Valid Probabilistic Predictions for Ginseng with Venn Machines Using Electronic Nose
Abstract
:1. Introduction
2. Probabilistic Prediction Methods
2.1. Venn Machine
2.2. Naive Bayes Classification
2.3. Softmax Regression
2.4. Platt’s Method
2.5. The Validity of Probabilistic Predictions
2.5.1. Loss Function
2.5.2. Cumulative Probability Values versus Cumulative Correct Predictions
3. Materials and Methods
3.1. Sample Preparation
3.2. E-Nose Equipment and Measurement
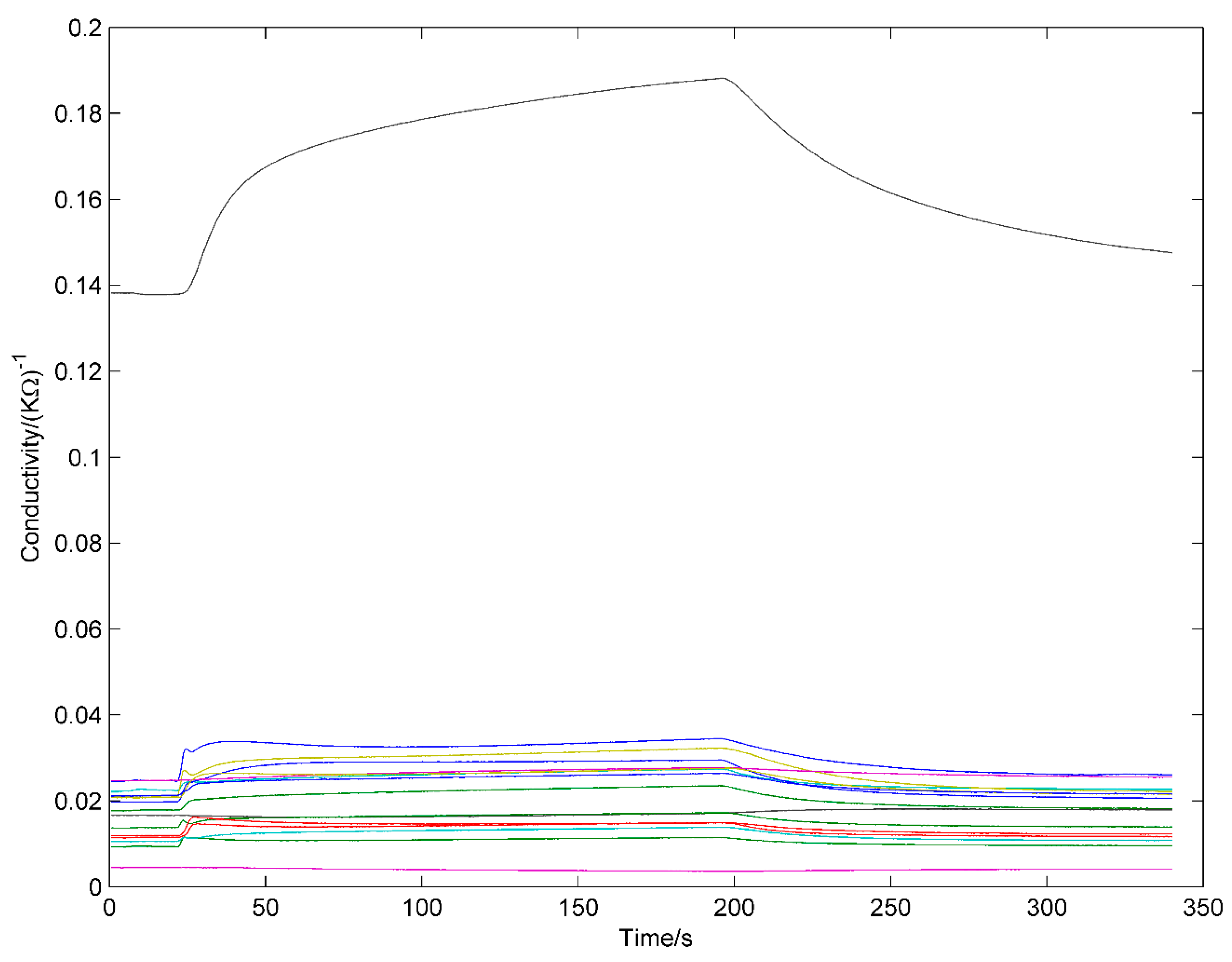
3.3. Data Processing
- 1.
- The maximal absolute response, , which is most efficient and widely-used steady feature.
- 2.
- The area under the full response curve, , where T (=340 s) is the total measurement time, which is also widely- used steady feature.
- 3–8.
- Exponential moving average of derivative [18,19] of V, , where the discretely sampled exponential moving average with smoothing factors . SR is the sampling rate, SR = 10 Hz. . For each smoothing factor, two features were extracted. A total of six transient feature were extracted. Besides steady features, transient features were considered to contain much effective information that should be made the best of [20].Finally, 16 × 8 = 128 features are extracted from each sample and all the features are scaled to [0 1]:where is the jth feature from ith sensor. is the feature after normalization.
4. Results and Discussion
4.1. Performance of Probabilistic Predictors in Offline Mode
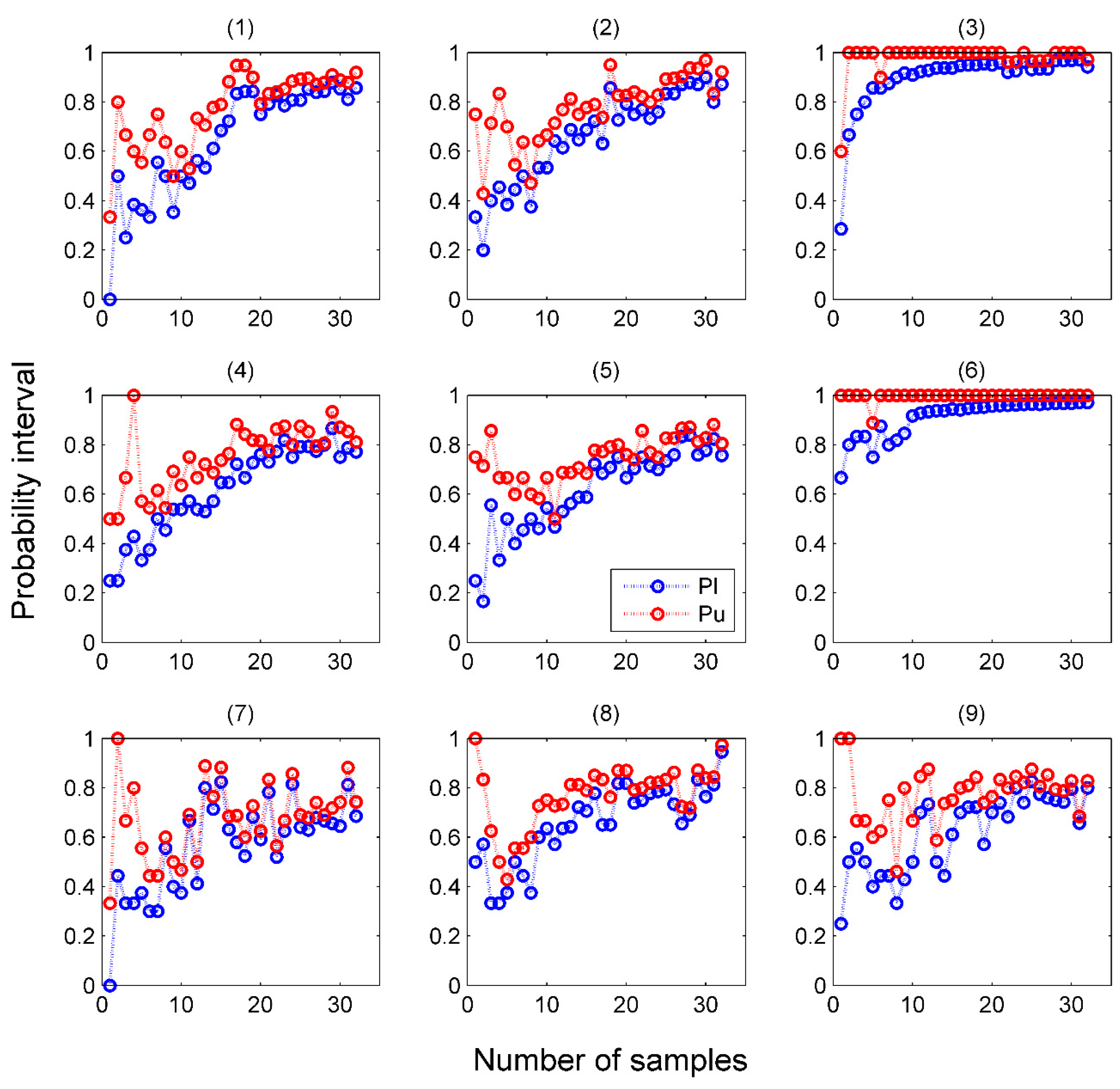
4.2. Performance of Venn Predictors in Online Mode
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- De Vito, S.; Piga, M.; Martinotto, L.; Di Francia, G. Co, NO 2 and NO x urban pollution monitoring with on-field calibrated electronic nose by automatic bayesian regularization. Sens. Actuators B Chem. 2009, 143, 182–191. [Google Scholar] [CrossRef]
- Musatov, V.Y.; Sysoev, V.V.; Sommer, M.; Kiselev, I. Assessment of meat freshness with metal oxide sensor microarray electronic nose: A practical approach. Sens. Actuators B Chem. 2010, 144, 99–103. [Google Scholar] [CrossRef]
- Baldwin, E.A.; Bai, J.H.; Plotto, A.; Dea, S. Electronic noses and tongues: Applications for the food and pharmaceutical industries. Sensors 2011, 11, 4744–4766. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Huang, C.Q.; Xing, J.G.; Zheng, J.B. Pattern classification using an olfactory model with pca feature selection in electronic noses: Study and application. Sensors 2012, 12, 2818–2830. [Google Scholar] [CrossRef] [PubMed]
- Schmekel, B.; Winquist, F.; Vikstrom, A. Analysis of breath samples for lung cancer survival. Anal. Chim. Acta 2014, 840, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Montuschi, P.; Mores, N.; Trove, A.; Mondino, C.; Barnes, P.J. The electronic nose in respiratory medicine. Respiration 2013, 85, 72–84. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.; Castro, M.; Feller, J.F. An e-nose made of carbon nanotube based quantum resistive sensors for the detection of eighteen polar/nonpolar voc biomarkers of lung cancer. J. Mater. Chem. B 2013, 1, 4563–4575. [Google Scholar] [CrossRef]
- Moon, J.; Han, H.; Park, S.; Dong, H.; Han, K.-Y.; Kim, H.; Bang, K.-H.; Choi, J.; Noh, B. Discrimination of the origin of commercial red ginseng concentrates using LC-MS/MS and electronic nose analysis based on a mass spectrometer. Food Sci. Biotechnol. 2014, 23, 1433–1440. [Google Scholar] [CrossRef]
- Lee, D.Y.; Cho, J.G.; Lee, M.K.; Lee, J.W.; Lee, Y.H.; Yang, D.C.; Baek, N.I. Discrimination of panax ginseng roots cultivated in different areas in Korea using hplc-elsd and principal component analysis. J. Ginseng. Res. 2011, 35, 31–38. [Google Scholar] [CrossRef]
- Lin, H.-T.; Lin, C.-J.; Weng, R. A note on platt’s probabilistic outputs for support vector machines. Mach. Learn. 2007, 68, 267–276. [Google Scholar] [CrossRef]
- Williams, C.K.I.; Barber, D. Bayesian classification with gaussian processes. IEEE Trans. Pattern Anal. 1998, 20, 1342–1351. [Google Scholar] [CrossRef]
- Nouretdinov, I.; Devetyarov, D.; Vovk, V.; Burford, B.; Camuzeaux, S.; Gentry-Maharaj, A.; Tiss, A.; Smith, C.; Luo, Z.Y.; Chervonenkis, A.; et al. Multiprobabilistic prediction in early medical diagnoses. Ann. Math. Artif. Int. 2015, 74, 203–222. [Google Scholar] [CrossRef]
- Vork, V.; Gammerman, A.; Shafer, G. Algorithmic Learning in a Random World; Springer: New York, NY, USA, 2005; p. 16. [Google Scholar]
- Papadopoulos, H. Reliable probabilistic classification with neural networks. Neurocomputing 2013, 107, 59–68. [Google Scholar] [CrossRef]
- Platt, J.C. Probabilistic outputs for support vector machines and comparison to regularized likelihood methods. In Advances in Large Margin Classiers; MIT Press: Cambridge, MA, USA, 2000. [Google Scholar]
- Vovk, V.; Petej, I. Venn-abers predictors. In Proceedings of the 30th Conference on Uncertainty in Artificial Intelligence, Quebec City, QC, Canada, 23–27 July 2014.
- Miao, J.C.; Zhang, T.L.; Wang, Y.; Li, G. Optimal sensor selection for classifying a set of ginsengs using metal-oxide sensors. Sensors 2015, 15, 16027–16039. [Google Scholar] [CrossRef] [PubMed]
- Muezzinoglu, M.K.; Vergara, A.; Huerta, R.; Rulkov, N.; Rabinovich, M.I.; Selverston, A.; Abarbanel, H.D.I. Acceleration of chemo-sensory information processing using transient features. Sens. Actuators B Chem. 2009, 137, 507–512. [Google Scholar] [CrossRef]
- Rodriguez-Lujan, I.; Fonollosa, J.; Vergara, A.; Homer, M.; Huerta, R. On the calibration of sensor arrays for pattern recognition using the minimal number of experiments. Chemometr. Intell. Lab. 2014, 130, 123–134. [Google Scholar] [CrossRef]
- Fonollosa, J.; Vergara, A.; Huerta, R. Algorithmic mitigation of sensor failure: Is sensor replacement really necessary? Sens. Actuators B Chem. 2013, 183, 211–221. [Google Scholar] [CrossRef]
- Chang, C.C.; Lin, C.J. Libsvm: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2011, 2, 1–27. [Google Scholar] [CrossRef]


| No. | Ginseng Species | Places of Production |
|---|---|---|
| 1 | Chinese red ginseng | Ji’an |
| 2 | Chinese red ginseng | Fusong |
| 3 | Korean red ginseng | Ji’an |
| 4 | Chinese white ginseng | Ji’an |
| 5 | Chinese white ginseng | Fusong |
| 6 | American ginseng | Fusong |
| 7 | American ginseng | USA |
| 8 | American ginseng | Canada |
| 9 | American ginseng | Tonghua |
| Methods | Classification Rate | Assessment Criteria of Validity | ||
|---|---|---|---|---|
| dln | dsq | d1 | ||
| VM-SVM | 86.35% | 0.3862 | 0.3419 | 0.0373 |
| Platt’s method | 88.57% | 0.3876 | 0.3439 | 0.1480 |
| VM-SR | 77.78% | 0.4690 | 0.3938 | 0.1085 |
| SR | 76.19% | Inf a | 0.4376 | 0.1853 |
| VM-NB | 60.32% | 0.5683 | 0.4475 | 0.0266 |
| NB | 40.32% | 0.5851 | 0.4510 | 0.0332 |
| Method/Category | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | |
|---|---|---|---|---|---|---|---|---|---|---|
| VM-SVM | Sensitivity | 0.9714 | 0.8857 | 1 | 0.8571 | 0.8286 | 1 | 0.6857 | 0.8286 | 0.7143 |
| Specificity | 0.9857 | 0.9893 | 0.9964 | 0.9821 | 0.9786 | 1 | 0.9536 | 0.9857 | 0.9750 | |
| Platt’s method | Sensitivity | 0.9714 | 0.8857 | 1 | 0.8571 | 0.8857 | 1 | 0.7143 | 0.8571 | 0.7714 |
| Specificity | 0.9857 | 0.9893 | 0.9964 | 0.9893 | 0.9821 | 1 | 0.9607 | 0.9857 | 0.9786 | |
| VM-SR | Sensitivity | 0.8000 | 0.8286 | 0.9429 | 0.7429 | 0.5714 | 1 | 0.5143 | 0.8000 | 0.8000 |
| Specificity | 0.9714 | 0.9714 | 1 | 0.9536 | 0.9643 | 0.9964 | 0.9571 | 0.9750 | 0.9607 | |
| SR | Sensitivity | 0.7429 | 0.7429 | 0.9714 | 0.7143 | 0.6000 | 1 | 0.5429 | 0.8000 | 0.7429 |
| Specificity | 0.9714 | 0.9714 | 1 | 0.9536 | 0.9643 | 0.9964 | 0.9571 | 0.9750 | 0.9607 | |
| VM-NB | Sensitivity | 0.8000 | 0 | 0.9714 | 0.6571 | 0.4286 | 0.9429 | 0.4286 | 0.4857 | 0.7143 |
| Specificity | 0.8464 | 0.9786 | 0.9929 | 0.9500 | 0.9321 | 1 | 0.9429 | 0.9714 | 0.9393 | |
| NB | Sensitivity | 0 | 0.3143 | 0.9714 | 0.2857 | 0.286 | 0.6857 | 0.571 | 0.3714 | 0.1429 |
| Specificity | 1 | 0.9607 | 0.9929 | 0.8750 | 0.8786 | 0.9107 | 0.8929 | 0.8679 | 0.9500 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Miao, J.; Lyu, X.; Liu, L.; Luo, Z.; Li, G. Valid Probabilistic Predictions for Ginseng with Venn Machines Using Electronic Nose. Sensors 2016, 16, 1088. https://doi.org/10.3390/s16071088
Wang Y, Miao J, Lyu X, Liu L, Luo Z, Li G. Valid Probabilistic Predictions for Ginseng with Venn Machines Using Electronic Nose. Sensors. 2016; 16(7):1088. https://doi.org/10.3390/s16071088
Chicago/Turabian StyleWang, You, Jiacheng Miao, Xiaofeng Lyu, Linfeng Liu, Zhiyuan Luo, and Guang Li. 2016. "Valid Probabilistic Predictions for Ginseng with Venn Machines Using Electronic Nose" Sensors 16, no. 7: 1088. https://doi.org/10.3390/s16071088
APA StyleWang, Y., Miao, J., Lyu, X., Liu, L., Luo, Z., & Li, G. (2016). Valid Probabilistic Predictions for Ginseng with Venn Machines Using Electronic Nose. Sensors, 16(7), 1088. https://doi.org/10.3390/s16071088





