Electrochemical Sunset Yellow Biosensor Based on Photocured Polyacrylamide Membrane for Food Dye Monitoring
Abstract
1. Introduction
2. Materials and Methods
2.1. Apparatus and Reagents
2.2. Preparation of Poly(AAm) and Poly(AAm-co-EMA)
2.3. Preparation of Biosensor Membrane
2.4. Water Absorption Test
2.5. Electrochemical Measurement of Sunset Yellow Using Laccase Immobilized in a Poly(AAm-co-EMA) Membrane
2.6. Optimization of Poly(AAm-co-EMA)/Lac/GCE Biosensor Membrane
2.7. Linear Range and Limit of Detection
2.8. Interference Study of Biosensor
2.9. Determination of Sunset Yellow in Soft Drink
3. Results and Discussion
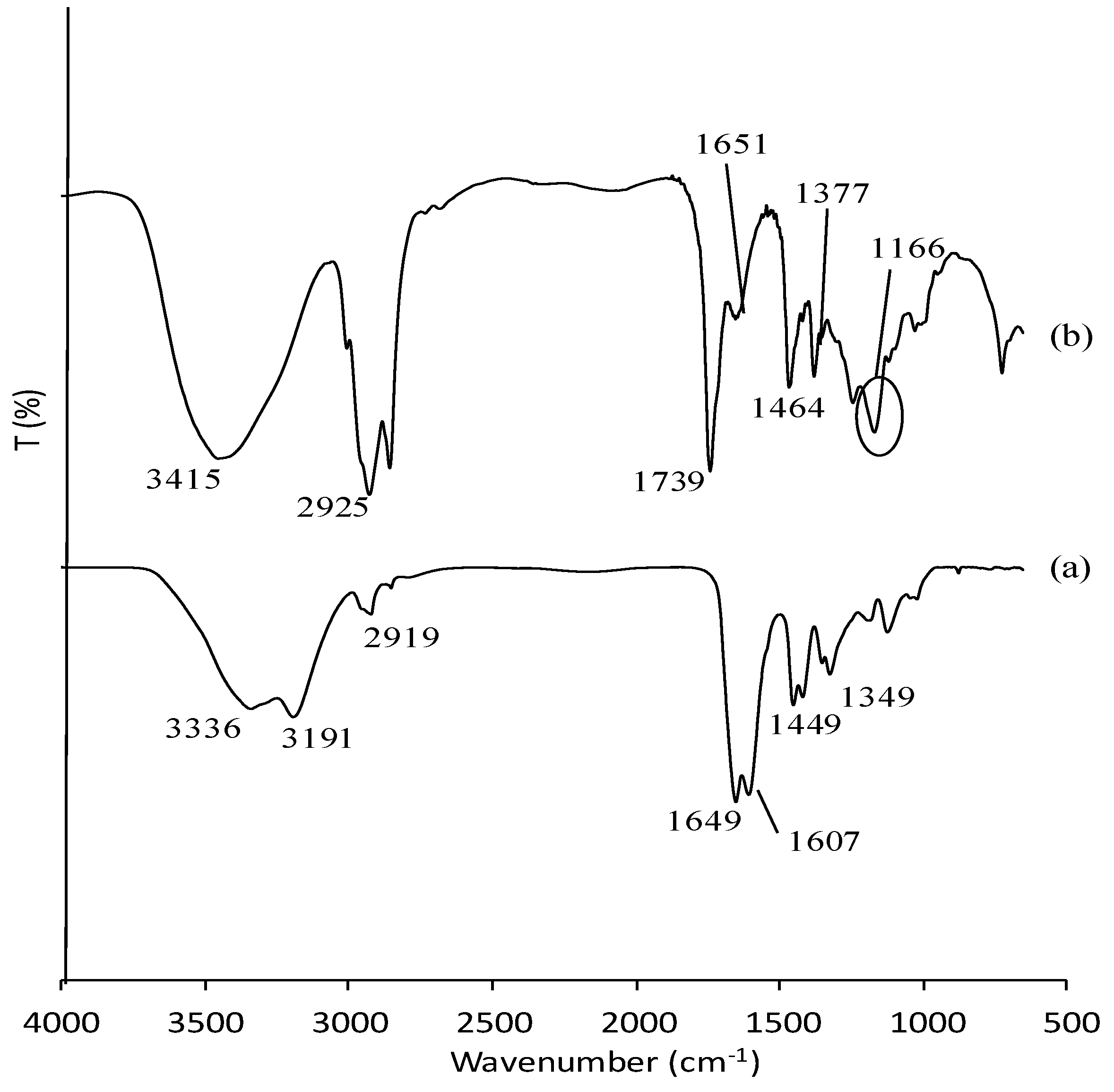
3.1. Characterization of Poly(AAm) and Poly(AAm-co-EMA) by Fourier-Transform Infrared Red (FTIR)
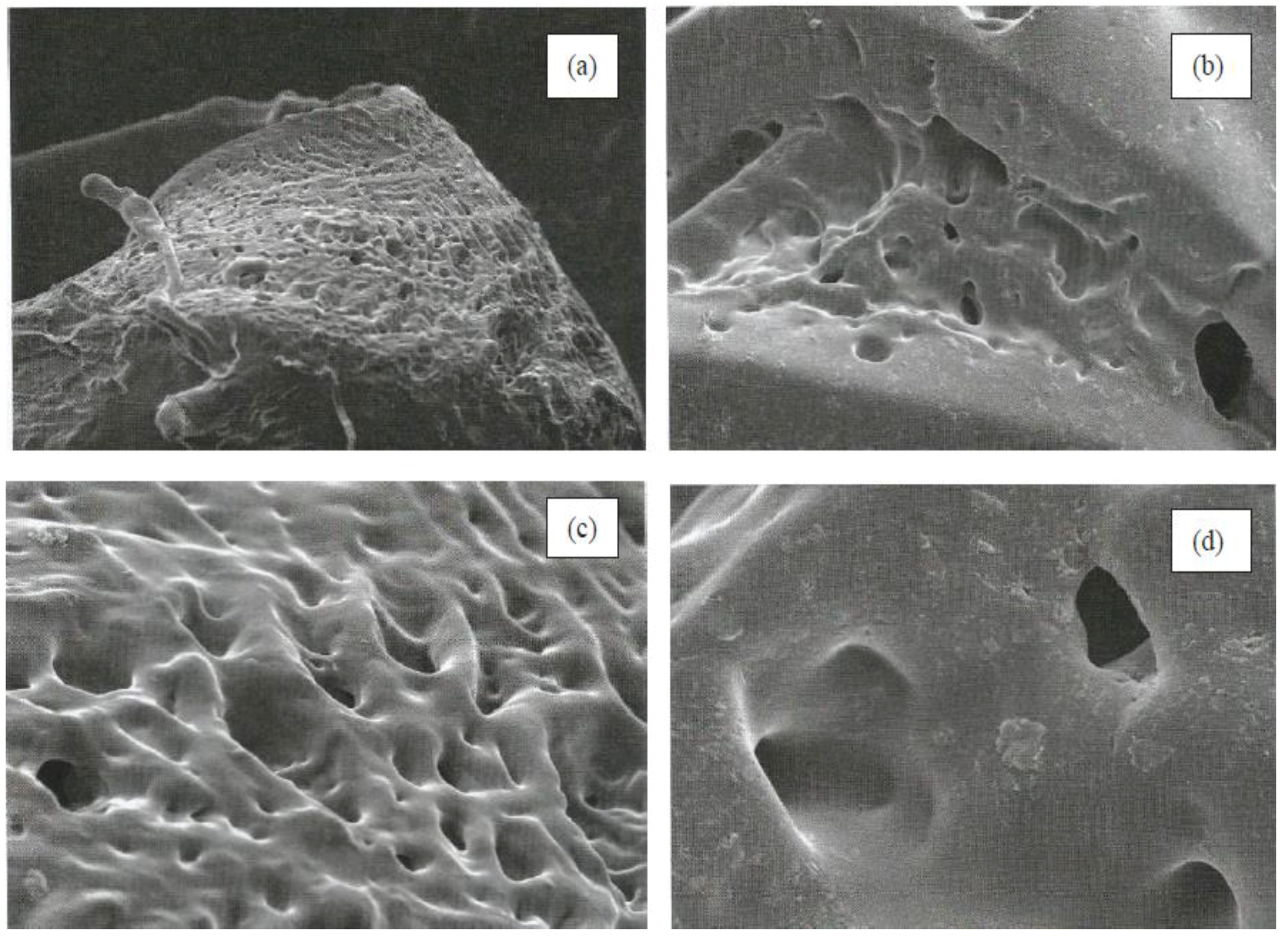
3.2. Morphological Characterization
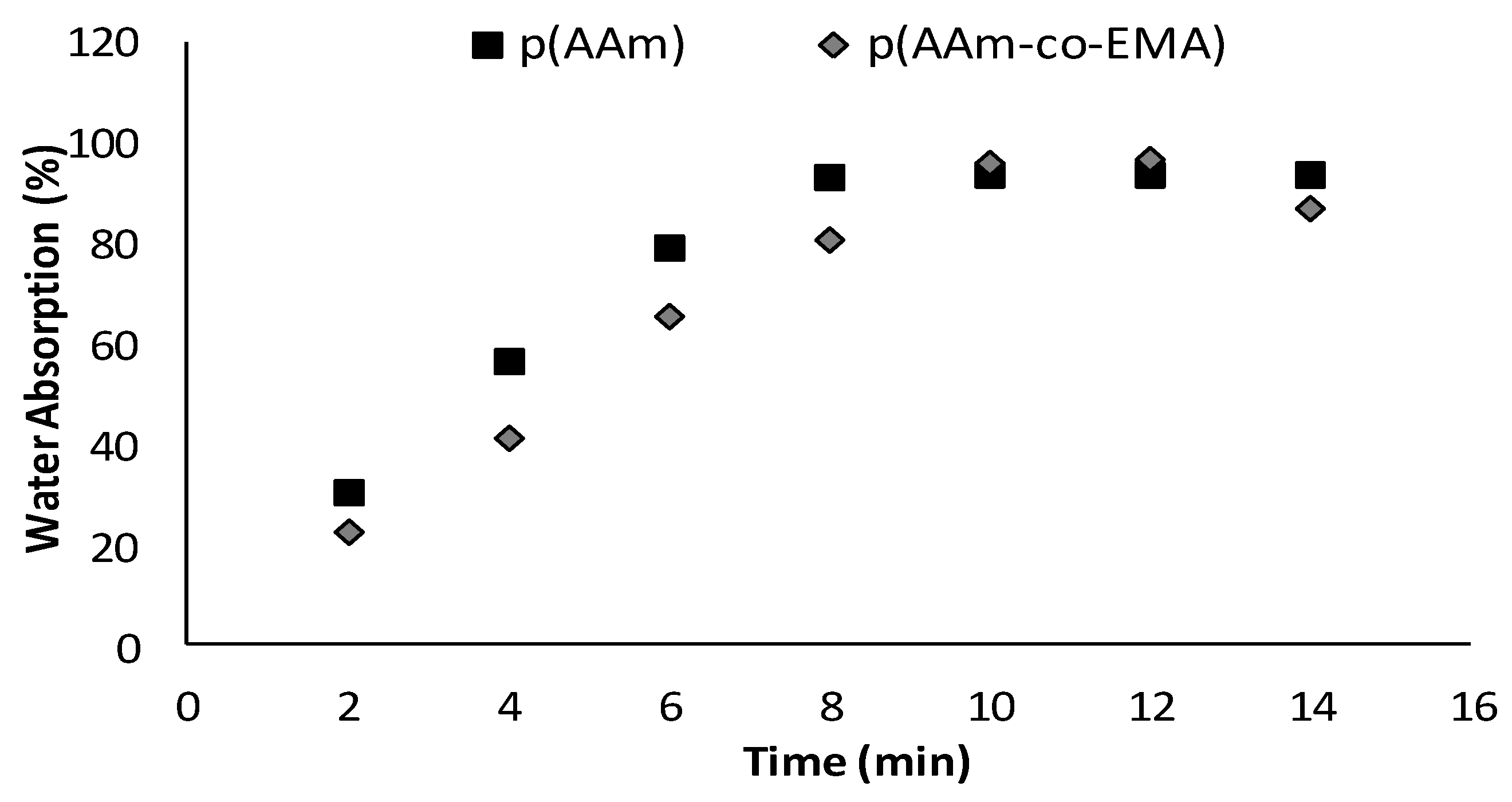
3.3. Water Absorption Test
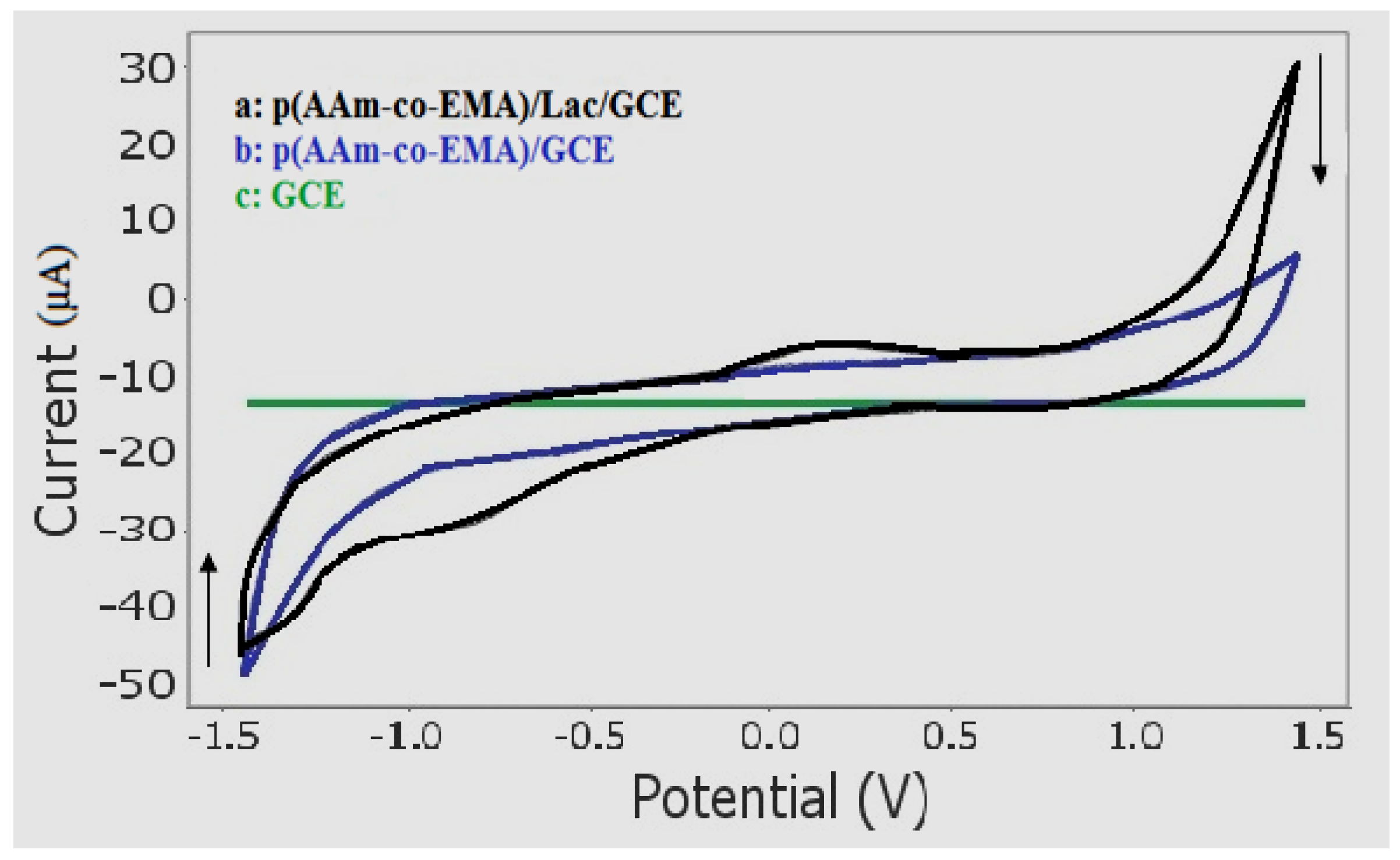
3.4. Electrochemical Behaviour of Sunset Yellow Using Poly(AAm-co-EMA)/Lac/GC Electrode
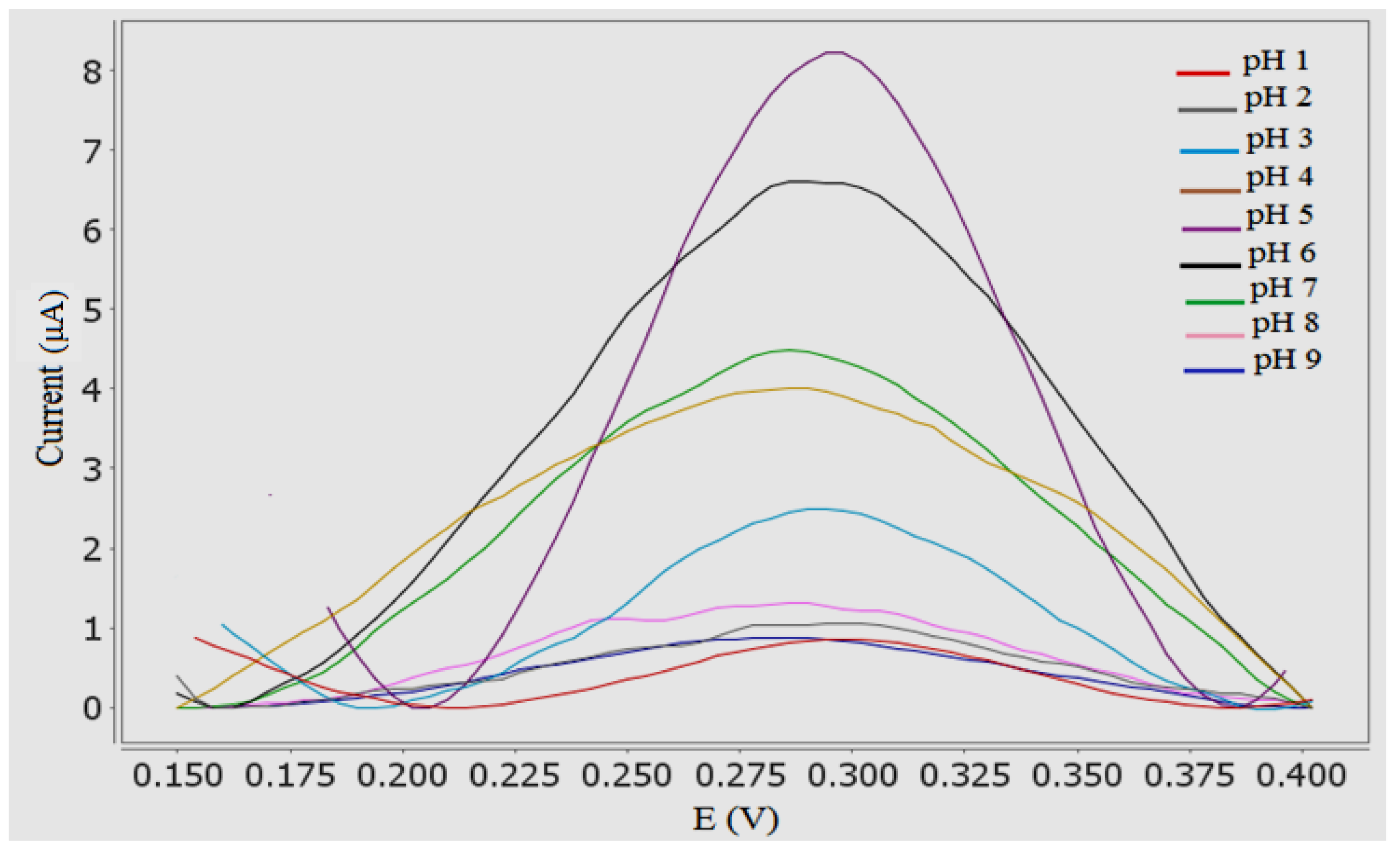
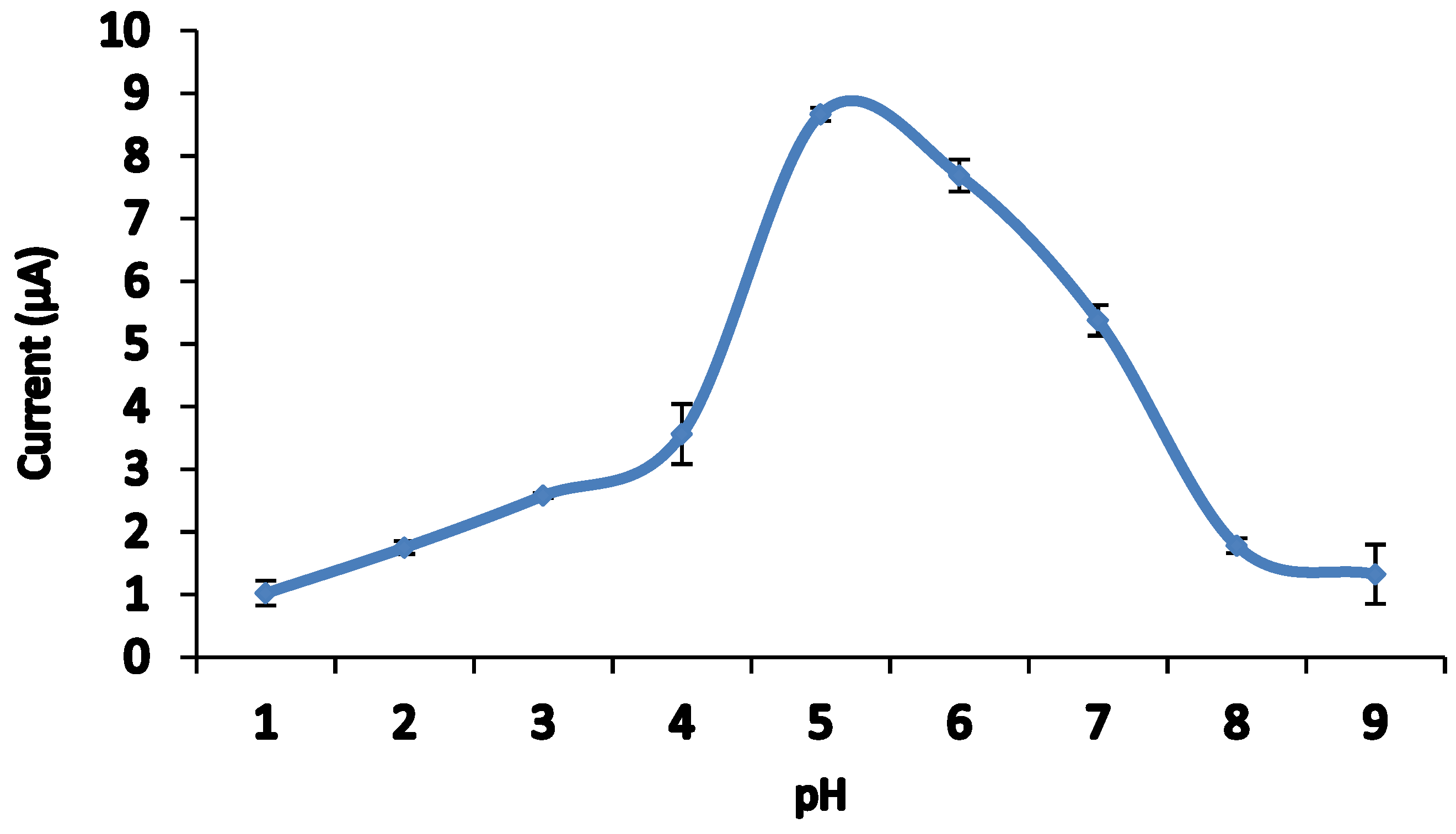
3.5. Effect of pH
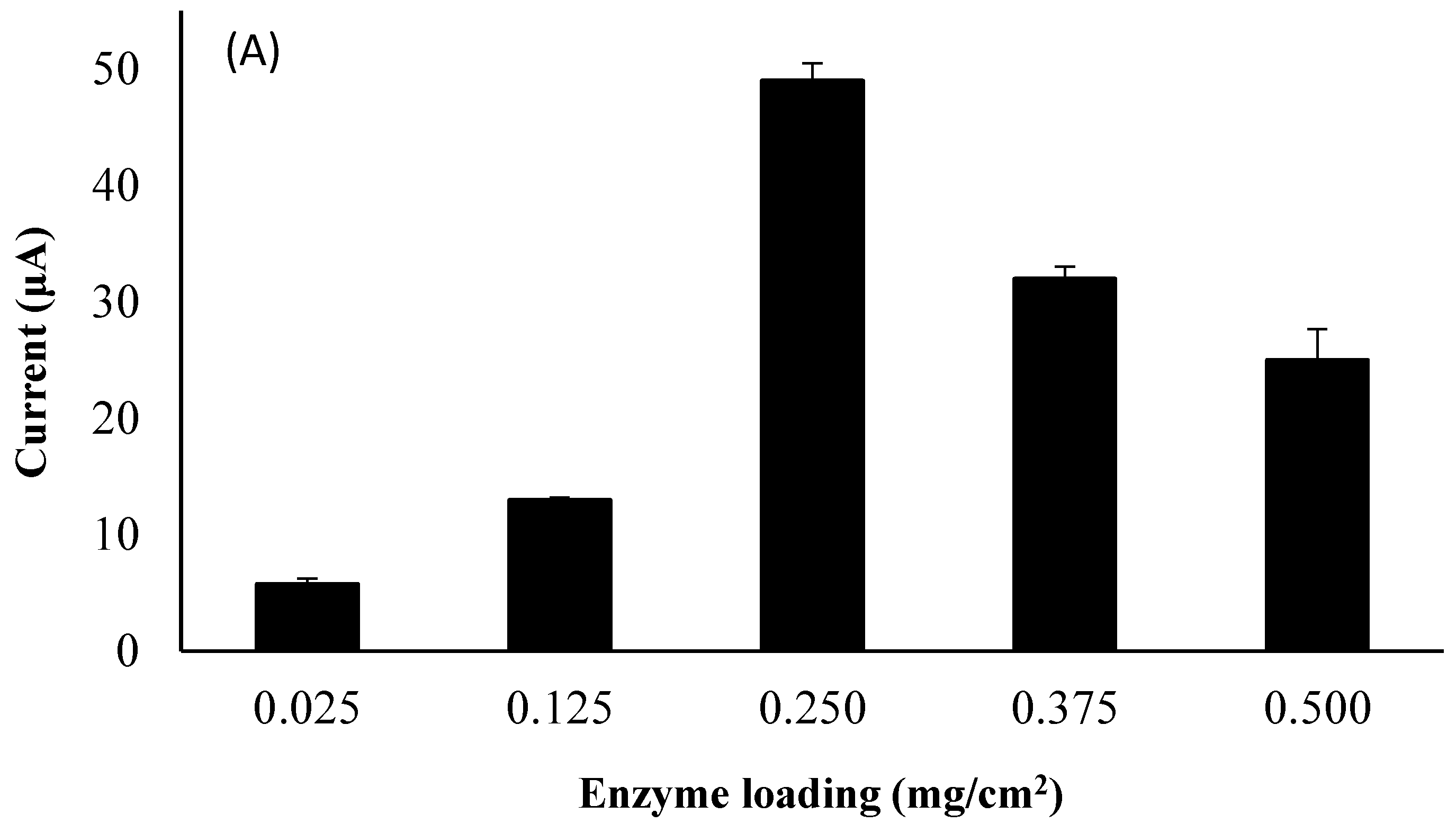
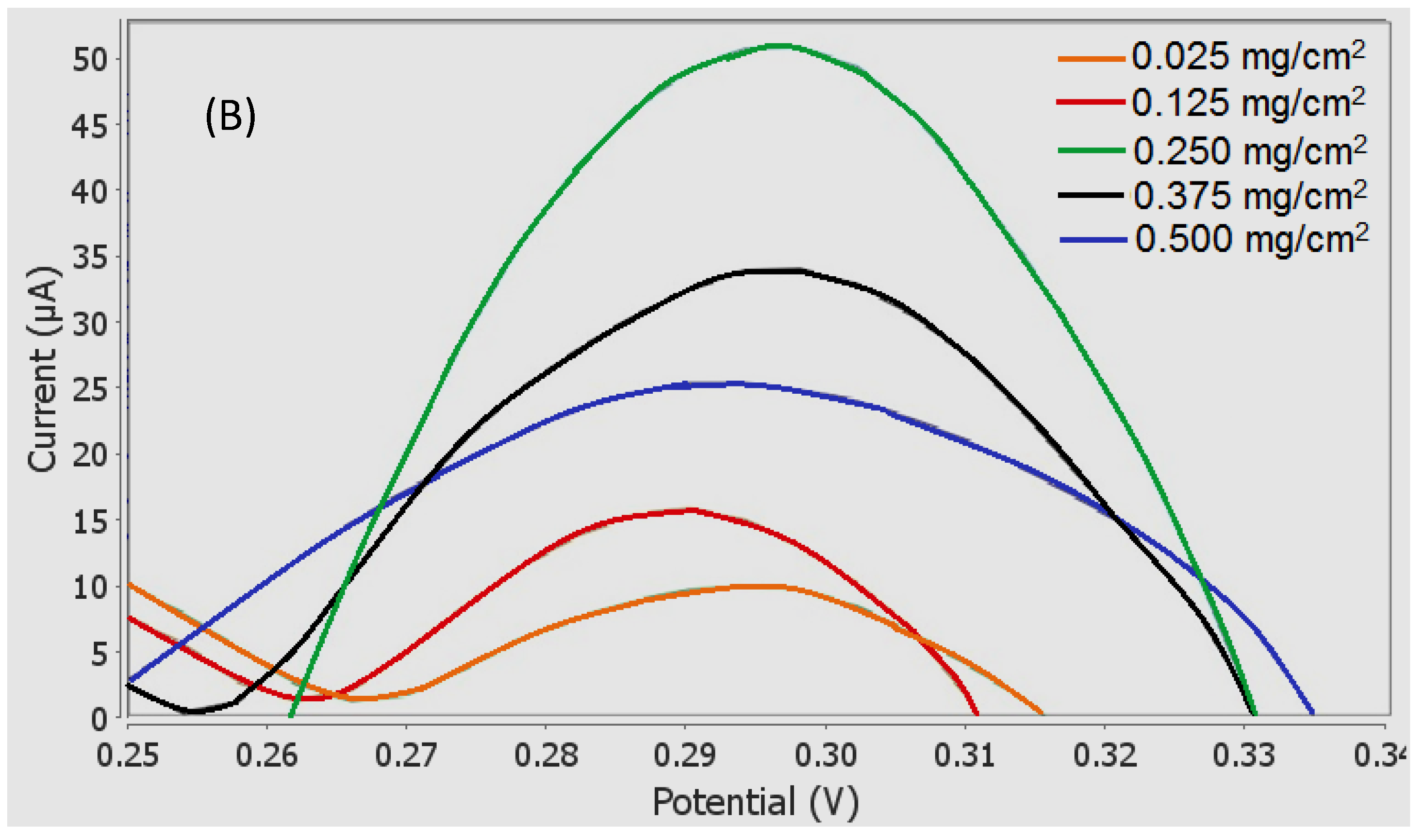
3.6. Influence of Laccase Loading
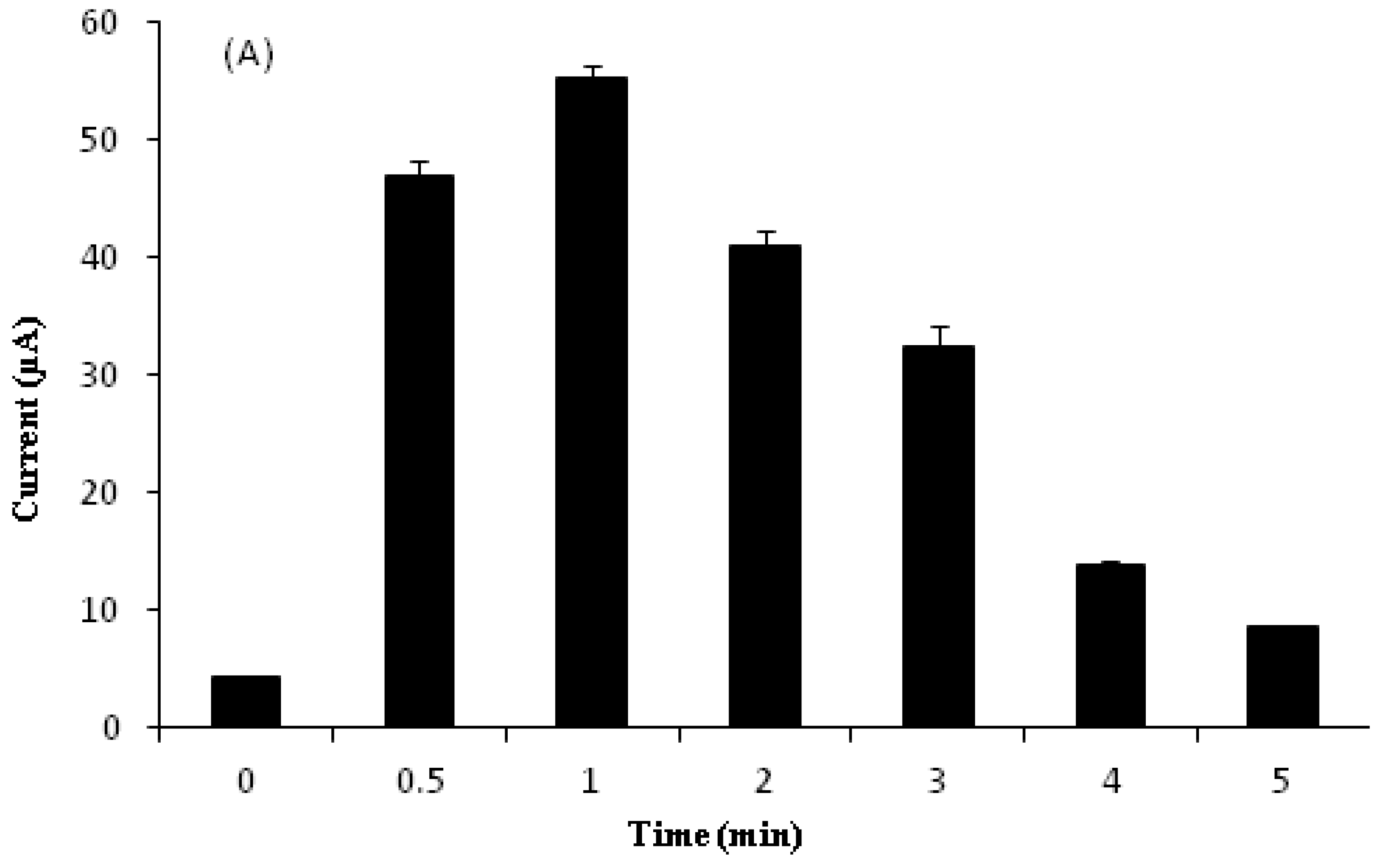
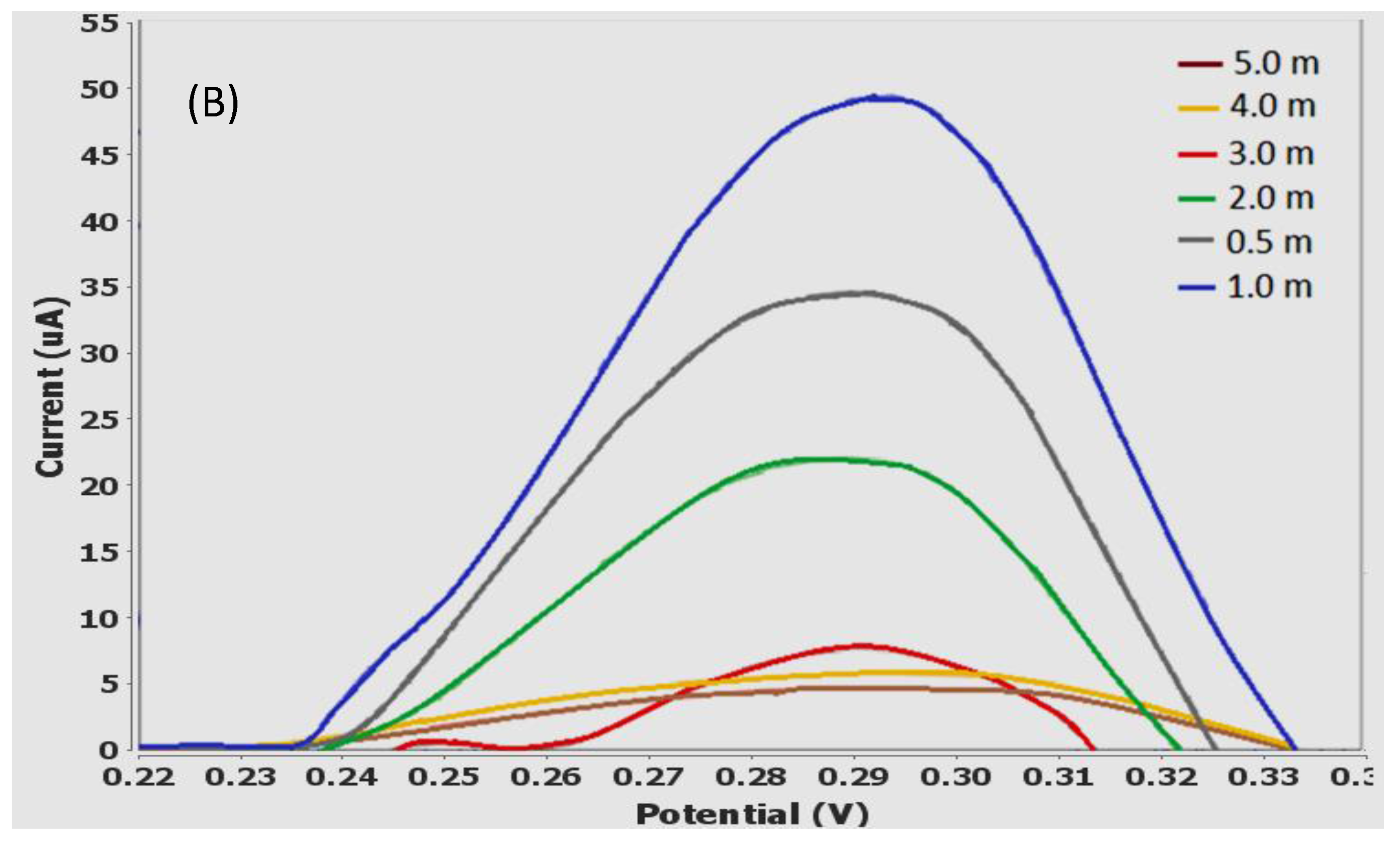
3.7. Effect of Accumulation Time
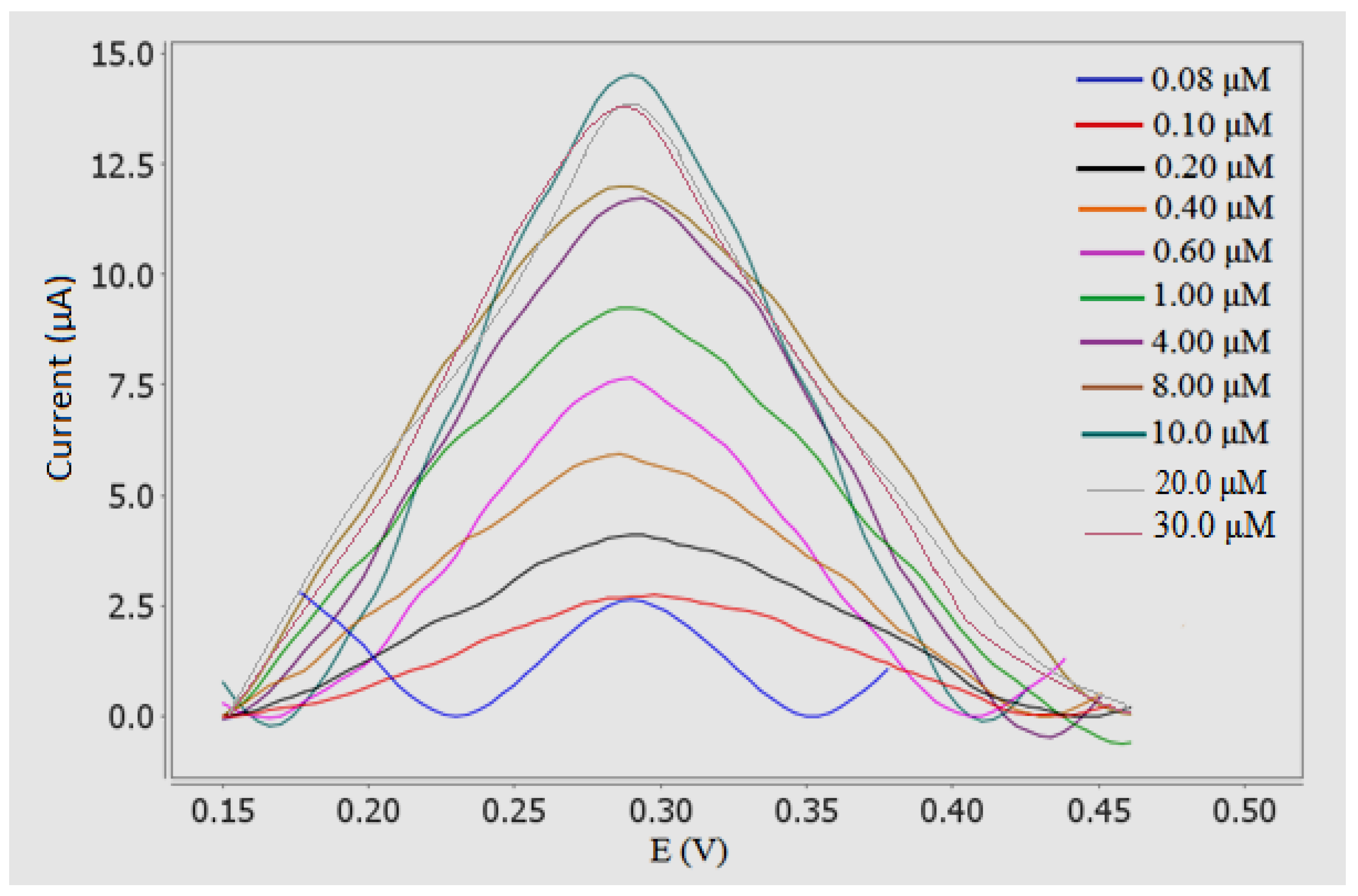
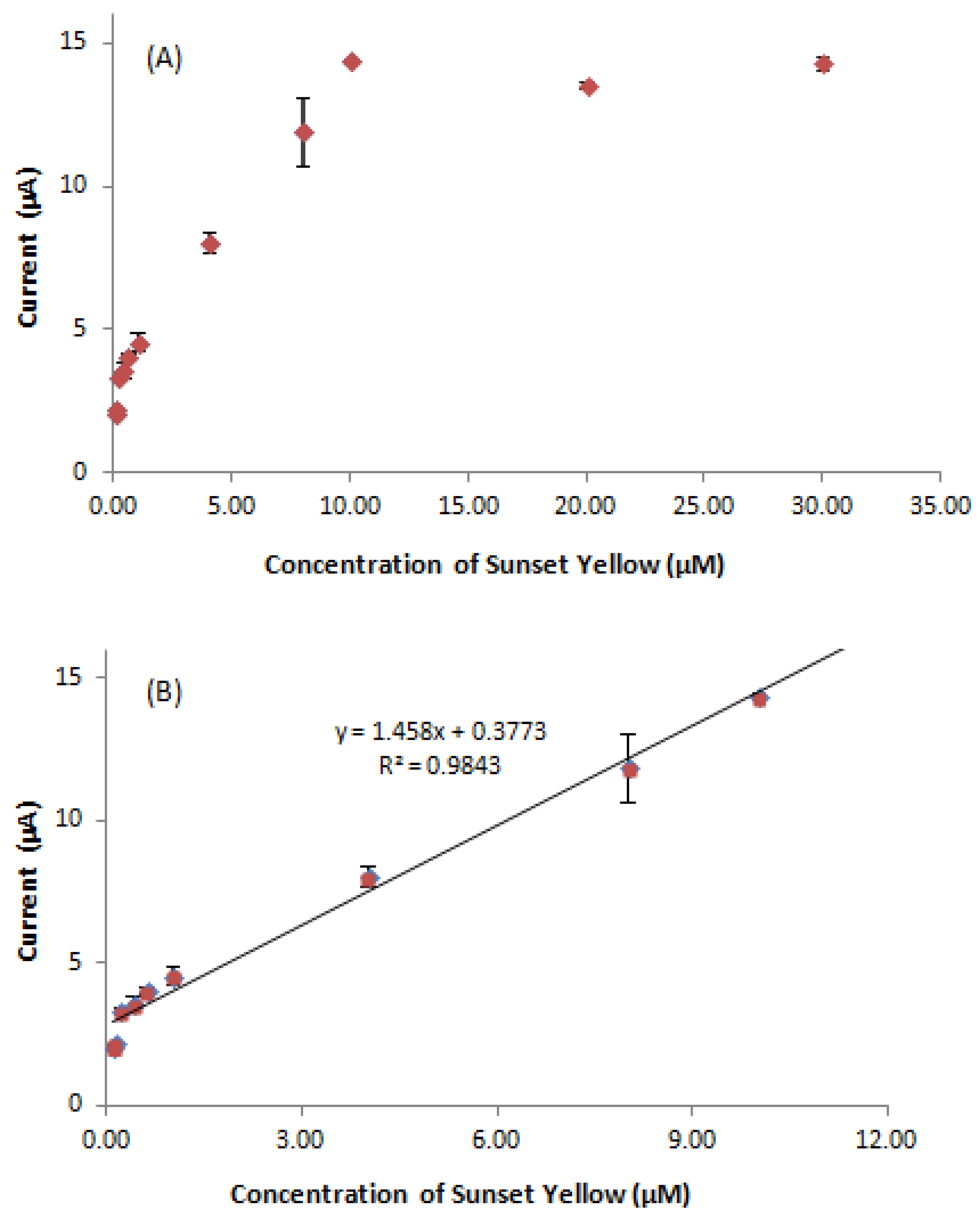
3.8. Determination of Sunset Yellow
3.9. Comparison with Previous Reported Sunset Yellow Sensors
3.10. Reproducibility, Stability and Repeatability
3.11. Interference Study
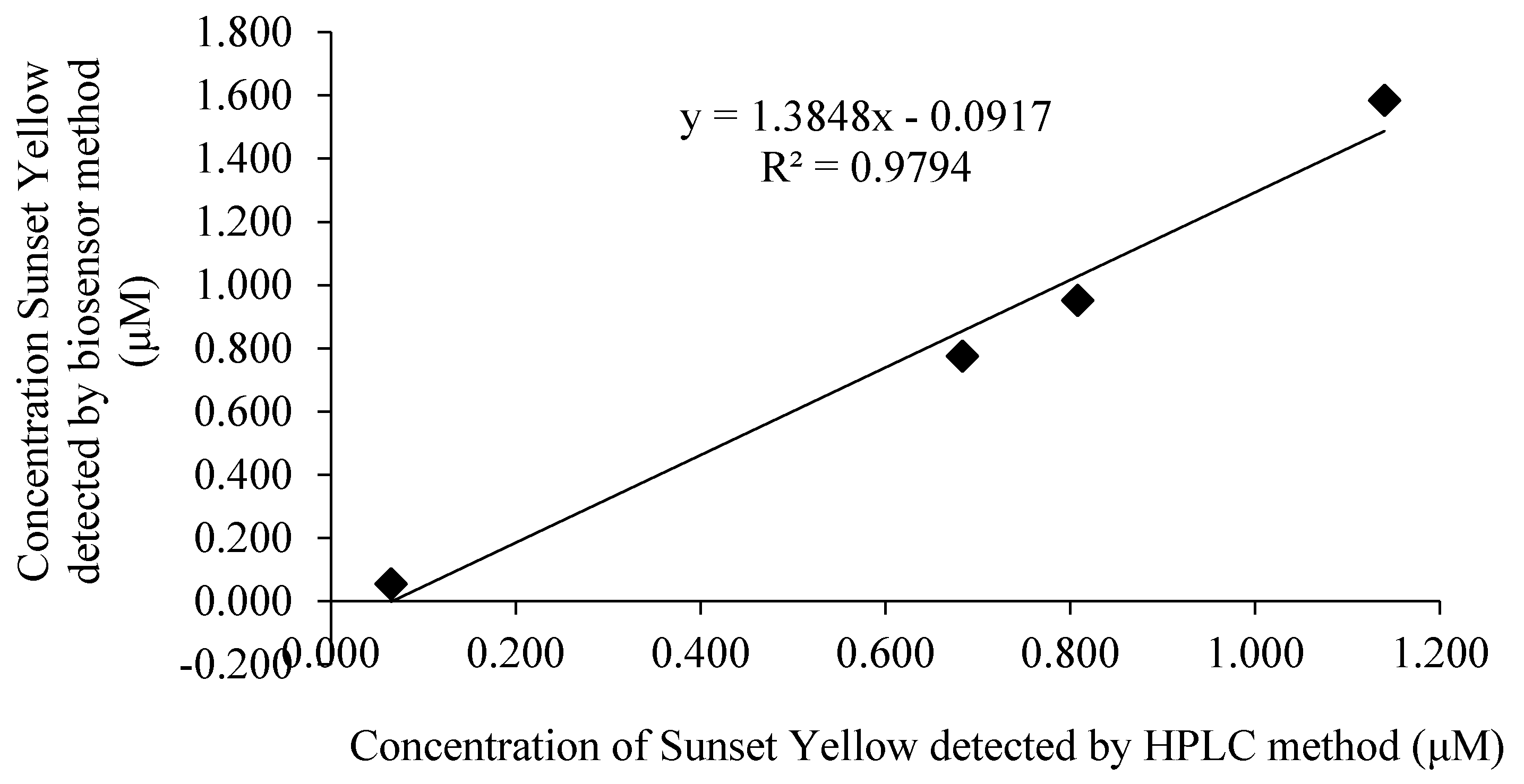
3.12. Sunset Yellow Determination in Soft Drink
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Llamas, N.E.; Garrido, M.; Di Nezio, M.S.; Fernandez Band, B.S. Second order advantage in the determination of amaranth, sunset yellow fcf and tatrazine by uv-vis and multivariate curve resolution-alternating least squares. Anal. Chim. Acta 2009, 655, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Ghoreishi, S.M.; Behpour, M.; Golestaneh, M. Simultaneous determination of Sunset Yellow and Tartrazine in soft drinks using gold nanoparticles carbon paste electrode. Food Chem. 2012, 132, 637–641. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wu, K.; Sun, Y.; Song, X. Highly sensitive electrochemical sensor for sunset yellow based on the enhancement effect of alumina microfibers. Sens. Actuators B 2013, 185, 582–586. [Google Scholar] [CrossRef]
- Wang, M.; Sun, Q.; Gao, Y.; Yang, X.; Zhao, J. Determination of sunset yellow in foods based on a facile electrochemical sensor. Anal. Methods 2014, 6, 8760–8766. [Google Scholar] [CrossRef]
- Wang, M.G.; Sun, Q.; Zhao, J. Sensitively simultaneous determination of sunset yellow and tatrazine in foods based on polyrrole modifies oxidized single-walled carbon nanotubes. J. Electrochem. Soc. 2014, 161, 297–304. [Google Scholar] [CrossRef]
- Xu, J.; Zhang, Y.; Zhou, H.; Wang, M.; Xu, P.; Zhang, J. An amperometric sensor for sunset yellow fcf detection based on molecularly imprinted polypyrrole. Sci. Res. Eng. 2012, 5, 159–162. [Google Scholar] [CrossRef]
- Zhang, W.; Liu, T.; Zheng, X.; Huang, W.; Wan, C. Surfaced-enhanced oxidation and detection of sunset yellow and tatrazine using multi-walled carbon nanotubes film-modified electrode. Colloids Surf. B 2009, 74, 28–31. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Du, Y.; Lu, D.; Wang, C. Fabrication of β-cyclodextrin-coated poly(diallyldimethylammonium chloride)-functionalized graphene composite film modified glassy carbon-rotating disk electrode and its application for simultaneous electrochemical determination colorants of sunset yellow and tatrazine. Anal. Chim. Acta 2013, 779, 22–34. [Google Scholar] [PubMed]
- Srivinivasan, G.P.; Sikkanthar, A.; Elamaran, A.; Delma, C.R.; Subramaniyam, K.; Somasundaran, S.T. Biodegradation of carcinogetic textile azo dye using bacterial isolates of mangrove sediment. J. Coast. Life Med. 2014, 2, 154–162. [Google Scholar]
- Gan, T.; Sun, J.; Cao, S.; Gao, F.; Zhang, Y.; Yang, Y. One-step electrochemical approach fo the preparation of graphene wrapped-phosphotungstic acids hydrid and its application for simulataneous determination of sunset yellow and tatrazine. Electrochim. Acta 2012, 74, 151–157. [Google Scholar] [CrossRef]
- Aguilar, F.; Boon, P.E.; Crebelli, R.; Dusemund, B.; Gott, D.; Hallas-Moller, T.; Konig, J.; Lindtner, O.; Marzin, D.; Meyland, I.; et al. Reconsideration of the temporary ADI and refined exposure assessmet for sunset yellow FCF (E 110). EFSA J. 2014, 12, 1–39. [Google Scholar] [CrossRef]
- Medeiros, R.A.; Lourencao, B.C.; Rocha-Filho, R.C.; Fatibello-Filho, O. Simulataneous voltammetric determination of synthetic colorants in food using a cathodically pretreated boron-doped diamond electrode. Talanta 2012, 97, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Gosetti, F.; Gianotti, V.; Polati, S.; Gennaro, M.C. HPLC-MS degradation study of E110 Sunset Yellow FCF in acommercial beverage. J. Chrom. A 2005, 1090, 107–115. [Google Scholar] [CrossRef]
- Sharma, Y.C. Optimization of parameters for adsorption of methylene blue on a low cost activated carbon. J. Chem. Eng. 2010, 55, 435–439. [Google Scholar] [CrossRef]
- Zhang, Y.; Chang, C.; Guo, Q.; Cao, H.; Bai, Y.; Liu, H. Analysis of tartrazine and sunset yellow aluminium lake in foods by capillary zone electrophoresis. Chin. J. Chrom. 2014, 32, 438–442. [Google Scholar] [CrossRef]
- Song, Y.Z. Electrochemical reduction of sunset yellow at a multi-walled carbon nanotube (MWCNT)-modified glassy carbon electrode and its analytical applications. Can. J. Chem. 2010, 88, 676–681. [Google Scholar] [CrossRef]
- Yu, L.; Shi, M.; Yue, X.; Qu, L. A novel and sensitive hexadecyltrimethyl ammonium bromide functionalized graphene supported platinum nanoparticles composite modified glassy carbon electrode for determination of sunset yellow. Sens. Actuators B 2015, 209, 1–8. [Google Scholar] [CrossRef]
- Munteanu, F.D.; Paulo-Cavaro, A. Biosensors based on laccase for detection of commercially reactive dyes. Anal. Lett. 2010, 43, 1126–1131. [Google Scholar] [CrossRef]
- Hanifah, S.A.; Heng, L.Y.; Ahmad, M. Effect of gold nanoparticles on the response of phenol biosensor containing photocurable membrane with tyrosinase. Sensors 2008, 8, 6407–6416. [Google Scholar]
- Zhang, K.; Luo, P.; Wu, J.; Wang, W.; Ye, B. Highly sensitive determination of sunset yellow in drink using a poly(L-cysteine) modified glassy carbon electrode. Anal. Methods 2013, 5, 5044–5050. [Google Scholar] [CrossRef]
- Yang, J.Y.; Li, W. CTAB functionalized graphene oxide/multi-walled carbon nanotube composite modified electrode for the simultaneous determination of sunset yellow and tartrazine. Russ. J. Electrochem. 2015, 51, 218–226. [Google Scholar] [CrossRef]
- Manole, A.; Herea, D.; Chiriac, H.; Melnig, V. Laccase activity determination. Cuza Uni. Sci. Ann. Lasi 2008, 4, 17–24. [Google Scholar]
- Portaccio, M.; Tuoro, D.D.; Arduini, F.; Moscone, D.; Cammarota, M.; Mita, D.G.; Lepore, M. Laccase biosensor based on screen-printed electrode modified with thionine-carbon black nanocomposite for Bisphenol A detection. Electrochim. Acta 2013, 109, 340–347. [Google Scholar] [CrossRef]
- Li, Y.; Li, Z.; Li, M.; Pan, Z.; Li, D. A disposable biosensor based on immobilization of laccase with silica spheres on the MWCNTs-doped screen printed electrode. Chem. Cent. J. 2012, 6, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Palanisamy, S.; Ramaraj, S.K.; Chen, S.M.; Yang, T.C.K.; Yi-Fan, P.; Chen, T.W.; Velusamy, V.; Selvam, S. A novel laccase biosensor based on laccase immobilized graphene-cellulose microfiber composite modified screen-printed carbon electrode for sensitive determination of catechol. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira Neto, J.R.; Rezende, S.G.; Lobon, G.S.; Garcia, T.A.; Macedo, L.F.; Garcia, L.F.; Alves, V.F.; Torres, I.M.S.; Santiago, M.F.; Schmidt, F.; et al. Electroanalysis and laccase-based biosensor on the determination of phenolic content and antioxidant power of honey samples. Food Chem. 2017, 237, 1118–1123. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.D.; Park, M.H.; Byun, H.S. Mechanical and water barrier properties of starch/PVA composite films by adding nanosized poly(methyl methacrylate-co-acrylamide) particle. Carbohydr. Polym. 2012, 87, 676–686. [Google Scholar] [CrossRef]
- Cai, Y.; Jessop, J.L.P. Decreased oxygen inhibition in photopolumerized acrylate/epoxide hydrid polymer coatings as demonstrated by Raman spectroscopy. Polymer 2006, 47, 6560–6566. [Google Scholar] [CrossRef]
- Rifa, A.A.; Hashim, S.; Muhamad, I.I. Synthesis and characterization of poly(acrylamide-co-acrylic acid)-grafted-poly(styrene-co-methyl) microgels by emulsion polymerization. J. Adv. Mater. Sci. 2015, 5, 21–26. [Google Scholar]
- Karthikeyan, S.; Anandan, C.; Subramanian, J.; Sekaran, G. Characterization of iron impregnated polyacrylamide catalyst and its application to the treatment of municipal wastewater. RSC Adv. 2013, 35, 15044–15057. [Google Scholar] [CrossRef]
- Pavia, D.L.; Lampman, G.M.; Kriz, G.S.; Vyvyan, J.R. Spectroscopy, 4th ed.; Brook/Cole Cengage Learning: Belmont, CA, USA, 2010; ISBN 13 978-0-495-11478-9. [Google Scholar]
- Rozi, N.; Hanifah, S.A.; Heng, L.Y.; Loh, K.S. Characterization of ultraviolet-photocured acrylamide based polymer. AIP Conf. Proc. 2016, 1784, 030017. [Google Scholar] [CrossRef]
- Li, S.; Liu, X. Synthesis, characterization and evaluation of semi-IPN hydrogels consisted of poly(methacrylic acid) and guar gum for colon-specific drug delivery. Poly. Adv. Tech. 2008, 19, 371–376. [Google Scholar] [CrossRef]
- Gowda, V.; Betageri, G.V. Water soluble polymers for pharmaceutical applications. Polymers 2011, 3, 1972–2009. [Google Scholar]
- Nesrinne, S.; Djamel, A. Synthesis, characterization and rheological behavior of pH sensitive poly(acrylamide-co-acrylic acid) hydrogels. Arab. J. Chem. 2013, 10, 1–9. [Google Scholar] [CrossRef]
- Makas, Y.G.; Kalkan, N.A.; Aksoy, S.; Altinok, H.; Hasirci, N. Immobilization of laccase in k-carrageenan based semi-interpenetrating networks. J. Biotechnol. 2010, 148, 216–220. [Google Scholar] [CrossRef] [PubMed]
- Morozova, O.V.; Shumakovich, G.P.; Gorbacheva, M.A.; Shieev, S.V.; Yaropolov, A.I. Blue laccase. Biochemistry 2007, 72, 11–36. [Google Scholar] [CrossRef]
- Rajesh; Kaneto, K. A new tyrosinase biosensor based on covalent immobilization of enzyme on N-(3-aminopropyl) pyrrole polymer film. Curr. Appl. Phys. 2005, 5, 178–183. [Google Scholar] [CrossRef]
- Yin, Z.Z.; Cheng, S.W.; Xu, L.B.; Liu, H.Y.; Huang, K.; Li, L.; Zhai, Y.Y.; Zeng, Y.B.; Liu, H.Q.; Shao, Y.; et al. Highly sensitive and selective sensor for sunset yellow based on molecularly imprinted polydopamine-coated multi-walled carbon nanotubes. Biosens. Bioelectron. 2017, 100, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Rovina, K.; Siddiquee, S.; Shaarani, S.M. Higly sensitive determination of sunset yellow FCF(E110) in food products based on chitosan/nanoparticles/MWCNTs with modified gold electrode. IOP Sci. 2016, 36, 1–7. [Google Scholar] [CrossRef]
- Chao, M.; Ma, X. Convenient electrochemical determination of sunset yellow and tatrazine in food sample using a poly(L-Phenylalanine)-modified glassy carbon electrode. Food Anal. Meth. 2015, 8, 130–138. [Google Scholar] [CrossRef]
- Konig, J. Chapter 2: Food colour additives of synthetic origin (pp. 35–60). In Colour Additives for Foods and Beverages, 1st ed.; Scotter, M.J., Ed.; Woodhead Publishing: Cambridge, UK, 2015; pp. 1–260. ISBN 9781782420118. [Google Scholar]
- Ya, Y.; Jiang, C.; Li, T.; Liao, J.; Fan, Y.; Wei, Y.; Yan, F.; Xie, L. A zinc oxide nanoflower-based electrochemical sensor for trace detection of sunset yellow. Sensors 2017, 17, 545. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.H.; Yuwen, L.X.; Peng, L.J. Parameters affecting the performance of immobilized enzyme. J. Chem. 2013, 2013, 946248. [Google Scholar] [CrossRef]
- Mohammad, R.; Ahmad, M.; Heng, L.Y. An amperometric biosensor utilizing a ferrocene-mediated horseradish peroxidase preaction for the determination of capsaicin. Sensors 2013, 12, 10014–10026. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.; Petrie, K.; Shim, J.; Abildshov, K.M.; Westhoff, C.L.; Cremers, S. A UPLC-MS/MS method for therapeutic drug monitoring of etonogestrel. Ther. Drug. Monit. 2013, 35. [Google Scholar] [CrossRef] [PubMed]
- Toffoli, G.; Rothman, N.; Ferguson, R.; Nagourney, S.; Robinson, D.; Sanguiliano, J.; Worby, P. Data Quality Assessment and Data Usability Evaluation Technical; New Jersey Department of Environment Protection: Trenton, NJ, USA, 2014; pp. 1–138. [Google Scholar]
| Types of Matrix Used | pH | Potential (V) | Linear Range (µM) | LOD (µM) | Sensitivity (μA/μM) | References |
|---|---|---|---|---|---|---|
| Screen-printed electrode modified with gold nanoparticles. | 4.0 | 0.750 (ox) | 0.10–2.00 | 0.0300 | 1.490 | [2] |
| β-cyclodextrin-layered poly(diallyldimethylammonium chloride)-graphene composite membrane. | 5.0 | 0.820 (ox) | 0.05–200.0 | 0.0120 | 0.476 | [8] |
| Platinum nanoparticle and CTAB/graphene-composite-modified GCE. | 3.0 | 0.811 (ox) * | 0.08–10.00 | 0.0042 | 0.749 | [17] |
| CTAB/Graphene/MWCNT-modified GCE. | 6.0 | −0.019 (ox) * | 0.01–20.00 | 0.0100 | 0.260 | [21] |
| Chitosan/CaONP/MWCNT/gold electrode. | 7.0 | 1.00 (ox) | 1.99–22.11 | 1.7685 | 1.326 | [40] |
| Polydopamine-coated-MWCNT/GCE. | 6.0 | 0.619 (ox) * | 0.0022–4.64 | 0.0014 | 17.112 | [39] |
| ZnONF/CPE. | 5.0 | 0.691 (ox) * | 0.001–0.02 0.02–0.15 | 0.0002 | 0.0046 | [43] |
| Poly(AAm-co-EMA)/Lac/GCE. | 5.0 | 0.296 (ox) | 0.08–10.00 | 0.0200 | 1.458 | This work |
| Fold-Concentration of Interference Substance | Interference Percentage (%) ± RSD | |||
|---|---|---|---|---|
| Citric Acid | Glucose | Tatrazine | Ascorbic Acid | |
| 1 | 97.1 ± 1.23 | 95.2 ± 0.02 | 101.8 ± 0.140 | 106.6 ± 0.55 |
| 10 | 104.1± 0.38 | 98.5 ± 0.02 | 99.1 ± 1.26 | 100.4 ± 0.76 |
| 50 | 100.3 ±1.25 | 107.2 ± 0.19 | 105.9 ± 0.74 | 95.0 ± 0.44 |
| 100 | 97.8 ± 0.94 | 101.5 ± 0.25 | 96.3 ± 0.30 | 94.4 ± 1.82 |
| Sample | Spiked (μM) | Expected (μM) | Found (μM) | Recovery (%) |
|---|---|---|---|---|
| Soft Drink | 0.00 | - | 14.431 | - |
| 0.08 | 14.511 | 14.745 | 101.6 | |
| 0.80 | 15.231 | 15.466 | 101.5 | |
| 1.00 | 15.431 | 15.642 | 101.4 | |
| 2.00 | 16.431 | 16.274 | 99.0 |
| Sunset Yellow Concentration (μM) | HPLC Method (μM) ± SD | Biosensor Method (μM) ± SD | Relative Error (%) | Correlation Curve |
|---|---|---|---|---|
| 0.08 | 0.065 ± 0.12 | 0.055 ± 0.60 | 18.18 | y = 1.3848x ‒ 0.0917 R2 = 0.9794 |
| 0.80 | 0.683 ± 0.13 | 0.776 ± 0.60 | −11.98 | |
| 1.00 | 0.808 ± 0.34 | 0.952 ± 0.64 | −15.13 | |
| 2.00 | 1.140 ± 0.04 | 1.584 ± 1.20 | −28.03 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rozi, N.; Ahmad, A.; Yook Heng, L.; Shyuan, L.K.; Hanifah, S.A. Electrochemical Sunset Yellow Biosensor Based on Photocured Polyacrylamide Membrane for Food Dye Monitoring. Sensors 2018, 18, 101. https://doi.org/10.3390/s18010101
Rozi N, Ahmad A, Yook Heng L, Shyuan LK, Hanifah SA. Electrochemical Sunset Yellow Biosensor Based on Photocured Polyacrylamide Membrane for Food Dye Monitoring. Sensors. 2018; 18(1):101. https://doi.org/10.3390/s18010101
Chicago/Turabian StyleRozi, Normazida, Amalina Ahmad, Lee Yook Heng, Loh Kee Shyuan, and Sharina Abu Hanifah. 2018. "Electrochemical Sunset Yellow Biosensor Based on Photocured Polyacrylamide Membrane for Food Dye Monitoring" Sensors 18, no. 1: 101. https://doi.org/10.3390/s18010101
APA StyleRozi, N., Ahmad, A., Yook Heng, L., Shyuan, L. K., & Hanifah, S. A. (2018). Electrochemical Sunset Yellow Biosensor Based on Photocured Polyacrylamide Membrane for Food Dye Monitoring. Sensors, 18(1), 101. https://doi.org/10.3390/s18010101




