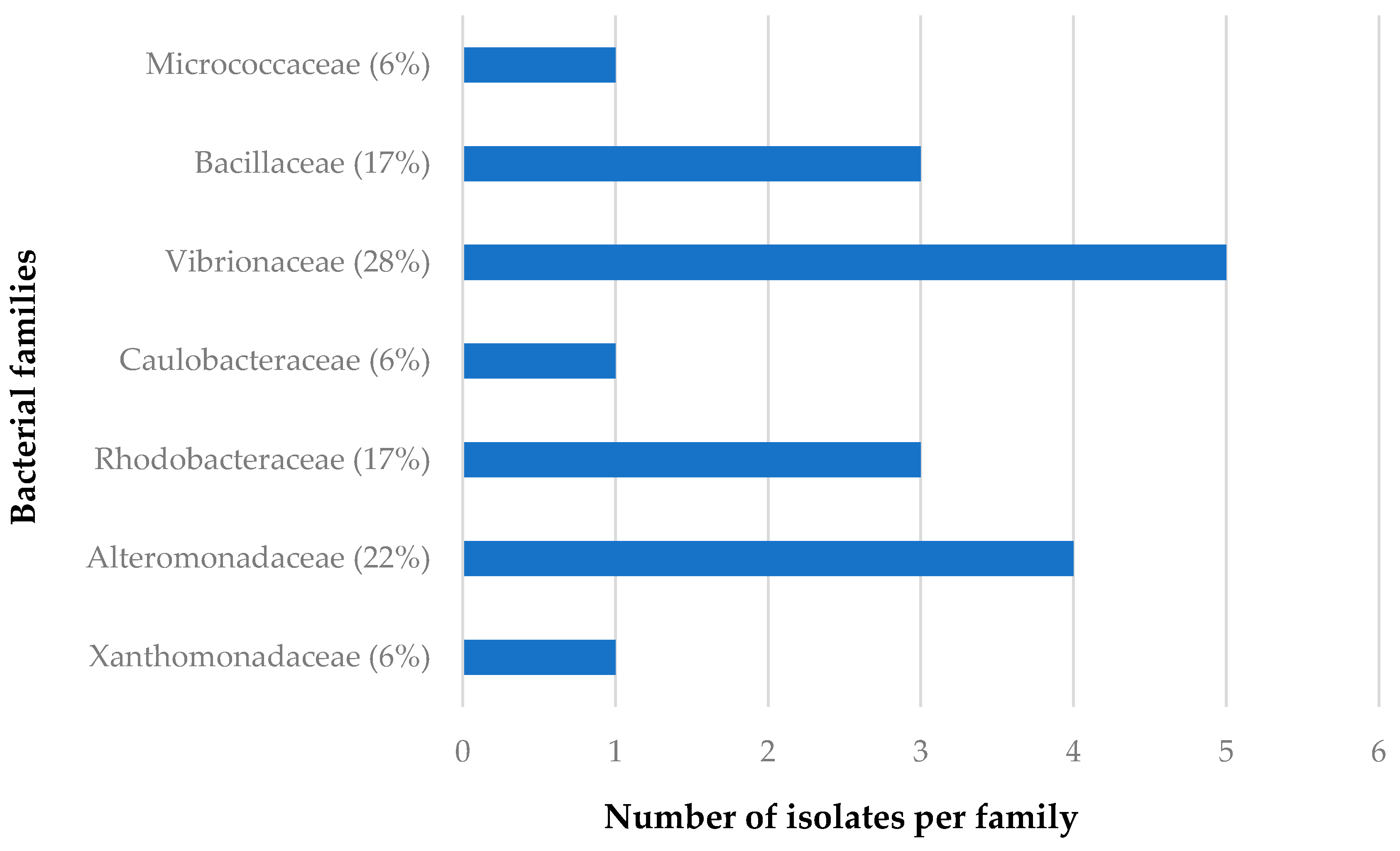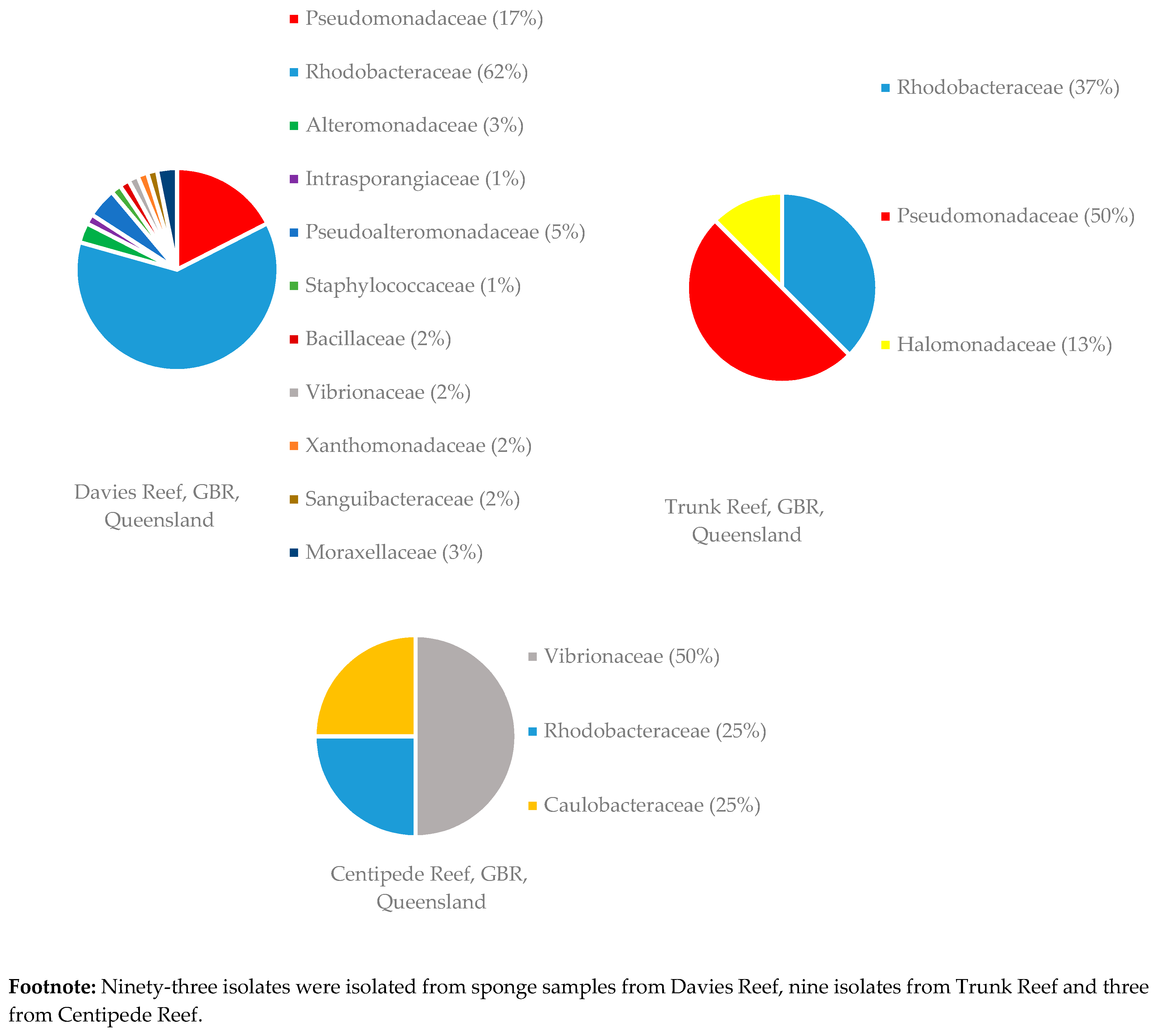Abstract
Sponges and their associated microbial communities have sparked much interest in recent decades due on the abundant production of chemically diverse metabolites that in nature serve as functional compounds required by the marine sponge host. These compounds were found to carry therapeutic importance for medicinal applications. In the presented study, 123 bacterial isolates from the culture collection of the Australian Institute of Marine Science (AIMS) previously isolated from two different sponge species, namely Candidaspongia flabellata and Rhopaloeides odorabile, originating from different locations on the Great Barrier Reef in Queensland, Australia, were thus studied for their bioactivity. The symbiotic bacterial isolates were first identified using 16S rRNA gene analysis and they were found to belong to five different dominating classes of Domain Bacteria, namely Alphaproteobacteria, Gammaproteobacteria, Flavobacteria, Bacilli and Actinobacteria. Following their taxonomical categorization, the isolates were screened for their antimicrobial activity against human pathogenic microbial reference strains: Escherichia coli (ATCC® BAA-196™), E. coli (ATCC® 13706™), E. coli (ATCC® 25922™), Klebsiella pneumoniae (ATCC® BAA-1705™), Enterococcus faecalis (ATCC® 51575™), Bacillus subtilis (ATCC® 19659™), Staphylococcus aureus (ATCC® 29247™), Candida albicans (ATCC® 10231™) and Aspergillus niger (ATCC® 16888™). Over 50% of the isolates displayed antimicrobial activity against one or more of the reference strains tested. The subset of these bioactive bacterial isolates was further investigated to identify their biosynthetic genes such as polyketide synthase (PKS) type I and non-ribosomal peptide synthetase (NRPS) genes. This was done using polymerase chain reaction (PCR) with degenerate primers that have been previously used to amplify PKS-I and NRPS genes. These specific genes have been reported to be possibly involved in bacterial secondary metabolite production. In 47% of the bacterial isolates investigated, the PKS and NRPS genes were located. Some of the bacterial isolates were found to possess both gene types, which agrees with the previous reported biosynthetic ability of certain sponge-symbiotic bacteria such as the Actinobacteria or Gammaproteobacteria to produce secondary metabolites with antimicrobial activity. All these reported activities further confirm that sponge-symbiotic bacteria hold significant bioactivity with medicinal and biotechnological importance.
1. Introduction
The marine environment has become an important source for natural product discovery due to the uniqueness and complexity of marine-derived metabolites that may contribute significantly towards discovery of novel and potent antibiotics [1]. Extremity of the environmental factors present in these marine environments (e.g., varying salt concentration, hydrostatic pressure, the range of available marine nutrients present in these environments) and the symbiosis with marine macro-organisms (e.g., the invertebrates) [2] would, in turn, induce changes in microbial metabolism, resulting in the production of chemically diverse compounds [3].
Sponges, of the phylum Porifera, have been the largest source of bioactive compounds providing a greater number of novel metabolites than any other marine taxon each year, contributing nearly 30% of all of the natural marine compounds discovered [4,5,6,7,8,9,10,11].
Sponges are among the oldest and most stable metazoans known to inhabit this planet [12] and appear in most tropical and temperate aquatic habitats. They exist in various shapes, sizes and colours [13] and are highly evolved and successfully adapted organisms [14,15,16].
Microorganisms are associated with marine sponges either transiently and come into contact through the host’s filter feeding mechanism or live symbiotically within the host and are responsible for a range of functional metabolic activities aiding the survival of the host including the production of defense metabolites [5,12,17]. These sponge-associated microorganisms can make up 35–60% of the total sponge mass [5,7,11,18,19]. Previously, compounds isolated from sponges were thought to be produced by the sponge itself however, further research identified abundant evidence that microorganisms living within the sponge are in most cases responsible for the production of most of these metabolites of interest [4,5,20,21]. Examples include the production of the glycoglycerolipid with anti-tumour properties by the sponge Halichondria panacea as well as from a Microbacterium species found within the sponge. Several quinolones with cytotoxic and antimicrobial properties were also isolated from the sponge Homophymia sp. as well as from a pseudomonad species isolated from the same sponge [5].
Microorganisms, including the sponge-associated bacteria, may produce biologically active compounds which requires the activation of specific gene clusters encoding multi-modular enzymes. These gene clusters may be in the form of non-ribosomal peptide synthetases (NRPS) and polyketide synthases (PKS), however, other gene clusters may also be responsible for biological activity. The variations of these enzymatic modules (PKS and NRPS) results in the production of numerous biologically active compounds [8,22,23]. In order to produce these types of secondary metabolites, a set of domains are required within the bacterial biosynthetic pathways; these include ketosynthase (KS), acyltransferase (AT) and acyl carrier proteins in PKSs and adenylation (A), condensation (C) and peptidyl carrier proteins (PCPs) for peptide elongation in NRPSs [8]. Furthermore, antibiotic compounds such as tetracycline and erythromycin, anticancer agent, e.g., bleomycin and the immunosuppressive agent, e.g., rapamycin [24,25] were reported to be produced from PKS and NRPS pathways. Accordingly, complex polyketides isolated from sponges and their associated microorganisms are claimed to be the most promising biologically active compounds with therapeutic applications [8,26]. Examples of these important polyketides include swinholide from the sponge Theonella swinhoei [27,28] and mayamycin produced by Streptomyces sp. HB202 from the sponge Haliclona simulans and H. panicea [29,30] that have potent antibiotic activities. Halichondrin B isolated from the sponge Halichondria okadai also displays potent anticancer activity [31,32].
The sponge species Candidaspongia flabellata is a rare and biochemically active Dictyoceratida sponge [7], noteworthy for the diversity and potency of its homosesterterpene and bishomosclarane secondary metabolite production. Rhopaloeides odorabile is a common Dictyoceratida sponge species to the Great Barrier Reef and is known to house an uncommon group of C20 diterpenes [33]. Both these sponge species are known to house a variety of bacterial genera [7,34]. Previously, the full cultivatable heterotrophic bacterial community associated with C. flabellata was identified by Burja and Hill [7] and the main bioactive compound isolated from this sponge species was fanolide, which has been shown to inhibit the growth of some tumour cells [7]. Phylogenetic analysis of the cultivatable community of R. odorabile has also been carried out by Webster et al. [33] which was found to be dominated by a single bacterial strain of the class alphaproteobacteria [33] with bioactivity. Therefore, by isolating and fermenting in conditions conducive to triggering biosynthetic gene pathways (e.g., PKS and NRPS) involved in metabolite production in these marine sponge bacteria, novel compounds with antimicrobial activity may be recovered.
In light of the information presented above, the aims of this study were (1) molecular level identification of previously isolated sponge-associated bacteria by the Australian Institute of Marine Science (AIMS); (2) examination of their potential to produce antimicrobial compounds active against human pathogenic and antibiotic-resistant bacteria; and (3) detection of their biosynthetic genes potentially involved in the production of some of these biologically-active metabolites.
2. Materials and Methods
2.1. Sponge Collection and Isolation of Bacteria
A set of sponge associated bacteria from the marine microbial culture collection of the Australian Institute of Marine Science (AIMS, http://aims.gov.au/) was supplied to the University of the Sunshine Coast (USC) for further analysis. Sponge samples were collected by the AIMS by SCUBA as described by Webster and Hill [35]. Sections of the sponge were removed using sterile scalpel blades and the tissue transferred directly into a plastic sampling bags that contained seawater. Sponge tissue were processed within 15 min of collection. A 1 cm3 portion of the sponge was excised and rinsed briefly in 70% ethanol and quickly transferred to sterile artificial seawater (ASW). The sponge tissue was then removed from the ASW and cut into sections using a sterile scalpel and finely ground using a mortar and pestle [35]. Isolates (n = 105) collected from 12 different sponge samples from Candidaspongia flabellata and 18 isolates from the four different sponge samples of Rhopaloeides odorabile (n = 123) were collected from the Great Barrier Reef (see Appendix A, Table 1 and Table 2 for locations and numbers of sponge samples) over a two-year period. These bacterial isolates were grown on Marine agar 2216 (BD Difco™) at temperatures of 22 °C, 26 °C and 27 °C, respectively (under similar conditions to their tropical marine environment), for 2 to 21 days (Appendix A, Table A1 and Table A2). Bacterial isolates were then stored at 80 °C in a cryoprotectant solution (tryptone soy broth and 30% glycerol) and used for further testing as needed.

Table 1.
Presence of potential PKS-I and NRPS genes, closest relative strain and similarity in isolates from R. odorabile sponge samples exhibiting biological activity.

Table 2.
Presence of potential PKS-I and NRPS genes, closest relative strain and similarity in isolates from C. flabellata sponge samples exhibiting biological activity.
2.2. Molecular Identification of the Bacterial Isolates
2.2.1. DNA Extraction and 16S rRNA Gene Sequencing
Firstly, DNA extraction for Gram-negative isolates was carried out using the FastPrep®-24 Instrument and FastDNA® kit (MP Biomedicals, Irvine, CA, USA), according to the FastDNA® kit instruction manual protocol for bacterial DNA extraction. Gram-positive isolates were extracted using the DNeasy Blood and Tissue kit (Qiagen Inc., Frederick, MD, USA) according to the manual for the extraction of DNA from Gram-positive isolates. Isolates were tested for their cell wall types using the Gram-staining technique. DNA extracts were run on a 1% agarose gel (1 h at 110 V) to ensure successful DNA extraction and stored at −20 °C for extended periods and 4 °C until use.
All bacterial isolates were identified using Polymerase Chain Reaction (PCR) amplification of 16S rRNA genes using the HotstarTaq Plus Master Mix kit (Qiagen Inc.) following the instruction manual: PCR Using HotStarTaq Plus Master Mix. Universal primers B27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and U1492R (5′-GGTTACCTTGTTACGACTT-3′) were used to obtain an amplicon length of 1500 base pairs (bp). Cycling parameters for the amplification of the genes were carried out per the HotstarTaq Plus Master Mix instruction manual. Negative controls using sterile distilled water were used to ensure the amplified gene was not a result of contamination. PCR was performed in a T100 Thermal Cycler (BioRad) and the PCR products were viewed on a 1% agarose gel in Gel Doc™ XR+ Imager (BioRad). Amplified PCR products were sent to Macrogen Inc. (Seoul, Korea) for sequencing and the same universal primers were used.
2.2.2. Sequence Alignment and Phylogenetic Analysis
The 16S rDNA sequences were prepared using CLC genomic work bench (Qiagen Inc.) and were aligned using the standard nucleotide Basic Local Alignment Search Tool (BLAST) analysis with the National Centre for Biotechnology Information (NCBI) database (http://www.ncbi.nlm.nih.gov/) [36].
The ARB program [37] was used for sequence alignment and phylogenetic analyses. Only sequences >1300 (bp) were used. Phylogenetic trees were constructed using the maximum likelihood algorithm, with bootstrap analysis using 1000 data re-samplings. Phylogenetic trees were used to determine the relatedness of the isolates to their nearest relative. The sequences from bacterial isolates in this study were deposited to GenBank (See Appendix A, Table A1 and Table A2 for GenBank accession numbers).
2.3. Antimicrobial Assay
All bacterial extracts were tested for antimicrobial activity. Isolates were first grown in 20 mL of Marine broth (Benton Dickinson Difco™) at 28 °C on a floor-shaker (Bioline Global) at 150 rpm to obtain seed cultures. An amount of 5 mL of each bacterial sample was transferred into flasks containing 50 mL Marine broth (BD Difco™) and incubated at 28 °C on a floor-shaker (Bioline Global) at 150 rpm for 7 days [38]. The liquid fermentation medium was then centrifuged at 7000× g for 20 min [18] to obtain the cell-free supernatant (CFS). The CFSs were then extracted twice with ethyl acetate (EtOAc, Honeywell Research Chemicals) [18,38] and the organic phase was evaporated to dryness on a rotary evaporator (BÜCHI™ Rotavapor R-205) at 37 °C to obtain a crude extract [11,39,40,41]. In this instance, bioactivity was tested using the method by Dashti, et al. [42]. Crude extracts were weighed and dissolved in known volumes (500 μL) of dimethyl sulfoxide (DMSO, Sigma-Aldrich, St. Louis, MO, USA). 50 μL of a 500–1000 μg/mL solution of extract was loaded onto sterile commercial blank discs (Oxoid).
The reference strains were: American Type Culture Collection (ATCC) reference strains, Escherichia coli (ATCC® BAA-196™), E. coli (ATCC® 13706™), E. coli (ATCC® 25922™), Klebsiella pneumoniae (ATCC® BAA-1705™), Enterococcus faecalis (ATCC® 51575™), Bacillus subtilis (ATCC® 19659™), Staphylococcus aureus (ATCC® 29247™), Candida albicans (ATCC® 10231™) and Aspergillus niger (ATCC® 16888™).
Reference strains and the yeast C. albicans were grown for 18 to 24 h in Mueller–Hinton (MH) broth at 37 °C and their optical density (OD) measured at 600 nm to obtain an OD reading of 0.063 corresponding to a McFarland tube density of 0.5 [43]. An amount of 200 µL of each reference strain was then spread onto MH agar plates into which wells (6 mm in diameter) were made and the CFS from the different isolates inoculated. Whereas, for the antifungal activity testing, the fungal inoculum was prepared on a potato-dextrose agar (PDA) plate. Following full growth, an agar plug was taken and placed in the center of a new PDA plate to allow the growth in a concentric ring fashion. After 48 h of growth on the new plate, wells were made on the PDA plate for the inoculation of the CFSs. Discs loaded with crude extracts were also placed evenly apart on the MH agar inoculated with the ATCC pathogenic reference strains. Plates were incubated for 24 to 48 h at 37 °C for bacteria and yeast [44] and for 48 to 96 h at 28 °C for the fungal strain. Marine broth and dimethyl sulfoxide (DMSO) were used as negative controls. Vancomycin (Oxoid) and gentamicin (Oxoid) were used as the positive controls against the different bacteria and cycloheximide (Oxoid) against fungi. Following incubation at the above given conditions, the plates were examined for zones of inhibition.
2.4. Amplification of Type 1 Polyketide Synthase and Non-Ribosomal Peptide Synthetase Genes
The occurrence of specific genes involved in the reported production of secondary metabolites [45] was screened in all isolates. For all the heterotrophic bacteria (except the actinobacteria), degenerate primers MDPQQRf (5′-RTRGAYCCNCAGCAICG-3′) and HGTGTr (5′-VGTNCCNGTGCCRTG-3′) [45] were used to amplify the B-ketosynthase (KS) domain fragment within the Type I polyketide synthase genes (PKS-I). Amplification of the non-ribosomal peptide synthetase (NRPS) gene was carried out using primers MTF (5′-CCNCGDATYTTNACYTG-3′) and MTR (5′-GCNGGYGGYGCNTAYGTNCC-3′) to amplify the conserved A domain [46]. The actinobacteria required a different set of primers and K1 (5′-TSAAGTCSAACATCGGBCA-3′) and M6R (5′-CGCAGGTTSCSGTACCAGTA-3′) were used to amplify PKS-I ketosynthase and methyl-malonyl-CoA transferase sequences and A3F (5′-GCSTACSYSATSTACACSTCSGG-3′) and A7R (5′-SASGTCVCCSGTSCGGTAS-3′) were used to amplify NRPS adenylation sequences [47,48]. An amount of 10% DMSO was also added to each reaction for the actinobacteria [49]. Amplification was carried out using the HotStarTaq Plus Master Mix Kit (Qiagen) as per cycling conditions given in the HotStarTaq Plus Manual. An annealing temperature of 56 °C was used for amplification of genes for the actinobacteria. PCR was performed as described above for Section 2.2.1.
3. Results
3.1. Molecular Identification of the Isolates
3.1.1. Phylogenetic Analysis Based on 16S rRNA Gene Sequencing
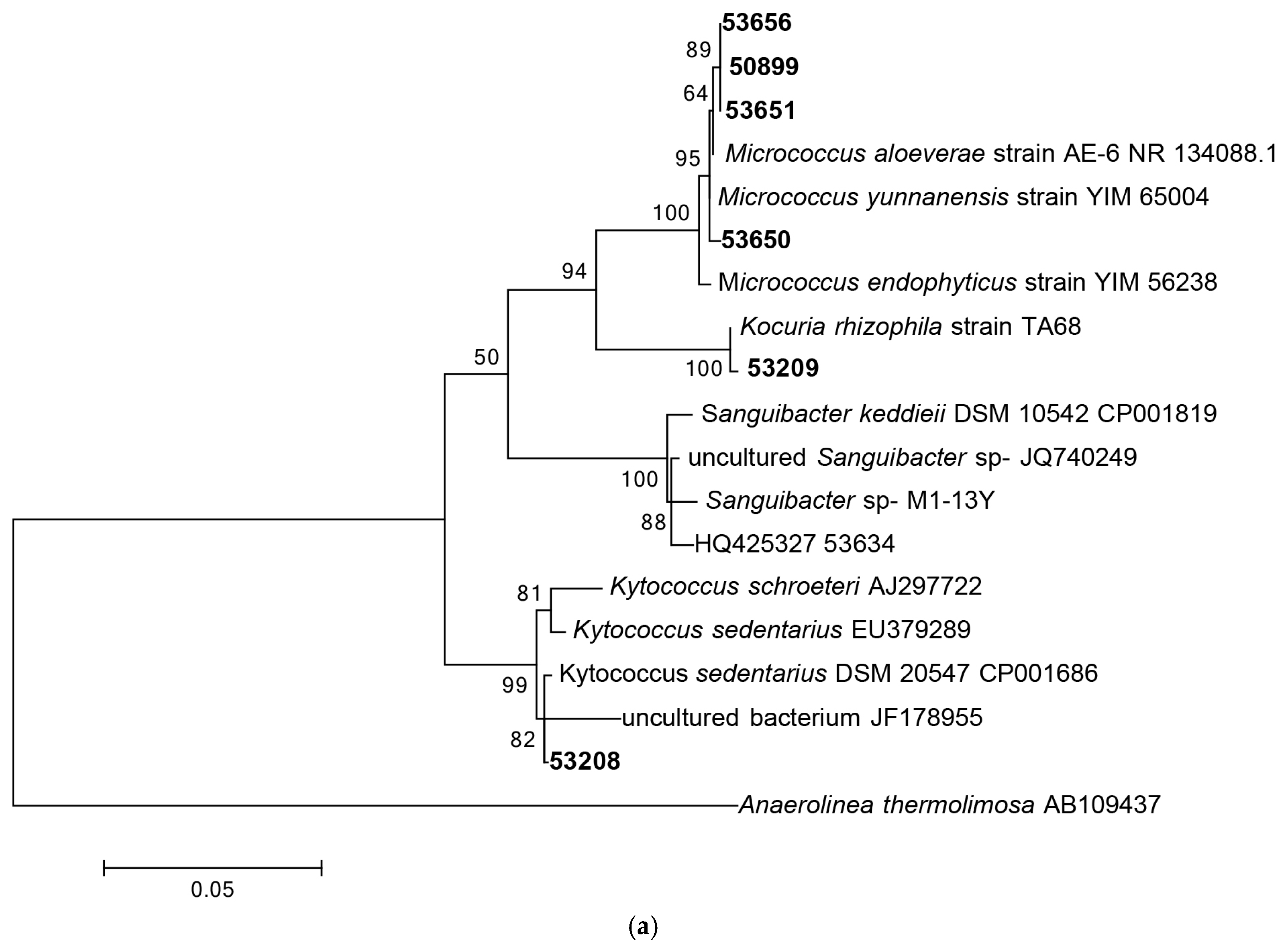
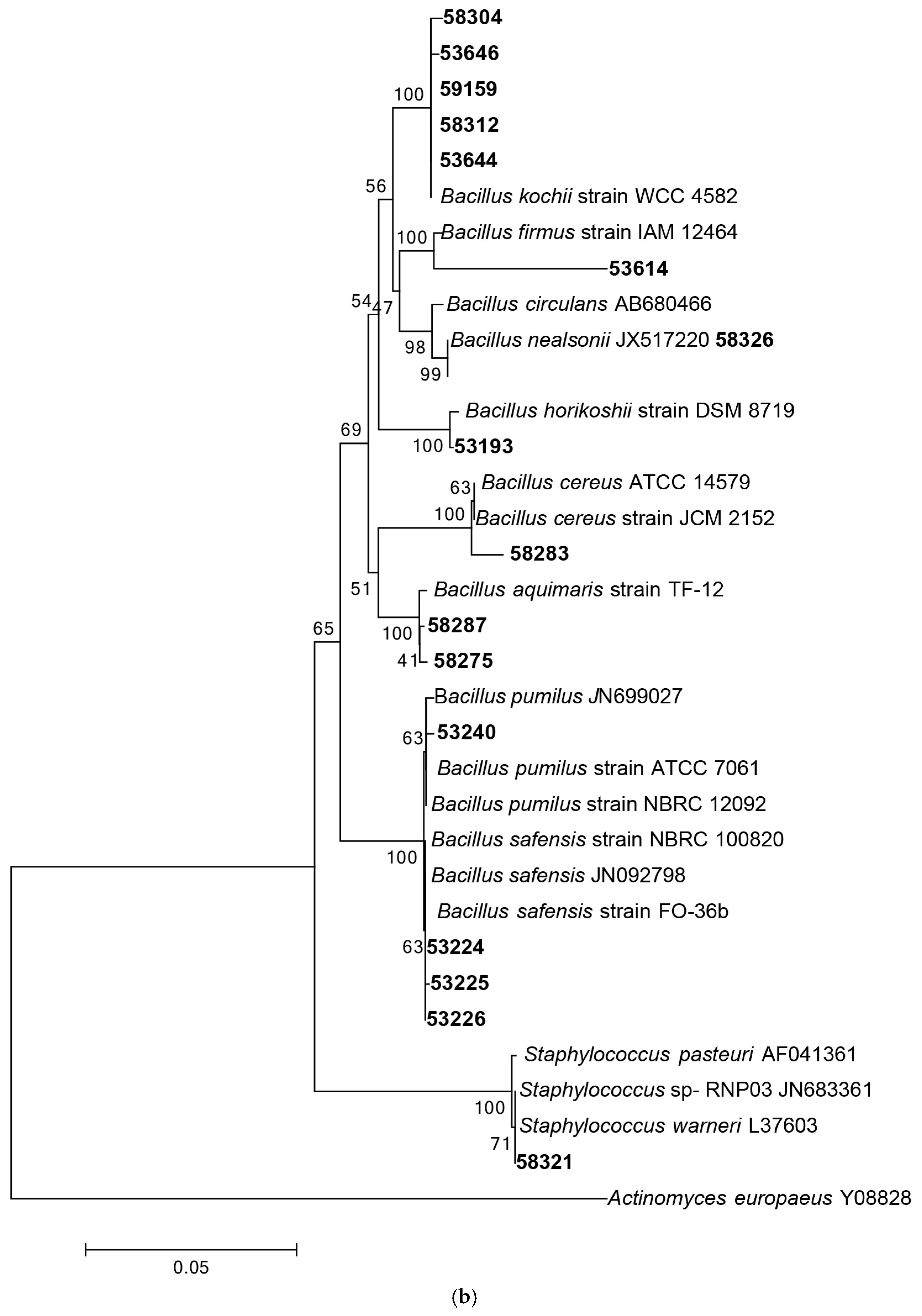
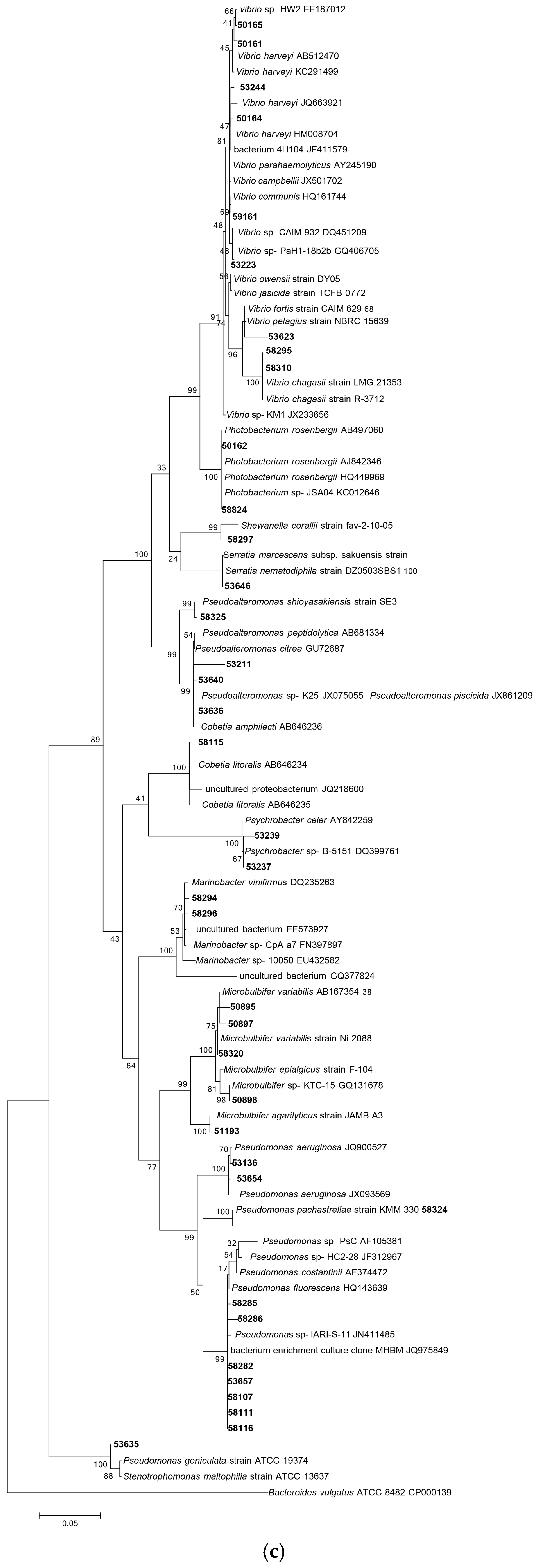
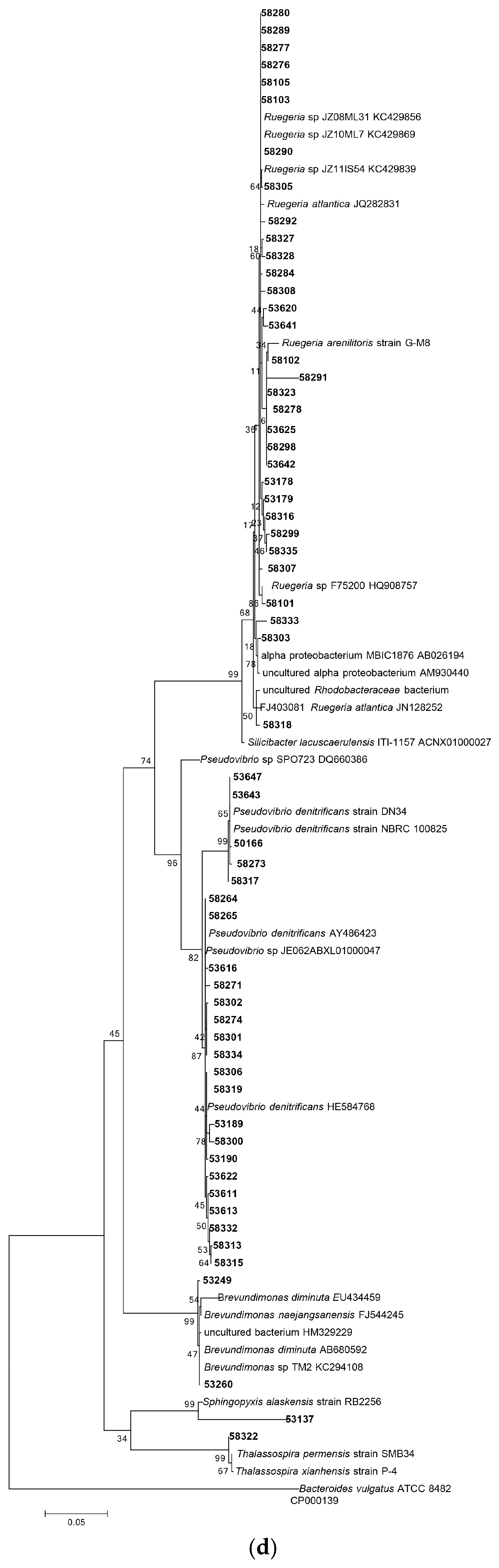
Following 16S rRNA gene sequencing, the isolates were found to belong to five major taxonomic classes of Domain Bacteria that were namely Gammaproteobacteria, Alphaproteobacteria, Bacilli, Actinobacteria and Flavobacteria. Analysis of the 16S rRNA gene sequences revealed the taxonomic position of each isolate in relation to their closest relative strains and the phylogenetic trees (Figure 1) were constructed according to these major classes. However, there was only one isolate belonging to the class Flavobacteria (isolate 58330), isolated from C. flabellata and its closest relative strain was Aquimarina spongiae strain A6 [50]. Closest relative species for each isolate as well as the percentage of similarity is given in Table 1 and Table 2.
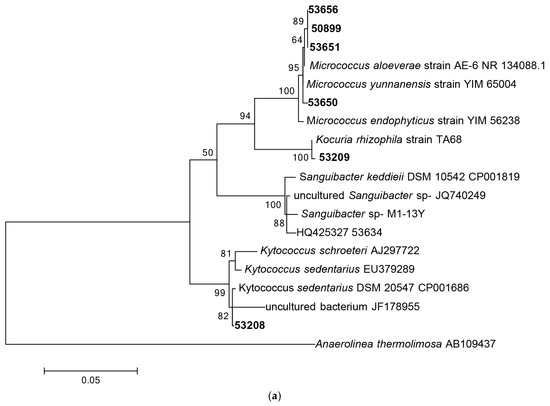
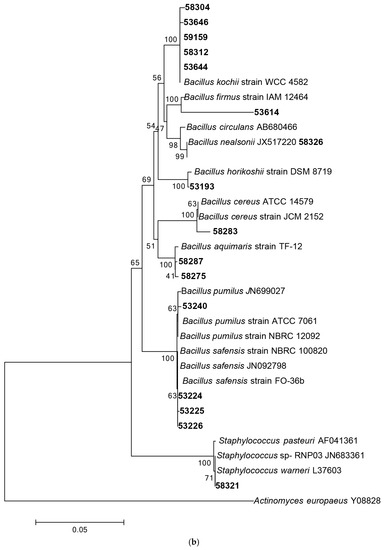
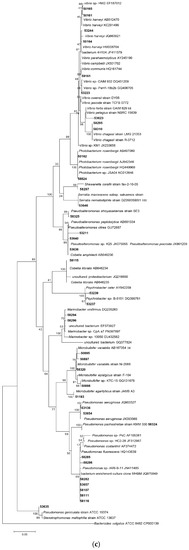
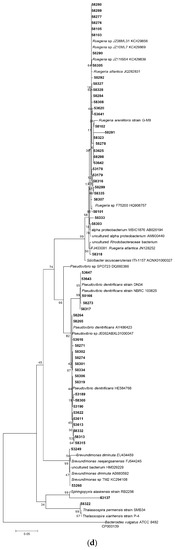
Figure 1.
Phylogenetic diversity of bacteria associated with sponge species C. flabellata and R. odorabile located on the Great Barrier Reef in Queensland, Australia. Phylogenetic trees were constructed using the maximum likelihood algorithm with bootstrap analysis using 1000 data re-samplings within Mega 6 [51]. Trees represent the phylogenetic diversity of the classes Actinobacteria (a); Bacilli (b); Gammaproteobacteria (c) and Alphaproteobacteria (d). Bacteria used as an out-group to root the different trees include, Anaerolinea thermolimosa for Actinobacteria, Actinomyces europaeus for Bacilli and Bacteriodes vulgatus for the Alpha- and Gammaproteobacteria. The scale bar represents 5% sequence divergence.
3.1.2. Distribution of Isolates in Relation to Their Sponge Hosts and Sponge Collection Locations
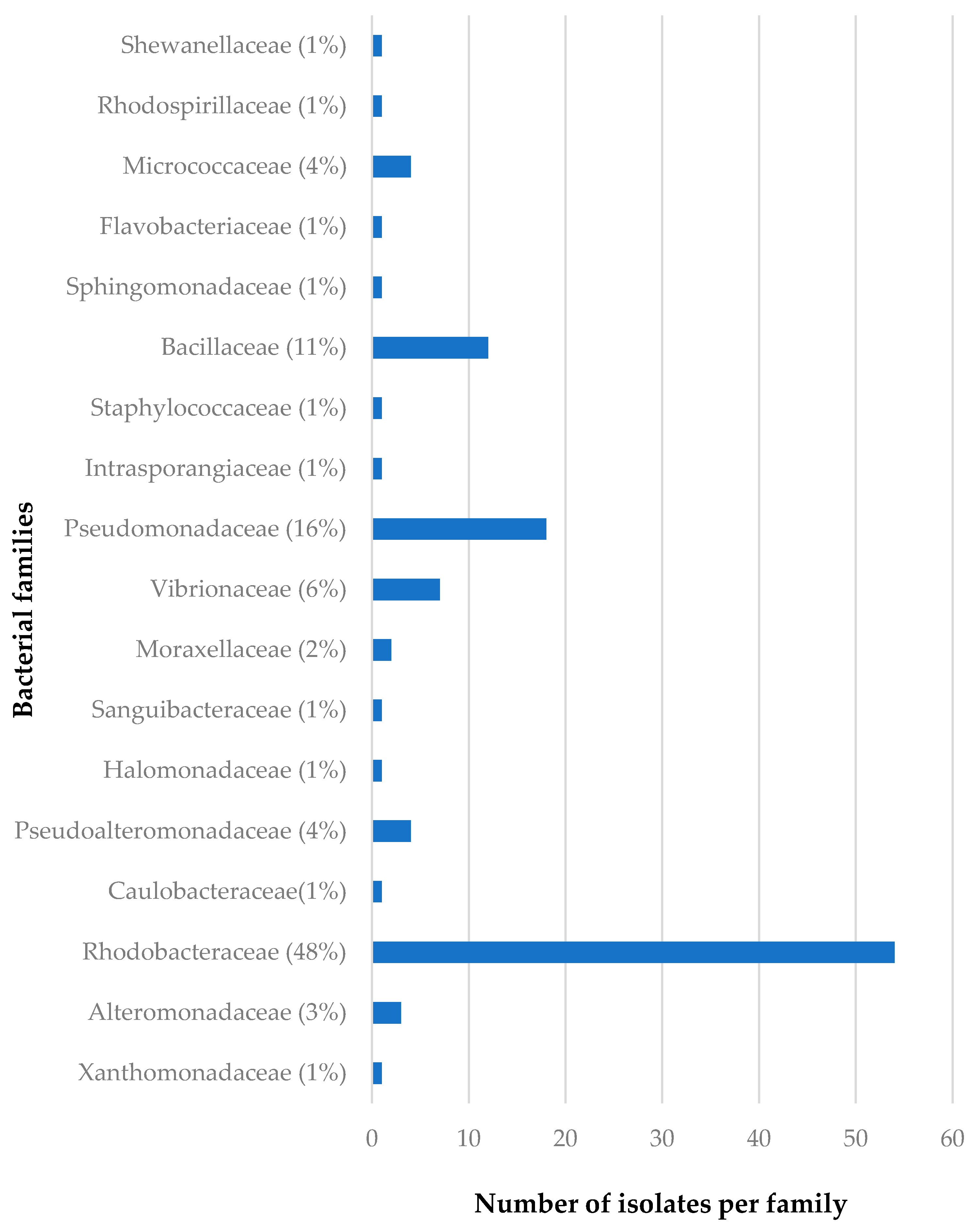
The distribution of the bacterial families isolated from C. flabellata and R. odorabile in relation to sponge species as well as to geographical locations of these sponges are given in Figure 2, Figure 3, Figure 4 and Figure 5. The main bacterial families isolated from Candidaspongia flabellata belonged to Rhodobacteraceae (48%) and Pseudomonadaceae (16%, Figure 2); from Rhopaloeides odorabile, they were from Vibrionaceae (28%) and Alteromonadaceae (22%, Figure 3). Some of these isolates were found to be related to the members of families known to produce bioactive compounds [34,52,53,54].
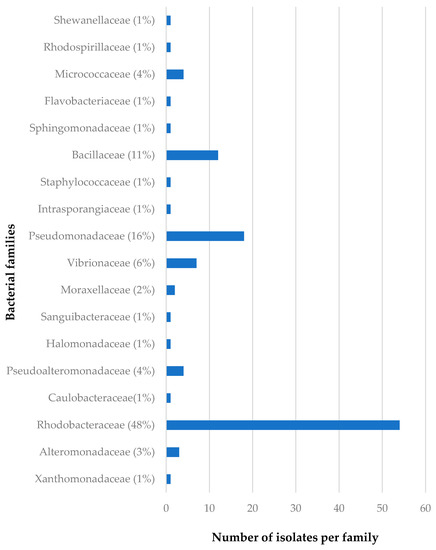
Figure 2.
The diversity of the bacterial families isolated from twelve different sponge samples of C. flabellata.
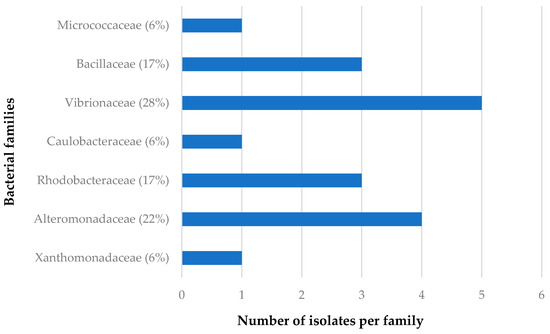
Figure 3.
The diversity of the bacterial families isolated from four different sponge samples of R. odorabile.
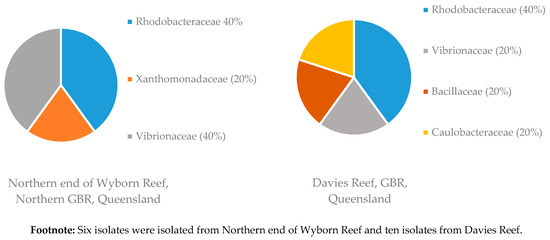
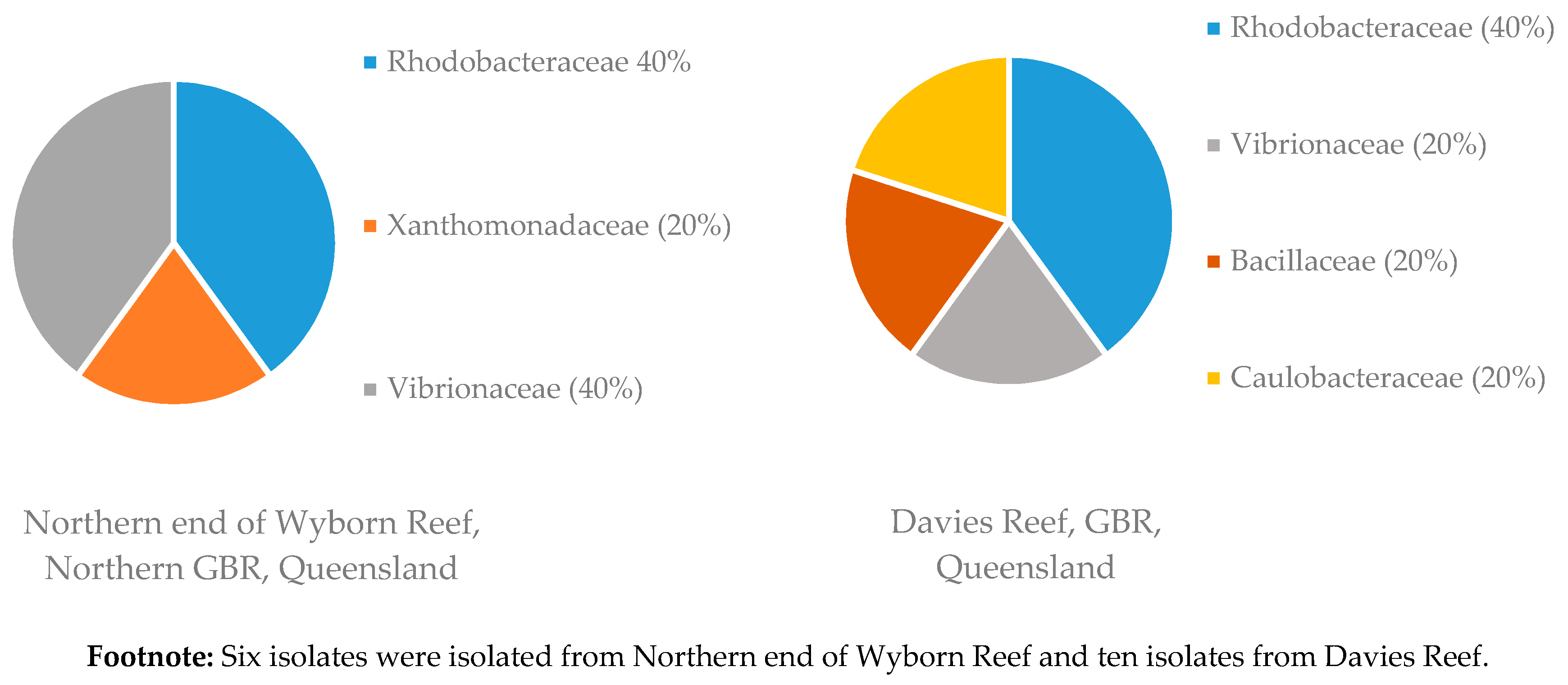
Figure 4.
Diversity of bacterial families isolated from two sponge samples of R. odorabile collected at different locations on the Great Barrier Reef (GBR). Footnote: Six isolates were isolated from Northern end of Wyborn Reef and ten isolates from Davies Reef.
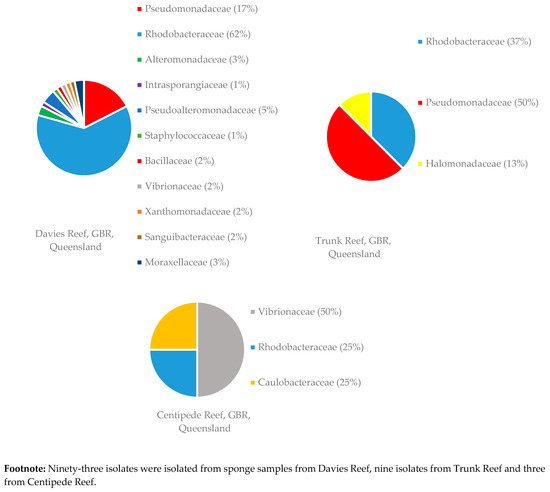
Figure 5.
Diversity of bacterial families isolated from 12 sponge samples of C. flabellata collected at various locations on the GBR. Footnote: Ninety-three isolates were isolated from sponge samples from Davies Reef, nine isolates from Trunk Reef and three from Centipede Reef.
Sponge samples of R. odorabile were collected from three different locations on the Great Barrier Reef, one sponge from Wyborn Reef, two sponges from the Davies Reef and a sponge from North East Percy Island. The familial diversity of two of these locations is given in Figure 4. From the sponge sample taken at North East Percy Island in the North-West corner of the Great Barrier Reef, only one family (two species of Alteromonadaceae were isolated) was able to be cultivated within the laboratory. Sponge samples of C. flabellata were collected from three locations on the Great Barrier Reef, ten sponges from the Davies Reef, one from the Trunk Reef and one from Centipede Reef. The familial diversity of these samples collected at the different locations is given in Figure 5. The reason for the increased number of bacterial isolates isolated from C. flabellata compared to R. odorabile is due to the increased number of C. flabellata sponge samples collected.
3.2. Antimicrobial Screening
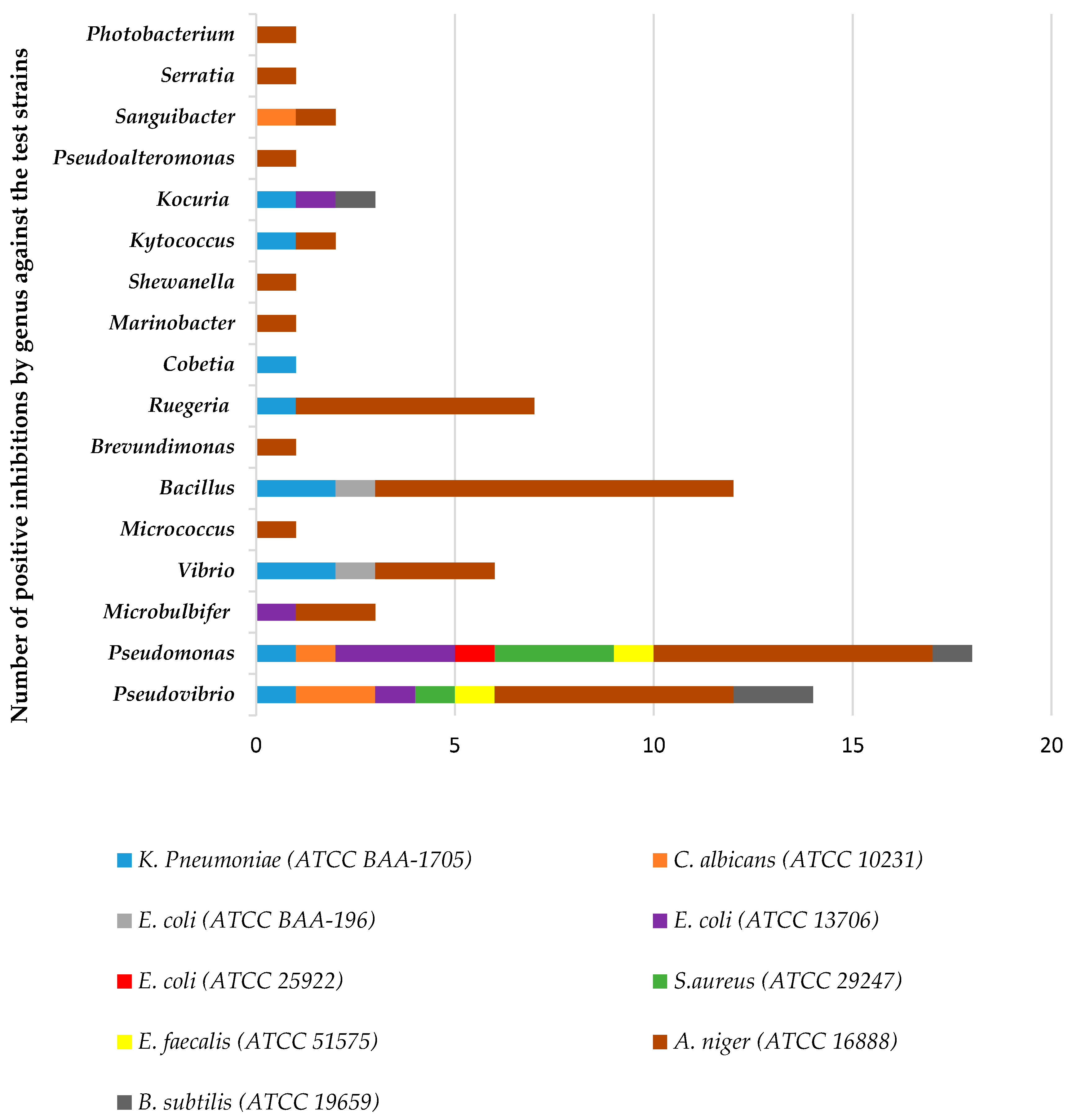
Out of the total isolates from both sponges, 51% exhibited inhibitory activity against one or more of the pathogenic test strains. The comparison of antimicrobial activity exhibited by the isolates at genus level is given in Figure 6. The isolates belonging to the genus Pseudomonas produced extracts with the highest activity against eight out of the nine test strains followed by Pseudovibrio (active against seven out of the nine test strains) and Bacillus strains (active against three of the test strains). Isolate 53654 with its closest relative strain being Pseudomonas aeruginosa PAO1 strain PAO1 was active against eight of the nine strains. Isolate 58264, closest relative strain being Pseudovibrio sp. FO-BEG1 strain FO-BEG1 was also active against six out of the nine strains with other isolates with this closest relative strain being highly active too. Among all the isolates belonging to the class actinobacteria, isolates belonging to genera Kocuria and Kytococcus produced extracts with the highest antimicrobial activity. Isolates with their closest relative strain, similarity and activity spectrum are given in Table 1 and Table 2.
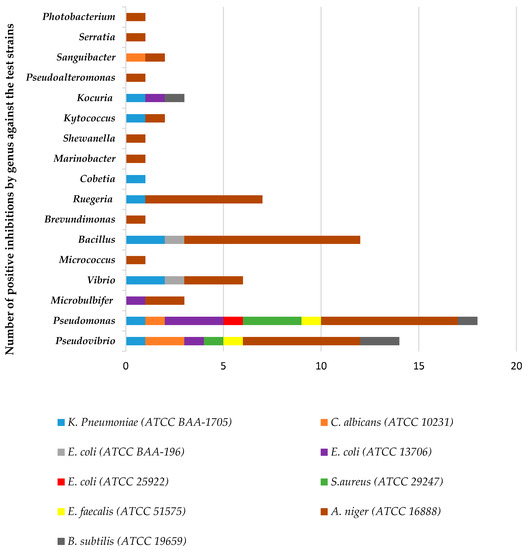
Figure 6.
Number of antimicrobial hits from sponge associated-genera obtained against pathogenic test organisms used in the antimicrobial assays.
3.3. Amplification of PKS-I and NRPS Genes
Detection of PKS-I and NRPS genes in all isolates was indicated by the presence of the corresponding fragment size range: 700 bp for PKS-I and 750–1000 bp for NRPS in non-actinobacteria and 1250–1400 bp for PKS-I and 700 bp for NRPS in actinobacteria. Of all isolates (n = 123) analyzed, 47% had at least one of the gene types. In total, 22% of isolates from R. odorabile possessed one gene type, PKS-I. 51% of isolates from C. flabellata possessed at least one of the gene types with PKS-I dominating and 5% of these C. flabellata isolates possessed both PKS-I and NRPS genes. The presence of these genes as well as the activity spectrum and closest relative strain for all isolates is given in Table 1 and Table 2.
4. Discussion
Due to the emergence and re-emergence of multi-drug resistant microorganisms, it is vital that new antimicrobial compounds are discovered that counteract the resistant mechanisms exhibited by these microorganisms. These include methicillin-resistance exhibited by S. aureus and multi-drug resistance exhibited by E. coli, K. pneumoniae, E. faecalis and C. albicans [24,55,56]. Currently, one of the most effective ways of discovering novel therapeutic agents is the cultivation and fermentation of novel or under-studied microorganisms isolated from diverse environments such as the marine environment [1]. In the presented study, antimicrobial activity was detected when sponge-associated isolates were fermented in marine broth, indicating their potential to produce bioactive compounds under marine sponge conditions. Due to the marine broth having a high salt content and other nutrients that help simulate the sea water environment, it is a highly suitable medium for marine bacterial growth. Marine broth may possess greater levels of nutrients compared to the surrounding sea water; however, due to constant seawater filtration by the sponge, these isolates may come into contact with more nutrients within the host environment as well as having access to other nutrients that may be provided by the sponge host [57]. Under diverse fermentation conditions with different nutrients and parameters, these bacteria may have the ability to produce more than one type of active metabolite. Here, the simple and previously successful one-strain-many-active-compounds (OSMAC) approach may be used which aims to activate metabolic pathways to produce different types of metabolites in order to identify the range of compounds that may be produced by a specific bacterial isolate [58]. The use of different fermentation nutrients and parameters as well as co-culturing of competing or antagonistic microorganisms may aid in the search for important active metabolites that may be used as antimicrobial or anticancer agents [34,59]. A study carried out by Graça et al. (2015) reported that the use of liquid fermentation, the same technique used in this study, over solid media fermentation to obtain antimicrobial extracts is advantageous in its ability to be quantitative, simple, quick to carry out, reproducible, less expensive than other methods used and able to be conducted in a high throughput way [8]. For this study, liquid fermentation was chosen as the main medium type as this was less time consuming for extraction, quick and simple and less expensive than when solid-state fermentation was trialed.
Detection of bioactive actinobacterial isolates from the genera Kocuria and Kytococcus once again confirmed that actinobacteria are prolific producers of compounds with antibiotic properties [44]. Previous examples include the production of Kocurin by sponge-derived Kocuria and Micrococcus sp. active against methicillin-resistant S. aureus [44]. Other well known-bioactive compound producers from different genera include Pseudoalteromonas [8,21,60], Bacillus [61], Pseudovibrio [62] and Pseudomonas [8,63] which is consistent with the findings of the presented study that isolates belonging to these genera were also found to produce antibacterial activity against the pathogenic reference strains.
Previously, a strain from the genus Pseudoalteromonas, Pseudoalteromonas piscida, isolated from a sponge species from the China Sea, Hymeniacidon perlevis, has been identified as a producer of the wide spectrum antimicrobial metabolite, norharman, a β-carboline alkaloid [21,64]. This genus has been well studied and out of the 41 species hosted by Pseudoalteromonas 16 of them were illustrated to produce antimicrobial metabolites [65]. Bacillus species from a range of marine macroorganisms have also been found to produce metabolites with antimicrobial and antifouling properties [66]. One Bacillus sp. in particular isolated from the sponge Halichondria sp. was found to have potent activity against a range of clinically pathogenic microorganisms. This species was found to be closely related to Bacillus licheniformis HNL09 and produce compounds such as indole, 3-phenylpropionic acid and dimer 4,4′-oxybis [3-phenylpropionic acid] [38]. The genus Pseudovibrio is known as the second most prolific bacterial genus that has been isolated from sponges and produces antimicrobial activities. This genus produces the antibacterial compound tropodithietic acid [29]. Pseudomonas sp. from sponges have also been found to produce potent antimicrobial activity against a range of bacteria, including resistant ones. Pseudomonas sp. from the sponge Callyspongia sp. was found to produce a chomophore substance against methicillin resistant S. aureus [67].
The antimicrobial assays in this study allowed for the observation of a high number of bacterial extracts active against three main reference strains: the fungal strain A. niger, the bacterial strain K. pneumoniae and the yeast C. albicans. Furthermore, the greatest level of bioactivity detected in the presented study against K. pneumoniae, S. aureus and A. niger originated from the genera Pseudomonas, Pseudovibrio and Bacillus. These genera have also been identified as important producers of biologically active compounds from many other studies [17,34,68].
Molecular identification of the sponge-associated bacteria involved in this study indicated that isolates belonged to five different bacterial classes. The main two groups of bacteria identified were alpha- and gammaproteobacteria and this finding was consistent with a previous study carried out by Thomas et al. [12] who investigated the sponge microbiome across 81 different sponge species from 20 different countries. They also reported that the main phylum identified was Proteobacteria across all the samples studies, with the divisions’ alpha- and gammaproteobacteria of this phylum being the most dominant [12]. These two bacterial classes were consistent across both sponge species (C. flabellata and R. odorabile) as being the most abundant with gammaproteobacteria dominating in R. odoabile. The dominant class from C. flabellata sponge samples was alphaproteobacteria which is comparable to the high number of isolates identified with this class from a previous study carried out on C. flabellata [7]. A previous study investigating the bacterial diversity on another Great Barrier Reef sponge, Cinachyrella sp., also found that these two classes were the most abundant [69]. Bacterial isolates identified to their closest relative that were common to both sponge species include Pseudovibrio sp., a Micrococcus sp., a Brevundimonas sp., a Vibrio sp., a Microbulbifer and a Photobacterium sp. The genera that prevailed among the sponge samples from C. flabellata were Pseudovibrio, Ruegeria and Bacilli and from R. odorabile, Pseudovibrio, Vibrio and Bacilli suggesting a similar diversity between the two Great Barrier Reef sponge species even though the cultivable community isolate numbers from R. odorabile was considerably lower than that of C. flabellata. The fact that these bacterial species have also been found to be associated with other sponge species from different reef locations around the world [8,18,70,71,72,73] suggests that these symbiotic bacterial classes may be common associates of marine sponges, even in the distantly related ones.
Amplification of specific genes, such as PKS-I and NRPS genes, within bacterial biosynthetic pathways is a valued method in the search for new bioactive metabolites as the presence of these genes might indicate the ability of bacterial isolates to produce metabolites with bioactivity of medical importance such as the antibacterial or anticancer activities if they are switched on using the right fermentation conditions [8,22,74]. PKS-I and NRPS genes were amplified in both bioactive isolates as well as isolates that did not produce antimicrobial. Of all isolates analyzed from both sponge species, 47% potentially had at least one of the gene types. The dominant bacterial genera, Pseudovibrio, Bacillus and Ruegeria, across all of the sponge samples tested in this study were found to carry these genes. This finding was in line with the previous studies that reported the existence of such genes in these genera, however, to the best of our knowledge, this is the first time the PKS-I gene has been potentially located in a member of the genera Microbulbifer isolated from R. odorabile but not from C. flabellata. Moreover, the isolates that had not displayed any activity but possessed one or both gene types may be due to inadequate activation of the gene biosynthetic pathway or the fact that not all PKS-I and NRPS genes might result in antimicrobial activity but may produce other activity such as anticancer activity [48]. Isolates may possess these biosynthetic genes, however, the required parameters and nutrients to induce activation [75,76] may have not met with the marine broth fermentation alone. Therefore, further fermentation trialing a range of parameters and nutrients is required. One way to trigger these silent biosynthetic gene pathways may be the use of rare earth elements (REEs) in fermentation media. These REEs consist of 17 elements which include scandium, yttrium and the lanthanides (15 elements from lanthanum to lutetium) and their use in fermentation media has been shown to cause the overproduction of antibiotics and the activation of silent genes within bacteria. An example includes the addition of scandium added to different Streptomyces (S. coelicolor, S. griseus, S. antibioticus and S. lividans) cultures that enhanced the antibiotic production by 2–25-fold [76,77]. The use of these elements also eliminates the need to use gene engineering technology or strain genomic information due to them being scattered throughout the global environment which suggests that microorganisms may have acquired the ability to respond to low levels of REEs as a way of adapting to the surrounding environment [76].
5. Conclusions
The isolation and molecular identification of these cultivatable bacteria from the two different sponge species, C. falbaellata and R. odorabile, revealed broad diversity of taxa. The results also illustrate the potential of bacteria associated with these two different sponge species to produce bioactive metabolites against pathogenic microorganisms which indicate that these bacterial strains might be of greater value if further studies are conducted on them, such as, genome mining, different fermentation conditions, silent gene activation and so on, that in turn may result in the discovery of novel metabolites of therapeutic value. Chemical dereplication of extracts will also be conducted to evaluate the number of known and novel compounds.
Acknowledgments
The authors would like to thank the Australian Institute of Marine Science for providing the sponge bacterial isolates and for their expert support. First Author gratefully acknowledges the Australian Government Research Training Program scholarship.
Author Contributions
Candice M. Brinkmann and D. İpek Kurtbӧke designed and conceived the experiments. Candice M. Brinkmann performed the experiments, carried out the antimicrobial assays, conducted the molecular work and analysed the microbiological and molecular data. Philip S. Kearns and Elizabeth Evans-Illidge supplied the bacterial samples that were collected by AIMS. Candice M. Brinkmann wrote the manuscript. D. İpek Kurtbӧke and Philip S. Kearns supervised Candice M. Brinkmann’s PhD project and provided feedback on the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
Appendix A

Table A1.
Incubation conditions, Genbank accession numbers and location of R. odorabile collection for bacterial isolates.
Table A1.
Incubation conditions, Genbank accession numbers and location of R. odorabile collection for bacterial isolates.
| Isolate Code | Closest Relative Strain | Sponge Sample Number | Purification Temperature (°C) | Days Grown | GenBank Accession Number | Latitude | Longitude | Location of Sponge Samples at the Great Barrier Reef (GBR), Queensland, Australia |
|---|---|---|---|---|---|---|---|---|
| 50161 | Vibrio harveyi strain NBRC 15634 | 16595 | 22 | 14 | KX418475 | −10.823 | 142.743 | NORTHERN END OF WYBORN REEF, NORTHERN GBR |
| 50162 | Photobacterium rosenbergii strain CC1 | 16595 | 22 | 14 | KX418476 | −10.823 | 142.743 | NORTHERN END OF WYBORN REEF, NORTHERN GBR |
| 50163 | Stenotrophomonas maltophilia strain ATCC 19861 | 16595 | 22 | 14 | KX418463 | −10.823 | 142.743 | NORTHERN END OF WYBORN REEF, NORTHERN GBR |
| 50164 | Vibrio campbellii strain ATCC 25920 | 16595 | 22 | 14 | KX418477 | −10.823 | 142.743 | NORTHERN END OF WYBORN REEF, NORTHERN GBR |
| 50165 | Vibrio harveyi strain NBRC 15634 | 16595 | 22 | 14 | KX418478 | −10.823 | 142.743 | NORTHERN END OF WYBORN REEF, NORTHERN GBR |
| 50166 | Pseudovibrio denitrificans strain NBRC 100825 | 16595 | 22 | 14 | KX418552 | −10.823 | 142.743 | NORTHERN END OF WYBORN REEF, NORTHERN GBR |
| 50895 | Microbulbifer variabilis strain Ni-2088 | 17643 | 27 | 14 | KX418479 | −18.826 | 147.64 | DAVIES REEF |
| 50897 | Microbulbifer variabilis strain Ni-2088 | 17643 | 27 | 14 | KX418480 | −18.826 | 147.64 | DAVIES REEF |
| 50898 | Microbulbifer variabilis strain Ni-2088 | 17643 | 27 | 14 | KX418464 | −18.826 | 147.64 | DAVIES REEF |
| 50899 | Micrococcus aloeverae strain AE-6 | 17643 | 27 | 14 | KX418589 | −18.826 | 147.64 | DAVIES REEF |
| 51193 | Microbulbifer agarilyticus strain JAMB A3 | 17766 | 27 | 14 | KX418502 | −21.659 | 150.324 | DAVIES REEF |
| 53189 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 19500 | 27 | 3 | KX418515 | −18.833 | 147.617 | NE-PERCY IS., NW CORNER |
| 53190 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 19500 | 27 | 3 | KX418516 | −18.833 | 147.617 | NE-PERCY IS., NW CORNER |
| 53223 | Vibrio owensii strain DY05 | 19500 | 30 | 2 | KX418481 | −18.833 | 147.617 | DAVIES REEF, GBR |
| 53224 | Bacillus safensis strain NBRC 100820 | 19500 | 30 | 2 | KX418582 | −18.833 | 147.617 | DAVIES REEF, GBR |
| 53225 | Bacillus safensis strain NBRC 100820 | 19500 | 30 | 2 | KX418572 | −18.833 | 147.617 | DAVIES REEF, GBR |
| 53226 | Bacillus safensis strain NBRC 100820 | 19500 | 30 | 2 | KX418577 | −18.833 | 147.617 | DAVIES REEF, GBR |
| 53249 | Brevundimonas diminuta strain NBRC 12697 | 19500 | 30 | 6 | KX418517 | −18.833 | 147.617 | DAVIES REEF, GBR |

Table A2.
Incubation conditions, Genbank accession numbers and location of C. flabellata collection for bacterial isolates.
Table A2.
Incubation conditions, Genbank accession numbers and location of C. flabellata collection for bacterial isolates.
| Isolate Code | Closest Relative Strain | Sponge Sample Number | Purification Temperature (°C) | Days Grown | GenBank Accession Number | Latitude | Longitude | Location of Sponge Samples at the Great Barrier Reef (GBR), Queensland, Australia |
|---|---|---|---|---|---|---|---|---|
| 53136 | Pseudomonas aeruginosa PAO1 strain PAO1 | 19496 | 27 | 10 | KX418482 | −18.820 | 147.63 | DAVIES REEF, GBR |
| 53137 | Sphingopyxis alaskensis strain RB2256 | 19496 | 27 | 6 | KX418562 | −18.820 | 147.63 | DAVIES REEF, GBR |
| 53178 | Ruegeria arenilitoris strain G-M8 | 19497 | 27 | 5 | KX418522 | −18.846 | 147.63 | DAVIES REEF, GBR |
| 53179 | Ruegeria arenilitoris strain G-M8 | 19497 | 27 | 5 | KX418523 | −18.846 | 147.63 | DAVIES REEF, GBR |
| 53651 | Micrococcus aloeverae strain AE-6 | 21250 | 26 | 19 | KX418483 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53654 | Pseudomonas aeruginosa PAO1 strain PAO1 | 21250 | 26 | 19 | KX418585 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53656 | Micrococcus aloeverae strain AE-6 | 21250 | 26 | 19 | KX418588 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53657 | Pseudomonas azotoformans strain NBRC 12693 | 21250 | 26 | 19 | KX418490 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 58101 | Ruegeria arenilitoris strain G-M8 | 21556 | 26 | 21 | KX418524 | −18.838 | 147.642 | TRUNK REEF, GBR |
| 58102 | Ruegeria arenilitoris strain G-M8 | 21556 | 26 | 21 | KX418555 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58103 | Ruegeria arenilitoris strain G-M8 | 21556 | 26 | 21 | KX418508 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58104 | Pseudomonas azotoformans strain NBRC 12693 | 21556 | 26 | 21 | KX418491 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58105 | Ruegeria arenilitoris strain G-M8 | 21556 | 26 | 21 | KX418509 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58107 | Pseudomonas azotoformans strain NBRC 12693 | 21556 | 26 | 21 | KX418492 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58111 | Pseudomonas azotoformans strain NBRC 12693 | 21556 | 26 | 21 | KX418493 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58115 | Cobetia amphilecti strain 46-2 | 21556 | 26 | 21 | KX418494 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58116 | Pseudomonas azotoformans strain NBRC 12693 | 21556 | 26 | 21 | KX418495 | −18.332 | 146.829 | TRUNK REEF, GBR |
| 58264 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21717 | 26 | 5 | KX418510 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58265 | Pseudomonas azotoformans strain NBRC 12693 | 21717 | 26 | 5 | KX418465 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58266 | Pseudomonas azotoformans strain NBRC 12693 | 21717 | 26 | 5 | KX418496 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58271 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21717 | 26 | 5 | KX418511 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58273 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21717 | 26 | 10 | KX418565 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58274 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21717 | 26 | 10 | KX418512 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58275 | Bacillus aquimaris strain TF-12 | 21717 | 26 | 5 | KX418579 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58276 | Ruegeria arenilitoris strain G-M8 | 21717 | 26 | 5 | KX418513 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58277 | Ruegeria arenilitoris strain G-M8 | 21717 | 26 | 5 | KX418514 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58278 | Ruegeria arenilitoris strain G-M8 | 21717 | 26 | 5 | KX418557 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58279 | Pseudomonas azotoformans strain NBRC 12693 | 21717 | 26 | 5 | KX418497 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58280 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418518 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58281 | Pseudomonas azotoformans strain NBRC 12693 | 21722 | 26 | 5 | KX418466 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58282 | Pseudomonas azotoformans strain NBRC 12693 | 21722 | 26 | 5 | KX418467 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58283 | Bacillus cereus ATCC 14579 | 21722 | 26 | 5 | KX418567 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58285 | Pseudomonas azotoformans strain NBRC 12693 | 21722 | 26 | 5 | KX418468 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58286 | Pseudomonas azotoformans strain NBRC 12693 | 21722 | 26 | 5 | KX418469 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58287 | Bacillus aquimaris strain TF-12 | 21722 | 26 | 5 | KX418568 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58289 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418525 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58290 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418526 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58291 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418566 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58292 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418519 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58293 | Vibrio owensii strain DY05 | 21717 | 26 | 5 | KX418505 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58294 | Marinobacter vinifirmus strain FB1 | 21722 | 26 | 5 | KX418470 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58295 | Vibrio chagasii strain LMG 21353 | 21722 | 26 | 5 | KX418498 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58296 | Marinobacter vinifirmus strain FB1 | 21722 | 26 | 5 | KX418471 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58297 | Shewanella corallii strain fav-2-10-05 | 21722 | 26 | 5 | KX418503 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58298 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418560 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58299 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418527 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58300 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21722 | 26 | 5 | KX418528 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 53193 | Bacillus horikoshii strain DSM 8719 | 19496 | 27 | 4 | KX418578 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 53208 | Kytococcus sedentarius strain DSM 20547 | 19496 | 27 | 4 | KX418583 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 53209 | Kocuria rhizophilia strain TA68 | 19496 | 27 | 4 | KX418587 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 53211 | Pseudoalteromonas piscicida strain NBRC 103038 | 19496 | 27 | 4 | KX418484 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 58301 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21722 | 26 | 5 | KX418529 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58302 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21722 | 26 | 5 | KX418530 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58303 | Ruegeria atlantica strain NBRC 15792 | 21722 | 26 | 5 | KX418531 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58304 | Bacillus kochii strain WCC 4582 | 21722 | 26 | 5 | KX418569 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58305 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418532 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58306 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21722 | 26 | 5 | KX418533 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58307 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418564 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58308 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418534 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58310 | Vibrio chagasii strain LMG 21353 | 21722 | 26 | 5 | KX418507 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58312 | Bacillus kochii strain WCC 4582 | 21722 | 26 | 5 | KX418574 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58313 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21722 | 26 | 5 | KX418535 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58315 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21722 | 26 | 5 | KX418536 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58316 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418537 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58317 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21727 | 26 | 5 | KX418558 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58318 | Ruegeria arenilitoris strain G-M8 | 21727 | 26 | 5 | KX418538 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58319 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21727 | 26 | 5 | KX418539 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58320 | Microbulbifer variabilis strain Ni-2088 | 21727 | 26 | 5 | KX418501 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58321 | Staphylococcus warneri SG1 strain SG1 | 21727 | 26 | 5 | KX418580 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58322 | Thalassospira permensis strain SMB34 | 21727 | 26 | 5 | KX418554 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58323 | Ruegeria arenilitoris strain G-M8 | 21727 | 26 | 5 | KX418556 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58324 | Pseudomonas pachastrellae strain KMM 330 | 21727 | 26 | 5 | KX418499 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58325 | Pseudoalteromonas shioyasakiensis strain SE3 | 21727 | 26 | 5 | KX418500 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58326 | Bacillus nealsonii strain DSM 15077 | 21727 | 26 | 5 | KX418581 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58327 | Ruegeria arenilitoris strain G-M8 | 21727 | 26 | 5 | KX418540 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58328 | Ruegeria arenilitoris strain G-M8 | 21727 | 26 | 5 | KX418541 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58330 | Aquimarina spongiae strain A6 | 21727 | 26 | 5 | KX418462 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58332 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21727 | 26 | 5 | KX418520 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58333 | Ruegeria atlantica strain NBRC 15792 | 21727 | 26 | 5 | KX418542 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58334 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21727 | 26 | 5 | KX418543 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 58335 | Ruegeria arenilitoris strain G-M8 | 21727 | 26 | 5 | KX418544 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 53244 | Vibrio campbellii strain ATCC 25920 | 19496 | 30 | 2 | KX418485 | −18.82 | 147.63 | DAVIES REEF, GBR |
| 53611 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21241 | 26 | 18 | KX418545 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53613 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21241 | 26 | 18 | KX418546 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53614 | Bacillus oceanisediminis strain H2 | 21241 | 26 | 18 | KX418576 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53616 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21241 | 26 | 18 | KX418547 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53620 | Ruegeria arenilitoris strain G-M8 | 21241 | 26 | 18 | KX418548 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53622 | Pseudovibrio sp. FO-BEG1 strain FO-BEG1 | 21241 | 26 | 18 | KX418549 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53623 | Vibrio fortis strain CAIM 629 | 21241 | 26 | 18 | KX418504 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53624 | Stenotrophomonas maltophilia R551-3 strain R551-3 | 21241 | 26 | 18 | KX418472 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53625 | Ruegeria arenilitoris strain G-M8 | 21241 | 26 | 18 | KX418561 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53634 | Sanguibacter inulinus strain ST50 | 21241 | 26 | 19 | KX418584 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53635 | Pseudomonas geniculate strain ATCC 19374 | 21241 | 26 | 19 | KX418506 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53636 | Pseudoalteromonas piscicida strain NBRC 103038 | 21250 | 26 | 19 | KX418486 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53640 | Pseudoalteromonas piscicida strain NBRC 103038 | 21250 | 26 | 19 | KX418487 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53641 | Ruegeria arenilitoris strain G-M8 | 21250 | 26 | 19 | KX418550 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53650 | Micrococcus yunnanensis strain YIM 65004 | 21250 | 26 | 19 | KX418586 | −18.838 | 147.642 | DAVIES REEF, GBR |
| 53240 | Bacillus safensis strain NBRC 100820 | 19496 | 30 | 2 | KX418571 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 53237 | Psychrobacter celer strain SW-238 | 19496 | 30 | 2 | KX418473 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 53239 | Psychrobacter celer strain SW-238 | 19496 | 30 | 2 | KX418474 | −18.820 | 147.630 | DAVIES REEF, GBR |
| 59159 | Bacillus kochii strain WCC 4582 | 22821 | 26 | 14 | KX418575 | −18.732 | 147.519 | CENTIPEDE REEF : BACK BOMMIE, GBR |
| 59161 | Vibrio alginolyticus strain ATCC 17749 | 22821 | 26 | 14 | KX418488 | −18.732 | 147.519 | CENTIPEDE REEF : BACK BOMMIE, GBR |
| 58824 | Photobacterium rosenbergii strain CC1 | 22821 | 28 | 15 | KX418489 | −18.732 | 147.519 | CENTIPEDE REEF : BACK BOMMIE, GBR |
| 58284 | Ruegeria arenilitoris strain G-M8 | 21722 | 26 | 5 | KX418551 | −18.833 | 147.626 | DAVIES REEF, GBR |
| 53260 | Brevundimonas diminuta strain NBRC 12697 | 19496 | 30 | 5 | KX418521 | −18.820 | 147.630 | DAVIES REEF, GBR |
References
- Hughes, C.C.; Fenical, W. Antibacterials from the sea. Chem. Eur. J. 2010, 16, 12512–12525. [Google Scholar] [CrossRef] [PubMed]
- Fenical, W.; Jensen, P.R. Developing a new resource for drug discovery: Marine actinomycete bacteria. Nat. Chem. Biol. 2006, 2, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Datta, D.; Talapatra, S.; Swarnakar, S. Bioactive compounds from marine invertebrates for potential medicines-an overview. Int. Lett. Nat. Sci. 2015, 7, 42–61. [Google Scholar] [CrossRef]
- Mehbub, M.F.; Lei, J.; Franco, C.; Zhang, W. Marine sponge derived natural products between 2001 and 2010: Trends and opportunities for discovery of bioactives. Mar. Drugs 2014, 12, 4539–4577. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.W.; Radax, R.; Steger, D.; Wagner, M. Sponge-associated microorganisms: Evolution, ecology, and biotechnological potential. Microbiol. Mol. Biol. Rev. 2007, 71, 295–347. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, P.; Balasubramaniam, M.; Dam Roy, S.; Sarma, K.; Hairun, R.; Sunder, J. Characterization of the antibacterial activity of bacteria associated with stylissa sp, a marine sponge. Adv. Anim. Vet. Sci. 2014, 2, 20–25. [Google Scholar] [CrossRef]
- Burja, A.M.; Hill, R.T. Microbial symbionts of the Australian Great Barrier Reef sponge, candidaspongia flabellata. Hydrology 2001, 461, 41–47. [Google Scholar]
- Graça, A.P.; Viana, F.; Bondoso, J.; Correia, M.I.; Gomes, L.; Humanes, M.; Reis, A.; Xavier, J.R.; Gaspar, H.; Lage, O.M. The antimicrobial activity of heterotrophic bacteria isolated from the marine sponge erylus deficiens (astrophorida, geodiidae). Front. Microbiol. 2015, 6, 389. [Google Scholar] [PubMed]
- Sagar, S.; Kaur, M.; Minneman, K.P. Antiviral lead compounds from marine sponges. Mar. Drugs 2010, 8, 2619–2638. [Google Scholar] [CrossRef] [PubMed]
- Atikana, A.; Naim, M.A.; Sipkema, D. Detection of keto synthase (ks) gene domain in sponges and bacterial sponges. Ann. Bogor. 2013, 17, 27–33. [Google Scholar]
- Aguila-Ramírez, R.N.; Hernández-Guerrero, C.J.; González-Acosta, B.; Id-Daoud, G.; Hewitt, S.; Pope, J.; Hellio, C. Antifouling activity of symbiotic bacteria from sponge aplysina gerardogreeni. Int. Biodeter. Biodegr. 2014, 90, 64–70. [Google Scholar] [CrossRef]
- Thomas, T.; Moitinho-Silva, L.; Lurgi, M.; Bjork, J.R.; Easson, C.; Astudillo-Garcia, C.; Olson, J.B.; Erwin, P.M.; Lopez-Legentil, S.; Luter, H.; et al. Diversity, structure and convergent evolution of the global sponge microbiome. Nat. Commun. 2016, 7, 11870. [Google Scholar] [CrossRef] [PubMed]
- Hentschel, U.; Usher, K.M.; Taylor, M.W. Marine sponges as microbial fermenters. FEMS Microbiol. Ecol. 2006, 55, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Bell, J.J. The functional roles of marine sponges. Estuar. Coast. Shelf Sci. 2008, 79, 341–353. [Google Scholar] [CrossRef]
- Van Soest, R.W.M.; Boury-Esnault, N.; Vacelet, J.; Dohrmann, M.; Erpenbeck, D.; de Voogd, N.J.; Santodomingo, N.; Vanhoorne, B.; Kelly, M.; Hooper, J.N.A. Global diversity of sponges (porifera). PLoS ONE 2012, 7, e35105. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, W.; Feeney, R.J. Contributions to the study of marine products. XXXII. The nucleosides of sponges. I. J. Org. Chem. 1951, 16, 981–987. [Google Scholar] [CrossRef]
- Kennedy, J.; Baker, P.; Piper, C.; Cotter, P.; Walsh, M.; Mooij, M.; Bourke, M.; Rea, M.; O’Connor, P.; Ross, R.P.; et al. Isolation and analysis of bacteria with antimicrobial activities from the marine sponge haliclona simulans collected from irish waters. Mar. Biotechnol. 2009, 11, 384–396. [Google Scholar] [CrossRef] [PubMed]
- Anand, T.P.; Bhat, A.W.; Shouche, Y.S.; Roy, U.; Siddharth, J.; Sarma, S.P. Antimicrobial activity of marine bacteria associated with sponges from the waters off the coast of south east india. Microbiol. Res. 2006, 161, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Webster, N.S.; Luter, H.M.; Soo, R.M.; Botte, E.S.; Simister, R.L.; Abdo, D.; Whalan, S. Same, same but different: Symbiotic bacterial associations in gbr sponges. Front. Microbiol. 2012, 3, 444. [Google Scholar] [CrossRef] [PubMed]
- Santos, O.C.S.; Pontes, P.V.M.L.; Santos, J.F.M.; Muricy, G.; Giambiagi-deMarval, M.; Laport, M.S. Isolation, characterization and phylogeny of sponge-associated bacteria with antimicrobial activities from brazil. Res. Microbiol. 2010, 161, 604–612. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.R.A.; Kavlekar, D.P.; LokaBharathi, P.A. Marine drugs from sponge-microbe association—A review. Mar. Drugs 2010, 8, 1417–1468. [Google Scholar] [CrossRef] [PubMed]
- Amoutzias, D.G.; Chaliotis, A.; Mossialos, D. Discovery strategies of bioactive compounds synthesized by nonribosomal peptide synthetases and type-i polyketide synthases derived from marine microbiomes. Mar. Drugs 2016, 14, 80. [Google Scholar] [CrossRef] [PubMed]
- Borchert, E.; Jackson, S.A.; O’Gara, F.; Dobson, A.D.W. Diversity of natural product biosynthetic genes in the microbiome of the deep sea sponges inflatella pellicula, poecillastra compressa, and stelletta normani. Front. Microbiol. 2016, 7, 1027. [Google Scholar] [CrossRef] [PubMed]
- Santos, O.C.S.; Soares, A.R.; Machado, F.L.S.; Romanos, M.T.V.; Muricy, G.; Giambiagi-deMarval, M.; Laport, M.S. Investigation of biotechnological potential of sponge-associated bacteria collected in brazilian coast. Lett. Appl. Microbiol. 2015, 60, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Williams, G.J. Engineering polyketide synthases and nonribosomal peptide synthetases. Curr. Opin. Struct. Biol. 2013, 23, 603–612. [Google Scholar] [CrossRef] [PubMed]
- Hochmuth, T.; Piel, J. Polyketide synthases of bacterial symbionts in sponges—Evolution-based applications in natural products research. Phytochemistry 2009, 70, 1841–1849. [Google Scholar] [CrossRef] [PubMed]
- Müller, W.E.G.; Grebenjuk, V.A.; Le Pennec, G.; Schröder, H.-C.; Brümmer, F.; Hentschel, U.; Müller, I.M.; Breter, H.-J. Sustainable production of bioactive compounds by sponges—cell culture and gene cluster approach: A review. Mar. Biotechnol. 2004, 6, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Carmely, S.; Kashman, Y. Structure of swinholide-a, a new macrolide from the marine sponge theonella swinhoei. Tetrahedron. Lett. 1985, 26, 511–514. [Google Scholar] [CrossRef]
- Indraningrat, A.A.G.; Smidt, H.; Sipkema, D. Bioprospecting sponge-associated microbes for antimicrobial compounds. Mar. Drugs 2016, 14, 87. [Google Scholar] [CrossRef] [PubMed]
- Schneemann, I.; Kajahn, I.; Ohlendorf, B.; Zinecker, H.; Erhard, A.; Nagel, K.; Wiese, J.; Imhoff, J.F. Mayamycin, a cytotoxic polyketide from astreptomycesstrain isolated from the marine spongehalichondria panicea. J. Nat. Prod. 2010, 73, 1309–1312. [Google Scholar] [CrossRef] [PubMed]
- Hirata, Y.; Uemura, D. Halichondrins-antitumor polyether macrolides from a marine sponge. Pure Appl. Chem. 1986, 58, 701–710. [Google Scholar] [CrossRef]
- Bai, R.L.; Paull, K.D.; Herald, C.L.; Malspeis, L.; Pettit, G.R.; Hamel, E. Halichondrin b and homohalichondrin b, marine natural products binding in the vinca domain of tubulin. Discovery of tubulin-based mechanism of action by analysis of differential cytotoxicity data. J. Biol. Chem. 1991, 266, 15882–15889. [Google Scholar] [PubMed]
- Webster, N.S.; Wilson, K.J.; Blackall, L.L.; Hill, R.T. Phylogenetic diversity of bacteria associated with the marine sponge rhopaloeides odorabile. Appl. Environ. Microbiol. 2001, 67, 434–444. [Google Scholar] [CrossRef] [PubMed]
- Graça, A.P.; Bondoso, J.; Gaspar, H.; Xavier, J.R.; Monteiro, M.C.; de la Cruz, M.; Oves-Costales, D.; Vicente, F.; Lage, O.M. Antimicrobial activity of heterotrophic bacterial communities from the marine sponge erylus discophorus (astrophorida, geodiidae). PLoS ONE 2013, 8, e78992. [Google Scholar] [CrossRef] [PubMed]
- Webster, N.S.; Hill, R.T. The culturable microbial community of the Great Barrier Reef sponge rhopaloeides odorabile is dominated by an α-proteobacterium. Mar. Biol. 2001, 138, 843–851. [Google Scholar] [CrossRef]
- Wheeler, D.; Bhagwat, M. Blast quickstart: Example-driven web-based blast tutorial. In Comparative Genomics: Volumes 1 and 2; Bergman, N., Ed.; Humana Press: Totowa, NJ, USA, 2008. [Google Scholar]
- Ludwig, W.; Strunk, O.; Westram, R.; Richter, L.; Meier, H.; Yadhukumar; Buchner, A.; Lai, T.; Steppi, S.; Jobb, G.; et al. ARB: A software environment for sequence data. Nucleic Acids Res. 2004, 32, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Elfalah, H.W.A.; Ahmad, A.; Usup, G. Anti-microbial properties of secondary metabolites of marine gordonia tearrae extract. J. Agric. Sci. 2013, 5, 94. [Google Scholar] [CrossRef]
- Devi, P.; Wahidullah, S.; Rodrigues, C.; Souza, L.D. The sponge-associated bacterium bacillus licheniformis sab1: A source of antimicrobial compounds. Mar. Drugs 2010, 8, 1203–1212. [Google Scholar] [CrossRef] [PubMed]
- Kavitha, A.; Prabhakar, P.; Vijayalakshmi, M.; Venkateswarlu, Y. Purification and biological evaluation of the metabolites produced by Streptomyces sp. Tk-vl_333. Res. Microbiol. 2010, 161, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Gohar, Y.M.; El-Naggar, M.M.; Soliman, M.K.; Barakat, K.M. Characterization of marine burkholderia cepacia antibacterial agents. J. Nat. Prod. 2010, 3, 86–94. [Google Scholar]
- Dashti, Y.; Grkovic, T.; Abdelmohsen, U.R.; Hentschel, U.; Quinn, R.J. Production of induced secondary metabolites by a co-culture of sponge-associated actinomycetes, Actinokineospora sp. Eg49 and Nocardiopsis sp. Rv163. Mar. Drugs 2014, 12, 3046–3059. [Google Scholar] [CrossRef] [PubMed]
- McFarland, J. The nephelometer: An instrument for estimating the number of bacteria in suspensions used for calculating the opsonic index and for vaccines. J. Am. Med. Assoc. 1907, 49, 1176–1178. [Google Scholar] [CrossRef]
- Bauer, A.W.; Kirby, W.M.; Sherris, J.C.; Turck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [PubMed]
- Palomo, S.; González, I.; de la Cruz, M.; Martín, J.; Tormo, J.R.; Anderson, M.; Hill, R.T.; Vicente, F.; Reyes, F.; Genilloud, O. Sponge-derived kocuria and micrococcus spp. As sources of the new thiazolyl peptide antibiotic kocurin. Mar. Drugs 2013, 11, 1071–1086. [Google Scholar] [CrossRef] [PubMed]
- Neilan, B.A.; Dittmann, E.; Rouhiainen, L.; Bass, R.A.; Schaub, V.; Sivonen, K.; Borner, T. Nonribosomal peptide synthesis and toxigenicity of cyanobacteria. J. Bacteriol. 1999, 181, 4089–4097. [Google Scholar] [PubMed]
- Kim, T.K.; Garson, M.J.; Fuerst, J.A. Marine actinomycetes related to the ‘salinospora’ group from the Great Barrier Reef sponge pseudoceratina clavata. Environ. Microbiol. 2005, 7, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Abdelmohsen, U.R.; Yang, C.; Horn, H.; Hajjar, D.; Ravasi, T.; Hentschel, U. Actinomycetes from red sea sponges: Sources for chemical and phylogenetic diversity. Mar. Drugs 2014, 12, 2771–2789. [Google Scholar] [CrossRef] [PubMed]
- Ayuso-Sacido, A.; Genilloud, O. New pcr primers for the screening of nrps and pks-i systems in actinomycetes: Detection and distribution of these biosynthetic gene sequences in major taxonomic groups. Microb. Ecol. 2005, 49, 10–24. [Google Scholar] [CrossRef] [PubMed]
- Yoon, B.J.; You, H.S.; Lee, D.H.; Oh, D.C. Aquimarina spongiae sp. Nov., isolated from marine sponge halichondria oshoro. Int. J. Syst. Evol. Microbiol. 2011, 61, 417–421. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. Mega6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Bondarev, V.; Richter, M.; Romano, S.; Piel, J.; Schwedt, A.; Schulz-Vogt, H.N. The genus pseudovibrio contains metabolically versatile bacteria adapted for symbiosis. Environ. Microbiol. 2013, 15, 2095–2113. [Google Scholar] [CrossRef] [PubMed]
- Pospisil, S.; Benada, O.; Kofronova, O.; Petricek, M.; Janda, L.; Havlicek, V. Kytococcus sedentarius (formerly micrococcus sedentarius) and dermacoccus nishinomiyaensis (formerly micrococcus nishinomiyaensis) produce monensins, typical streptomyces cinnamonensis metabolites. Can. J. Microbiol. 1998, 44, 1007–1011. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Yan, X.; Han, X.; Chen, H.; Lin, W.; Lee, F.S.; Wang, X. Identification of norharman as the cytotoxic compound produced by the sponge (Hymeniacidon perleve)-associated marine bacterium pseudoalteromonas piscicida and its apoptotic effect on cancer cells. Biotechnol. Appl. Biochem. 2006, 44, 135–142. [Google Scholar] [PubMed]
- Ventola, C.L. The antibiotic resistance crisis: Part 1: Causes and threats. Pharm. Ther. 2015, 40, 277. [Google Scholar]
- Cita, Y.P.; Suhermanto, A.; Radjasa, O.K.; Sudharmono, P. Antibacterial activity of marine bacteria isolated from sponge xestospongia testudinaria from sorong, papua. Asian. Pac. J. Trop. Med. 2017, 7, 450–454. [Google Scholar] [CrossRef]
- Zobell, C.E. Studies on marine bacteria. I. The cultural requirements of heterotrophic aerobes. J. Mar. Sci. 1941, 4, 42–75. [Google Scholar]
- Wei, H.; Lin, Z.; Li, D.; Gu, Q.; Zhu, T. OSMAC (one strain many compounds) approach in the research of microbial metabolites--a review. Acta Microbiol. Sin. 2010, 50, 701–709. [Google Scholar]
- Marmann, A.; Aly, A.H.; Lin, W.; Wang, B.; Proksch, P. Co-cultivation—A powerful emerging tool for enhancing the chemical diversity of microorganisms. Mar. Drugs 2014, 12, 1043–1065. [Google Scholar] [CrossRef] [PubMed]
- Hentschel, U.; Schmid, M.; Wagner, M.; Fieseler, L.; Gernert, C.; Hacker, J. Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the mediterranean sponges aplysina aerophoba and aplysina cavernicola. FEMS Microbiol. Ecol. 2001, 35, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.S.; Krishna, E.R.; Sujatha, P.; Kumar, B.V. Screening and isolation of associated bioactive microorganisms from fasciospongia cavernosa from of visakhapatnam coast, bay of bengal. J. Chem. 2012, 9, 2166–2176. [Google Scholar]
- Crowley, S.P.; O’Gara, F.; O’Sullivan, O.; Cotter, P.D.; Dobson, A.D.W. Marine pseudovibrio sp. As a novel source of antimicrobials. Mar. Drugs 2014, 12, 5916–5929. [Google Scholar] [CrossRef] [PubMed]
- Michel-Briand, Y.; Baysse, C. The pyocins of pseudomonas aeruginosa. Biochimie 2002, 84, 499–510. [Google Scholar] [CrossRef]
- Zheng, L.; Chen, H.; Han, X.; Lin, W.; Yan, X. Antimicrobial screening and active compound isolation from marine bacterium nj6-3-1 associated with the sponge hymeniacidon perleve. World J. Microbiol. Biotechnol. 2005, 21, 201–206. [Google Scholar] [CrossRef]
- Offret, C.; Desriac, F.; Le Chevalier, P.; Mounier, J.; Jégou, C.; Fleury, Y. Spotlight on antimicrobial metabolites from the marine bacteria pseudoalteromonas: Chemodiversity and ecological significance. Mar. Drugs 2016, 14, 129. [Google Scholar] [CrossRef] [PubMed]
- Satheesh, S.; Ba-akdah, M.A.; Al-Sofyani, A.A. Natural antifouling compound production by microbes associated with marine macroorganisms—A review. Electron. J. Biotechn. 2016, 21, 26–35. [Google Scholar] [CrossRef]
- Skariyachan, S.; G Rao, A.; Patil, M.R.; Saikia, B.; Bharadwaj Kn, V.; Rao Gs, J. Antimicrobial potential of metabolites extracted from bacterial symbionts associated with marine sponges in coastal area of gulf of mannar biosphere, India. Lett. Appl. Microbiol. 2014, 58, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Flemer, B.; Kennedy, J.; Margassery, L.M.; Morrissey, J.P.; O’Gara, F.; Dobson, A.D.W. Diversity and antimicrobial activities of microbes from two irish marine sponges, Suberites carnosus and Leucosolenia sp. Lett. Appl. Microbiol. 2012, 112, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Cuvelier, M.L.; Blake, E.; Mulheron, R.; McCarthy, P.J.; Blackwelder, P.; Thurber, R.L.V.; Lopez, J.V. Two distinct microbial communities revealed in the sponge cinachyrella. Front. Microbiol. 2014, 5, 581. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, C.C.P.; Costa, R.; Araujo, F.V.; Hajdu, E.; Peixoto, R.; Lins, U.; Rosado, A.S.; Van Elsas, J.D. Diversity of bacteria in the marine sponge aplysina fulva in brazilian coastal waters. Appl. Environ. Microbiol. 2009, 75, 3331–3343. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; He, L.; Miao, X. Cultivable bacterial community from south china sea sponge as revealed by dgge fingerprinting and 16S rDNA phylogenetic analysis. Curr. Microbiol. 2007, 55, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Turque, A.S.; Cardoso, A.M.; Silveira, C.B.; Vieira, R.P.; Freitas, F.A.D.; Albano, R.M.; Gonzalez, A.M.; Paranhos, R.; Muricy, G.; Martins, O.B. Bacterial communities of the marine sponges hymeniacidon heliophila and polymastia janeirensis and their environment in rio de janeiro, brazil. Mar. Biol. 2008, 155, 135. [Google Scholar] [CrossRef]
- Kim, T.K.; Fuerst, J.A. Diversity of polyketide synthase genes from bacteria associated with the marine sponge pseudoceratina clavata: Culture-dependent and culture-independent approaches. Environ. Microbiol. 2006, 8, 1460–1470. [Google Scholar] [CrossRef] [PubMed]
- Boddy, C.N. Bioinformatics tools for genome mining of polyketide and non-ribosomal peptides. J. Ind. Microbiol. Biot. 2014, 41, 443–450. [Google Scholar] [CrossRef] [PubMed]
- Reen, F.J.; Romano, S.; Dobson, A.D.W.; O’Gara, F. The sound of silence: Activating silent biosynthetic gene clusters in marine microorganisms. Mar. Drugs 2015, 13, 4754–4783. [Google Scholar] [CrossRef] [PubMed]
- Ochi, K.; Hosaka, T. New strategies for drug discovery: Activation of silent or weakly expressed microbial gene clusters. Appl. Microbiol. Biotechnol. 2013, 97, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Giddings, L.-A.; Newman, D.J. Bioactive Compounds from Extremophiles: Genomic Studies, Biosynthetic Gene Clusters, and New Dereplication Methods; Briefs in Microbiology; Springer: Berlin/Heidelberg, Germany, 2015; pp. 1–124. [Google Scholar]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).