV2-Cre in hCAR x dTomato mice (retrograde-amygdala) | Bitter | [207] | |||||
| Mouse T2R5/T1R2 knockout | AVV2-GFP (anterograde-thalamus) | GCaMP6s OGB-AM + sulforhodamine 101 | Sweet, bitter, umami, salt | [202] |
| Host Cell | Stimuli | Ca2+ Sensor | Readout | Introduced Genes | Source |
|---|---|---|---|---|---|
| HEK293 | Bitter | Fura-2 | Microscopy | Gα15+T2R3-5-10-16 | [47] |
| Fura-AM | Gα16gust44/ Gα16gust37+T2R5-16 | [259] | |||
| Plate reader | Gα16gust44+T2R46 | [288] | |||
| Gα16gust44+T2R46-43-31 | [265] | ||||
| Fluo-AM | Gα16gust44+T2R14 | [289] | |||
| Gα16gust44+variants of T2R16 | [276] | ||||
| Gα16gust44+T2R43-44-4-46-50 | [290] | ||||
| Gα16gust44+T2R31 | [279] | ||||
| Gα16gust44+T2R16 | [291] | ||||
| Gα16gust44+T2R43-44 | [72] | ||||
| Gα15T2R16 | [292] | ||||
| Fluo-4 | Gα16gust44+hT2R31 | [279] | |||
| Gα16gust44+T2Rs (25 different types) | [73] | ||||
| Sweet | Fura-AM | V1R | [261] | ||
| Fluo-AM | Gα15+T1R2+T1R3 | [293] | |||
| Gα16gust44+fT1R2+T1R3 | [266,271] | ||||
| Fura-2 | Gα16gust44+T1R2/R3 or T2R44 | [294] | |||
| Microscopy | Gα15+T1R2+T1R3 | [83] | |||
| Sweet, umami | Fluo-AM | Gα15+T1R2+T1R3 Gα15+T1R1+T1R3 | [93] | ||
| Acid | Fura-2-AM | PKD1L3+PKD2L1 | [260] | ||
| [264] | |||||
| Fura-2 | [150] | ||||
| Fluo-AM | [154] | ||||
| Kokumi | Fluo-8, Flamindo (for cAMP) | CaSR | [263] |
| Promoter | Reporter Gene | Sensor | Readout | Stimuli | Source | |
|---|---|---|---|---|---|---|
| Gustducin | lacZ β–galactosidase | No Ca2+ imaging | Ca2+ | Microscopy | Bitter | [111] |
| GFP | No Ca2+ imaging | [79] | ||||
| Fura-2 | [141] | |||||
| PLCβ2 | CaOD | [297] | ||||
| KCl | [49] | |||||
| PLCβ2, GAD | Sweet, bitter, umami, sour, salt | [114] | ||||
| Sweet, bitter, umami | [33] | |||||
| Sweet, bitter, umami, ACh | [42] | |||||
| PLCβ2, GAD OXTR | Fura-2-AM | Oxytocin | [142] | |||
| T1R3 GAD | Glutamate | [311] | ||||
| IP3R | Bitter, KCl | [50] | ||||
| TRPM5 T1R3 | Fura-2-AM | KCl | [113] | |||
| TRPM5 | CFP | Fluo-5F | Ca2+ | [112] | ||
| T2R32 | GFP Sapphire | CaGD | KCl | [46] | ||
| PKD2L1 | YFP | Carboxi-DFFDA + H+ uncaging | pH | Sour | [312] | |
| Fura-2-AM pHrodo-Red-AM | pH, Ca2+ | [69] | ||||
| GAD | GFP | No Ca2+ imaging | Ca2+ | [67] | ||
| [313] | ||||||
| Fura-2 | [150] | |||||
| IBMX-forskolin | [149] | |||||
| PYY | No Ca2+ imaging | Bitter, lipids, sour, sweet, bitter, umami, salt | [314] | |||
| BC | Ca2+ Sensor | Readout | Stimuli | Receptor | Neurotransmitter | Source |
|---|---|---|---|---|---|---|
| CHO | Fura-2-AM | Microscopy | KCl, sour, sweet, bitter, ATP | 5HT2c | 5HT | [36,328] |
| 5HT2c or P2X2/P2X3 | 5HT ATP | [37,41,56,105,143] | ||||
| Adenosine | [325] | |||||
| Calcitonin gene-related peptide | NA 5-HT | [324] | ||||
| KCl, sour, sweet, bitter | 5HT2c or α1A or dual | NA 5HT | [39] | |||
| Sweet, bitter, KCl | P2X2/P2X3 | ATP | [329] | |||
| KCl, sour, sweet, bitter | GABAB+ Gαqo5 | GABA | [40] | |||
| Sweet, umami, bitter (fluorescein) | M3r P2X2/P2X3 | ACh ATP | [42] | |||
| KCl, taste mix, substance P | P GABAB+ Gαqo5 or 2X2/P2X3 | GABA, ATP | [104] | |||
| COS-1 | Fluo-4 | Bitter, sour, depolarization, ACh, 5HT, NA, glutamate | P2Y endogenous | ATP ACh | [32,323] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
von Molitor, E.; Riedel, K.; Hafner, M.; Rudolf, R.; Cesetti, T. Sensing Senses: Optical Biosensors to Study Gustation. Sensors 2020, 20, 1811. https://doi.org/10.3390/s20071811
von Molitor E, Riedel K, Hafner M, Rudolf R, Cesetti T. Sensing Senses: Optical Biosensors to Study Gustation. Sensors. 2020; 20(7):1811. https://doi.org/10.3390/s20071811
Chicago/Turabian Stylevon Molitor, Elena, Katja Riedel, Mathias Hafner, Rüdiger Rudolf, and Tiziana Cesetti. 2020. "Sensing Senses: Optical Biosensors to Study Gustation" Sensors 20, no. 7: 1811. https://doi.org/10.3390/s20071811
APA Stylevon Molitor, E., Riedel, K., Hafner, M., Rudolf, R., & Cesetti, T. (2020). Sensing Senses: Optical Biosensors to Study Gustation. Sensors, 20(7), 1811. https://doi.org/10.3390/s20071811








