Dimension Reduction Aided Hyperspectral Image Classification with a Small-sized Training Dataset: Experimental Comparisons
Abstract
:1. Introduction
- (a)
- Well defined training samples: the training data should well represent the data to be classified;
- (b)
- Feature determination: features should maximally reflect the data information while with an appropriate dimension;
- (c)
- Appropriate classification algorithms: the adopted algorithms should accommodate the volume of training data and the dimension of pattern vector.
- (1)
- There are generally hundreds of bands in HSI (although not all of them are useful) while only a limited number of training samples are available;
- (2)
- (3)
- Certain bands are dominated by sensor noises or may be not relevant to specific features so may contain little useful information for a task of concern.
- (1)
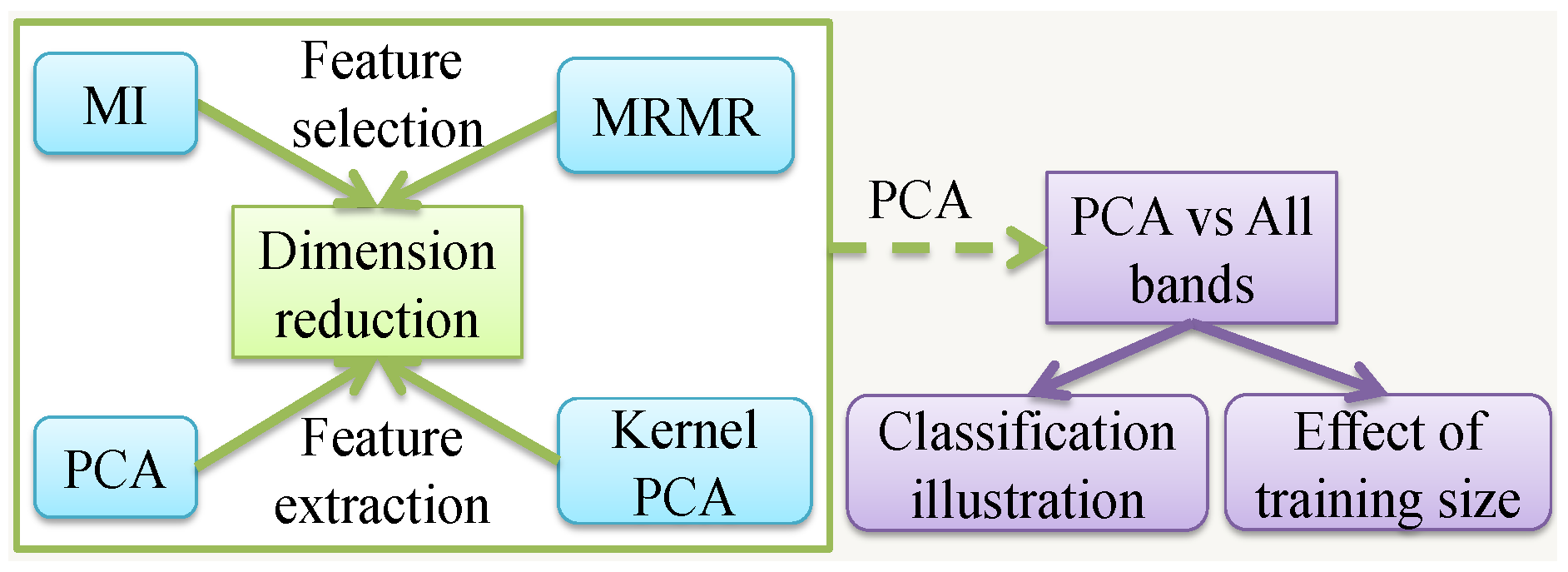
- Different dimension reduction techniques including feature selection and feature extraction algorithms are compared for HSI classification, where the most suitable one is identified, namely SVM with PCA.
- (2)
- Comparatively experimental results on a real HSI dataset demonstrate that by augmenting SVM with dimension reduction techniques (i.e., PCA), good or even better classification performance can be achieved, particularly when the size of training data is small.
- (3)
- More importantly, it is discovered that reducing feature dimension can substantially simplify the SVM models and so reduce classification time while preserving performance, which is vital for real-time remote sensing applications.
2. Problem Formulation and Motivations
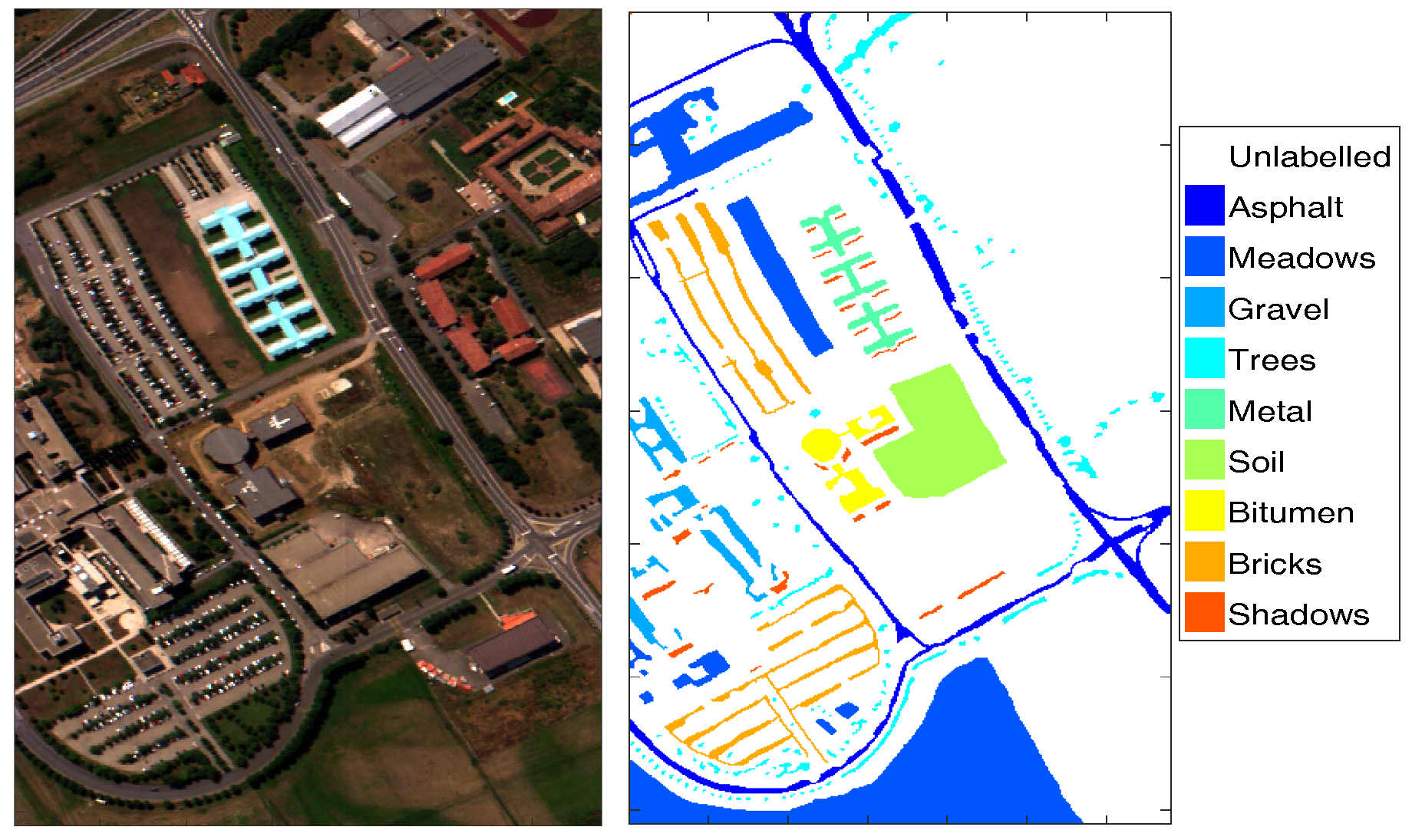
2.1. HSI Dataset
2.2. Problem Formulation
2.2.1. Classification Problem
- (1)
- Overall Accuracy (OA): the percentage of correctly classified pixels;
- (2)
- Average Accuracy (AA): the mean of the percentages of correctly classified pixels for each class;
- (3)
- Kappa coefficient: the percentage of correctly classified pixels corrected by the number of agreements that would be expected purely by chance.
2.2.2. SVM with Kernel
2.3. Motivations Via Band Analysis
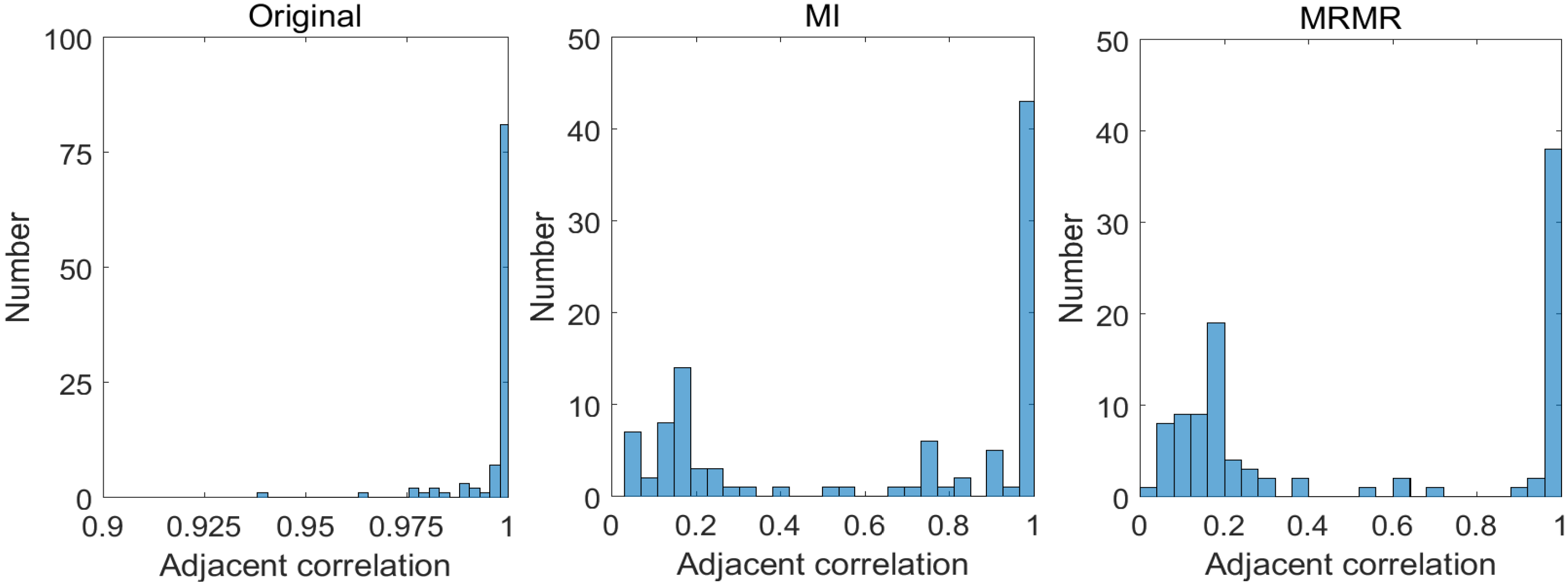
2.3.1. Correlations between Bands
- (1)
- Bands that are spectrally “near” to each other (i.e., close to matrix diagonal) tend to be highly correlated (brighter);
- (2)
- There is an obvious dark rectangle (x-coordinate: bands No. 70–No. 103, y-coordinate: bands No. 1–No. 70), this is because bands No. 1 to No. 70 correspond to wavelength interval 430 nm to 700 nm (i.e., RGB visible light region) while No. 70 to No. 103 are wavelength interval 700 nm to 860 nm (i.e., NIR region), and the correlations between visible light and NIR are quite low;
- (3)
- There are some grey areas showing the correlations for Red/Green, Red/Blue, Green/Blue, where the correlations of Red/Green and Red/Blue are slightly lower than Green/Blue. There are also four “bright” squares showing the correlations of bands within different channels.
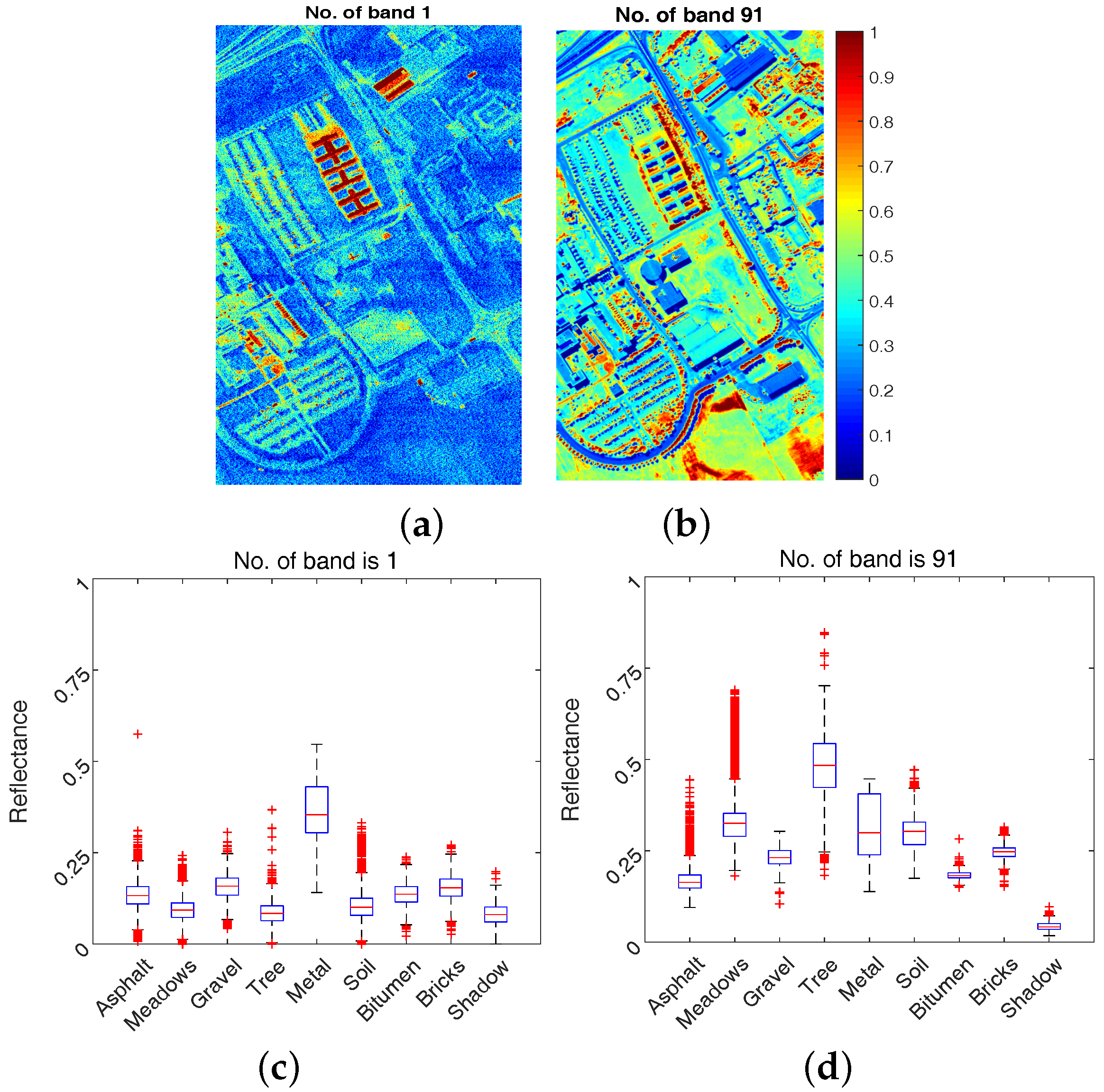
2.3.2. Sample Bands with Box Plot
2.3.3. Motivations
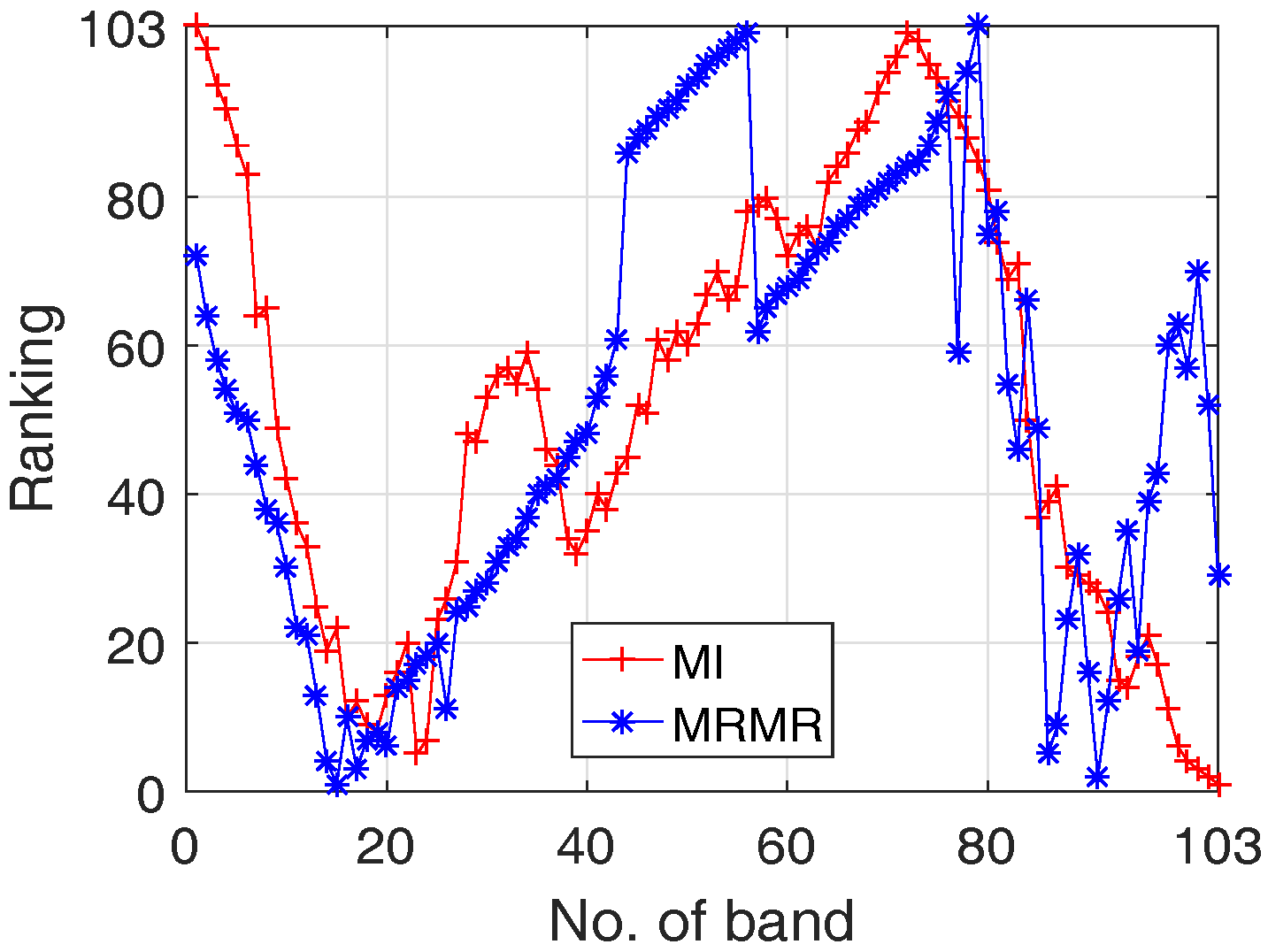
- In HSI data, there are a large number of bands (e.g., 100 bands or more), where different bands have various differentiating ability in classification (see, Figure 4);
- When the dimension of features is high, generally large volume of training data are required, which may not be realistic in some applications such as crop health monitoring, since in such fields, it is hard, expensive and time-consuming for experts to collect comparatively sufficient labelled data, particularly for early disease detection;
- With an increased number of features, the model parameter estimation of classification algorithms becomes very difficult as discussed in Section 1.
- Training and classification time for algorithms with a large number of features is generally high.
3. Feature Selection and Extraction
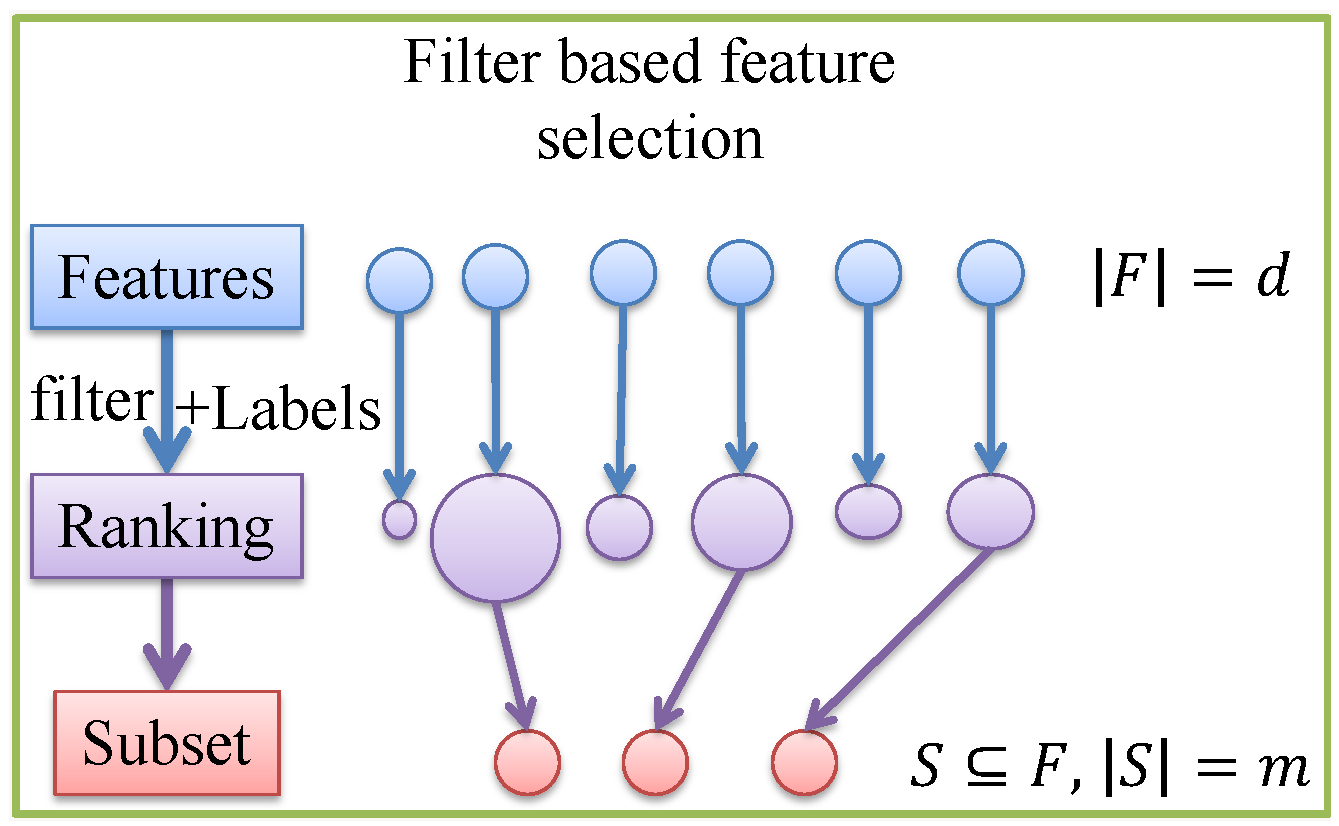
3.1. Feature Selections
3.1.1. MI Approach
3.1.2. MRMR Approach
3.2. Feature Extractions
3.2.1. Principal Components Analysis
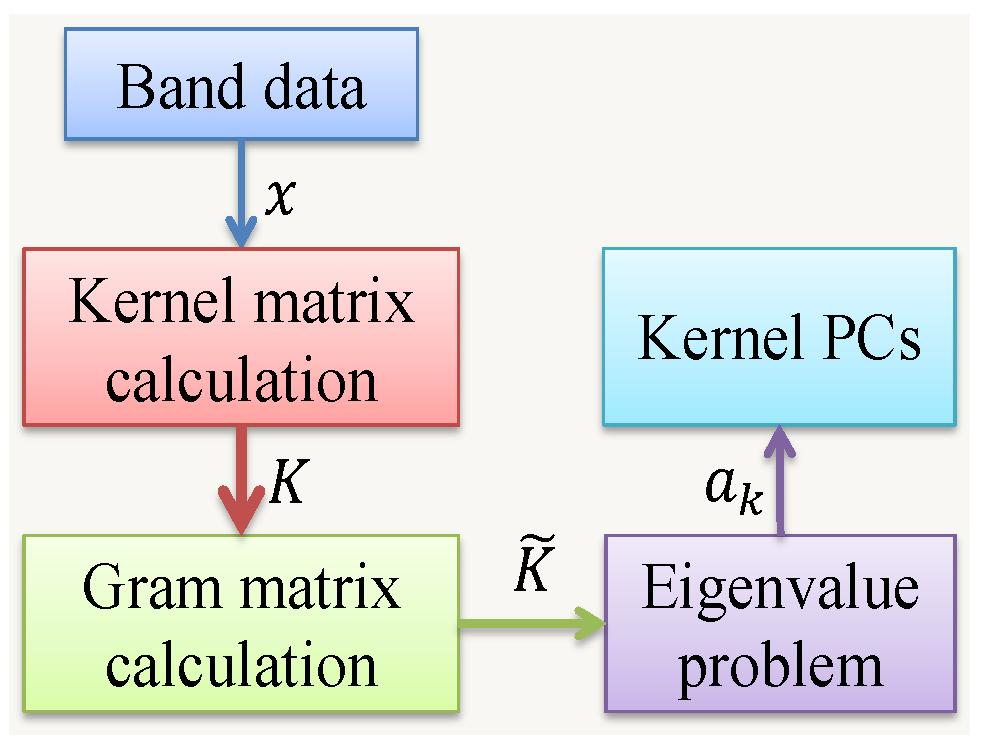
3.2.2. Kernel PCA
| Algorithm 1 Steps for Kernel PCA |
|
4. Experimental Results and Discussions
4.1. Feature Selection vs. Feature Extraction
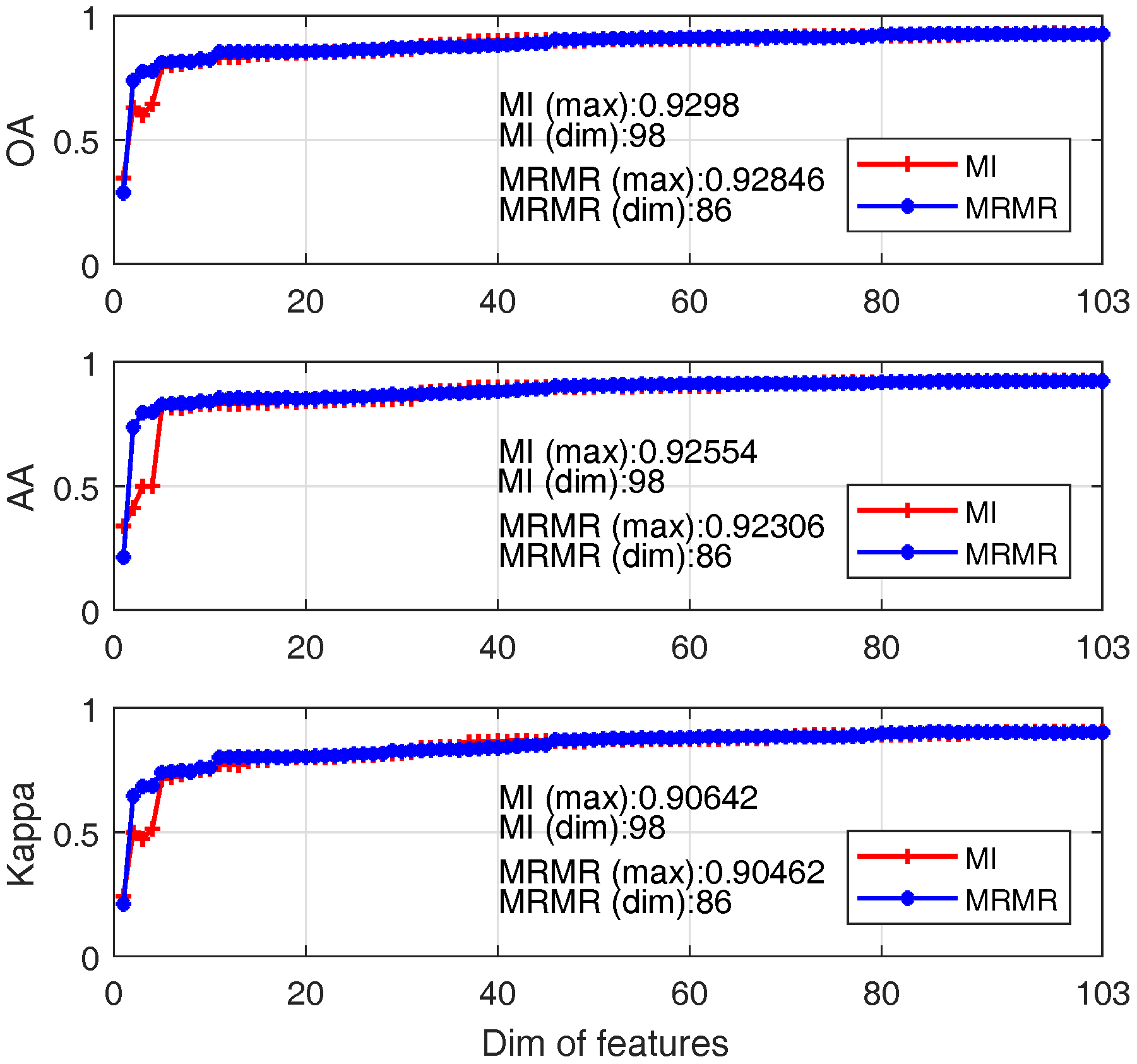
4.1.1. Feature Selection
- (1)
- For both approaches, the classification accuracy reaches 80% when about first 10 significant bands are adopted;
- (2)
- When band number is less than 10; MRMR outperforms MI for most cases; this is mainly due to the better features produced by MRMR by considering both feature relevance and redundancy;
- (3)
- The best OA of MI and MRMR based SVM are 93% and 92.8% with band numbers 98 and 86.
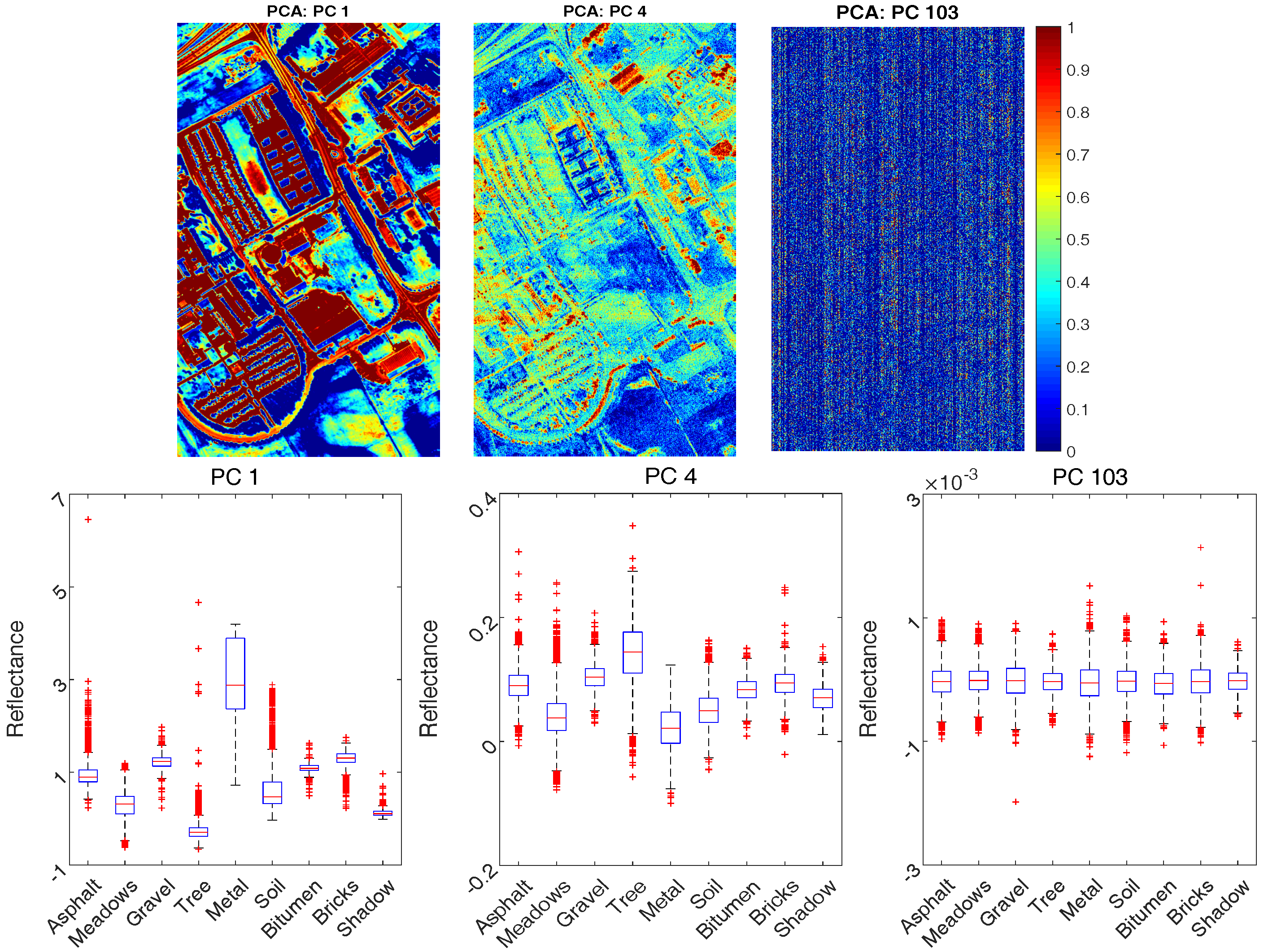
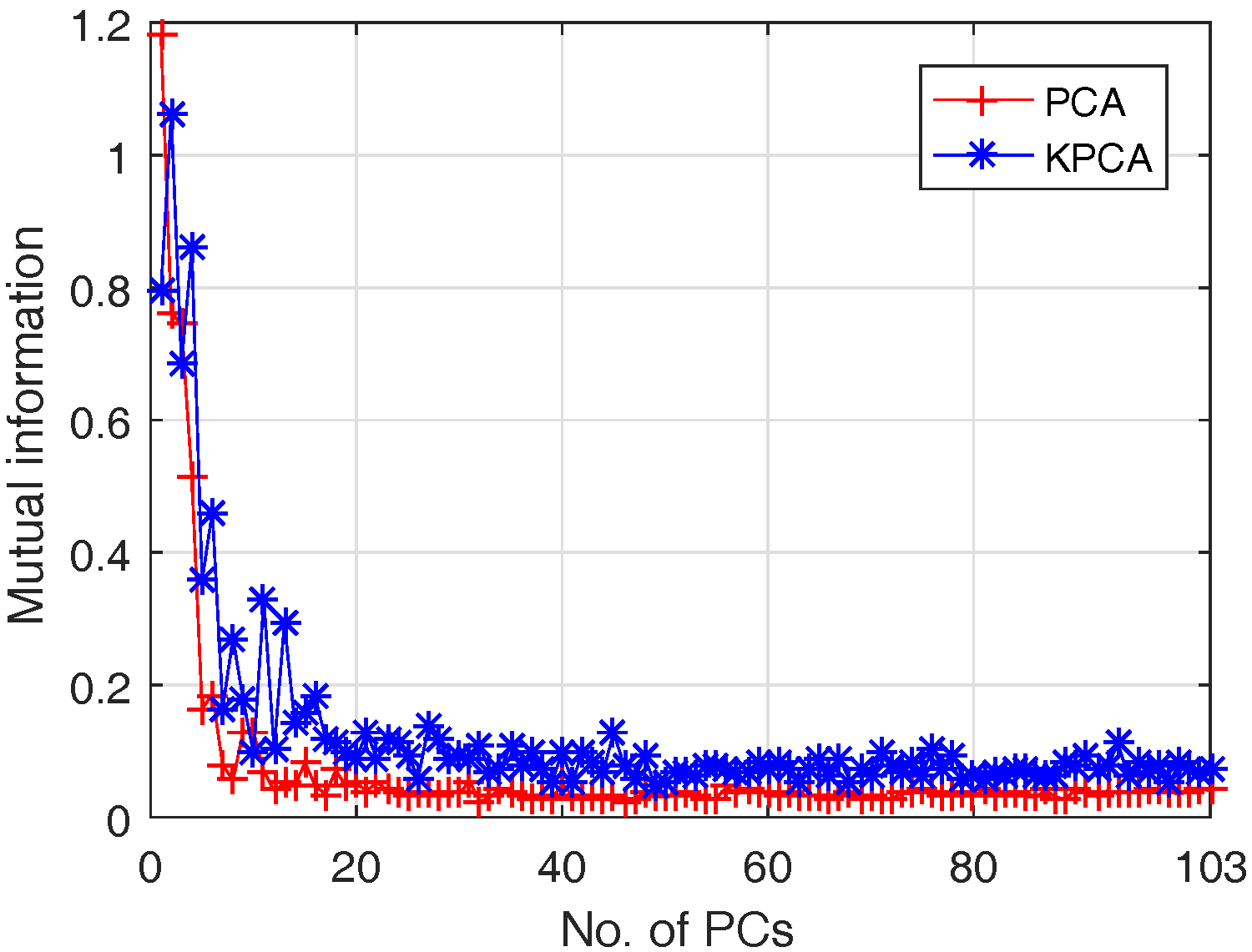
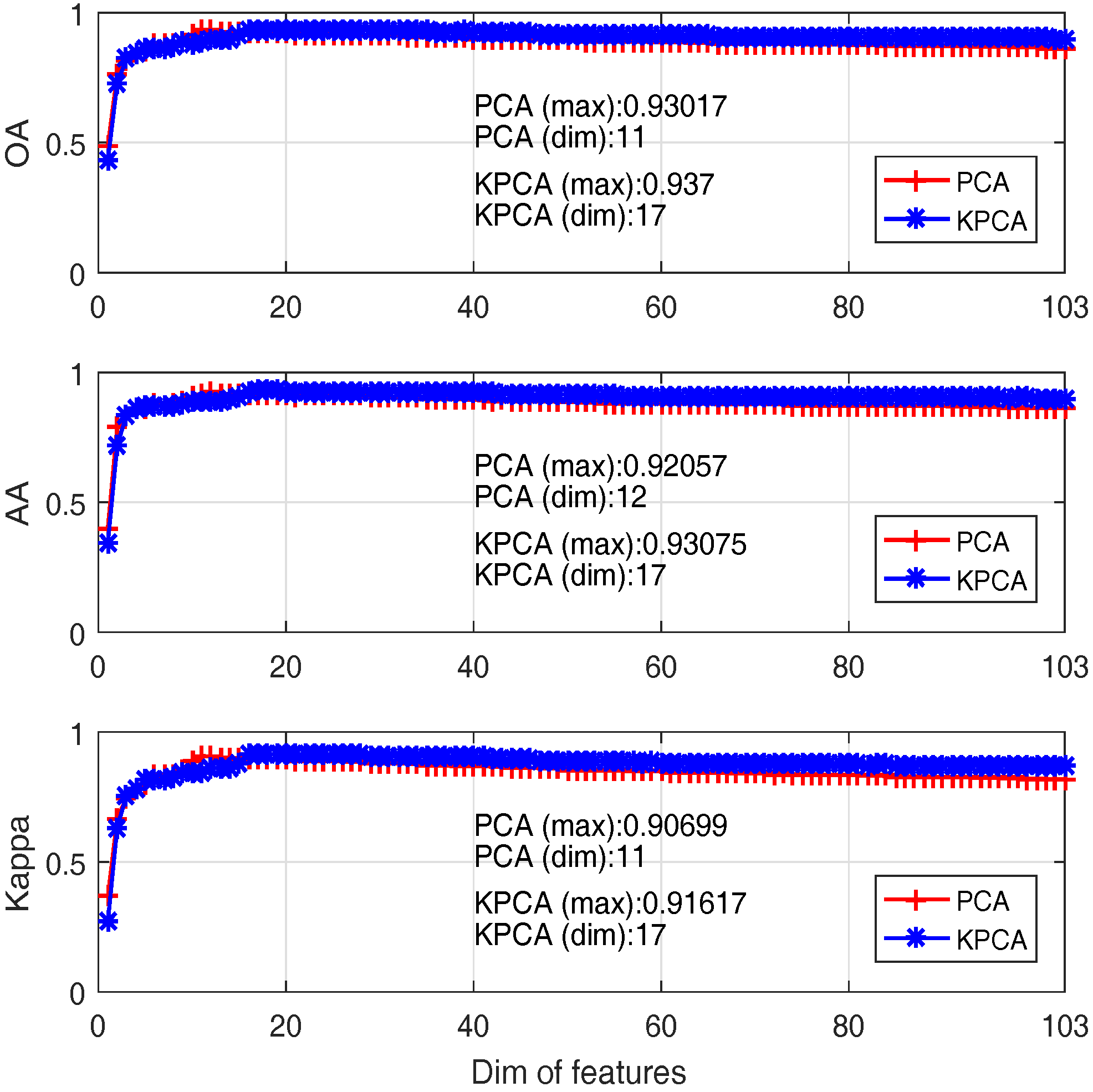
4.1.2. Feature Extraction
- (i)
- Classification accuracy reaches 80% when only the first 3 PCs are adopted, although the first 3 PCs account for 99% of the cumulative variance; this is because data variance is not exactly a good reflection of the data structure when it comes to classification;
- (ii)
- The best OA of PCA and Kernel PCA are 93% and 93.7%, when the number of bands are 11 and 17 respectively. That means Kernel PCA obtains similar (or slightly better) performance than PCA, but require a larger number of PCs.
4.1.3. Feature Selection vs. Feature Extraction
- (1)
- Only selected bands are needed in MI and MRMR approaches, however, all original bands are needed in PCA and Kernel PCA so that new features can be extracted;
- (2)
- The best performance of all four algorithms are quite close (no statistical difference);
- (3)
- To achieve the best performance, generally a larger number of features are required in MI and MRMR based feature selection approaches in comparison with PCA and Kernel PCA based feature extraction approaches;
- (4)
- Most importantly, linear PCA based approach is recommended since it is relatively simple, with low computational cost and a smaller number of features required to achieve its best performance.
4.2. PCA vs All Bands
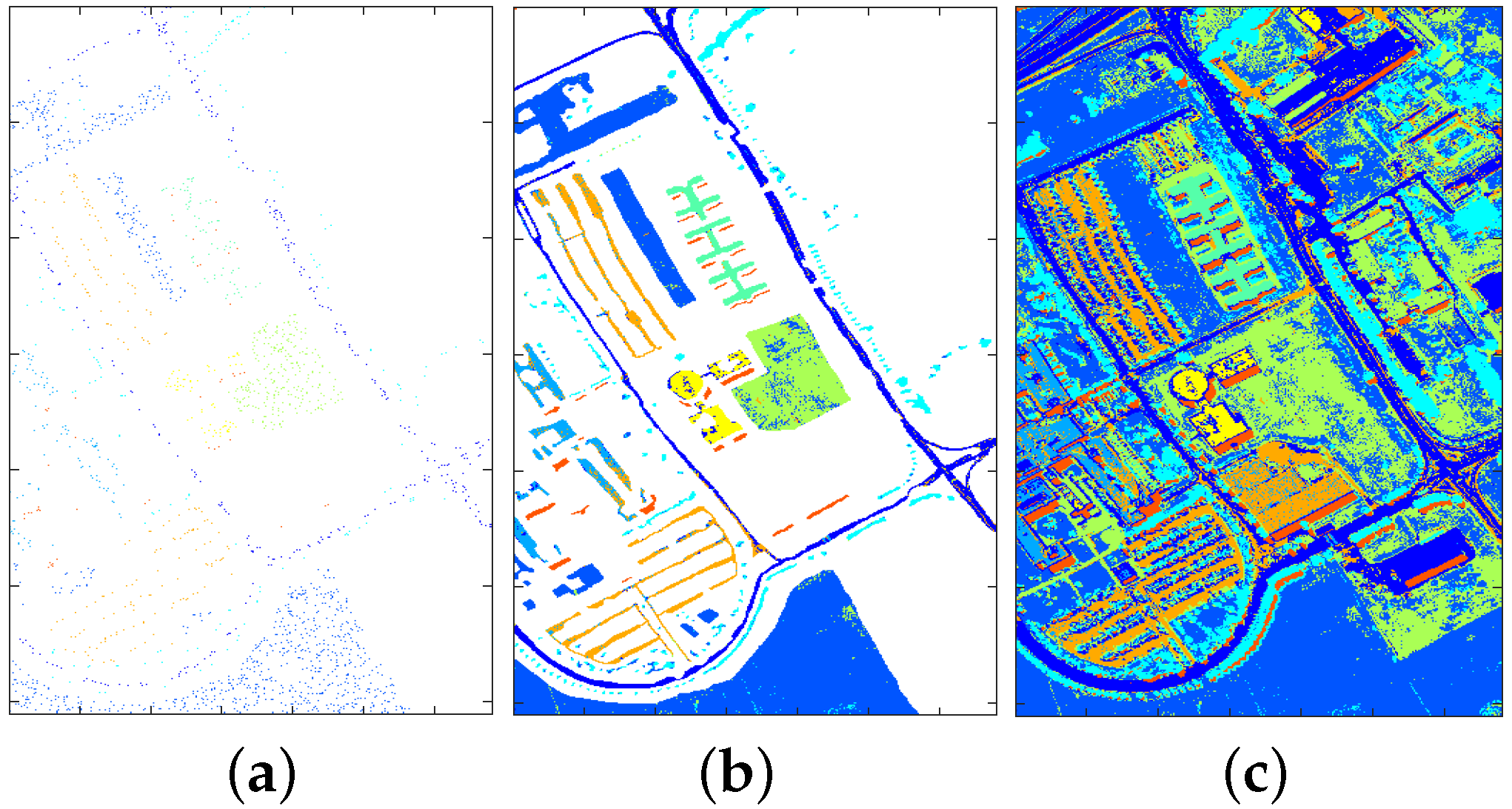
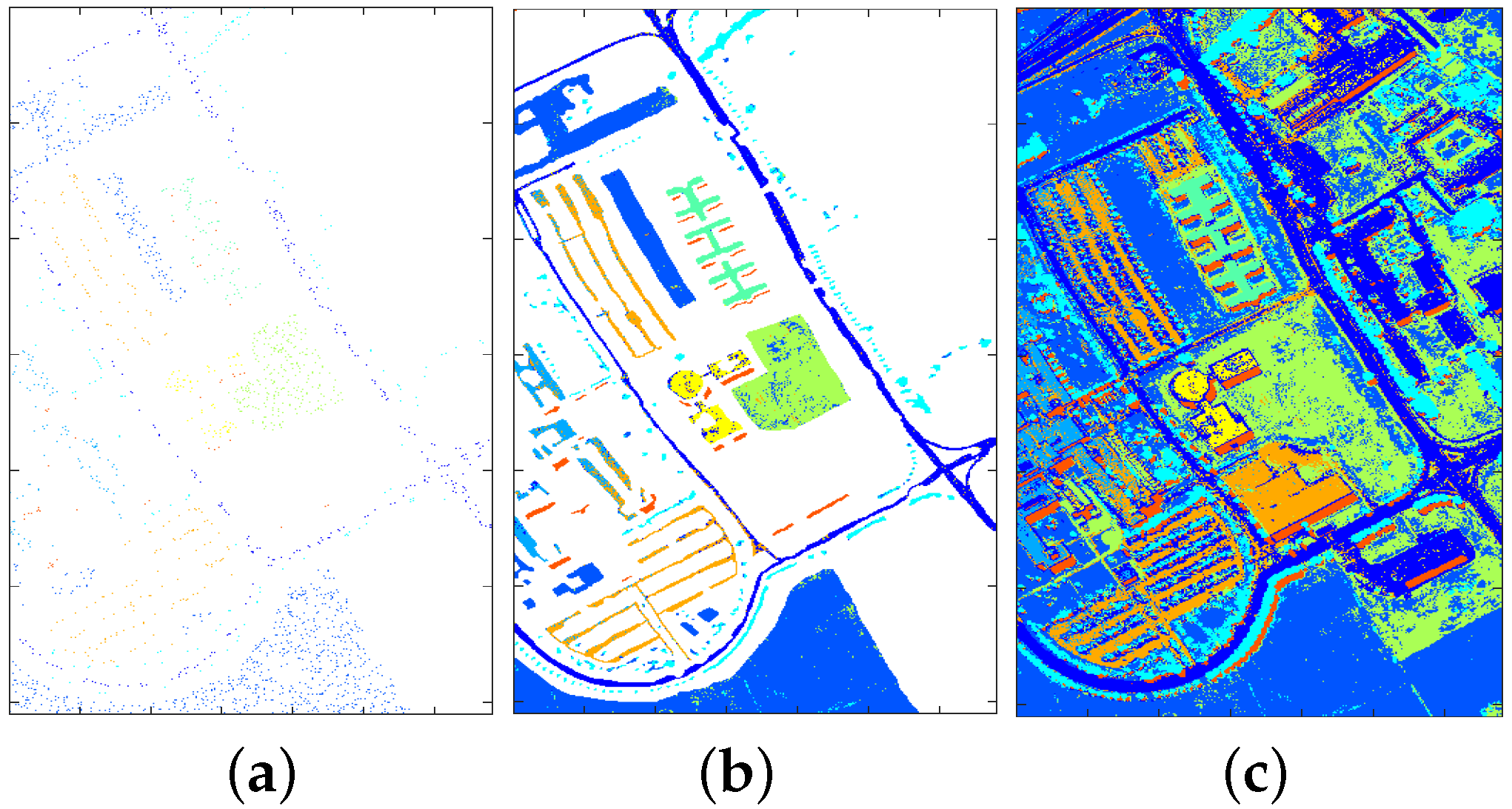
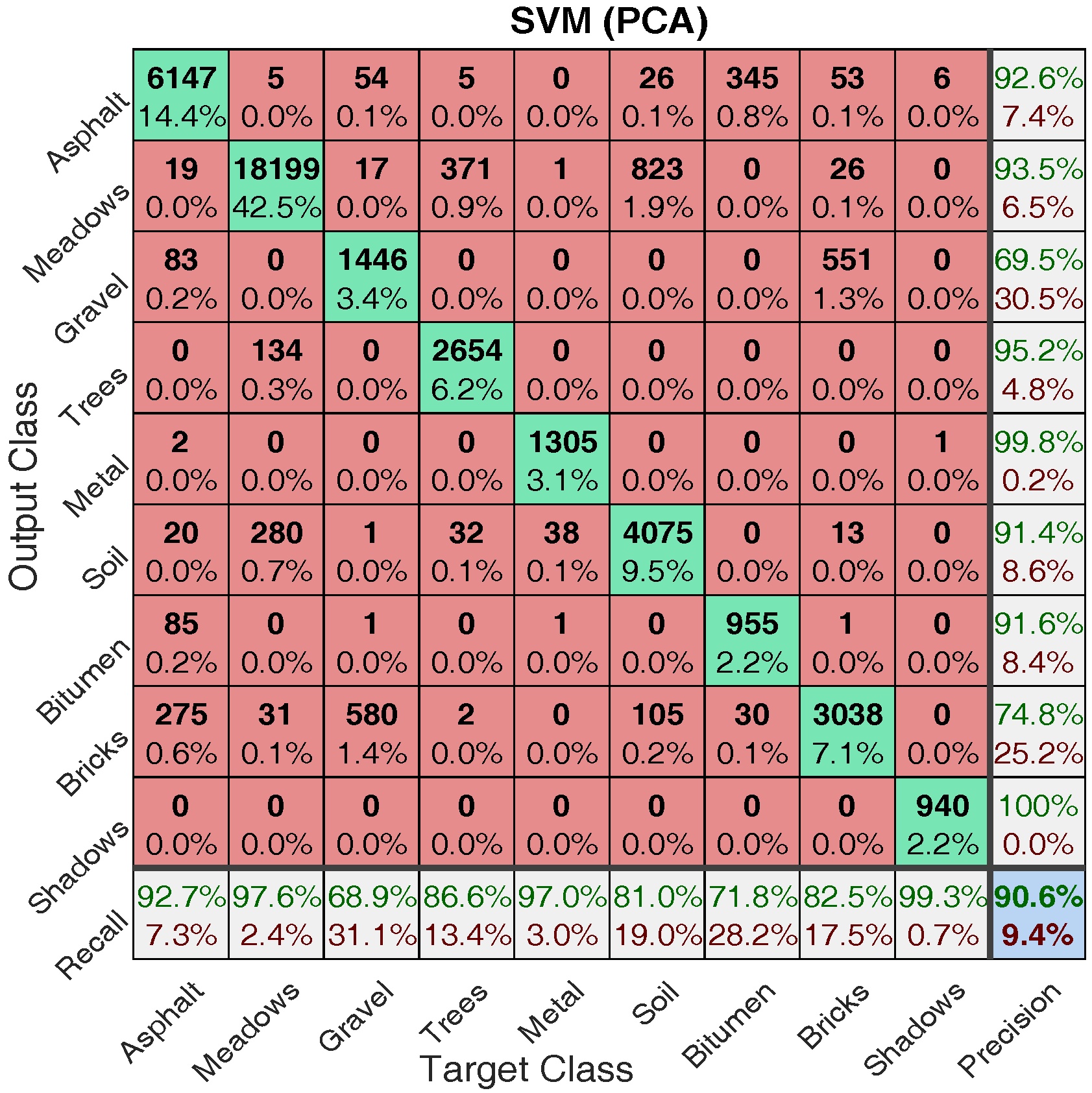
4.2.1. Classification Illustration
4.2.2. The Effect of Training Size
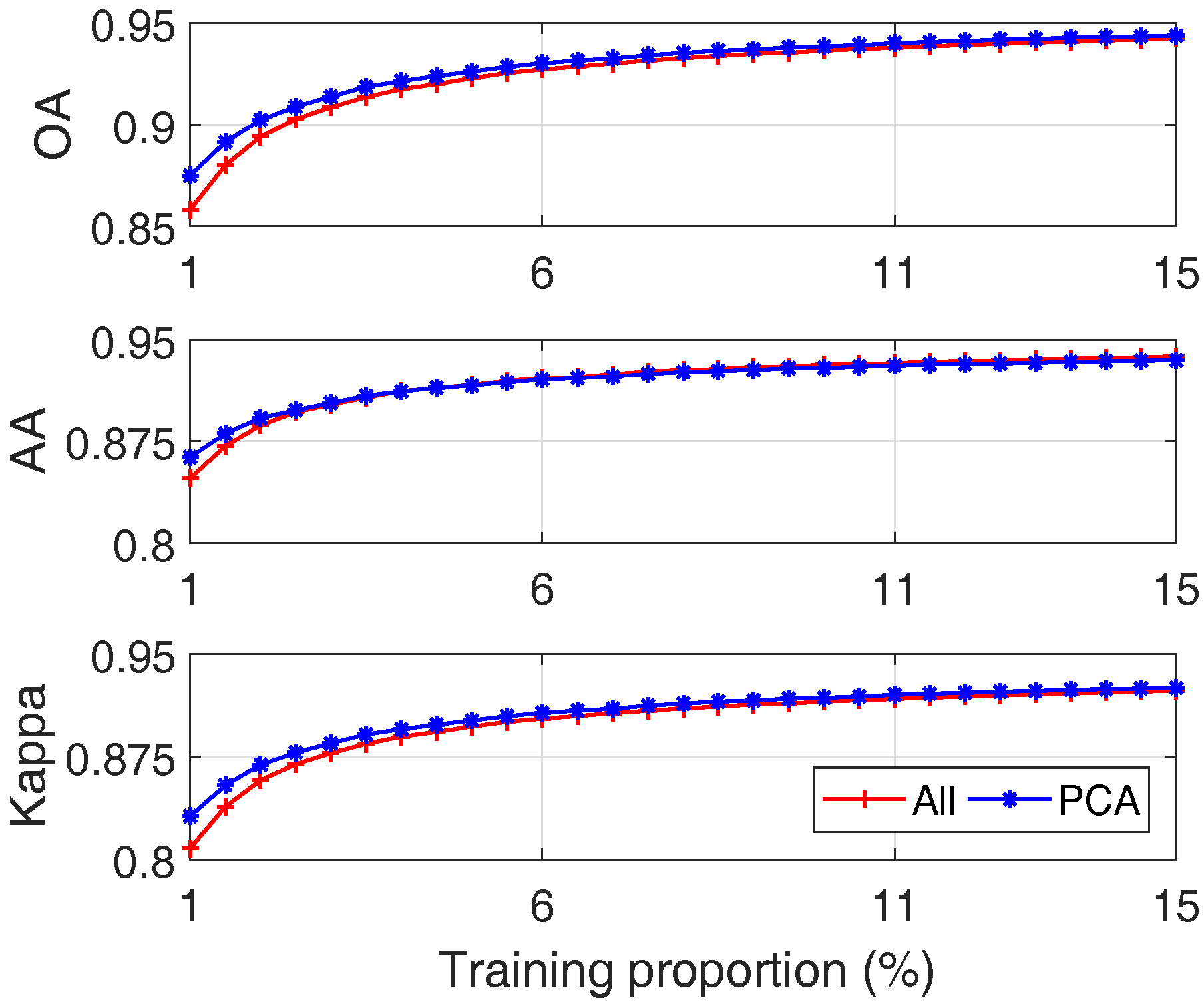
- (1)
- With an increased number of training samples, both approaches can get better results, which is consistent with the common sense in machine learning;
- (2)
- When the size of training data is small, both approaches degrade; this is due to the sparseness of training data. However, SVM with PCA outperforms SVM with all bands as shown by the left part of the plots of Figure 18. This is because by reducing the number of features using PCA, the effect of data sparseness can be attenuated to some extent.
- (3)
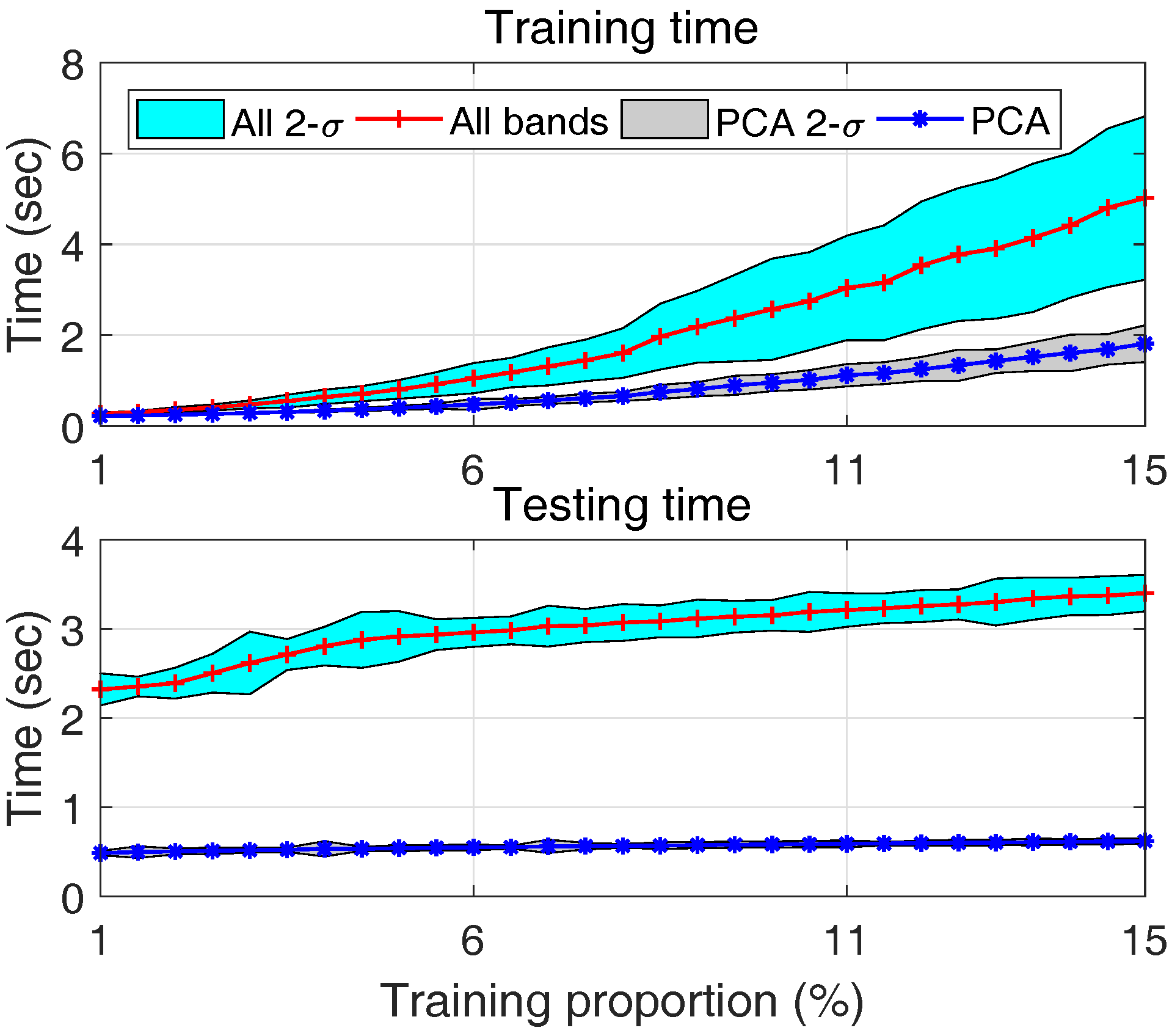
- With an increased size of training dataset, the training time for both approaches increases as shown by the upper plot of Figure 19. However, PCA based approach always requires less time, about 1/3 of all bands based approach. This is mainly due to the substantially reduced number of features in PCA based approach.
- (4)
- This is even more significant in real-time classification, it can be seen from the lower plot of Figure 19 that the computation time of all bands approach is about 6 times that of PCA approach, and this ratio keeps increasing with more training samples. This is because, on the one hand given a number of training data the SVM model using all bands is more complex due to the larger number of features; on the other hand, given a number of testing samples, the SVM model trained using a larger number of training samples is usually more complex due to an increased number of support vectors. The testing time of PCA based approach, due to the reduced number of features, is less sensitive to the size of training dataset.
- (5)
- Considering the classification performance (similar or even better) and computation load for both training (1/3) and testing (1/6), one can conclude that compared with SVM by using all bands, SVM with selected features using PCA is more appropriate for real-time applications, particularly when the training data are very limited.
5. Conclusions and Future Work
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Chen, Y.; Jiang, H.; Li, C.; Jia, X.; Ghamisi, P. Deep feature extraction and classification of hyperspectral images based on convolutional neural networks. IEEE Trans. Geosci. Remote Sens. 2016, 54, 6232–6251. [Google Scholar] [CrossRef]
- Jin, X.; Kumar, L.; Li, Z.; Xu, X.; Yang, G.; Wang, J. Estimation of Winter Wheat Biomass and Yield by Combining the AquaCrop Model and Field Hyperspectral Data. Remote Sens. 2016, 8, 972. [Google Scholar] [CrossRef]
- Li, L.; Zhang, Q.; Huang, D. A review of imaging techniques for plant phenotyping. Sensors 2014, 14, 20078–20111. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Yang, G.; Li, C.; Wang, Y.; Liu, J.; Yu, H.; Feng, H.; Xu, B.; Zhao, X.; Yang, X. Retrieving Soybean Leaf Area Index from Unmanned Aerial Vehicle Hyperspectral Remote Sensing: Analysis of RF, ANN, and SVM Regression Models. Remote Sens. 2017, 9, 309. [Google Scholar] [CrossRef]
- Fu, Y.; Zhao, C.; Wang, J.; Jia, X.; Yang, G.; Song, X.; Feng, H. An Improved Combination of Spectral and Spatial Features for Vegetation Classification in Hyperspectral Images. Remote Sens. 2017, 9, 261. [Google Scholar] [CrossRef]
- Nevalainen, O.; Honkavaara, E.; Tuominen, S.; Viljanen, N.; Hakala, T.; Yu, X.; Hyyppä, J.; Saari, H.; Pölönen, I.; Imai, N.N.; et al. Individual tree detection and classification with UAV-based photogrammetric point clouds and hyperspectral imaging. Remote Sens. 2017, 9, 185. [Google Scholar] [CrossRef]
- Ghamisi, P.; Plaza, J.; Chen, Y.; Li, J.; Plaza, A. Advanced Supervised Spectral Classifiers for Hyperspectral Images: A Review. IEEE Geosci. Remote Sens. Mag. 2017, 8–32. [Google Scholar] [CrossRef]
- Fauvel, M.; Tarabalka, Y.; Benediktsson, J.A.; Chanussot, J.; Tilton, J.C. Advances in spectral-spatial classification of hyperspectral images. Proc. IEEE 2013, 101, 652–675. [Google Scholar] [CrossRef]
- Hughes, G. On the mean accuracy of statistical pattern recognizers. IEEE Trans. Inf. Theory 1968, 14, 55–63. [Google Scholar] [CrossRef]
- Chi, M.; Feng, R.; Bruzzone, L. Classification of hyperspectral remote-sensing data with primal SVM for small-sized training dataset problem. Adv. Space Res. 2008, 41, 1793–1799. [Google Scholar] [CrossRef]
- Mountrakis, G.; Im, J.; Ogole, C. Support vector machines in remote sensing: A review. ISPRS J. Photogramm. Remote Sens. 2011, 66, 247–259. [Google Scholar] [CrossRef]
- Wang, J.; Chang, C.I. Independent component analysis-based dimensionality reduction with applications in hyperspectral image analysis. IEEE Trans. Geosci. Remote Sens. 2006, 44, 1586–1600. [Google Scholar] [CrossRef]
- Jia, X.; Kuo, B.C.; Crawford, M.M. Feature mining for hyperspectral image classification. Proc. IEEE 2013, 101, 676–697. [Google Scholar] [CrossRef]
- Kumar, S.; Ghosh, J.; Crawford, M.M. Best-bases feature extraction algorithms for classification of hyperspectral data. IEEE Trans. Geosci. Remote Sens. 2001, 39, 1368–1379. [Google Scholar] [CrossRef]
- Guyon, I.; Elisseeff, A. An introduction to variable and feature selection. J. Mach. Learn. Res. 2003, 3, 1157–1182. [Google Scholar]
- Yuan, Y.; Zhu, G.; Wang, Q. Hyperspectral band selection by multitask sparsity pursuit. IEEE Trans. Geosci. Remote Sens. 2015, 53, 631–644. [Google Scholar] [CrossRef]
- Chandrashekar, G.; Sahin, F. A survey on feature selection methods. Comput. Electr. Eng. 2014, 40, 16–28. [Google Scholar] [CrossRef]
- Andrew, A.M.; Zakaria, A.; Mad Saad, S.; Md Shakaff, A.Y. Multi-stage feature selection based intelligent classifier for classification of incipient stage fire in building. Sensors 2016, 16, 31. [Google Scholar] [CrossRef] [PubMed]
- Mather, P.M.; Koch, M. Computer Processing of Remotely-Sensed Images: An Introduction; John Wiley & Sons: Hoboken, NJ, USA, 2011. [Google Scholar]
- Li, S.; Qiu, J.; Yang, X.; Liu, H.; Wan, D.; Zhu, Y. A novel approach to hyperspectral band selection based on spectral shape similarity analysis and fast branch and bound search. Eng. Appl. Artif. Intell. 2014, 27, 241–250. [Google Scholar] [CrossRef]
- Guyon, I.; Weston, J.; Barnhill, S.; Vapnik, V. Gene selection for cancer classification using support vector machines. Mach. Learn. 2002, 46, 389–422. [Google Scholar] [CrossRef]
- Pal, M.; Foody, G.M. Feature selection for classification of hyperspectral data by SVM. IEEE Trans. Geosci. Remote Sens. 2010, 48, 2297–2307. [Google Scholar] [CrossRef]
- Lu, S.; Oki, K.; Shimizu, Y.; Omasa, K. Comparison between several feature extraction/classification methods for mapping complicated agricultural land use patches using airborne hyperspectral data. Int. J. Remote Sens. 2007, 28, 963–984. [Google Scholar] [CrossRef]
- Plaza, A.; Benediktsson, J.A.; Boardman, J.W.; Brazile, J.; Bruzzone, L.; Camps-Valls, G.; Chanussot, J.; Fauvel, M.; Gamba, P.; Gualtieri, A.; et al. Recent advances in techniques for hyperspectral image processing. Remote Sens. Environ. 2009, 113, S110–S122. [Google Scholar] [CrossRef]
- Kang, X.; Li, S.; Benediktsson, J.A. Spectral—Spatial hyperspectral image classification with edge-preserving filtering. IEEE Trans. Geosci. Remote Sens. 2014, 52, 2666–2677. [Google Scholar] [CrossRef]
- Boser, B.E.; Guyon, I.M.; Vapnik, V.N. A training algorithm for optimal margin classifiers. In Proceedings of the Fifth Annual Workshop on Computational Learning Theory, Pittsburgh, PA, USA, 27–29 July 1992; pp. 144–152. [Google Scholar]
- Vapnik, V. Statistical Learning Theory; Wiley: New York, NY, USA, 1998; Volume 1. [Google Scholar]
- Gevaert, C.M.; Persello, C.; Vosselman, G. Optimizing Multiple Kernel Learning for the Classification of UAV Data. Remote Sens. 2016, 8, 1025. [Google Scholar] [CrossRef]
- Rifkin, R. MIT—Multiclass Classification. Available online: http://www.mit.edu/~9.520/spring09/Classes/multiclass.pdf (accessed on 25 June 2017).
- Kohavi, R.; John, G.H. Wrappers for feature subset selection. Artif. Intell. 1997, 97, 273–324. [Google Scholar] [CrossRef]
- Koller, D.; Sahami, M. Toward Optimal Feature Selection; Technical Report; Stanford InfoLab: Southampton, UK, 1996. [Google Scholar]
- Marill, T.; Green, D. On the effectiveness of receptors in recognition systems. IEEE Trans. Inf. Theory 1963, 9, 11–17. [Google Scholar] [CrossRef]
- Roffo, G. Feature Selection Library (MATLAB Toolbox). arXiv, 2016; arXiv:1607.01327. [Google Scholar]
- Gu, Q.; Li, Z.; Han, J. Generalized fisher score for feature selection. arXiv, 2012; arXiv:1202.3725. [Google Scholar]
- Cover, T.M.; Thomas, J.A. Elements of Information Theory; John Wiley & Sons: Hoboken, NJ, USA, 2012. [Google Scholar]
- Pohjalainen, J.; Räsänen, O.; Kadioglu, S. Feature selection methods and their combinations in high-dimensional classification of speaker likability, intelligibility and personality traits. Comput. Speech Lang. 2015, 29, 145–171. [Google Scholar] [CrossRef]
- Peng, H.; Long, F.; Ding, C. Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans. Pattern Anal. Mach. Intell. 2005, 27, 1226–1238. [Google Scholar] [CrossRef] [PubMed]
- Schölkopf, B.; Smola, A.; Müller, K.R. Kernel principal component analysis. In Proceedings of the International Conference on Artificial Neural Networks, Lausanne, Switzerland, 8–10 October 1997; Springer: New York, NY, USA, 1997; pp. 583–588. [Google Scholar]
- Bishop, C.M. Pattern recognition. Mach. Learn. 2006, 128, 1–58. [Google Scholar]
- Wang, Q. Kernel principal component analysis and its applications in face recognition and active shape models. arXiv, 2012; arXiv:1207.3538. [Google Scholar]
- Rodarmel, C.; Shan, J. Principal component analysis for hyperspectral image classification. Surv. Land Inf. Sci. 2002, 62, 115–122. [Google Scholar]




| # | MI | MRMR | PCA | KPCA |
|---|---|---|---|---|
| Type | FS | FS | FE | FE |
| All bands | No | No | Yes | Yes |
| Best OA | 92.98% | 92.85% | 93.02% | 93.70% |
| Best AA | 92.55% | 92.31% | 92.06% | 93.08% |
| Best Kappa | 0.906 | 0.905 | 0.907 | 0.910 |
| No. features | 98 | 86 | 11 | 17 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, J.; Yi, D.; Liu, C.; Guo, L.; Chen, W.-H. Dimension Reduction Aided Hyperspectral Image Classification with a Small-sized Training Dataset: Experimental Comparisons. Sensors 2017, 17, 2726. https://doi.org/10.3390/s17122726
Su J, Yi D, Liu C, Guo L, Chen W-H. Dimension Reduction Aided Hyperspectral Image Classification with a Small-sized Training Dataset: Experimental Comparisons. Sensors. 2017; 17(12):2726. https://doi.org/10.3390/s17122726
Chicago/Turabian StyleSu, Jinya, Dewei Yi, Cunjia Liu, Lei Guo, and Wen-Hua Chen. 2017. "Dimension Reduction Aided Hyperspectral Image Classification with a Small-sized Training Dataset: Experimental Comparisons" Sensors 17, no. 12: 2726. https://doi.org/10.3390/s17122726
APA StyleSu, J., Yi, D., Liu, C., Guo, L., & Chen, W.-H. (2017). Dimension Reduction Aided Hyperspectral Image Classification with a Small-sized Training Dataset: Experimental Comparisons. Sensors, 17(12), 2726. https://doi.org/10.3390/s17122726






