The Rhizosphere of Coffea Arabica in Its Native Highland Forests of Ethiopia Provides a Niche for a Distinguished Diversity of Trichoderma
Abstract
:1. Introduction
2. Results
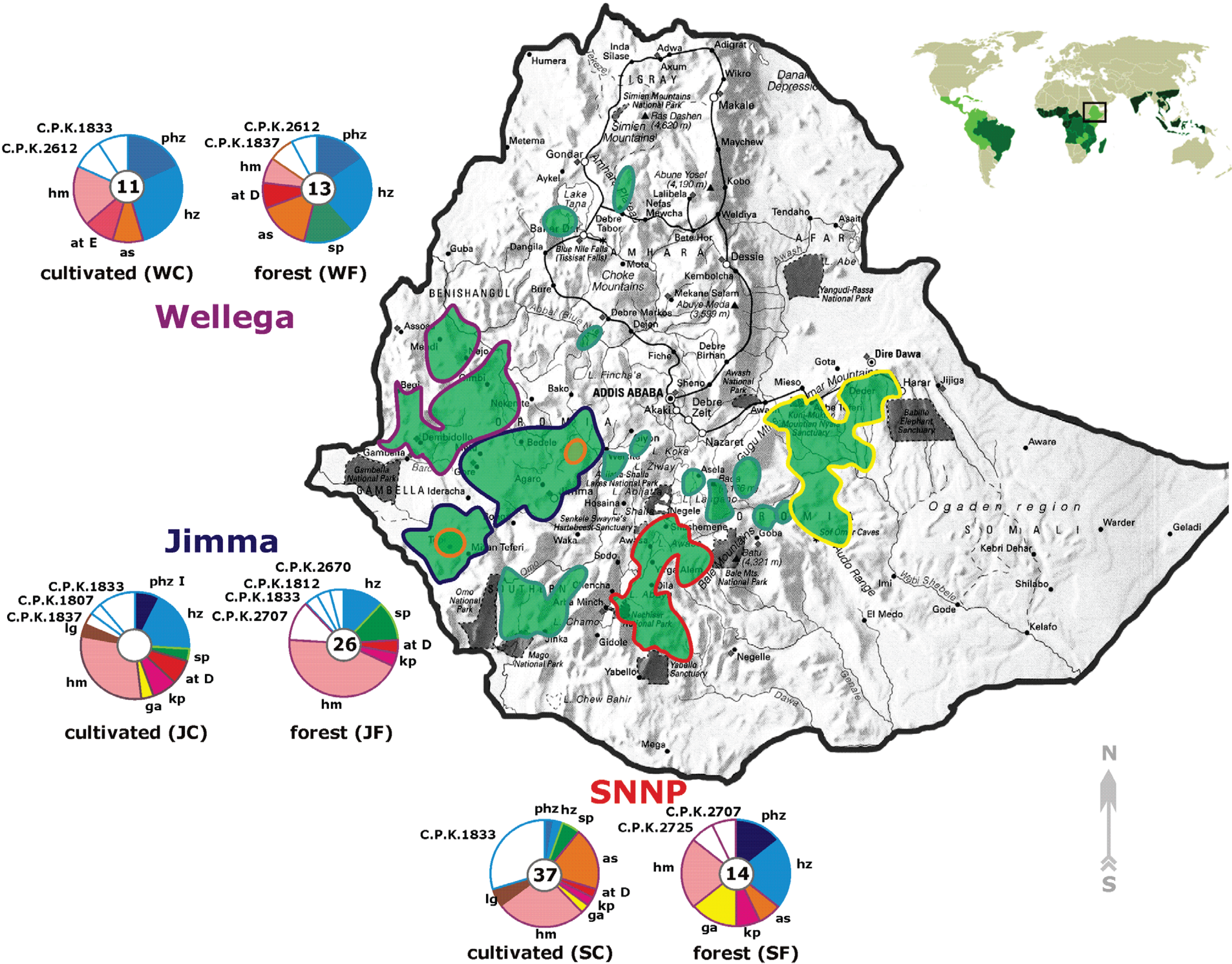
2.1. Habitats of C. arabica and the Sampling Strategy
| Region | Ecosystem | Code | Alt. range (m) | Soil properties | No. of samples | No. of Trichoderma isolates | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| Color | Texture | C% | N% | pH | ||||||
| Jimma | native forest | JF | 1320–2150 | Reddish brown | Sandy loam | 2.7 | 0.27 | 4.80 | 30 | 26 |
| semi-forest | JC | 1370–2400 | Reddish brown | Silt loam | 2.7 | 0.6 | 4.72 | 29 | 27 | |
| Wellega | native forest | WF | 1500–2300 | Reddish brown | Silt loam | 2.4 | 0.34 | 4.97 | 15 | 13 |
| semi-forest | WC | 1570–2400 | Reddish brown | Sandy loam | 1.6 | 0.24 | 4.67 | 53 | 11 | |
| SNNP | native forest | SF | 1650–2050 | Reddish brown | Sandy loam | 2.3 | 0.26 | 6.01 | 13 | 14 |
| semi-forest | SC | 1670–2080 | Reddish brown | Silt loam | 2.3 | 0.32 | 4.81 | 12 | 37 | |
| Hararghe | native forest | HF | 2150 | Reddish brown | Sandy loam | 2.5 | 0.23 | 6.53 | 2 | 2 |
| semi-forest | HC | 1580–2350 | Reddish brown | Sandy loam | 2 | 0.19 | 4.01 | 6 | 4 | |
2.2. Occurrence of Known Trichoderma Species
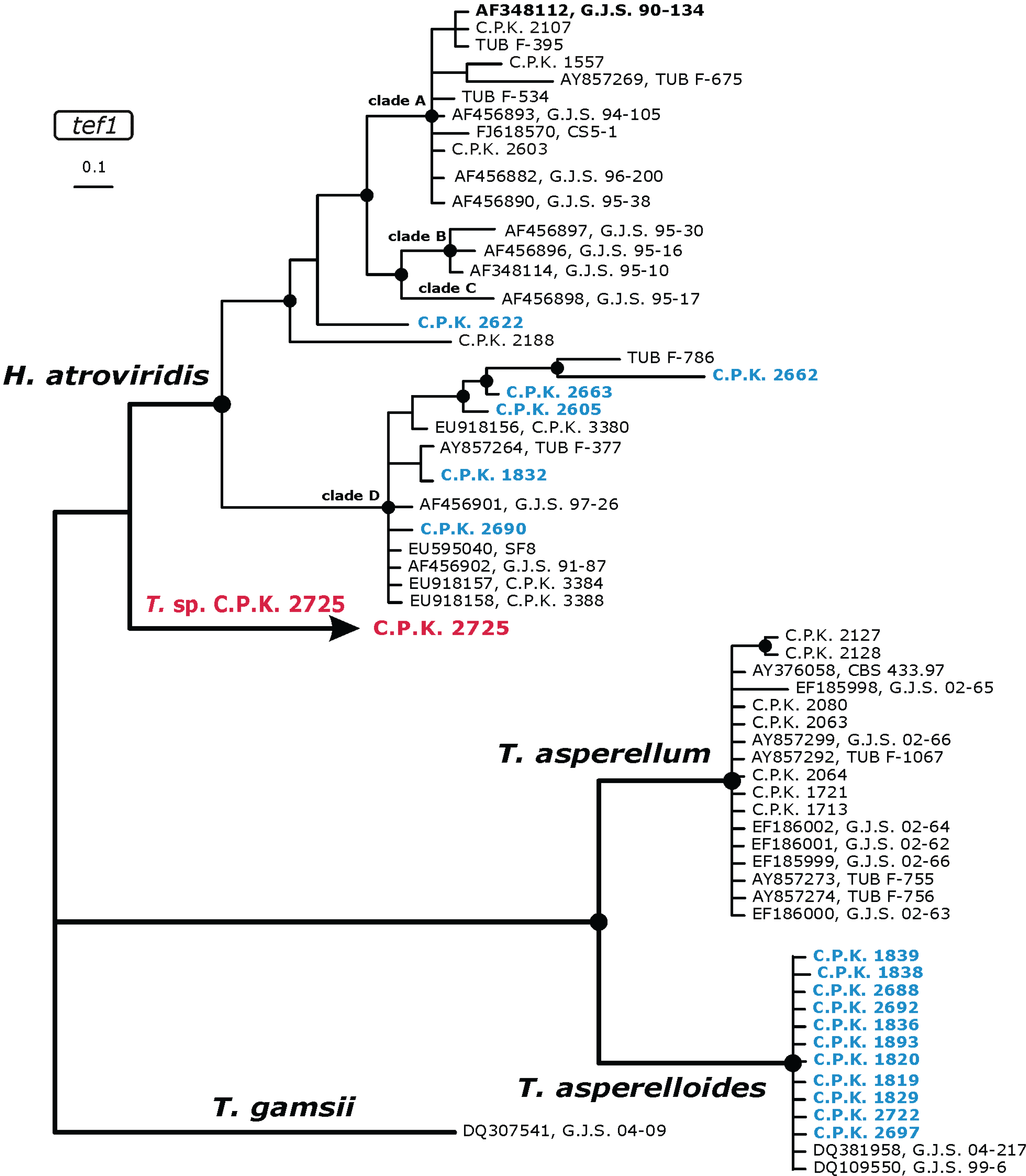
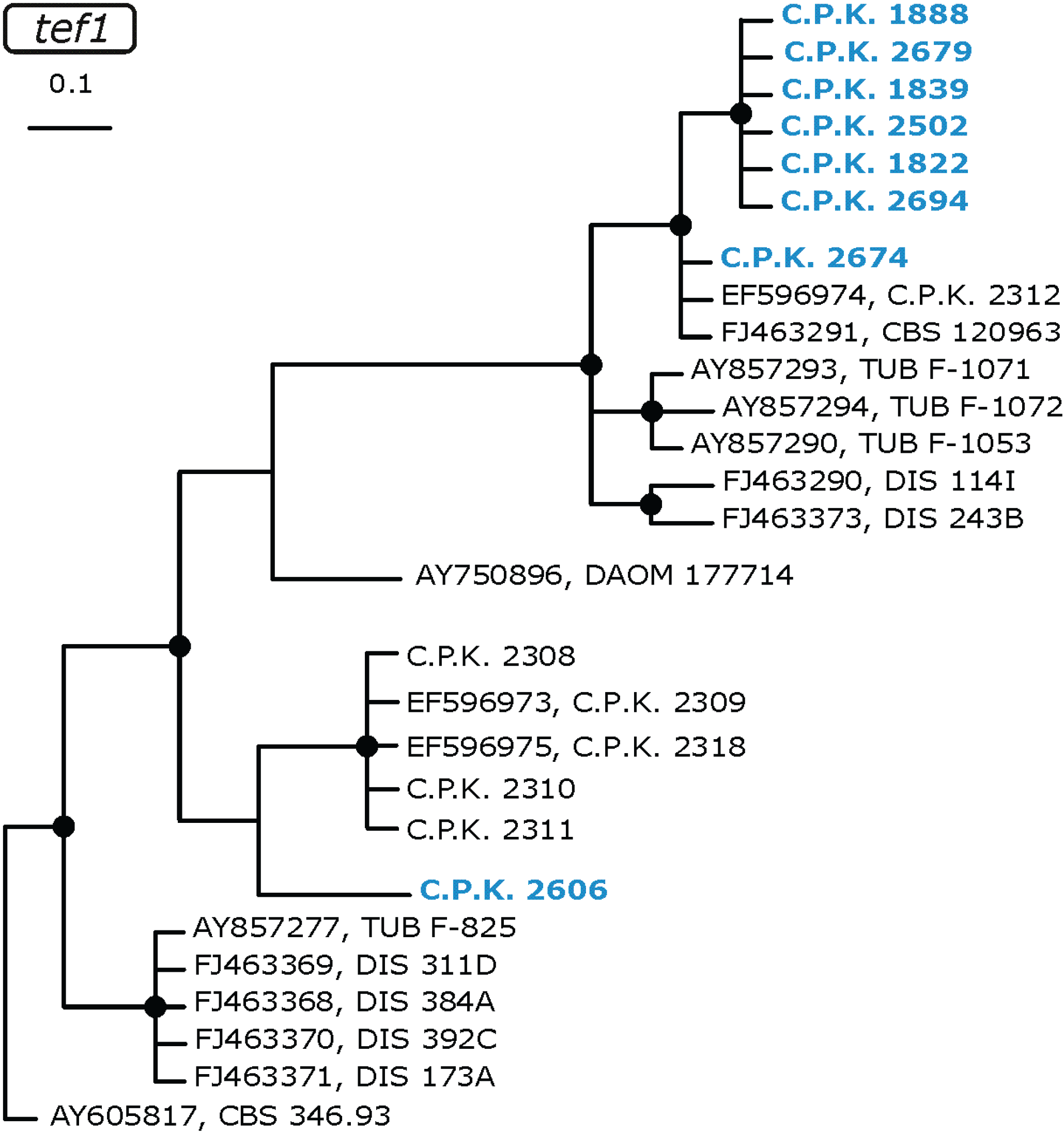
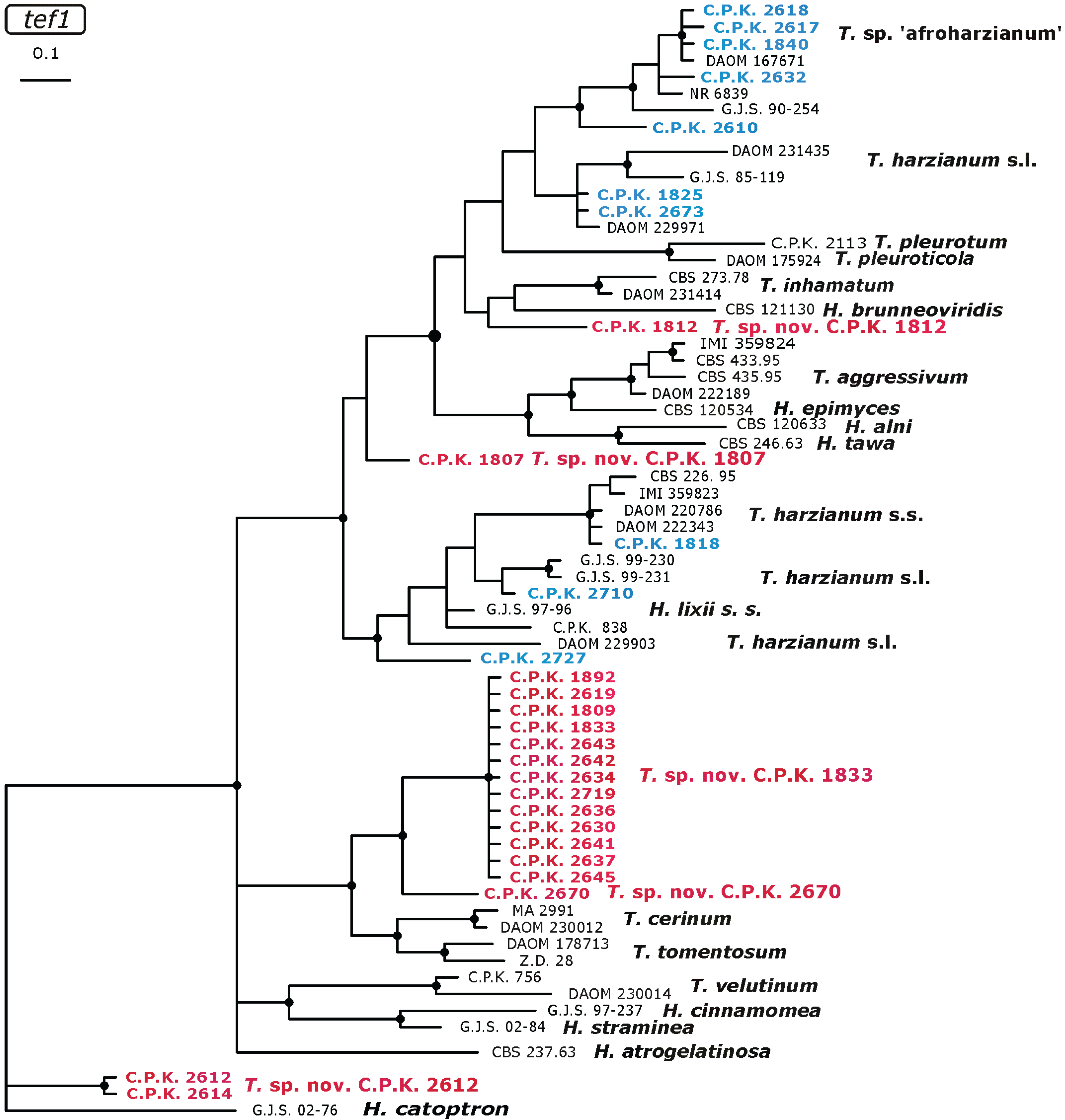
2.3. Occurrence of Potentially Endemic Trichoderma Taxa
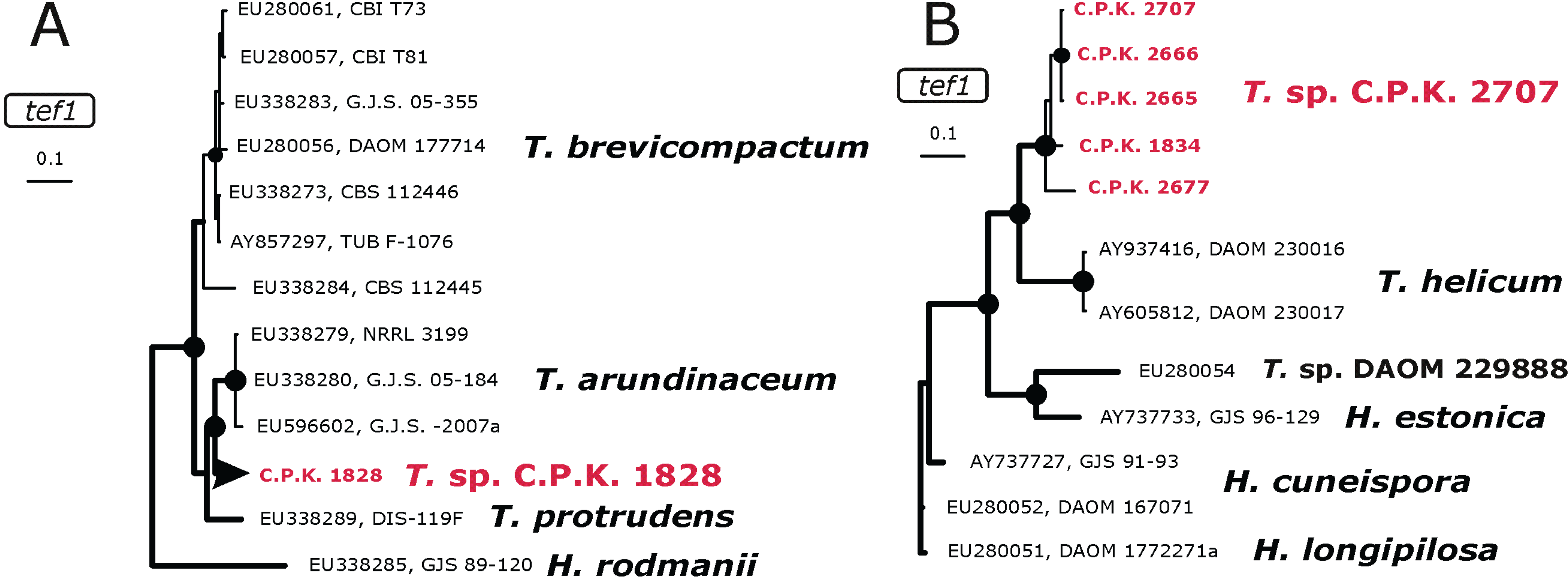
2.4. Habitat Preference of Trichoderma
| Index | Average | Ecosystem | Sampled regions | |||||
|---|---|---|---|---|---|---|---|---|
| Forest | Semi-forest | Wellega | Jimma | SNNP | Hararghe | |||
| Shannon diversity | H | 2.47 | 2.34 | 2.29 | 2.14 | 2.19 | 2.15 | 1.56 |
| Simpson | D | 0.88 | 0.87 | 0.87 | 0.9 | 0.84 | 0.86 | 0.93 |
2.5. Antagonistic Activity of Trichoderma Isolates against Gibberella xylarioides
3. Discussion
| Isolate No. | Species | PIRG1 | Clear Zone |
|---|---|---|---|
| C.P.K. 2612 | T. sp. C.P.K.2612 | 76a | - |
| C.P.K. 2614 | T. sp. C.P.K.2612 | 72b | - |
| C.P.K. 1808 | T. sp. C.P.K.1833 | 64c | - |
| C.P.K. 2619 | T. sp. C.P.K.1833 | 62c | - |
| C.P.K. 1817 | T. sp. C.P.K.1837 | 62c | 1.5 |
| C.P.K. 2698 | T. hamatum | 60c | - |
| C.P.K. 2622 | H. atroviride | 56d | 2 |
| C.P.K. 1819 | T. asperelloides | 56d | - |
| C.P.K. 1888 | T. spirale | 55d | - |
| C.P.K. 1834 | T .sp. C.P.K.2707 | 55d | 1.5 |
| F. xylarioides (Control) | 0 | - | |
4. Experimental Section
4.1. Soil Sampling and Characterization
4.2. Strain Isolation and Purification
4.3. DNA Extraction, PCR Amplification and Sequencing
4.4. Identification of Trichoderma
4.5. Molecular Phylogenetic Analysis
4.6. Antagonism of Trichoderma Isolates against Gibberella xylarioides
4.7. Statistical Analysis
5. Conclusions
Acknowledgements
References
- Coffee. 2009. Available online: http://www.coffeeresearch.org/ (accessed on 5 May 2009).
- Davids, K. Alternative Africas: Rwanda, Tanzania, Zimbabwe. Available online: http://www.coffeereview.com/ (accessed on 5 May 2009).
- Fair Trade Farmers in Ethiopia. 2004. Available online: http://www.globalexchange.org/campaigns/fairtrade/coffee/EthiopiaFlyer.pdf (accessed on 5 May 2009).
- Wellman, F.L. Coffee: Botany, Cultivation and Utilization. World Crop Series; Inter Science: New York, NY, USA, 1961. [Google Scholar]
- Meyer, F. Notes on wild Coffea arabica from southwestern Ethiopia, with some historical considerations. Econ. Bot. 1965, 19, 136–151. [Google Scholar] [CrossRef]
- Friis, I. The wild populations of Coffea arabica L., and cultivated coffee. In Taxonomic Aspects of African Economic Botany, Proceedings of the IX plenary meeting of AETFAT, Las Palmas de Gran Canaria, 18–23 March, 1978; Kunkel, G., Ed.; Bentham-Moxon Trust: Les Palmas de Gran Canaria, Spain, 1979. [Google Scholar]
- Adugna, G.; Hulluka, M.; Hindorf, H. Incidence of tracheomycosis, Gibberella xylarioides (Fusarium xylarioides), on Arabica coffee in Ethiopia. Ztschr. Pflanzenkrankh. Pflanzenschutz 2001, 108, 136–142. [Google Scholar]
- Geiser, D.M.; Ivey, M.L.L.; Hakiza, G.; Juba, J.H.; Miller, S.A. Gibberella xylarioides (anamorph: Fusarium xylarioides), a causative agent of coffee wilt disease in Africa, is a previously unrecognized member of the G. fujikuroi species complex. Mycologia 2005, 97, 191–201. [Google Scholar] [CrossRef]
- Serani, S.; Taligoola, H.K.; Hakiza, G.J. An investigation into Fusarium spp. associated with coffee and banana plants as potential pathogens of robusta coffee. Afric. J. Ecol. 2007, 45, 91–95. [Google Scholar]
- Silva, M.C.; Várzea, V.; Guerra-Guimarães, L.; Azinheira, H.G.; Fernandez, D.; Petitot, A.-S.; Bertrand Lashermes, B.P.; Nicole, M. Coffee resistance to the main diseases: Leaf rust and coffee berry disease. Braz. J. Plant Physiol. 2006, 18, 119–147. [Google Scholar] [CrossRef]
- Strange, R.N. Plant Disease Control: Towards Environmentally Acceptable Methods; Chapman and Hall: New York, NY, USA, 1993. [Google Scholar]
- Van der Vossen, H.A.M. A critical analysis of the agronomic and economic sustainability of organic coffee production. Exper. Agricult. 2005, 41, 449–473. [Google Scholar] [CrossRef]
- Cook, R.J.; Bruckart, W.L.; Coulson, J.R.; Goettel, M.S.; Humber, R.A.; Lumsden, R.D.; Maddox, J.V.; McManus, M.L.; Moore, L.; Meyer, S.F.; Quimby, P.C.; Stack, J.P.; Vaughn, J.L. Safety of microorganisms intended for pest and plant disease control: A framework for scientific evaluation. Biol. Control 1996, 7, 333–351. [Google Scholar] [CrossRef]
- Harman, G.E.; Howell, C.R.; Viterbo, A.; Chet, I.; Lorito, M. Trichoderma species—opportunistic, avirulent plant symbionts. Nature Rev. Microbiol. 2004, 2, 43–56. [Google Scholar]
- Jensen, D.F.; Wolffhechel, H. The use of fungi, particularly Trichoderma spp. and Gliocladium spp to control root rot and damping-off diseases. In Biological Control: Benefits and Risks; Hokkanen, H.M.T., Lynch, J.M., Eds.; Cambridge University Press: Cambridge, UK, 1993; pp. 177–189. [Google Scholar]
- Bailey, B.A.; Lumsden, R.D. Direct effects of Trichoderma and Gliocladium on plant growth and resistance to pathogens. In Trichoderma and Gliocladium Vol. 2; Harman, G.E., Kubicek, C.P., Eds.; Taylor and Francis: London, UK, 1998; pp. 185–204. [Google Scholar]
- Whipps, J.M. Microbial interactions and biocontrol in the rhizosphere. J. Exp. Bot. 2001, 52, 487–511. [Google Scholar] [CrossRef]
- Bowen, G.D.; Rovira, A.D. Microbial colonization of plant roots. Annu. Rev. Phytopathol. 1976, 14, 121–144. [Google Scholar] [CrossRef]
- Weller, D.M. Biological control of soilborne plant pathogens in the rhizosphere with bacteria. Annu. Rev. Phytopathol. 1988, 26, 379–407. [Google Scholar] [CrossRef]
- Rubio-Pérez, E.; Molinero-Ruiz, M.L.; Melero-Vara, J.M.; Basallote-Ureba, M.J. Selection of potential antagonists against asparagus crown and root rot caused by Fusarium spp. Commun. Agric. Appl. Biol. Sci. 2008, 73, 203–206. [Google Scholar]
- Gilardi, G.; Baudino, M.; Gullino, M.L.; Garibaldi, A. Attempts to control Fusarium root rot of bean by seed dressing. Commun. Agric. Appl. Biol. Sci. 2008, 73, 75–80. [Google Scholar]
- Shanmugam, V.; Sharma, V.; Ananthapadmanaban, B. Genetic relatedness of Trichoderma isolates antagonistic against Fusarium oxysporum f.sp. dianthi inflicting carnation wilt. Folia. Microbiol. 2002, 53, 130–138. [Google Scholar]
- Ojiambo, P.S.; Scherm, H. Biological and application-oriented factors influencing plant disease suppression by biological control: a meta-analytical review. Phytopathology 2006, 96, 1168–1174. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Kopchinskiy, A.G.; Komon, M.; Bissett, J.; Szakacs, G.; Kubicek, C.P. An oligonucleotide barcode for species identification in Trichoderma and Hypocrea. Fungal Genet. Biol. 2005, 42, 813–828. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Kubicek, C.P.; Komon-Zelazowska, M.; Mulaw, T.B.; Bissett, J. The Trichoderma harzianum demon: complex speciation history resulting in coexistence of hypothetical biological species, recent agamospecies and numerous relict lineages. BMC Evol. Biol. 2010, in press. [Google Scholar]
- Nagy, V.; Seidl, V.; Szakacs, G.; Komon-Zelazowska, M.; Kubicek, C.P.; Druzhinina, I.S. Application of DNA BarCodes for screening of industrially important fungi: the haplotype of Trichoderma harzianum sensu stricto indicates superior chitinase formation. Appl. Environ. Microbiol. 2007, 73, 7048–7058. [Google Scholar] [CrossRef]
- Samuels, G.J.; Ismaiel, A.; Bon, M.-C.; De Respinis, S.; Petrini, O. Trichoderma asperellum sensu lato consists of two cryptic species. Mycologia 2009. [Google Scholar] [CrossRef]
- Dodd, S.L.; Lieckfeldt, E.; Samuels, G.J. Hypocrea atroviridis sp. nov., the teleomorph of Trichoderma atroviride. Mycologia 2003, 95, 27–40. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Komoń-Zelazowska, M.; Kredics, L.; Hatvani, L.; Antal, Z.; Belayneh, T.; Kubicek, C.P. Alternative reproductive strategies of Hypocrea orientalis and genetically close but clonal Trichoderma longibrachiatum, both capable to cause invasive mycoses of humans. Microbiology 2008, 154, 3447–3459. [Google Scholar] [CrossRef]
- Degenkolb, T.; Dieckmann, R.; Nielsen, K.F.; Grafenhan, T.; Theis, C.; Zafari, D.; Chaverri, P.; Ismaiel, A.; Bruckner, H.; Von Döhren, H.; Thrane, U.; Petrini, O.; Samuels, G.J. The Trichoderma brevicompactum clade: a separate lineage with new species, new peptaibiotics, and mycotoxins. Mycol. Prog. 2008, 7, 177–219. [Google Scholar] [CrossRef]
- Mittermeier, R.A.; Robles, G.P.; Hoffman, M.; Pilgrim, J.; Brooks, T.; Mittermeier, C.G.; Lamourex, J.; da Fonseca, G.A.B. Hotspots Revisited: Earth’s Biologically Richest and Most Endangered Terrestrial Ecoregions; CEMEX: Mexico City, Mexico, 2004. [Google Scholar]
- Migheli, Q.; Balmas, V.; Komoń-Zelazowska, M.; Scherm, B.; Caria, R.; Kopchinskiy, A.; Kubicek, C.P.; Druzhinina, I.S. Soils of a Mediterranean hotspot of biodiversity and endemism (Sardinia, Tyrrhenian Islands) are inhabited by pan-European and likely invasive species of Hypocrea/Trichoderma. Environ. Microbiol. 2009, 11, 35–46. [Google Scholar] [CrossRef]
- Zachow, C.; Berg, C.; Müller, H.; Meincke, R.; Komon-Zelazowska, M.; Druzhinina, I.S.; Kubicek, C.P.; Berg, G. Fungal biodiversity in the soils/rhizospheres of Tenerife (Canary Islands): relationship to vegetation zones and environmental factors. ISME J. 2009, 3, 79–92. [Google Scholar] [CrossRef]
- Hoyos-Carvajala, L.; Orduz, S.; Bissett, J. Genetic and metabolic biodiversity of Trichoderma from Colombia and adjacent neotropic regions. Fungal Genet. Biol. 2009, 46, 615–631. [Google Scholar] [CrossRef]
- Kullnig, C.; Szakacs, G.; Kubicek, C.P. Molecular identification of Trichoderma species from Russia, Siberia and the Himalaya. Mycol. Res. 2000, 104, 1117–1125. [Google Scholar] [CrossRef]
- Kubicek, C.P.; Bissett, J.; Kullnig-Gradinger, C.M.; Druzhinina, I.S.; Szakacs, G. Genetic and metabolic diversity of Trichoderma: a case study on South-East Asian isolates. Fungal Genet. Biol. 2003, 38, 310–317. [Google Scholar] [CrossRef]
- Zhang, C.L.; Druzhinina, I.S.; Kubicek, C.P.; Xu, T. Biodiversity of Trichoderma in China: evidence for a North to South difference of species distribution in East Asia. FEMS Microbiol. Letts. 2005, 251, 251–257. [Google Scholar] [CrossRef]
- Batjes, N.H. Total carbon and nitrogen in the soils of the world. Eur. J. Soil Sci. 1996, 47, 151. [Google Scholar] [CrossRef]
- Tondje, P.R.; Roberts, D.P.; Bon, M.C.; Widmer, T.; Samuels, G.J.; Ismaiel, A.; Begoude, A..; Tchana, T.; Nyemb-Tshomb, E.; Ndoumbe-Nkeng, M.; Bateman, R.; Fontem, D.; Hebbar, K.P. Isolation and identification of mycoparasitic isolates of Trichoderma asperellum with potential for suppression of black pod disease of cacao in Cameroon. Biol. Control 2007, 43, 202–212. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Komon, M.; Kubicek, C.P.; Druzhinina, I.S. Hypocrea crystalligena sp. nov., a common European species with a white spored Trichoderma anamorph. Mycologia 2006, 98, 500–514. [Google Scholar]
- Kopchinskiy, A.; Komon, M.; Kubicek, C.P.; Druzhinina, I.S. TRICHOBLAST: a mulilocus database for Trichoderma and Hypocrea identifications. Mycol. Res. 2005, 109, 658–660. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl. Acid. Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef]
- Nicholas, K.B.; Nicholas, H.B. Genedoc, a Tool for Editing and Annotating Multiple Sequence Alignments. 1996. Available online: http://www.cris.com/ketchup/genedoc.html (accessed on 1 January 2008).
- Rannala, B.; Yang, Z. Probability distribution of molecular evolutionary trees: A new method of phylogenetic inference. J. Mol. Evol. 1996, 43, 304–311. [Google Scholar] [CrossRef]
- Yang, Z.; Rannala, B. Bayesian phylogenetic inference using DNA sequences: a Markov chain Monte Carlo method. Mol. Biol. Evol. 1997, 14, 717–724. [Google Scholar] [CrossRef]
- Huelsenbeck, J.P.; Ronquist, F. MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Chaverri, P.; Fallah, P.; Kubicek, C.P.; Samuels, G.J. Hypocrea flaviconidia, a new species from Costa Rica with yellow conidia. Stud. Mycol. 2004, 50, 401–407. [Google Scholar]
- Rita, N.; Tricita, H.Q. Soil mycoflora of black pepper rhizosphere in the Philippines and their In-vitro antagonism against phytophthora capsici L. Indo. J. Agric. Sci. 2004, 5, 1–10. [Google Scholar]
- Shannon, C.E.; Weiner, W. The Mathematical Theory of Communication; University of Illionis Press: Urbana, IL, USA, 1963. [Google Scholar]
- Simpson, E.H. Measurement of diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Giraud, T.; Refrégier, G.; de Vienne, D.; Le Gac, M. Speciation in fungi. Fungal Genet. Biol. 2008, 45, 791–802. [Google Scholar] [CrossRef]
- Aga, E.; Bryngelsson, T.; Bekele, E.; Salomon, B. Genetic diversity of forest arabica coffee (Coffea arabica L.) in Ethiopia as revealed by random amplified polymorphic DNA (RAPD) analysis. Hereditas 2003, 138, 36–46. [Google Scholar] [CrossRef]
- Masumbuko, L.I.; Bryngelsson, T.; Mneney, E.E.; Salomon, B. Genetic diversity in Tanzanian Arabica coffee using random amplified polymorphic DNA (RAPD) markers. Hereditas 2003, 139, 56–63. [Google Scholar] [CrossRef]
- Anthony, F.; Combes, C.; Astorga, C.; Bertrand, B.; Graziosi, G.; Lashermes, P. The origin of cultivated Coffea arabica L. varieties revealed by AFLP and SSR markers. Theor. Appl. Genet. 2002, 104, 894–900. [Google Scholar] [CrossRef]
Appendix
| Region | Ecosystem | Alt. (m) | Species | No. of Isolates | Strain No. | GenBank accession No. | ||
|---|---|---|---|---|---|---|---|---|
| C.P.K. | ITS 1 and 2 | tef1 | ||||||
| Wellega | disturbed semi-forest | 2500 | T. sp. ‘afroharzianum’ | 2 | 2618 | FJ412028 | FJ436158 | |
| 2400 | 2617 | FJ412059 | FJ436157 | |||||
| 2250 | T. harzianum sensu lato | 3 | 2620 | FJ412029 | ||||
| 2500 | 2616 | FJ412026 | ||||||
| 2800 | 2615 | FJ412025 | ||||||
| 1900 | T. asperelloides | 1 | 1839 | FJ412074 | FJ436141 | |||
| 2800 | H. atroviridis/T. atroviride clade E | 1 | 2622 | FJ412094 | FJ436160 | |||
| 1570 | T. hamatum | 2 | 1826 | FJ411979 | FJ436130 | |||
| 2400 | 2613 | FJ411986 | ||||||
| 2500 | T. sp. C.P.K.2612 | 1 | 2614 | FJ412013 | FJ436156 | |||
| 2400 | T. sp. C.P.K.1833 | 1 | 2619 | FJ412060 | FJ436159 | |||
| native forest | 1750 | T. harzianum sensu lato | 5 | 2610 | FJ412024 | FJ436154 | ||
| 1900 | 1825 | FJ412017 | FJ436129 | |||||
| 1750 | 3 | 2611 | FJ412027 | |||||
| 1750 | 2608 | FJ412023 | ||||||
| 1750 | 2609 | FJ412032 | ||||||
| 1750 | T. spirale | 2 | 2606 | FJ412078 | FJ436153 | |||
| 1750 | 2607 | FJ412079 | ||||||
| 1560 | T. asperelloides | 2 | 1893 | FJ412049 | FJ436148 | |||
| 1500 | 1838 | FJ412047 | FJ436140 | |||||
| 1950 | H. atroviridis/T. atroviride clade D | 1 | 2605 | FJ412093 | FJ436152 | |||
| 2300 | T. hamatum | 1 | 1827 | FJ411980 | ||||
| 1750 | T. sp. C.P.K.2612 | 1 | 2612 | FJ412011 | FJ436155 | |||
| 1740 | T. sp. C.P.K. 1837 | 1 | 1837 | FJ412090 | FJ436139 | |||
| Hararghe | disturbed semi-forest | 2300 | T. sp. ‘afroharzianum’ | 1 | 1840 | FJ412018 | FJ436142 | |
| 2100 | T. asperelloides | 1 | 1829 | FJ412048 | FJ436132 | |||
| 2080 | T. sp. C.P.K.1828 | 1 | 1828 | FJ412085 | FJ436131 | |||
| 1580 | T. sp. C.P.K. 1837 | 1 | 1841 | FJ436143 | ||||
| native forest | 2150 | T. harzianum sensu lato | 2 | 2623 | FJ412030 | |||
| 2150 | 2624 | FJ412031 | ||||||
| Jimma | disturbed semi-forest | 1950 | T. harzianum sensu lato | 1 | 2673 | FJ412038 | FJ436175 | |
| 2200 | T. harzianum sensu stricto | 1 | 1818 | FJ412016 | FJ436123 | |||
| 1750 | T. harzianum sensu lato | 5 | 2664 | FJ412034 | ||||
| 1800 | 2506 | FJ412020 | ||||||
| 1850 | 2507 | FJ412021 | ||||||
| 1950 | 2672 | FJ412037 | ||||||
| 1590 | 2684 | FJ412036 | ||||||
| 1660 | T. spirale | 1 | 1888 | FJ412075 | FJ436144 | |||
| 1650 | H. atroviridis/T. atroviride clade D | 2 | 2662 | FJ412095 | FJ436170 | |||
| 1700 | 2663 | FJ412096 | FJ436171 | |||||
| 1360 | H. koningiopsis/T. koningiopsis | 2 | 1816 | FJ412100 | FJ436121 | |||
| 1850 | T. gamsii | 1 | 2508 | FJ412103 | FJ436151 | |||
| 1600 | T. hamatum | 8 | 2682 | FJ411994 | ||||
| 1500 | 2686 | FJ411996 | ||||||
| 1500 | 2687 | FJ411997 | FJ436180 | |||||
| 2400 | 1810 | FJ411977 | FJ436116 | |||||
| 1770 | 1814 | FJ411981 | FJ436119 | |||||
| 1500 | 2685 | FJ411995 | FJ436179 | |||||
| 1680 | 1811 | FJ411978 | ||||||
| 2000 | 2505 | FJ411985 | FJ436150 | |||||
| 1330 | H. koningiopsis/T. koningiopsis | 1 | 1831 | FJ412091 | FJ436133 | |||
| 1950 | T. longibrachiatum | 1 | 1815 | FJ412086 | FJ436120 | |||
| 2300 | T. sp. C.P.K. 1837 | 1 | 1817 | FJ412089 | FJ436122 | |||
| 2150 | T. sp. C.P.K.1807 | 1 | 1807 | FJ412014 | FJ436113 | |||
| 2250 | T. sp. C.P.K.1833 | 3 | 1808 | FJ412055 | FJ436114 | |||
| 2300 | 1809 | FJ412056 | FJ436115 | |||||
| 1800 | 2671 | FJ412069 | ||||||
| native forest | 2100 | T. harzianum sensu lato | 3 | 2510 | FJ412022 | |||
| 1750 | 2504 | FJ412019 | ||||||
| 1700 | 2668 | FJ412035 | ||||||
| 1670 | T. spirale | 3 | 2674 | FJ412071 | FJ436176 | |||
| 1880 | 2679 | FJ412072 | FJ436178 | |||||
| 1800 | 2502 | FJ412076 | FJ436149 | |||||
| 1800 | H. atroviridis/T. atroviride clade D | 1 | 1832 | FJ412092 | FJ436134 | |||
| 1950 | H. koningiopsis/T. koningiopsis | 1 | 1813 | FJ412099 | FJ436118 | |||
| 1750 | T. hamatum | 11 | 2499 | FJ411982 | ||||
| 1880 | 2678 | FJ411991 | ||||||
| 1900 | 2680 | FJ411992 | ||||||
| 1780 | 2675 | FJ411989 | ||||||
| 1600 | 2500 | FJ411983 | ||||||
| 1700 | 2501 | FJ411984 | ||||||
| 1800 | 2667 | FJ411987 | ||||||
| 1640 | 2669 | FJ411988 | ||||||
| 1780 | 2676 | FJ411990 | ||||||
| 1900 | 2681 | FJ411993 | ||||||
| 1950 | 1835 | FJ436137 | ||||||
| 1960 | T. sp. C.P.K.1833 | 1 | 1833 | FJ412058 | FJ436135 | |||
| 2150 | T. sp. C.P.K.1812 | 1 | 1812 | FJ412015 | FJ436117 | |||
| 1720 | T. sp. C.P.K.2670 | 1 | 2670 | FJ412107 | FJ436174 | |||
| 1830 | T. sp. C.P.K.2707 | 3 | 2677 | FJ412083 | FJ436177 | |||
| 1600 | 2666 | FJ412082 | FJ436173 | |||||
| 1800 | 1834 | FJ412080 | FJ436136 | |||||
| SNNP | disturbed semi-forest | 2070 | T. sp. ‘afroharzianum’ | 1 | 2632 | FJ412033 | FJ436162 | |
| 2000 | T. harzianum sensu lato | 1 | 2718 | FJ412039 | ||||
| 1750 | T. spirale | 2 | 2694 | FJ412077 | FJ436184 | |||
| 1670 | 1822 | FJ412073 | FJ436127 | |||||
| 1670 | T. asperelloides | 7 | 1836 | FJ412046 | FJ436138 | |||
| 1750 | 2692 | FJ412051 | FJ436183 | |||||
| 2000 | 2722 | FJ412053 | FJ436191 | |||||
| 1750 | 2697 | FJ412052 | FJ436185 | |||||
| 1750 | 1819 | FJ412044 | FJ436124 | |||||
| 1700 | 1820 | FJ412045 | FJ436125 | |||||
| 1750 | 2688 | FJ412050 | FJ436181 | |||||
| 1750 | H. atroviridis/T. atroviride clade D | 1 | 2690 | FJ412097 | FJ436182 | |||
| 1780 | H. koningiopsis/T. koningiopsis | 1 | 1823 | FJ412102 | FJ436128 | |||
| 2000 | T. gamsii | 1 | 2723 | FJ412106 | FJ436192 | |||
| 1750 | T. hamatum | 10 | 2689 | FJ411998 | ||||
| 1750 | 2700 | FJ412004 | ||||||
| 1750 | 2703 | FJ412007 | ||||||
| 1750 | 2691 | FJ411999 | ||||||
| 1750 | 2695 | FJ412000 | ||||||
| 1750 | 2696 | FJ412001 | ||||||
| 1750 | 2698 | FJ412002 | ||||||
| 1750 | 2699 | FJ412003 | ||||||
| 1750 | 2702 | FJ412005 | ||||||
| 1750 | 2704 | FJ412006 | ||||||
| 1800 | T. longibrachiatum | 2 | 1889 | FJ412087 | FJ436145 | |||
| 1770 | 1890 | FJ412088 | FJ436146 | |||||
| 1690 | T. sp. C.P.K.1833 | 11 | 2645 | FJ412068 | FJ436169 | |||
| 2000 | 2719 | FJ412070 | FJ436190 | |||||
| 2200 | 2634 | FJ412063 | FJ436163 | |||||
| 1750 | 1892 | FJ412057 | FJ436147 | |||||
| 1850 | 2626 | FJ412061 | ||||||
| 1970 | 2630 | FJ412062 | FJ436161 | |||||
| 2200 | 2636 | FJ412064 | FJ436164 | |||||
| 2260 | 2637 | FJ412065 | FJ436165 | |||||
| 1790 | 2641 | FJ412066 | FJ436166 | |||||
| 1690 | 2642 | FJ412067 | FJ436167 | |||||
| 1700 | 2643 | FJ436168 | ||||||
| native forest | 1850 | T. harzianum sensu lato | 5 | 2710 | FJ412040 | FJ436189 | ||
| 1900 | 2727 | FJ412012 | FJ436194 | |||||
| 1850 | 2712 | FJ412042 | ||||||
| 1850 | 2711 | FJ412041 | ||||||
| 1900 | 2728 | FJ412043 | ||||||
| 2000 | T. asperelloides | 1 | 2724 | FJ412054 | ||||
| 1750 | H. koningiopsis/T. koningiopsis | 1 | 1821 | FJ412101 | FJ436126 | |||
| 1650 | T. gamsii | 2 | 2706 | FJ412104 | FJ436186 | |||
| 2000 | 2709 | FJ412105 | FJ436188 | |||||
| 1650 | T. hamatum | 3 | 2705 | FJ412008 | ||||
| 2050 | 2715 | FJ412009 | ||||||
| 2050 | 2717 | FJ412010 | ||||||
| 2050 | T. sp. C.P.K. 2725 | 1 | 2725 | FJ412098 | FJ436193 | |||
| 1950 | T. sp. C.P.K.2707 | 1 | 2707 | FJ412084 | FJ436187 | |||
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Belayneh Mulaw, T.; Kubicek, C.P.; Druzhinina, I.S. The Rhizosphere of Coffea Arabica in Its Native Highland Forests of Ethiopia Provides a Niche for a Distinguished Diversity of Trichoderma. Diversity 2010, 2, 527-549. https://doi.org/10.3390/d2040527
Belayneh Mulaw T, Kubicek CP, Druzhinina IS. The Rhizosphere of Coffea Arabica in Its Native Highland Forests of Ethiopia Provides a Niche for a Distinguished Diversity of Trichoderma. Diversity. 2010; 2(4):527-549. https://doi.org/10.3390/d2040527
Chicago/Turabian StyleBelayneh Mulaw, Temesgen, Christian P. Kubicek, and Irina S. Druzhinina. 2010. "The Rhizosphere of Coffea Arabica in Its Native Highland Forests of Ethiopia Provides a Niche for a Distinguished Diversity of Trichoderma" Diversity 2, no. 4: 527-549. https://doi.org/10.3390/d2040527
APA StyleBelayneh Mulaw, T., Kubicek, C. P., & Druzhinina, I. S. (2010). The Rhizosphere of Coffea Arabica in Its Native Highland Forests of Ethiopia Provides a Niche for a Distinguished Diversity of Trichoderma. Diversity, 2(4), 527-549. https://doi.org/10.3390/d2040527




