An Automated Field Phenotyping Pipeline for Application in Grapevine Research
Abstract
:1. Introduction
2. Material and Methods
2.1. Plant Material
2.2. Automated Image Acquisition
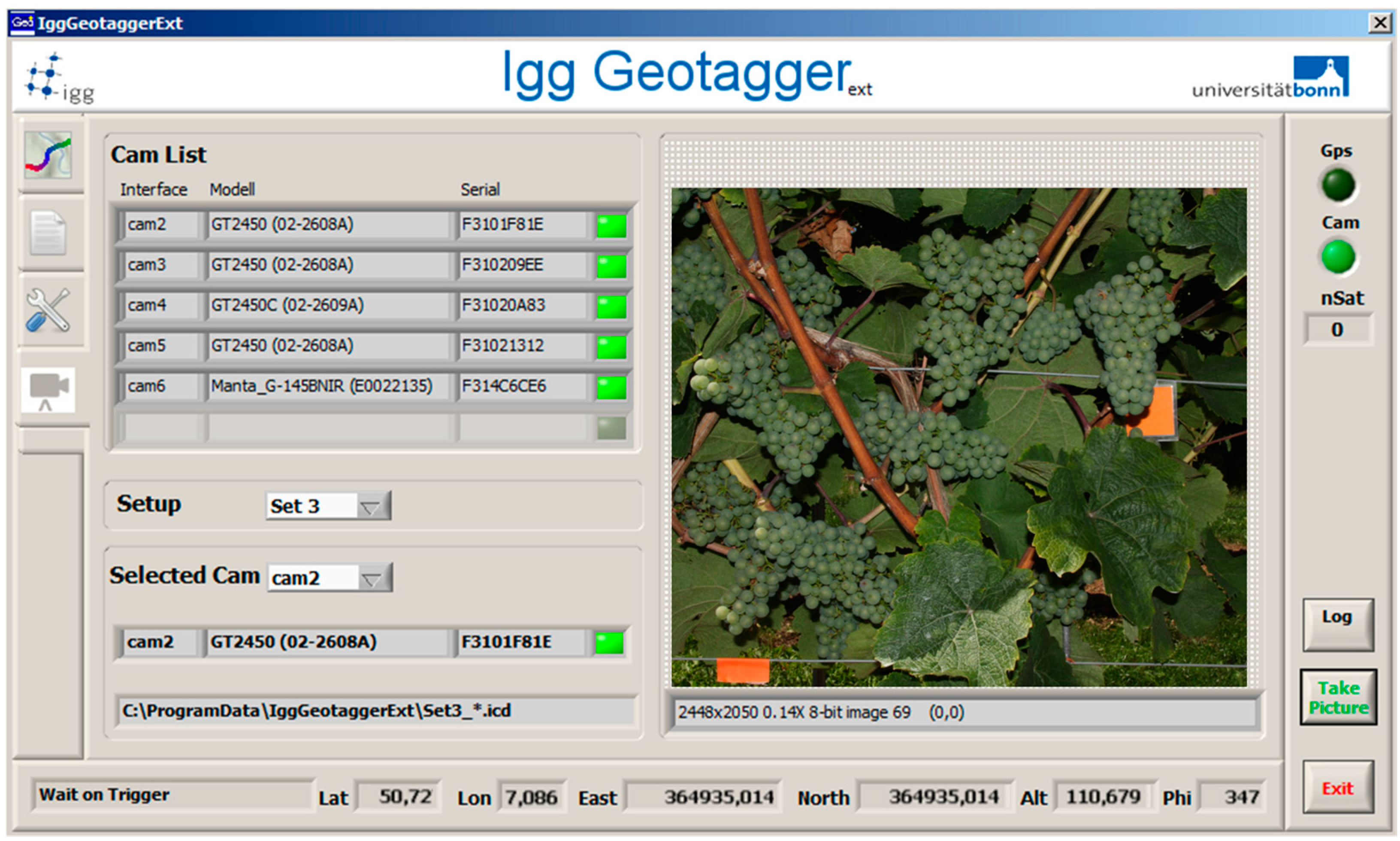
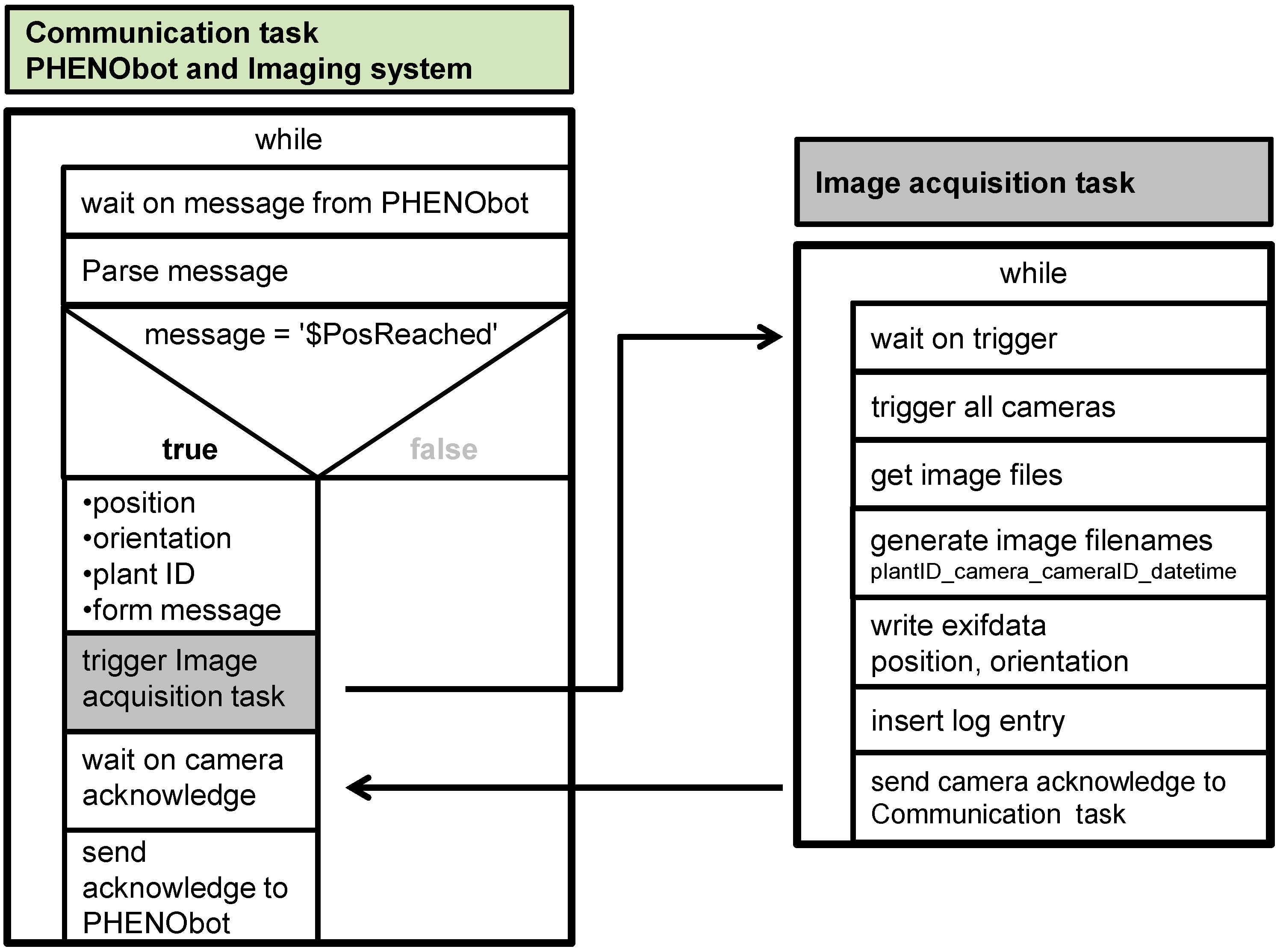
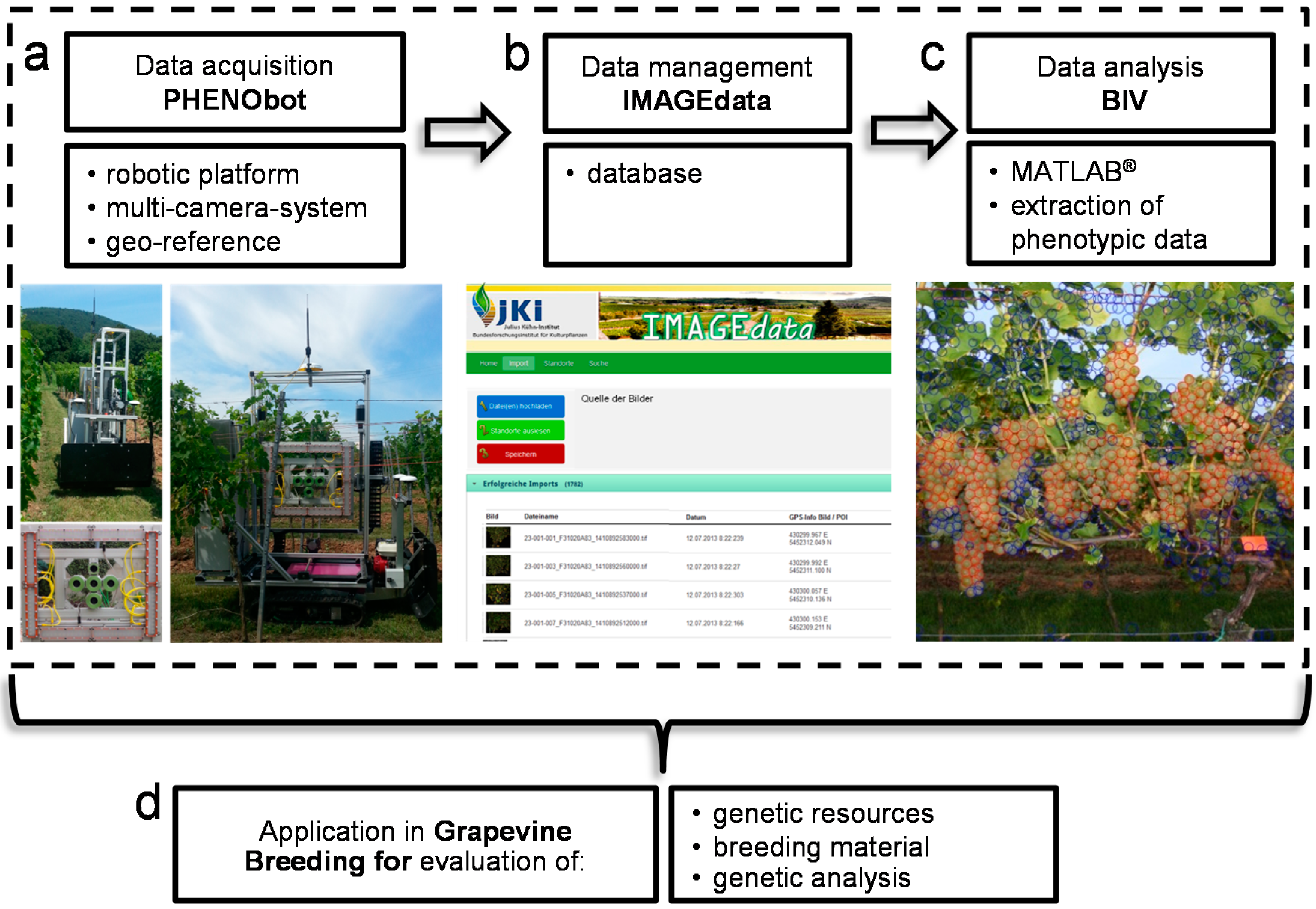
2.3. Data Management
2.4. Image Analysis
2.5. Statistical Analysis
3. Results and Discussion
3.1. Field Application of the Phenotyping Robot

3.2. Image Analysis
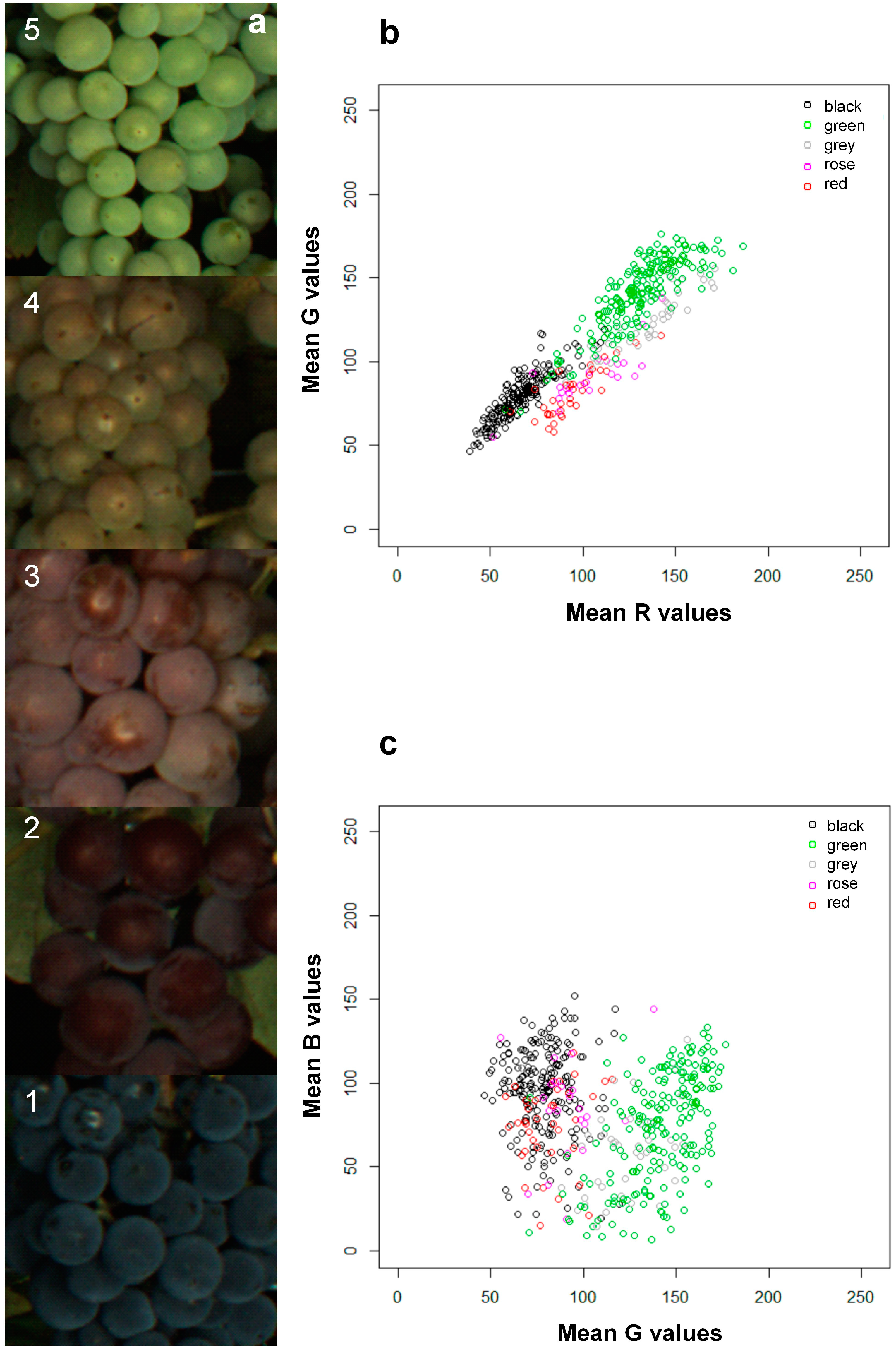
| Real Color Classes | |||||
|---|---|---|---|---|---|
| Predicted Color Classes | Black | Green | Grey | Red | Rose |
| black | 197 | 7 | 2 | 5 | 3 |
| green | 5 | 178 | 7 | 0 | 0 |
| grey | 0 | 15 | 28 | 2 | 3 |
| red | 0 | 0 | 1 | 26 | 13 |
| rose | 0 | 0 | 1 | 4 | 3 |
3.3. Future Work
4. Conclusions
Acknowledgments
Author Contributions
Appendix
Conflicts of Interest
References
- Thomas, M.R.; Scott, N.S. Microsatellite repeats in grapevine reveal DNA polymorphisms when analysed as sequence-tagged sites (STSS). Theoret. Appl. Genetics 1993, 86, 985–990. [Google Scholar] [CrossRef]
- Eibach, R.; Zyprian, E.; Welter, L.; Töpfer, R. The use of molecular markers for pyramiding resistance genes in grapevine breeding. Vitis 2007, 46, 120–124. [Google Scholar]
- Barba, P.; Cadle-Davidson, L.; Harriman, J.; Glaubitz, J.; Brooks, S.; Hyma, K.; Reisch, B. Grapevine powdery mildew resistance and susceptibility loci identified on a high-resolution SNP map. Theoret. Appl. Genet. 2014, 127, 73–84. [Google Scholar] [CrossRef]
- Granier, C.; Aguirrezabal, L.; Chenu, K.; Cookson, S.J.; Dauzat, M.; Hamard, P.; Thioux, J.J.; Rolland, G.; Bouchier-Combaud, S.; Lebaudy, A.; et al. Phenopsis, an automated platform for reproducible phenotyping of plant responses to soil water deficit in arabidopsis thaliana permitted the identification of an accession with low sensitivity to soil water deficit. New Phytol. 2006, 169, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Walter, A.; Scharr, H.; Gilmer, F.; Zierer, R.; Nagel, K.; Ernst, M.; Wiese, A.; Virnich, O.; Christ, M.; Uhlig, B.; et al. Dynamics of seedling growth acclimation towards altered light conditions can be quantified via growscreen: A setup and procedure designed for rapid optical phenotyping of different plant species. New Phytol. 2007, 174, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Reuzeau, C.; Pen, J.; Frankard, V.; de Wolf, J.; Peerbolte, R.; Broekaert, W.; van Camp, W. Traitmilltm: A discovery engine for identifying yield-enhancement genes in cereals. Mol. Plant Breed. 2005, 5, 753–759. [Google Scholar]
- Clark, R.T.; MacCurdy, R.B.; Jung, J.K.; Shaff, J.E.; McCouch, S.R.; Aneshansley, D.J.; Kochian, L.V. Three-dimensional root phenotyping with a novel imaging and software platform. Plant Physiol. 2011, 156, 455–465. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.; Czauderna, T.; Hoffmann, R.; Stein, N.; Schreiber, F. Htpheno: An image analysis pipeline for high-throughput plant phenotyping. BMC Bioinf. 2011, 12, 148–156. [Google Scholar] [CrossRef]
- Bellin, D.; Peressotti, E.; Merdinoglu, D.; Wiedemann-Merdinoglu, S.; Adam-Blondon, A.F.; Cipriani, G.; Morgante, M.; Testolin, R.; di Gaspero, G. Resistance to plasmopara viticola in grapevine “bianca” is controlled by a major dominant gene causing localised necrosis at the infection site. Theor. Appl. Genet. 2009, 120, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Mazzetto, F.; Calcante, A.; Mena, A.; Vercesi, A. Integration of optical and analogue sensors for monitoring canopy health and vigour in precision viticulture. Precis. Agric. 2010, 11, 636–649. [Google Scholar] [CrossRef]
- Llorens, J.; Gil, E.; Llop, J.; Queralto, M. Georeferenced lidar 3D vine plantation map generation. Sensors 2011, 11, 6237–6256. [Google Scholar] [CrossRef] [PubMed]
- Braun, T.; Koch, H.; Strub, O.; Zolynski, G.; Berns, K. Improving pesticide spray application in vineyards by automated analysis of the foliage distribution pattern in the leaf wall. In Commercial Vehicle Technology 2010, Proceedings of the 1st Commercial Vehicle Technology Symposium (CVT 2010), Kaiserslautern, Germany, 16–18 March 2010; Berns, K., Schindler, C., Dreßler, K., Jörg, B., Kalmar, R., Hirth, J., Eds.; pp. 539–548.
- Berenstein, R.; Shahar, O.B.; Shapiro, A.; Edan, Y. Grape clusters and foliage detection algorithms for autonomous selective vineyard sprayer. Intell. Serv. Robot. 2010, 3, 233–243. [Google Scholar] [CrossRef]
- Corbett-Davies; Sam Botterill; Tom Green, R.; Saxton, V. An expert system for automatically pruning vines. In Proceedings of the 27th Conference on Image and Vision Computing New Zealand; Dunedin, New Zealand: 26–28 November 2012; pp. 55–60.
- Nuske, S.; Achar, S.; Bates, T.; Narasimhan, S.; Singh, S. Yield estimation in vineyards by visual grape detection. In Proceedings of the International Conference on Itelligent Robots and Systems, San Francisco, CA, USA, 25–30 September 2011; IEEE: San Francisco, CA, USA, 2011. [Google Scholar]
- Diago, M.P.; Correa, C.; Millán, B.; Barreiro, P.; Valero, C.; Tardaguila, J. Grapevine yield and leaf area estimation using supervised classification methodology on rgb images taken under field conditions. Sensors 2012, 12, 16988–17006. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Marden, S.; Whitty, M. Towards automated yield estimation in viticulture. In Proceedings of the Australasian Conference on Robotics and Automation, Sydney, Australia, 2–4 December 2013; UNSW: Sydney, Australia, 2013. [Google Scholar]
- Nuske, S.; Wilshusen, K.; Achar, S.; Yoder, L.; Narasimhan, S.; Singh, S. Automated visual yield estimation in vineyards. J. Field Robot. 2014, 31, 837–860. [Google Scholar] [CrossRef]
- Diago, M.P.; Sanz-Garcia, A.; Millan, B.; Blasco, J.; Tardaguila, J. Assessment of flower number per inflorescence in grapevine by image analysis under field conditions. J. Sci. Food Agric. 2014, 94, 1981–1987. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, S.; Bei, R.; Pech, J.; Tyerman, S. Computational water stress indices obtained from thermal image analysis of grapevine canopies. Irrig. Sci. 2012, 30, 523–536. [Google Scholar] [CrossRef]
- Fuentes, S.; Poblete-Echeverría, C.; Ortega-Farias, S.; Tyerman, S.; de Bei, R. Automated estimation of leaf area index from grapevine canopies using cover photography, video and computational analysis methods. Aust. J. Grape Wine Res. 2014, 20, 465–473. [Google Scholar] [CrossRef]
- Möller, M.; Alchanatis, V.; Cohen, Y.; Meron, M.; Tsipris, J.; Naor, A.; Ostrovsky, V.; Sprintsin, M.; Cohen, S. Use of thermal and visible imagery for estimating crop water status of irrigated grapevine. J. Exp. Bot. 2007, 58, 827–838. [Google Scholar] [CrossRef] [PubMed]
- Calcante, A.; Mena, A.; Mazzetto, F. Evaluation of “ground sensing” optical sensors for diagnosis of plasmopara viticola on vines. Span. J. Agric. Res. 2012, 10, 619–630. [Google Scholar] [CrossRef]
- Mazzetto, F.; Calcante, A.; Mena, A.; Sacco, P. Test of ground-sensing devices for monitoring canopy vigour and downy mildew presence in vineyards: First qualitative traits. J. Agric. Eng. Riv. Ing. Agrar. 2011, 42, 1–9. [Google Scholar]
- Grocholsky, B.; Nuske, S.; Aasted, M.; Achar, S.; Bates, T. A camera and laser system for automatic vine balance assessment. In Proceedings of the ASABE Annual International Meeting, Louisville, Kentucky, 7–10 August 2011; ASABE: Louisville, Kentucky, 2011. [Google Scholar]
- Ruckelshausen, A.; Biber, P.; Dorna, M.; Gremmes, H.; Klose, R.; Linz, A.; Rahe, F.; Resch, R.; Thiel, M.; Trautz, D.; et al. Bonirob—An autonomous field robot platform for individual plant phenotyping. In Proceedings of the 7th European Conference on Precision Agriculture, Wageningen, The Netherlands, 6–8 July 2009; Academic Publishers: Wageningen, The Netherlands, 2009; pp. 841–847. [Google Scholar]
- Busemeyer, L.; Mentrup, D.; Moller, K.; Wunder, E.; Alheit, K.; Hahn, V.; Maurer, H.P.; Reif, J.C.; Wurschum, T.; Muller, J.; et al. Breedvision—A multi-sensor platform for non-destructive field-based phenotyping in plant breeding. Sensors 2013, 13, 2830–2847. [Google Scholar] [CrossRef] [PubMed]
- Longo, D.; Pennissi, A.; Bonsignore, R.; Muscato, G.; Schillaci, G. A multifunctional tracked vehicle able to operate in vineyards using gps and laser range-finder technology. In Proceedings of the International Conference Ragusa SHWA2010, Ragusa Ibla Campus, Italy, 16–18 September 2010; pp. 487–492.
- Longo, D.; Muscato, G.; Caruso, L.; Conti, A.; Schillaci, G. Design of a remotely operable sprayer for precision farming application. In Proceedings of the International Conference RAGUSA SHWA, Ragusa, Italy, 3–6 September 2012.
- Lorenz, D.H.; Eichhorn, K.W.; Bleiholder, H.; Klose, R.; Meier, U.; Weber, E. Growth stages of the grapevine: Phenological growth stages of the grapevine (Vitis vinifera L. ssp. Vinifera)—Codes and descriptions according to the extended BBCH scale. Aust. J. Grape Wine Res. 1995, 1, 100–103. [Google Scholar] [CrossRef]
- Anonymous. 2nd Edition of the Oiv Descriptor List for Grape Varieties and Vitis Species. Available online: http://www.oiv.int/oiv/info/enplubicationoiv#grape (accessed on 6 January 2015).
- Herzog, K.; Roscher, R.; Wieland, M.; Kicherer, A.; Läbe, T.; Förstner, W.; Kuhlmann, H.; Töpfer, R. Initial steps for high-throughput phenotyping in vineyards. Vitis 2014, 53, 1–8. [Google Scholar]
- Schwarz, H.P.; Rüger, P.; Kicherer, A.; Töpfer, R. Development of an autonomous driven robotic platform used for ht-phenotyping in viticulture. Mech. Eng. Lett. Szent István Univ. 2013, 10, 153–160. [Google Scholar]
- Roscher, R.; Herzog, K.; Kunkel, A.; Kicherer, A.; Töpfer, R.; Förstner, W. Automated image analysis framework for high-throughput determination of grapevine berry sizes using conditional random fields. Comput. Electron. Agric. 2014, 100, 148–158. [Google Scholar] [CrossRef]
- Doligez, A.; Bouquet, A.; Danglot, Y.; Lahogue, F.; Riaz, S.; Meredith, C.P.; Edwards, K.J.; This, P. Genetic mapping of grapevine (Vitis vinifera L.) applied to the detection of QTLs for seedlessness and berry weight. Theor. Appl. Genet. 2002, 105, 780–795. [Google Scholar] [CrossRef] [PubMed]
- Doligez, A.; Audiot, E.; Baumes, R.; This, P. QTLs for muscat flavor and monoterpenic odorant content in grapevine (Vitis vinifera L.). Mol. Breed. 2006, 18, 109–125. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kicherer, A.; Herzog, K.; Pflanz, M.; Wieland, M.; Rüger, P.; Kecke, S.; Kuhlmann, H.; Töpfer, R. An Automated Field Phenotyping Pipeline for Application in Grapevine Research. Sensors 2015, 15, 4823-4836. https://doi.org/10.3390/s150304823
Kicherer A, Herzog K, Pflanz M, Wieland M, Rüger P, Kecke S, Kuhlmann H, Töpfer R. An Automated Field Phenotyping Pipeline for Application in Grapevine Research. Sensors. 2015; 15(3):4823-4836. https://doi.org/10.3390/s150304823
Chicago/Turabian StyleKicherer, Anna, Katja Herzog, Michael Pflanz, Markus Wieland, Philipp Rüger, Steffen Kecke, Heiner Kuhlmann, and Reinhard Töpfer. 2015. "An Automated Field Phenotyping Pipeline for Application in Grapevine Research" Sensors 15, no. 3: 4823-4836. https://doi.org/10.3390/s150304823
APA StyleKicherer, A., Herzog, K., Pflanz, M., Wieland, M., Rüger, P., Kecke, S., Kuhlmann, H., & Töpfer, R. (2015). An Automated Field Phenotyping Pipeline for Application in Grapevine Research. Sensors, 15(3), 4823-4836. https://doi.org/10.3390/s150304823





