Estimating Biomechanical Time-Series with Wearable Sensors: A Systematic Review of Machine Learning Techniques
Abstract
1. Introduction
1.1. Physical Models
1.2. Regression Techniques
1.3. Relevant Reviews
2. Methods
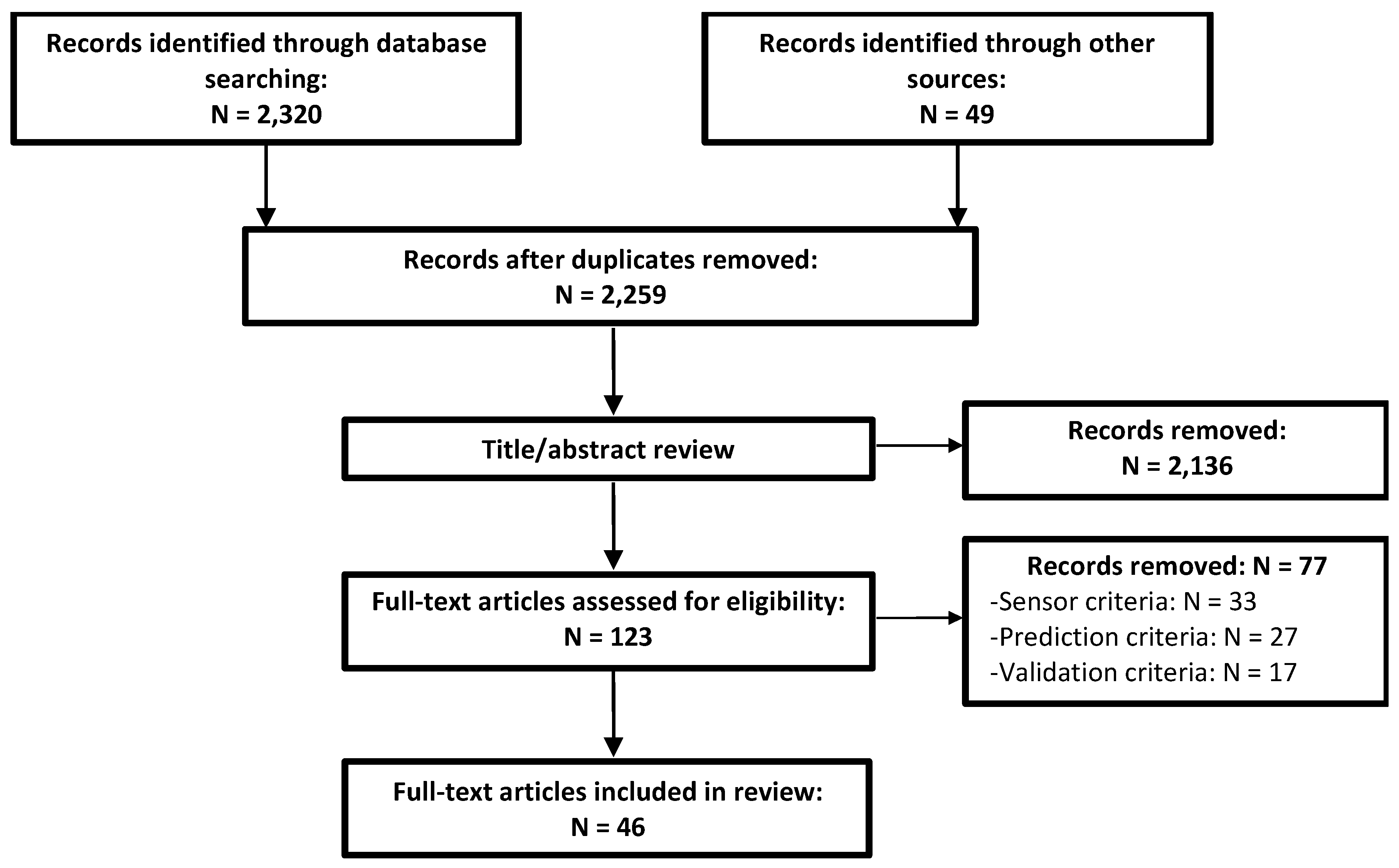
2.1. Search Strategy
2.2. Inclusion/Exclusion Criteria
- (1)
- Sensor criteria: clear use of data for estimation from a sensor that is currently deployable as a wearable. Studies investigating model inputs dependent on virtual wearable sensor data derived from a non-wearable sensor were excluded. Studies using exoskeletons were excluded if the wearable sensor is only feasibly deployed using the exoskeleton.
- (2)
- Prediction criteria: use of non-physical regression (not classification, regressed parameters must not be physical constructs). The estimated variable must have been a biomechanical time-series describing either the kinetics or kinematics of a joint, segment, or muscle. Studies were excluded if they estimated only grip or pinch forces unless the contact forces of each involved segment were estimated separately. Finally, studies estimating only ground reaction forces and moments were excluded as methods for this purpose have recently been reviewed [40].
- (3)
- Validation criteria: all studies reviewed must have reported the objective (i.e., numerical) quantification of testing error using their estimation method. Studies were excluded if they report statistics for the training error only or if the only description of performance was given graphically. Studies utilizing inappropriate validation were excluded (e.g., one that could not be repeated or one using an invalid gold standard for validation).
2.3. Data Analysis
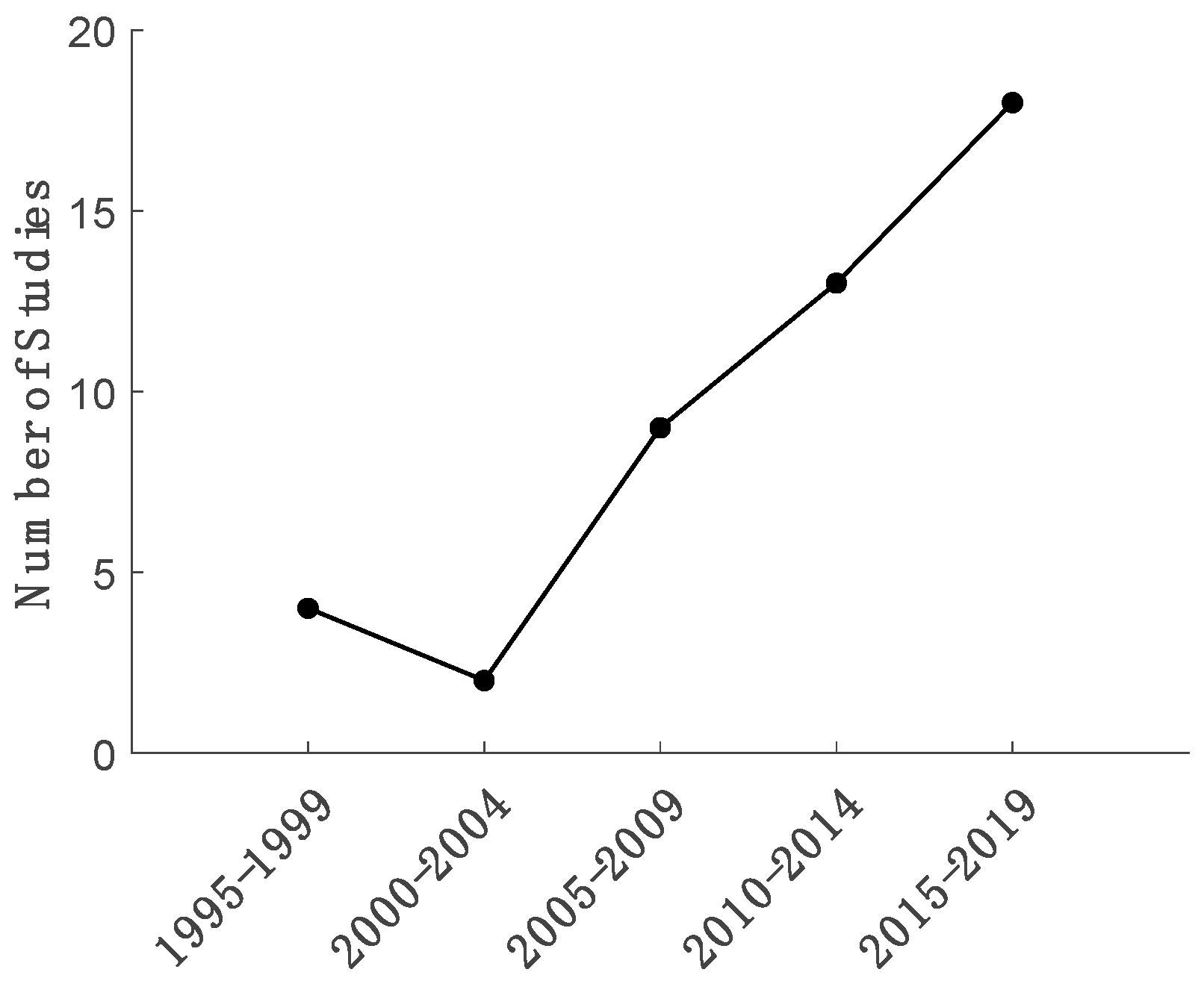
3. Results
3.1. Subject Demographics
3.2. Wearable Sensors
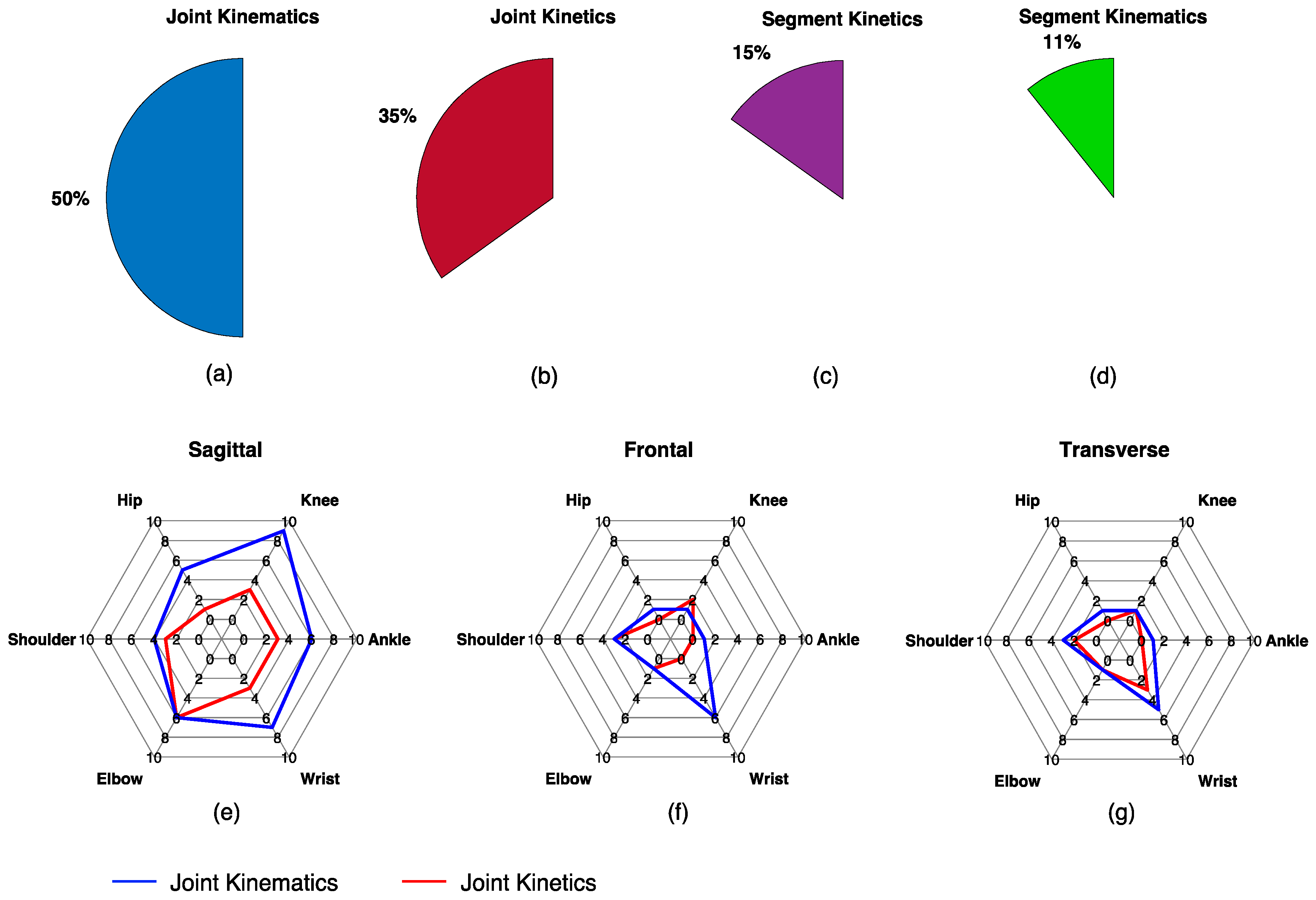
3.3. Biomechanical Variables
3.4. Prediction Equations
3.4.1. Prediction Equation Classification
3.4.2. Descriptive Statistics of Prediction Equations
4. Discussion
4.1. Overview of Techniques
4.2. Concerns for Practical Implementation
4.3. Incorporating Domain Knowledge
4.3.1. Reference to Muscle Synergies
4.3.2. Towards a Hybrid Approach
5. Conclusions
- ▪
- Development of methods using hardware specifications that can be implemented remotely and for a full 24-h capture.
- ▪
- Development of subject-general models or real-time calibration.
- ▪
- Development of hybrid machine learning and physics-based estimation.
- ▪
- Open-source algorithms.
- ▪
- Development of regression models for estimating muscle forces and joint contact forces.
- ▪
- Validation of models on impaired populations.
Author Contributions
Funding
Conflicts of Interest
References
- Coravos, A.; Khozin, S.; Mandl, K.D. Developing and adopting safe and effective digital biomarkers to improve patient outcomes. Npj Digit. Med. 2019, 2. [Google Scholar] [CrossRef] [PubMed]
- Frechette, M.L.; Meyer, B.M.; Tulipani, L.J.; Gurchiek, R.D.; McGinnis, R.S.; Sosnoff, J.J. Next Steps in Wearable Technology and Community Ambulation in Multiple Sclerosis. Curr. Neurol. Neurosci. Rep. 2019, 19, 80. [Google Scholar] [CrossRef] [PubMed]
- Espay, A.J.; Bonato, P.; Nahab, F.B.; Maetzler, W.; Dean, J.M.; Klucken, J.; Eskofier, B.M.; Merola, A.; Horak, F.; Lang, A.E.; et al. Technology in Parkinson’s disease: Challenges and opportunities: Technology in PD. Mov. Disord. 2016, 31, 1272–1282. [Google Scholar] [CrossRef] [PubMed]
- McGinnis, E.W.; McGinnis, R.S.; Muzik, M.; Hruschak, J.; Lopez-Duran, N.L.; Perkins, N.C.; Fitzgerald, K.; Rosenblum, K.L. Movements indicate threat response phases in children at-risk for anxiety. IEEE J. Biomed. Health Inform. 2017, 21, 1460–1465. [Google Scholar] [CrossRef]
- McGinnis, R.S.; McGinnis, E.W.; Hruschak, J.; Lopez-Duran, N.L.; Fitzgerald, K.; Rosenblum, K.L.; Muzik, M. Rapid detection of internalizing diagnosis in young children enabled by wearable sensors and machine learning. PLoS ONE 2019, 14, e0210267. [Google Scholar] [CrossRef]
- McGinnis, R.S.; Cain, S.M.; Tao, S.; Whiteside, D.; Goulet, G.C.; Gardner, E.C.; Bedi, A.; Perkins, N.C. Accuracy of Femur Angles Estimated by IMUs During Clinical Procedures Used to Diagnose Femoroacetabular Impingement. IEEE Trans. Biomed. Eng. 2015, 62, 1503–1513. [Google Scholar] [CrossRef]
- McGinnis, R.S.; Patel, S.; Silva, I.; Mahadevan, N.; DiCristofaro, S.; Jortberg, E.; Ceruolo, M.; Aranyosi, A.J. Skin mounted accelerometer system for measuring knee range of motion. In Proceedings of the 2016 IEEE 38th Annual International Conference of the Engineering in Medicine and Biology Society (EMBC), Orlando, FL, USA, 17–20 August 2016; pp. 5298–5302. [Google Scholar]
- Gurchiek, R.D.; Choquette, R.H.; Beynnon, B.D.; Slauterbeck, J.R.; Tourville, T.W.; Toth, M.J.; McGinnis, R.S. Remote Gait Analysis Using Wearable Sensors Detects Asymmetric Gait Patterns in Patients Recovering from ACL Reconstruction. In Proceedings of the 2019 IEEE 16th International Conference on Wearable and Implantable Body Sensor Networks (BSN), Chicago, IL, USA, 19–22 May 2019; pp. 1–4. [Google Scholar]
- Sigward, S.M.; Chan, M.-S.M.; Lin, P.E. Characterizing knee loading asymmetry in individuals following anterior cruciate ligament reconstruction using inertial sensors. Gait Posture 2016, 49, 114–119. [Google Scholar] [CrossRef]
- Takayanagi, N.; Sudo, M.; Yamashiro, Y.; Lee, S.; Kobayashi, Y.; Niki, Y.; Shimada, H. Relationship between Daily and In-laboratory Gait Speed among Healthy Community-dwelling Older Adults. Sci. Rep. 2019, 9, 3496. [Google Scholar] [CrossRef]
- Prajapati, S.K.; Gage, W.H.; Brooks, D.; Black, S.E.; McIlroy, W.E. A Novel Approach to Ambulatory Monitoring: Investigation Into the Quantity and Control of Everyday Walking in Patients With Subacute Stroke. Neurorehabil. Neural Repair 2011, 25, 6–14. [Google Scholar] [CrossRef]
- Del Din, S.; Godfrey, A.; Galna, B.; Lord, S.; Rochester, L. Free-living gait characteristics in ageing and Parkinson’s disease: Impact of environment and ambulatory bout length. J. Neuroeng. Rehabil. 2016, 13, 46. [Google Scholar] [CrossRef]
- Richards, R.E.; van den Noort, J.C.; van der Esch, M.; Booij, M.J.; Harlaar, J. Effect of real-time biofeedback on peak knee adduction moment in patients with medial knee osteoarthritis: Is direct feedback effective? Clin. Biomech. 2018, 57, 150–158. [Google Scholar] [CrossRef] [PubMed]
- Andriacchi, T.P.; Mündermann, A. The role of ambulatory mechanics in the initiation and progression of knee osteoarthritis. Curr. Opin. Rheumatol. 2006, 18, 514–518. [Google Scholar] [CrossRef] [PubMed]
- Carbone, A.; Rodeo, S. Review of current understanding of post-traumatic osteoarthritis resulting from sports injuries. J. Orthop. Res. 2017, 35, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Delp, S.L.; Loan, J.P.; Hoy, M.G.; Zajac, F.E.; Topp, E.L.; Rosen, J.M. An interactive graphics-based model of the lower extremity to study orthopaedic surgical procedures. IEEE Trans. Biomed. Eng. 1990, 37, 757–767. [Google Scholar] [CrossRef]
- Cofré Lizama, L.E.; Khan, F.; Lee, P.V.; Galea, M.P. The use of laboratory gait analysis for understanding gait deterioration in people with multiple sclerosis. Mult. Scler. Houndmills Basingstoke Engl. 2016, 22, 1768–1776. [Google Scholar] [CrossRef]
- Van Veen, B.; Montefiori, E.; Modenese, L.; Mazzà, C.; Viceconti, M. Muscle recruitment strategies can reduce joint loading during level walking. J. Biomech. 2019. [Google Scholar] [CrossRef]
- Myers, C.A.; Laz, P.J.; Shelburne, K.B.; Judd, D.L.; Winters, J.D.; Stevens-Lapsley, J.E.; Davidson, B.S. Simulated hip abductor strengthening reduces peak joint contact forces in patients with total hip arthroplasty. J. Biomech. 2019, 93, 18–27. [Google Scholar] [CrossRef]
- Decker, M.J.; Torry, M.R.; Noonan, T.J.; Sterett, W.I.; Steadman, J.R. Gait retraining after anterior cruciate ligament reconstruction. Arch. Phys. Med. Rehabil. 2004, 85, 848–856. [Google Scholar] [CrossRef]
- Sabatini, A.M. Estimating Three-Dimensional Orientation of Human Body Parts by Inertial/Magnetic Sensing. Sensors 2011, 11, 1489–1525. [Google Scholar] [CrossRef]
- Bergamini, E.; Ligorio, G.; Summa, A.; Vannozzi, G.; Cappozzo, A.; Sabatini, A. Estimating Orientation Using Magnetic and Inertial Sensors and Different Sensor Fusion Approaches: Accuracy Assessment in Manual and Locomotion Tasks. Sensors 2014, 14, 18625–18649. [Google Scholar] [CrossRef]
- McGinnis, R.S.; Hough, J.; Perkins, N.C. Accuracy of Wearable Sensors for Estimating Joint Reactions. J. Comput. Nonlinear Dyn. 2017, 12, 041010. [Google Scholar] [CrossRef]
- Lloyd, D.G.; Besier, T.F. An EMG-driven musculoskeletal model to estimate muscle forces and knee joint moments in vivo. J. Biomech. 2003, 36, 765–776. [Google Scholar] [CrossRef]
- Sartori, M.; Reggiani, M.; Farina, D.; Lloyd, D.G. EMG-Driven Forward-Dynamic Estimation of Muscle Force and Joint Moment about Multiple Degrees of Freedom in the Human Lower Extremity. PLoS ONE 2012, 7, e52618. [Google Scholar] [CrossRef]
- Winters, J.M. Hill-Based Muscle Models: A Systems Engineering Perspective. In Multiple Muscle Systems: Biomechanics and Movement Organization; Winters, J.M., Woo, S.L.-Y., Eds.; Springer: New York, NY, USA, 1990; pp. 69–93. ISBN 978-1-4613-9030-5. [Google Scholar]
- Dowling, J.J. The Use of Electromyography for the Noninvasive Prediction of Muscle Forces: Current Issues. Sports Med. 1997, 24, 82–96. [Google Scholar] [CrossRef] [PubMed]
- Gurchiek, R.D.; McGinnis, R.S.; Needle, A.R.; McBride, J.M.; van Werkhoven, H. The use of a single inertial sensor to estimate 3-dimensional ground reaction force during accelerative running tasks. J. Biomech. 2017, 61, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Blemker, S.S.; Pinsky, P.M.; Delp, S.L. A 3D model of muscle reveals the causes of nonuniform strains in the biceps brachii. J. Biomech. 2005, 38, 657–665. [Google Scholar] [CrossRef]
- Fernandez, J.W.; Buist, M.L.; Nickerson, D.P.; Hunter, P.J. Modelling the passive and nerve activated response of the rectus femoris muscle to a flexion loading: A finite element framework. Med. Eng. Phys. 2005, 27, 862–870. [Google Scholar] [CrossRef]
- Röhrle, O.; Sprenger, M.; Schmitt, S. A two-muscle, continuum-mechanical forward simulation of the upper limb. Biomech. Model. Mechanobiol. 2017, 16, 743–762. [Google Scholar] [CrossRef]
- Staudenmann, D.; Roeleveld, K.; Stegeman, D.F.; van Dieën, J.H. Methodological aspects of SEMG recordings for force estimation—A tutorial and review. J. Electromyogr. Kinesiol. 2010, 20, 375–387. [Google Scholar] [CrossRef]
- Dumas, R.; Barré, A.; Moissenet, F.; Aissaoui, R. Can a reduction approach predict reliable joint contact and musculo-tendon forces? J. Biomech. 2019, 95. [Google Scholar] [CrossRef]
- Dorschky, E.; Nitschke, M.; Seifer, A.-K.; van den Bogert, A.J.; Eskofier, B.M. Estimation of gait kinematics and kinetics from inertial sensor data using optimal control of musculoskeletal models. J. Biomech. 2019, 95, 109278. [Google Scholar] [CrossRef] [PubMed]
- Clancy, E.A.; Bida, O.; Rancourt, D. Influence of advanced electromyogram (EMG) amplitude processors on EMG-to-torque estimation during constant-posture, force-varying contractions. J. Biomech. 2006, 39, 2690–2698. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, C.E.; Williams, K.I. Gaussian Processes for Machine Learning; The MIT Press: Cambridge, MA, USA, 2006. [Google Scholar]
- Stetter, B.J.; Ringhof, S.; Krafft, F.C.; Sell, S.; Stein, T. Estimation of Knee Joint Forces in Sport Movements Using Wearable Sensors and Machine Learning. Sensors 2019, 19, 3690. [Google Scholar] [CrossRef] [PubMed]
- Faisal, A.I.; Majumder, S.; Mondal, T.; Cowan, D.; Naseh, S.; Deen, M.J. Monitoring Methods of Human Body Joints: State-of-the-Art and Research Challenges. Sensors 2019, 19, 2629. [Google Scholar] [CrossRef] [PubMed]
- Trinler, U.; Hollands, K.; Jones, R.; Baker, R. A systematic review of approaches to modelling lower limb muscle forces during gait: Applicability to clinical gait analyses. Gait Posture 2018, 61, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Ancillao, A.; Tedesco, S.; Barton, J.; O’Flynn, B. Indirect Measurement of Ground Reaction Forces and Moments by Means of Wearable Inertial Sensors: A Systematic Review. Sensors 2018, 18, 2564. [Google Scholar] [CrossRef]
- Schöllhorn, W.I. Applications of artificial neural nets in clinical biomechanics. Clin. Biomech. 2004, 19, 876–898. [Google Scholar] [CrossRef]
- Shull, P.B.; Jirattigalachote, W.; Hunt, M.A.; Cutkosky, M.R.; Delp, S.L. Quantified self and human movement: A review on the clinical impact of wearable sensing and feedback for gait analysis and intervention. Gait Posture 2014, 40, 11–19. [Google Scholar] [CrossRef]
- Caldas, R.; Mundt, M.; Potthast, W.; Buarque de Lima Neto, F.; Markert, B. A systematic review of gait analysis methods based on inertial sensors and adaptive algorithms. Gait Posture 2017, 57, 204–210. [Google Scholar] [CrossRef]
- Suryanarayanan, S.; Reddy, N.P.; Gupta, V. An intelligent system with EMG-based joint angle estimation for telemanipulation. Stud. Health Technol. Inform. 1996, 29, 546–552. [Google Scholar]
- Clancy, E.A.; Liu, L.; Liu, P.; Moyer, D.V.Z. Identification of Constant-Posture EMG–Torque Relationship About the Elbow Using Nonlinear Dynamic Models. IEEE Trans. Biomed. Eng. 2012, 59, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Song, R.; Tong, K.Y. Using recurrent artificial neural network model to estimate voluntary elbow torque in dynamic situations. Med. Biol. Eng. Comput. 2005, 43, 473–480. [Google Scholar] [CrossRef] [PubMed]
- Youn, W.; Kim, J. Estimation of elbow flexion force during isometric muscle contraction from mechanomyography and electromyography. Med. Biol. Eng. Comput. 2010, 48, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Xiloyannis, M.; Gavriel, C.; Thomik, A.A.C.; Faisal, A.A. Gaussian Process Autoregression for Simultaneous Proportional Multi-Modal Prosthetic Control With Natural Hand Kinematics. IEEE Trans. Neural Syst. Rehabil. Eng. 2017, 25, 1785–1801. [Google Scholar] [CrossRef] [PubMed]
- Howell, A.M.; Kobayashi, T.; Hayes, H.A.; Foreman, K.B.; Bamberg, S.J.M. Kinetic Gait Analysis Using a Low-Cost Insole. IEEE Trans. Biomed. Eng. 2013, 60, 3284–3290. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, D.A.; Ferris, D.P. Estimation of ground reaction forces and ankle moment with multiple, low-cost sensors. J. NeuroEng. Rehabil. 2015, 12, 90. [Google Scholar] [CrossRef]
- Wang, J.; Wang, L.; Miran, S.M.; Xi, X.; Xue, A. Surface Electromyography Based Estimation of Knee Joint Angle by Using Correlation Dimension of Wavelet Coefficient. IEEE Access 2019, 7, 60522–60531. [Google Scholar] [CrossRef]
- Xia, P.; Hu, J.; Peng, Y. EMG-Based Estimation of Limb Movement Using Deep Learning With Recurrent Convolutional Neural Networks. Artif. Organs 2018, 42, E67–E77. [Google Scholar] [CrossRef]
- Xu, L.; Chen, X.; Cao, S.; Zhang, X.; Chen, X. Feasibility Study of Advanced Neural Networks Applied to sEMG-Based Force Estimation. Sensors 2018, 18, 3226. [Google Scholar] [CrossRef]
- Farmer, S.; Silver-Thorn, B.; Voglewede, P.; Beardsley, S.A. Within-socket myoelectric prediction of continuous ankle kinematics for control of a powered transtibial prosthesis. J. Neural Eng. 2014, 11, 056027. [Google Scholar] [CrossRef]
- Stulp, F.; Sigaud, O. Many regression algorithms, one unified model: A review. Neural Netw. 2015, 69, 60–79. [Google Scholar] [CrossRef] [PubMed]
- Michieletto, S.; Tonin, L.; Antonello, M.; Bortoletto, R.; Spolaor, F.; Pagello, E.; Menegatti, E. GMM-Based Single-Joint Angle Estimation Using EMG Signals. In Intelligent Autonomous Systems 13; Menegatti, E., Michael, N., Berns, K., Yamaguchi, H., Eds.; Springer International Publishing: Cham, Switzerland, 2016; Volume 302, pp. 1173–1184. ISBN 978-3-319-08337-7. [Google Scholar]
- Ziai, A.; Menon, C. Comparison of regression models for estimation of isometric wrist joint torques using surface electromyography. J. NeuroEng. Rehabil. 2011, 8, 56. [Google Scholar] [CrossRef] [PubMed]
- Hahne, J.M.; Biebmann, F.; Jiang, N.; Rehbaum, H.; Farina, D.; Meinecke, F.C.; Muller, K.-R.; Parra, L.C. Linear and Nonlinear Regression Techniques for Simultaneous and Proportional Myoelectric Control. IEEE Trans. Neural Syst. Rehabil. Eng. 2014, 22, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Mijovic, B.; Popovic, M.B.; Popovic, D.B. Synergistic control of forearm based on accelerometer data and artificial neural networks. Braz. J. Med. Biol. Res. 2008, 41, 389–397. [Google Scholar] [CrossRef]
- Koike, Y.; Kawato, M. Estimation of dynamic joint torques and trajectory formation from surface electromyography signals using a neural network model. Biol. Cybern. 1995, 73, 291–300. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, 2nd ed.; Springer: New York, NY, USA, 2009. [Google Scholar]
- Ngeo, J.G.; Tamei, T.; Shibata, T. Continuous and simultaneous estimation of finger kinematics using inputs from an EMG-to-muscle activation model. J. NeuroEng. Rehabil. 2014, 11, 122. [Google Scholar] [CrossRef]
- Findlow, A.; Goulermas, J.Y.; Nester, C.; Howard, D.; Kenney, L.P.J. Predicting lower limb joint kinematics using wearable motion sensors. Gait Posture 2008, 28, 120–126. [Google Scholar] [CrossRef]
- Goulermas, J.Y.; Findlow, A.H.; Nester, C.J.; Liatsis, P.; Zeng, X.J.; Kenney, L.; Tresadern, P.; Thies, S.B.; Howard, D. An Instance-Based Algorithm With Auxiliary Similarity Information for the Estimation of Gait Kinematics From Wearable Sensors. IEEE Trans. Neural Netw. 2008, 19, 1574–1582. [Google Scholar] [CrossRef]
- Wouda, F.; Giuberti, M.; Bellusci, G.; Veltink, P. Estimation of Full-Body Poses Using Only Five Inertial Sensors: An Eager or Lazy Learning Approach? Sensors 2016, 16, 2138. [Google Scholar] [CrossRef]
- Sun, W.; Zhu, J.; Jiang, Y.; Yokoi, H.; Huang, Q. One-Channel Surface Electromyography Decomposition for Muscle Force Estimation. Front. Neurorobotics 2018, 12, 20. [Google Scholar] [CrossRef]
- Shih, P.-S.; Patterson, P.E. Predicting Joint Moments and Angles from EMG Signals. Biomed. Sci. Instrum. 1997, 33, 191–196. [Google Scholar] [PubMed]
- Van Dieën, J.H.; Visser, B. Estimating net lumbar sagittal plane moments from EMG data. The validity of calibration procedures. J. Electromyogr. Kinesiol. 1999, 9, 309–315. [Google Scholar] [CrossRef]
- Au, A.T.C.; Kirsch, R.F. EMG-based prediction of shoulder and elbow kinematics in able-bodied and spinal cord injured individuals. IEEE Trans. Rehabil. Eng. 2000, 8, 471–480. [Google Scholar] [CrossRef] [PubMed]
- Dipietro, L.; Sabatini, A.M.; Dario, P. Artificial neural network model of the mapping between electromyographic activation and trajectory patterns in free-arm movements. Med. Biol. Eng. Comput. 2003, 41, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Dosen, S.; Popovic, D.B. Accelerometers and Force Sensing Resistors for Optimal Control of Walking of a Hemiplegic. IEEE Trans. Biomed. Eng. 2008, 55, 1973–1984. [Google Scholar] [CrossRef] [PubMed]
- Hahn, M.E.; O’Keefe, K.B. A NEURAL NETWORK MODEL FOR ESTIMATION OF NET JOINT MOMENTS DURING NORMAL GAIT. J. Musculoskelet. Res. 2008, 11, 117–126. [Google Scholar] [CrossRef]
- Delis, A.L.; Carvalho, J.L.A.; da Rocha, A.F.; Ferreira, R.U.; Rodrigues, S.S.; Borges, G.A. Estimation of the knee joint angle from surface electromyographic signals for active control of leg prostheses. Physiol. Meas. 2009, 30, 931–946. [Google Scholar] [CrossRef] [PubMed]
- Jiang, N.; Englehart, K.B.; Parker, P.A. Extracting Simultaneous and Proportional Neural Control Information for Multiple-DOF Prostheses From the Surface Electromyographic Signal. IEEE Trans. Biomed. Eng. 2009, 56, 1070–1080. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, J.L.G.; Holmgaard, S.; Jiang, N.; Englehart, K.B.; Farina, D.; Parker, P.A. Simultaneous and Proportional Force Estimation for Multifunction Myoelectric Prostheses Using Mirrored Bilateral Training. IEEE Trans. Biomed. Eng. 2011, 58, 681–688. [Google Scholar] [CrossRef]
- De Vries, W.H.K.; Veeger, H.E.J.; Baten, C.T.M.; van der Helm, F.C.T. Determining a long term ambulatory load profile of the shoulder joint: Neural networks predicting input for a musculoskeletal model. Hum. Mov. Sci. 2012, 31, 419–428. [Google Scholar] [CrossRef]
- Jiang, N.; Vest-Nielsen, J.L.; Muceli, S.; Farina, D. EMG-based simultaneous and proportional estimation of wrist/hand kinematics in uni-lateral trans-radial amputees. J. NeuroEng. Rehabil. 2012, 9, 42. [Google Scholar] [CrossRef] [PubMed]
- Muceli, S.; Farina, D. Simultaneous and Proportional Estimation of Hand Kinematics From EMG During Mirrored Movements at Multiple Degrees-of-Freedom. IEEE Trans. Neural Syst. Rehabil. Eng. 2012, 20, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Kamavuako, E.N.; Scheme, E.J.; Englehart, K.B. Wrist torque estimation during simultaneous and continuously changing movements: surface vs. untargeted intramuscular EMG. J. Neurophysiol. 2013, 109, 2658–2665. [Google Scholar] [CrossRef] [PubMed]
- Jiang, N.; Muceli, S.; Graimann, B.; Farina, D. Effect of arm position on the prediction of kinematics from EMG in amputees. Med. Biol. Eng. Comput. 2013, 51, 143–151. [Google Scholar] [CrossRef] [PubMed]
- De Vries, W.H.K.; Veeger, H.E.J.; Baten, C.T.M.; van der Helm, F.C.T. Can shoulder joint reaction forces be estimated by neural networks? J. Biomech. 2016, 49, 73–79. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, R.; Chen, W.; Xiong, C. Simultaneous and Continuous Estimation of Shoulder and Elbow Kinematics from Surface EMG Signals. Front. Neurosci. 2017, 11, 280. [Google Scholar] [CrossRef]
- Ding, Q.; Han, J.; Zhao, X. Continuous Estimation of Human Multi-Joint Angles From sEMG Using a State-Space Model. IEEE Trans. Neural Syst. Rehabil. Eng. 2017, 25, 1518–1528. [Google Scholar] [CrossRef]
- Clancy, E.A.; Martinez-Luna, C.; Wartenberg, M.; Dai, C.; Farrell, T.R. Two degrees of freedom quasi-static EMG-force at the wrist using a minimum number of electrodes. J. Electromyogr. Kinesiol. 2017, 34, 24–36. [Google Scholar] [CrossRef]
- Wouda, F.J.; Giuberti, M.; Bellusci, G.; Maartens, E.; Reenalda, J.; van Beijnum, B.-J.F.; Veltink, P.H. Estimation of Vertical Ground Reaction Forces and Sagittal Knee Kinematics During Running Using Three Inertial Sensors. Front. Physiol. 2018, 9, 218. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, X.; Cheng, Y.; Xi, N. Surface EMG based continuous estimation of human lower limb joint angles by using deep belief networks. Biomed. Signal Process. Control 2018, 40, 335–342. [Google Scholar] [CrossRef]
- Dai, C.; Hu, X. Finger Joint Angle Estimation Based on Motoneuron Discharge Activities. IEEE J. Biomed. Health Inform. 2019. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Zhu, Z.; Martinez-Luna, C.; Hunt, T.R.; Farrell, T.R.; Clancy, E.A. Two degrees of freedom, dynamic, hand-wrist EMG-force using a minimum number of electrodes. J. Electromyogr. Kinesiol. 2019, 47, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Kapelner, T.; Vujaklija, I.; Jiang, N.; Negro, F.; Aszmann, O.C.; Principe, J.; Farina, D. Predicting wrist kinematics from motor unit discharge timings for the control of active prostheses. J. NeuroEng. Rehabil. 2019, 16, 47. [Google Scholar] [CrossRef] [PubMed]
- McGinnis, R.S.; Mahadevan, N.; Moon, Y.; Seagers, K.; Sheth, N.; Wright, J.A.; DiCristofaro, S.; Silva, I.; Jortberg, E.; Ceruolo, M.; et al. A machine learning approach for gait speed estimation using skin-mounted wearable sensors: From healthy controls to individuals with multiple sclerosis. PLoS ONE 2017, 12, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Gurchiek, R.D.; Choquette, R.H.; Beynnon, B.D.; Slauterbeck, J.R.; Tourville, T.W.; Toth, M.J.; McGinnis, R.S. Open-Source Remote Gait Analysis: A Post-Surgery Patient Monitoring Application. Sci. Rep. 2019. [Google Scholar] [CrossRef]
- Tang, W.; Sazonov, E.S. Highly Accurate Recognition of Human Postures and Activities Through Classification With Rejection. IEEE J. Biomed. Health Inform. 2014, 18, 309–315. [Google Scholar] [CrossRef]
- Fregly, B.J.; Reinbolt, J.A.; Rooney, K.L.; Mitchell, K.H.; Chmielewski, T.L. Design of patient-specific gait modifications for knee osteoarthritis rehabilitation. IEEE Trans. Biomed. Eng. 2007, 54, 1687–1695. [Google Scholar] [CrossRef]
- Kern, D.S.; Semmler, J.G.; Enoka, R.M. Long-term activity in upper- and lower-limb muscles of humans. J. Appl. Physiol. 2001, 91, 2224–2232. [Google Scholar] [CrossRef]
- Tikkanen, O.; Haakana, P.; Pesola, A.J.; Häkkinen, K.; Rantalainen, T.; Havu, M.; Pullinen, T.; Finni, T. Muscle Activity and Inactivity Periods during Normal Daily Life. PLoS ONE 2013, 8, e52228. [Google Scholar] [CrossRef]
- Finni, T.; Haakana, P.; Pesola, A.J.; Pullinen, T. Exercise for fitness does not decrease the muscular inactivity time during normal daily life: Inactivity time is independent of exercise. Scand. J. Med. Sci. Sports 2014, 24, 211–219. [Google Scholar] [CrossRef]
- Gao, Y.; Melin, M.; Mäkäräinen, K.; Rantalainen, T.; Pesola, A.J.; Laukkanen, A.; Sääkslahti, A.; Finni, T. Children’s physical activity and sedentary time compared using assessments of accelerometry counts and muscle activity level. PeerJ 2018, 6, e5437. [Google Scholar] [CrossRef] [PubMed]
- McGinnis, R.S.; Slauterbeck, J.R.; Tourville, T.W.; Toth, M.J. Wearable Sensors Capture Differences in Muscle Activity and Gait Patterns During Daily Activity in Patients Recovering from ACL Reconstruction. In Proceedings of the 15th International Conference on Wearable and Implantable Body Sensor Networks, Las Vegas, NV, USA, 3–7 March 2018; pp. 38–41. [Google Scholar]
- Clancy, E.A.; Morin, E.L.; Merletti, R. Sampling, noise-reduction and amplitude estimation issues in surface electromyography. J. Electromyogr. Kinesiol. 2002, 12, 1–16. [Google Scholar] [CrossRef]
- Buchanan, T.S.; Lloyd, D.G.; Manal, K.; Besier, T.F. Neuromusculoskeletal Modeling: Estimation of Muscle Forces and Joint Moments and Movements from Measurements of Neural Command. J. Appl. Biomech. 2004, 20, 367–395. [Google Scholar] [CrossRef] [PubMed]
- Neptune, R.R.; Clark, D.J.; Kautz, S.A. Modular control of human walking: A simulation study. J. Biomech. 2009, 42, 1282–1287. [Google Scholar] [CrossRef] [PubMed]
- Tresch, M.C.; Jarc, A. The case for and against muscle synergies. Curr. Opin. Neurobiol. 2009, 19, 601–607. [Google Scholar] [CrossRef]
- Tresch, M.C.; Cheung, V.C.K.; d’Avella, A. Matrix Factorization Algorithms for the Identification of Muscle Synergies: Evaluation on Simulated and Experimental Data Sets. J. Neurophysiol. 2006, 95, 2199–2212. [Google Scholar] [CrossRef]
- Bianco, N.A.; Patten, C.; Fregly, B.J. Can Measured Synergy Excitations Accurately Construct Unmeasured Muscle Excitations? J. Biomech. Eng. 2018, 140. [Google Scholar] [CrossRef]
- Wang, L.; Buchanan, T.S. Prediction of joint moments using a neural network model of muscle activations from EMG signals. IEEE Trans. Neural Syst. Rehabil. Eng. 2002, 10, 30–37. [Google Scholar] [CrossRef]
- Gurchiek, R.D.; Rupasinghe Arachchige Don, H.S.; Pelawa Watagoda, L.C.R.; McGinnis, R.S.; van Werkhoven, H.; Needle, A.R.; McBride, J.M.; Arnholt, A.T. Sprint Assessment Using Machine Learning and a Wearable Accelerometer. J. Appl. Biomech. 2019, 35, 164–169. [Google Scholar] [CrossRef]
- Reichstein, M.; Camps-Valls, G.; Stevens, B.; Jung, M.; Denzler, J.; Carvalhais, N. Prabhat Deep learning and process understanding for data-driven Earth system science. Nature 2019, 566, 195–204. [Google Scholar] [CrossRef]
- Xiong, Y.; Zhang, Y.; Guo, X.; Wang, C.; Shen, C.; Li, J.; Tang, J.; Liu, J. Seamless global positioning system/inertial navigation system navigation method based on square-root cubature Kalman filter and random forest regression. Rev. Sci. Instrum. 2019, 90, 015101. [Google Scholar] [CrossRef] [PubMed]
- Pathak, J.; Wikner, A.; Fussell, R.; Chandra, S.; Hunt, B.R.; Girvan, M.; Ott, E. Hybrid forecasting of chaotic processes: Using machine learning in conjunction with a knowledge-based model. Chaos Interdiscip. J. Nonlinear Sci. 2018, 28, 041101. [Google Scholar] [CrossRef]
- Artificial intelligence alone won’t solve the complexity of Earth sciences. Nature 2019, 566, 153. [CrossRef] [PubMed]
- Gui, K.; Liu, H.; Zhang, D. A Practical and Adaptive Method to Achieve EMG-Based Torque Estimation for a Robotic Exoskeleton. IEEEASME Trans. Mechatron. 2019, 24, 483–494. [Google Scholar] [CrossRef]
| Review Relevant Item | Search Terms |
|---|---|
| Regression | regress* OR “machine learning” OR “artificial intelligence” OR “statistical learning” OR “supervised learning” OR “unsupervised learning” OR “neural network” OR perceptron OR “support vector” OR “gaussian process” |
| AND | |
| Biomechanical Time-Series | joint OR limb OR segment OR ankle OR knee OR hip OR wrist OR elbow OR shoulder OR muscle AND angle OR velocity OR acceleration OR moment OR torque OR force OR kinematic* OR kinetic* OR biomechanics OR mechanics OR dynamics |
| AND | |
| Wearable Sensors | wearable OR accelerometer OR gyroscope OR electromyo* OR EMG OR sEMG OR “inertial sensor” OR “inertial measurement unit” OR IMU OR insole OR goniometer |
| Reference (Year) | Variable (Location): Plane(s) | Tasks | Inputs | Model | Performance Summary | |
|---|---|---|---|---|---|---|
| Koike and Kawato [60] (1995) | sEMG (2 kHz, 10) | (elbow): S (shoulder): F | ISO, OC | TS | NN (FB, dyn) | CD: 0.89 |
| Suryanarayanan et al. [44] (1996) | sEMG (2 kHz, 1) | (elbow): S | OC | TS | NN (dyn) | RMSE 15% |
| Shih and Patterson [67] (1997) | sEMG (900 Hz, 4) | (elbow): S (wrist): S (shoulder): S (elbow): S (wrist): S (shoulder): S | WCP | TS | NN | RMSE: 0.67–5.76 Nm, 0.64–5.62 Nm RMSE: 4.78–13.76°, 4.73–14.33° |
| van Dieën and Visser [68] (1999) | sEMG (600 Hz, 6) | (lumbo-sacral): S | ISO, LOC | TS | (dyn) | RMSE: 26–54 Nm, 49–160 Nm |
| Au and Kirsch [69] (2000) | sEMG (500 Hz, 6) | (shoulder): S, F, T (elbow): S (shoulder): S, F, T (elbow): S (shoulder): S, F, T (elbow): S | OC, LOC | TS | NN (dyn) | RMSE: 14.2–19.6° RMSE: 8–17.2° (impaired subjects) |
| Dipietro et al. [70] (2003) | sEMG (1 kHz, 5) | (hand): T | OC | TS | NN (FB) | RMSE: 7.3–11.5% |
| Song and Tong [46] (2005) | sEMG (1 kHz, 3) goni (1 kHz, 2) | (elbow): S | LOC | TS | NN (FB) | nRMSE: 4.53–8.45% nRMSE: 10.56–16.20% (sEMG only) |
| Clancy et al. [35] (2006) | sEMG (4096 Hz, 8) | (elbow): S | ISO | TS | (dyn) | MAE: 7.3% |
| Došen and Popovič [71] (2008) | 2D ACC (200 Hz, 4) | (ankle): S (knee): S (hip): S (hip joint center): S | MSW | TS | NN (dyn) | RMSE: 1.19–3.60°, 1.18–2.62° RMSE: 0.26–0.39 m/s2, 0.29–0.46 m/s2 CC (): 0.97–0.998, 0.97–0.998 CC (): 0.96–0.99, 0.91–0.99 |
| Findlow et al. [63] (2008) | IMU (100 Hz, 4) | (ankle): S (knee): S (hip): S | Normal Walk | TS | NP (KS) | MAE: 1.69–2.30°, 4.91–9.06° MAE: 1.78–5.32° (reduced sensor array) CC: 0.93–0.99, 0.70–0.89 CC: 0.87–0.99 (reduced sensor array) |
| Goulermas et al. [64] (2008) | IMU (--, 4) | (ankle): S (knee): S (hip): S | MSW | TS | NP (KS) | CC: 0.97, 0.96, 0.83 |
| Hahn and O’Keefe [72] (2008) | sEMG (1 kHz, 7) | (ankle): S (knee): S (hip): S | Normal Walk | TS | NN | CD: 0.54–0.84 (sEMG only) CD: 0.77–0.92 (sEMG with demographics & anthropometrics) |
| Mijovic et al. [59] (2008) | 2D ACC (50 Hz, 2) | (forearm): S | OC | TS | NN (RBF) | CD: 0.841–0.998, 0.75–0.99, 0.03–0.88 |
| Delis et al. [73] (2009) | sEMG (1744.25 Hz, 2) | (knee): S | Normal Walk | DISC (TD) | NN (SOM) | CC: 0.59–0.84 |
| Jiang et al. [74] (2009) | sEMG (1 kHz, 8) | CF (hand) | ISO | DISC (TD) | (1) NN (2) | (1) CD: 0.86 (2) CD: 0.78 |
| Youn and Kim [47] (2010) | sEMG (1 kHz, 2) MMG (1 kHz, 2) | CF (hand) | ISO | DISC (TD) | NN | nRMSE 16% (MMG only) nRMSE 13% (sEMG only) nRMSE 10% (sEMG + MMG) |
| Ziai and Menon [57] (2011) | sEMG (1 kHz, 8) | (wrist): S | ISO | TS | (1) (2) (lasso) (3) (LWPR) (4) NP (SVR) (5) NN (2L) | (1) nRMSE: 2.88% (2) nRMSE: 2.83% (3) nRMSE: 3.03% (4) nRMSE: 2.85% (5) nRMSE: 2.82% |
| Nielsen et al. [75] (2011) | sEMG (1024 Hz, 7) | CF (hand) | ISO | DISC (TD) | NN | RMSE: 0.16 N RMSE: 0.10 N (impaired subjects) CD: 0.93 CD: 0.82 (impaired subjects) |
| de Vries et al. [76] (2012) | MIMU (50 Hz, 4) sEMG (1 kHz, 13) | ISF (SC): S, F, T ISF (AC): S, F, T ISF (shoulder): S, F, T ISF (elbow): S, F, T | LOC, ADL | TS | NN | nRMSE: 7–17% |
| Jiang et al. [77] (2012) | sEMG (2048 Hz, 7) | (wrist): S, F, T | OC | DISC (TD) | NN | CD: 0.74–0.78 |
| Muceli and Farina [78] (2012) | HD-sEMG 128 (2048 Hz, 2) | (wrist): S, F, T | OC | TS | NN | CD: 0.79–0.89 |
| Clancy et al. [45] (2012) | sEMG (4096 Hz, 2) | (elbow): S | ISO | TS | , , , (dyn) | nMAE: 4.65–6.38% nMAE: 5.55–7.97% (reduced training set) |
| Howell et al. [49] (2013) | FSR (118 Hz, 12) | (ankle): S (knee): S, F | Normal Walk | TS | nRMSE: 5.9–17.1% CC: 0.82–0.97 | |
| Kamavuako et al. [79] (2013) | sEMG (10 kHz, 6) | (wrist): S, T | ISO | DISC (TD) | NN | nRMSE: 6.1–13.5% CD: 0.87–0.91 |
| Jiang et al. [80] (2013) | sEMG (2048 Hz, 7) | (wrist): S, F, T | OC | DISC (TD) | NN | CD: 0.63–0.86, 0.34–0.74 CD: 0.61–0.77, 0.46–0.59 (impaired subjects) |
| Farmer et al. [54] (2014) | sEMG (1 kHz, 4) | (ankle): S | Normal Walk | TS | NN (FB, dyn) | RMSE: 1.2–5.4° |
| Ngeo et al. [62] (2014) | sEMG (2 kHz, 8) | (MCPs): S | OC | TS DISC (TD) | (1) NN (dyn) (2) NP (GPR, dyn) | (1) CC: 0.71 (TS inputs only) (2) CC: 0.84 (TS inputs only) |
| Hahne et al. [58] (2014) | HD-sEMG 192 (2048 Hz, 1) | (wrist): S, F | OC | DISC (TD) | (1) (ridge) (2) (3) NN (4) NP (KRR) | (4) CD: 0.8 (reduced sensor array) CD: 0.8–0.9 (range across all models) |
| Jacobs and Ferris [50] (2015) | FSR (1 kHz, 8) Load Cell (1 kHz, 1) | (ankle): S | MSW, Calf Raises | TS | NN | nRMSE: 7.04–13.78% nRMSE: 8.72–16.52% (FSR only) nRMSE: 20.47–46.02% (Load Cell only) |
| de Vries et al. [81] (2016) | MIMU (50 Hz, 4) sEMG (1 kHz, 13) | ISF (shoulder): S, F, T | LOC, ADL | TS | NN | nSEM: 4–1% nSEM: 3–21% (reduced sensor array) |
| Wouda et al. [65] (2016) | MIMU (240 Hz, 5) | (ankle): S, F, T (knee): S, F, T (hip): S, F, T (shoulder): S, F, T (elbow): S, F, T (wrist): S, F, T (spine): S, F, T | OC, ADL, MSW, MSR, sport | TS | (1) NN (2) NP (k-NN) | (1) Mean Error: 7° (2) Mean Error: 8° |
| Michieletto et al. [56] (2016) | sEMG (1 kHz, 8) | (knee): S | Seated Kick | TS | (GMR) | Custom error statistic (see paper) |
| Xiloyannis et al. [48] (2017) | sEMG (--, 5) MMG (--, 5) | (MCPs): S | OC, ADL, ISO | TS | (FB) (2) NP (GPR, FB) | (1) CC: 0.54 (2) CC: 0.79, 0.62, 0.67 (sEMG only) |
| Zhang et al. [82] (2017) | sEMG (1 kHz, 8) | (shoulder): S, F, T (elbow): S | OC | DISC (TD) | NN | CD: 0.90–0.91, 0.86–0.87 |
| Ding et al. [83] (2017) | sEMG (2 kHz, 8) | (elbow): S (humerus): S, F, T | OC, ADL | TS | (1) NN (2) NN (FB) (3) NN (FB, UKF) | (1) RMSE: 11–14°, CC: 0.88–0.90 (2) RMSE: 11–15°, CC: 0.87–0.89 (3) RMSE: 7–9°, CC: 0.95–0.96 |
| Clancy et al. [84] (2017) | sEMG (2048 Hz, 16) | CF (hand): S, F (wrist): T | ISO | TS | RMSE: 6.7–10.6%, 11.0–15.7 (4 sensors) | |
| Xia et al. [52] (2018) | sEMG (2 kHz, 5) | (hand): S, F, T | OC | DISC (FD) DISC (TD) | 1) NN (CNN) 2) NN (C-LSTM, FB) | (1) CD: 0.78 (2) CD: 0.90 |
| Wouda et al. [85] (2018) | MIMU (240 Hz, 3) | (knee): S | MSR | TS | NN | RMSE: 2.27–8.41°, 6.29–25.05° CC: 0.98–0.99, 0.77–0.99 |
| Sun et al. [66] (2018) | sEMG (16 kHz, 1) | CF (forearm) | ISO | DISC (MUAP-TD) | CD: 0.72–0.89 | |
| Chen et al. [86] (2018) | sEMG (1.2 kHz, 10) | (ankle): S (knee): S (hip): S | MSW | TS | NN (DBN) | RMSE: 2.45–3.96° CC: 0.95–0.97 |
| Xu et al. [53] (2018) | HD-sEMG 128 (1 kHz, 1) | CF (forearm) | ISO | TS | (1) NN (CNN) (2) NN (LSTM, FB) (3) NN (C-LSTM, FB) | (1) nRMSE: 7.33–10.93% (2) nRMSE: 6.16–9.33% (3) nRMSE: 5.95–9.74% |
| Wang et al. [51] (2019) | sEMG (1.6 kHz, 5) | (knee): S | LOC | DISC (FD) | NN (FB) | nRMSE: 3.55–5.13% |
| Dai and Hu [87] (2019) | HD-sEMG 160 (2048 Hz, 1) | (MCPs): S | OC | TS, DISC (MUAP-FD) | CD: 0.66–0.81 (TS inputs) CD: 0.69–0.86 (MUAP-FD inputs) | |
| Dai et al. [88] (2019) | sEMG (2048 Hz, 16) | CF (hand): S, F (wrist): T | ISO | TS | (dyn) | RMSE: 7.3–9.2%, 11.5–13.0% (4 sensors) |
| Kapelner et al. [89] (2019) | HD-sEMG 192 (2048 Hz, 3) | (wrist): S, F, T | OC | DISC (TD, MUAP-TD) | CD: 0.77 (MUAP-TD inputs) CD: 0.70 (TD inputs) | |
| Stetter et al. [37] (2019) | IMU (1.5 kHz, 2) | ISF (knee): S, F, T | MSW, MSR, sport | TS | NN (2L) | nRMSE: 14.2–45.9% CC: 0.25–0.94 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gurchiek, R.D.; Cheney, N.; McGinnis, R.S. Estimating Biomechanical Time-Series with Wearable Sensors: A Systematic Review of Machine Learning Techniques. Sensors 2019, 19, 5227. https://doi.org/10.3390/s19235227
Gurchiek RD, Cheney N, McGinnis RS. Estimating Biomechanical Time-Series with Wearable Sensors: A Systematic Review of Machine Learning Techniques. Sensors. 2019; 19(23):5227. https://doi.org/10.3390/s19235227
Chicago/Turabian StyleGurchiek, Reed D., Nick Cheney, and Ryan S. McGinnis. 2019. "Estimating Biomechanical Time-Series with Wearable Sensors: A Systematic Review of Machine Learning Techniques" Sensors 19, no. 23: 5227. https://doi.org/10.3390/s19235227
APA StyleGurchiek, R. D., Cheney, N., & McGinnis, R. S. (2019). Estimating Biomechanical Time-Series with Wearable Sensors: A Systematic Review of Machine Learning Techniques. Sensors, 19(23), 5227. https://doi.org/10.3390/s19235227






