The Application of Whole Cell-Based Biosensors for Use in Environmental Analysis and in Medical Diagnostics
Abstract
:1. Introduction
2. Using Cells That Can Act as Sensors Against Environmental Analytes
2.1. Detection of Bioavailability
2.2. Reporter Genes
2.3. Regulatory Proteins
2.4. Host Cells
2.5. Multifunctionalization
3. The Use of Sensors Based on Whole Cells in Medical Diagnostics
3.1. Precision Medicine
3.2. Detection of Micronutrients
3.3. Diagnosis of Diseases
4. A Summary on the Use of Whole Cell-Based Biosensors and Their Prospect
Acknowledgments
Conflicts of Interest
References
- Daunert, S.; Barrett, G.; Feliciano, J.S.; Shetty, R.S.; Shrestha, S.; Smithspencer, W. Genetically engineered whole-cell sensing systems: Coupling biological recognition with reporter genes. Cheminform 2000, 100, 2705–2738. [Google Scholar]
- Zhang, W.; Li, X.; Zou, R.; Wu, H.; Shi, H.; Yu, S.; Liu, Y. Multifunctional glucose biosensors from Fe3O4 nanoparticles modified chitosan/graphene nanocomposites. Sci. Rep. 2015, 5, 11129. [Google Scholar] [CrossRef] [PubMed]
- Cheng, R.; Liu, Y.; Ou, S.; Pan, Y.; Zhang, S.; Chen, H.; Dai, L.; Qu, J. Optical turn-on sensors based on graphene oxide for selective detection of D-glucosamine. Anal. Chem. 2012, 84, 5641–5644. [Google Scholar] [CrossRef] [PubMed]
- Wen, W.; Chen, W.; Ren, Q.; Hu, X.; Xiong, H.; Zhang, X.; Wang, S.; Zhao, Y. A highly sensitive nitric oxide biosensor based on hemoglobin–chitosan/graphene–hexadecyltrimethylammonium bromide nanomatrix. Sens. Actuators B Chem. 2012, 166–167, 444–450. [Google Scholar] [CrossRef]
- Zhang, W.; Chen, C.; Yang, D.; Dong, G.; Jia, S.; Zhao, B.; Yan, L.; Yao, Q.; Sunna, A.; Liu, Y. Optical biosensors based on nitrogen-doped graphene functionalized with magnetic nanoparticles. Adv. Mater. Interfaces 2016, 3, 1600590. [Google Scholar] [CrossRef]
- Pancrazio, J.J.; Whelan, J.P.; Borkholder, D.A.; Ma, W.; Stenger, D.A. Development and application of cell-based biosensors. Ann. Biomed. Eng. 1999, 27, 697–711. [Google Scholar] [CrossRef] [PubMed]
- Bousse, L. Whole cell biosensors. Sens. Actuators B Chem. 1996, 34, 270–275. [Google Scholar] [CrossRef]
- Du, H.; Strohsahl, C.M.; Camera, J.; Miller, B.L.; Krauss, T.D. Sensitivity and specificity of metal surface-immobilized “molecular beacon” biosensors. J. Am. Chem. Soc. 2005, 127, 7932–7940. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Wu, C.; Cai, H.; Hu, N.; Zhou, J.; Wang, P. Cell-based biosensors and their application in biomedicine. Chem. Rev. 2014, 114, 6423–6461. [Google Scholar] [CrossRef] [PubMed]
- Raut, N.; O’Connor, G.; Pasini, P.; Daunert, S. Engineered cells as biosensing systems in biomedical analysis. Anal. Bioanal. Chem. 2012, 402, 3147–3159. [Google Scholar] [CrossRef] [PubMed]
- Ben-Yoav, H.; Biran, A.; Pedahzur, R.; Belkin, S.; Buchinger, S.; Reifferscheid, G.; Shacham-Diamand, Y. A whole cell electrochemical biosensor for water genotoxicity bio-detection. Electrochim. Acta 2009, 54, 6113–6118. [Google Scholar] [CrossRef]
- Tian, Y.; Lu, Y.; Xu, X.; Wang, C.; Zhou, T.; Li, X. Construction and comparison of yeast whole-cell biosensors regulated by two RAD54 promoters capable of detecting genotoxic compounds. Toxicol. Mech. Methods 2017, 27, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Behzadian, F.; Barjeste, H.; Hosseinkhani, S.; Zarei, A.R. Construction and characterization of Escherichia coli whole-cell biosensors for toluene and related compounds. Curr. Microbiol. 2011, 62, 690–696. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Stüben, D.; Berner, Z.; Albers, J.; Hintsche, R.; Jantzen, E. Development of an ultramicroelectrode arrays (UMEAs) sensor for trace heavy metal measurement in water. Sens. Actuators B Chem. 2004, 97, 168–173. [Google Scholar] [CrossRef]
- Gardner, G.T.; Stern, P.C. Environmental Problems and Human Behavior; Allyn & Bacon: Boston, MA, USA, 1996. [Google Scholar]
- Pulido, M.D.; Parrish, A.R. Metal-induced apoptosis: Mechanisms. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2003, 533, 227–241. [Google Scholar] [CrossRef]
- Derraik, J.G. The pollution of the marine environment by plastic debris: A review. Mar. Pollut. Bull. 2002, 44, 842–852. [Google Scholar] [CrossRef]
- Turdean, G.L. Design and development of biosensors for the detection of heavy metal toxicity. Int. J. Electrochem. 2011. [Google Scholar] [CrossRef]
- Tüzen, M. Determination of heavy metals in soil, mushroom and plant samples by atomic absorption spectrometry. Microchem. J. 2003, 74, 289–297. [Google Scholar] [CrossRef]
- Noll, M.R. Trace elements in terrestrial environments: Biogeochemistry, bioavailability, and risks of metals. J. Environ. Qual. 2003, 32, 374. [Google Scholar] [CrossRef]
- Bosma, T.N.; Middeldorp, P.J.; Schraa, G.; Zehnder, A.J. Mass transfer limitation of biotransformation: Quantifying bioavailability. Environ. Sci. Technol. 1996, 31, 248–252. [Google Scholar] [CrossRef]
- Sanseverino, J.; Dunbar, P.; Larimer, F.; Sayler, G. Rapid, sensitive bioluminescent reporter technology for naphthalene exposure and biodegradation. Biophys. J. 1986, 49, 79. [Google Scholar]
- Andreescu, S.; Sadik, O.A. Trends and challenges in biochemical sensors for clinical and environmental monitoring. Pure Appl. Chem. 2004, 76, 861–878. [Google Scholar] [CrossRef]
- Belkin, S. Microbial whole-cell sensing systems of environmental pollutants. Curr. Opin. Microbiol. 2003, 6, 206–212. [Google Scholar] [CrossRef]
- Amaro, F.; Turkewitz, A.P.; Gutiérrez, J.C. Whole-cell biosensors for detection of heavy metal ions in environmental samples based on metallothionein promoters from Tetrahymena thermophila. Microb. Biotechnol. 2011, 4, 513–522. [Google Scholar] [CrossRef] [PubMed]
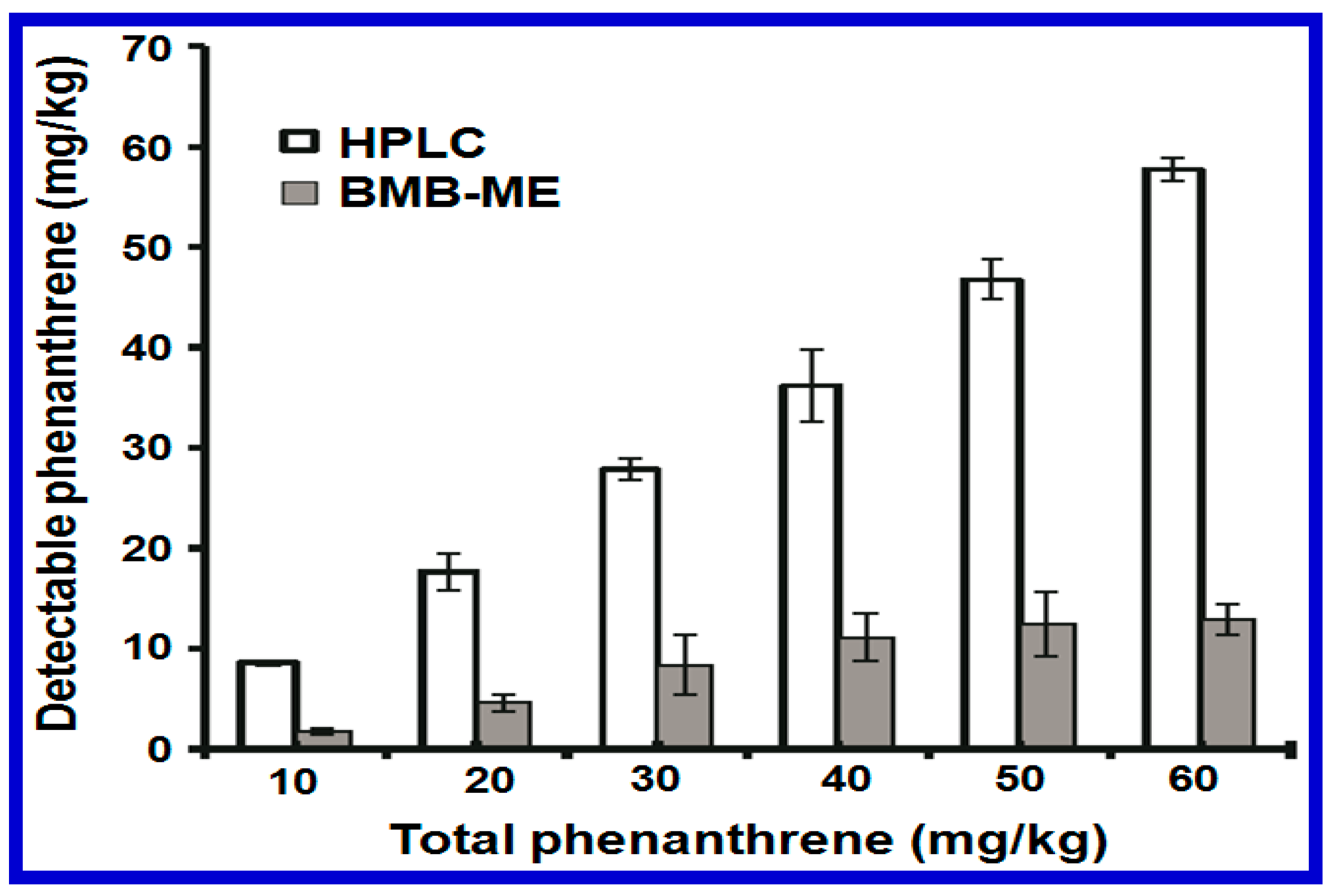
- Wei, H.; Ze-Ling, S.; Le-Le, C.; Wen-Hui, Z.; Chuan-Chao, D. Specific detection of bioavailable phenanthrene and mercury by bacterium reporters in the red soil. Int. J. Environ. Sci. Technol. 2014, 11, 685–694. [Google Scholar] [CrossRef]
- Peltola, P.; Ivask, A.; Åström, M.; Virta, M. Lead and Cu in contaminated urban soils: Extraction with chemical reagents and bioluminescent bacteria and yeast. Sci. Total Environ. 2005, 350, 194–203. [Google Scholar] [CrossRef] [PubMed]
- Ecken, H.; Ingebrandt, S.; Krause, M.; Richter, D.; Hara, M.; Offenhäusser, A. 64-Channel extended gate electrode arrays for extracellular signal recording. Electrochim. Acta 2003, 48, 3355–3362. [Google Scholar] [CrossRef]
- Hakkila, K.; Maksimow, M.; Karp, M.; Virta, M. Reporter genes lucFF, luxCDABE, gfp, and dsred have different characteristics in whole-cell bacterial sensors. Anal. Biochem. 2000, 301, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.C.; Amaro, F.; Martín-González, A. Heavy metal whole-cell biosensors using eukaryotic microorganisms: An updated critical review. Front. Microbiol. 2015, 6, 1–8. [Google Scholar]
- Sagi, E.; Hever, N.; Rosen, R.; Bartolome, A.J.; Premkumar, J.R.; Ulber, R.; Lev, O.; Scheper, T.; Belkin, S. Fluorescence and bioluminescence reporter functions in genetically modified bacterial sensor strains. Sens. Actuators B Chem. 2003, 90, 2–8. [Google Scholar] [CrossRef]
- Mascher, T.; Zimmer, S.L.; Smith, T.A.; Helmann, J.D. Antibiotic-inducible promoter regulated by the cell envelope stress-sensing two-component system LiaRS of Bacillus subtilis. Antimicrob. Agents Chemother. 2004, 48, 2888–2896. [Google Scholar] [CrossRef] [PubMed]
- Chong, H.; Ching, C.B. Development of colorimetric-based whole-cell biosensor for organophosphorus compounds by engineering transcription regulator DmpR. ACS Synth. Biol. 2016, 5, 1290–1298. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.K.; Grimaud, R.; Hoskins, J.R.; Wickner, S.; Maurizi, M.R. Unfolding and internalization of proteins by the ATP-dependent proteases ClpXP and ClpAP. Proc. Natl. Acad. Sci. USA 2000, 97, 8898–8903. [Google Scholar] [CrossRef] [PubMed]
- Stocker, J.; Balluch, D.; Gsell, M.; Harms, H.; Feliciano, J.; Daunert, S.; van der Meer, J.R. Development of a set of simple bacterial biosensors for quantitative and rapid measurements of arsenite and arsenate in potable water. Environ. Sci. Technol. 2003, 37, 4743–4750. [Google Scholar] [CrossRef] [PubMed]
- Yagi, K. Applications of whole-cell bacterial sensors in biotechnology and environmental science. Appl. Microbiol. Biotechnol. 2007, 73, 1251–1258. [Google Scholar] [CrossRef] [PubMed]
- Naylor, L.H. Reporter gene technology: The future looks bright. Biochem. Pharmacol. 1999, 58, 749–757. [Google Scholar] [CrossRef]
- Joyeux, A.; Balaguer, P.; Germain, P.; Boussioux, A.M.; Pons, M.; Nicolas, J.C. Engineered cell lines as a tool for monitoring biological activity of hormone analogs. Anal. Biochem. 1997, 249, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Welsh, S.; Kay, S.A. Reporter gene expression for monitoring gene transfer. Curr. Opin. Biotechnol. 1997, 8, 617–622. [Google Scholar] [CrossRef]
- Jain, V.K.; Magrath, I.T. A chemiluminescent assay for quantitation of β-galactosidase in the femtogram range: Application to quantitation of β-galactosidase in lacZ-transfected cells. Anal. Biochem. 1991, 199, 119–124. [Google Scholar] [CrossRef]
- Biran, I.; Klimentiy, L.; Hengge-Aronis, R.; Ron, E.Z.; Rishpon, J. On-line monitoring of gene expression. Microbiology 1999, 145, 2129–2133. [Google Scholar] [CrossRef] [PubMed]
- Karttunen, J.; Shastri, N. Measurement of ligand-induced activation in single viable T cells using the lacZ reporter gene. Proc. Natl. Acad. Sci. USA 1991, 88, 3972–3976. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, H.; Wakabayashi, M.; Yamashiro, H.; Maeda, I.; Isoda, K.; Kondoh, M.; Kawase, M.; Miyasaka, H.; Yagi, K. Whole-cell arsenite biosensor using photosynthetic bacterium Rhodovulum sulfidophilum. Appl. Microbiol. Biotechnol. 2006, 73, 332–338. [Google Scholar] [CrossRef] [PubMed]
- Yeliseev, A.A.; Eraso, J.M.; Kaplan, S. Differential carotenoid composition of the B875 and B800-850 photosynthetic antenna complexes in Rhodobacter sphaeroides 2.4.1: Involvement of spheroidene and spheroidenone in adaptation to changes in light intensity and oxygen availability. J. Bacteriol. 1996, 178, 5877–5883. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Asad, S.; Ali, A. Bioluminescent bioreporter for assessment of arsenic contamination in water samples of India. J. Biosci. 2013, 38, 251–258. [Google Scholar] [CrossRef] [PubMed]
- De Mora, K.; Joshi, N.; Balint, B.L.; Ward, F.B.; Elfick, A.; French, C.E. A pH-based biosensor for detection of arsenic in drinking water. Anal. Bioanal. Chem. 2011, 400, 1031–1039. [Google Scholar] [CrossRef] [PubMed]
- Joe, M.H.; Lee, K.H.; Lim, S.Y.; Im, S.H.; Song, H.P.; Lee, I.S.; Kim, D.H. Pigment-based whole-cell biosensor system for cadmium detection using genetically engineered Deinococcus radiodurans. Bioprocess Biosyst. Eng. 2012, 35, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Branco, R.; Cristóvão, A.; Morais, P.V. Highly sensitive, highly specific whole-cell bioreporters for the detection of chromate in environmental samples. PLoS ONE 2013, 8, e54005. [Google Scholar] [CrossRef] [PubMed]
- Ravikumar, S.; Ganesh, I.; Yoo, I.K.; Hong, S.H. Construction of a bacterial biosensor for zinc and copper and its application to the development of multifunctional heavy metal adsorption bacteria. Process Biochem. 2012, 47, 758–765. [Google Scholar] [CrossRef]
- Willardson, B.M.; Wilkins, J.F.; Rand, T.A.; Schupp, J.M.; Hill, K.K.; Keim, P.; Jackson, P.J. Development and testing of a bacterial biosensor for toluene-based environmental contaminants. Appl. Environ. Microbiol. 1998, 64, 1006–1012. [Google Scholar] [PubMed]
- Tecon, R.; Beggah, S.; Czechowska, K.; Sentchilo, V.; Chronopoulou, P.M.; McGenity, T.J.; Van der Meer, J.R. Development of a multistrain bacterial bioreporter platform for the monitoring of hydrocarbon contaminants in marine environments. Environ. Sci. Technol. 2009, 44, 1049–1055. [Google Scholar] [CrossRef] [PubMed]
- Shingler, V.; Moore, T. Sensing of aromatic compounds by the DmpR transcriptional activator of phenol-catabolizing Pseudomonas sp. strain CF600. J. Bacteriol. 1994, 176, 1555–1560. [Google Scholar] [CrossRef] [PubMed]
- Sticher, P.; Jaspers, M.C.; Stemmler, K.; Harms, H.; Zehnder, A.J.; Van Der Meer, J.R. Development and characterization of a whole-cell bioluminescent sensor for bioavailable middle-chain alkanes in contaminated groundwater samples. Appl. Environ. Microbiol. 1997, 63, 4053–4060. [Google Scholar] [PubMed]
- Korpela, M.T.; Kurittu, J.S.; Karvinen, J.T.; Karp, M.T. A recombinant Escherichia coli sensor strain for the detection of tetracyclines. Anal. Chem. 1998, 70, 4457–4462. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, S.I.; Oda, Y.; Shimada, T.; Oki, I.; Sugimoto, K. SOS-inducing activity of chemical carcinogens and mutagens in Salmonella typhimurium TA1535/pSK1002: Examination with 151 chemicals. Mutat. Res. Lett. 1987, 192, 239–246. [Google Scholar] [CrossRef]
- Pontel, L.; Audero, M.M.; Checa, S.; Soncini, F. GolS controls the response to gold by the hierarchical induction of Salmonella-specific genes that include a CBA efflux-coding operon. Mol. Microbiol. 2007, 66, 814–825. [Google Scholar] [CrossRef] [PubMed]
- Cerminati, S.; Soncini, F.C.; Checa, S.K. A sensitive whole-cell biosensor for the simultaneous detection of a broad-spectrum of toxic heavy metal ions. Chem. Commun. 2015, 51, 5917–5920. [Google Scholar] [CrossRef] [PubMed]
- Shang, Y.; Song, X.; Bowen, J.; Corstanje, R.; Gao, Y.; Gaertig, J.; Gorovsky, M.A. A robust inducible-repressible promoter greatly facilitates gene knockouts, conditional expression, and overexpression of homologous and heterologous genes in Tetrahymena thermophila. Proc. Natl. Acad. Sci. USA 2002, 99, 3734. [Google Scholar] [CrossRef] [PubMed]
- Díaz, S.; Amaro, F.; Rico, D.; Campos, V.; Benítez, L.; Martíngonzález, A.; Hamilton, E.P.; Orias, E.; Gutiérrez, J.C. Tetrahymena metallothioneins fall into two discrete subfamilies. PLoS ONE 2007, 2, e291. [Google Scholar] [CrossRef] [PubMed]
- Whangsuk, W.; Dubbs, J.M.; Sallabhan, R.; Somsongkul, K.; Mongkolsuk, S.; Loprasert, S. ChpR is a chlorpyrifos-responsive transcription regulator in Sinorhizobium meliloti. J. Mol. Microbiol. Biotechnol. 2010, 18, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Whangsuk, W.; Thiengmag, S.; Dubbs, J.; Mongkolsuk, S.; Loprasert, S. Specific detection of the pesticide, chlorpyrifos, by a sensitive genetic-based whole cell biosensor. Anal. Biochem. 2015, 493, 11–13. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Rosen, B.P. The ArsR protein is a trans-acting regulatory protein. Mol. Microbiol. 1991, 5, 1331–1336. [Google Scholar] [CrossRef] [PubMed]
- Gross, M. Antibiotics in crisis. Curr. Biol. CB 2013, 23, R1063. [Google Scholar] [CrossRef] [PubMed]
- Espinosa-Urgel, M.; Serrano, L.; Ramos, J.L.; Fernández-Escamilla, A.M. Engineering biological approaches for detection of toxic compounds: A new microbial biosensor based on the pseudomonas putida TtgR repressor. Mol. Biotechnol. 2015, 57, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Magrisso, S.; Erel, Y.; Belkin, S. Microbial reporters of metal bioavailability. Microb. Biotechnol. 2008, 1, 320–330. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Sánchez, V.; Molina, L.; Ramos, J.L.; Segura, A. New family of biosensors for monitoring BTX in aquatic and edaphic environments. Microb. Biotechnol. 2016, 9, 858–867. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; He, Y.; Wang, Y.; Wang, H.; Wu, L.; Aries, E.; Huang, W.E. Whole-cell bacterial bioreporter for actively searching and sensing of alkanes and oil spills. Microb. Biotechnol. 2012, 5, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Throne-Holst, M.; Wentzel, A.; Ellingsen, T.E.; Kotlar, H.K.; Zotchev, S.B. Identification of novel genes involved in long-chain n-alkane degradation by Acinetobacter sp. strain DSM 17874. Appl. Environ. Microbiol. 2007, 73, 3327–3332. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak, A.; Geissdörfer, W.; Hillen, W. Expression of alkane hydroxylase from Acinetobacter sp. Strain ADP1 is induced by a broad range of n-alkanes and requires the transcriptional activator AlkR. J. Bacteriol. 1998, 180, 5822–5827. [Google Scholar] [PubMed]
- Sevilla, E.; Yuste, L.; Rojo, F. Marine hydrocarbonoclastic bacteria as whole-cell biosensors for n-alkanes. Microb. Biotechnol. 2015, 8, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Brutesco, C.; Prévéral, S.; Escoffier, C.; Descamps, E.C.T.; Prudent, E.; Cayron, J.; Dumas, L.; Ricquebourg, M.; Adryanczyk-Perrier, G.; Groot, A.D. Bacterial host and reporter gene optimization for genetically encoded whole cell biosensors. Environ. Sci. Pollut. Res. Int. 2016, 24, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Cayron, J.; Prudent, E.; Escoffier, C.; Gueguen, E.; Mandrandberthelot, M.A.; Pignol, D.; Garcia, D.; Rodrigue, A. Pushing the limits of nickel detection to nanomolar range using a set of engineered bioluminescent Escherichia coli. Environ. Sci. Pollut. Res. 2017, 24, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Aisyah, W.N.; Jusoh, W.; Wong, L.S. Exploring the Potential of Whole Cell Biosensor: A Review in Environmental Applications. Int. J. Chem. Environ. Biol. Sci. 2014, 2, 52–56. [Google Scholar]
- Bazin, I.; Seo, H.B.; Suehs, C.M.; Ramuz, M.; De, W.M.; Gu, M.B. Profiling the biological effects of wastewater samples via bioluminescent bacterial biosensors combined with estrogenic assays. Environ. Sci. Pollut. Res. 2016, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Tauriainen, S.; Karp, M.; Chang, W.; Virta, M. Recombinant luminescent bacteria for measuring bioavailable arsenite and antimonite. Appl. Environ. Microbiol. 1997, 63, 4456–4461. [Google Scholar] [PubMed]
- Anne, K.; Taisia, R.; Angela, I. A suite of recombinant luminescent bacterial strains for the quantification of bioavailable heavy metals and toxicity testing. BMC Biotechnol. 2009, 9, 41. [Google Scholar]
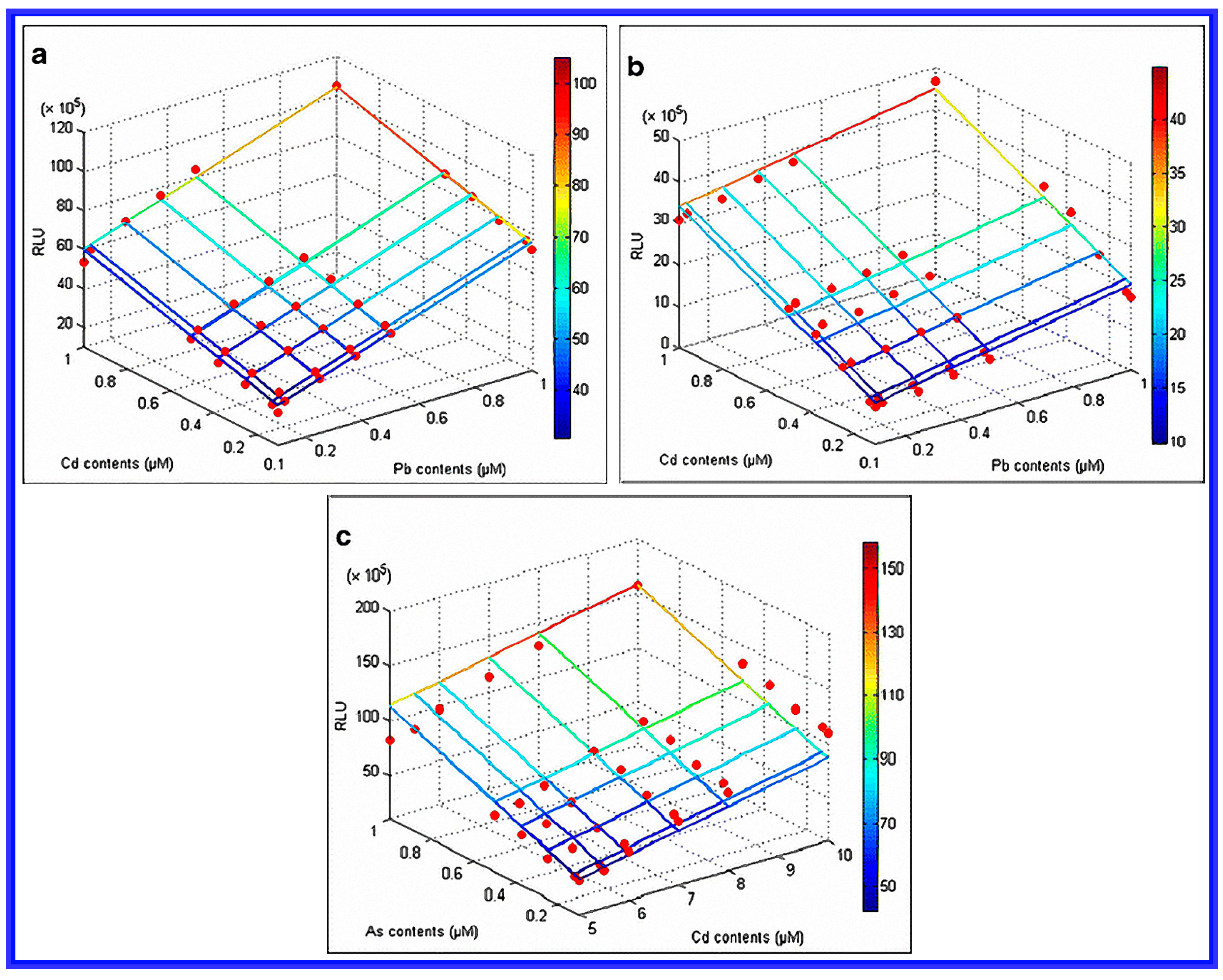
- Hou, Q.; Ma, A.; Wang, T.; Lin, J.; Wang, H.; Du, B.; Zhuang, X.; Zhuang, G. Detection of bioavailable cadmium, lead, and arsenic in polluted soil by tailored multiple Escherichia coli whole-cell sensor set. Anal. Bioanal. Chem. 2015, 407, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.C.; Youn, C.H.; Ahn, J.M.; Gu, M.B. Screening of Target-Specific Stress-Responsive Genes for the Development of Cell-Based Biosensors Using a DNA Microarray. Anal. Chem. 2005, 77, 8020–8026. [Google Scholar] [CrossRef] [PubMed]
- Reed, B.; Blazeck, J.; Alper, H. Evolution of an alkane-inducible biosensor for increased responsiveness to short-chain alkanes. J. Biotechnol. 2012, 158, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Camanzi, L.; Bolelli, L.; Maiolini, E.; Girotti, S.; Matteuzzi, D. Optimal conditions for stability of photoemission and freeze drying of two luminescent bacteria for use in a biosensor. Environ. Toxicol. Chem. 2011, 30, 801. [Google Scholar] [CrossRef] [PubMed]
- Prévéral, S.; Brutesco, C.; Descamps, E.C.T.; Escoffier, C.; Pignol, D.; Ginet, N.; Garcia, D. A bioluminescent arsenite biosensor designed for inline water analyzer. Environ. Sci. Pollut. Res. 2017, 24, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Elad, T.; Almog, R.; Yagur-Kroll, S.; Levkov, K.; Melamed, S.; Shacham-Diamand, Y.; Belkin, S. Online monitoring of water toxicity by use of bioluminescent reporter bacterial biochips. Environ. Sci. Technol. 2011, 45, 8536–8544. [Google Scholar] [CrossRef] [PubMed]
- Wolf, D.; Mascher, T. The applied side of antimicrobial peptide-inducible promoters from Firmicutes bacteria: Expression systems and whole-cell biosensors. Appl. Microbiol. Biotechnol. 2016, 100, 4817–4829. [Google Scholar] [CrossRef] [PubMed]
- Hillger, J.M.; Schoop, J.; Boomsma, D.I.; Slagboom, P.E.; Ijzerman, A.P.; Heitman, L.H. Whole-cell biosensor for label-free detection of GPCR-mediated drug responses in personal cell lines. Biosens. Bioelectron. 2015, 74, 233–242. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Castracane, J.; Tokranova, N.; Gracias, A.; Lnenicka, G.; Szaro, B. A living cell-based biosensor utilizing G-protein coupled receptors: Principles and detection methods. Biosens. Bioelectron. 2007, 22, 3230–3237. [Google Scholar] [CrossRef] [PubMed]
- Zager, V.; Cemazar, M.; Hreljac, I.; Lah, T.T.; Sersa, G.; Filipic, M. Development of human cell biosensor system for genotoxicity detection based on DNA damage-induced gene expression. Radiol. Oncol. 2010, 44, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Powers, H.J. Riboflavin (vitamin B-2) and health. Am. J. Clin. Nutr. 2003, 77, 1352. [Google Scholar] [PubMed]
- Prchal, J.T.; Conrad, M.E.; Skalka, H.W. Association of presenile cataracts with heterozygosity for galactosaemic states and with riboflavin deficiency. Lancet 1978, 1, 12. [Google Scholar] [CrossRef]
- Cardoso, D.R.; Libardi, S.H.; Skibsted, L.H. Riboflavin as a photosensitizer. Effects on human health and food quality. Food Funct. 2012, 3, 487. [Google Scholar] [CrossRef] [PubMed]
- Besaratinia, A.; Kim, S.I.; Bates, S.E.; Pfeifer, G.P. Riboflavin activated by ultraviolet A1 irradiation induces oxidative DNA damage-mediated mutations inhibited by Vitamin C. Proc. Natl. Acad. Sci. USA 2007, 104, 5953–5958. [Google Scholar] [CrossRef] [PubMed]
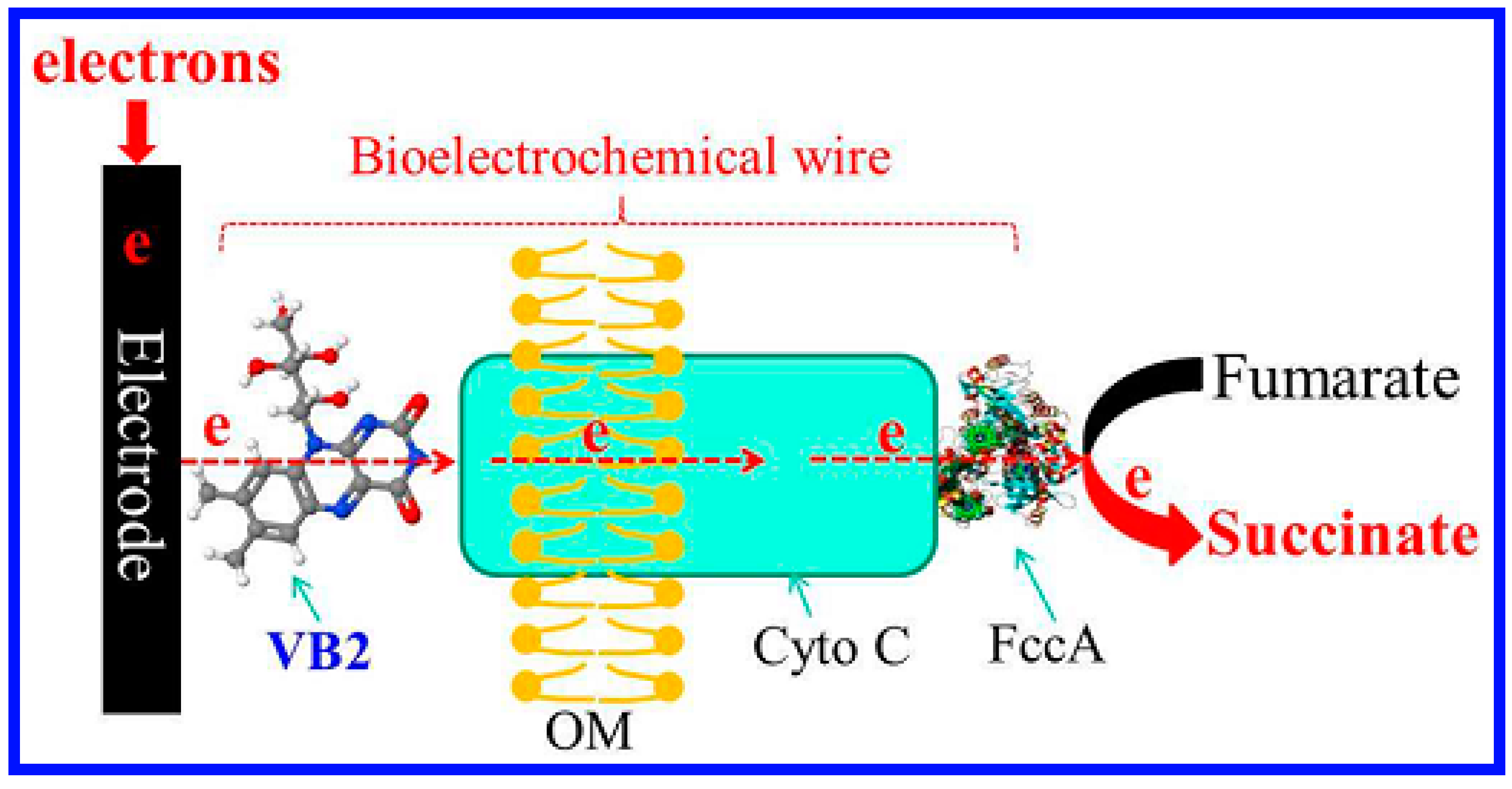
- Si, R.; Yang, Y.; Yu, Y.; Han, S.; Zhang, C.; Sun, D.; Zhai, D.; Liu, X.; Yong, Y. Wiring bacterial electron flow for sensitive whole-cell amperometric detection of riboflavin. Anal. Chem. 2016, 88, 11222–11228. [Google Scholar] [CrossRef] [PubMed]
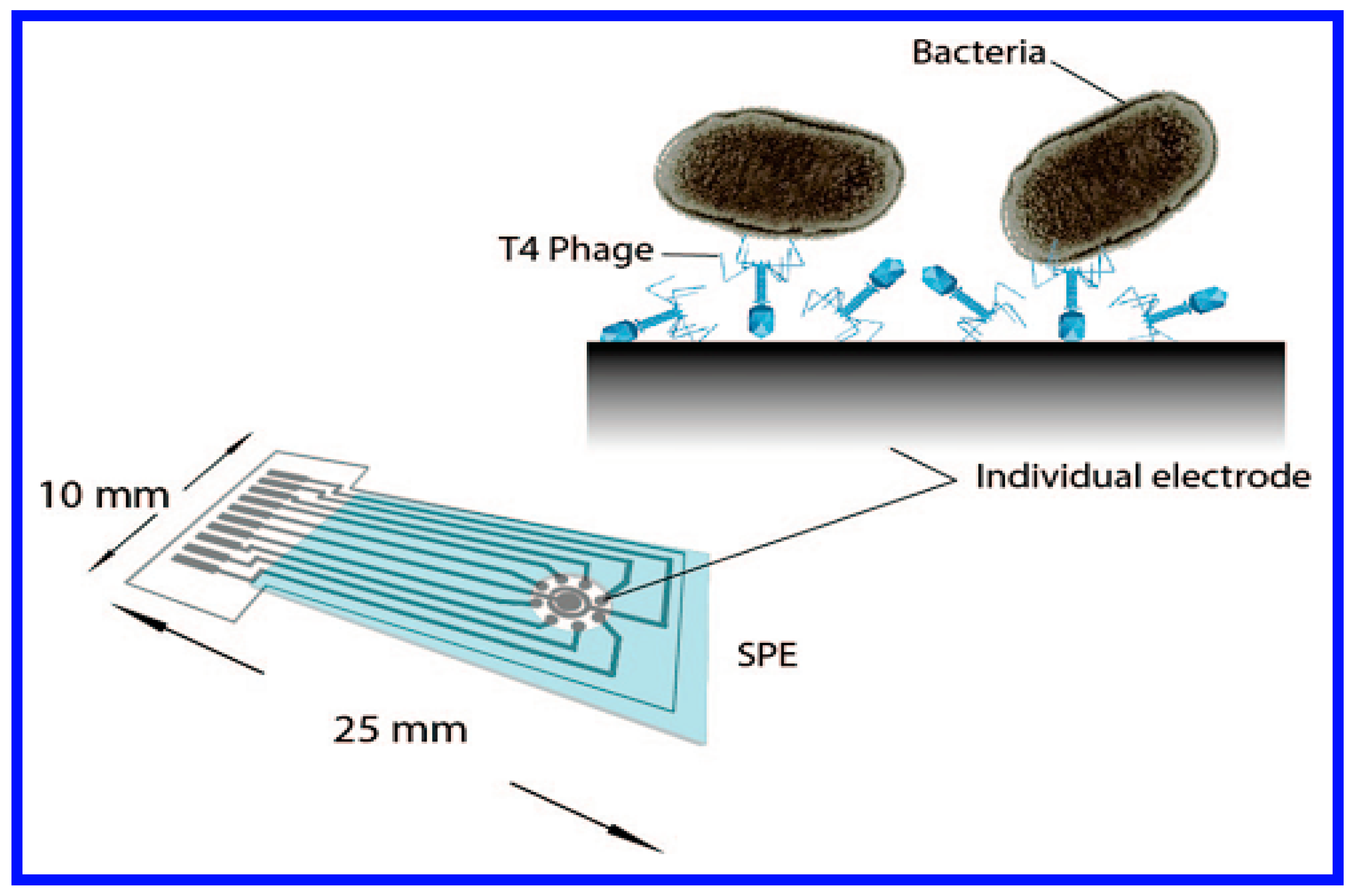
- Shabani, A.; Zourob, M.; Allain, B.; Marquette, C.A.; Lawrence, M.F.; Mandeville, R. Bacteriophage-modified microarrays for the direct impedimetric detection of bacteria. Anal. Chem. 2008, 80, 9475–9482. [Google Scholar] [CrossRef] [PubMed]
- Sifri, C.D. Healthcare epidemiology: Quorum sensing: Bacteria talk sense. Clin. Infect. Dis. 2008, 47, 1070–1076. [Google Scholar] [CrossRef] [PubMed]
- Winson, M.K. Construction and analysis of luxCDABE-based plasmid sensors for investigating N-acyl homoserine lactone-mediated quorum sensing. Fems Microbiol. Lett. 1998, 163, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Kumari, A.; Pasini, P.; Deo, S.K.; Flomenhoft, D.; Shashidhar, H.; Daunert, S. Biosensing systems for the detection of bacterial quorum signaling molecules. Anal. Chem. 2006, 78, 7603–7609. [Google Scholar] [CrossRef] [PubMed]
- Pearson, J.P.; Passador, L.; Iglewski, B.H.; Greenberg, E.P. A second N-acylhomoserine lactone signal produced by pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 1995, 92, 1490–1494. [Google Scholar] [CrossRef] [PubMed]
- Susan, C.; Morpeth, H.O.R.; Crump, J.A. Invasive non-Typhi Salmonella disease in Africa. Clin. Infect. Dis. 2009, 49, 606–611. [Google Scholar]
- Venkatesh, A.G.; Sun, A.; Brickner, H.; Looney, D.; Hall, D.A.; Aronoff-Spencer, E. Yeast dual-affinity biobricks: Progress towards renewable whole-cell biosensors. Biosens. Bioelectron. 2015, 70, 462–468. [Google Scholar] [CrossRef] [PubMed]
- Bereza-Malcolm, L.T.; Mann, G.; Franks, A.E. Environmental sensing of heavy metals through whole cell microbial biosensors: A synthetic biology approach. ACS Synth. Biol. 2004, 4, 535–546. [Google Scholar] [CrossRef] [PubMed]
- Ismail, A.B.M.; Yoshinobu, T.; Iwasaki, H.; Sugihara, H.; Yukimasa, T.; Hirata, I.; Iwata, H. Investigation on light-addressable potentiometric sensor as a possible cell–semiconductor hybrid. Biosens. Bioelectron. 2003, 18, 1509–1514. [Google Scholar] [CrossRef]
- Horry, H.; Charrier, T.; Durand, M.J.; Vrignaud, B.; Picart, P.; Daniel, P.; Thouand, G. Technological conception of an optical biosensor with a disposable card for use with bioluminescent bacteria. Sens. Actuators B Chem. 2007, 122, 527–534. [Google Scholar] [CrossRef]
- Lu, Y.; Macias, D.; Dean, Z.S.; Kreger, N.R. A UAV-mounted whole cell biosensor system for environmental monitoring applications. IEEE Trans. Nanobiosci. 2015, 14, 811–817. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, S.F. Microbial biosensors. Biosens. Bioelectron. 2001, 16, 337–353. [Google Scholar] [CrossRef]



| Gene | Detection Method | Advantages | Disadvantages |
|---|---|---|---|
| lux [29] | Bioluminescence | Easy measurement, rapid response | Thermal lability, O2 requirement |
| luc [30] | Bioluminescence | High sensitivity, rapid response, thermal stability | O2 and ATP requirements, low permeability |
| gfp [31] | Fluorescence | No substrate requirement, high stability | Low sensitivity, lag-time for stable fluorescence, autofluorescence |
| lacZ [32] | Bioluminescence, Fluorescence, Colorimetry, Electrochemistry | High stability, wide variety of detection methods, detection by naked eyes | Substrate dependent, low permeability |
| crtA [33] | Colorimetry | Detection by naked eyes | Substrate dependent |
| Host Chassis | Reporter Gene | Target Analyte | Detection Sensitivity | Reference |
|---|---|---|---|---|
| E. coli | luxCDABE | arsenic | 0.74–69 μg/L | [45] |
| E. coli | lacZ | arsentate | <10 μg/L | [46] |
| D. radiodurans | lacZ | cadmium | 1–10 mM | [47] |
| crtI | 50 nM–1 mM | |||
| E. coli | Gap | chromate | 100 nM | [48] |
| E. coli | Gfp | zinc | 16 μM | [49] |
| copper | 26 μM | |||
| Escherichia coli | Luc | benzene, toluene and xylene | 40 μM | [50] |
| E. coli | luxAB | benzene, toluene and xylene | 0.24 μM | [51] |
| P. putida | luxAB | phenol | 3 μM | [52] |
| B. sartisoli | luxAB | naphthalene and phenanthrene | 0.17 μM | [51] |
| E. coli | luxAB | C6–C10 alkanes | 10 nM | [53] |
| E. coli | luxCDABE | tetracyclines | 45 nM | [54] |
| S. typhimurium | lacZ | single-stranded DNA | 10 nM mitomycinC | [55] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gui, Q.; Lawson, T.; Shan, S.; Yan, L.; Liu, Y. The Application of Whole Cell-Based Biosensors for Use in Environmental Analysis and in Medical Diagnostics. Sensors 2017, 17, 1623. https://doi.org/10.3390/s17071623
Gui Q, Lawson T, Shan S, Yan L, Liu Y. The Application of Whole Cell-Based Biosensors for Use in Environmental Analysis and in Medical Diagnostics. Sensors. 2017; 17(7):1623. https://doi.org/10.3390/s17071623
Chicago/Turabian StyleGui, Qingyuan, Tom Lawson, Suyan Shan, Lu Yan, and Yong Liu. 2017. "The Application of Whole Cell-Based Biosensors for Use in Environmental Analysis and in Medical Diagnostics" Sensors 17, no. 7: 1623. https://doi.org/10.3390/s17071623
APA StyleGui, Q., Lawson, T., Shan, S., Yan, L., & Liu, Y. (2017). The Application of Whole Cell-Based Biosensors for Use in Environmental Analysis and in Medical Diagnostics. Sensors, 17(7), 1623. https://doi.org/10.3390/s17071623




